Evaluation of the Persistence and Characterization of Listeria monocytogenes in Foodservice Operations
Abstract
:1. Introduction
2. Materials and Methods
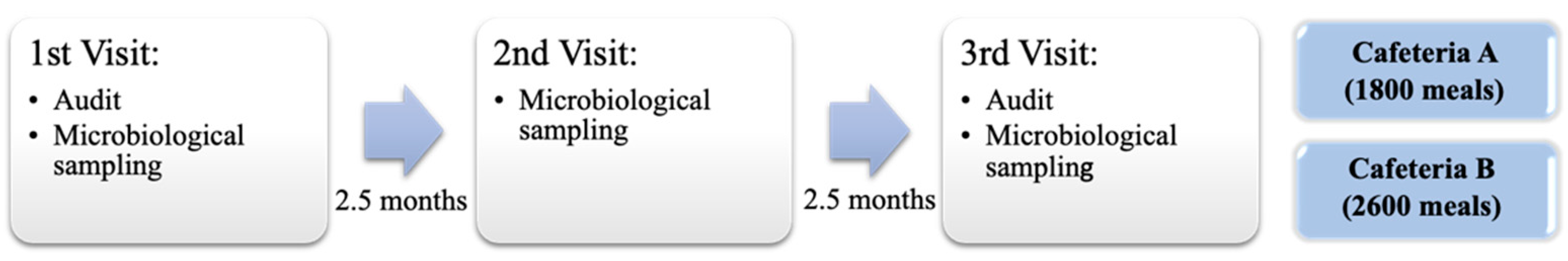
2.1. Description of Food Services Establishments
2.2. Microbiological Sampling
2.2.1. Food Products Sampling
2.2.2. Environmental Sampling
2.3. Microbiological Analysis and Bacterial Identification
2.3.1. Microbiological Analysis of Food Samples
2.3.2. Microbiological Analysis of Environmental Samples
2.4. L. monocytogenes Isolate Characterization
2.4.1. Whole Genome Sequencing
2.4.2. Genome Assembly and Genomic Characterization
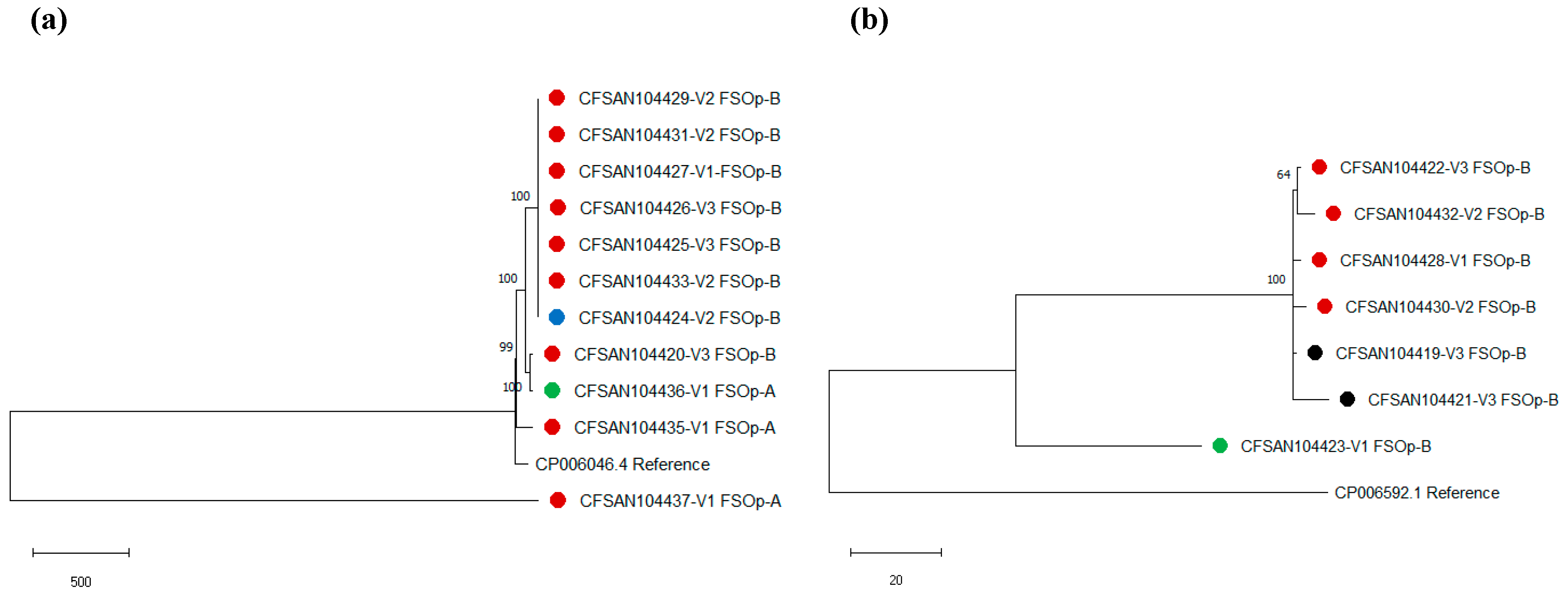
2.4.3. Phylogenetic Relatedness among L. monocytogenes Isolates
2.4.4. Determination of the Minimal Inhibitory Concentration for Quaternary Ammonium (MIC-QA)
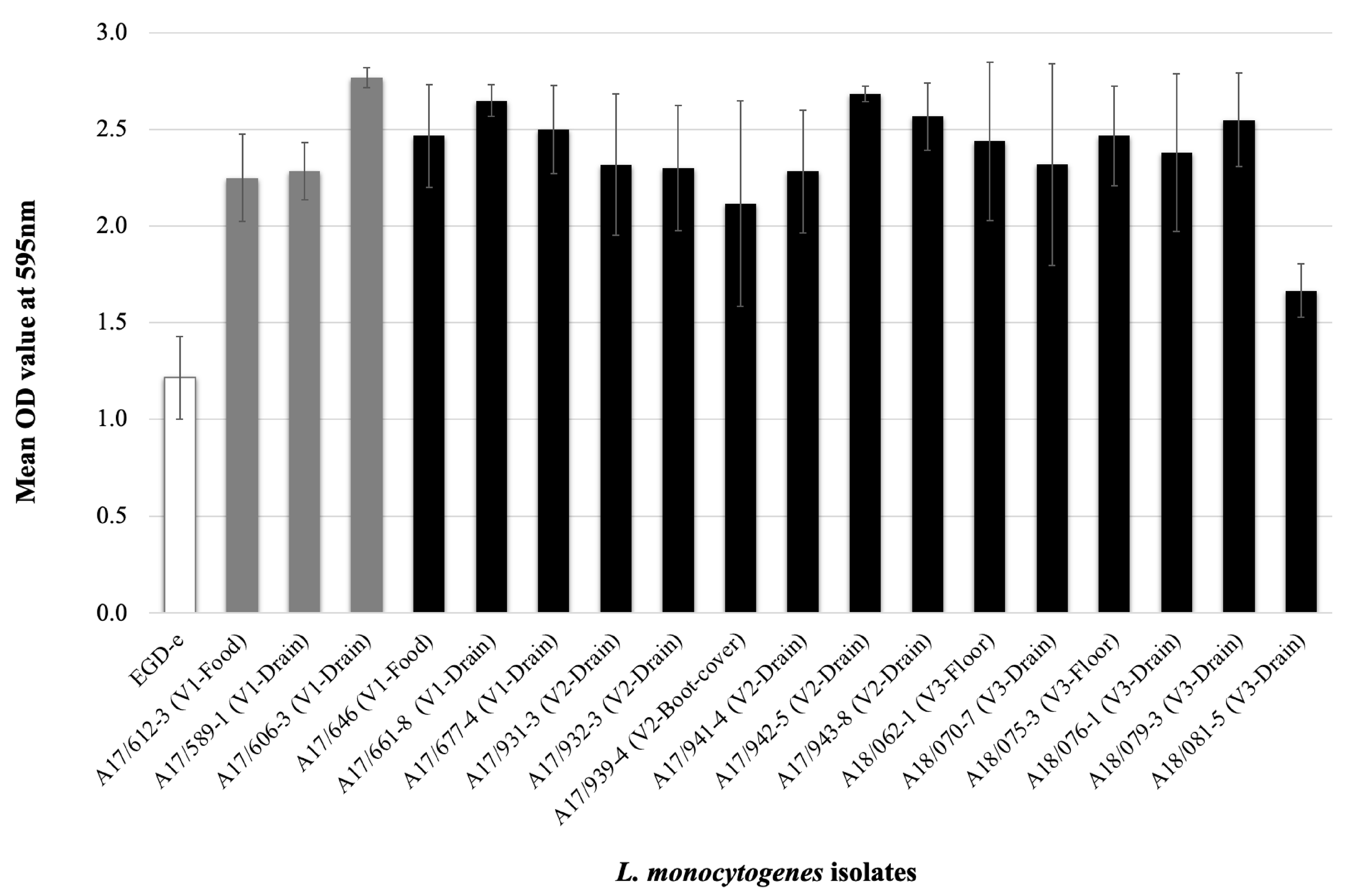
2.4.5. Evaluation of Biofilm Formation
2.5. Audits and Structure of the Checklist
2.6. Statistical Analysis
3. Results
3.1. Food Microbiological Analysis
3.2. Evaluation of Contact Surfaces Contamination
3.3. L. monocytogenes Genome Characterization
3.4. L. monocytogenes Phenotype Characterization
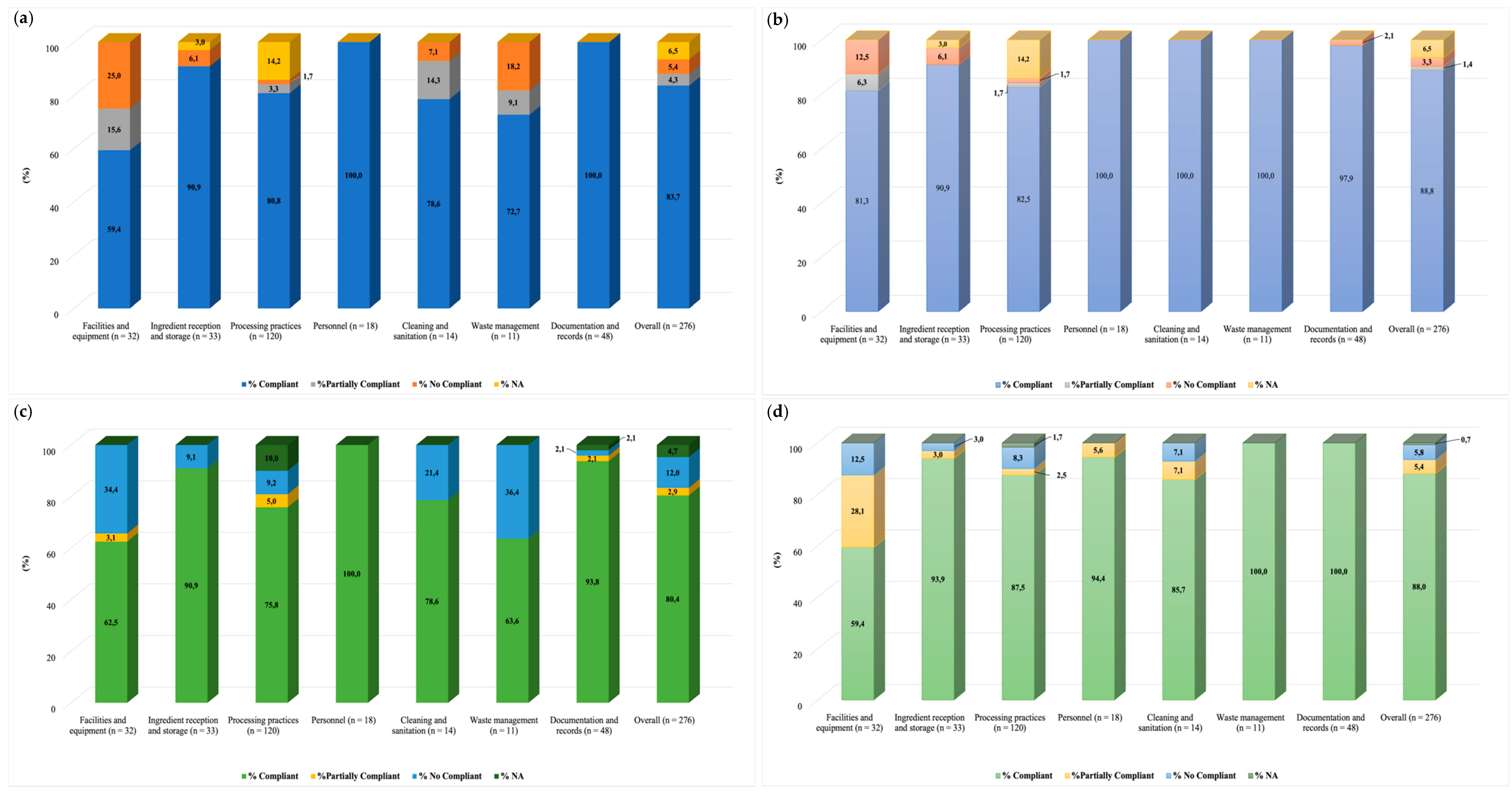
3.5. Audits: Evaluation of Good Manufacturing Practices
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Buchanan, R.L.; Gorris, L.G.M.; Hayman, M.M.; Jackson, T.C.; Whiting, R.C. A review of Listeria monocytogenes: An update on outbreaks, virulence, dose-response, ecology, and risk assessments. Food Control 2017, 75, 13. [Google Scholar] [CrossRef]
- Charlier, C.; Perrodeau, É.; Leclercq, A.; Cazenave, B.; Pilmis, B.; Henry, B.; Lopes, A.; Maury, M.M.; Moura, A.; Goffinet, F.; et al. Clinical features and prognostic factors of listeriosis: The MONALISA national prospective cohort study. Lancet Infect. Dis. 2017, 17, 510–519. [Google Scholar] [CrossRef]
- Ooi, S.T.; Lorber, B. Gastroenteritis due to Listeria monocytogenes. Clin. Infect. Dis. 2005, 40, 1327–1332. [Google Scholar] [CrossRef]
- Schlech, W.F. Epidemiology and clinical manifestations of Listeria monocytogenes infection. Microbiol. Spectr. 2019, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Silva, S.; Teixeira, P.; Oliveira, R.; Azeredo, J. Adhesion to and viability of Listeria monocytogenes on food contact surfaces. J. Food Prot. 2008, 71, 1379–1385. [Google Scholar] [CrossRef]
- Carpentier, B.; Cerf, O. Review—Persistence of Listeria monocytogenes in food industry equipment and premises. Int. J. Food Microbiol. 2011, 145, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, V.; Wiedmann, M.; Teixeira, P.; Stasiewicz, M.J. Listeria monocytogenes persistence in food-associated environments: Epidemiology, strain characteristics, and implications for public health. J. Food Prot. 2014, 77, 150–170. [Google Scholar] [CrossRef]
- Rodríguez-López, P.; Rodríguez-Herrera, J.J.; Vázquez-Sánchez, D.; López Cabo, M. Current knowledge on Listeria monocytogenes biofilms in food-related environments: Incidence, resistance to biocides, ecology and biocontrol. Foods 2018, 7, 85. [Google Scholar] [CrossRef] [Green Version]
- Pan, Y.; Breidt, F.; Kathariou, S. Resistance of Listeria monocytogenes biofilms to sanitizing agents in a simulated food processing environment. Appl. Environ. Microbiol. 2006, 72, 7711–7717. [Google Scholar] [CrossRef] [Green Version]
- Nakamura, H.; Takakura, K.; Sone, Y.; Itano, Y.; Nishikawa, Y. Biofilm formation and resistance to benzalkonium chloride in Listeria monocytogenes isolated from a fish processing plant. J. Food Prot. 2013, 76, 1179–1186. [Google Scholar] [CrossRef]
