DNA Analysis Detects Different Mislabeling Trend by Country in European Cod Fillets
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sampling and Labelling
2.2. DNA Extraction
2.3. PCR and DNA Sequencing
- FISHCO1LBC: 5′-TCAACYAAT CAYAAAGATATYGGCAC-3′
- FISHCO1HBC: 5′-ACTTCYGGGTGRCCR AARAATCA-3′
2.4. Data Analysis
2.5. Statistical Analysis
3. Results
3.1. DNA Analysis
3.2. Catch Location and Catch Method
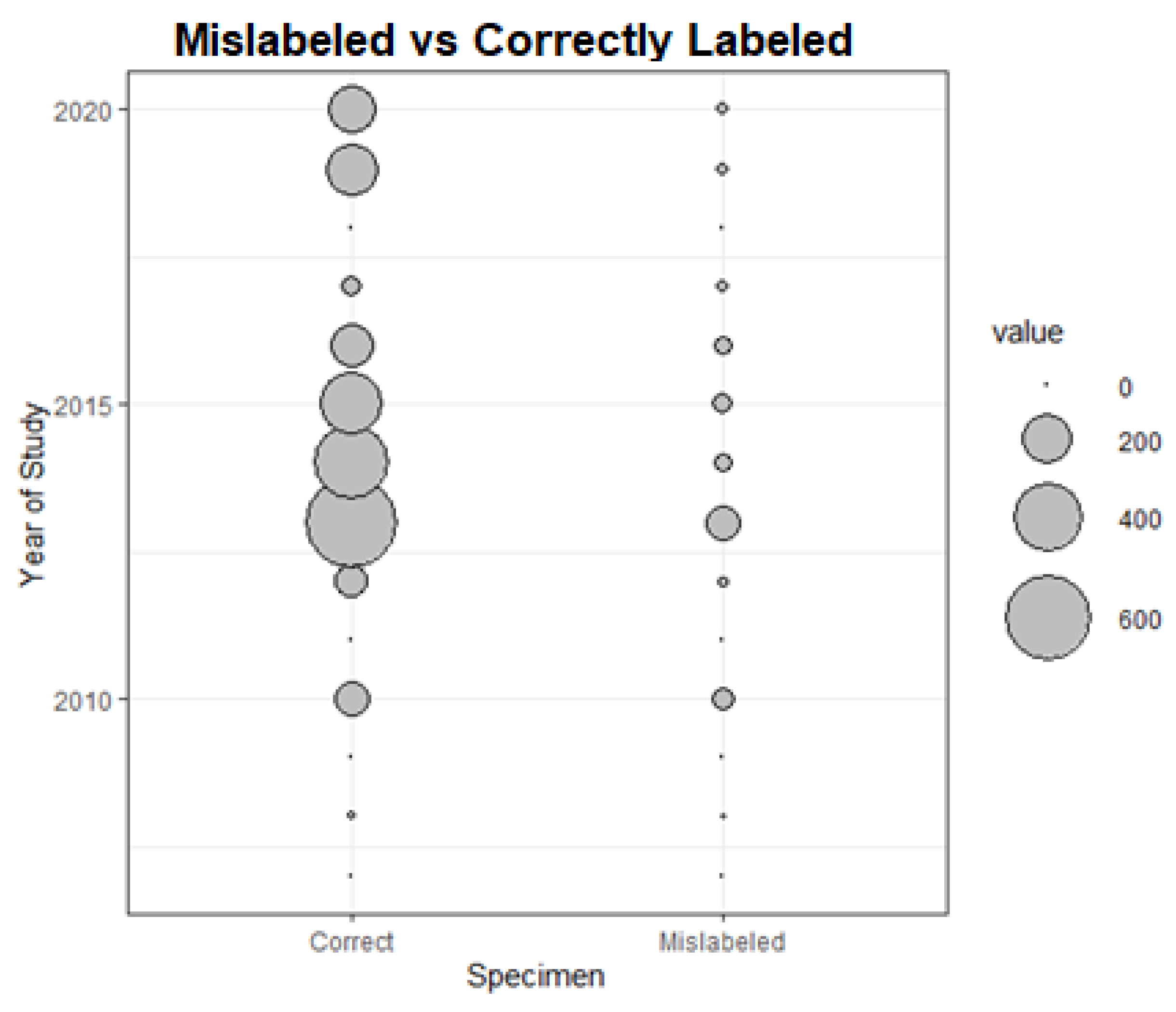
3.2.1. Proportion of Mislabeling over Time
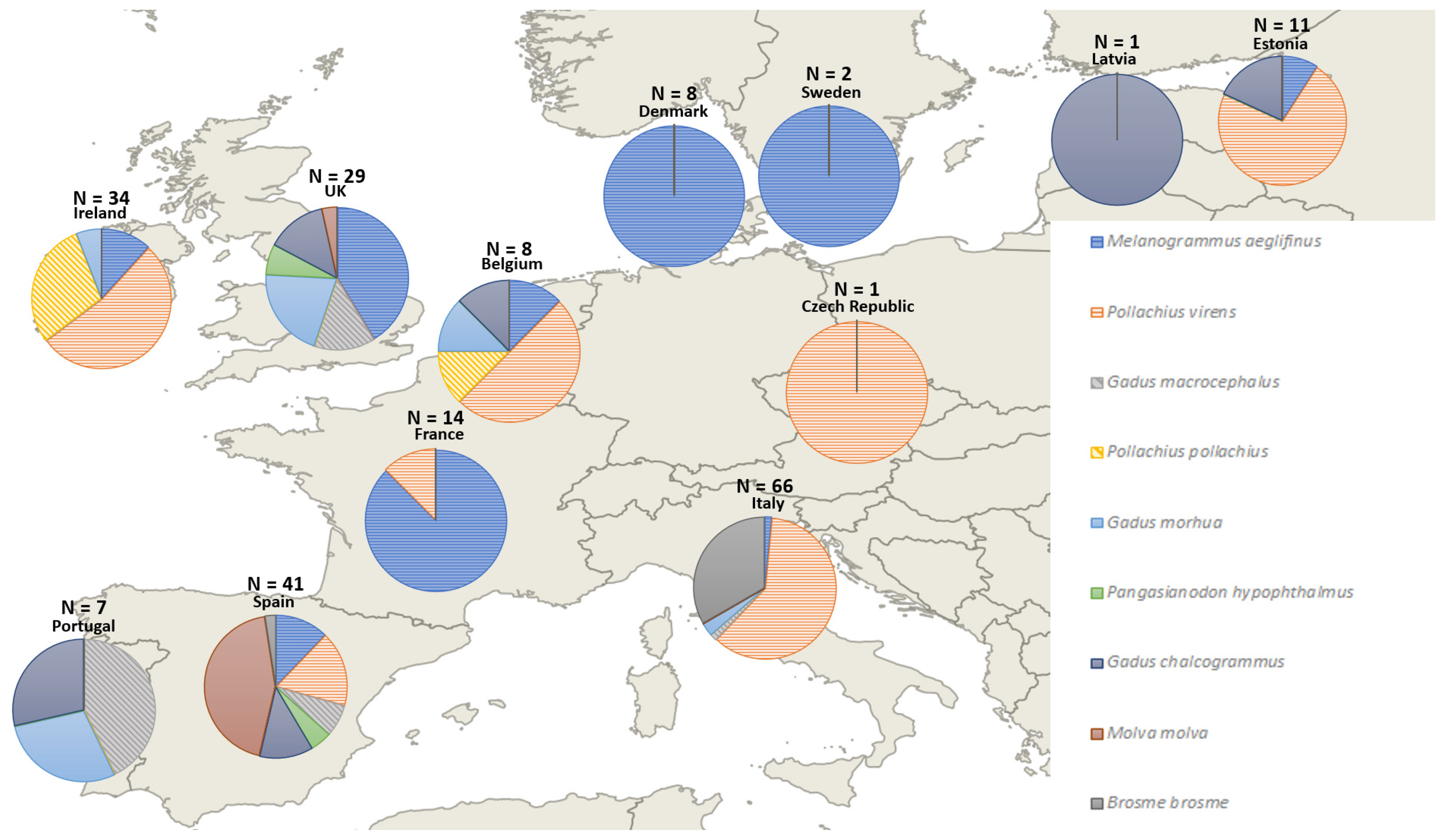
3.2.2. Substitute Species
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Publication | Sequence Accession Number |
|---|---|
| Filonzi et al., 2010 [38] | GenBank: EU7521, DQ173995, EU752090, EF211102 |
| Miller and Mariani, 2010 [32] | not provided |
| Miller, Jessel, and Mariani, 2012 [28] | not provided |
| Pinto et al., 2013 [29] | not provided |
| Helgoe et al., 2020 [30] | not provided |
| Cutarelli et al., 2013 [39] | not provided |
| Benard-Capelle et al. 2015 [33] | Barcode of Life: FCSF001-14 to FCSF291-14; FCSF292-14 to FCSF404-14 |
| Harris et al. 2016 [40] | not provided |
| Mariani et al., 2015 [32] | GenBank: KJ510424–KJ531384; KJ563141–KJ645864 |
| Helyar et al., 2014 [41] | GenBank: KJ614671--KJ615069 |
