A Review of Intrinsic Optical Imaging Serial Blockface Histology (ICI-SBH) for Whole Rodent Brain Imaging
Abstract
1. Introduction
2. Intrinsic Optical Contrast Imaging (ICI) of Brain Tissue
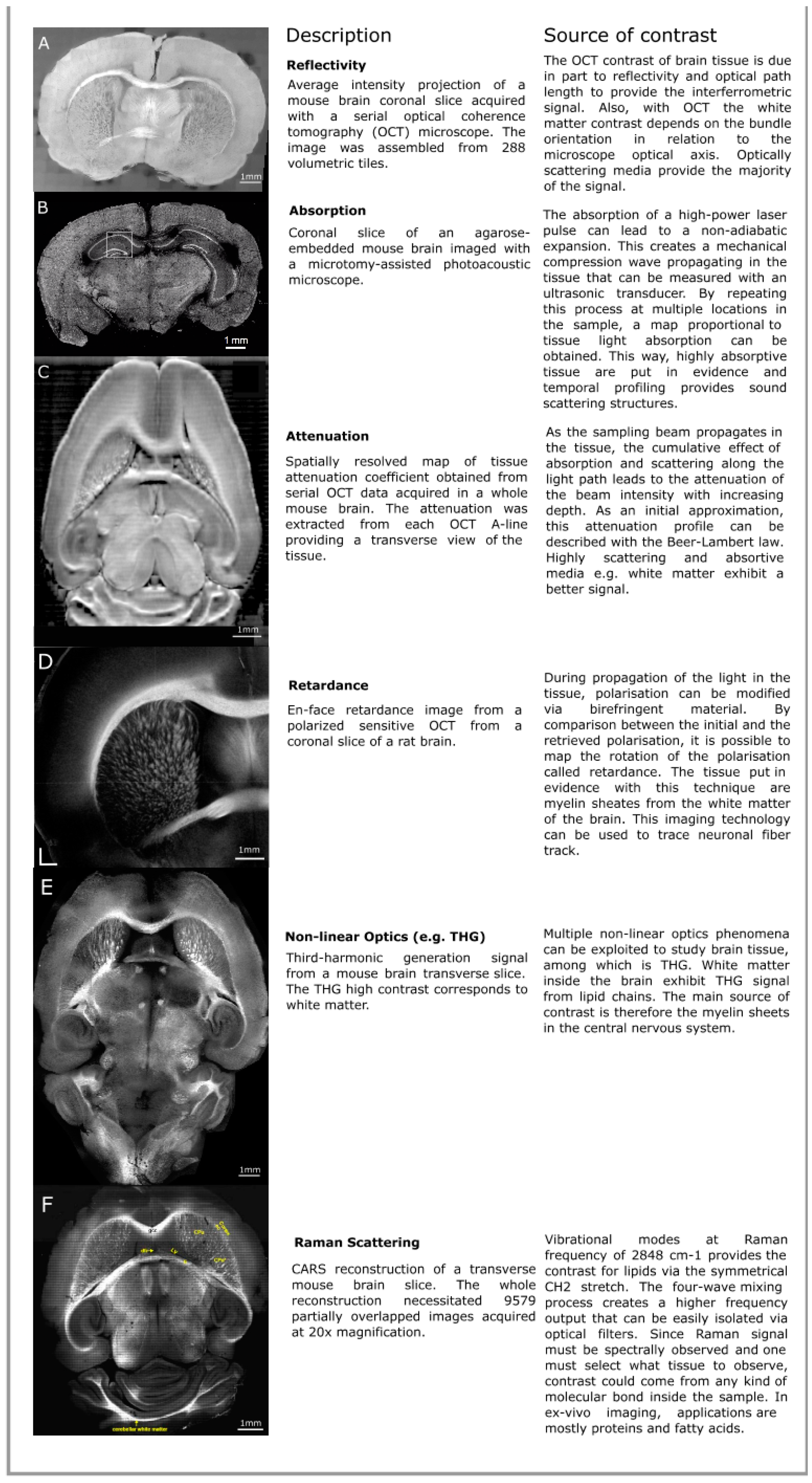
2.1. Reflectivity
2.2. Attenuation
2.3. Birefringence
2.4. Nonlinear Optical Processes
2.5. Raman Scattering
3. Serial Blockface Histology
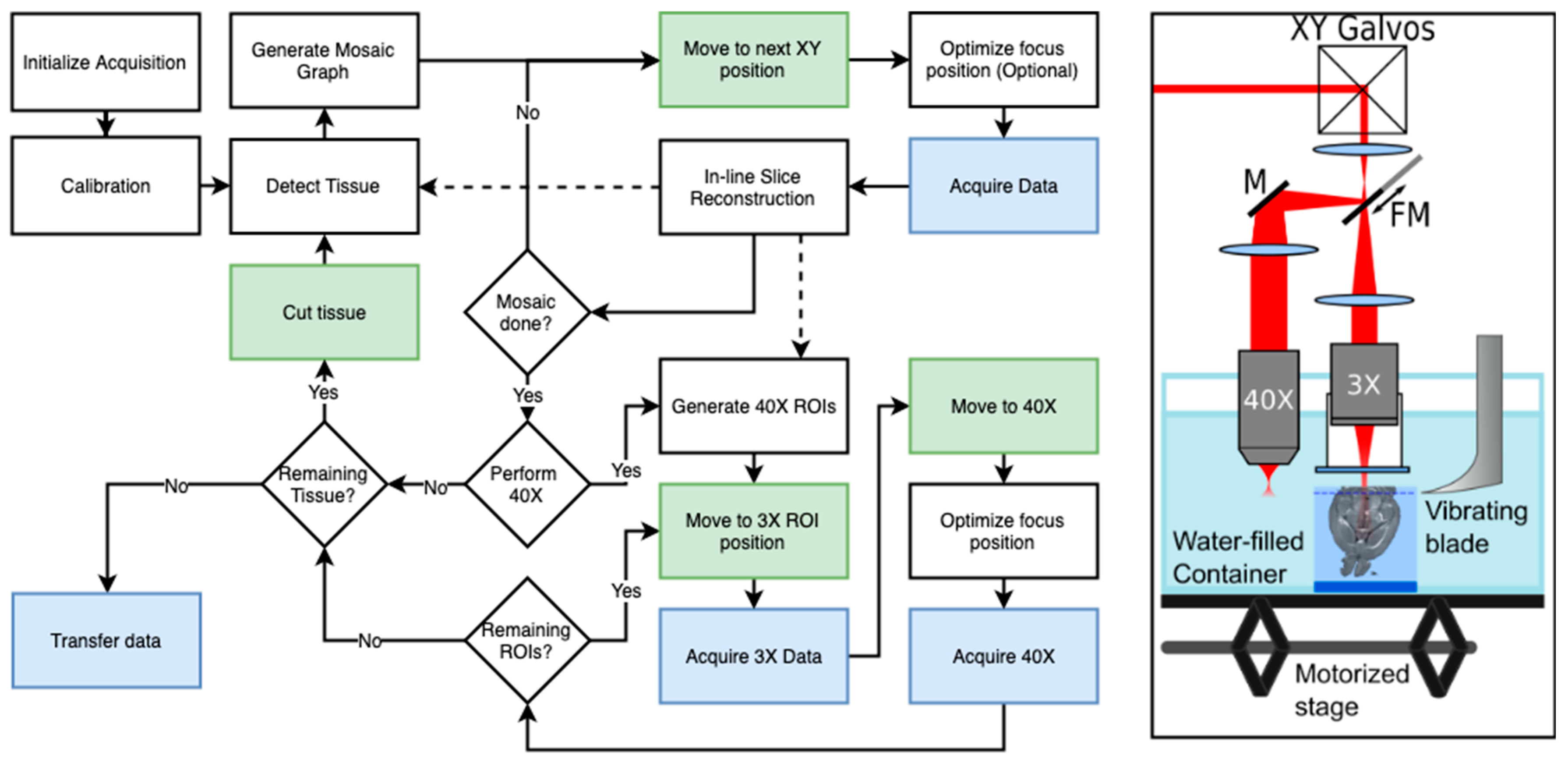
3.1. SBH Acquisition Automation
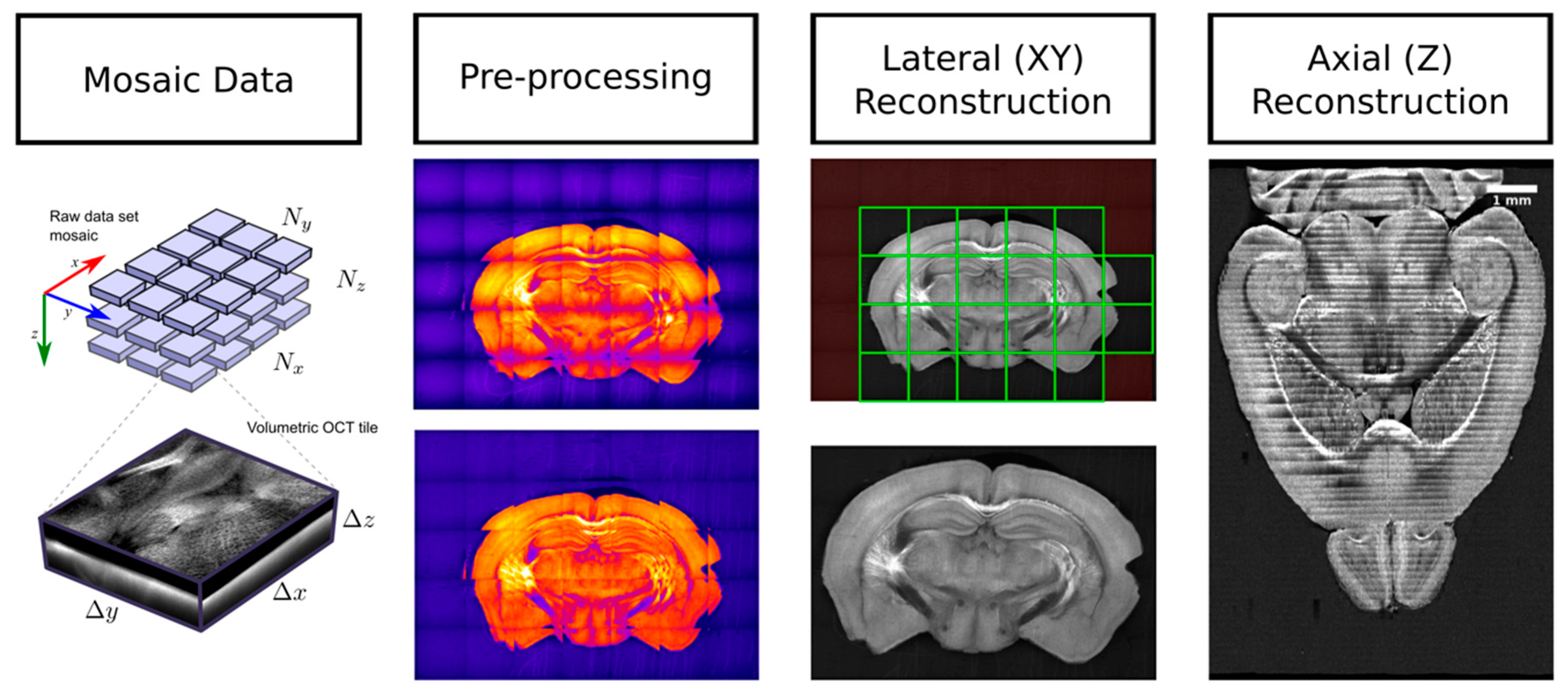
3.2. SBH Data Processing
3.2.1. SBH Data Preprocessing
3.2.2. Intra-Slice Registration and Stitching
3.2.3. Inter-Slice Registration and Stitching
3.3. Alternatives to SBH
4. Applications
4.1. Validation Studies
4.2. Multi-Modal Brain Atlases
5. Discussion and Conclusions
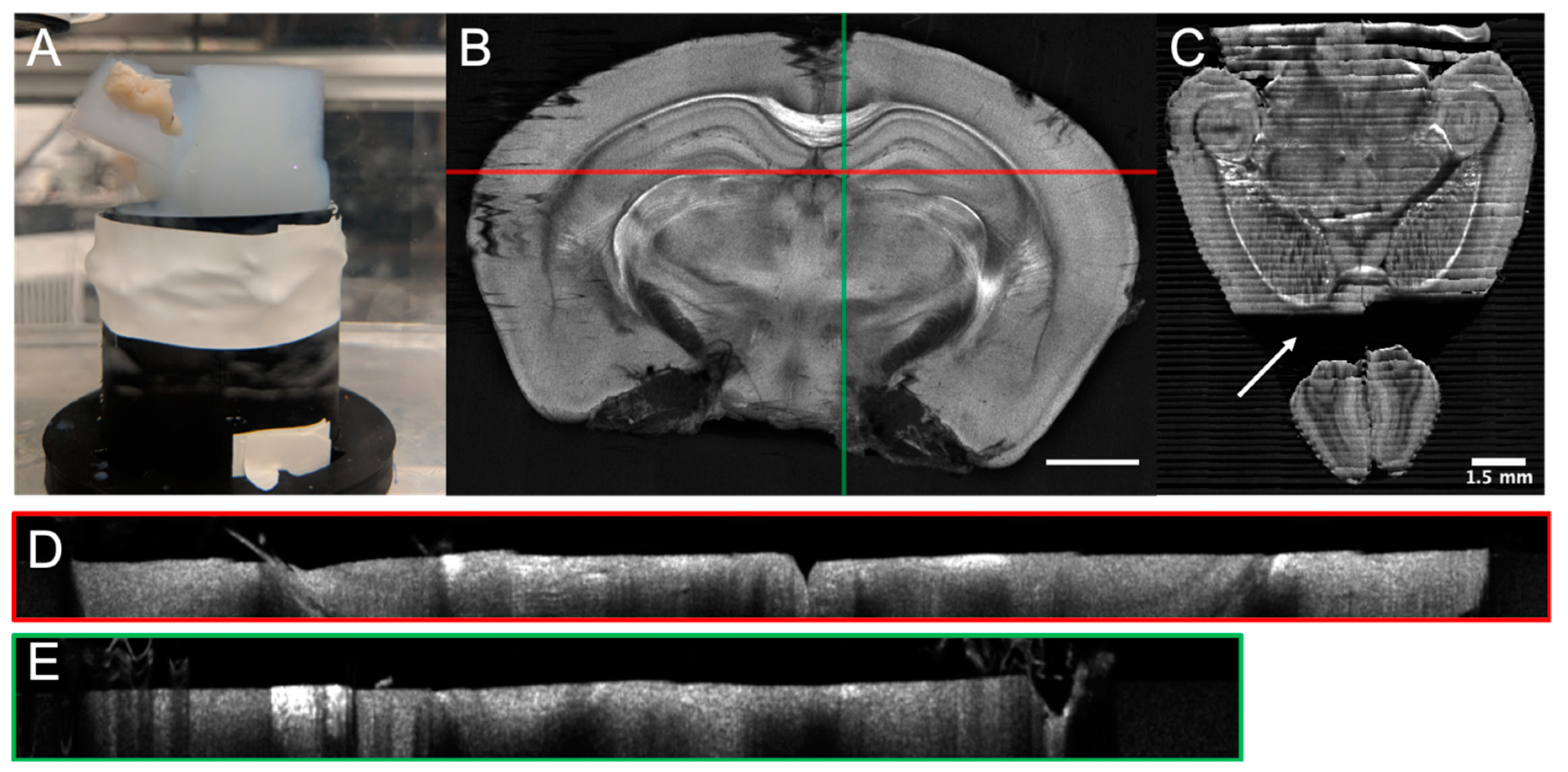
5.1. Tissue Preparation and Cutting Artefacts
5.2. Machine Learning and Serial Histology
5.3. Open-Source SBH Platform
Author Contributions
Funding
Conflicts of Interest
References
- Raichle, M.E. A Brief History of Human Brain Mapping. Trends Neurosci. 2009, 32, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Filippi, M. Oxford Textbook of Neuroimaging; Oxford University Press: New York, NY, USA, 2015. [Google Scholar] [CrossRef]
- Doronina-Amitonova, L.V.; Fedotov, I.V.; Fedotov, A.B.; Anokhin, K.V.; Zheltikov, A.M. Neurophotonics: Optical Methods to Study and Control the Brain. Uspekhi Fiz. Nauk 2015, 185, 371–392. [Google Scholar] [CrossRef]
- Cho, Y.K.; Zheng, G.; Augustine, G.J.; Hochbaum, D.; Cohen, A.; Knöpfel, T.; Pisanello, F.; Pavone, F.S.; Vellekoop, I.M.; Booth, M.J.; et al. Roadmap on Neurophotonics. J. Opt. 2016, 18, 093007. [Google Scholar] [CrossRef] [PubMed]