- Ortiz, S.; López, V.; Martínez-Suárez, J.V. The influence of subminimal inhibitory concentrations of benzalkonium chloride on biofilm formation by Listeria monocytogenes. Int. J. Food Microbiol. 2014, 189, 106–112. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Melcón, C.; Capita, R.; Rodríguez-Jerez, J.J.; Martínez-Suárez, J.V.; Alonso-Calleja, C. Effect of low doses of disinfectants on the biofilm-forming ability of Listeria monocytogenes. Foodborne Pathog. Dis. 2019, 16, 262–268. [Google Scholar] [CrossRef] [PubMed]
- Tamburro, M.; Ripabelli, G.; Vitullo, M.; Dallman, T.J.; Pontello, M.; Amar, C.F.; Sammarco, M.L. Gene expression in Listeria monocytogenes exposed to sublethal concentration of benzalkonium chloride. Comp. Immunol. Microbiol. Infect. Dis. 2015, 40, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.; Yu, T.; Xu, Y.; Wang, H.; Korkeala, H.; Shi, L. MdrL, a major facilitator superfamily efflux pump of Listeria monocytogenes involved in tolerance to benzalkonium chloride. Appl. Microbiol. Biotechnol. 2019, 103, 1339–1350. [Google Scholar] [CrossRef]
- Minarovičová, J.; Véghová, A.; Mikulášová, M.; Chovanová, R.; Šoltýs, K.; Drahovská, H.; Kaclíková, E. Benzalkonium chloride tolerance of Listeria monocytogenes strains isolated from a meat processing facility is related to presence of plasmid-borne bcrABC cassette. Antonie Van Leeuwenhoek 2018, 111, 1913–1923. [Google Scholar] [CrossRef]
- Malley, T.J.; Butts, J.; Wiedmann, M. Seek and destroy process: Listeria monocytogenes process controls in the ready-to-eat meat and poultry industry. J. Food Prot. 2015, 78, 436–445. [Google Scholar] [CrossRef]
- Véghová, A.; Minarovičová, J.; Koreňová, J.; Drahovská, H.; Kaclíková, E. Prevalence and tracing of persistent Listeria monocytogenes strains in meat processing facility production chain. J. Food Saf. 2017, 37, e12315. [Google Scholar] [CrossRef]
- Camargo, A.C.; Woodward, J.J.; Call, D.R.; Nero, L.A. Listeria monocytogenes in food-processing facilities, food contamination, and human listeriosis: The Brazilian scenario. Foodborne Pathog. Dis. 2017, 14, 623–636. [Google Scholar] [CrossRef]
- Leong, D.; Alvarez-Ordóñez, A.; Jordan, K. Monitoring occurrence and persistence of Listeria monocytogenes in foods and food processing environments in the Republic of Ireland. Front. Microbiol. 2014, 5, 436. [Google Scholar] [CrossRef]
- Leong, D.; NicAogáin, K.; Luque-Sastre, L.; McManamon, O.; Hunt, K.; Alvarez-Ordóñez, A.; Scollard, J.; Schmalenberger, A.; Fanning, S.; O’Byrne, C.; et al. A 3-year multi-food study of the presence and persistence of Listeria monocytogenes in 54 small food businesses in Ireland. Int J. Food Microbiol. 2017, 249, 18–26. [Google Scholar] [CrossRef]
- Stasiewicz, M.J.; Oliver, H.F.; Wiedmann, M.; den Bakker, H.C. Whole-genome sequencing allows for improved identification of persistent Listeria monocytogenes in food-associated environments. Appl. Environ. Microbiol. 2015, 81, 6024–6037. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.; Burall, L.S.; Luo, Y.; Timme, R.; Melka, D.; Muruvanda, T.; Payne, J.; Wang, C.; Kastanis, G.; Maounounen-Laasri, A.; et al. Listeria monocytogenes in stone fruits linked to a multistate outbreak: Enumeration of cells and whole-genome sequencing. Appl. Environ. Microbiol. 2016, 82, 7030–7040. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vongkamjan, K.; Roof, S.; Stasiewicz, M.J.; Wiedmann, M. Persistent Listeria monocytogenes subtypes isolated from a smoked fish processing facility included both phage susceptible and resistant isolates. Food Microbiol. 2013, 35, 38–48. [Google Scholar] [CrossRef] [PubMed]
- Castro, H.; Jaakkonen, A.; Hakkinen, M.; Korkeala, H.; Lindström, M. Occurrence, persistence, and contamination routes of Listeria monocytogenes genotypes on three finnish dairy cattle Farms: A longitudinal study. Appl. Environ. Microbiol. 2018, 84, e02000-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harrand, A.S.; Jagadeesan, B.; Baert, L.; Wiedmann, M.; Orsi, R.H. Evolution of Listeria monocytogenes in a food processing plant involves limited single-nucleotide substitutions but considerable diversification by gain and loss of prophages. Appl. Environ. Microbiol. 2020, 86, e02493-19. [Google Scholar] [CrossRef] [PubMed]