| Brechon et al., 2016 [25] | not provided |
| Muñoz-Colmenero et al., 2016 [42] | GenBank: KF597025–KF597028; KF597030; KF597040–KF597046; KF597048; KP330299–KP330373; KP330374–KP330387 |
| Christiansen et al., 2018 [43] | not provided |
| Pardo and Jimenez, 2020 [44] | GenBank: MT266987-MT267272 |
| Pardo et al., 2018 [11] | not provided |
| Minoudi et al., 2020 [45] | not provided |
| Barendse et al., 2019 [46] | not provided |
| Deconinck et al., 2020 [24] | not provided |
References
- Galal-Khallaf, A.; Ardura, A.; Mohammed-Geba, K.; Borrell, Y.J.; Garcia-Vazquez, E. DNA Barcoding Reveals a High Level of Mislabelling in Egyptian Fish Fillets. Food Control 2014, 46, 441–445. [Google Scholar] [CrossRef]
- Khaksar, R.; Carlson, T.; Schaffner, D.W.; Ghorashi, M.; Best, D.; Jandhyala, S.; Traverso, J.; Amini, S. Unmasking Seafood Mislabelling in U.S. Markets: DNA Barcoding as a Unique Technology for Food Authentication and Quality Control. Food Control 2015, 56, 71–76. [Google Scholar] [CrossRef]
- Carvalho, D.C.; Guedes, D.; da Gloria Trindade, M.; Coelho, R.M.S.; de Lima Araujo, P.H. Nationwide Brazilian Governmental Forensic Programme Reveals Seafood Mislabelling Trends and Rates Using DNA Barcoding. Fish. Res. 2017, 191, 30–35. [Google Scholar] [CrossRef]