- Keiser, G. Biophotonics; Graduate Texts in Physics; Springer: Singapore, 2016. [Google Scholar] [CrossRef]
- Farkas, E.; Luiten, P.G.M. Cerebral Microvascular Pathology in Aging and Alzheimer’s Disease. Prog. Neurobiol. 2001, 64, 575–611. [Google Scholar] [CrossRef]
- De La Torre, J.C. Is Alzheimer’s Disease a Neurodegenerative or a Vascular Disorder? Data, Dogma, and Dialectics. Lancet Neurol. 2004, 3, 184–190. [Google Scholar] [CrossRef]
- de la Torre, J.C. Cerebrovascular and Cardiovascular Pathology in Alzheimer’s Disease. Int. Rev. Neurobiol. 2009, 84, 35–48. [Google Scholar] [PubMed]
- Grammas, P.; Martinez, J.; Miller, B. Cerebral Microvascular Endothelium and the Pathogenesis of Neurodegenerative Diseases. Expert Rev. Mol. Med. 2011, 13. [Google Scholar] [CrossRef] [PubMed]
- Tsai, P.S.; Friedman, B.; Ifarraguerri, A.I.; Thompson, B.D.; Lev-Ram, V.; Schaffer, C.B.; Xiong, Q.; Tsien, R.Y.; Squier, J.A.; Kleinfeld, D. All-Optical Histology Using Ultrashort Laser Pulses. Neuron 2003, 39, 27–41. [Google Scholar] [CrossRef]
- Sands, G.B.; Gerneke, D.A.; Hooks, D.A.; Green, C.R.; Smaill, B.H.; Legrice, I.J. Automated Imaging of Extended Tissue Volumes Using Confocal Microscopy. Microsc. Res. Tech. 2005, 67, 227–239. [Google Scholar] [CrossRef] [PubMed]
- Ragan, T.; Kadiri, L.R.; Venkataraju, K.U.; Bahlmann, K.; Sutin, J.; Taranda, J.; Arganda-Carreras, I.; Yongsoo, K.; Seung, H.S.; Osten, P. Serial two-photon tomography for automated ex vivo mouse brain imaging. Nat. Methods 2012, 9, 255–258. [Google Scholar] [CrossRef] [PubMed]