- Hurley, D.; Luque-Sastre, L.; Parker, C.T.; Huynh, S.; Eshwar, A.K.; Nguyen, S.V.; Andrews, N.; Moura, A.; Fox, E.M.; Jordan, K.; et al. Whole-genome sequencing-based characterization of 100 Listeria monocytogenes isolates collected from food processing environments over a four-year period. MSphere 2019, 4, e00252-19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Camargo, A.C.; Moura, A.; Avillan, J.; Herman, N.; McFarland, A.P.; Sreevatsan, S.; Call, D.R.; Woodward, J.J.; Lecuit, M.; Nero, L.A. Whole-genome sequencing reveals Listeria monocytogenes diversity and allows identification of long-term persistent strains in Brazil. Environ. Microbiol. 2019, 21, 4478–4487. [Google Scholar] [CrossRef]
- Smith, A.; Hearn, J.; Taylor, C.; Wheelhouse, N.; Kaczmarek, M.; Moorhouse, E.; Singleton, I. Listeria monocytogenes isolates from ready to eat plant produce are diverse and have virulence potential. Int. J. Food Microbiol. 2019, 299, 23–32. [Google Scholar] [CrossRef] [Green Version]
- Schirone, M.; Visciano, P.; Di Serafino, G.; Tofalo, R.; Ciccarelli, A.; Suzzi, G. Assessment of knowledge and applications of hygiene practices in the food service sector. J. Food Saf. 2018, 38, e12457. [Google Scholar] [CrossRef]
- Spanu, C.; Jordan, K. Listeria monocytogenes environmental sampling program in ready-to-eat processing facilities: A practical approach. Compr. Rev. Food Sci. Food Saf. 2020, 19, 2843–2861. [Google Scholar] [CrossRef]
- Henriques, A.R.; Gama, L.T.; Fraqueza, M.J. Tracking Listeria monocytogenes contamination and virulence-associated characteristics in the ready-to-eat meat-based food products industry according to the hygiene level. Int. J. Food Microbiol. 2017, 242, 101–106. [Google Scholar] [CrossRef] [PubMed]
- Kotsanopoulos, K.V.; Arvanitoyannis, I.S. The role of auditing, food safety, and food quality standards in the food industry: A review. Compr. Rev. Food Sci. Food Saf. 2017, 16, 760–775. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Center for Food Safety and Applied Nutrition. Draft Guidance for Industry: Control of Listeria Monocytogenes in Ready-to-Eat Foods; FDA-2008-D-0096; Center for Food Safety and Applied Nutrition: College Park, MD, USA, 2017. [Google Scholar]
- Food Standards Chile. Reglamento Sanitario de los Alimentos (RSA). 2019. Available online: https://www.minsal.cl/reglamento-sanitario-de-los-alimentos/ (accessed on 6 May 2021).
- ISO 4833-2; Microbiology of the Food Chain—Horizontal Method for the Enumeration of Microorganisms—Part 2: Colony Count at 30 °C by the Surface Plating Technique. International Organization for Standardization: Geneva, Switzerland, 2013.
- ISO 21528; Microbiology of the Food Chain—Horizontal Methods for the Detection and Enumeration of Enterobacteriaceae—Part 2: Colony-Count Technique. International Organization for Standardization: Geneva, Switzerland, 2017.
- Feng, P.; Weagant, S.; Grant, M.; Burkhardt, W. BAM Chapter 4: Enumeration of Escherichia coli and the Coliform bacteria. In Bacteriological Analytical Manual; US Food and Drug Administration: White Oak, MD, USA, 2020. [Google Scholar]
- ISO 6888; Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of Coagulase-Positive Staphylococci (Staphylococcus aureus and Other Species)—Part 1: Technique Using Baird-Parker Agar Medium. International Organization for Standardization: Geneva, Switzerland, 2021.
- ISO 7937; Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of Clostridium perfringens—Colony-Count Technique. International Organization for Standardization: Geneva, Switzerland, 2004.
