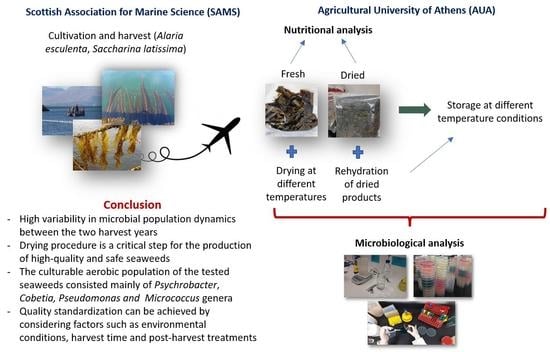
Quality and Safety Assessment of Edible Seaweeds Alaria esculenta and Saccharina latissima Cultivated in Scotland
Abstract
:1. Introduction
2. Materials and Methods
2.1. Seaweed Collection and Treatments
2.2. Determination of Nutritional Parameters (Protein, Fat, Fatty Acid Profile, Carbohydrates, Moisture and Ash)
2.3. Microbiological Analysis
2.4. Identification of Bacterial Species
2.5. Statistical Analysis
3. Results and Discussion
3.1. Nutritional Analysis
3.2. Microbial Profile of Fresh Seaweeds
3.3. Microbiological Quality of Dried Products
3.4. Rehydrated Products
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ferdouse, F.; Holdt, S.L.; Smith, R.; Murúa, P.; Yang, Z.; FAO Consultants. The Global Status of Seaweed Production, Trade and Utilization; FAO Globefish Research Programme: Rome, Italy, 2018; Volume 124. [Google Scholar]
- Delaney, A.; Frangoudes, K.; Li, S.A. Society and seaweed: Understanding the past and present. In Seaweed in Health and Disease Prevention; Levine, I., Fleurence, J., Eds.; Academic Press: Cambridge, MA, USA, 2016; pp. 7–40. [Google Scholar]
- Mouritsen, O.G.; Rhatigan, P.; Pérez-Lloréns, J.L. The rise of seaweed gastronomy: Phycogastronomy. Bot. Mar. 2018, 62, 195–209. [Google Scholar] [CrossRef]
- Rioux, L.-E.; Beaulieu, L.; Turgeon, S.L. Seaweeds: A traditional ingredients for new gastronomic sensation. Food Hydrocoll. 2017, 68, 255–265. [Google Scholar] [CrossRef]
- Perry, J.J.; Brodt, A.; Skonberg, D.I. Influence of dry salting on quality attributes of farmed kelp (Alaria esculenta) during long-term refrigerated storage. LWT 2019, 114, 108362. [Google Scholar] [CrossRef]
- Stévant, P.; Marfaing, H.; Rustad, T.; Sandbakken, I.; Fleurence, J.; Chapman, A. Nutritional value of the kelps Alaria esculenta and Saccharina latissima and effects of short-term storage on biomass quality. Environ. Boil. Fishes 2017, 29, 2417–2426. [Google Scholar] [CrossRef]
- Handå, A.; Forbord, S.; Wang, X.; Broch, O.J.; Dahle, S.W.; Størseth, T.R.; Reitan, K.I.; Olsen, Y.; Skjermo, J. Seasonal and depth-dependent growth of cultivated kelp (Saccharina latissima) in close proximity to salmon (Salmo salar) aquaculture in Norway. Aquaculture 2013, 414–415, 191–201. [Google Scholar] [CrossRef]
- Kraan, S.; Tramullas, A.V.; Guiry, M.D. The edible brown seaweed Alaria esculenta (Phaeophyceae, Laminariales): Hybridization, growth and genetic comparisons of six Irish populations. Environ. Boil. Fishes 2000, 12, 577–583. [Google Scholar] [CrossRef]
- Neto, R.T.; Marçal, C.; Queirós, A.S.; Abreu, H.; Silva, A.M.S.; Cardoso, S.M. Screening of Ulva rigida, Gracilaria sp., Fucus vesiculosus and Saccharina latissima as functional ingredients. Int. J. Mol. Sci. 2018, 19, 2987. [Google Scholar] [CrossRef] [Green Version]
- Stévant, P.; Indergård, E.; Ólafsdóttir, A.; Marfaing, H.; Larssen, W.E.; Fleurence, J.; Roleda, M.; Rustad, T.; Slizyte, R.; Nordtvedt, T. Effects of drying on the nutrient content and physico-chemical and sensory characteristics of the edible kelp Saccharina latissima. Environ. Boil. Fishes 2018, 30, 2587–2599. [Google Scholar] [CrossRef]
