Characterization of Antibiotic Resistant Coliform Bacteria and Resistance Genes Isolated from Samples of Smoothie Drinks and Raw Milk
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. Identification of Antibiotic-Resistant Strains
2.3. Susceptibility Testing
2.4. Biofilm Production Testing
2.5. Efflux Pumps Overproduction Testing with Ethidium Bromide (EtBr)
2.6. Antibiotic Resistance Genes Detection
3. Results and Discussion
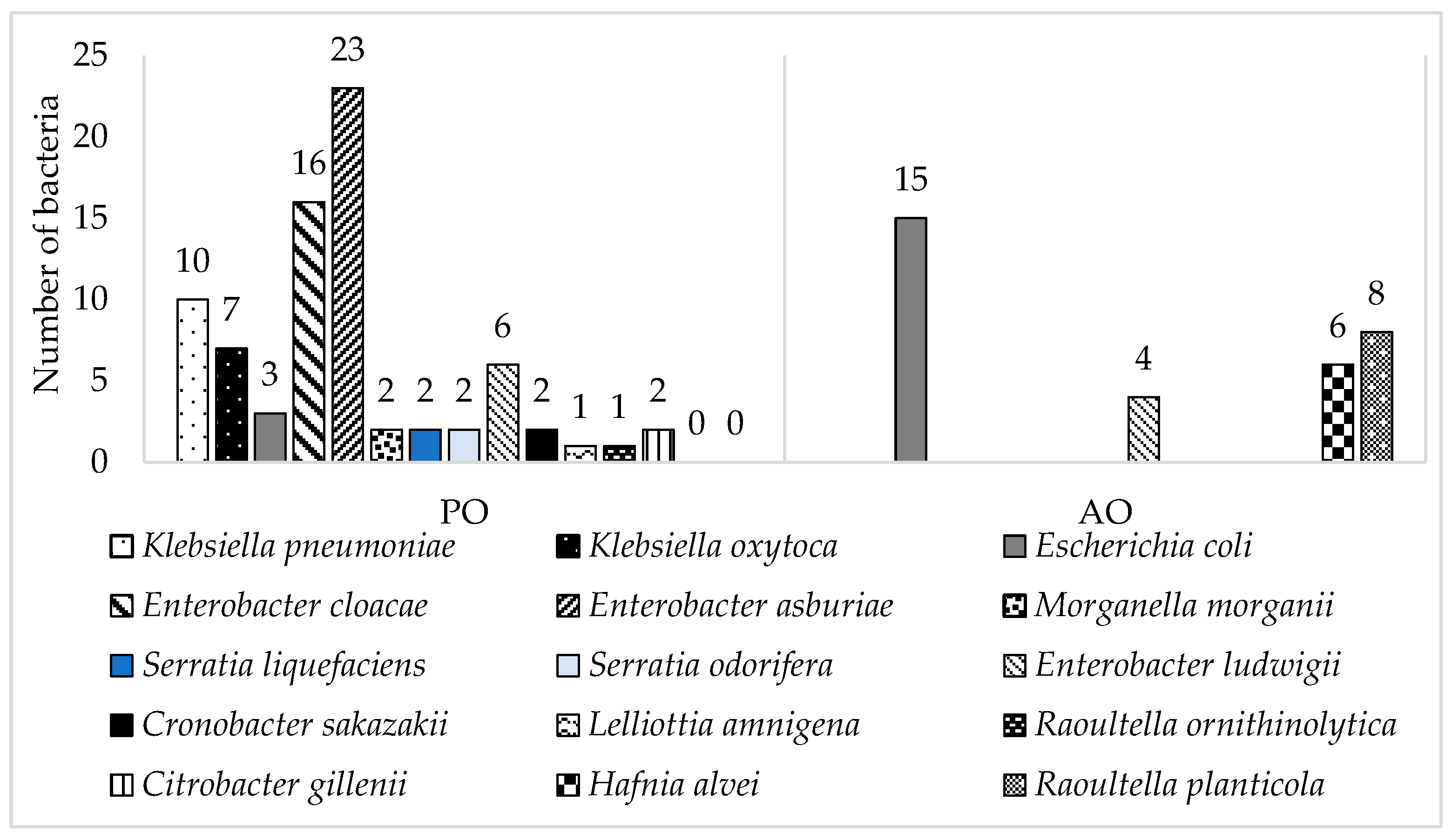
3.1. Antibiotic-Resistant Isolates Identification