- ISO 21871; Microbiology of Food and Animal Feeding Stuffs: Horizontal Method for the Determination of Low Numbers of Presumptive Bacillus cereus—Most Probable Number Technique and Detection Method. International Organization for Standardization: Geneva, Switzerland, 2006.
- ISO21527-1; Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of Yeasts and Molds—Part 1: Colony Count Technique in Products with Water Activity Greater than 0,95. International Organization for Standardization: Geneva, Switzerland, 2008.
- ISO6579; Microbiology of the Food Chain—Horizontal Method for the Detection, Enumeration and Serotyping of Salmonella—Part 1: Detection of Salmonella spp. International Organization for Standardization: Geneva, Switzerland, 2017.
- Kim, J.S.; Lee, G.G.; Park, J.S.; Jung, Y.H.; Kwak, H.S.; Kim, S.B.; Nam, Y.S.; Kwon, S.T. A novel multiplex PCR assay for rapid and simultaneous detection of five pathogenic bacteria: Escherichia coli O157:H7, Salmonella, Staphylococcus aureus, Listeria monocytogenes, and Vibrio parahaemolyticus. J. Food Prot. 2007, 70, 1656–1662. [Google Scholar] [CrossRef] [PubMed]
- Hitchins, A.D.; Jinneman, K.; Chen, Y. Bacteriological Analytical Manual (BAM) Chapter 10: Detection of Listeria monocytogenes in Foods and Environmental Samples, and Enumeration of Listeria monocytogenes in Foods; US Food and Drug Administration: White Oak, MD, USA, 2017. Available online: https://www.fda.gov/food/laboratory-methods-food/bam-chapter-10-detection-listeria-monocytogenes-foods-and-environ-585mental-samples-and-enumeration (accessed on 7 May 2021).
- Parra, A.; Toro, M.; Jacob, R.; Navarrete, P.; Troncoso, M.; Figueroa, G.; Reyes-Jara, A. Antimicrobial effect of copper surfaces on bacteria isolated from poultry meat. Braz. J. Microbiol. 2018, 49 (Suppl. S1), 113–118. [Google Scholar] [CrossRef] [PubMed]
- Losito, P.; Visciano, P.; Genualdo, M.; Satalino, R.; Migailo, M.; Ostuni, A.; Luisi, A.; Cardone, G. Evaluation of hygienic conditions of food contact surfaces in retail outlets: Six years of monitoring. LWT 2017, 77, 67–71. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [Green Version]
- Gurevich, A.; Saveliev, V.; Vyahhi, N.; Tesler, G. QUAST: Quality assessment tool for genome assemblies. Bioinformatics 2013, 29, 1072–1075. [Google Scholar] [CrossRef]
- Chen, Y.; Pouillot, R.; Dennis, S.; Xian, Z.; Luchansky, J.B.; Porto-Fett, A.C.S.; Lindsay, J.A.; Hammack, T.S.; Allard, M.; Van Doren, J.M.; et al. Genetic diversity and profiles of genes associated with virulence and stress resistance among isolates from the 2010–2013 interagency Listeria monocytogenes market basket survey. PLoS ONE 2020, 15, e0231393. [Google Scholar] [CrossRef]
- Kaas, R.S.; Leekitcharoenphon, P.; Aarestrup, F.M.; Lund, O. Solving the problem of comparing whole bacterial genomes across different sequencing platforms. PLoS ONE 2014, 9, e104984. [Google Scholar] [CrossRef] [Green Version]
- Stecher, G.; Tamura, K.; Kumar, S. Molecular Evolutionary Genetics Analysis (MEGA) for macOS. Mol. Biol. Evol. 2020, 37, 1237–1239. [Google Scholar] [CrossRef] [PubMed]
- Toledo, V.; den Bakker, H.C.; Hormazábal, J.C.; González-Rocha, G.; Bello-Toledo, H.; Toro, M.; Moreno-Switt, A.I. Genomic diversity of Listeria monocytogenes isolated from clinical and non-clinical samples in Chile. Genes 2018, 9, 396. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hammons, S.R.; Stasiewicz, M.J.; Roof, S.; Oliver, H.F. Aerobic plate counts and ATP levels correlate with Listeria monocytogenes detection in retail delis. J. Food Prot. 2015, 78, 825–830. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; ISBN 978-3-319-24277-4. Available online: https://ggplot2.tidyverse.org (accessed on 15 December 2021).