- Blikra, M.J.; Wang, X.; James, P.; Skipnes, D. Saccharina latissima cultivated in northern Norway: Reduction of potentially toxic elements during processing in relation to cultivation depth. Foods 2021, 10, 1290. [Google Scholar] [CrossRef]
- López-Pérez, O.; Picon, A.; Nuñez, M. Volatile compounds and odour characteristics of seven species of dehydrated edible seaweeds. Food Res. Int. 2017, 99, 1002–1010. [Google Scholar] [CrossRef]
- Banach, J.L.; Hil, E.H.; Van Der Fels-Klerx, H.J. Food safety hazards in the European seaweed chain. Compr. Rev. Food Sci. Food Saf. 2020, 19, 332–364. [Google Scholar] [CrossRef]
- del Olmo, A.; Picon, A.; Nuñez, M. Preservation of five edible seaweeds by high pressure processing: Effect on microbiota, shelf life, colour, texture and antioxidant capacity. Algal Res. 2020, 49, 101938. [Google Scholar] [CrossRef]
- Singh, R.; Reddy, C. Seaweed-microbial interactions: Key functions of seaweed-associated bacteria. FEMS Microbiol. Ecol. 2014, 88, 213–230. [Google Scholar] [CrossRef]
- Egan, S.; Harder, T.; Burke, C.; Steinberg, P.; Kjelleberg, S.; Thomas, T. The seaweed holobiont: Understanding seaweed-bacteria interactions. FEMS Microbiol. Rev. 2013, 37, 462–476. [Google Scholar] [CrossRef] [Green Version]
- Blikra, M.; Løvdal, T.; Vaka, M.R.; Roiha, I.S.; Lunestad, B.T.; Lindseth, C.; Skipnes, D. Assessment of food quality and microbial safety of brown macroalgae (Alaria esculenta and Saccharina latissima). J. Sci. Food Agric. 2019, 99, 1198–1206. [Google Scholar] [CrossRef]
- Nayyar, D.; Skonberg, D.I. Contrasting effects of two storage temperatures on the microbial, physicochemical, and sensory properties of two fresh red seaweeds, Palmaria palmata and Gracilaria tikvahiae. Environ. Boil. Fishes 2018, 31, 731–739. [Google Scholar] [CrossRef]
- Sánchez-García, F.; Hernández, I.; Palacios, V.M.; Roldán, A.M. Freshness quality and shelf-life evaluation of the seaweed Ulva rigida through physical, chemical, microbiological, and sensory methods. Foods 2021, 10, 181. [Google Scholar] [CrossRef]
- Picon, A.; Del Olmo, A.; Nuñez, M. Bacterial diversity in six species of fresh edible seaweeds submitted to high pressure processing and long-term refrigerated storage. Food Microbiol. 2020, 94, 103646. [Google Scholar] [CrossRef]
- ISO. Food and Feed Products—General Guidelines for the Determination of Nitrogen by the Kjeldahl Method (1871:2009); International Organization for Standardization: Geneva, Switzerland, 2009. [Google Scholar]
- Stévant, P.; Ólafsdóttir, A.; Déléris, P.; Dumay, J.; Fleurence, J.; Ingadóttir, B.; Jónsdóttir, R.; Ragueneau, E.; Rebours, C.; Rustad, T. Semi-dry storage as a maturation process for improving the sensory characteristics of the edible red seaweed dulse (Palmaria palmata). Algal Res. 2020, 51, 102048. [Google Scholar] [CrossRef]
- Angell, A.R.; Mata, L.; de Nys, R.; Paul, N. The protein content of seaweeds: A universal nitrogen-to-protein conversion factor of five. Environ. Boil. Fishes 2015, 28, 511–524. [Google Scholar] [CrossRef]
- ISO. Meat and Meat Products—Determination of Total Fat Content (1443:1973); International Organization for Standardization: Geneva, Switzerland, 2016. [Google Scholar]
