The Environment, Farm Animals and Foods as Sources of Clostridioides difficile Infection in Humans
Abstract
1. Introduction
2. C. difficile Infection (CDI) in Humans
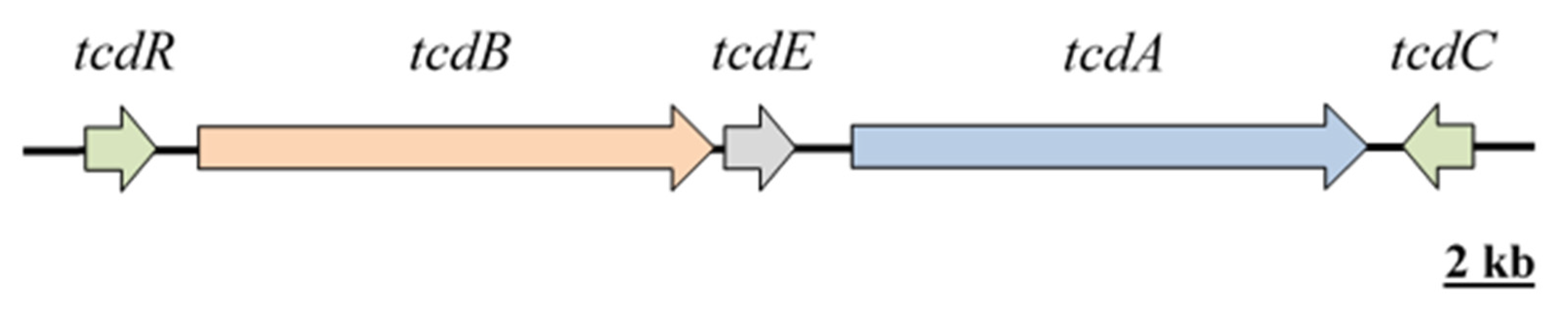
3. Virulence
| Virulence Factor | Encoding Genes | Role in CDI | References |
|---|---|---|---|
| Toxin A | tcdA | Multiple cytopathic and cytotoxic effects on the targeted cells include disruption of Rho, Rac and Cdc42-dependent signalling, the actin cytoskeleton and the tight adherence junctions, increasing epithelial permeability, allowing commensal bacterial translocation, inflammation, diarrhoea and sometimes death. | [66,67,68,75] |
| Toxin B | tcdB | ||
| TcdR | tcdR | TcdR is a positive regulator (produced in response environmental conditions) that triggers the induction of transcription of the toxin genes (tcdA and tcdB). | [76,77] |
| TcdC | tcdC | TcdC is a negative regulator that inhibits the expression of tcdA and tcdB. Mutations may cause increased production of toxins A and B. | [62,64] |
| TcdE | tcdE | TcdE may function as a lytic protein to facilitate the release of toxins A and B to the extracellular environment by a phage-like system, as these toxins lack signal peptides. | [78,79] |
| CDT | cdtA & cdtB | C. difficile binary toxin (CDT) is a transferase that disrupts the normal cytoskeletal function of cells by inhibiting the protein actin. The altered actin cytoskeleton causes an imbalance between actin and microtubules. | [69,70,71] |
4. Ribotypes
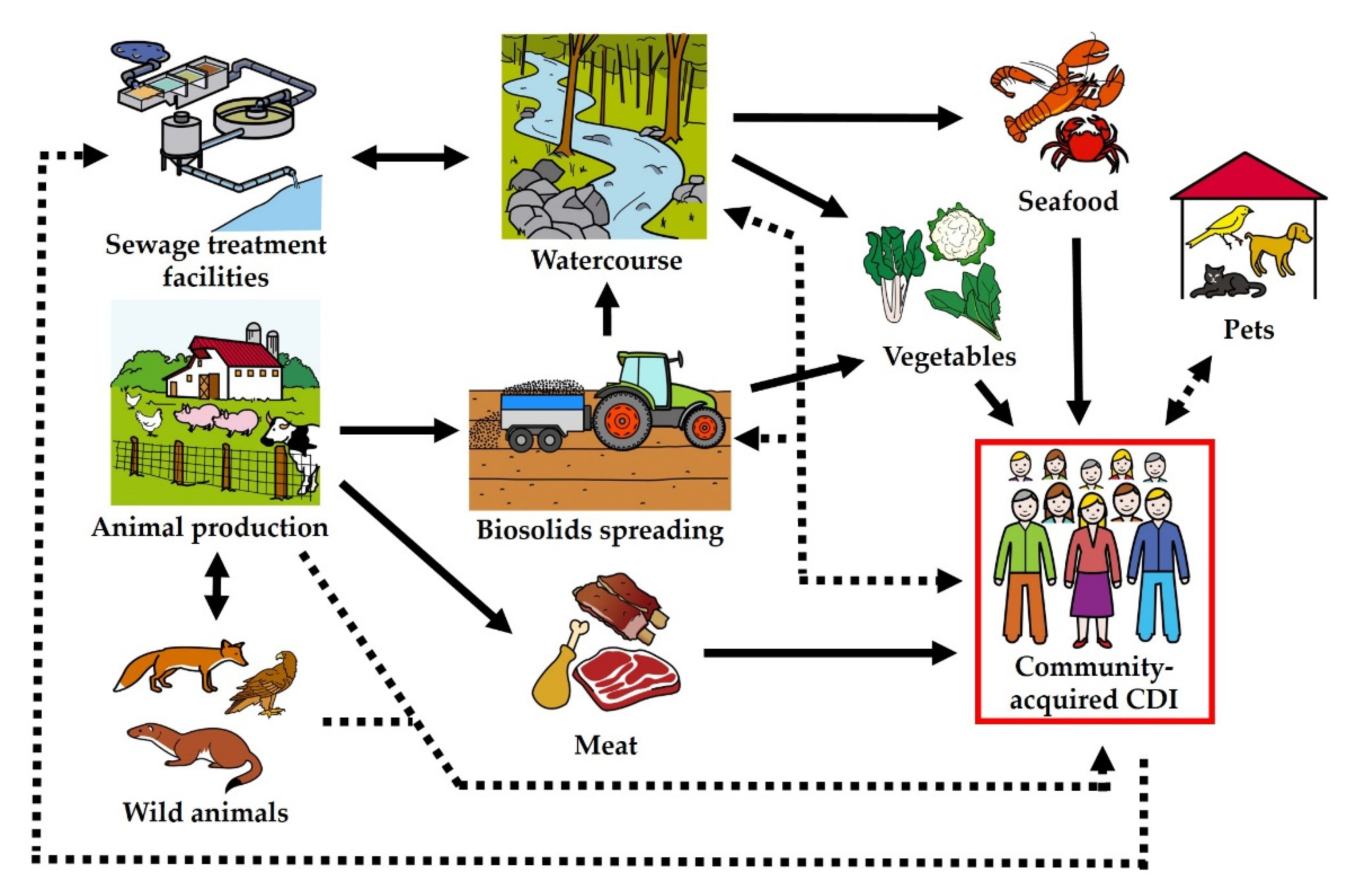
5. C. difficile in the Environment, Farm Animals and Food
5.1. Water
5.2. Soil, Manure and Silage
5.3. Farm Environment and Animals
5.4. C. difficile at the Animal Slaughter Stage
5.5. C. difficile in Retail Foods
5.6. C. difficile in Meat and Seafood
5.7. C. difficile in Vegetables
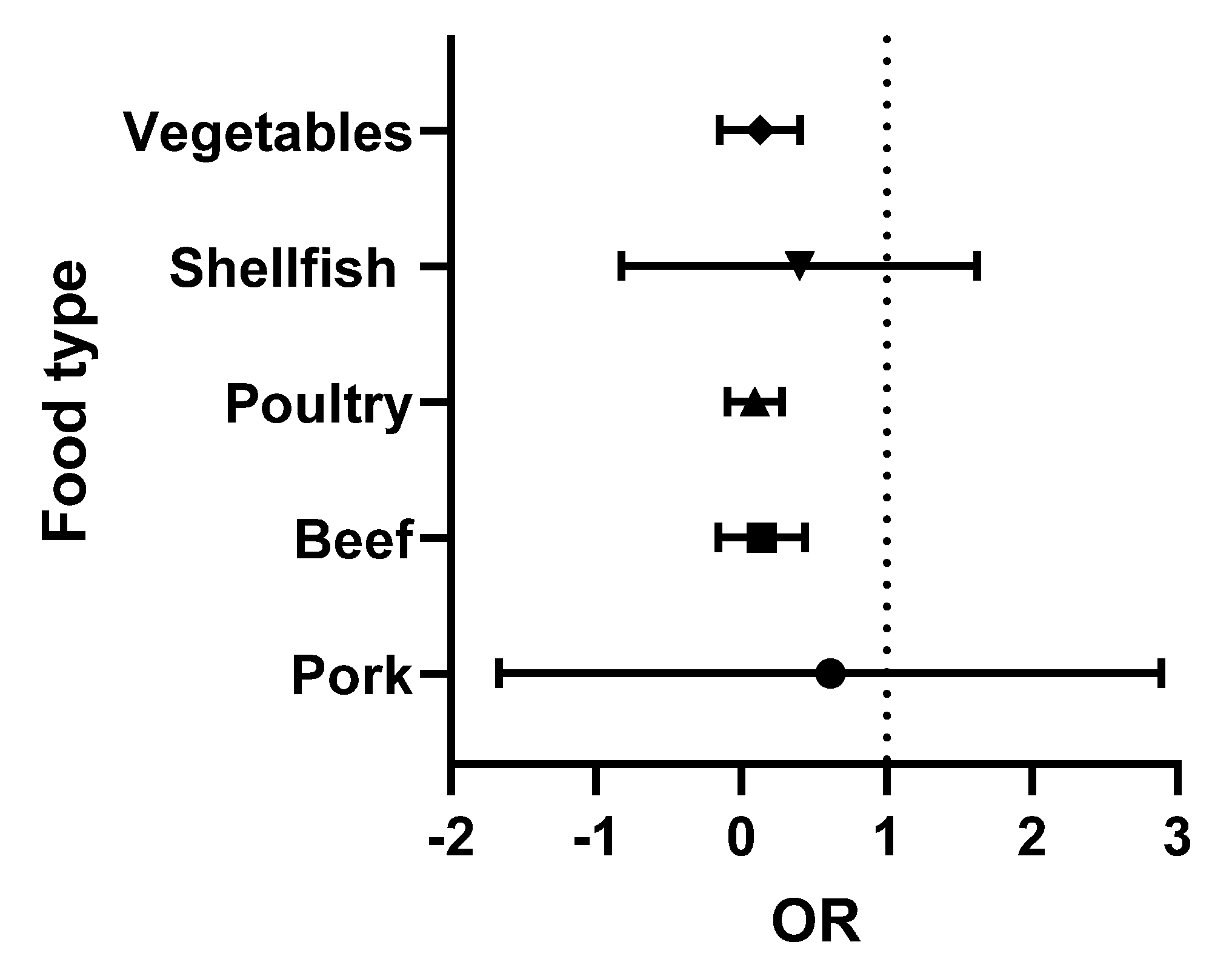
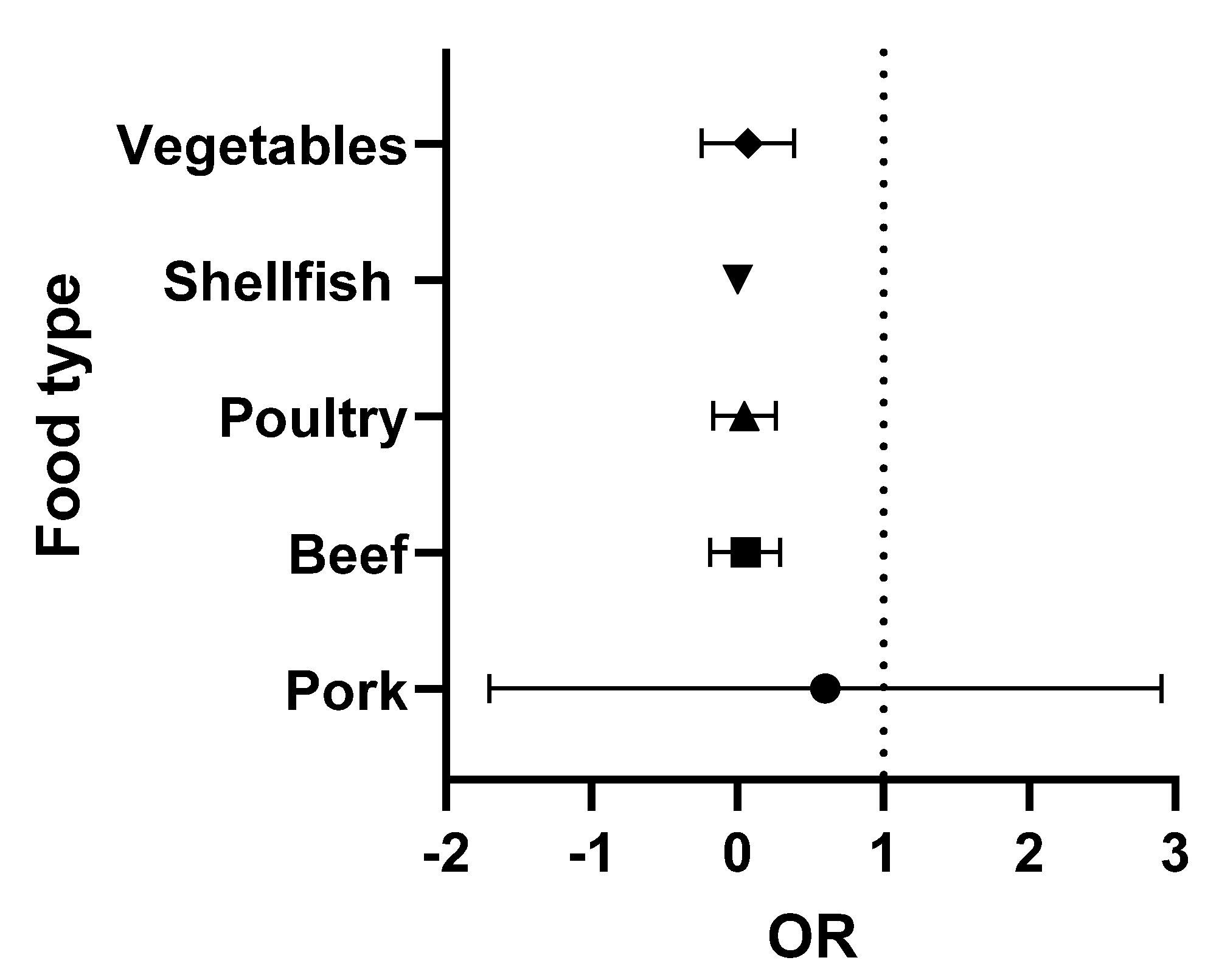
5.8. Meta-Analysis
6. The Epidemiology of Foodborne Infection
7. Control Strategies
8. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Heinlen, L.; Ballard, J.D. Clostridium difficile infection. Am. J. Med. Sci. 2010, 340, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Rousseau, C.; Poilane, I.; De Pontual, L.; Maherault, A.C.; Le Monnier, A.; Collignon, A. Clostridium difficile carriage in healthy infants in the community: A potential reservoir for pathogenic strains. Clin. Inf. Dis. 2012, 55, 1209–1215. [Google Scholar] [CrossRef] [PubMed]
- Galdys, A.L.; Curry, S.R.; Harrison, L.H. Asymptomatic Clostridium difficile colonization as a reservoir for Clostridium difficile infection. Expert Rev. Anti-Infect. Ther. 2014, 12, 967–980. [Google Scholar] [CrossRef]
- Martin, J.S.; Monaghan, T.M.; Wilcox, M.H. Clostridium difficile infection: Epidemiology, diagnosis and understanding transmission. Nat. Rev. Gastroenterol. Hepatol. 2016, 13, 206–216. [Google Scholar] [CrossRef]
- Czepiel, J.; Dróżdż, M.; Pituch, H.; Kuijper, E.J.; Perucki, W.; Mielimonka, A.; Goldman, S.; Wultańska, D.; Garlicki, A.; Biesiada, G. Clostridium difficile infection: Review. Eur. J. Clin. Microbiol. Infect. Dis. 2019, 38, 1211–1221. [Google Scholar] [CrossRef]
- Vedantam, G.; Clark, A.; Chu, M.; McQuade, R.; Mallozzi, M.; Viswanathan, V.K. Clostridium difficile infection: Toxins and non-toxin virulence factors, and their contributions to disease establishment and host response. Gut Microbes 2012, 3, 121–134. [Google Scholar] [CrossRef]
- Manabe, Y.C.; Vinetz, J.M.; Moore, R.D.; Merz, C.; Charache, P.; Bartlett, J.G. Clostridium difficile colitis: An efficient clinical approach to diagnosis. Ann. Intern. Med. 1995, 123, 835–840. [Google Scholar] [CrossRef]
- Borriello, S.P. Pathogenesis of Clostridium difficile infection. J. Antimicrobial. Chemo. 1998, 41, 13–19. [Google Scholar] [CrossRef]
- Brazier, J.S. Clostridium difficile: From obscurity to superbug. Brit. J. Biomed. Sci. 2008, 65, 39–44. [Google Scholar] [CrossRef]
- Peng, Z.; Jin, D.; Kim, H.B.; Stratton, C.W.; Wu, B.; Tang, Y.-W.; Sun, X. Update on Antimicrobial Resistance in Clostridium difficile. J. Clin. Microbiol. 2017, 55, 1998–2008. [Google Scholar] [CrossRef]
- Owen, R.C.; Donskey, C.J.; Gaynes, R.P.; Loo, V.G.; Muto, C.A. Antimicrobial-associated risk factors for Clostridium difficile infection. Clin. Infect. Dis. 2008, 46, 19–31. [Google Scholar]
- Ananthakrishnan, A. Clostridium difficile infection: Epidemiology, risk factors and management. Nat. Rev. Gastroenterol. Hepatol. 2011, 8, 17–26. [Google Scholar] [CrossRef] [PubMed]
- Enoch, D.A.; Aliyu, S.H. Is Clostridium difficile infection still a problem for hospitals? Can. Med. Assoc. J. 2012, 184, 17–18. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.; Zhu, N.A. Community-acquired Clostridium difficile infection. Can Fam Physician. 2017, 63, 131–132. [Google Scholar]
- Lim, S.C.; Knight, D.R.; Riley, T.V. Clostridium difficile and One Health. Clin. Microbiol. Infect. 2020, 26, 857–863. [Google Scholar] [CrossRef]
- Koene, M.; Mevius, D.; Wagenaar, J.; Harmanus, C.; Hensgens, M.; Meetsma, A.; Putirulan, F.; van Bergen, M.; Kuijper, E. Clostridium difficile in Dutch animals: Their presence, characteristics and similarities with human isolates. Clin. Microbiol. Infect. 2012, 18, 778–784. [Google Scholar] [CrossRef]
- Schneeberg, A.; Rupnik, M.; Neubauer, H.; Seyboldt, C. Prevalence and distribution of Clostridium difficile PCR ribotypes in cats and dogs from animal shelters in Thuringia, Germany. Anaerobe 2012, 18, 484–488. [Google Scholar] [CrossRef]
- Indra, A.; Lassnig, H.; Baliko, N.; Much, P.; Fiedler, A.; Huhulescu, S. Clostridium difficile: A new zoonotic agent? Wien. Klin. Wochenschr. 2009, 121, 91–95. [Google Scholar] [CrossRef]
- Schmid, A.; Messelhäusser, U.; Hörmansdorfer, S.; Sauter-Louis, C.; Mansfeld, R. Occurrence of zoonotic Clostridia and Yersinia in healthy cattle. J. Food Protect. 2013, 76, 1697–1703. [Google Scholar] [CrossRef]
- Noren, T.; Johansson, K.; Unemo, M. Clostridium difficile PCR-ribotype 046 is common among neonatal pigs and humans in Sweden. Clin. Microbiol. Infect. 2014, 20, O2–O6. [Google Scholar] [CrossRef]