3.2. Antibiotic Susceptibility Profile of Isolated Coliform Bacteria
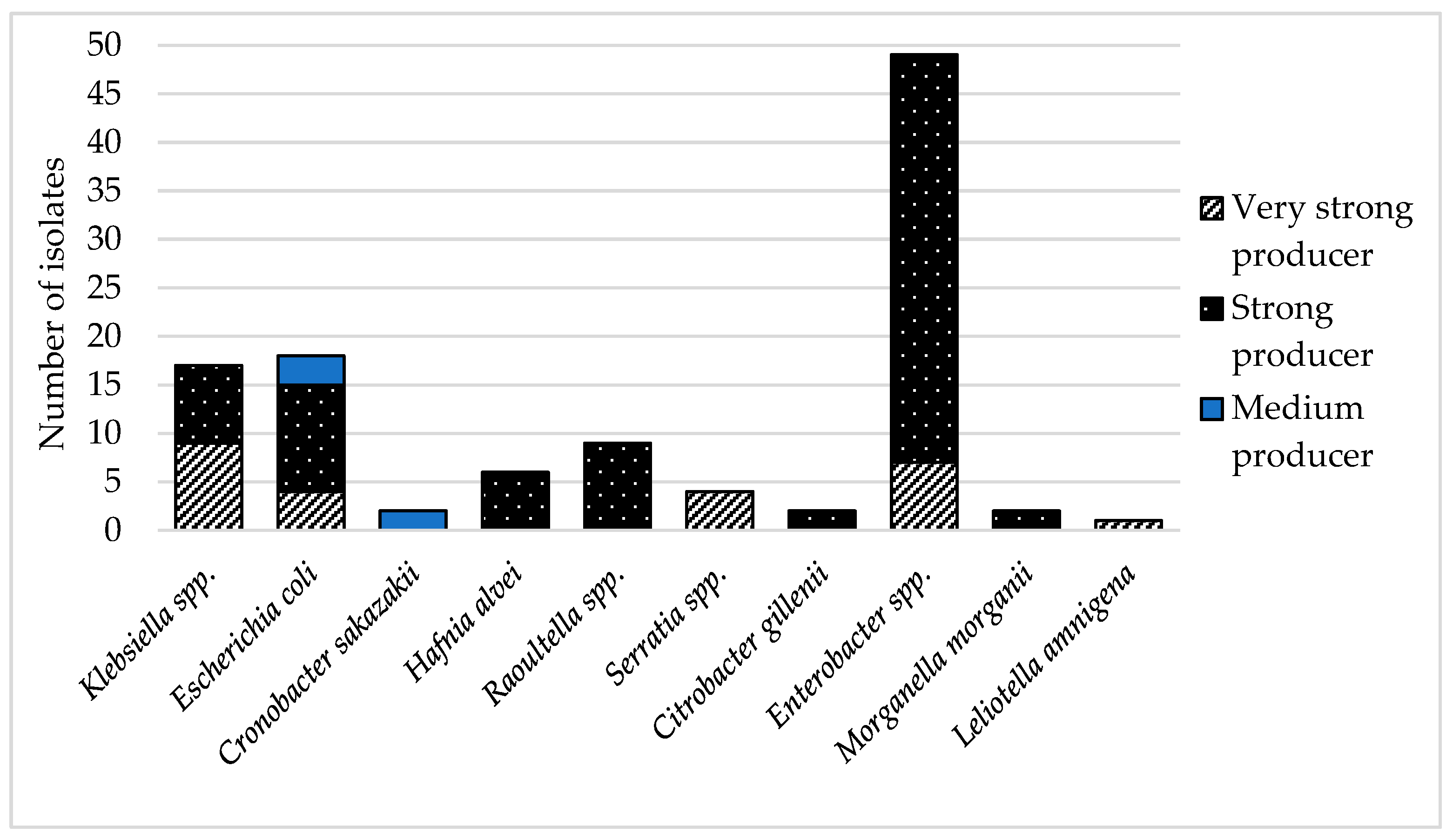
3.3. Antibiotic Resistance Mechanism
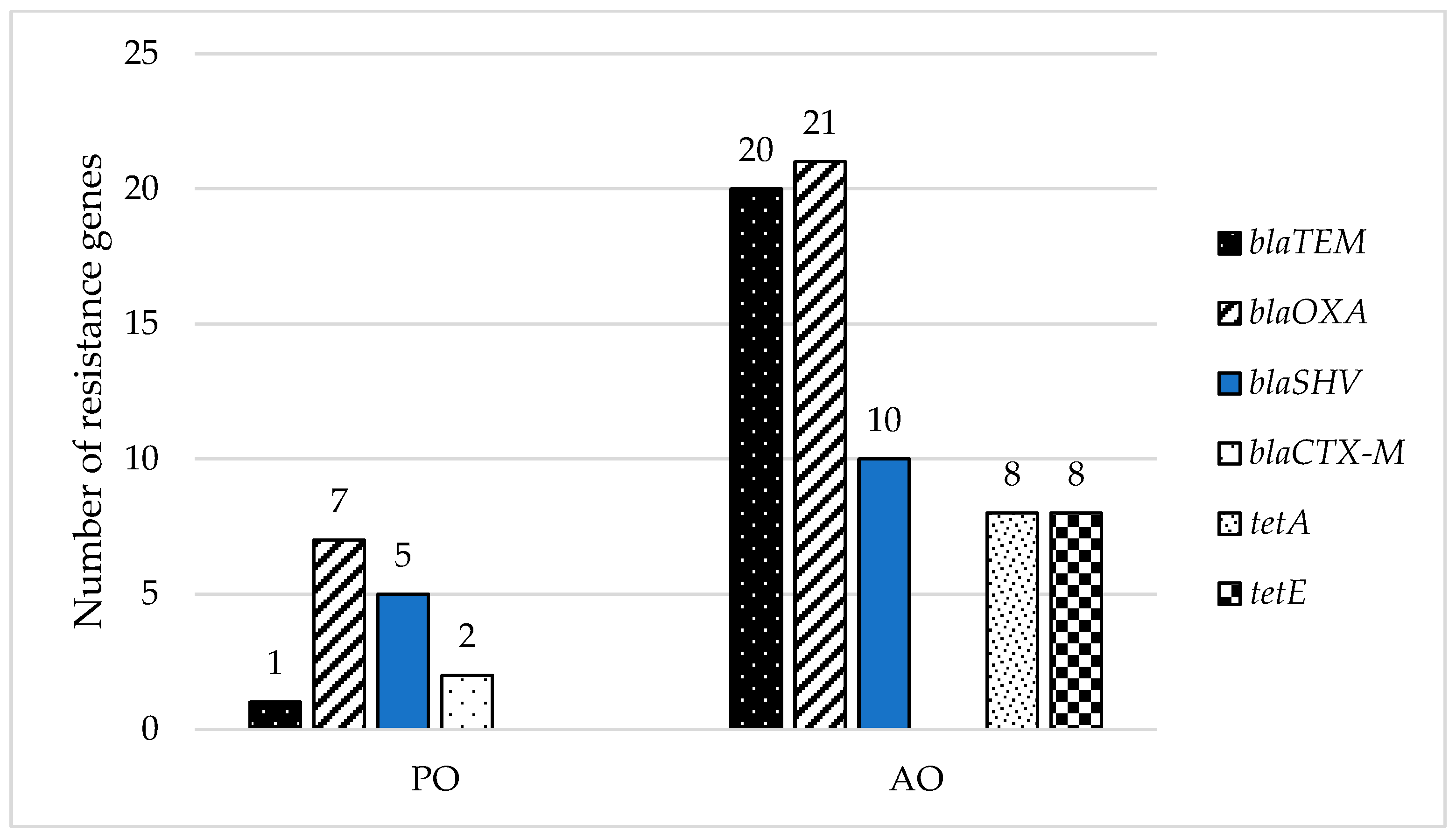
3.4. Important Antibiotic Resistance Genes in Isolated Coliform Bacteria
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- WHO. Better Food and Nutrition in Europe: A Progress Report. Available online: https://www.euro.who.int/en/health-topics/disease-prevention/nutrition/news/news/2018/01/better-food-and-nutrition-in-europe-a-progress-report (accessed on 23 March 2022).