- ISO. Meat and Meat Products—Determination of Free Fat Content (1444:1996); International Organization for Standardization: Geneva, Switzerland, 2018. [Google Scholar]
- ISO. Animal and Vegetable Fats and Oils—Gas Chromatography of Fatty Acid Methyl Esters—Part 2: Preparation of Methyl Esters of Fatty Acids (12966-2:2017); International Organization for Standardization: Geneva, Switzerland, 2017. [Google Scholar]
- Nielsen, S.S. Food analysis. In Food Science Text Series, 4th ed.; Springer: Midtown Manhattan, NY, USA, 2010; p. 602. [Google Scholar]
- Wei, S.; Zhao, H.; Xian, Y.; Hussain, M.A.; Wu, X. Multiplex PCR assays for the detection of Vibrio alginolyticus, Vibrio parahaemolyticus, Vibrio vulnificus, and Vibrio cholerae with an internal amplification control. Diagn. Microbiol. Infect. Dis. 2014, 79, 115–118. [Google Scholar] [CrossRef]
- Doulgeraki, A.I.; Paramithiotis, S.; Nychas, G.-J. Characterization of the Enterobacteriaceae Community that developed during storage of minced beef under aerobic or modified atmosphere packaging conditions. Int. J. Food Microbiol. 2011, 145, 77–83. [Google Scholar] [CrossRef]
- Tzamourani, A.P.; Di Napoli, E.; Paramithiotis, S.; Oikonomou-Petrovits, G.; Panagiotidis, S.; Panagou, E.Z. Microbiological and physicochemical characterisation of green table olives of Halkidiki and Conservolea varieties processed by the Spanish method on industrial scale. Int. J. Food Sci. Technol. 2021, 56, 3845–3857. [Google Scholar] [CrossRef]
- Schiener, P.; Black, K.D.; Stanley, M.S.; Green, D. The seasonal variation in the chemical composition of the kelp species Laminaria digitata, Laminaria hyperborea, Saccharina latissima and Alaria esculenta. Environ. Boil. Fishes 2014, 27, 363–373. [Google Scholar] [CrossRef]
- Mæhre, H.K.; Malde, M.K.; Eilertsen, K.E.; Elvevoll, E. Characterization of protein, lipid and mineral contents in common Norwegian seaweeds and evaluation of their potential as food and feed. J. Sci. Food Agri. 2014, 94, 3281–3290. [Google Scholar] [CrossRef]
- Holdt, S.L.; Kraan, S. Bioactive compounds in seaweed: Functional food applications and legislation. Environ. Boil. Fishes 2011, 23, 543–597. [Google Scholar] [CrossRef]
- López-López, I.; Bastida, S.; Ruiz-Capillas, C.; Bravo, L.; Larrea, M.; Sánchez-Muniz, F.; Cofrades, S.; Jiménez-Colmenero, F. Composition and antioxidant capacity of low-salt meat emulsion model systems containing edible seaweeds. Meat Sci. 2009, 83, 492–498. [Google Scholar] [CrossRef]
- Rocha, C.; Pacheco, D.; Cotas, J.; Marques, J.; Pereira, L.; Gonçalves, A. Seaweeds as valuable sources of essential fatty acids for human nutrition. Int. J. Environ. Res. Public Health 2021, 18, 4968. [Google Scholar] [CrossRef]
- Biohaz, E.P.; Ricci, A.; Allende, A.; Bolton, D.; Chemaly, M.; Davies, R.; Escámez, P.S.F.; Girones, R.; Herman, L.; Koutsoumanis, K.; et al. Guidance on the requirements for the development of microbiological criteria. EFSA J. 2017, 15. [Google Scholar] [CrossRef] [Green Version]
- Kerrison, P.D.; Innes, M.; Macleod, A.; McCormick, E.; Elbourne, P.D.; Stanley, M.S.; Hughes, A.D.; Kelly, M.S. Comparing the effectiveness of twine and binder-seeding in the Laminariales species Alaria esculenta and Saccharina latissima. Environ. Boil. Fishes 2020, 32, 2173–2181. [Google Scholar] [CrossRef] [Green Version]