- Lein, E.S.; Hawrylycz, M.J.; Ao, N.; Ayres, M.; Bensinger, A.; Bernard, A.; Boe, A.F.; Boguski, M.S.; Brockway, K.S.; Byrnes, E.J.; et al. Genome-Wide Atlas of Gene Expression in the Adult Mouse Brain. Nature 2007, 445, 168–176. [Google Scholar] [CrossRef] [PubMed]
- Kleinfeld, D.; Bharioke, A.; Blinder, P.; Bock, D.D.; Briggman, K.L.; Chklovskii, D.B.; Denk, W.; Helmstaedter, M.; Kaufhold, J.P.; Lee, W.-C.A.; et al. Large-Scale Automated Histology in the Pursuit of Connectomes. J. Neurosci. 2011, 31, 16125–16138. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.W.; Harris, J.A.; Ng, L.; Winslow, B.; Cain, N.; Mihalas, S.; Wang, Q.; Lau, C.; Kuan, L.; Henry, A.M.; et al. A Mesoscale Connectome of the Mouse Brain. Nature 2014, 508, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, J.; Castonguay, A.; Pouliot, P.; Descoteaux, M.; Lesage, F. Whole Mouse Brain Imaging Using Optical Coherence Tomography: Reconstruction, Normalization, Segmentation, and Comparison with Diffusion MRI. Neurophotonics 2017, 4, 41501. [Google Scholar] [CrossRef] [PubMed]
- Wong, T.T.W.; Zhang, R.; Zhang, C.; Hsu, H.-C.; Maslov, K.I.; Wang, L.V.L.; Shi, J.; Chen, R.; Shung, K.K.; Zhou, Q.; et al. Label-Free Automated Three-Dimensional Imaging of Whole Organs by Microtomy-Assisted Photoacoustic Microscopy. Nat. Commun. 2017, 8, 1386. [Google Scholar] [CrossRef] [PubMed]
- Farrar, M.J.; Wise, F.W.; Fetcho, J.R.; Schaffer, C.B. In Vivo Imaging of Myelin in the Vertebrate Central Nervous System Using Third Harmonic Generation Microscopy. Biophys. J. 2011, 100, 1362–1371. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Zhu, J.; Akkin, T. Serial Optical Coherence Scanner for Large-Scale Brain Imaging at Microscopic Resolution. Neuroimage 2014, 84, 1007–1017. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Huff, T.B.; Wang, H.-W.; Cheng, J.-X.; Wang, H. Ex Vivo and in Vivo Imaging of Myelin Fibers in Mouse Brain by Coherent Anti-Stokes Raman Scattering Microscopy. Opt. Express 2008, 16, 19396. [Google Scholar] [CrossRef]
- Yaqoob, Z.; Wu, J.; Yang, C. Spectral Domain Optical Coherence Tomography: A Better OCT Imaging Strategy. Biotechniques 2005, 39 (Suppl. 6). [Google Scholar] [CrossRef]
- Zipfel, W.R.; Williams, R.M.; Christie, R.; Nikitin, A.Y.; Hyman, B.T.; Webb, W.W. Live Tissue Intrinsic Emission Microscopy Using Multiphoton-Excited Native Fluorescence and Second Harmonic Generation. Proc. Natl. Acad. Sci. USA 2003, 100, 7075–7080. [Google Scholar] [CrossRef]
- Wang, L.V. Tutorial on Photoacoustic Microscopy and Computed Tomography. IEEE J. Sel. Top. Quantum Electron. 2008, 14, 171–179. [Google Scholar] [CrossRef]
- Wang, X.; Pang, Y.; Ku, G.; Xie, X.; Stoica, G.; Wang, L.V. Noninvasive Laser-Induced Photoacoustic Tomography for Structural and Functional in Vivo Imaging of the Brain. Nat. Biotechnol. 2003, 21, 803–806. [Google Scholar] [CrossRef] [PubMed]
- Schlup, P.; Masihzadeh, O.; Bartels, R.A. High-Speed, Label-Free Second Harmonic Generation Holographic Microscopy of Biological Specimens. Multiphot. Microsc. Biomed. Sci. XI 2011, 7903, 790308. [Google Scholar] [CrossRef]
- Witte, S.; Negrean, A.; Lodder, J.C.; de Kock, C.P.J.; Testa Silva, G.; Mansvelder, H.D.; Louise Groot, M. Label-Free Live Brain Imaging and Targeted Patching with Third-Harmonic Generation Microscopy. Proc. Natl. Acad. Sci. USA 2011, 108, 5970–5975. [Google Scholar] [CrossRef] [PubMed]