- Koebnick, C.; Garcia, A.L.; Dagnelie, P.C.; Strassner, C.; Lindemans, J.; Katz, N.; Leitzmann, C.; Hoffmann, I. Human Nutrition and Metabolism Long-Term Consumption of a Raw Food Diet Is Associated with Favorable Serum LDL Cholesterol and Triglycerides but Also with Elevated Plasma Homocysteine and Low Serum HDL Cholesterol in Humans. ASN 2005, 135, 2372–2378. [Google Scholar]
- Link, L.B.; Jacobson, J.S. Factors affecting adherence to a raw vegan diet. Complement. Ther. Clin. Pract. 2008, 1, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Olaimat, A.N.; Holley, R.A. Factors in Fl Uencing the Microbial Safety of Fresh Produce: A Review. Food Microbiol. 2012, 32, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Havelaar, A.H.; Brul, S.; De Jong, A.; De Jonge, R.; Zwietering, M.H.; Kuile, H.B. Future challenges to microbial food safetyInternational. Int. J. Food Microbiol. 2010, 139, S79–S94. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. Outbreak of E. coli Infections Linked to Leafy Greens. Available online: https://www.cdc.gov/ecoli/2020/o157h7-10-20b/index.html (accessed on 23 March 2022).
- Centers for Disease Control and Prevention. Outbreak of E. coli Infections Linked to Clover Sprouts. Available online: https://www.cdc.gov/ecoli/2020/o103h2-02-20/index.html (accessed on 23 March 2022).
- Centers for Disease Control and Prevention. E. coli Outbreak Linked to Packaged Salads. Available online: https://www.cdc.gov/ecoli/2021/o157h7-12-21/index.html (accessed on 23 March 2022).
- Centers for Disease Control and Prevention. E. coli Outbreak Linked to Baby Spinach. Available online: https://www.cdc.gov/ecoli/2021/o157h7-11-21/index.html (accessed on 23 March 2022).
- Centers for Disease Control and Prevention. Outbreak of E. coli Infections Linked to Romaine Lettuce. Available online: https://www.cdc.gov/ecoli/2019/o157h7-11-19/index.html (accessed on 23 March 2022).
- Centers for Disease Control and Prevention. Outbreak of E. coli Infections Linked to Romaine Lettuce. Available online: https://www.cdc.gov/ecoli/2018/o157h7-11-18/index.html (accessed on 23 March 2022).
- Centers for Disease Control and Prevention. Multistate Outbreak of E. coli O157:H7 Infections Linked to Romaine Lettuce (Final Update). Available online: https://www.cdc.gov/ecoli/2018/o157h7-04-18/index.html (accessed on 23 March 2022).
- Krahulcová, M.; Micajová, B.; Olejníková, P.; Cverenkárová, K.; Bírošová, L. Microbial Safety of Smoothie Drinks from Fresh Bars Collected in Slovakia. Foods 2021, 10, 551. [Google Scholar] [CrossRef]
- Krahulcová, M.; Lépesová, K.; Bírošová, L. Occurence of Antibiotic Resistant Bacteria in Raw Cow Milk from Vending Machines. Acta Chim. Slov. 2018, 11, 55–59. [Google Scholar] [CrossRef]
- Sebastianski, M.; Bridger, N.A.; Featherstone, R.M.; Robinson, J.L. Disease Outbreaks Linked to Pasteurized and Unpasteurized Dairy Products in Canada and the United States: A Systematic Review. Can. J. Public Health 2022, 1–10. [Google Scholar] [CrossRef]
- WHO. Surveillance of Antimicrobial Resistance in Europe, 2020 Data; WHO: Geneva, Switzerland, 2021; pp. 1–5. ISBN 978-92-890-5629-8. [Google Scholar]
- WHO. New Report Calls for Urgent Action to Avert Antimicrobial Resistance Crisis. Available online: https://www.who.int/news/item/29-04-2019-new-report-calls-for-urgent-action-to-avert-antimicrobial-resistance-crisis (accessed on 23 March 2022).