- Lachnit, T.; Blümel, M.; Imhoff, J.F.; Wahl, M. Specific epibacterial communities on macroalgae: Phylogeny matters more than habitat. Aquat. Biol. 2009, 5, 181–186. [Google Scholar] [CrossRef]
- Bondoso, J.; Balagué, V.; Gasol, J.M.; Lage, O. Community composition of the Planctomycetes associated with different macroalgae. FEMS Microbiol. Ecol. 2014, 88, 445–456. [Google Scholar] [CrossRef] [Green Version]
- Gupta, S.; Cox, S.; Abu-Ghannam, N. Effect of different drying temperatures on the moisture and phytochemical constituents of edible Irish brown seaweed. LWT 2011, 44, 1266–1272. [Google Scholar] [CrossRef] [Green Version]
- Del Olmo, A.; Picon, A.; Nuñez, M. The microbiota of eight species of dehydrated edible seaweeds from North West Spain. Food Microbiol. 2018, 70, 224–231. [Google Scholar] [CrossRef]
- Ozturkoglu-Budak, S.; Wiebenga, A.; Bron, P.A.; de Vries, R.P. Protease and lipase activities of fungal and bacterial strains derived from an artisanal raw ewe’s milk cheese. Int. J. Food Microbiol. 2016, 237, 17–27. [Google Scholar] [CrossRef]
- Antunes-Rohling, A.; Calero, S.; Halaihel, N.; Marquina, P.; Raso, J.; Calanche, J.; Beltrán, J.A.; Álvarez, I.; Cebrián, G. Characterization of the spoilage microbiota of Hake Fillets packaged under a modified atmosphere (MAP) rich in CO2 (50% CO2/50% N2) and stored at different temperatures. Foods 2019, 8, 489. [Google Scholar] [CrossRef] [Green Version]
- Broekaert, K.; Noseda, B.; Heyndrickx, M.; Vlaemynck, G.; Devlieghere, F. Volatile compounds associated with Psychrobacter spp. and Pseudoalteromonas spp., the dominant microbiota of brown shrimp (Crangon crangon) during aerobic storage. Int. J. Food Microbiol. 2013, 166, 487–493. [Google Scholar] [CrossRef]
- Albakosh, M.A.; Naidoo, R.K.; Kirby, B.; Bauer, R. Identification of epiphytic bacterial communities associated with the brown alga Splachnidium rugosum. Environ. Boil. Fishes 2015, 28, 1891–1901. [Google Scholar] [CrossRef]
- Arahal, D.R.; Castillo, A.M.; Ludwig, W.; Schleifer, K.H.; Ventosa, A. Proposal of Cobetia marina gen. nov., comb. nov., within the family Halomonadaceae, to include the species Halomonas marina. Syst. Appl. Microbiol. 2002, 25, 207–211. [Google Scholar] [CrossRef]
- Martin, M.; Barbeyron, T.; Martin, R.; Portetelle, D.; Michel, G.; Vandenbol, M. The cultivable surface microbiota of the brown alga Ascophyllum nodosum is enriched in macroalgal-polysaccharide-degrading bacteria. Front. Microbiol. 2015, 6, 1487. [Google Scholar] [CrossRef]
- Mulaosmanovic, E.; Lindblom, T.; Windstam, S.; Bengtsson, M.; Rosberg, A.; Mogren, L.; Alsanius, B. Processing of leafy vegetables matters: Damage and microbial community structure from field to bag. Food Control 2021, 125, 107894. [Google Scholar] [CrossRef]
