Research on the Genetic Polymorphism and Function of inlA with Premature Stop Codons in Listeria monocytogenes
Abstract
1. Introduction
2. Materials and Methods
2.1. Strain and Cultivation Conditions
2.2. Whole Genome Sequence (WGS)
2.3. Phylogenetic Analysis of inlA with PMSC
2.4. Construction of inlA Deletion Strain
2.5. Growth Experiment
2.6. Biofilm Formation Assay
2.7. Simulated Gastrointestinal Tract Experiment
2.8. In Vitro Virulence Assay
2.9. Data Analysis
3. Results
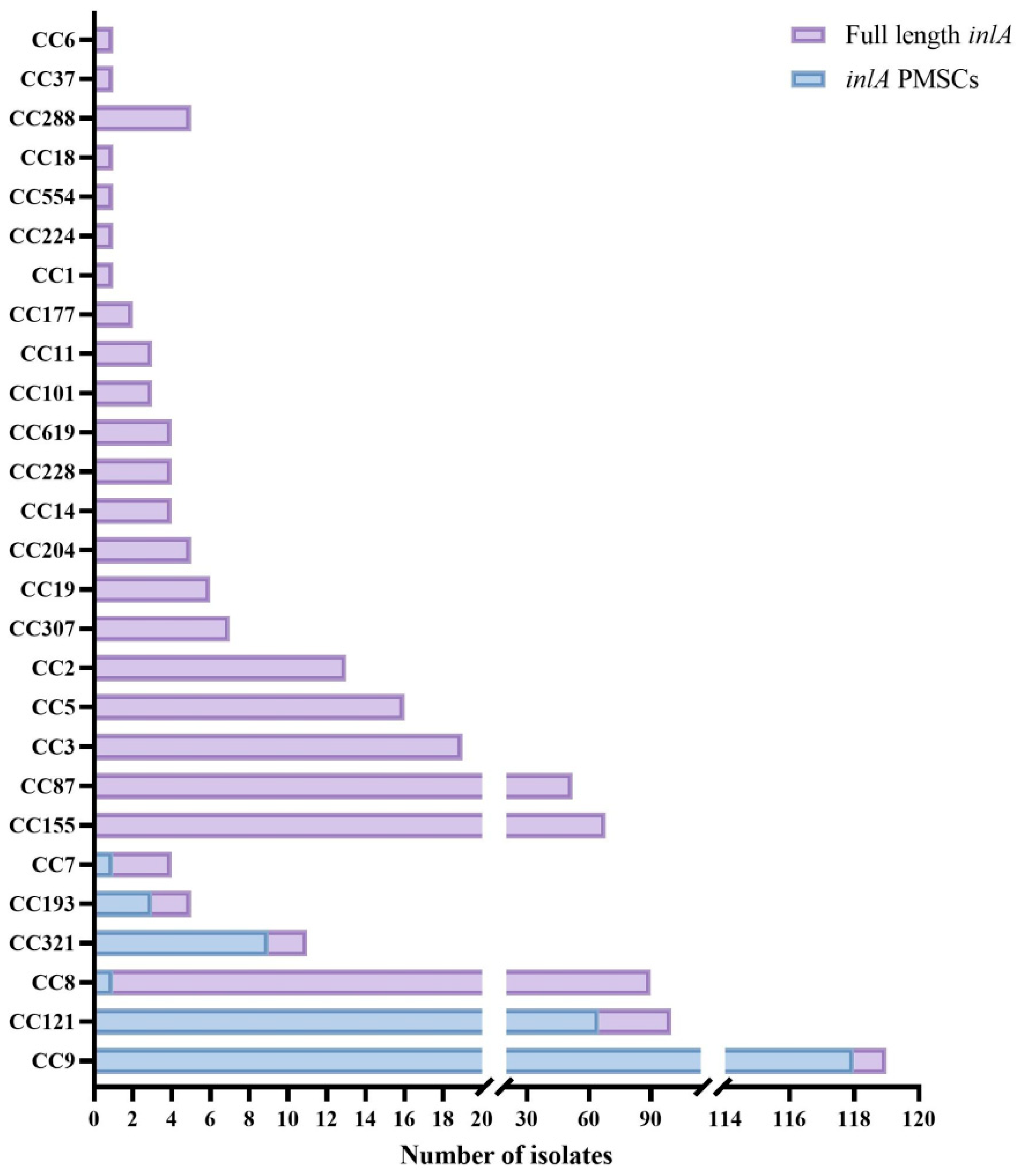
3.1. Distribution of inlA with PMSC
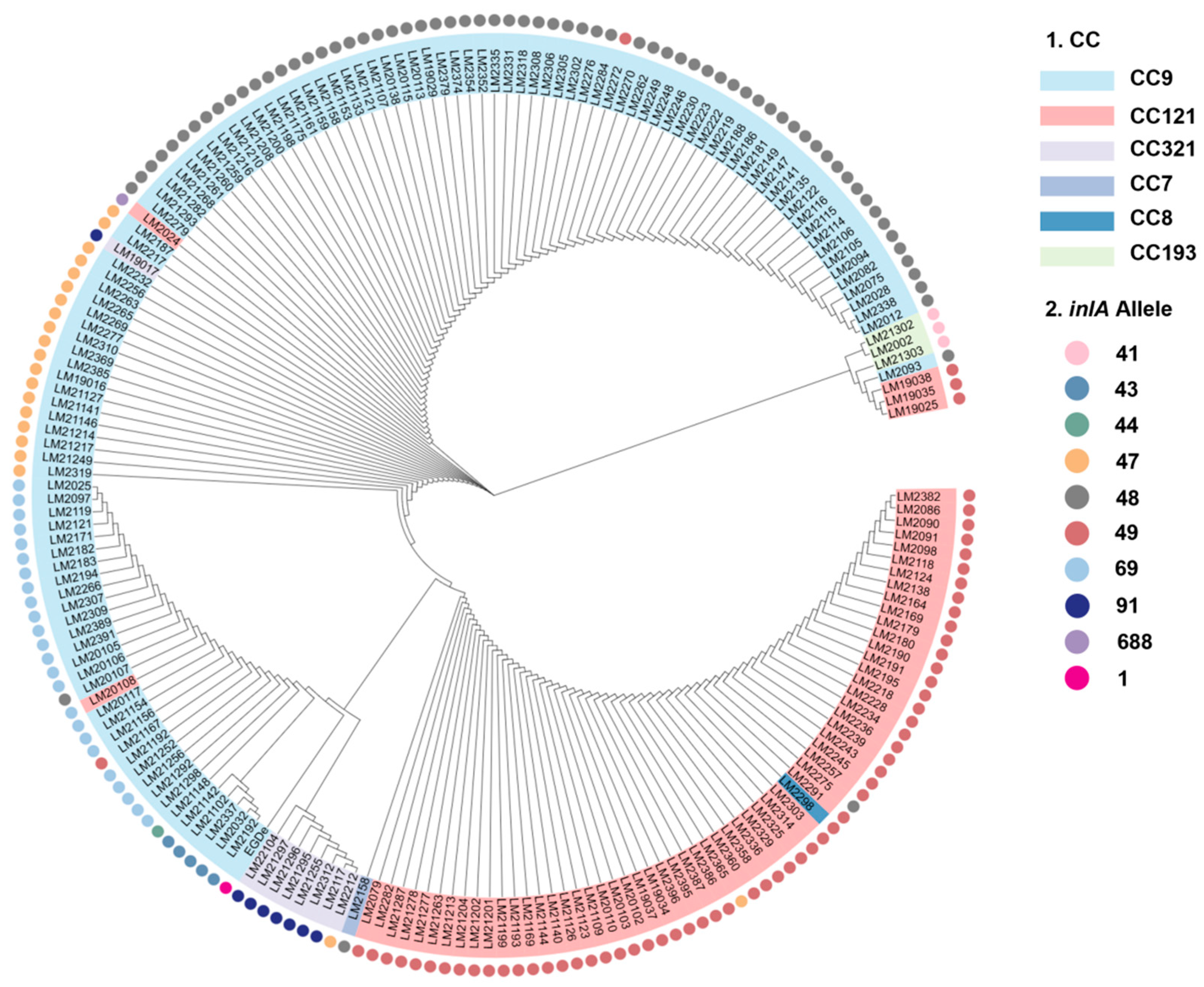
3.2. Phylogenetic Tree Analysis Based on inlA Gene
3.3. Relationship Between CCs and Types of inlA with PMSC
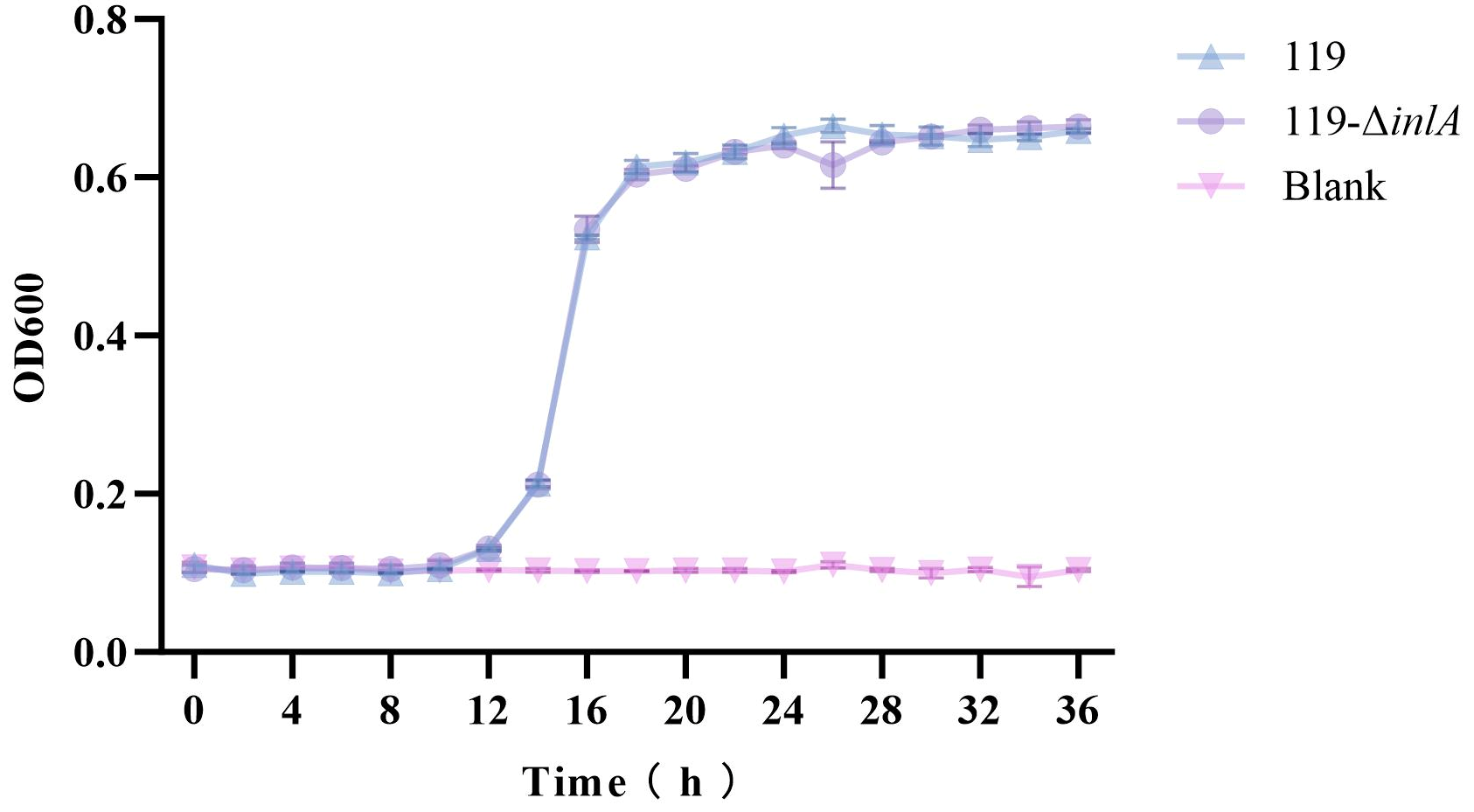