- Tortorello, M.L. Indicator organisms for safety and quality—Uses and methods for detection: Minireview. J. AOAC Int. 2003, 86, 1208–1217. [Google Scholar] [CrossRef] [PubMed]
- Zoellner, C.; Venegas, F.; Churey, J.J.; Dávila-Aviña, J.; Grohn, Y.T.; García, S.; Heredia, N.; Worobo, R.W. Microbial dynamics of indicator microorganisms on fresh tomatoes in the supply chain from Mexico to the USA. Int. J. Food Microbiol. 2016, 238, 202–207. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority, European Centre for Disease Prevention and Control. Multi-country outbreak of Listeria monocytogenes serogroup IVb, multi-locus sequence type 6, infections linked to frozen corn and possibly to other frozen vegetables—First update. EFSA Supporting Publ. 2018, 15, 1448E. [Google Scholar] [CrossRef]
- Koutsoumanis, K.; Alvarez-Ordóñez, A.; Bolton, D.; Bover-Cid, S.; Chemaly, M.; Davies, R.; De Cesare, A.; Herman, L.; Hilbert, F.; Lindqvist, R.; et al. The public health risk posed by Listeria monocytogenes in frozen fruit and vegetables including herbs, blanched during processing. EFSA J. 2020, 18, e06092. [Google Scholar] [CrossRef] [Green Version]
- Centers for Disease Control and Prevention (CDC). Multistate Outbreak of Listeriosis Linked to Frozen Vegetables (Final Update). 2016. Available online: https://www.cdc.gov/listeria/outbreaks/frozen-vegetables-05-16/index.html (accessed on 29 October 2021).
- Willis, C.; McLauchlin, J.; Aird, H.; Amar, C.; Barker, C.; Dallman, T.; Elviss, N.; Lai, S.; Sadler-Reeves, L. Occurrence of Listeria and Escherichia coli in frozen fruit and vegetables collected from retail and catering premises in England 2018–2019. Int. J. Food Microbiol. 2020, 334, 108849. [Google Scholar] [CrossRef]
- Mir, S.A.; Shah, M.A.; Mir, M.M.; Dar, B.N.; Greiner, R.; Roohinejad, S. Microbiological contamination of ready-to-eat vegetable salads in developing countries and potential solutions in the supply chain to control microbial pathogens. Food Control 2018, 85, 235–244. [Google Scholar] [CrossRef]
- Ricci, A.; Allende, A.; Bolton, D.; Chemaly, M.; Davies, R.; Fernández Escámez, P.S.; Girones, R.; Herman, L.; Koutsoumanis, K.; Nørrung, B.; et al. Listeria monocytogenes contamination of ready-to-eat foods and the risk for human health in the EU. EFSA J. 2018, 16, e05134. [Google Scholar] [CrossRef]
- Cordano, A.M.; Jacquet, C. Listeria monocytogenes isolated from vegetable salads sold at supermarkets in Santiago, Chile: Prevalence and strain characterization. Int. J. Food Microbiol. 2009, 132, 176–179. [Google Scholar] [CrossRef] [PubMed]
- Scollon, A.M.; Wang, H.; Ryser, E.T. Transfer of Listeria monocytogenes during mechanical slicing of onions. Food Control 2016, 65, 160–167. [Google Scholar] [CrossRef] [Green Version]
- Garner, D.; Kathariou, S. Fresh produce-associated listeriosis outbreaks, sources of concern, teachable moments, and insights. J. Food Prot. 2016, 79, 337–344. [Google Scholar] [CrossRef] [PubMed]
- Self, J.L.; Conrad, A.; Stroika, S.; Jackson, A.; Whitlock, L.; Jackson, K.A.; Beal, J.; Wellman, A.; Fatica, M.K.; Bidol, S.; et al. Multistate outbreak of listeriosis associated with packaged leafy green salads, United States and Canada, 2015–2016. Emerg. Infect. Dis. 2019, 25, 1461–1468. [Google Scholar] [CrossRef]
- Chen, M.; Chen, Y.; Wu, Q.; Zhang, J.; Cheng, J.; Li, F.; Zeng, H.; Lei, T.; Pang, R.; Ye, Q.; et al. Genetic characteristics and virulence of Listeria monocytogenes isolated from fresh vegetables in China. BMC Microbiol. 2019, 19, 119. [Google Scholar] [CrossRef]
- Sneed, J.; Strohbehn, C.; Gilmore, S.A.; Mendonca, A. Microbiological evaluation of foodservice contact surfaces in Iowa assisted-living facilities. J. Am. Diet. Assoc. 2004, 104, 1722–1724. [Google Scholar] [CrossRef]
- Barria, C.; Malecki, M.; Arraiano, C.M. Bacterial adaptation to cold. Microbiology 2013, 159, 2437–2443. [Google Scholar] [CrossRef] [Green Version]
- Tasara, T.; Stephan, R. Cold stress tolerance of Listeria monocytogenes: A review of molecular adaptive mechanisms and food safety implications. J. Food Prot. 2006, 69, 1473–1484. [Google Scholar] [CrossRef]
- Chan, Y.C.; Wiedmann, M. Physiology and genetics of Listeria monocytogenes survival and growth at cold temperatures. Crit. Rev. Food Sci. Nutr. 2009, 49, 237–253. [Google Scholar] [CrossRef]
- Bucur, F.I.; Grigore-Gurgu, L.; Crauwels, P.; Riedel, C.U.; Nicolau, A.I. Resistance of Listeria monocytogenes to stress conditions encountered in food and food processing environments. Front. Microbiol. 2018, 9, 2700. [Google Scholar] [CrossRef] [Green Version]
- Montero, D.; Bodero, M.; Riveros, G.; Lapierre, L.; Gaggero, A.; Vidal, R.M.; Vidal, M. Molecular epidemiology and genetic diversity of Listeria monocytogenes isolates from a wide variety of ready-to-eat foods and their relationship to clinical strains from listeriosis outbreaks in Chile. Front. Microbiol. 2015, 6, 384. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Magdovitz, B.F.; Gummalla, S.; Thippareddi, H.; Harrison, M.A. Evaluating environmental monitoring protocols for Listeria spp. and Listeria monocytogenes in frozen food manufacturing facilities. J. Food Prot. 2020, 83, 172–187. [Google Scholar] [CrossRef] [PubMed]
- Simmons, C.K.; Wiedmann, M. Identification and classification of sampling sites for pathogen environmental monitoring programs for Listeria monocytogenes: Results from an expert elicitation. Food Microbiol. 2018, 75, 2–17. [Google Scholar] [CrossRef] [PubMed]