- Tatsika, S.; Karamanoli, K.; Karayanni, H.; Genitsaris, S. Metagenomic characterization of bacterial communities on ready-to-eat vegetables and effects of household washing on their diversity and composition. Pathogens 2019, 8, 37. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Söderqvist, K.; Osman, O.A.; Wolff, C.; Bertilsson, S.; Vågsholm, I.; Boqvist, S. Emerging microbiota during cold storage and temperature abuse of ready-to-eat salad. Infect. Ecol. Epidemiol. 2017, 7, 1328963. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Del Olmo, A.; Picon, A.; Nuñez, M. High pressure processing for the extension of Laminaria ochroleuca (kombu) shelf-life: A comparative study with seaweed salting and freezing. Innov. Food Sci. Emerg. Technol. 2019, 52, 420–428. [Google Scholar] [CrossRef]
- Odeyemi, O.A.; Alegbeleye, O.O.; Strateva, M.; Stratev, D. Understanding spoilage microbial community and spoilage mechanisms in foods of animal origin. Compr. Rev. Food Sci. Food Saf. 2020, 19, 311–331. [Google Scholar] [CrossRef] [Green Version]
- Fu, K.; Li, J.; Wang, Y.; Liu, J.; Yan, H.; Shi, L.; Zhou, L. An innovative method for rapid identification and detection of vibrio alginolyticus in different infection models. Front. Microbiol. 2016, 7, 651. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, Y.; Zhao, P.; Chen, L.; Wu, H.; Si, X.; Shen, X.; Shen, H.; Qiao, Y.; Zhu, S.; Chen, Q.; et al. Fast, simple and highly specific molecular detection of Vibrio alginolyticus pathogenic strains using a visualized isothermal amplification method. BMC Veter. Res. 2020, 16, 76. [Google Scholar] [CrossRef] [PubMed]
- Harrysson, H.; Krook, J.L.; Larsson, K.; Tullberg, C.; Oerbekke, A.; Toth, G.; Pavia, H.; Undeland, I. Effect of storage conditions on lipid oxidation, nutrient loss and colour of dried seaweeds, Porphyra umbilicalis and Ulva fenestrata, subjected to different pretreatments. Algal Res. 2021, 56, 102295. [Google Scholar] [CrossRef]
- Santiago, A.; Moreira, R. Drying of edible seaweeds. In Sustainable Seaweed Technologies; Torres, M.D., Kraan, S., Dominguez, H., Eds.; Elsevier: Amsterdam, The Netherlands, 2020; pp. 131–154. [Google Scholar]
- Badmus, U.O.; Taggart, M.; Boyd, K.G. The effect of different drying methods on certain nutritionally important chemical constituents in edible brown seaweeds. Environ. Boil. Fishes 2019, 31, 3883–3897. [Google Scholar] [CrossRef] [Green Version]
- Tello-Ireland, C.; Lemus-Mondaca, R.; Vega-Galvez, A.; López, J.; Di Scala, K. Influence of hot-air temperature on drying kinetics, functional properties, colour, phycobiliproteins, antioxidant capacity, texture and agar yield of alga Gracilaria chilensis. LWT 2011, 44, 2112–2118. [Google Scholar] [CrossRef]
- Gupta, S.; Rajauria, G.; Abu-Ghannam, N. Study of the microbial diversity and antimicrobial properties of Irish edible brown seaweeds. Int. J. Food Sci. Technol. 2010, 45, 482–489. [Google Scholar] [CrossRef] [Green Version]
- Cox, S.; Gupta, S.; Abu-Ghannam, N. Effect of different rehydration temperatures on the moisture, content of phenolic compounds, antioxidant capacity and textural properties of edible Irish brown seaweed. LWT 2012, 47, 300–307. [Google Scholar] [CrossRef]





| A | Alaria esculenta | |||||
| Harvest year | 2019 | 2020 | ||||
| Form | Fresh/Frozen | Dried | Rehydrated | Fresh/Frozen | Dried | Rehydrated |
| Storage temperatures (°C) | 5, 15 | 25 | 5, 10 | 0, 5, 10, 15 | 25 | 5, 10 |