- Florez-Cuadrado, D.; Moreno, M.A.; Ugarte-Ruíz, M.; Domínguez, L. Antimicrobial Resistance in the Food Chain in the European Union. Adv. Food Nutr. Res. 2018, 86, 115–136. [Google Scholar] [CrossRef]
- Martins, M.; McCusker, M.P.; Viveiros, M.; Couto, I.; Fanning, S.; Pagès, J.; Amaral, L. A Simple Method for Assessment of MDR Bacteria for Over-Expressed Efflux Pumps. Open Microbiol. J. 2013, 7, 72–82. [Google Scholar] [CrossRef]
- Founou, L.L.; Founou, R.C.; Essack, S.Y. Antibiotic Resistance in the Food Chain: A Developing Country-Perspective. Front. Microbiol. 2016, 7, 1–19. [Google Scholar] [CrossRef]
- Verraes, C.; Van Boxstael, S.; Van Meervenne, E.; Van Coillie, E.; Butaye, P.; Catry, B.; Herman, L. Antimicrobial Resistance in the Food Chain: A Review. Int. J. Environ. Res. Public Health 2013, 10, 2643–2669. [Google Scholar] [CrossRef] [PubMed]
- Sharma, D.; Misba, L.; Khan, A.U. Antibiotics Versus Biofilm: An Emerging Battleground in Microbial Communities. Antimicrob. Resist. Infect. Control. 2019, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Van Den Bogaard, A.E.; Stobberingh, E.E. Epidemiology of Resistance to Antibiotics Links between Animals and Humans. Int. J. Antimicrob. Agents 2000, 4, 327–335. [Google Scholar] [CrossRef]
- Guerra, S.T.; Orsi, H.; Joaquim, S.F.; Guimarães, F.F.; Lopes, B.C.; Dalanezi, F.M.; Leite, D.S.; Langoni, H.; Pantoja, J.C.F.; Rall, V.L.M.; et al. Short Communication: Investigation of Extra-Intestinal Pathogenic Escherichia coli Virulence Genes, Bacterial Motility, and Multidrug Resistance Pattern of Strains Isolated from Dairy Cows with Different Severity Scores of Clinical Mastitis. Int. J. Dairy Sci. 2020, 103, 3606–3614. [Google Scholar] [CrossRef] [PubMed]
- Elbehiry, A.; Marzouk, E.; Moussa, I.M.; Alenzi, A.; Al-Maary, K.S.; Kabli, S.A.; Attala, O.A.; Mubarak, A.S. Multidrug-resistant Escherichia coli in Raw Milk: Molecular Characterization and the potential impact of camel’s Urine as an Antibacterial Agent. Saudi J. Biol. Sci. 2021, 28, 2091–2097. [Google Scholar] [CrossRef] [PubMed]
- Wanjala, W.N.; Nduko, J.M.; Mwende, M.C. Coliforms Contamination and Hygienic Status of Milk Chain in Emerging Economies. J. Food Qual. Hazards Control. 2018, 5, 3–10. [Google Scholar] [CrossRef]
- Ozbek, B.; Mataracı-kara, E.; Er, S.; Ozdamar, M.; Yilmaz, M. In Vitro Activities of Colistin, Tigecycline and Tobramycin, Alone or in Combination, against Carbapenem-Resistant Enterobacteriaceae Strains. J. Glob. Antimicrob. Resist. 2015, 3, 278–282. [Google Scholar] [CrossRef]
- Nteimam, J. Screening for Extended-Spectrum Beta-Lactamase-Producing Pathogenic Enterobacteria in District General Hospitals. J. Clin. Microbiol. 2005, 43, 1488–1490. [Google Scholar] [CrossRef]
- Liptakova, A.; Siegfried, L.; Rosocha, J.; Podracka, L.; Bogyiova, E.; Kotulova, D. A Family Outbreak of Haemolytic Uraemic Syndrome and Haemorrhagic Colitis Caused by Verocytotoxigenic Escherichia coli O157 from Unpasteurised Cow’s Milk in Slovakia. Clin. Microbiol. Infect. 2004, 10, 576–578. [Google Scholar] [CrossRef][Green Version]
- Mylius, M.; Dreesman, J.; Pulz, M.; Pallasch, G.; Beyrer, K.; Claußen, K.; Allerberger, F.; Fruth, A.; Lang, C.; Prager, R.; et al. Shiga toxin-producing Escherichia coli O103:H2 outbreak in Germany after school trip to Austria due to raw cow milk, 2017—The important role of international collaboration for outbreak investigations. Int. J. Med. Microbiol. 2018, 308, 539–544. [Google Scholar] [CrossRef]
- WHO. Antimicrobial Resistance: Global Report on Surveillance 2014. Available online: https://apps.who.int/iris/handle/10665/112642 (accessed on 17 April 2022).
- Lépesová, K.; Olejníková, P.; Mackuľak, T.; Tichý, J.; Birošová, L. Annual Changes in the Occurrence of Antibiotic-Resistant Coliform Bacteria and Enterococci in Municipal Wastewater. Environ. Sci. Pollut. Res. 2019, 26, 18470–18483. [Google Scholar] [CrossRef] [PubMed]
- Beenken, K.E.; Blevins, J.S.; Smeltzer, M.S. Mutation of SarA in Staphylococcus aureus Limits Biofilm Formation. Infect. Immun. 2003, 71, 4206–4211. [Google Scholar] [CrossRef] [PubMed]
- Lépesová, K.; Olejníková, P.; Mackuľak, T.; Cverenkárová, K.; Krahulcová, M.; Bírošová, L. Hospital Wastewater—Important Source of Multidrug Resistant Coliform Bacteria with ESBL-Production. Int. J. Environ. Res. Public Health 2020, 17, 7827. [Google Scholar] [CrossRef] [PubMed]
- Dallenne, C.; Da Costa, A.; Decre, D.; Favier, C.H.; Arlet, G. Development of a Set of Multiplex PCR Assays for the Detection of Genes Encoding Important b -Lactamases in Enterobacteriaceae. J. Antimicrob. Chemother. 2018, 65, 490–495. [Google Scholar] [CrossRef] [PubMed]
- Ng, L.; Martin, I.; Alfa, M.; Mulvey, M. Multiplex PCR for the Detection of Tetracycline Resistant Genes. Mol. Cell. Probes. 2001, 15, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Iversen, C. Physiological Description: Enterobacter. In Encyclopedia of Food Microbiology; Academic Press: Amsterdam, The Netherlands, 2014; Volume 1, pp. 598–603. [Google Scholar] [CrossRef]