- Xiong, X.; Guardone, L.; Cornax, M.J.; Tinacci, L.; Guidi, A.; Gianfaldoni, D.; Armani, A. DNA barcoding reveals substitution of sablefish (Anoplopoma fimbria) with Patagonian and Antarctic toothfish (Dissostichus eleginoides and Dissostichus mawsoni) in online market in China: How mislabeling opens door to IUU fishing. Food Control 2016, 70, 380–391. [Google Scholar] [CrossRef]
- Mariani, S.; Griffiths, A.M.; Velasco, A.; Kappel, K.; Jerome, M.; Perez-Martin, R.I.; Schroder, U.; Verrez-Bagnis, V.; Silva, H.; Vandamme, S.G.; et al. Low Mislabelling Rates Indicate Marked Improvements in European Seafood Market Operations. Front. Ecol. Environ. 2015, 13, 536–540. [Google Scholar] [CrossRef]
- PDF4PRO. Available online: https://pdf4pro.com/view/the-state-of-world-fisheries-and-aquaculture-187a.html (accessed on 11 March 2021).
- European Commission. Available online: https://ec.europa.eu/fisheries/press/eu-fish-market-2019-edition-out-everything-you-wanted-know-about-eu-market-fish-and-seafood_en (accessed on 11 March 2021).
- EUMOFA. Available online: https://www.eumofa.eu/market-analysis#yearly (accessed on 11 March 2021).
- European Commission. Available online: https://ec.europa.eu/fisheries/eumofa-monthly-highlights-12018_en (accessed on 11 March 2021).