- Orsi, R.H.; den Bakker, H.C.; Wiedmann, M. Listeria monocytogenes lineages: Genomics, evolution, ecology, and phenotypic characteristics. Int. J. Med. Microbiol. 2011, 301, 79–96. [Google Scholar] [CrossRef] [PubMed]
- Raschle, S.; Stephan, R.; Stevens, M.J.A.; Cernela, N.; Zurfluh, K.; Muchaamba, F.; Nüesch-Inderbinen, M. Environmental dissemination of pathogenic Listeria monocytogenes in flowing surface waters in Switzerland. Sci. Rep. 2021, 11, 9066. [Google Scholar] [CrossRef] [PubMed]
- Moura, A.; Lefrancq, N.; Wirth, T.; Leclercq, A.; Borges, V.; Gilpin, B.; Dallman, T.J.; Frey, J.; Franz, E.; Nielsen, E.M.; et al. Emergence and global spread of Listeria monocytogenes main clinical clonal complex. Sci. Adv. 2021, 7, eabj9805. [Google Scholar] [CrossRef]
- Lachtara, B.; Osek, J.; Wieczorek, K. Molecular Typing of Listeria monocytogenes IVb serogroup isolated from food and food production environments in Poland. Pathogens 2021, 10, 482. [Google Scholar] [CrossRef]
- Hingston, P.; Chen, J.; Dhillon, B.K.; Laing, C.; Bertelli, C.; Gannon, V.; Tasara, T.; Allen, K.; Brinkman, F.S.; Truelstrup Hansen, L.; et al. Genotypes associated with Listeria monocytogenes Isolates displaying impaired or enhanced tolerances to cold, salt, acid, or desiccation stress. Front. Microbiol. 2017, 8, 369. [Google Scholar] [CrossRef] [Green Version]
- Gadea, R.; Fernández Fuentes, M.; Pérez Pulido, R.; Gálvez, A.; Ortega, E. Effects of exposure to quaternary-ammonium-based biocides on antimicrobial susceptibility and tolerance to physical stresses in bacteria from organic foods. Food Microbiol. 2017, 63, 58–71. [Google Scholar] [CrossRef]
- Müller, A.; Rychli, K.; Zaiser, A.; Wieser, C.; Wagner, M.; Schmitz-Esser, S. The Listeria monocytogenes transposon Tn6188 provides increased tolerance to various quaternary ammonium compounds and ethidium bromide. FEMS Microbiol. Lett. 2014, 361, 166–173. [Google Scholar] [CrossRef] [Green Version]
- Haubert, L.; Zehetmeyr, M.L.; da Silva, W.P. Resistance to benzalkonium chloride and cadmium chloride in Listeria monocytogenes isolates from food and food-processing environments in southern Brazil. Can. J. Microbiol. 2019, 65, 429–435. [Google Scholar] [CrossRef] [PubMed]
- Lake, F.B.; van Overbeek, L.S.; Baars, J.J.P.; Koomen, J.; Abee, T.; den Besten, H.M.W. Genomic characteristics of Listeria monocytogenes isolated during mushroom (Agaricus bisporus) production and processing. Int. J. Food Microbiol. 2021, 360, 109438. [Google Scholar] [CrossRef] [PubMed]
- Møretrø, T.; Schirmer, B.C.T.; Heir, E.; Fagerlund, A.; Hjemli, P.; Langsrud, S. Tolerance to quaternary ammonium compound disinfectants may enhance growth of Listeria monocytogenes in the food industry. Int. J. Food Microbiol. 2017, 241, 215–224. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mullapudi, S.; Siletzky, R.M.; Kathariou, S. Heavy-metal and benzalkonium chloride resistance of Listeria monocytogenes isolates from the environment of turkey-processing plants. Appl. Environ. Microbiol. 2008, 74, 1464–1468. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martínez-Suárez, J.V.; Ortiz, S.; López-Alonso, V. Potential impact of the resistance to quaternary ammonium disinfectants on the persistence of Listeria monocytogenes in food processing environments. Front. Microbiol. 2016, 7, 638. [Google Scholar] [CrossRef] [Green Version]
- Beresford, M.R.; Andrew, P.W.; Shama, G. Listeria monocytogenes adheres to many materials found in food-processing environments. J. Appl. Microbiol. 2001, 90, 1000–1005. [Google Scholar] [CrossRef]
- Bridier, A.; Sanchez-Vizuete, P.; Guilbaud, M.; Piard, J.C.; Naïtali, M.; Briandet, R. Biofilm-associated persistence of food-borne pathogens. Food Microbiol. 2015, 45, 167–178. [Google Scholar] [CrossRef]
- Colagiorgi, A.; Bruini, I.; Di Ciccio, P.A.; Zanardi, E.; Ghidini, S.; Ianieri, A. Listeria monocytogenes biofilms in the wonderland of food industry. Pathogens 2017, 6, 41. [Google Scholar] [CrossRef] [Green Version]
- Dzieciol, M.; Schornsteiner, E.; Muhterem-Uyar, M.; Stessl, B.; Wagner, M.; Schmitz-Esser, S. Bacterial diversity of floor drain biofilms and drain waters in a Listeria monocytogenes contaminated food processing environment. Int. J. Food Microbiol. 2016, 223, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, H.; Wu, C.; Deng, W.; Wang, D.; Zhao, G.; Song, J.; Jiang, Y. Molecular analysis of dominant species in Listeria monocytogenes-positive biofilms in the drains of food processing facilities. Appl. Microbiol. Biotechnol. 2016, 100, 3165–3175. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.H.; Cole, S.; Badel-Berchoux, S.; Guillier, L.; Felix, B.; Krezdorn, N.; Hébraud, M.; Bernardi, T.; Sultan, I.; Piveteau, P. Biofilm formation of Listeria monocytogenes strains under food processing environments and pan-genome-wide association study. Front. Microbiol. 2019, 10, 2698. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nowak, J.; Cruz, C.D.; Palmer, J.; Fletcher, G.C.; Flint, S. Biofilm formation of the L. monocytogenes strain 15G01 is influenced by changes in environmental conditions. J. Microbiol. Methods 2015, 119, 189–195. [Google Scholar] [CrossRef]
- Kadam, S.R.; den Besten, H.M.; van der Veen, S.; Zwietering, M.H.; Moezelaar, R.; Abee, T. Diversity assessment of Listeria monocytogenes biofilm formation: Impact of growth condition, serotype and strain origin. Int. J. Food Microbiol. 2013, 165, 259–264. [Google Scholar] [CrossRef] [PubMed]