3.4. Growth of the inlA Deletion Strain
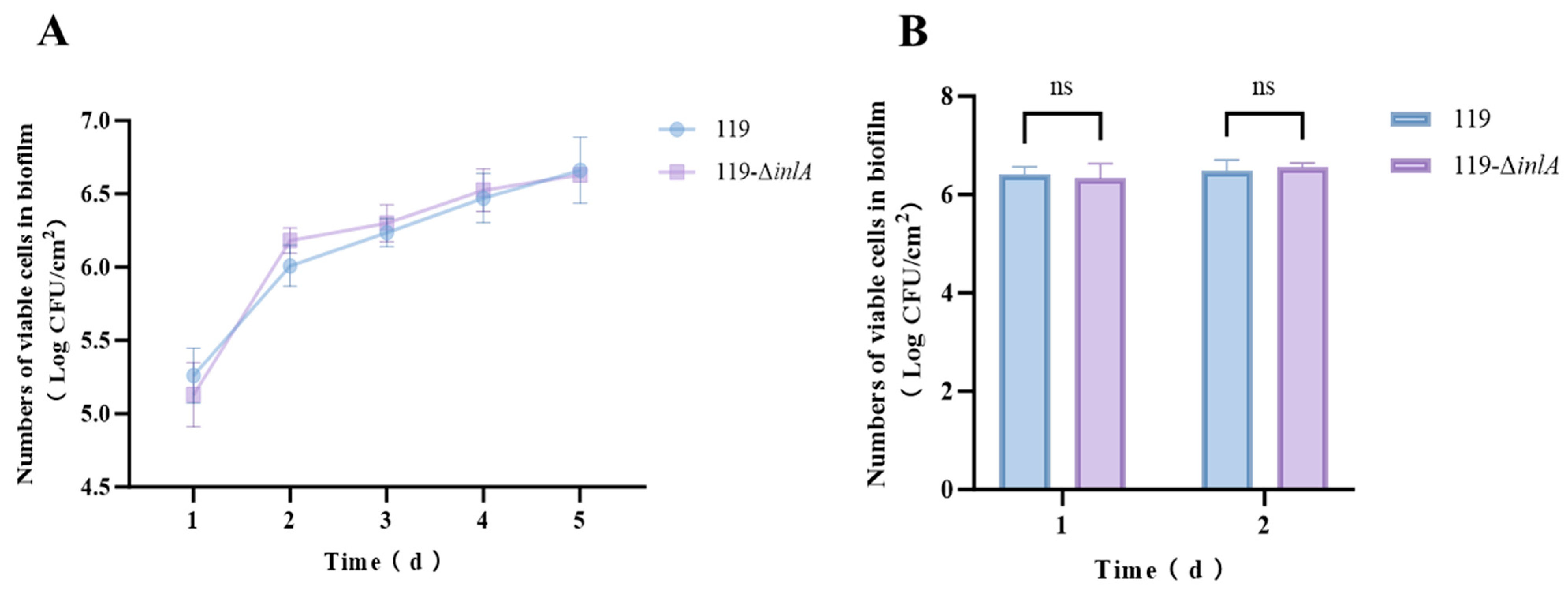
3.5. Biofilm Formation Abilities of the inlA Deletion Strain
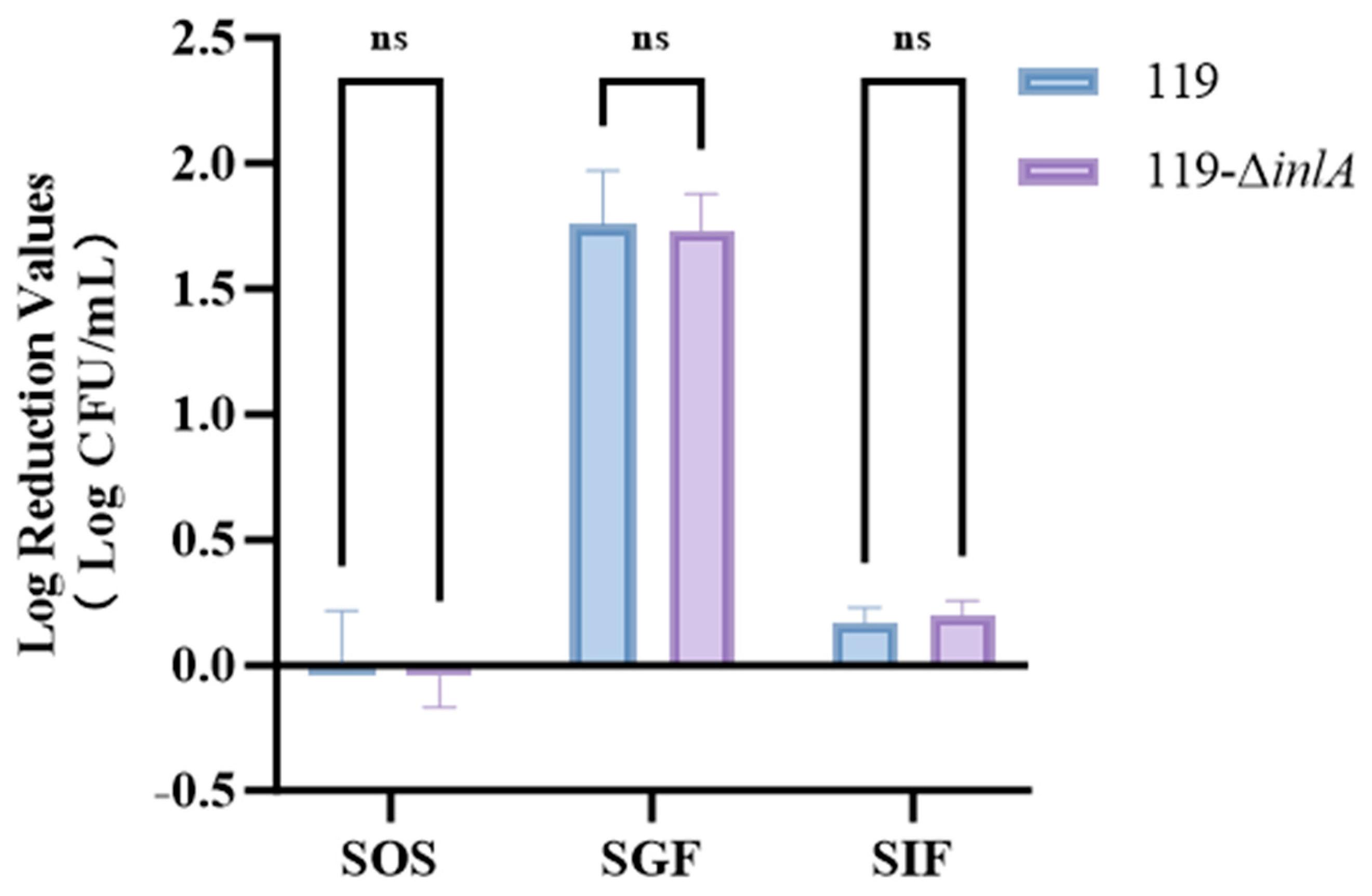
3.6. Results of Continuous GI Treatment on the inlA Deletion Strain
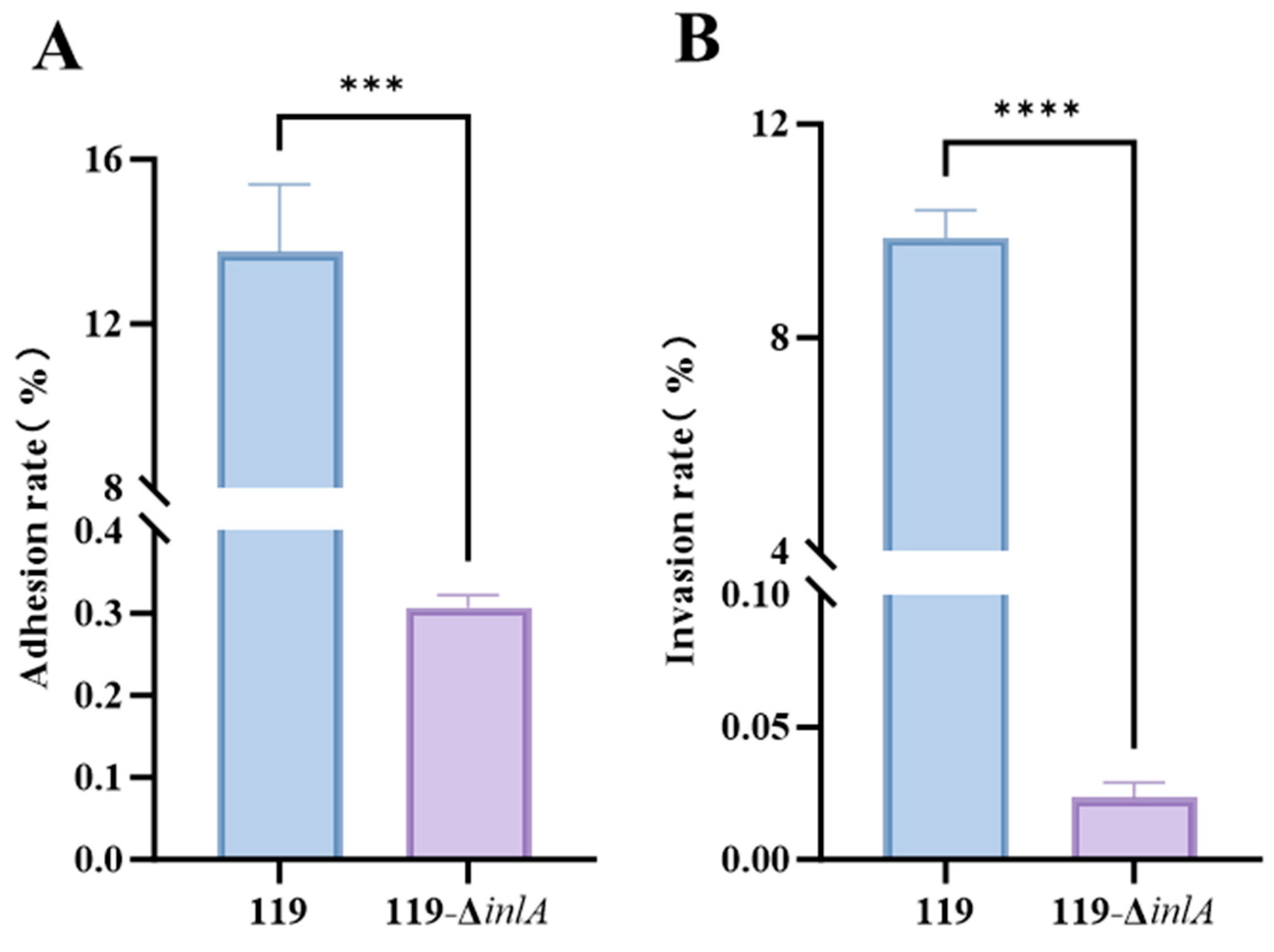
3.7. Virulence Assay of the inlA Deletion Strain
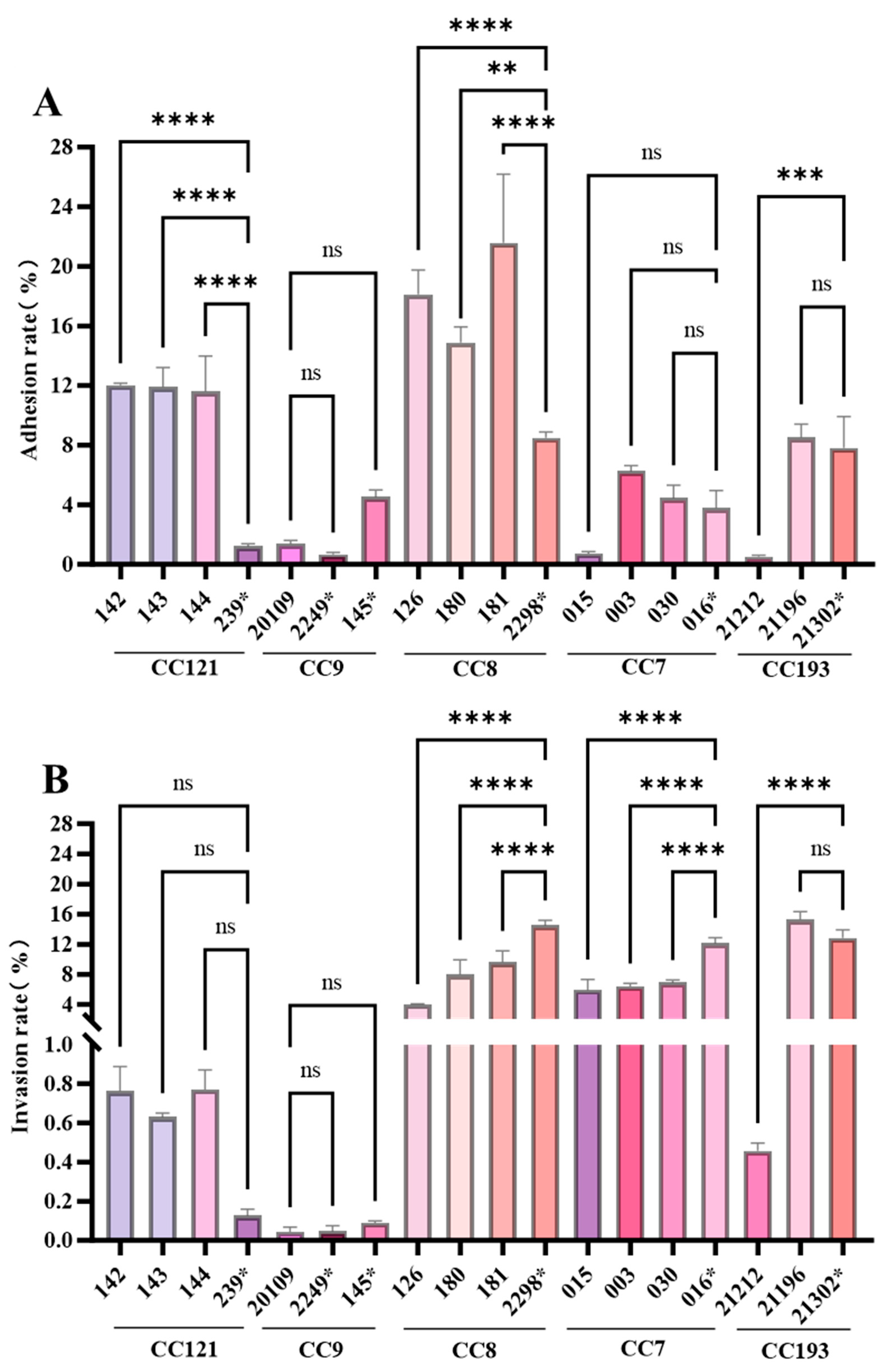
3.8. Virulence Assay of inlA with PMSC Strains
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ji, S.; Song, Z.; Luo, L.; Wang, Y.; Li, L.; Mao, P.; Ye, C.; Wang, Y. Whole-genome sequencing reveals genomic characterization of Listeria monocytogenes from food in China. Front. Microbiol. 2023, 13, 1049843. [Google Scholar] [CrossRef]
- McCollum, J.T.; Cronquist, A.B.; Silk, B.J.; Jackson, K.A.; O’Connor, K.A.; Cosgrove, S.; Gossack, J.P.; Parachini, S.S.; Jain, N.S.; Ettestad, P. Multistate outbreak of listeriosis associated with cantaloupe. N. Engl. J. Med. 2013, 369, 944–953. [Google Scholar] [CrossRef]
- CDC (Centers for Disease Control and Prevention), 2023. Listeria (Listeriosis). Available online: https://www.cdc.gov/listeria/index.html (accessed on 14 December 2023).