- Olsgard, F.; Schaanning, M.T.; Widdicombe, S.; Kendall, M.A.; Austen, M.C. Effects of Bottom Trawling on Ecosystem Functioning. J. Exp. Mar. Biol. Ecol. 2008, 366, 123–133. [Google Scholar] [CrossRef]
- Pardo, M.Á.; Jiménez, E.; Viðarsson, J.R.; Ólafsson, K.; Ólafsdóttir, G.; Daníelsdóttir, A.K.; Pérez-Villareal, B. DNA Barcoding Revealing Mislabelling of Seafood in European Mass Caterings. Food Control 2018, 92, 7–16. [Google Scholar] [CrossRef]
- Pardo, M.Á.; Jiménez, E.; Pérez-Villarreal, B. Misdescription Incidents in Seafood Sector. Food Control 2016, 62, 277–283. [Google Scholar] [CrossRef]
- Kroetz, K.; Luque, G.M.; Gephart, J.A.; Jardine, S.L.; Lee, P.; Moore, K.C.; Cole, C.; Steinkruger, A.; Josh Donlan, C. Consequences of Seafood Mislabelling for Marine Populations and Fisheries Management. Proc. Natl. Acad. Sci. USA 2020, 117, 30318–30323. [Google Scholar] [CrossRef]
- Triantafyllidis, A.; Karaiskou, N.; Perez, J.; Martinez, J.L.; Roca, A.; Lopez, B.; Garcia-Vazquez, E. Fish Allergy Risk Derived from Ambiguous Vernacular Fish Names: Forensic DNA-Based Detection in Greek Markets. Food Res. Int. 2010, 43, 2214–2216. [Google Scholar] [CrossRef]
- Barcaccia, G.; Lucchin, M.; Cassandro, M. DNA Barcoding as a Molecular Tool to Track down Mislabelling and Food Piracy. Diversity 2016, 8, 2. [Google Scholar] [CrossRef] [Green Version]
- Kochzius, M.; Seidel, C.; Antoniou, A.; Botla, S.K.; Campo, D.; Cariani, A.; Vazquez, E.G.; Hauschild, J.; Hervet, C.; Hjörleifsdottir, S.; et al. Identifying Fishes through DNA Barcodes and Microarrays. PLoS ONE 2010, 5, e12620. [Google Scholar] [CrossRef] [Green Version]
- Huxley-Jones, E.; Shaw, J.L.A.; Fletcher, C.; Parnell, J.; Watts, P.C. Use of DNA Barcoding to Reveal Species Composition of Convenience Seafood. Conserv. Biol. 2012, 26, 367–371. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; de Waard, J.R. Biological identifications through DNA barcodes. Proc. Natl. Acad. Sci. USA 2003, 270, 313–321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ward, R.; Hanner, R.; Hebert, P. The campaign to DNA barcode all fishes, FISH-BOL. J. Fish Biol. 2009, 74, 329–356. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Ratnasingham, S.; DeWaard, J.R. Barcoding Animal Life: Cytochrome c Oxidase Subunit 1 Divergences among Closely Related Species. Proc. R. Soc. B Biol. Sci. 2003, 270 (Suppl. 1), 96–99. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naaum, A.M.; Warner, K.; Mariani, S.; Hanner, R.H.; Carolin, C.D. Seafood Mislabelling Incidence and Impacts; Elsevier Inc.: Amsterdam, The Netherlands, 2016. [Google Scholar] [CrossRef]
- Standal, D.; Bouwer Utne, I. Can Cod Farming Affect Cod Fishing? A System Evaluation of Sustainability. Mar. Policy 2007, 31, 527–534. [Google Scholar] [CrossRef]
- Halldórsson, J.E.; Björnsson, B.; Gunnlaugsson, S.B. Feasibility of Ranching Coastal Cod (Gadus morhua) Compared with on-Growing, Full-Cycle Farming and Fishing. Mar. Policy 2012, 36, 11–17. [Google Scholar] [CrossRef]
- Deconinck, D.; Volckaert, F.A.M.; Hostens, K.; Panicz, R.; Eljasik, P.; Faria, M.; Monteiro, C.S.; Robbens, J.; Derycke, S. A High-Quality Genetic Reference Database for European Commercial Fishes Reveals Substitution Fraud of Processed Atlantic Cod (Gadus morhua) and Common Sole (Solea solea) at Different Steps in the Belgian Supply Chain. Food Chem. Toxicol. 2020, 141, 111417. [Google Scholar] [CrossRef]
- Bréchon, A.L.; Hanner, R.; Mariani, S. A Systematic Analysis across North Atlantic Countries Unveils Subtleties in Cod Product Labelling. Mar. Policy 2016, 69, 124–133. [Google Scholar] [CrossRef]
- Kappel, K.; Schröder, U. Substitution of High-Priced Fish with Low-Priced Species: Adulteration of Common Sole in German Restaurants. Food Control 2016, 59, 478–486. [Google Scholar] [CrossRef]
- Günther, B.; Raupach, M.J.; Knebelsberger, T. Full-length and mini-length DNA barcoding for the identification of seafood commercially traded in Germany. Food Control 2017, 73, 922–929. [Google Scholar] [CrossRef]
- Miller, D.; Jessel, A.; Mariani, S. Seafood Mislabelling: Comparisons of Two Western European Case Studies Assist in Defining Influencing Factors, Mechanisms and Motives. Fish Fish. 2012, 13, 345–358. [Google Scholar] [CrossRef]
- Di Pinto, A.; Di Pinto, P.; Terio, V.; Bozzo, G.; Bonerba, E.; Ceci, E.; Tantillo, G. DNA Barcoding for Detecting Market Substitution in Salted Cod Fillets and Battered Cod Chunks. Food Chem. 2013, 141, 1757–1762. [Google Scholar] [CrossRef]