- Rodriguez-Palacios, A.; Borgmann, S.; Kline, T.R.; LeJeune, J.T. Clostridium difficile in foods and animals: History and measures to reduce exposure. Animal Health Res. Rev. 2013, 14, 11–29. [Google Scholar] [CrossRef]
- Metcalf, D.; Costa, M.C.; Dew, W.M.; Weese, J.S. Clostridium difficile in vegetables, Canada. Lett. Appl. Microbiol. 2010, 51, 600–602. [Google Scholar] [CrossRef] [PubMed]
- Weese, J. Clostridium difficile in food—Innocent bystander or serious threat? Clin. Microbiol. Infect. 2010, 16, 3–10. [Google Scholar] [CrossRef] [PubMed]
- McBee, R.H. Intestinal flora of some antarctic birds and mammals. J. Bacteriol. 1960, 79, 311–312. [Google Scholar] [CrossRef] [PubMed]
- Hafiz, S. Clostridium difficile and Its Toxins. Ph.D. Thesis, Department of Microbiology, University of Leeds, Leeds, UK, 1974. [Google Scholar]
- Princewell, T.J.T.; Agba, M.I. Examination of bovine faeces for the isolation and identification of Clostridium species. J. Appl. Bacteriol. 1982, 52, 97–102. [Google Scholar] [CrossRef]
- Nagy, J.; Bilkei, G. Neonatal piglet losses associated with Escherichia coli and Clostridium difficile infection in a Slovakian outdoor production unit. Vet. J. 2003, 166, 98–100. [Google Scholar] [CrossRef]
- Borriello, S.P.; Honour, P.; Turner, T.; Barclay, F. Household pets as a potential reservoir for Clostridium difficile infection. J. Clin. Pathol. 1983, 36, 84–87. [Google Scholar] [CrossRef]
- Weber, A.; Kroth, P.; Heil, G. Domestic animals as excreters of Clostridium difficile. Dtsch. Med. Wochenschr. 1988, 113, 1617–1618. [Google Scholar]
- Goorhuis, A.; Debast, S.B.; van Leengoed, L.A.; Harmanus, C.; Notermans, D.W.; Bergwerff, A.A.; Kuijper, E.J. Clostridium difficile PCR ribotype 078: An emerging strain in humans and in pigs? J. Clin. Microbiol. 2008, 46, 1157. [Google Scholar] [CrossRef]
- Jhung, M.A.; Thompson, A.D.; Killgore, G.E.; Zukowski, W.E.; Songer, G.; Warny, M.; Johnson, S.; Gerding, D.N.; McDonald, L.C.; Limbago, B.M. Toxinotype V Clostridium difficile in humans and food animals. Emer. Infect. Dis. 2008, 14, 1039–1045. [Google Scholar] [CrossRef]
- Rupnik, M.; Widmer, A.; Zimmermann, O.; Eckert, C.; Barbut, F. Clostridium difficile toxinotype V, ribotype 078, in animals and humans. J. Clin. Microbiol. 2008, 46, 2146. [Google Scholar] [CrossRef] [PubMed]
- Debast, S.B.; van Leengoed, L.A.; Goorhuis, A.; Harmanus, C.; Kuijper, E.J.; Bergwerff, A.A. Clostridium difficile PCR ribotype 078 toxinotype V found in diarrhoeal pigs identical to isolates from affected humans. Environ. Microbiol. 2009, 11, 505–511. [Google Scholar] [CrossRef] [PubMed]
- Susick, E.K.; Putnam, M.; Bermudez, D.M.; Thakur, S. Longitudinal study comparing the dynamics of Clostridium difficile in conventional and antimicrobial free pigs at farm and slaughter. Vet. Microbiol. 2012, 157, 172–178. [Google Scholar] [CrossRef]
- Songer, J.G.; Trinh, H.T.; Killgore, G.E.; Thompson, A.D.; McDonald, L.C.; Limbago, B.M. Clostridium difficile in Retail Meat Products, USA, 2007. Emer. Infect. Dis. 2009, 15, 819–821. [Google Scholar] [CrossRef]
- Weese, J.; Avery, B.P.; Rousseau, J.; Reid-Smith, J. Detection and Enumeration of Clostridium difficile Spores in Retail Beef and Pork. Appl. Environ. Microbiol. 2009, 15, 5009–5011. [Google Scholar] [CrossRef]
- Bouttier, S.; Barc, M.C.; Felix, B.; Lambert, S.; Collignon, A.; Barbut, F. Clostridium difficile in ground meat, France. Emer. Infect. Dis. 2010, 16, 733–735. [Google Scholar] [CrossRef]
- de Boer, E.; Zwartkruis-Nahuis, A.; Heuvelink, A.E.; Harmanus, C.; Kuijper, E.J. Prevalence of Clostridium difficile in retailed meat in The Netherlands. Int. J. Food Microbiol. 2011, 144, 561–564. [Google Scholar] [CrossRef] [PubMed]
- Quesada-Gomez, C.; Mulvey, M.R.; Vargas, P.; Gamboa-Coronado Mde, l.M.; Rodriguez, C.; Rodriguez-Cavillini, E. Isolation of a toxigenic and clinical 25 genotype of Clostridium difficile in retail meats in Costa Rica. J. Food Pro. 2013, 76, 348–351. [Google Scholar] [CrossRef]
- Kouassi, K.A.; Dadie, A.T.; N’Guessan, K.F.; Dje, K.M.; Loukou, Y.G. Clostridium perfringens and Clostridium difficile in cooked beef sold in Côte d’Ivoire and their antimicrobial susceptibility. Anaerobe 2014, 28, 90–94. [Google Scholar] [CrossRef]
- Primavilla, S.; Farneti, S.; Petruzzelli, A.; Drigo, I.; Scuota, S. Contamination of hospital food with Clostridium difficile in Central Italy. Anaerobe 2019, 55, 8–10. [Google Scholar] [CrossRef]
- Miller, M.; Gravel, D.; Mulvey, M.; Taylor, G.; Boyd, D.; Simor, A.; Gardam, M.; McGeer, A.; Hutchinson, J.; Moore, D.; et al. Health Care-Associated Clostridium difficile Infection in Canada: Patient Age and Infecting Strain Type Are Highly Predictive of Severe Outcome and Mortality. Clin. Infect. Dis 2010, 50, 194–201. [Google Scholar] [CrossRef] [PubMed]
- Kelly, C.P. Immune response to Clostridium difficile infection. Eur. J. Gastroenterol. Hepatol 1996, 8, 1048–1053. [Google Scholar] [CrossRef] [PubMed]
- Ogra, P.L. Ageing and its possible impact on mucosal immune responses. Ageing Res. Rev. 2010, 9, 101–106. [Google Scholar] [CrossRef] [PubMed]
- Cole, S.A.; Stahl, T.J. Persistent and Recurrent Clostridium difficile Colitis. Clin. Colon Rectal Sur. 2015, 28, 65–69. [Google Scholar] [CrossRef] [PubMed]
- Dharbhamulla, N.; Abdelhady, A.; Domadia, M.; Patel, S.; Gaughan, J.; Roy, S. Risk Factors Associated with Recurrent Clostridium difficile Infection. J. Clin. Med. Res. 2019, 11, 1–6. [Google Scholar] [CrossRef]
- Song, J.H.; Kim, Y.S. Recurrent Clostridium difficile Infection: Risk Factors, Treatment, and Prevention. Gut Liver 2019, 13, 16–24. [Google Scholar] [CrossRef]
- Britton, R.A.; Young, V.B. Role of the intestinal microbiota in resistance to colonization by Clostridium difficile. Gastroenterology 2014, 146, 1547–1553. [Google Scholar] [CrossRef]
- Min, J.H.; Kim, Y.S. Proton pump inhibitors should be used with caution in critically Ill patients to prevent the risk of Clostridium difficile infection. Gut Liver 2016, 10, 493–494. [Google Scholar] [CrossRef]
- Tariq, R.; Singh, S.; Gupta, A.; Pardi, D.S.; Khanna, S. Association of gastric acid suppression with recurrent Clostridium difficile infection: A systematic review and meta-analysis. JAMA Internal Med. 2017, 177, 784–791. [Google Scholar] [CrossRef]