- The European Union One Health 2023 Zoonoses report. EFSA J. 2024, 22, 544. [CrossRef]
- Feng, Y.; Wu, S.; Varma, J.K.; Klena, J.D.; Angulo, F.J.; Ran, L. Systematic review of human listeriosis in China, 1964–2010. Trop. Med. Int. Health 2013, 18, 1248–1256. [Google Scholar] [CrossRef]
- Fan, Z.; Xie, J.; Li, Y.; Wang, H. Listeriosis in mainland China: A systematic review. Int. J. Infect. Dis. 2019, 81, 17–24. [Google Scholar] [CrossRef]
- Niu, Y.; Li, W.; Xu, B.; Chen, W.; Qi, X.; Zhou, Y.; Fu, P.; Ma, X.; Guo, Y. Risk factors associated with food consumption and food-handling habits for sporadic listeriosis: A case-control study in China from 2013 to 2022. Emerg. Microbes Infect. 2024, 13, 2307520. [Google Scholar] [CrossRef]
- Ireton, K.; Mortuza, R.; Gyanwali, G.C.; Gianfelice, A.; Hussain, M. Role of internalin proteins in the pathogenesis of Listeria monocytogenes. Mol. Microbiol. 2021, 116, 1407–1419. [Google Scholar] [CrossRef]
- Kim, H.; Marquis, H.; Boor, K.J. σ B contributes to Listeria monocytogenes invasion by controlling expression of inlA and inlB. Microbiology 2005, 151, 3215–3222. [Google Scholar] [CrossRef]
- Phelps, C.C.; Vadia, S.; Arnett, E.; Tan, Y.; Zhang, X.; Pathak-Sharma, S.; Gavrilin, M.A.; Seveau, S. Relative roles of Listeriolysin O, InlA, and InlB in Listeria monocytogenes uptake by host cells. Infect Immun. 2018, 86, e00555-18. [Google Scholar] [CrossRef]
- Li, L.; Wang, Y.; Pu, J.; Chen, J.; Liu, L.; Mao, P.; Sun, H.; Luo, X.; Ye, C. Unveiling the mutations and conservation of InlA in Listeria monocytogenes. Microorganisms 2024, 12, 485. [Google Scholar] [CrossRef]
- Su, X.; Cao, G.; Zhang, J.; Pan, H.; Zhang, D.; Kuang, D.; Yang, X.; Xu, X.; Shi, X.; Meng, J. Characterization of internalin genes in Listeria monocytogenes from food and humans, and their association with the invasion of Caco-2 cells. Gut Pathog. 2019, 11, 30. [Google Scholar] [CrossRef]
- Van Stelten, A.; Simpson, J.M.; Chen, Y.; Scott, V.N.; Whiting, R.C.; Ross, W.H.; Nightingale, K.K. Significant shift in median guinea pig infectious dose shown by an outbreak-associated Listeria monocytogenes epidemic clone strain and a strain carrying a premature stop codon mutation in inlA. Appl. Environ. Microbiol. 2011, 77, 2479–2487. [Google Scholar] [CrossRef]
- Gelbíčová, T.; Koláčková, I.; Pantůček, R.; Karpíšková, R. A novel mutation leading to a premature stop codon in inlA of Listeria monocytogenes isolated from neonatal listeriosis. New Microbiol. 2015, 38, 293–296. [Google Scholar]
- Holch, A.; Ingmer, H.; Licht, T.R.; Gram, L. Listeria monocytogenes strains encoding premature stop codons in inlA invade mice and guinea pig fetuses in orally dosed dams. J. Med. Microbiol. 2013, 62, 1799–1806. [Google Scholar] [CrossRef]
- Medeiros, M.; Castro, V.H.L.; Mota, A.; Pereira, M.G.; De Martinis, E.C.P.; Perecmanis, S.; Santana, A.P. Assessment of internalin a gene sequences and cell adhesion and invasion capacity of Listeria monocytogenes strains isolated from foods of animal and related origins. Foodborne Pathog. Dis. 2021, 18, 243–252. [Google Scholar] [CrossRef]
- Magagna, G.; Gori, M.; Russini, V.; De Angelis, V.; Spinelli, E.; Filipello, V.; Tranquillo, V.M.; De Marchis, M.L.; Bossù, T.; Fappani, C. Evaluation of the virulence potential of Listeria monocytogenes through the characterization of the truncated forms of internalin A. Int. J. Mol. Sci. 2023, 24, 10141. [Google Scholar] [CrossRef]
- Van Stelten, A.; Simpson, J.M.; Ward, T.J.; Nightingale, K.K. Revelation by single-nucleotide polymorphism genotyping that mutations leading to a premature stop codon in inlA are common among Listeria monocytogenes isolates from ready-to-eat foods but not human listeriosis cases. Appl. Environ. Microbiol. 2010, 76, 2783–2790. [Google Scholar] [CrossRef]
- GB4789.30-2016; National Food Safety Standard-Food Microbiology Inspection Listeria monocytogenes Inspection. National Standard: Beijing, China, 2016.