- Bénard-Capelle, J.; Guillonneau, V.; Nouvian, C.; Fournier, N.; Le Loët, K.; Dettai, A. Fish mislabelling in France: Substitution rates and retail types. PeerJ 2015, 2, e714. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Helgoe, J.; Oswald, K.J.; Quattro, J.M. A Comprehensive Analysis of the Mislabelling of Atlantic Cod (Gadus morhua) Products in Spain. Fish. Res. 2020, 222, 105400. [Google Scholar] [CrossRef]
- Miller, D.D.; Mariani, S. Smoke, Mirrors, and Mislabelled Cod: Poor Transparency in the European Seafood Industry. Front. Ecol. Environ. 2010, 8, 517–521. [Google Scholar] [CrossRef]
- Estoup, A.; Largiader, C.R.; Perrot, E.; Chourrout, D. Rapid one-tube DNA extraction for reliable PCR dectection of fish polymorphic markers and transgenes. Mol. Mar. Biol. Biotechnol. 1996, 5, 295–298. [Google Scholar]
- Handy, S.M.; Deeds, J.R.; Ivanova, N.V.; Hebert, P.D.; Hanner, R.H.; Ormos, A.; Yancy, H.F. A single-laboratory validated method for the generation of DNA barcodes for the identification of fish for regulatory compliance. J. AOAC Int. 2011, 94, 201–210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ward, R.D.; Zemlak, T.S.; Innes, B.H.; Last, P.R.; Hebert, P.D. DNA barcoding Australia’s fish species. Philos. Trans. R. Soc. B Biol. Sci. 2005, 360, 1847–1857. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In Nucleic Acids Symposium Series; Information Retrieval Ltd.: London, UK, 1999; Volume 41, pp. 95–98, c1979–c2000. [Google Scholar]
- Filonzi, L.; Chiesa, S.; Vaghi, M.; Nonnis Marzano, F. Molecular Barcoding Reveals Mislabelling of Commercial Fish Products in Italy. Food Res. Int. 2010, 43, 1383–1388. [Google Scholar] [CrossRef]
- Cutarelli, A.; Amoroso, M.G.; De Roma, A.; Girardi, S.; Galiero, G.; Guarino, A.; Corrado, F. Italian Market Fish Species Identification and Commercial Frauds Revealing by DNA Sequencing. Food Control 2014, 37, 46–50. [Google Scholar] [CrossRef]
- Harris, D.J.; Rosado, D.; Xavier, R. DNA Barcoding Reveals Extensive Mislabelling in Seafood Sold in Portuguese Supermarkets. J. Aquat. Food Prod. Technol. 2016, 25, 1375–1380. [Google Scholar] [CrossRef]
- Helyar, S.J.; Lloyd, H.D.; De Bruyn, M.; Leake, J.; Bennett, N.; Carvalho, G.R. Fish Product Mislabelling: Failings of Traceability in the Production Chain and Implications for Illegal, Unreported and Unregulated (IUU) Fishing. PLoS ONE 2014, 9, e98691. [Google Scholar] [CrossRef] [Green Version]
- Muñoz-Colmenero, M.; Blanco, O.; Arias, V.; Martinez, J.L.; Garcia-Vazquez, E. L’authentification ADN Des Produits Halieutiques Révèle Un Mauvais Étiquetage Associé Au Traitement Des Fruits de Mer. Fisheries 2016, 41, 128–138. [Google Scholar] [CrossRef]
- Christiansen, H.; Fournier, N.; Hellemans, B.; Volckaert, F.A.M. Seafood Substitution and Mislabelling in Brussels’ Restaurants and Canteens. Food Control 2018, 85, 66–75. [Google Scholar] [CrossRef]
- Pardo, M.Á.; Jiménez, E. DNA Barcoding Revealing Seafood Mislabelling in Food Services from Spain. J. Food Compos. Anal. 2020, 91, 103521. [Google Scholar] [CrossRef]
- Minoudi, S.; Karaiskou, N.; Avgeris, M.; Gkagkavouzis, K.; Tarantili, P.; Triantafyllidou, D.; Palilis, L.; Avramopoulou, V.; Tsikliras, A.; Barmperis, K.; et al. Seafood Mislabelling in Greek Market Using DNA Barcoding. Food Control 2020, 113, 107213. [Google Scholar] [CrossRef]
- Barendse, J.; Roel, A.; Longo, C.; Andriessen, L.; Webster, L.M.I.; Ogden, R.; Neat, F. DNA Barcoding Validates Species Labelling of Certified Seafood. Curr. Biol. 2019, 29, R198–R199. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mariani, S.; Ellis, J.; O’Reilly, A.; Bréchon, A.L.; Sacchi, C.; Miller, D.D. Mass Media Influence and the Regulation of Illegal Practices in the Seafood Market. Conserv. Lett. 2014, 7, 478–483. [Google Scholar] [CrossRef] [Green Version]
- Horreo, J.L.; Fitze, P.S.; Jiménez-Valverde, A.; Noriega, J.A.; Pelaez, M.L. Amplification of 16S RDNA Reveals Important Fish Mislabelling in Madrid Restaurants. Food Control 2019, 96, 146–150. [Google Scholar] [CrossRef]
- Gilman, E.; Legorburu, G.; Fedoruk, A.; Heberer, C.; Zimring, M.; Barkai, A. Increasing the Functionalities and Accuracy of Fisheries Electronic Monitoring Systems. Aquat. Conserv. Mar. Freshw. Ecosyst. 2019, 29, 901–926. [Google Scholar] [CrossRef]
- Vlachopoulou, E.I.; Wilson, A.M.; Miliou, A. Disconnects in EU and Greek Fishery Policies and Practices in the Eastern Aegean Sea and Impacts on Posidonia Oceanica Meadows. Ocean Coast. Manag. 2013, 76, 105–113. [Google Scholar] [CrossRef]
- Food and Agriculture Organization of the United Nations. Available online: http://www.fao.org/fishery/area/Area27/en (accessed on 11 March 2021).