- Mehta, P.; Nahass, R.G.; Brunetti, L. Acid suppression medications during hospitalization as a risk factor for recurrence of Clostridioides difficile infection: Systematic review and meta-analysis. Clin. Infect. Dis. 2021, 545, 1–7. [Google Scholar] [CrossRef]
- Surawicz, C.M.; Brandt, L.J.; Binion, D.G.; Ananthakrishnan, A.N.; Curry, S.R.; Gilligan, P.H.; McFarland, L.V.; Mellow, M.; Zuckerbraun, B.S. Guidelines for diagnosis, treatment, and prevention of Clostridium difficile infections. Am. J. Gastroenterol. 2013, 108, 478–498. [Google Scholar] [CrossRef] [PubMed]
- Louie, T.J.; Miller, M.A.; Mullane, K.M.; Weiss, K.; Lentnek, A.; Golan, Y.; Gorbach, S.; Sears, P.; Shue, Y.-K. Fidaxomicin versus vancomycin for Clostridium difficile infection. N. Eng. J. Med. 2011, 364, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Chaparro-Rojas, F.; Mullane, K.M. Emerging therapies for Clostridium difficile infection—Focus on fidaxomicin. Infect. Drug Res. 2013, 6, 41–53. [Google Scholar]
- Iv, E.C.O.; Iii, E.C.O.; Johnson, D.A. Clinical update for the diagnosis and treatment of Clostridium difficile infection. World J. Gastro. Pharmacol. Ther. 2014, 5, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Dyer, C.; Hutt, L.P.; Burky, R.; Joshi, L.T. Biocide Resistance and Transmission of Clostridium difficile Spores Spiked onto Clinical Surfaces from an American Health Care Facility. Appl. Environ. Microbiol. 2019, 85, e01090-19. [Google Scholar] [CrossRef] [PubMed]
- Depestel, D.D.; Aronoff, D.M. Epidemiology of Clostridium difficile infection. J. Pharm. Pract. 2013, 26, 464–475. [Google Scholar] [CrossRef] [PubMed]
- Barbut, F.; Petit, J.C. Epidemiology of Clostridium difficile-associated infections. Clin. Microbiol. Infect. 2001, 7, 405–410. [Google Scholar] [CrossRef]
- Akerlund, T.; Svenungsson, B.; Lagergren, A.; Burman, L.G. Correlation of disease severity with fecal toxin levels in patients with Clostridium difficile-associated diarrhea and distribution of PCR ribotypes and toxin yields in vitro of corresponding isolates. J. Clin. Microbiol. 2006, 44, 353–358. [Google Scholar] [CrossRef]
- Evans, C.T.; Safdar, N. Current trends in the epidemiology and outcomes of Clostridium difficile infection. Clin Infect Dis. 2015, 60 (Suppl. S2), S66–S71. [Google Scholar] [CrossRef]
- Czepiel, J.; Krutova, M.; Mizrahi, A.; Khanafer, N.; Enoch, D.A.; Patyi, M.; Deptuła, A.; Agodi, A.; Nuvials, X.; Pituch, H.; et al. Mortality Following Clostridioides difficile Infection in Europe: A Retrospective Multicenter Case-Control Study. Antibiotic 2021, 10, 299. [Google Scholar] [CrossRef]
- Voth, D.; Ballard, J.D. Clostridium difficile Toxins: Mechanism of Action and Role in Disease. Clin. Microbiol. Rev. 2005, 2, 247–263. [Google Scholar] [CrossRef] [PubMed]
- Rupnik, M.; Janezic, S. An update on Clostridium difficile toxinotyping. J. Clin. Microbiol. 2016, 54, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Hundsberger, T.; Braun, V.; Weidmann, M.; Leuke, P.; Sauerborn, M.; von Eichel-Streiber, C. Transcription analysis of the genes tcdA-E of the pathogenicity locus of Clostridium difficile. Eur. J. Biochem. 1997, 244, 735–742. [Google Scholar] [CrossRef] [PubMed]
- Matamouros, S.; England, P.; Dupuy, B. Clostridium difficile toxin expression is inhibited by the novel regulator TcdC. Mol. Microbiol. 2007, 64, 1274–1288. [Google Scholar] [CrossRef] [PubMed]
- Donta, S.T.; Sullivan, N.; Wilkins, T.D. Differential effects of Clostridium difficile toxins on tissue-cultured cells. J. Clin. Microbiol. 1982, 15, 1157–1158. [Google Scholar] [CrossRef]
- Feltis, B.A.; Wiesner, S.M.; Kim, A.S.; Erlandsen, S.L.; Lyerly, D.L.; Wilkins, T.D.; Wells, C.L. Clostridium difficile toxins A and B can alter epithelial permeability and promote bacterial paracellular migration through HT-29 enterocytes. Shock 2000, 14, 629–634. [Google Scholar] [CrossRef]
- Riggs, M.M.; Sethi, A.K.; Zabarsky, T.F.; Eckstein, E.C.; Jump, R.L.P.; Donskey, C.J. Asymptomatic carriers are a potential source for transmission of epidemic and nonepidemic Clostridium difficile strains among long-term care facility residents. Clin. Infect. Dis. 2007, 45, 992–998. [Google Scholar] [CrossRef]
- Stiles, B.G.; Wigelsworth, D.J.; Popoff, M.R.; Barth, H. Clostridial binary toxins: Iota and c2 family portraits. Front. Cell. Infect. Microbiol. 2011, 1, 11. [Google Scholar] [CrossRef]
- Gerding, D.; Johnson, S.; Rupnik, M.; Aktories, K. Clostridium difficile binary toxin CDT: Mechanism, epidemiology, and potential clinical importance. Gut Microbes 2014, 5, 15–27. [Google Scholar] [CrossRef]
- Aktories, K.; Papatheodorou, P.; Schwan, C. Binary Clostridium difficile toxin (CDT)—A virulence factor disturbing the cytoskeleton. Anaerobe 2018, 53, 21–29. [Google Scholar] [CrossRef]
- Kuehne, S.A.; Collery, M.M.; Kelly, M.L.; Cartman, S.T.; Cockayne, A.; Minton, N.P. Importance of toxin A, toxin B, and CDT in virulence of an epidemic Clostridium difficile strain. J. Infect. Dis. 2014, 209, 83–86. [Google Scholar] [CrossRef] [PubMed]
- Wilson, V.; Cheek, L.; Satta, G.; Walker-Bone, K.; Cubbon, M.; Citron, D. Predictors of death after Clostridium difficile infection: A report on 128 strain-typed cases from a teaching hospital in the United Kingdom. Clin. Infect. Dis. 2010, 50, 77–81. [Google Scholar] [CrossRef] [PubMed]
- Goldenberg, S.D.; French, G.L. Lack of association of tcdC type and binary toxin status with disease severity and outcome in toxigenic Clostridium Difficile. J. Infect. 2011, 62, 355–362. [Google Scholar] [CrossRef] [PubMed]
- Di Bella, S.; Ascenzi, P.; Siarakas, S.; Petrosillo, N.; di Masi, A. Clostridium difficile Toxins A and B: Insights into Pathogenic Properties and Extraintestinal Effects. Toxins 2016, 8, 134. [Google Scholar] [CrossRef] [PubMed]
- Mizuno, T.; Kaibuchi, K.; Yamamoto, T.; Kawamura, M.; Sakoda, T.; Fujioka, H.; Matsuura, Y.; Takai, Y. A stimulatory GDP/GTP exchange protein for smg p21 is active on the post-translationally processed form of c-Ki-ras p21 and rhoA p21. Proc. Natl. Acad. Sci. USA 1991, 88, 6442–6446. [Google Scholar] [CrossRef] [PubMed]