| Storage time | 7 days | 6 months | 5 days | 7 days | 6 months | 5 days |
| Microbiological analysis | 7 time points | 2 time points | 6 time points | 9 time points | 2 time points | 6 time points |
| Number of replicates | 4 * | 4 | 4 | 4 | 4 | 4 |
| Nutritional analysis | Day 0 | Day 0 | - | Day 0 | Day 0 | - |
| Number of replicates | 3 | 3 | - | 3 | 3 | - |
| B | Saccharina latissima | |||||
| Harvest year | 2019 | 2020 | ||||
| Form | Fresh/Frozen | Dried | Rehydrated | Fresh/Frozen | Dried | Rehydrated |
| Storage temperatures (°C) | 5, 15 | 25 | 5, 10 | 0, 5, 10, 15 | 25 | 5, 10 |
| Storage time | 7 days | 6 months | 5 days | 60 (0 °C), 25 (5 °C, 13 days (at 10 and 15 °C) | 6 months | 5 days |
| Microbiological analysis | 7 time points | 2 time points | 6 time points | 8 time points (0, 5 °C), 9 time points (10, 15 °C) | 2 time points | 6 time points |
| Number of replicates | 4 | 4 | 4 | 4 | 4 | 4 |
| Nutritional analysis | Day 0 | Day 0 | - | Day 0 | Day 0 | - |
| Number of replicates | 3 | 3 | - | 3 | 3 | - |
| g/100 g | A. esculenta | S. latissima | ||||||
|---|---|---|---|---|---|---|---|---|
| 2019 | 2020 | 2019 | 2020 | |||||
| Fresh | Dried | Fresh | Dried | Fresh | Dried | Fresh | Dried | |
| Protein | 2.44 ± 0.10 a * | 11.30 ± 0.06 A * | 1.99 ± 0.08 a * | 9.13 ± 0.35 A * | 1.40 ± 0.14 a * | 9.93 ± 0.14 A * | 0.87 ± 0.01 b * | 7.44 ± 0.12 B * |
| Fat | 0.21 ± 0.02 a * | 0.82 ± 0.10 A * | 0.13 ± 0.08 a | 0.60 ± 0.09 A | 0.11 ± 0.06 a * | 0.55 ± 0.08 A * | 0.08 ± 0.01 a | 0.73 ± 0.10 A |
| Content (%) | ||||||||
| Saturated | - | 49.53 ± 1.80 | - | 73.97 ± 1.00 | - | 62.72 ± 1.30 | - | 79.91 ± 1.00 |
| MUFA | - | 26.67 ± 1.50 | - | 13.60 ± 0.80 | - | 24.44 ± 1.00 | - | 8.74 ± 0.10 |
| PUFA | - | 23.79 ± 0.96 | - | 12.43 ± 0.94 | - | 12.85 ± 0.91 | - | 11.35 ± 0.70 |
| Carbohydrates | 14.64 ± 1.00 a * | 50.72 ± 1.00 A * | 14.41 ± 0.90 a * | 53.28 ± 1.20 A * | 6.93 ± 1.00 a * | 42.16 ± 1.42 A * | 6.91 ± 0.91 a * | 43.97 ± 1.00 B * |
| Moisture | 78.83 ± 1.20 a * | 11.16 ± 0.74 A | 80.55 ± 1.30 a * | 10.64 ± 0.01 A | 88.30 ± 1.60 a * | 9.70 ± 0.95 A | 89.40 ± 0.50 a * | 8.92 ± 0.50 A |
| Ash | 3.88 ± 0.64 a | 26.00 ± 1.40 A * | 2.92 ± 0.18 a | 26.34 ± 0.78 A * | 3.26 ± 0.98 a | 37.66 ± 1.30 A * | 2.74 ± 0.17 a | 38.94 ± 1.90 A * |
| NaCl | 2.00 ± 0.40 a | 15.00 ± 0.58 A * | 1.80 ± 0.10 a | 16.00 ± 1.00 A * | 2.00 ± 0.64 a | 28.00 ± 0.50 A * | 1.85 ± 0.10 a | 24.00 ± 0.80 B * |
| Isolate Code | Temperature °C | Days of Storage | TVC (Log CFU/g) | Closest Relative Microorganism | GenBank Accession Number of Closest Relative | % Similarity |
|---|---|---|---|---|---|---|
| SAE18 | 5 | 3 | 5.6 | Cobetia crustatorum | NR_116500.1 | 99.13 |
| SAE19 | Psychrobacter fozii | NR_025531.1 | 98.30 | |||
| SAE20 | Pseudoalteromonas tetraodonis GFC | NR_119142.1 | 98.08 | |||
| SAE37 | 5 | 5 | 7.7 | Psychrobacter fozii | NR_025531.1 | 99.83 |
| SAE40 | Cobetia crustatorum | NR_116500.1 | 99.31 | |||
| SAE60 | 5 | 7 | 8.0 | Corynebacterium tapiri | NR_145582.1 | 95.81 |
| SAE61 | Lelliottia amnigena | NR_024642.1 | 99.83 | |||
| SAE62 | Jeotgalicoccus psychrophilus | NR_025644.1 | 99.14 | |||
| SAE67 | Cobetia litoralis | NR_113403.1 | 86.68 | |||