- Ozeki, Y.; Dake, F.; Kajiyama, S.; Fukui, K.; Itoh, K. Analysis and Experimental Assessment of the Sensitivity of Stimulated Raman Scattering Microscopy. Opt. Express 2009, 17, 3651. [Google Scholar] [CrossRef] [PubMed]
- Cote, D.; Lin, C.P.; Evans, C.L.; Puoris’haag, M.; Potma, E.O.; Xie, X.S. Chemical Imaging of Tissue in Vivo with Video-Rate Coherent Anti-Stokes Raman Scattering Microscopy. Proc. Natl. Acad. Sci. USA 2005, 102, 16807–16812. [Google Scholar] [CrossRef]
- Huang, D.; Swanson, E.A.; Lin, C.P.; Schuman, J.S.; Stinson, W.G.; Chang, W.; Hee, M.R.; Flotte, T.; Gregory, K.; Puliafito, C.A.; et al. Optical Coherence Tomography. Science 1991, 254, 1178–1181. [Google Scholar] [CrossRef]
- Bourdieu, L.; Léger, J.-F.; Boccara, C.; Ben Arous, J.; Binding, J.; Gigan, S. Brain Refractive Index Measured in Vivo with High-NA Defocus-Corrected Full-Field OCT and Consequences for Two-Photon Microscopy. Opt. Express 2011, 19, 4833. [Google Scholar] [CrossRef]
- Sun, J.; Jin Lee, S.; Wu, L.; Sarntinoranont, M.; Xie, H. Refractive Index Measurement of Acute Rat Brain Tissue Slices Using Optical Coherence Tomography. Opt. Express 2012, 20, 1084–1095. [Google Scholar] [CrossRef]
- Bachmann, L.; Zezell, D.M.; Ribeiro, A.D.C.; Gomes, L.; Ito, A.S. Fluorescence Spectroscopy of Biological Tissues—A Review. Appl. Spectrosc. Rev. 2006, 41, 575–590. [Google Scholar] [CrossRef]
- Petry, R.; Schmitt, M.; Popp, J. Raman Spectroscopy—A Prospective Tool in the Life Sciences. ChemPhysChem 2003, 4, 14–30. [Google Scholar] [CrossRef] [PubMed]