- Wassinger, A.; Zhang, L.; Tracy, E.; Munson, R.S.; Jr Kathariou, S.; Wang, H.H. Role of a GntR-family response regulator LbrA in Listeria monocytogenes biofilm formation. PLoS ONE 2013, 8, e70448. [Google Scholar] [CrossRef]
- Liu, X.; Chen, W.; Fang, Z.; Yu, Y.; Bi, J.; Wang, J.; Dong, Q.; Zhang, H. Persistence of Listeria monocytogenes ST5 in ready-to-eat food processing environment. Foods 2022, 11, 2561. [Google Scholar] [CrossRef]
- Cheng, Y.; Wang, X.; Liu, Y.; Qin, X.; Li, Z.; Dong, Q. Growth and survival characteristics of Listeria monocytogenes of different sources and subtypes. Lwt 2023, 184, 115114. [Google Scholar] [CrossRef]
- Dong, Q.; Lu, X.; Gao, B.; Liu, Y.; Aslam, M.Z.; Wang, X.; Li, Z. Lactiplantibacillus plantarum subsp. plantarum and fructooligosaccharides combination inhibits the growth, adhesion, invasion, and virulence of Listeria monocytogenes. Foods 2022, 11, 170. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Dong, Q.; Liu, Y.; Liu, H.; Zhang, H.; Wang, X. Systematic review of Listeria monocytogenes from food and clinical samples in Chinese mainland from 2010 to 2019. Food Qual. Saf. 2022, 6, fyac021. [Google Scholar] [CrossRef]
- Gao, B.; Cai, H.; Xu, B.; Yang, F.; Dou, X.; Dong, Q.; Yan, H.; Bu, X.; Li, Z. Growth, biofilm formation, and motility of Listeria monocytogenes strains isolated from food and clinical samples located in Shanghai (China). Food Res. Int. 2024, 184, 114232. [Google Scholar] [CrossRef]
- Liu, X.; Shi, T.; Li, J.; Wu, H.; Zhao, Q.; Fang, Z.; Liang, Y.; Xiao, Q.; Chen, M.; Dong, Q. pLM33 provides tolerance of persistent Listeria monocytogenes ST5 to various stress conditions and also enhances its virulence. Food Microbiol. 2025, 126, 104675. [Google Scholar] [CrossRef]
- Shi, Q.; Zhang, X.; Wang, D.; Zhang, W.; Jin, X.; Sun, Y.; Huang, A. Genetic diversity, antimicrobial resistance, and virulence profiles of Listeria monocytogenes isolates from Nantong, China (2020–2023). Foodborne Pathog. Dis. 2025. [Google Scholar] [CrossRef]
- Zhang, H.; Chen, W.; Wang, J.; Xu, B.; Liu, H.; Dong, Q.; Zhang, X. 10-Year molecular surveillance of Listeria monocytogenes using whole-genome sequencing in Shanghai, China, 2009–2019. Front. Microbiol. 2020, 11, 551020. [Google Scholar] [CrossRef]
- Ferreira da Silva, M.; Ferreira, V.; Magalhães, R.; Almeida, G.; Alves, A.; Teixeira, P. Detection of premature stop codons leading to truncated internalin A among food and clinical strains of Listeria monocytogenes. Food Microbiol. 2017, 63, 6–11. [Google Scholar] [CrossRef]
- Hingston, P.; Chen, J.; Dhillon, B.K.; Laing, C.; Bertelli, C.; Gannon, V.; Tasara, T.; Allen, K.; Brinkman, F.S.L.; Hansen, L.T. Genotypes associated with Listeria monocytogenes isolates displaying impaired or enhanced tolerances to cold, salt, acid, or desiccation stress. Front. Microbiol. 2017, 8, 369. [Google Scholar] [CrossRef]
- Liu, X.; Xia, X.; Liu, Y.; Li, Z.; Shi, T.; Zhang, H.; Dong, Q. Recent advances on the formation, detection, resistance mechanism, and control technology of Listeria monocytogenes biofilm in food industry. Food Res Int. 2024, 180, 114067. [Google Scholar] [CrossRef]
- Luo, Q.; Shang, J.; Feng, X.; Guo, X.; Zhang, L.; Zhou, Q. PrfA led to reduced biofilm formation and contributed to altered gene expression patterns in biofilm-forming Listeria monocytogenes. Curr. Microbiol. 2013, 67, 372–378. [Google Scholar] [CrossRef] [PubMed]
- Di Ciccio, P.; Rubiola, S.; Panebianco, F.; Lomonaco, S.; Allard, M.; Bianchi, D.M.; Civera, T.; Chiesa, F. Biofilm formation and genomic features of Listeria monocytogenes strains isolated from meat and dairy industries located in Piedmont (Italy). Int. J. Food Microbiol. 2022, 378, 109784. [Google Scholar] [CrossRef]
- Franciosa, G.; Maugliani, A.; Scalfaro, C.; Floridi, F.; Aureli, P. Expression of internalin A and biofilm formation among Listeria monocytogenes clinical isolates. Int. J. Immunopathol. Pharmacol. 2009, 22, 183–193. [Google Scholar] [CrossRef]