- Khalifa, A.Y.Z.; Saleh, F.A. Characterization of the Plant Growth Promoting Bacterium, Enterobacter Cloacae MSR1, Isolated from Roots of Non-Nodulating Medicago Sativa. Saudi J. Biol. Sci. 2016, 23, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Mezzatesta, M.L.; Gona, F.; Stefani, S. Enterobacter Cloacae Complex: Clinical Impact and Emerging Antibiotic Resistance. Future Microbiol. 2012, 7, 887–902. [Google Scholar] [CrossRef]
- Salmanov, A.G.; Kolesnik, A.V.; Andriuschenko, D.V. Epidemiology of Intra-Abdominal Infections in Ukraine: Results of a Multicenter Study (2014–2015). Wiad. Lek. 2019, 72, 513–518. [Google Scholar] [CrossRef]
- Nalepa, B.; Anna, M.; Hanna, L. Seasonal Variances in Bacterial Microbiota and Volatile Organic Compounds in Raw Milk. Int. J. Food Microbiol. 2018, 267, 70–76. [Google Scholar] [CrossRef]
- Ruangwittayanusorn, K.; Promket, D.; Chantiratikul, A. Monitoring the Hygiene of Raw Milk from Farms to Milk Retailers. Ital. Oral Surg. 2016, 11, 95–99. [Google Scholar] [CrossRef]
- Júnior, J.C.R.; Silva, F.F.; Lima, J.B.A.; Ossugui, E.H.; Junior, P.I.T.; Campos, A.C.L.P.; Navarro, A.; Tamanini, R.; Ribeiro, J.; Alfieri, A.A.; et al. Short Communication: Molecular Characterization and Antimicrobial Resistance of Pathogenic Escherichia coli Isolated from Raw Milk and Minas Frescal Cheeses in Brazil. J. Dairy Sci. 2019, 102, 10850–10854. [Google Scholar] [CrossRef] [PubMed]
- Rios, E.A.; Santos, J.; García-Meniño, I.; Flament-Simon, S.C.; Blanco, J.; García-lópez, M.; Otero, A.; Rodríguez-Calleja, J.M. Antimicrobial Resistance and Diversity of Atypical EPEC and STEC Isolated from Cow’s Milk, Cheese and Dairy Cattle Farm Environments. LWT—Food Sci. Technol. 2019, 108, 319–325. [Google Scholar] [CrossRef]
- Yu, Z.N.; Wang, J.; Ho, H.; Wang, Y.T.; Huang, S.N.; Han, R.W. Prevalence and Antimicrobial-Resistance Phenotypes and Genotypes of Escherichia coli Isolated from Raw Milk Samples from Mastitis Cases in Four Regions of China. J. Glob. Antimicrob. Resis. 2020, 22, 94–101. [Google Scholar] [CrossRef]
- Abraham, S.; Jordan, D.; Wong, H.S.; Johnson, J.R.; Toleman, M.A.; Wakeham, D.L.; Gordon, D.M.; Turnidge, J.D.; Mollinger, J.L.; Gibson, J.S.; et al. First Detection of Extended-Spectrum Cephalosporin- and Fluoroquinolone-Resistant Escherichia coli in Australian Food-Producing Animals. J. Glob. Antimicrob. Resist. 2015, 3, 273–277. [Google Scholar] [CrossRef]
- Westerveld, D.; Hussain, J.; Aljaafareh, A.; Ataya, A. A Rare Case of Raoultella planticola Pneumonia: An Emerging Pathogen. Respir. Med. Case Rep. 2017, 21, 69–70. [Google Scholar] [CrossRef]
- Puerta-Fernandez, S.; Miralles-Linares, F.; Sanchez-Simonet, M.V.; Bernal-Lopez, M.R.; Gomez-Huelgas, G. Raoultella Planticola Bacteraemia Secondary to Gastroenteritis. Clin. Microbiol. Infect. 2013, 19, E236–E237. [Google Scholar] [CrossRef]
- Sitaula, S.; Shahrrava, A.; Al Zoubi, M.; Malow, J. The First Case Report of Raoultella planticola Liver Abscess. IDCases 2016, 5, 69–71. [Google Scholar] [CrossRef]
- Cooney, S.; Brien, S.O.; Iversen, C.; Fanning, S. Other Pathogenic Enterobacteriaceae—Enterobacter and Other Genera; Elsevier: Amsterdam, The Netherlands, 2014; Volume 1, ISBN 9780123786128. [Google Scholar]
- Litrenta, J.; Oetgen, M. Hafnia Alvei: A New Pathogen in Open Fractures. Trauma Case Rep. 2017, 8, 41–45. [Google Scholar] [CrossRef]
- Ruppé, É.; Woerther, P.L.; Barbier, F. Mechanisms of Antimicrobial Resistance in Gram—Negative Bacilli. Ann. Intensive Care 2015, 5, 1–15. [Google Scholar] [CrossRef]
- Caroff, N.; Espaze, E.; Gautreau, D.; Richet, H.; Reynaud, A. Analysis of the Effects of—42 and—32 AmpC Promoter Mutations in Clinical Isolates of Escherichia coli Hyperproducing AmpC. J. Antimicrob. Chemother. 2000, 45, 783–788. [Google Scholar] [CrossRef] [PubMed]
- Alpay-Karaoglu, S.; Ozgumus, O.B.; Sevim, E.; Kolayli, F.; Sevim, A.; Yesilgil, P. Investigation of Antibiotic Resistance Profile and TEM-Type β-Lactamase Gene Carriage of Ampicillin-Resistant Escherichia coli Strains Isolated from Drinking Water. Ann. Microbiol. 2007, 57, 281–288. [Google Scholar] [CrossRef]
- Koo, H.; Woo, G. Characterization of Antimicrobial Resistance of Escherichia coli Recovered from Foods of Animal and Fish Origin in Korea. J. Food Prot. 2012, 75, 966–972. [Google Scholar] [CrossRef] [PubMed]
- Antunes, N.T.; Frase, H.; Toth, M.; Mobashery, S.; Vakulenko, S.B. Resistance to the Third-Generation Cephalosporin Ceftazidime by a Deacylation-Deficient Mutant of the TEM β-Lactamase by the Uncommon Covalent-Trapping Mechanism. Biochemistry 2011, 50, 6387–6395. [Google Scholar] [CrossRef] [PubMed]
- Potz, N.A.C.; Hope, R.; Warner, M.; Johnson, A.P.; Livermore, D.M. Prevalence and Mechanisms of Cephalosporin Resistance in Enterobacteriaceae in London and South-East England. J. Antimicrob. Chemother. 2006, 58, 320–326. [Google Scholar] [CrossRef]
- Paterson, D.L. Resistance in Gram-Negative Bacteria: Enterobacteriaceae. Am. J. Med. 2006, 119, 20–28. [Google Scholar] [CrossRef]
- Godziszewska, J.; Pogorzelska-Nowicka, E.; Brodowska, M.; Jagura-Burdzy, G.; Wierzbicka, A. Detection in Raw Cow’s Milk of Coliform Bacteria—Reservoir of Antibiotic Resistance. LWT 2018, 93, 634–640. [Google Scholar] [CrossRef]
- EFSA. Scientific Opinion on Chloramphenicol in Food and Feed. Available online: https://www.efsa.europa.eu/en/efsajournal/pub/3907 (accessed on 17 April 2022).