| SAE69 | Psychrobacter cryohalolentis K5 | NR_075055.1 | 99.30 | |||
| SAE70 | Psychrobacter fozii | NR_025531.1 | 100.0 | |||
| SAE72 | Cobetia crustatorum | NR_116500.1 | 99.13 | |||
| SAE03 | 15 | 1 | 6.0 | Psychrobacter piscatorii | NR_112807.1 | 100.0 |
| SAE09 | Psychrobacter fozii | NR_025531.1 | 100.0 | |||
| SAE10 | Cobetia crustatorum | NR_116500.1 | 99.13 | |||
| SAE22 | 15 | 3 | 9.5 | Cobetia crustatorum | NR_116500.1 | 99.30 |
| SAE23 | Pseudomonas psychrophila | NR_028619.1 | 99.65 | |||
| SAE36 | Pseudomonas weihenstephanensis | NR_148764.1 | 99.65 | |||
| SAE50 | 15 | 5 | 10.0 | Lelliottia amnigena | NR_024642.1 | 99.83 |
| SAE51 | Pseudomonas psychrophila | NR_028619.1 | 99.65 | |||
| SAE59 | Pseudomonas monteilii | NR_112073.1 | 99.48 | |||
| SAE75 | 15 | 7 | 9.5 | Psychrobacter adeliensis | NR_117632.1 | 93.82 |
| SAE76 | Psychrobacter fozii | NR_025531.1 | 100.0 | |||
| SAE85 | Pseudomonas weihenstephanensis | NR_148764.1 | 99.65 |
| Isolate Code | Temperature °C | Days of Storage | TVC (Log CFU/g) | Closest Relative Microorganism | GenBank Accession Number of Closest Relative | % Similarity |
|---|---|---|---|---|---|---|
| SSL01 | 5 | 0 | 1.5 | Mesobacillus subterraneus | MT515815.1 | 99.67 |
| SSL03 | Micrococcus luteus | NR_037113.1 | 99.48 | |||
| SSL06 | Staphylococcus hominis | NR_036956.1 | 94.55 | |||
| SSL10 | Staphylococcus epidermidis | NR_036904.1 | 99.83 | |||
| SSL11 | 5 | 2 | 2.2 | Micrococcus aloeverae | NR_134088.1 | 99.10 |
| SSL12 | Acinetobacteriwoffii | NR_113346.1 | 100.0 | |||
| SSL14 | Roseomonas aestuarii | NR_114285.1 | 100.0 | |||
| SSL18 | 5 | 3 | 2.0 | Micrococcus luteus | NR_037113.1 | 99.82 |
| SSL43 | 5 | 13 | 2.2 | Alkalihalobacillus hwajinpoensis | NR_025264.1 | 96.14 |
| SSL47 | Bacillus cereus | R_115526.1 | 100.0 | |||
| SSL24 | 15 | 7 | 2.0 | Lelliottia amnigena | NR_024642.1 | 99.83 |
| SSL26 | Psychrobacter pacificensis | NR_027187.1 | 100.0 | |||
| SSL30 | 15 | 8 | 4.0 | Psychrobacter fozii | NR_025531.1 | 99.65 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lytou, A.E.; Schoina, E.; Liu, Y.; Michalek, K.; Stanley, M.S.; Panagou, E.Z.; Nychas, G.-J.E. Quality and Safety Assessment of Edible Seaweeds Alaria esculenta and Saccharina latissima Cultivated in Scotland. Foods 2021, 10, 2210. https://doi.org/10.3390/foods10092210
Lytou AE, Schoina E, Liu Y, Michalek K, Stanley MS, Panagou EZ, Nychas G-JE. Quality and Safety Assessment of Edible Seaweeds Alaria esculenta and Saccharina latissima Cultivated in Scotland. Foods. 2021; 10(9):2210. https://doi.org/10.3390/foods10092210
Chicago/Turabian StyleLytou, Anastasia E., Eirini Schoina, Yunge Liu, Kati Michalek, Michele S. Stanley, Efstathios Z. Panagou, and George-John E. Nychas. 2021. "Quality and Safety Assessment of Edible Seaweeds Alaria esculenta and Saccharina latissima Cultivated in Scotland" Foods 10, no. 9: 2210. https://doi.org/10.3390/foods10092210
APA StyleLytou, A. E., Schoina, E., Liu, Y., Michalek, K., Stanley, M. S., Panagou, E. Z., & Nychas, G.-J. E. (2021). Quality and Safety Assessment of Edible Seaweeds Alaria esculenta and Saccharina latissima Cultivated in Scotland. Foods, 10(9), 2210. https://doi.org/10.3390/foods10092210