- Mishchenko, M.I.; Travis, L.D.; Lacis, A.A. Multiple Scattering of Light by Particles: Radiative Transfer and Coherent Backscattering; Cambridge University Press: New York, NY, USA, 2006. [Google Scholar]
- Jacques, S.L. Optical Properties of Biological Tissues: A Review. Phys. Med. Biol. 2013, 58, R37. [Google Scholar] [CrossRef] [PubMed]
- Faber, D.J.; van der Meer, F.J.; Aalders, M.C.G.; van Leeuwen, T.G. Quantitative Measurement of Attenuation Coefficients of Weakly Scattering Media Using Optical Coherence Tomography. Opt. Express 2004, 12, 4353–4365. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Wang, L.V. Photoacoustic Imaging in Biomedicine. Rev. Sci. Instrum. 2006, 77, 41101. [Google Scholar] [CrossRef]
- Li, W.; Chen, R.; Lv, J.; Wang, H.; Liu, Y.; Peng, Y.; Qian, Z.; Fu, G.; Nie, L. In Vivo Photoacoustic Imaging of Brain Injury and Rehabilitation by High-Efficient Near-Infrared Dye Labeled Mesenchymal Stem Cells with Enhanced Brain Barrier Permeability. Adv. Sci. 2018, 5. [Google Scholar] [CrossRef]
- Yang, S.; Xing, D.; Lao, Y.; Yang, D.; Zeng, L.; Xiang, L.; Chen, W.R. Noninvasive Monitoring of Traumatic Brain Injury and Post-Traumatic Rehabilitation with Laser-Induced Photoacoustic Imaging. Appl. Phys. Lett. 2007, 90, 243902. [Google Scholar] [CrossRef]
- Guevara, E.; Berti, R.; Londono, N.; Xie, N.; Bellec, P.; Lesage, R.; Lodygensky, G.A. Imaging of an Inflammatory Injury in the Newborn Rat Brain with Photoacoustic Tomography. PLoS ONE 2013, 8, e83045. [Google Scholar] [CrossRef]
- Ku, G.; Wang, X.; Xie, X.; Stoica, G.; Wang, L.V. Imaging of Tumor Angiogenesis in Rat Brains in Vivo by Photoacoustic Tomography. Appl. Opt. 2005, 44, 770–775. [Google Scholar] [CrossRef]