- Birošová, L.; Bodík, I.; Ryba, J.; Jaroslav, Š.; Grabic, R. Pilot Study of Seasonal Occurrence and Distribution of Antibiotics and Drug Resistant Bacteria in Wastewater Treatment Plants in Slovakia. Sci. Total Environ. 2014, 490, 440–444. [Google Scholar] [CrossRef]
- Štefunková, A.; Cverenkárová, K.; Krahulcová, M. Antibiotic Resistant Bacteria in Surface Waters in Slovakia. Acta Chim. Slov. 2020, 13, 63–71. [Google Scholar] [CrossRef]
- Chen, D.; Song, J.; Tang, H.; Yang, L.; Wu, A.; Lin, Y.; Lin, Y. Extensively Drug-Resistant Raoultella planticola Carrying Multiple Resistance Genes Including blaNDM-1. JMMCR 2014, 1, 1–3. [Google Scholar] [CrossRef][Green Version]
- Martin, R.M.; Bachman, M.A. Colonization, Infection, and the Accessory Genome of Klebsiella pneumoniae. Front. Cell Infect. Microbiol. 2018, 8, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Yin, Z.; Yuan, C.; Du, Y.; Yang, P.; Qian, C.; Wei, Y.; Zhang, S. Comparative Genomic Analysis of the Hafnia Genus Reveals an Explicit Evolutionary Relationship between the Species Alvei and Paralvei and Provides Insights into Pathogenicity. BMC Genom. 2019, 20, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Davin-Regli, A.; Lavigne, J.; Pagés, J. Enterobacter spp.: Update on Taxonomy, Clinical Aspects, and Emerging Antimicrobial Resistance. Clin. Microbiol. Rev. 2019, 32, e00002-19. [Google Scholar] [CrossRef] [PubMed]
- dos Santos, G.S.; Solidônio, E.G.; Costa, M.C.V.V.; Melo, R.O.A.; de Souza, I.F.A.C.; Silva-Lecerda, G.R.; Sena, K.X.F.R. Study of the Enterobacteriaceae Group CESP (Citrobacter, Enterobacter, Serratia, Providencia, Morganella and Hafnia): A Review. In The Battle against Microbial Pathogens: Basic Science, Technological Advances and Educational Programs, 1st ed.; Méndez-Vilas, A., Ed.; Formatex Research Center S.L.: Badajoz, Spain, 2015; Volume 1, pp. 794–805. [Google Scholar]
- Stress, O.; Shirshikova, T.V.; Sierra-bakhshi, C.G.; Kamaletdinova, L.K.; Matrosova, L.E.; Khabipova, N.N.; Evtugyn, V.G.; Khilyas, I.V.; Danilova, I.V.; Mardanova, A.M.; et al. The ABC-Type Efflux Pump MacAB Is Involved in Protection of Serratia marcescens against Aminoglycoside Antibiotics, Polymyxins, and Oxidative Stress. mSphere 2021, 6, e00033-21. [Google Scholar] [CrossRef]
- Henriques, I.S.; Fonseca, F.; Alves, A.; José, M.; Correia, A. Occurrence and diversity of integrons and beta-lactamase genes among ampicillin-resistant isolates from estuarine waters. Res. Microbiol. 2006, 10, 938–947. [Google Scholar] [CrossRef]
- Aziz, Z.S.; Albukhaty, S.; Abbood, H.K. Prevalence and antibiotic resistance pattern of certain types of bacterial flora in uterine ewe’s samples. Karbala Int. J. Mod. Sci. 2017, 3, 259–266. [Google Scholar] [CrossRef]
- Butaye, P.; Cloeckaert, A.; Schwarz, S. Mobile Genes Coding for Efflux-Mediated Antimicrobial Resistance in Gram-Positi v e and Gram-Negative Bacteria. Int. J. Antimicrob. Agents 2003, 22, 205–210. [Google Scholar] [CrossRef]
- Overdevest, I.; Willemsen, I.; Rijnsburger, M.; Eustace, A.; Xu, L.; Hawkey, P.; Heck, M.; Savelkoul, P.; Vandenbroucke-grauls, C.; Van Der Zwaluw, K.; et al. Extended-spectrum β-lactamase genes of Escherichia coli in chicken meat and humans, The Netherlands. Emerg. Infect. Dis. 2011, 17, 1216–1222. [Google Scholar] [CrossRef]
- Su, Y.; Yu, C.; Tsai, Y.; Wang, S.; Lee, C.; Chu, C. Fluoroquinolone-resistant and extended-spectrum β-lactamase-producing Escherichia coli from the milk of cows with clinical mastitis in Southern Taiwan. J. Microbiol. Immunol. Infect. 2016, 49, 892–901. [Google Scholar] [CrossRef]
- Helsens, N.; Calvez, S.; Prevost, H.; Bouju-albert, A.; Maillet, A.; Rossero, A.; Hurtaud-pessel, D.; Zagorec, M.; Magras, C. Antibiotic Resistance Genes and Bacterial Communities of Farmed Rainbow Trout Fillets (Oncorhynchus mykiss). Front. Microbiol. 2020, 11, 1–17. [Google Scholar] [CrossRef]
- Tyrrell, J.M.; Wootton, M.; Toleman, M.A.; Howe, R.A.; Woodward, M.; Walsh, T.R. Genetic & virulence profiling of ESBL-positive E. coli from nosocomial & veterinary sources. Vet. Microbiol. 2016, 186, 37–43. [Google Scholar] [CrossRef] [PubMed]
| ATB Class | Antimicrobial | EUCAST (mg/L) > | CLSI (mg/L) ≥ | Higher Than CLSI (mg/L) |
|---|---|---|---|---|
| Penicillins | Ampicillin | 8 | 32 | 50 |
| Cephalosporins | Ceftazidime | 4 | 16 | 32 |
| Carbapenems | Meropenem | 8 | 4 | 12 |
| Fluoroquinolones | Ciprofloxacin | 0,5 | 1 | 2 |
| Aminoglycosides | Gentamicin | 2 | 16 | 20 |