- Mani, N.; Dupuy, B. Regulation of toxin synthesis in Clostridium difficile by an alternative RNA polymerase sigma factor. Proc. Natl. Acad. Sci. USA 2001, 98, 5844–5849. [Google Scholar] [CrossRef]
- Tan, K.S.; Wee, B.Y.; Song, K.P. Evidence for holin function of tcdE gene in the pathogenicity of Clostridium difficile. J. Med. Microbiol. 2001, 50, 613–619. [Google Scholar] [CrossRef]
- Govind, R.; Dupuy, B. Secretion of Clostridium difficile toxins A and B requires the holin-like protein TcdE. PLoS Pathog. 2012, 8, e1002727. [Google Scholar] [CrossRef]
- Dayananda, P.; Wilcox, M.H. A Review of Mixed Strain Clostridium difficile colonization and infection. Front. Microbiol. 2019, 10, 692. [Google Scholar] [CrossRef]
- Azimirad, M.; Krutova, M.; Yadegar, A.; Shahrokh, S.; Olfatifar, M.; Aghdaei, H.A.; Fawley, W.N.; Wilcox, M.H.; Zali, M.R. Clostridioides difficile ribotypes 001 and 126 were predominant in Tehran healthcare settings from 2004 to 2018: A 14-year-long cross-sectional study. Emerg. Microbes Infect. 2020, 1, 1432–1443. [Google Scholar] [CrossRef]
- Neely, F.; Lambert, M.L.; Van Broeck, J.; Delmée, M. Clinical and laboratory features of the most common Clostridium difficile ribotypes isolated in Belgium. J. Hosp. Infect. 2017, 4, 394–399. [Google Scholar] [CrossRef] [PubMed]
- Stein, K.; Egan, S.; Lynch, H.; Harmanus, C.; Kyne, L.; Herra, C.; McDermott, S.; Kuijper, E.; Fitzpatrick, F.; FitzGerald, S.; et al. PCR-ribotype distribution of Clostridium difficile in Irish pigs. Anaerobe 2017, 48, 237–241. [Google Scholar] [CrossRef]
- Kuijper, E.J.; Coignard, B.; Tüll, P. Emergence of Clostridium difficile-associated disease in North America and Europe. Clinical microbiology and infection: The official publication of the European Society of Clinical Microbiology and Infectious Diseases. Clin. Microbiol. Infect. 2006, 6, 2–18. [Google Scholar] [CrossRef] [PubMed]
- Hookman, P.; Barkin, J.S. Clostridium difficile associated infection, diarrhea and colitis. World J. Gastroenterol. 2009, 15, 1554–1580. [Google Scholar] [CrossRef] [PubMed]
- Janezic, S.; Garneau, J.R.; Monot, M. Comparative Genomics of Clostridium difficile. Adv. Exp. Med. Biol. 2018, 1050, 59–75. [Google Scholar] [PubMed]
- Jones, A.M.; Kuijper, E.J.; Wilcox, M.H. Clostridium difficile: A European perspective. J. Infect. 2013, 66, 115–128. [Google Scholar] [CrossRef]
- Baldan, R.; Trovato, A.; Bianchini, V.; Biancardi, A.; Cichero, P.; Mazzotti, M.; Nizzero, P.; Moro, M.; Ossi, C.; Scarpellini, P.; et al. Clostridium difficile PCR ribotype 018, a successful epidemic genotype. J. Clin. Microbiol. 2015, 53, 2575–2580. [Google Scholar] [CrossRef]
- Knetsch, C.W.; Hensgens, M.P.M.; Harmanus, C.; van der Bijl, M.W.; Savelkoul, P.H.M.; Kuijper, E.J.; Corver, J.; van Leeuwen, H.C. Genetic markers for Clostridium difficile lineages linked to hypervirulence. Microbiology 2011, 157, 3113–3123. [Google Scholar] [CrossRef]
- Schneeberg, A.; Neubauer, H.; Schmoock, G.; Baier, S.; Harlizius, J.; Nienhoff, H.; Brase, K.; Zimmermann, S.; Seyboldt, C. Clostridium difficile genotypes in piglet populations in Germany. J. Clin. Microbiol. 2013, 51, 3796–3803. [Google Scholar] [CrossRef]
- Álvarez-Pérez, S.; Blanco, J.L.; Harmanus, C.; Kuijper, E.; Garcia, M.E. Subtyping and antimicrobial susceptibility of Clostridium difficile PCR ribotype 078/126 isolates of human and animal origin. Vet. Microbiol. 2017, 199, 15–22. [Google Scholar] [CrossRef]
- Cairns, M.D.; Stabler, R.A.; Shetty, N.; Wren, B.W. The continually evolving Clostridium difficile species. Future Microbiol. 2012, 7, 945–957. [Google Scholar] [CrossRef] [PubMed]
- Cairns, M.D.; Preston, M.D.; Lawley, T.D.; Clark, T.G.; Stabler, R.A.; Wren, B.W. Genomic epidemiology of a protracted hospital outbreak caused by a toxin A-negative Clostridium difficile sublineage PCR ribotype 017 strain in London, England. J. Clin. Microbiol. 2015, 53, 3141–3147. [Google Scholar] [CrossRef] [PubMed]
- Collins, D.A.; Hawkey, P.M.; Riley, T.V. Epidemiology of Clostridium difficile infection in Asia. Antimicrob. Resist. Infect. Control. 2013, 2, 21. [Google Scholar] [CrossRef] [PubMed]
- Spigaglia, P.; Barbanti, F.; Dionisi, A.M.; Mastrantonio, P. Clostridium difficile isolates resistant to fluoroquinolones in Italy: Emergence of PCR ribotype 018. J. Clin. Microbiol. 2010, 48, 2892–2896. [Google Scholar] [CrossRef]
- Bauer, M.P.; Notermans, D.W.; van Benthem, B.H.; Brazier, J.S.; Wilcox, M.H.; Rupnik, M.; Monnet, D.L.; van Dissel, J.T.; Kuijper, E.J. Clostridium difficile infection in Europe: A hospital-based survey. Lancet 2011, 377, 63–73. [Google Scholar] [CrossRef]
- Barbanti, F.; Spigaglia, P. Characterization of Clostridium difficile PCR-ribotype 018: A problematic emerging type. Anaerobe 2016, 42, 123–129. [Google Scholar] [CrossRef]
- Zidaric, V.; Beigot, S.; Lapajne, S.; Rupnik, M. The occurrence and high diversity of Clostridium difficile genotypes in rivers. Anaerobe 2010, 16, 371–375. [Google Scholar] [CrossRef]
- Bazaid, F. Distribution and Sources of Clostridium difficile Present in Water Sources; MSc University of Guelph: Guelph, Canada, 2012. [Google Scholar]
- Warriner, K.; Xu, C.; Habash, M.; Sultan, S.; Weese, S.J. Dissemination of Clostridium difficile in food and the environment: Significant sources of C. difficile community acquired infection? J. Appl. Microbiol. 2016, 122, 542–555. [Google Scholar] [CrossRef]
- Moradigaravand, D.; Gouliouris, T.; Ludden, C.; Reuter, S.; Jamrozy, D.; Blane, B.; Naydenova, P.; Judge, K.H.; Aliyu, S.F.; Hadjirin, N.A.; et al. Genomic survey of Clostridium difficile reservoirs in the East of England implicates environmental contamination of wastewater treatment plants by clinical lineages. Microb. Genom. 2018, 4, e000162. [Google Scholar] [CrossRef]
- Kotila, S.M.; Pitkänen, T.; Brazier, J.; Eerola, E.; Jalava, J.; Kuusi, M.; Könönen, E.; Laine, J.; Miettinen, I.T.; Vuento, R.; et al. Clostridium difficile contamination of public tap water distribution system during a waterborne outbreak in Finland. Scand. J. Public Health 2013, 41, 541–545. [Google Scholar] [CrossRef]