- Bai, X.; Liu, D.; Xu, L.; Tenguria, S.; Drolia, R.; Gallina, N.L.F.; Cox, A.D.; Koo, O.K.; Bhunia, A.K. Biofilm-isolated Listeria monocytogenes exhibits reduced systemic dissemination at the early (12–24 h) stage of infection in a mouse model. npj Biofilms Microbiomes 2021, 7, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Hadjilouka, A.; Gkolfakis, P.; Patlaka, A.; Grounta, A.; Vourli, G.; Paramithiotis, S.; Touloumi, G.; Triantafyllou, K.; Drosinos, E.H. In vitro gene transcription of Listeria monocytogenes after exposure to human gastric and duodenal aspirates. J. Food Prot. 2020, 83, 89–100. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Wu, Q.; Zhang, J.; Chen, M.; Guo, W. Analysis of multilocus sequence typing and virulence characterization of Listeria monocytogenes isolates from Chinese retail ready-to-eat Food. Front. Microbiol. 2016, 7, 168. [Google Scholar] [CrossRef] [PubMed]
- Jacquet, C.; Doumith, M.; Gordon, J.I.; Martin, P.M.V.; Cossart, P.; Lecuit, M. A molecular marker for evaluating the pathogenic potential of foodborne Listeria monocytogenes. J. Infect. Dis. 2004, 189, 2094–2100. [Google Scholar] [CrossRef] [PubMed]
- Van Stelten, A.; Nightingale, K.K. Development and implementation of a multiplex single-nucleotide polymorphism genotyping assay for detection of virulence-attenuating mutations in the Listeria monocytogenes virulence-associated gene inlA. Appl. Environ. Microbiol. 2008, 74, 7365–7375. [Google Scholar] [CrossRef]
- Pouillot, R.; Kiermeier, A.; Guillier, L.; Cadavez, V.; Sanaa, M. Updated parameters for Listeria monocytogenes dose–response model considering pathogen virulence and age and sex of consumer. Foods 2024, 13, 751. [Google Scholar] [CrossRef]
- Dellafiora, L.; Filipello, V.; Dall’Asta, C.; Finazzi, G.; Galaverna, G.; Losio, M.N. A structural study on the Listeria monocytogenes internalin A—Human E-cadherin interaction: A molecular tool to investigate the effects of missense mutations. Toxins 2020, 12, 60. [Google Scholar] [CrossRef]
- Van Stelten, A.; Roberts, A.R.; Manuel, C.S.; Nightingale, K.K. Listeria monocytogenes isolates carrying virulence-attenuating mutations in internalin A are Commonly isolated from ready-to-eat food processing plant and retail environments. J. Food Prot. 2016, 79, 1733–1740. [Google Scholar] [CrossRef] [PubMed]
- Camejo, A.; Carvalho, F.; Reis, O.; Leitão, E.; Sousa, S.; Cabanes, D. The arsenal of virulence factors deployed by Listeria monocytogenes to promote its cell infection cycle. Virulence 2011, 2, 379–394. [Google Scholar] [CrossRef] [PubMed]
| Number | Strains | CC Types | inlA Types |
|---|---|---|---|
| 1 | 119 | 87 | inlA |
| 2 | 119-ΔinlA | 87 | - |
| 3 | 142 | 121 | inlA |
| 4 | 143 | 121 | inlA |
| 5 | 144 | 121 | inlA |
| 6 | 239* | 121 | inlA-49 |
| 7 | 20109 | 9 | inlA |
| 8 | 2249* | 9 | inlA-49 |
| 9 | 145* | 9 | inlA-43 |
| 10 | 126 | 8 | inlA |
| 11 | 180 | 8 | inlA |
| 12 | 181 | 8 | inlA |
| 13 | 2298* | 8 | inlA-49 |
| 14 | 015 | 7 | inlA |
| 15 | 003 | 7 | inlA |
| 16 | 030 | 7 | inlA |
| 17 | 016* | 7 | inlA-48 |
| 18 | 21212 | 193 | inlA |
| 19 | 21196 | 193 | inlA |
| 20 | 21302* | 193 | inlA-41 |
| Allele | Mutant Site | InlA Amino Acid Length (aa) | CCs/Number |
|---|---|---|---|
| 48 | 12 (deletion A) | 8 | CC9/68, CC121/1, CC7/1 |
| 49 | 1474 (C → T) | 491 | CC9/2, CC121/61, CC8/1 |
| 47 | 1637 (deletion A) | 576 | CC9/19, CC121/1, CC321/1 |
| 69 | 976 (G → T) | 325 | CC9/23, CC121/1 |
| 43 | 1380 (G → A) | 459 | CC9/5 |
| 44 | 2054 (G→ A) | 684 | CC9/1 |
| 688 | 262 (C → G) | 55 | CC121/1 |
| 91 | 2100 (C → G) | 699 | CC321/8 |
| 41 | 12 (insertion A) | 25 | CC193/3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Gao, B.; Li, Z.; Liang, Y.; Shi, T.; Dong, Q.; Chen, M.; Wu, H.; Zhang, H. Research on the Genetic Polymorphism and Function of inlA with Premature Stop Codons in Listeria monocytogenes. Foods 2025, 14, 2955. https://doi.org/10.3390/foods14172955
Liu X, Gao B, Li Z, Liang Y, Shi T, Dong Q, Chen M, Wu H, Zhang H. Research on the Genetic Polymorphism and Function of inlA with Premature Stop Codons in Listeria monocytogenes. Foods. 2025; 14(17):2955. https://doi.org/10.3390/foods14172955
Chicago/Turabian StyleLiu, Xin, Binru Gao, Zhuosi Li, Yingying Liang, Tianqi Shi, Qingli Dong, Min Chen, Huanyu Wu, and Hongzhi Zhang. 2025. "Research on the Genetic Polymorphism and Function of inlA with Premature Stop Codons in Listeria monocytogenes" Foods 14, no. 17: 2955. https://doi.org/10.3390/foods14172955
APA StyleLiu, X., Gao, B., Li, Z., Liang, Y., Shi, T., Dong, Q., Chen, M., Wu, H., & Zhang, H. (2025). Research on the Genetic Polymorphism and Function of inlA with Premature Stop Codons in Listeria monocytogenes. Foods, 14(17), 2955. https://doi.org/10.3390/foods14172955