- Food and Agriculture Organization of the United Nations. Available online: http://www.fao.org/tempref/FI/maps/fig_h4_67_0.gif (accessed on 11 March 2021).
- Pedersen, E.J.; Thompson, P.L.; Ball, R.A.; Fortin, M.J.; Gouhier, T.C.; Link, H.; Moritz, C.; Nenzen, H.; Stanley, R.R.E.; Taranu, Z.E.; et al. Signatures of the Collapse and Incipient Recovery of an Overexploited Marine Ecosystem. R. Soc. Open Sci. 2017, 4, 170215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Årthun, M.; Bogstad, B.; Daewel, U.; Keenlyside, N.S.; Sandø, A.B.; Schrum, C.; Ottersen, G. Climate Based Multi-Year Predictions of the Barents Sea Cod Stock. PLoS ONE 2018, 13, e0206319. [Google Scholar] [CrossRef]
- Guggisberg, S. The EU’s Regulation on the Sustainable Management of External Fishing Fleets: International and European Law Perspectives. Int. J. Mar. Coast. Law 2019, 34, 291–324. [Google Scholar] [CrossRef] [Green Version]
| Sample Number | Species in Label | Country | Number of Samples | City | Selling Point | Catch Method | Catch Location |
|---|---|---|---|---|---|---|---|
| CodFreGerHW | G. morhua | Germany | 4 | Hamburg | Fish market | trawl nets | Barents Sea (FAO 27) |
| CodFreGerHF | G. morhua | Germany | 10 | Hamburg | Fish market | trawl nets | Norwegian Sea (FAO 27) |
| CodFreGerBS | G. morhua | Germany | 10 | Bremerhaven | Fish market | trawl nets | Great Belt (FAO 27) |
| CodFreGerHAF | G. morhua | Germany | 2 | Hamburg | Fish market | N/A | N/A |
| CodFreGerFN | G. morhua | Germany | 2 | Fulda | Fish market chain | seine fishing | North Atlantic Ocean (FAO 27) |
| CodFreGerFE | G. morhua | Germany | 3 | Fulda | Supermarket | N/A | Norwegian Sea (FAO 27) |
| CodFreGerFT | G. morhua | Germany | 4 | Fulda | Supermarket | N/A | Iceland (FAO 27) |
| CodFroGerHL | G. morhua | Germany | 6 | Hamburg | Supermarket | various | FAO 27 |
| CodFroGerHA | G. morhua | Germany | 6 | Hamburg | Supermarket | bottom otter trawl | Spitzbergen, Barents Sea (FAO 27) |
| CodFroGerFT | G. morhua | Germany | 10 | Fulda | Supermarket | various | Barents Sea/Norwegian Sea/North East Atlantic (FAO 27) |
| CodFroGerFR | G. morhua | Germany | 8 | Fulda | Supermarket | seine fishing/long lines | Barents Sea/Norwegian Sea (FAO 27) |
| CodFroGerFK | G. morhua | Germany | 10 | Fulda | Supermarket | trawl nets/NA | North East Atlantic/(FAO 27) |
| CodFroGerFL | G. morhua/G. macrocephalus | Germany | 16 | Fulda | Supermarket | various | North East Atlantic (FAO 27)/North East Pacific (FAO 67) |
| CodFroGerBA | G. morhua | Germany | 2 | Bremerhaven | Supermarket | bottom otter trawl | Barents Sea (FAO 27) Norwegian Sea |
| CodFroGerFA | G. morhua | Germany | 8 | Fulda | Supermarket | trawl nets | FAO 27 |
| CodFreNetRVg | G. morhua | The Netherlands | 10 | Rotterdam | Fish market | N/A | North Sea (FAO 27) |
| CodFreNetBd | G. morhua | The Netherlands | 14 | Rotterdam | Fish market | N/A | Norway (FAO 27) |
| CodFreNetRN | G. morhua | The Netherlands | 1 | Rotterdam | Fish market | N/A | N/A |
| CodFreNetRVb | G. morhua | The Netherlands | 5 | Rotterdam | Fish market | N/A | North Sea (FAO 27) |
| CodFroNetRA | G. morhua | The Netherlands | 21 | Rotterdam | Supermarket | seine fishing/long lines | North East Atlantic, Barents Sea, North Sea (FAO 27) |