- Al Saif, N.; Brazier, J.S. The distribution of Clostridium difficile in the environment of South Wales. J. Med. Microbiol. 1996, 45, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Baverud, V.; Gustafsson, A.; Franklin, A.; Aspan, A.; Gunnarsson, A. Clostridium difficile: Prevalence in horses and environment, and antimicrobial susceptibility. Equine Vet. J. 2003, 35, 465–471. [Google Scholar] [CrossRef] [PubMed]
- Higazi, T.B.; AL-Saghir, M.; Burkett, M.; Pusok, R. PCR detection of Clostridium difficile and its toxigenic strains in public places in Southeast Ohio. Int. J. Microbiol. Res. 2011, 2, 105–111. [Google Scholar]
- Rodriguez, C.; Bouchafa, L.; Soumillion, K.; Ngyuvula, E.; Taminiau, B.; VanBroeck, J.; Delmée, M.; Daube, G. Seasonality of Clostridium difficile in the natural environment. Transbound. Emerg. Dis. 2019, 66, 2440–2449. [Google Scholar] [CrossRef] [PubMed]
- Dharmasena, M.; Jiang, X. Isolation of toxigenic Clostridium difficile from animal manure and composts being used as biological soil amendments. Appl. Environ. Microbiol. 2018, 84, e00738-18. [Google Scholar] [CrossRef]
- Shivaperumal, N.; Chang, B.J.; Riley, T.V. High Prevalence of Clostridium difficile in Home Gardens in Western Australia. Appl. Environ. Microbiol. J. 2020, 87, e01572-20. [Google Scholar] [CrossRef]
- Fröschle, B.; Messelhäusser, U.; Höller, C.; Lebuhn, M. Fate of Clostridium botulinum and incidence of pathogenic clostridia in biogas processes. J. Appl. Microbiol. 2015, 119, 936–947. [Google Scholar] [CrossRef]
- Marcos, P.; Whyte, P.; Rogers, T.; McElroy, M.; Fanning, S.; Frias, J.; Bolton, D. The prevalence of Clostridioides difficile on farms, in abattoirs and in retail foods in Ireland. Food Microbiol. 2021, 98, 103781. [Google Scholar] [CrossRef]
- Rodriguez, C.; Taminiau, B.; Van Broeck, J.; Avesani, V.; Delmee, M.; Daube, G. Clostridium difficile in young farm animals and slaughter animals in Belgium. Anaerobe 2012, 18, 621–625. [Google Scholar] [CrossRef]
- Wu, Y.C.; Lee, J.J.; Tsai, B.Y.; Liu, Y.F.; Chen, C.M.; Tien, N.; Tsai, P.J.; Chen, T.H. Potentially hypervirulent Clostridium difficile PCR ribotype 078 lineage isolates in pigs and possible implications for humans in Taiwan. Int. J. Med. Microbiol. 2016, 306, 115–122. [Google Scholar] [CrossRef]
- Rodriguez, C.; Hakimi, D.E.; Vanleyssem, R.; Taminiau, B.; Van Broeck, J.; Delmee, M.; Korsak, N.; Daube, G. Clostridium difficile in beef cattle farms, farmers and their environment: Assessing the spread of the bacterium. Vet. Microbiol. 2017, 210, 183–187. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.Y.; Cho, A.; Kim, J.W.; Kim, H.; Kim, B. High prevalence of Clostridium difficile PCR ribotype 078 in pigs in Korea. Anaerobe 2018, 51, 42–46. [Google Scholar] [CrossRef]
- Krutova, M.; Zouharova, M.; Matejkova, J.; Tkadlec, J.; Krejci, J.; Faldyna, M.; Nyc, O.; Bernardy, J. The emergence of Clostridium difficile PCR ribotype 078 in piglets in the Czech Republic clusters with Clostridium difficile PCR ribotype 078 isolates from Germany, Japan and Taiwan. Int. J. Med. Microbiol. 2018, 308, 770–775. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, C.; Taminiau, B.; van Broeck, J.; Daube, G. Clostridium difficile in Food and Animals: A Comprehensive Review. Adv. Exp. Med. Biol. 2016, 932, 65–92. [Google Scholar] [PubMed]
- Keel, K.; Brazier, J.S.; Post, K.W.; Weese, S.; Songer, J.G. Prevalence of PCR Ribotypes among Clostridium difficile Isolates from Pigs, Calves, and Other Species. J. Clin. Microbiol. 2007, 6, 1963–1964. [Google Scholar] [CrossRef]
- Rodriguez-Palacios, A.; Staempfli, H.R.; Duffield, T.; Weese, J.S. Clostridium difficile in Retail Ground Meat, Canada. Emerg. Infect. Dis. 2007, 3, 485–487. [Google Scholar] [CrossRef]
- Knight, D.R.; Riley, T.V. Prevalence of Gastrointestinal Clostridium difficile Carriage in Australian Sheep and Lambs. Appl. Environ. Microbiol. 2013, 79, 5689–5692. [Google Scholar] [CrossRef]
- Harvey, R.; Norman, K.; Andrews, K.; Hume, M.; Scanlan, C.; Callaway, T. Clostridium difficile in poultry and poultry meat. Foodborne Path. Dis. 2011, 8, 1321–1323. [Google Scholar] [CrossRef]
- Abdel-Glil, M.Y.; Thomas, P.; Schmoock, G.; Abou-El-Azm, K.; Wieler, L.H.; Neubauer, H.; Seyboldt, C. Presence of Clostridium difficile in poultry and poultry meat in Egypt. Anaerobe 2018, 51, 21–25. [Google Scholar] [CrossRef]
- Hussain, I.; Borah, P.; Sharma, R.K.; Rajkhowa, S.; Rupnik, M.; Saikia, D.P. Molecular characteristics of Clostridium difficile isolates from human and animals in the North Eastern region of India. Mol. Cell Probes 2016, 30, 306–311. [Google Scholar] [CrossRef]
- Simango, C.; Mwakurudza, S. Clostridium difficile in broiler chickens sold at market places in Zimbabwe and their antimicrobial susceptibility. Int. J. Food Microbiol. 2008, 124, 268–270. [Google Scholar] [CrossRef] [PubMed]
- Zidaric, V.; Zemljic, M.; Janezic, S.; Kocuvan, A.; Rupnik, M. High diversity of Clostridium difficile genotypes isolated from a single poultry farm producing replacement laying hens. Anaerobe 2008, 14, 325–327. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority (EFSA). Scientific Opinion on the public health hazards to be covered by inspection of meat (bovine animals). Eur. Food Saf. Aut. J. 2013, 11, 1–261. [Google Scholar]
- Rodriguez, C.; Avesani, V.; Van Broeck, J.; Taminiau, B.; Delmée, M.; Daube, G. Presence of Clostridium difficile in pigs and cattle intestinal contents and carcass contamination at the slaughterhouse in Belgium. Int. J. Food Microbiol. 2013, 166, 256–262. [Google Scholar] [CrossRef]
- Hopman, N.E.; Oorburg, D.; Sanders, I.; Kuijper, E.J.; Lipman, L.J. High occurrence of various Clostridium difficile PCR ribotypes in pigs arriving at the slaughterhouse. Vet. Quart. 2011, 31, 179–181. [Google Scholar] [CrossRef]
- Hawken, P.; Weese, J.S.; Friendship, R.; Warriner, K. Carriage and dissemination of Clostridium difficile and methicillin resistant Staphylococcus aureus in pork processing. Food Control 2013, 31, 433–437. [Google Scholar] [CrossRef]
- Wu, Y.C.; Chen, C.M.; Kuo, C.J.; Lee, J.J.; Chen, P.C.; Chang, Y.C. Prevalence and molecular characterization of Clostridium difficile isolates from a pig slaughterhouse, pork, and humans in Taiwan. Int. J. Food Microbiol. 2013, 242, 37–44. [Google Scholar] [CrossRef]