- Doijad, S.P.; Barbuddhe, S.B.; Garg, S.; Poharkar, K.V.; Kalorey, D.R.; Kurkure, N.V.; Rawool, D.B.; Chakraborty, T. Biofilm-forming abilities of Listeria monocytogenes serotypes isolated from different sources. PLoS ONE 2015, 10, e0137046. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jordan, S.J.; Perni, S.; Glenn, S.; Fernandes, I.; Barbosa, M.; Sol, M.; Tenreiro, R.P.; Chambel, L.; Barata, B.; Zilhao, I.; et al. Listeria monocytogenes biofilm-associated protein (BapL) may contribute to surface attachment of L. monocytogenes but is absent from many field isolates. Appl. Environ. Microbiol. 2008, 74, 5451–5456. [Google Scholar] [CrossRef] [Green Version]
- Popowska, M.; Krawczyk-Balska, A.; Ostrowski, R.; Desvaux, M. InlL from Listeria monocytogenes is involved in biofilm formation and adhesion to mucin. Front. Microbiol. 2017, 8, 660. [Google Scholar] [CrossRef] [Green Version]
- Lourenço, A.; de Las Heras, A.; Scortti, M.; Vazquez-Boland, J.; Frank, J.F.; Brito, L. Comparison of Listeria monocytogenes Exoproteomes from biofilm and planktonic state: Lmo2504, a protein associated with biofilms. Appl. Environ. Microbiol. 2013, 79, 6075–6082. [Google Scholar] [CrossRef] [Green Version]
- Chang, Y.; Gu, W.; Fischer, N.; McLandsborough, L. Identification of genes involved in Listeria monocytogenes biofilm formation by mariner-based transposon mutagenesis. Appl. Microbiol. Biotechnol. 2012, 93, 2051–2062. [Google Scholar] [CrossRef]
- Sela, S.; Frank, S.; Belausov, E.; Pinto, R. A Mutation in the luxS gene influences Listeria monocytogenes biofilm formation. Appl. Environ. Microbiol. 2006, 72, 5653–5658. [Google Scholar] [CrossRef] [Green Version]
- Da Silva, D.A.L.; de Melo Tavares, R.; Camargo, A.C.; Yamatogi, R.S.; De Martinis, E.C.P.; Nero, L.A. Biofilm growth by Listeria monocytogenes on stainless steel and expression of biofilm-related genes under stressing conditions. World J. Microbiol. Biotechnol. 2021, 37, 119. [Google Scholar] [CrossRef]
- Santos, T.; Viala, D.; Chambon, C.; Esbelin, J.; Hébraud, M. Biofilm adaptation to different temperatures seen through shotgun proteomics. Front. Nutr. 2019, 6, 89. [Google Scholar] [CrossRef]
- Disanto, C.; Celano, G.; Dambrosio, A.; Quaglia, N.C.; Bozzo, G.; Tritto, A.; Celano, G.V. Food safety in collective catering: Knowledge, attitudes and correct application of GHP/GMP knowledge among foodservice workers. Ital. J. Food Saf. 2020, 9, 8453. [Google Scholar] [CrossRef] [PubMed]
- Chapman, B.; Eversley, T.; Fillion, K.; Maclaurin, T.; Powell, D. Assessment of food safety practices of food service food handlers (risk assessment data): Testing a communication intervention (evaluation of tools). J. Food Prot. 2010, 73, 1101–1107. [Google Scholar] [CrossRef] [PubMed]
- Ulloa-Bello, M. Enfermedades Transmitidas Por los Alimentos en Chile: Agentes Causantes y Factores Contribuyentes Asociados a Brotes Ocurridos Durante el Año 2013. Master’s Thesis, Universidad de Chile, Santiago, Chile, 2013. Available online: http://repositorio.uchile.cl/handle/2250/138263 (accessed on 29 October 2021).
- Lianou, A.; Sofos, J.N. A review of the incidence and transmission of Listeria monocytogenes in ready-to-eat products in retail and food service environments. J. Food Prot. 2007, 70, 2172–2198. [Google Scholar] [CrossRef] [PubMed]
- Doménech, E.; Amorós, J.A.; Escriche, I. Food safety objectives for Listeria monocytogenes in Spanish food sampled in cafeterias and restaurants. J. Food Prot. 2011, 74, 1569–1573. [Google Scholar] [CrossRef]
- Lockis, V.R.; Cruz, A.G.; Walter, E.H.; Faria, J.A.; Granato, D.; Sant’Ana, A.S. Prerequisite programs at schools: Diagnosis and economic evaluation. Foodborne Pathog. Dis. 2011, 8, 213–220. [Google Scholar] [CrossRef]
- Bartsch, S.M.; Asti, L.; Nyathi, S.; Spiker, M.L.; Lee, B.Y. Estimated cost to a restaurant of a foodborne illness outbreak. Public Health Rep. 2018, 133, 274–286. [Google Scholar] [CrossRef] [Green Version]
- Rebouças, L.; Santiago, L.; Martins, L.; Rios Menezes, A.; Araújo, M.; Almeida, R. Food safety knowledge and practices of food handlers, head chefs and managers in hotels’ restaurants of Salvador, Brazil. Food Control 2017, 73, 372–381. [Google Scholar] [CrossRef]
- Al-Maqbali, A.A.; Al-Abri, S.S.; Vidyanand, V.; Al-Abaidani, I.; Al-Balushi, A.S.; Bawikar, S.; El Amir, E.; Al-Azri, S.; Kumar, R.; Al-Rashdi, A.; et al. Community foodborne of Salmonella Weltevreden outbreak at Northern Governorate, Sultanate of Oman. J. Epidemiol. Glob. Health 2021, 11, 224–229. [Google Scholar] [CrossRef]
| FSOp | Zone | No Samples Analyzed | |||
|---|---|---|---|---|---|
| Visit 1 | Visit 2 | Visit 3 | Total Samples | ||
| A | 1 | 5 | 4 | 5 | 14 |
| 2 | 1 | 2 | 2 | 5 | |
| 3 | 13 | 14 | 13 | 40 | |
| 4 | 1 | 0 | 0 | 1 | |
| Total | 20 | 20 | 20 | 60 | |
| B | 1 | 6 | 6 | 6 | 18 |
| 2 | 5 | 6 | 6 | 17 | |
| 3 | 14 * | 18 | 18 | 50 | |
| 4 | 1 | 0 | 0 | 1 | |
| Total | 26 * | 30 | 30 | 86 | |
| APC | Enterobacteriaceae | E. coli | Salmonella spp. | S. aureus | C. perfringens | L. monocytogenes * | RSA Requirements | |
|---|---|---|---|---|---|---|---|---|
| FSOp-A | ||||||||
| Sampling Visit 1 | ||||||||
| Frozen veg mix: Corn, carrot, and string beans | 4.7 × 1036 | 10 | ND | P | APC: M = 5 × 105—Enterobacteriaceae: M = 5 × 104—Salmonella: m = 0 | |||
| Sampling Visit 2 | ||||||||
| Salad: Potato salad | 4.4 × 106 | <3 | ND | <10 | ND | APC: M = 106—E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0 | ||
| Sampling Visit 3 | ||||||||