- Li, M.-L.; Oh, J.-T.; Xie, X.; Ku, G.; Wang, W.; Li, C.; Lungu, G.; Stoica, G.; Wang, L.V. Simultaneous Molecular and Hypoxia Imaging of Brain Tumors In Vivo Using Spectroscopic Photoacoustic Tomography. Proc. IEEE 2008, 96, 481–489. [Google Scholar] [CrossRef]
- De Boer, J.F. Polarization Sensitive Optical Coherence Tomography: A Review of Technology and Applications. Appl. Sci. 2017, 7, 474. [Google Scholar] [CrossRef]
- Kasaragod, D.K.; Lu, Z.; Jacobs, J.; Matcher, S.J. Experimental validation of an extended Jones matrix calculus model to study the 3D structural orientation of the collagen fibers in articular cartilage using polarization-sensitive optical coherence tomography. Biomed. Opt. Express 2012, 3, 378–387. [Google Scholar] [CrossRef] [PubMed]
- Deber, C.M.; Reynolds, S.J. Central Nervous System Myelin: Structure, Function, and Pathology. Clin. Biochem. 1991, 24, 113–134. [Google Scholar] [CrossRef]
- Villiger, M.; Zhang, E.Z.; Nadkarni, S.K.; Oh, W.-Y.; Vakoc, B.J.; Bouma, B.E. Spectral Binning for Mitigation of Polarization Mode Dispersion Artifacts in Catheter-Based Optical Frequency Domain Imaging. Opt. Express 2013, 21, 16353. [Google Scholar] [CrossRef] [PubMed]
- van der Sijde, J.N.; Karanasos, A.; Villiger, M.; Bouma, B.E.; Regar, E. First-in-Man Assessment of Plaque Rupture by Polarization-Sensitive Optical Frequency Domain Imaging in Vivo. Eur. Heart J. 2016, 37, 1932. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y.-J.; Makita, S.; Sugiyama, S.; Yasuno, Y. Optically Buffered Jones-Matrix-Based Multifunctional Optical Coherence Tomography with Polarization Mode Dispersion Correction. Biomed. Opt. Express 2014, 6, 225. [Google Scholar] [CrossRef] [PubMed]