| CodFroNetRH | G. morhua | The Netherlands | 20 | Rotterdam | Supermarket | trawl nets, seine fishing/long lines | North East Atlantic, Barents Sea, North Sea (FAO 27) |
| CodFreFraMCf | G. morhua | France | 3 | Montpellier | Supermarket | hooks, long lines | Iceland, Faroe Islands (FAO 27) |
| CodFreFraMAu | G. morhua | France | 5 | Montpellier | Supermarket | trawls, hooks long lines | Faroe Islands, Barents Sea (FAO 27) |
| CodFreFraSCc | G. morhua | France | 7 | Sète | Fish market | trawls | FAO 27 |
| CodFreFraSCl | G. morhua | France | 8 | Sète | Fish market | trawls | FAO 27 |
| CodFreFraSPr | G. morhua | France | 5 | Sète | Fish market | trawls | FAO 27 |
| CodFreFraSCk | G. morhua | France | 3 | Sète | Fish market | trawls | FAO 27 |
| CodFroFraMCar | G. morhua | France | 23 | Montpellier | Supermarket | hooks, long lines, trawls | Barents Sea Faroe Islands FAO 27Iceland (FAO 27) |
| CodFroFraMSu | G. morhua | France | 12 | Montpellier | Supermarket | hooks, long lines, trawls | Barents Sea, Norwegian Sea, (FAO 27) |
| CodFroFraMAu | G. morhua | France | 14 | Montpellier | Supermarket | hooks, long lines, trawls | Barents Sea, Norwegian Sea, North Sea (FAO 27) |
| Year of Sampling | Correctly Labelled | Incorrectly Labelled | Sampling Country | Species on Label | Reference |
|---|---|---|---|---|---|
| 2008 | 4 | 2 | Italy | G. morhua | Filonzi et al. 2010 (Appendix A) [37] |
| 2010 | 94 | 37 | Ireland | Cod | Miller and Mariani 2010 (Appendix A) [32] |
| 2012 | 88 | 7 | UK | Cod | Miller, Jessel, and Mariani 2012 (Appendix A) [28] |
| 2013 | 43 | 62 | Italy | Cod | Pinto et al. 2013 (Appendix A) [29] |
| 2013 | 512 | 34 | Spain | G. morhua | Helgoe et al. 2020 (Appendix A) [31] |
| 2013 | 2 | 2 | Italy | Cod | Cutarelli et al. 2013 (Appendix A) [38] |
| 2013 | 136 | 6 | France | Cod | Benard-Capelle et al. 2015 (Appendix A) [30] |
| 2014 | 5 | 0 | Portugal | G. morhua/G. macrocephalus | Harris et al. 2016 (Appendix A) [39] |
| 2014 | 48 | 0 | France | Cod | Mariani et al. 2015 (Appendix A) [5] |
| 2014 | 42 | 0 | Germany | Cod | Mariani et al. 2015 (Appendix A) [5] |
| 2014 | 55 | 1 | Ireland | Cod | Mariani et al. 2015 (Appendix A) [5] |
| 2014 | 67 | 4 | Portugal | Cod, G. morhua/G. macrocephalus | Mariani et al. 2015 (Appendix A) [5] |
| 2014 | 118 | 4 | Spain | Cod | Mariani et al. 2015 (Appendix A) [5] |
| 2014 | 138 | 8 | UK | Cod | Mariani et al. 2015 (Appendix A) [5] |
| 2014 | 170 | 9 | UK | Cod | Helyar et al. 2014 (Appendix A) [40] |
| 2015 | 43 | 0 | Norway | Cod | Brechon et al. 2016 (Appendix A) [25] |
| 2015 | 44 | 0 | Netherlands | Cod | Brechon et al. 2016 (Appendix A) [25] |
| 2015 | 13 | 19 | Estonia | Cod | Brechon et al. 2016 (Appendix A) [25] |
| 2015 | 43 | 0 | Belgium | Cod | Brechon et al. 2016 (Appendix A) [25] |
| 2015 | 41 | 1 | UK | Cod | Brechon et al. 2016 (Appendix A) [25] |
| 2015 | 35 | 8 | Denmark | Cod | Brechon et al. 2016 (Appendix A) [25] |
| 2015 | 43 | 2 | Sweden | Cod | Brechon et al. 2016 (Appendix A) [25] |
| 2016 | 49 | 5 | Spain | Cod | Muñoz-Colmenero et al. 2016 (Appendix A) [41] |
| 2016 | 62 | 11 | Belgium | Cod | Christiansen et al. 2018 (Appendix A) [42] |
| 2016 | 41 | 6 | Spain | G. morhua | Pardo and Jimenez 2020 (Appendix A) [43] |