| Frozen broccoli | >106 | 4.9 × 105 | ND | ND | APC: M = 5 × 105—Enterobacteriaceae: M = 5 × 104—Salmonella: m = 0 | |||
| Frozen avocado purée | 1.2 × 106 | 10 | <3 | ND | ND | APC: M = 5 × 105—Enterobacteriaceae: M = 5 × 105 -E. coli: M 102—Salmonella: m = 0 | ||
| Salad: Celery | >106 | <3 | ND | <10 | ND | APC: M = 106—E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0 | ||
| FSOp-B | ||||||||
| Sampling Visit 1 | ||||||||
| Salad: Cabbage and carrot mix | 5 × 106 | <3 | ND | <10 | P | APC: M = 106—E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0 | ||
| Sampling Visit 2 | ||||||||
| Salad: Cabbage and carrot mix | 1.1 × 106 | <3 | ND | <10 | ND | APC: M = 106—E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0 | ||
| Salad: Boiled eggs and lettuce | 1 × 106 | <3 | ND | <10 | ND | APC: M = 106—E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0 | ||
| Salad: Boiled string bean | 2 × 106 | <3 | ND | <10 | ND | APC: M = 106—E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0 | ||
| Salad: Tomato and cilantro | 5.4 × 105 | <3 | ND | <10 | ND | APC: M = 106—E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0 | ||
| Salad: Beef and mix vegetables | 1.1 × 106 | <3 | ND | <10 | <10 | ND | APC: M = 106— E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0— C. perfringens: M = 5 × 102. | |
| Salad: Cucumber | 2.2 × 107 | <3 | ND | <10 | ND | APC: M = 106—E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0 | ||
| Sampling Visit 3 | ||||||||
| Dessert: Spanish custard (RTE) | 5 × 106 | <3 | ND | <10 | ND | APC: M = 106—E. coli: M = 5 × 102—S. aureus: M = 5 × 102—Salmonella: m = 0 | ||
| FSOpA | FSOpB | |||||
|---|---|---|---|---|---|---|
| Zone 1 | Zone 2 | Zone 3 | Zone 1 | Zone 2 | Zone 3 | |
| Total samples | 14 | 5 | 40 | 18 | 17 | 50 |
| APC 50–499 CFU/cm2 | 1 | 1 | NA | 2 | 1 | NA |
| APC >500 CFU/cm2 | 5 | 0 | NA | 3 | 2 | NA |
| L. monocytogenes | ND | ND | 4 | ND | ND | 14 |
| FSOp | Predictor Variable | Estimate (Odds Ratio) | Pr (>[z]) & |
|---|---|---|---|
| A | Log APC | 1.499 | 0.168 |
| B | Log APC | 2.488 | 0.013 * |
| QA Resistant Related Genes | Biofilm-Related Genes * | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| FSOp | Visit | Strain | CFSAN Number | SRA | Source | Lineage | MLST | Serogroup | MIC-QA (ppm) | bcrA | bcrB/bcrC | qacH | qacC | lmo0673/lmo2504 | luxS/recO | lmo2026/lmo0435 |
| A | 1 | A17/612-3 | CFSAN104436 | SRR12957137 | Frozen veg mi × 1 | I | ST2/CC2 | IVb | 5 | + | + | + | ||||
| 1 | A17/589-1 | CFSAN104437 | SRR12957136 | Drain | I | ST1/CC1 | IVb | 5 | + | + | + | |||||
| 1 | A17/606-3 | CFSAN104435 | SRR12957145 | Drain | I | ST2/CC2 | IVb | 5 | + | + | + | |||||
| B | 1 | A17/646 | CFSAN104423 | SRR12957144 | RTE Salad2 | I | ST2349 */CC5 | IIb | 5 | + | + | + | + | |||
| 1 | A17/661-8 | CFSAN104428 | SRR12957143 | Drain | I | ST5/CC5 | IIb | 10 | + | + | + | + | ||||
| 1 | A17/677-4 | CFSAN104427 | SRR12957142 | Drain | I | ST2/CC2 | IVb | 10 | + | + | + | + | ||||
| 2 | A17/931-3 | CFSAN104430 | SRR12957141 | Drain | I | ST5/CC5 | IIb | 10 | + | + | + | + | ||||
| 2 | A17/932-3 | CFSAN104431 | SRR12957140 | Drain | I | ST2/CC2 | IVb | 10 | + | + | + | + | ||||
| 2 | A17/939-4 | CFSAN104424 | SRR12957139 | Boot cover | I | ST2/CC2 | IVb | 10 | + | + | + | + | ||||
| 2 | A17/941-4 | CFSAN104432 | SRR12957138 | Drain | I | ST5/CC5 | IIb | 10 | + | + | + | + | ||||
| 2 | A17/942-5 | CFSAN104433 | SRR12957135 | Drain | I | ST2/CC2 | IVb | 10 | + | + | + | + | ||||
| 2 | A17/943-8 | CFSAN104429 | SRR12957134 | Drain | I | ST2/CC2 | IVb | 10 | + | + | + | + | ||||
| 3 | A18/062-1 | CFSAN104419 | SRR12957151 | Floor | I | ST5/CC5 | IIb | 10 | + | + | + | + | ||||
| 3 | A18/070-7 | CFSAN104420 | SRR12957150 | Drain | I | ST2/CC2 | IVb | 5 | + | + | + | |||||
| 3 | A18/075-3 | CFSAN104421 | SRR12957149 | Floor | I | ST5/CC5 | IIb | 10 | + | + | + | + | ||||
| 3 | A18/076-1 | CFSAN104422 | SRR12957148 | Drain | I | ST5/CC5 | IIb | 10 | + | + | + | + | ||||
| 3 | A18/079-3 | CFSAN104426 | SRR12957147 | Drain | I | ST2/CC2 | IVb | 10 | + | + | + | + | ||||
| 3 | A18/081-5 | CFSAN104425 | SRR12957146 | Drain | I | ST2/CC2 | IVb | 10 | + | + | + | + | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Toro, M.; Williams-Vergara, J.; Solar, C.; Quesille-Villalobos, A.M.; Kwon, H.J.; Navarrete, P.; Meng, J.; Chen, Y.; Reyes-Jara, A. Evaluation of the Persistence and Characterization of Listeria monocytogenes in Foodservice Operations. Foods 2022, 11, 886. https://doi.org/10.3390/foods11060886
Toro M, Williams-Vergara J, Solar C, Quesille-Villalobos AM, Kwon HJ, Navarrete P, Meng J, Chen Y, Reyes-Jara A. Evaluation of the Persistence and Characterization of Listeria monocytogenes in Foodservice Operations. Foods. 2022; 11(6):886. https://doi.org/10.3390/foods11060886
Chicago/Turabian StyleToro, Magaly, Jessica Williams-Vergara, Camila Solar, Ana María Quesille-Villalobos, Hee Jin Kwon, Paola Navarrete, Jianghong Meng, Yi Chen, and Angélica Reyes-Jara. 2022. "Evaluation of the Persistence and Characterization of Listeria monocytogenes in Foodservice Operations" Foods 11, no. 6: 886. https://doi.org/10.3390/foods11060886
APA StyleToro, M., Williams-Vergara, J., Solar, C., Quesille-Villalobos, A. M., Kwon, H. J., Navarrete, P., Meng, J., Chen, Y., & Reyes-Jara, A. (2022). Evaluation of the Persistence and Characterization of Listeria monocytogenes in Foodservice Operations. Foods, 11(6), 886. https://doi.org/10.3390/foods11060886