| Amphenicols | Chloramphenicol | 8 | 32 | 50 |
| Tetracyclines | Tetracycline | - | 16 | 32 |
| ATB Class | Gene | Primer | DNA Sequence 5′→3′ | AS (bp) | AT (°C) |
|---|---|---|---|---|---|
| β-Lactams | blaTEM * | fwd | CATTTCCGTGTCGCCCTTATTC | 800 | 54 |
| rev | CGTTCATCCATAGTTGCCTGAC | ||||
| blaSHV * | fwd | AGCCGCTTGAGCAAATTAAAC | 713 | 54 | |
| rev | ATCCCGCAGATAAATCACCAC | ||||
| blaOXA * | fwd | GGCACCAGATTCAACTTTCAAG | 564 | 54 | |
| rev | GACCCCAAGTTTCCTGTAAGTG | ||||
| blaCTX-M group 1 * | fwd | TTAGGAARTGTGCCGCTGYA | 688 | 54 | |
| rev | CGATATCGTTGGTGGTRCCAT | ||||
| Tetracyclines | tetA ** | fwd | GCTACATCCTGCTTGCCTTC | 210 | 55 |
| rev | CATAGATCGCCGTGAAGAGG | ||||
| tetE ** | fwd | AAACCACATCCTCCATACGC | 278 | 55 | |
| rev | AAATAGGCCACAACCGTCAG |
| Resistant Isolate | Sample | AMP | CIP | GEN | CHF | TET | CEF | MER | MDR | Resistant Isolate | Sample | AMP | CIP | GEN | CHF | TET | CEF | MER | MDR |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| E. coli | PO-GS | R | S | R1 | S | S | S | S | - | L. amnigena | PO-FS | R | S | S | S | R1 | S | S | - |
| E. cloacae | PO-GS | R | S | R1 | S | S | S | S | - | S. liquefaciens | PO-FS | R | S | S | S | S | S | S | - |
| K. pneumoniae | PO-GS | R | S | R1 | S | S | S | S | - | E. cloacae | PO-FS | R | R | R1 | R1 | S | S | S | + |
| K. pneumoniae | PO-GS | R | S | R1 | S | S | S | S | - | E. cloacae | PO-FS | R | S | S | R1 | S | S | S | - |
| K. pneumoniae | PO-GS | R | S | R1 | S | S | S | S | - | E. cloacae | PO-GS | R | S | R1 | R1 | S | R | S | + |
| K. pneumoniae | PO-GS | R | S | R1 | S | S | S | S | - | C. gillenii | PO-GS | R | S | R1 | S | S | S | S | - |
| K. pneumoniae | PO-GS | R | S | S | S | S | S | S | - | K. oxytoca | PO-GS | R | S | R1 | S | S | S | R | + |
| K. pneumoniae | PO-GS | R | S | S | R | S | S | S | - | R. ornithinolytica | PO-GS | R | S | S | S | S | S | S | - |
| K. pneumoniae | PO-GS | R | S | S | S | S | S | S | - | E. ludwigii | PO-GS | R | S | S | R1 | S | R | S | + |
| E. coli | PO-GS | R | S | S | R | R1 | S | S | + | E. cloacae | PO-GS | R | S | S | R1 | S | S | S | - |
| E. coli | PO-GS | R | S | S | R | R1 | S | S | + | E. cloacae | PO-GS | R | R | R1 | R1 | S | S | S | + |
| K. pneumoniae | PO-GS | R | S | S | S | S | S | S | - | C. gillenii | PO-GS | R | R | R | R | S | S | S | + |
| K. pneumoniae | PO-GS | R | S | S | S | S | S | S | - | E. cloacae | PO-GS | R | S | S | S | S | S | S | - |
| E. asburiae | PO-GS | R | S | S | S | S | S | S | - | E. cloacae | PO-GS | R | S | S | S | S | S | S | - |
| E. cloacae | PO-GS | R | S | S | S | S | S | S | - | E. cloacae | PO-FS | R | S | S | R1 | S | S | S | - |
| K. pneumoniae | PO-GS | R | S | S | S | S | S | S | - | E. cloacae | PO-FS | R | S | S | R1 | S | S | S | - |
| M. morganii | PO-GS | R | S | S | S | S | S | S | - | E. asburiae | PO-FS | R | S | S | S | S | S | S | - |
| M. morganii | PO-GS | R | S | S | R1 | R1 | S | S | + | E. asburiae | PO-FS | R | S | S | R1 | S | S | S | - |
| K. oxytoca | PO-FS | R | R | R1 | R1 | R1 | R | R | + | S. liquefaciens | PO-FS | R | S | S | S | S | S | S | - |
| K. oxytoca | PO-FS | R | S | S | S | S | S | S | - | E. asburiae | PO-FS | R | S | S | S | S | S | S | - |
| K. oxytoca | PO-FS | R | S | S | R1 | S | S | S | - | E. ludwigii | PO-FS | R | R1 | S | R1 | S | R | S | + |
| K. oxytoca | PO-FS | R | R | R1 | R | S | R | R | + | E. ludwigii | PO-FS | R | S | S | R1 | S | S | S | - |
| E. asburiae | PO-FS | R | S | S | S | S | R | S | - | E. coli | AO | R | S | S | S | S | S | S | - |
| E. asburiae | PO-FS | R | S | S | S | S | R | S | - | E. coli | AO | R | S | S | S | S | S | S | - |
| E. asburiae | PO-FS | R | S | S | S | S | R | S | - | E. coli | AO | R | S | S | S | S | S | S | - |
| E. asburiae | PO-GS | R | S | R1 | S | S | R | S | + | E. coli | AO | R | S | S | S | S | S | S | - |
| E. asburiae | PO-GS | R | S | R1 | S | S | R | S | + | H. alvei | AO | R | S | S | R1 | S | S | S | - |
| E. asburiae | PO-GS | R | S | S | S | S | R | S | - | H. alvei | AO | R | S | S | R1 | S | S | S | - |
| E. asburiae | PO-GS | R | S | S | S | S | R | S | - | H. alvei | AO | R | S | S | R1 | S | S | S | - |
| E. asburiae | PO-GS | R | R2 | R1 | R1 | S | R | R | + | H. alvei | AO | R | S | S | R1 | S | S | S | - |
| E. asburiae | PO-GS | R | S | S | S | S | S | S | - | H. alvei | AO | R | S | S | R1 | S | S | S | - |
| E. asburiae | PO-GS | R | S | R1 | R1 | S | R | S | + | H. alvei | AO | R | S | S | R1 | S | S | S | - |