- Hampikyan, H.; Bingol, E.B.; Muratoglu, K.; Akkaya, E.; Cetin, O.; Colak, H. The prevalence of Clostridium difficile in cattle and sheep carcasses and the antibiotic susceptibility of isolates. Meat Sci. 2018, 139, 120–124. [Google Scholar] [CrossRef]
- Esfandiari, Z.; Weese, J.S.; Ezzatpanah, H.; Chamani, M.; Shoaei, P.; Yaran, M.; Ataei, B.; Maracy, M.R.; Ansariyan, A.; Ebrahimi, F.; et al. Isolation and characterization of Clostridium difficile in farm animals from slaughterhouse to retail stage in Isfahan, Iran. Food. Path. Dis. 2015, 10, 864–866. [Google Scholar] [CrossRef]
- Candel-Pérez, C.; Santaella-Pascual, J.; Ros-Berruezo, G.; Martínez-Gracia, C. Occurrence of Clostridioides [Clostridium] difficile in poultry giblets at slaughter and retail pork and poultry meat in southeastern Spain. J. Food Prot. 2021, 84, 310–314. [Google Scholar] [CrossRef]
- Keessen, E.C.; van den Berkt, A.J.; Haasjes, N.H.; Hermanus, C.; Kuijper, E.J.; Lipman, L.J. The relation between farm specific factors and prevalence of Clostridium difficile in slaughter pigs. Vet. Microbiol. 2011, 54, 130–134. [Google Scholar] [CrossRef] [PubMed]
- Lund, B.M.; Peck, M.W. A possible route for foodborne transmission of Clostridium difficile? Food. Path. Dis. 2015, 12, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Harvey, R.; Norman, K.; Andrews, K.; Norby, B.; Hume, M.; Scanlan, C.; Hardin, M.; Scott, H. Clostridium difficile in retail meat and processing plants in Texas. J. Vet. Diag. Investig. 2011, 23, 807–811. [Google Scholar] [CrossRef]
- Varshney, J.B.; Very, K.J.; Williams, J.L.; Hegarty, J.P.; Stewart, D.B.; Lumadue, J. Characterization of Clostridium difficile isolates from human fecal samples and retail meat from Pennsylvania. Food. Path. Dis. 2014, 11, 822–829. [Google Scholar] [CrossRef]
- Guran, H.S.; Ilhak, O.I. Clostridium difficile in retail chicken meat parts and liver in the Eastern Region of Turkey. J. Con. Prot. Food Saf. 2015, 10, 359–364. [Google Scholar] [CrossRef]
- Metcalf, D.; Avery, B.P.; Janecko, N.; Matic, N.; Reid-Smith, R.; Weese, J.S. Clostridium difficile in seafood and fish. Anaerobe 2011, 17, 85–86. [Google Scholar] [CrossRef] [PubMed]
- Pasquale, V.; Romano, V.; Rupnik, M.; Capuano, F.; Bove, D.; Aliberti, F.; Krovacek, K.; Dumontet, S. Occurrence of toxigenic Clostridium difficile in edible bivalve molluscs. Food Microbiol. 2012, 31, 309–312. [Google Scholar] [CrossRef]
- Troiano, T.; Harmanus, C.; Sanders, I.; Pasquale, V.; Dumontet, S.; Capuano, F.; Romano, V.; Kuijper, E.J. Toxigenic Clostridium difficile PCR ribotypes in ediblemarine bivalve molluscs in Italy. Int. J. Food Microbiol. 2015, 208, 30–34. [Google Scholar] [CrossRef]
- Agnoletti, F.; Arcangelia, G.; Barbantib, F.; Barcoa, L.; Brunettaa, R.; Cocchia, M.; Conederaa, G.; D’Estea, L.; Drigoa, I.; Spigagliab, P.; et al. Survey, characterization and antimicrobial susceptibility of Clostridium difficile from marine bivalve shellfish of North Adriatic Sea. Int. J. Food Microbiol. 2019, 298, 74–80. [Google Scholar] [CrossRef]
- Eckert, C.; Burghoffer, B.; Barbut, F. Contamination of ready-to-eat raw vegetables with Clostridium difficile, France. J. Med. Microbiol. 2013, 62, 1435–1438. [Google Scholar] [CrossRef]
- Lim, S.C.; Foster, N.F.; Elliott, B.; Riley, T.V. High prevalence of Clostridium difficile on retail root vegetables, Western Australia. J. Appl. Microbiol. 2018, 124, 585–590. [Google Scholar] [CrossRef]
- Tkalec, V.; Janezica, S.; Skoka, B.; Simonica, T.; Mesarica, S.; Vrabica, T.; Rupnik, M. High Clostridium difficile contamination rates of domestic and imported potatoes compared to some other vegetables in Slovenia. Food Microbiol. 2019, 78, 194–200. [Google Scholar] [CrossRef]
- Marcos, P.; Whyte, P.; Burgess, C.; Ekhlas, D.; Bolton, D. Detection and Genomic Characterisation of Clostridioides difficile from Spinach Fields. Pathogens 2022, 11, 1310. [Google Scholar] [CrossRef]
- Bakri, M.M.; Brown, D.J.; Butcher, J.P.; Sutherland, A.D. Clostridium difficile in ready-to-eat salads, Scotland. Emerg. Infect. Dis. 2009, 15, 817–818. [Google Scholar] [CrossRef] [PubMed]
- Dharmasena, M.; Wang, H.; Wei, T.; Bridges, W.C., Jr.; Jiang, X. Survival of Clostridioides difficile in finished dairy compost under controlled conditions. J. Appl. Microbiol. 2021, 131, 996–1006. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Weese, J.S.; Flemming, C.; Odumeru, J.; Warriner, K. Fate of Clostridium difficile during wastewater treatment and incidence in Southern Ontario watersheds. J. Appl. Microbiol. 2014, 117, 891–904. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Sun, M.; Zhang, Y.; Gao, J.; Kong, F.; Liu, D.; Yu, H.; Du, J.; Tang, R. Prevalence, genotype and antimicrobial resistance of Clostridium difficile isolates from healthy pets in Eastern China. BMC Infect. Dis. 2019, 19, 46. [Google Scholar] [CrossRef]
- Weese, J.S. Clostridium (Clostridioides) difficile in animals. J. Vet. Diagn.Investig. Off. Publ. Am. Assoc. Vet. Lab. Diag. 2020, 32, 213–221. [Google Scholar] [CrossRef]
- Hammitt, M.C.; Bueschel, D.M.; Keel, M.K.; Glock, R.D.; Cuneo, P.; DeYoung, D.W.; Reggiardo, C.; Trinh, H.T.; Songer, J.G. A possible role for Clostridium difficile in the etiology of calf enteritis. Vet. Microbiol. 2008, 127, 343–352. [Google Scholar] [CrossRef]
- Costa, M.C.; Stämpfli, H.R.; Arroyo, L.G.; Pearl, D.L.; Weese, J.S. Epidemiology of Clostridium difficile on a veal farm: Prevalence, molecular characterization and tetracycline resistance. Vet. Microbiol. 2011, 152, 379–384. [Google Scholar] [CrossRef]
- Costa, M.C.; Reid-Smith, R.; Gow, S.; Hannon, S.J.; Booker, C.; Rousseau, J.; Benedict, K.M.; Morley, P.S.; Weese, J.S. Prevalence and molecular characterization of Clostridium difficile isolated from feedlot beef cattle upon arrival and mid-feeding period. BMC Vet. Res. 2012, 8, 38. [Google Scholar] [CrossRef] [PubMed]
- Keessen, E.C.; Harmanus, C.; Dohmen, W.; Kuijper, E.J.; Lipman, L.J. Clostridium difficile infection associated with pig farms. Emerg. Infect. Dis. 2013, 19, 1032–1034. [Google Scholar] [CrossRef] [PubMed]