| 2017 | 7 | 3 | Portugal | G. Morhua/Cod | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 2 | 1 | France | Cod | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 1 | 1 | Czech Republic | Cod | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 1 | 1 | Greece | Cod | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 4 | 1 | UK | G. morhua/G. macrocephalus | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 0 | 1 | Finland | G. morhua/G. macrocephalus | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 1 | 1 | Latvia | G. morhua/G. macrocephalus | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 1 | 1 | Switzerland | G. morhua/G. macrocephalus | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 1 | 0 | Germany | Cod | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 2 | 0 | Belgium | Cod | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 3 | 0 | Norway | Cod | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 2 | 0 | Italy | Cod | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 1 | 0 | Netherlands | Cod | Pardo et al. 2018 (Appendix A) [11] |
| 2017 | 3 | 0 | Ireland | G. morhua | Pardo et al. 2018 (Appendix A) [11] |
| 2018 | 10 | 0 | Greece | Cod | Minoudi et al. 2020 (Appendix A) [44] |
| 2019 | 70 | 4 | UK, Belgium | G. macrocephalus/G.morhua | Barendse et al. 2019 (Appendix A) [45] |
| 2019 | 42 | 1 | UK | Cod | Barendse et al. 2019 (Appendix A) [45] |
| 2019 | 111 | 3 | Belgium | G. morhua | Deconinck et al. 2020 (Appendix A) [24] |
| 2020 | 71 | 0 | Netherlands | G. morhua | This study |
| 2020 | 101 | 0 | Germany | G. morhua/G. macrocephalus | This study |
| 2020 | 27 | 9 | France | G. morhua | This study |
| Substitute Species | IUCN Status | Years of Occurrence |
|---|---|---|
| Gadus morhua | Vulnerable | 2010, 2012, 2014, 2019 |
| G. macrocephalus | Not evaluated | 2013, 2014, 2019 |
| Lates niloticus | Least concern | 2013, 2016 |
| G. chalcogrammus | Near threatened | 2013, 2015, 2016, 2017, 2019 |
| Molva molva | Data deficient | 2013, 2014, 2016, 2017 |
| Pangasianodon hypophtalmus | Endangered | 2012, 2013, 2017 |
| Pollachius pollachius | Least concern | 2010, 2016 |
| P. virens | Least concern | 2010, 2013, 2015, 2016, 2017, 2019 |
| Melanogrammus aeglefinus | Vulnerable | 2010, 2012, 2013, 2014, 2015, 2019, 2020 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Feldmann, F.; Ardura, A.; Blanco-Fernandez, C.; Garcia-Vazquez, E. DNA Analysis Detects Different Mislabeling Trend by Country in European Cod Fillets. Foods 2021, 10, 1515. https://doi.org/10.3390/foods10071515
Feldmann F, Ardura A, Blanco-Fernandez C, Garcia-Vazquez E. DNA Analysis Detects Different Mislabeling Trend by Country in European Cod Fillets. Foods. 2021; 10(7):1515. https://doi.org/10.3390/foods10071515
Chicago/Turabian StyleFeldmann, Frederik, Alba Ardura, Carmen Blanco-Fernandez, and Eva Garcia-Vazquez. 2021. "DNA Analysis Detects Different Mislabeling Trend by Country in European Cod Fillets" Foods 10, no. 7: 1515. https://doi.org/10.3390/foods10071515
APA StyleFeldmann, F., Ardura, A., Blanco-Fernandez, C., & Garcia-Vazquez, E. (2021). DNA Analysis Detects Different Mislabeling Trend by Country in European Cod Fillets. Foods, 10(7), 1515. https://doi.org/10.3390/foods10071515