| E. asburiae | PO-GS | R | S | S | S | S | R | S | - | E. ludwigii | AO | R | S | R1 | S | S | S | S | - |
| E. asburiae | PO-GS | R | S | S | S | S | S | S | - | E. ludwigii | AO | R | S | R1 | S | S | S | S | - |
| E. asburiae | PO-FS | R | S | S | S | S | S | S | - | E. ludwigii | AO | R | S | R1 | S | S | S | S | - |
| E. asburiae | PO-FS | R | S | S | S | S | R | S | - | E. coli | AO | R | S | R1 | S | S | S | S | - |
| E. asburiae | PO-FS | R | S | S | S | S | R | S | - | E. ludwigii | AO | R | S | S | S | S | S | S | - |
| E. asburiae | PO-FS | R | S | S | S | S | R | S | - | E. coli | AO | R | S | S | S | S | R1 | S | - |
| E. asburiae | PO-FS | R | S | S | S | S | R | R | + | E. coli | AO | R | S | S | S | S | R1 | S | - |
| E. asburiae | PO-FS | R | S | S | S | S | R | S | - | E. coli | AO | R | S | S | S | S | R1 | S | - |
| E. asburiae | PO-FS | R | S | R1 | S | R1 | R | S | + | R. planticola | AO | R | S | S | S | R2 | R2 | S | + |
| K. oxytoca | PO-FS | R | S | S | S | S | S | S | - | R. planticola | AO | R | S | S | S | R2 | R2 | S | + |
| K. oxytoca | PO-FS | R | S | S | S | S | S | S | - | R. planticola | AO | R | S | S | S | R2 | R2 | S | + |
| E. cloacae | PO-FS | R | S | R1 | S | S | R | S | + | R. planticola | AO | R | S | S | S | R2 | R2 | S | + |
| E. cloacae | PO-FS | R | R1 | R1 | S | S | S | R | + | R. planticola | AO | R | S | S | S | R2 | R2 | S | + |
| S. odorifera | PO-FS | R | S | R1 | S | S | R | S | + | R. planticola | AO | R | S | S | S | R1 | R1 | S | + |
| S. odorifera | PO-FS | R | S | R1 | S | S | S | S | - | R. planticola | AO | R | S | S | S | R1 | R1 | S | + |
| E. cloacae | PO-FS | R | S | S | S | S | R | S | - | R. planticola | AO | R | S | S | S | R1 | R1 | S | + |
| E. cloacae | PO-FS | R | S | S | S | S | S | S | - | E. coli | AO | R | S | S | R1 | S | R2 | S | + |
| E. ludwigii | PO-FS | R | S | S | S | S | S | S | - | E. coli | AO | R | S | S | R1 | S | R2 | S | + |
| E. ludwigii | PO-FS | R | S | S | S | S | S | S | - | E. coli | AO | R | S | S | R1 | S | R2 | S | + |
| E. cloacae | PO-FS | R | S | R1 | S | S | S | S | - | E. coli | AO | R | S | S | R1 | S | R2 | S | + |
| E. ludwigii | PO-FS | R | S | R | S | S | S | S | - | E. coli | AO | R | S | S | R1 | S | R2 | S | + |
| C. sakazakii | PO-FS | R | S | R1 | S | S | S | S | - | E. coli | AO | R | S | S | R1 | S | R2 | S | + |
| C. sakazakii | PO-FS | R | S | R1 | S | S | S | S | - | E. coli | AO | R | S | S | R1 | S | R2 | S | + |
| Isolates Positive for Resistance Genes | Sample | β-Lactamase Genes | ESBL | Tet Genes | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| blaTEM | blaSHV | blaOXA | blaTEM + blaSHV | blaTEM + blaOXA | blaSHV + blaOXA | blaCTX-M-1 | blaCTX-M-2 | tetA | tetE | ||
| E. coli (n = 2) | PO | 1 | 1 | ||||||||
| E. cloacae (n = 1) | PO | 1 | |||||||||
| K. pneumoniae (n = 5) | PO | 4 | 2 | 1 | |||||||
| K. oxytoca (n = 4) | PO | 4 | |||||||||
| E. asburiae (n = 2) | PO | 1 | 1 | ||||||||
| E. coli (n = 14) | AO | 6 | 1 | 13 | 1 | 5 | |||||
| H. alvei (n = 5) | AO | 2 | 5 | 2 | |||||||
| E. ludwigii (n = 4) | AO | 4 | 4 | 4 | |||||||
| R. planticola (n = 8) | AO | 8 | 8 | 8 | 8 | 8 | |||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Krahulcová, M.; Cverenkárová, K.; Olejníková, P.; Micajová, B.; Koreneková, J.; Bírošová, L. Characterization of Antibiotic Resistant Coliform Bacteria and Resistance Genes Isolated from Samples of Smoothie Drinks and Raw Milk. Foods 2022, 11, 1324. https://doi.org/10.3390/foods11091324
Krahulcová M, Cverenkárová K, Olejníková P, Micajová B, Koreneková J, Bírošová L. Characterization of Antibiotic Resistant Coliform Bacteria and Resistance Genes Isolated from Samples of Smoothie Drinks and Raw Milk. Foods. 2022; 11(9):1324. https://doi.org/10.3390/foods11091324
Chicago/Turabian StyleKrahulcová, Monika, Klára Cverenkárová, Petra Olejníková, Barbora Micajová, Júlia Koreneková, and Lucia Bírošová. 2022. "Characterization of Antibiotic Resistant Coliform Bacteria and Resistance Genes Isolated from Samples of Smoothie Drinks and Raw Milk" Foods 11, no. 9: 1324. https://doi.org/10.3390/foods11091324
APA StyleKrahulcová, M., Cverenkárová, K., Olejníková, P., Micajová, B., Koreneková, J., & Bírošová, L. (2022). Characterization of Antibiotic Resistant Coliform Bacteria and Resistance Genes Isolated from Samples of Smoothie Drinks and Raw Milk. Foods, 11(9), 1324. https://doi.org/10.3390/foods11091324