- Magistrali, C.F.; Maresca, C.; Cucco, L.; Bano, L.; Drigo, I.; Filippini, G.; Dettori, A.; Broccatelli, S.; Pezzotti, G. Prevalence and risk factors associated with Clostridium difficile shedding in veal calves in Italy. Anaerobe 2015, 33, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Romano, V.; Albanese, F.; Dumontet, S.; Krovacek, K.; Petrini, O.; Pasquale, V. Prevalence and genotypic characterization of Clostridium difficile from ruminants in Switzerland. Zoo. Pub. Health 2012, 59, 545–548. [Google Scholar] [CrossRef] [PubMed]
- Larson, H.E.; Price, A.B.; Honour, P.; Borriello, S.P. Clostridium difficile and the aetiology of pseudomembranous colitis. Lancet 1978, 1, 1063–1066. [Google Scholar] [CrossRef] [PubMed]
- Olson, M.M.; Shanholtzer, C.J.; Lee, J.T., Jr.; Gerding, D.N. Ten years of prospective Clostridium difficile-associated disease surveillance and treatment at the Minneapolis VA Medical Center, 1982–1991. Infect. Control Hosp. Epidemiol. 1994, 15, 371–381. [Google Scholar] [CrossRef]
- Slimings, C.; Riley, T. Antibiotics and hospital-acquired Clostridium difficile infection: Update of systematic review and meta-analysis. J. Antimicrob. Chemother. 2014, 69, 881–891. [Google Scholar] [CrossRef]
- Freeman, J.; Vernon, J.; Morris, K.; Nicholson, S.; Todhunter, S.; Longshaw, C.; Wilcox, M.H. Pan-European Longitudinal Surveillance of Antibiotic Resistance among Prevalent Clostridium difficile Ribotypes’ Study Group. Clin. Microbiol. Infect. 2015, 21, 248.e9–248.e16. [Google Scholar] [CrossRef]
- McDonald, L.C.; Coignard, B.; Dubberke, E.; Song, X.; Horan, T.; Kutty, P.K.; Ad Hoc Clostridium difficile Surveillance Working Group. Recommendations for surveillance of Clostridium difficile-associated disease. Infect. Control. Hosp. Epidemiol. 2007, 28, 140–145. [Google Scholar] [CrossRef]
- Lessa, F.C.; Mu, Y.; Bamberg, W.M.; Beldavs, Z.G.; Dumyati, G.K.; Dunn, J.R.; Farley, M.M.; Holzbauer, S.M.; Meek, J.I.; Phipps, E.C.; et al. Burden of Clostridium difficile infection in the United States. N. Engl. J. Med. 2015, 372, 825–834. [Google Scholar] [CrossRef]
- Khanna, S. Microbiota restoration for recurrent Clostridioides difficile: Getting one step closer every day! J. Intern. Med. 2021, 290, 294–309. [Google Scholar] [CrossRef] [PubMed]
- Kuntz, J.L.; Chrischilles, E.A.; Pendergast, J.F.; Herwaldt, L.A.; Polgreen, P.M. Incidence of and risk factors for community-associated Clostridium difficile infection: A nested case-control study. BMC Infect. Dis. 2011, 11, 194. [Google Scholar] [CrossRef] [PubMed]
- Moono, P.; Lim, S.C.; Riley, T.V. High prevalence of toxigenic Clostridium difficile in public space lawns in Western Australia. Sci. Rep. 2017, 7, 41196. [Google Scholar] [CrossRef]
- Rodriguez Diaz, C.; Seyboldt, C.; Rupnik, M. Non-human C. difficile reservoirs and sources: Animals, food, environment. Adv. Exp. Med. Biol. 2018, 1050, 227–243. [Google Scholar]
- Avberšek, J.; Pirš, T.; Pate, M.; Rupnik, M.; Ocepek, M. Clostridium difficile in goats and sheep in Slovenia: Characterisation of strains and evidence of age-related shedding. Anaerobe 2014, 28, 163–167. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Palacios, A.; Barman, T.; LeJeune, J.T. Three-week summer period prevalence of Clostridium difficile in farm animals in a temperate region of the United States (Ohio). Can. Vet. J. 2014, 55, 786–789. [Google Scholar] [PubMed]
- Cataldo, M.A.; Granata, G.; Petrosillo, N. Clostridium difficile infection: New approaches to prevention, non-antimicrobial treatment, and stewardship. Expert Rev. Anti-Infect. Ther. 2017, 15, 1027–1040. [Google Scholar] [CrossRef]
- Maron, D.F.; Smith, T.J.; Nachman, K.E. Restrictions on antimicrobial use in food animal production: An international regulatory and economic survey. Glob. Health 2013, 9, 48. [Google Scholar] [CrossRef]
- Squire, M.M.; Riley, T.V. Clostridium difficile infection in humans and piglets: A ‘One Health’ opportunity. Curr. Top. Microbiol. Immunol. 2013, 365, 299–314. [Google Scholar]
- Deng, K.; Plaza-Garrido, A.; Torres, J.A.; Paredes-Sabja, D. Survival of Clostridium difficile spores at low temperatures. Food Microbiol. 2015, 46, 218–221. [Google Scholar] [CrossRef]
- Flock, G.; Chen, C.-H.; Yin, H.-B.; Fancher, S.; Mooyottu, S.; Venkitanarayanan, K. Effect of chilling, freezing and cooking on survivability of Clostridium difficile spores in ground beef. Meat Sci. 2016, 112, 161. [Google Scholar] [CrossRef]
- Rodriguez-Palacios, A.; Reid-Smith, R.J.; Staempfli, H.R.; Weese, J.S. Clostridium difficile survives minimal temperature recommended for cooking ground meats. Anaerobe 2010, 16, 540–542. [Google Scholar] [CrossRef] [PubMed]
- Ojha, S.C.; Chankhamhaengdecha, S.; Singhakaew, S.; Ounjai, P.; Janvilisri, T. Inactivation of Clostridium difficile spores by microwave irradiation. Anaerobe 2016, 38, 14–20. [Google Scholar] [CrossRef] [PubMed]
- Doona, C.J.; Feeherry, F.E.; Setlow, B.; Wang, S.; Li, W.; Nichols, F.C. Effects of high-pressure treatment on spores of Clostridium species. Appl. Environ. Microbiol. 2016, 82, 5287–5297. [Google Scholar] [CrossRef]
- Deng, K.; Talukdar, P.K.; Sarker, M.R.; Paredes-Sabja, D.; Torres, J.A. Survival of Clostridium difficile spores at low water activity. Food Microbiol. 2017, 65, 274–278. [Google Scholar] [CrossRef]
- Broda, D.M.; Delacy, K.M.; Bell, R.G.; Terry, J.; Cook, R.L. Psychrotrophic Clostridium spp. associated with ‘blown pack’ spoilage of chilled vacuum-packed red meats and dog rolls in gas-impermeable plastic casings. Int. J. Food Microbiol. 1996, 29, 335–352. [Google Scholar] [CrossRef]
- Atasoy, F.; Gücükoğlu, A. Detection of Clostridium difficile and toxin genes in samples of modified atmosphere packaged (MAP) minced and cubed beef meat. Ank. ÜNiversitesi Vet. Fakültesi Derg. 2017, 64, 165–170. [Google Scholar]
- Lim, S.C.; Foster, N.F.; Riley, T.V. Susceptibility of Clostridium difficile to the food preservatives sodium nitrite, sodium nitrate and sodium metabisulphite. Anaerobe 2016, 37, 67–71. [Google Scholar] [CrossRef]
- Le Lay, C.; Dridi, L.; Bergeron, M.G.; Ouellette, M.; Fliss, I. Nisin is an effective inhibitor of Clostridium difficile vegetative cells and spore germination. J. Med. Microbiol. 2016, 65, 169–175. [Google Scholar] [CrossRef]
- Aljarallah, K.M. Inhibition of Clostridium difficile by natural herbal extracts. J. Taibah Univ. Med. Sci. 2016, 11, 427–431. [Google Scholar] [CrossRef]
- Roshan, N.; Riley, T.V.; Hammer, K.A. Antimicrobial activity of natural products against Clostridium difficile in vitro. J. Appl. Microbiol. 2017, 123, 92–103. [Google Scholar] [CrossRef] [PubMed]
| Product | Raw or RTE | Total No. (%) Positive | Toxin Gene Profile | Ribotype(s) | Reference |
|---|---|---|---|---|---|
| Ground pork | Raw | 3/7 (41.3%) | A+ B+ CDT+ | 027 078 | [35] |
| Ground pork | Raw | 14/115 (12%) | A+ B+ CDT+ | 027 078 | [36] |
| Ground pork | Raw | 2/66 (3.0%) | A+ B+ CDT− | 029 | [39] |
| Pork meat | Raw | 35/303 (11.5%) | A+ B+ CDT+ | 078 | [136] |
| Pork sausages | RTE | 10/16 (62.5%) | A+ B+ CDT+ | 027 078 | [35] |
| Ground beef | Raw | 13/26 (42.4%) | A+ B+ CDT+ | 027 078 | [35] |
| Ground beef | Raw | 11/53 (20.8%) | A+ B+ CDT+ | M31 | [118] |
| A+ B+ CDT− | 014 077 | ||||
| Ground beef | Raw | 14/115 (12%) | A+ B+ CDT+ | 027 078 | [36] |
| Ground beef | Raw | 2/105 (1.9%) | A+ B+ CDTND | 012 | [37] |
| Ground beef | Raw | 21/303 (6.9%) | A+ B+ CDT+ | PA22 | [136] |
| Beef | Raw | 1/67 (1.5%) | A+ B+ CDT− | 029 | [39] |
| Beef sausages | RTE | 1/7 (14.3%) | A+ B+ CDT+ | 027 | [35] |
| Corned beef | RTE | 1/10 (10%) | AND B+ CDTND | ND 1 | [110] |
| Ground veal | Raw | 1/7 (14.3%) | A+ B+ CDT+ | M31 | [118] |
| Turkey | Raw | 44/303 (14.5%) | A+ B+ CDT+ | PA01 PA05 PA16 | [136] |
| Ground turkey | Raw | 4/9 (44.4%) | A+ B+ CDT+ | 078 | [35] |
| Lamb | Raw | 1/16 (6.3%) | A+ B+ CDT+ | 045 | [38] |
| Chicken | Raw | 7/257 (2.7%) | A+ B+ CDT− | 001 003 071 087 | [38] |
| Chicken | Raw | 1/67 (1.5%) | A+ B+ CDT− | 029 | [39] |
| Chicken | Raw | 25/310 (8.0%) | A+ B+ CDT− | ND 1 | [137] |
| Chicken | Raw | 26/203 (12.8%) | A+ B+ CDT+ | 078 | [23] |
| Chicken | Raw | 24/303 (7.8%) | A+ B+ CDT+ | PA05 PA16 | [136] |
| Chicken | Raw | 4/32 (12.5%) | A+ B+ CDT+ | 078 | [110] |
| Chicken | RTE | 1/130 (0.8%) | A+ B+ CDT− | 014 020 | [41] |
| Shellfish | Raw | 118/702 (16.8%) | A+ B+ CDT+ | 126 475 | [141] |
| Bivalve molluscs | Raw | 26/53 (49%) | A+ B+ CDT+ | 078 | [139] |
| A+ B+ CDT− | 002 012 014/020 018 001 | ||||
| Bivalve molluscs | Raw | 36/925 (3.9%) | A+ B+ CDT+ | 078 126 010 | [140] |
| A− B+ CDT− | 017 001 |
| Product | Raw or RTE | Total No. (%) Positive | Toxin Gene Profile | Ribotype(s) | Reference |
|---|---|---|---|---|---|
| Root vegetables (potatoes, beetroots, onions and carrots) | Raw | 30/100 (30%) | A+ B+ CDT+ | QX 274 | [143] |
| A+ B+ CDT− | 002 137 QX519 QX049 101 | ||||
| A− B+ CDT+ | 070 237 584 | ||||
| A− B− CDT+ | 033 | ||||
| Root vegetables (potatoes, ginger) and leaf vegetables | Raw and RTE | 28/154 (18.2%) | A+ B+ CDT− | 001/072 011/049 014/020 012 070 150 394 SLO129 SLO187 SLO279 | [144] |
| A+B+CDT+ | 027 244 126 023 | ||||
| Lettuce | RTE | 1/54 (1.9%) | A+ B+ CDT+ | 126 | [41] |
| Vegetables (potato, onion, mushroom, carrot, radish and cucumber) | Raw | 7/300 (2.4%) | A+ BND CDT ND | ND 1 | [103] |
| Salad (lettuce, lamb’s lettuce) and vegetable (pea sprouts) | RTE | 3/104 (2.8%) | A+ B+ CDT− | 014/020 001 015 | [142] |
| Vegetables (carrots, potatoes, garlic, ginger, beets, mushrooms, lettuce, green onions, radishes, etc.) | Raw and RTE | 5/111 (4.5%) | A+ B+ CDT+ | 078 | [22] |
| A+ B+ CDT− | ND 1 | ||||
| Salad (baby leaf spinach) | RTE | 2/60 (3.3%) | A+ B+ CDT+ | 078 126 | [145] |
| Salad (baby leaf spinach, organic mixed leaf salad, organic lettuce) | RTE | 3/40 (7.5%) | A+ B+ CDT ND | 001 | [146] |
| A− B+ CDT ND | 017 | ||||
| Spinach leaves | RTE | 2/10 (20%) | A−B+ CDT− | ND 1 | [110] |
| Iceberg lettuce leaves | RTE | 1/10 (10%) | A−B+ CDT− | ND 1 | [110] |
| Little Gem lettuce leaves | RTE | 1/10 (10%) | A−B+ CDT− | ND 1 | [110] |
| Wild rocket leaves | RTE | 1/10 (10%) | A−B+ CDT+ | ND 1 | [110] |
| Coleslaw | RTE | 1/10 (10%) | A−B+ CDT− | ND 1 | [110] |
| Ribotype | Pathogenic | Hypervirulent | CA CDI 1 | Reference(s) | ||||
|---|---|---|---|---|---|---|---|---|
| yes | no | unk 2 | yes | no | unk | |||
| 001 | ✓ | ✓ | [81,99,147,148,149,150] | |||||
| 002 | ✓ | ✓ | ✓ | [81,99,148,149,151,152] | ||||
| 003 | ✓ | ✓ | ✓ | [81,99] | ||||
| 010 | ✓ | ✓ | [150] | |||||
| 011 | ✓ | ✓ | [148] | |||||
| 012 | ✓ | ✓ | ✓ | [81,148,149,150,153] | ||||
| 014 | ✓ | ✓ | ✓ | [81,99,148,151,153] | ||||
| 015 | ✓ | ✓ | [148,149,151] | |||||
| 017 | ✓ | ✓ | [145,148,149,154] | |||||
| 018 | ✓ | ✓ | [148,149] | |||||
| 020 | ✓ | ✓ | [148,149,151] | |||||
| 023 | ✓ | ✓ | [148,149,151] | |||||
| 027 | ✓ | ✓ | ✓ | [72,73,147,148,149,153,155] | ||||
| 029 | ✓ | ✓ | ✓ | [99,153] | ||||
| 070 | ✓ | ✓ | ✓ | [81,149] | ||||
| 071 | ✓ | ✓ | [149] | |||||
| 072 | ✓ | ✓ | [99,148,149,156] | |||||
| 077 | ✓ | ✓ | [149] | |||||
| 078 | ✓ | ✓ | ✓ | [72,73,148,149,153,154,155] | ||||
| 087 | ✓ | ✓ | [148,149] | |||||
| 101 | ✓ | ✓ | [149] | |||||
| 126 | ✓ | ✓ | ✓ | [6,80,81,99,149,153] | ||||
| 137 | ✓ | ✓ | [149] | |||||
| 150 | ✓ | ✓ | [149] | |||||
| 033, 045, 049, 237, 244, 394, 475, 584, M31, PA01, PA05, PA16, PA22, QX049, QX274, QX519, SLO129, SLO187, SLO279 | No information | |||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bolton, D.; Marcos, P. The Environment, Farm Animals and Foods as Sources of Clostridioides difficile Infection in Humans. Foods 2023, 12, 1094. https://doi.org/10.3390/foods12051094
Bolton D, Marcos P. The Environment, Farm Animals and Foods as Sources of Clostridioides difficile Infection in Humans. Foods. 2023; 12(5):1094. https://doi.org/10.3390/foods12051094
Chicago/Turabian StyleBolton, Declan, and Pilar Marcos. 2023. "The Environment, Farm Animals and Foods as Sources of Clostridioides difficile Infection in Humans" Foods 12, no. 5: 1094. https://doi.org/10.3390/foods12051094
APA StyleBolton, D., & Marcos, P. (2023). The Environment, Farm Animals and Foods as Sources of Clostridioides difficile Infection in Humans. Foods, 12(5), 1094. https://doi.org/10.3390/foods12051094