- Sayegh, R.G.; Zotter, S.; Roberts, P.K.; Kandula, M.M.; Sacu, S.; Kreil, D.P.; Baumann, B.; Pircher, M.; Hitzenberger, C.K.; Schmidt-Erfurth, U. Polarization-Sensitive Optical Coherence Tomography and Conventional Retinal Imaging Strategies in Assessing Foveal Integrity in Geographic Atrophy. Investig. Ophthalmol. Vis. Sci. 2015, 56, 5246–5255. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Zhu, J.; Reuter, M.; Vinke, L.N.; Yendiki, A.; Boas, D.A.; Fischl, B.; Akkin, T. Cross-Validation of Serial Optical Coherence Scanning and Diffusion Tensor Imaging: A Study on Neural Fiber Maps in Human Medulla Oblongata. Neuroimage 2014, 100, 395–404. [Google Scholar] [CrossRef]
- Baumann, B.; Woehrer, A.; Ricken, G.; Augustin, M.; Mitter, C.; Pircher, M.; Kovacs, G.G.; Hitzenberger, C.K. Visualization of Neuritic Plaques in Alzheimer’s Disease by Polarization-Sensitive Optical Coherence Microscopy. Sci. Rep. 2017, 7, 43477. [Google Scholar] [CrossRef] [PubMed]
- Boyd, R.W. Nonlinear Optics, 3rd ed.; Academic Press: Amsterdam, NL, USA, 2008. [Google Scholar]
- Nuriya, M.; Jiang, J.; Nemet, B.; Eisenthal, K.B.; Yuste, R. Imaging Membrane Potential in Dendritic Spines. Proc. Natl. Acad. Sci. USA 2006, 103, 786–790. [Google Scholar] [CrossRef]
- Cox, G.; Kable, E.; Jones, A.; Fraser, I.; Manconi, F.; Gorrell, M.D. 3-Dimensional Imaging of Collagen Using Second Harmonic Generation. J. Struct. Biol. 2003, 141, 53–62. [Google Scholar] [CrossRef]
- Stoller, P.; Celliers, P.M.; Reiser, K.M.; Rubenchik, A.M. Quantitative Second-Harmonic Generation Microscopy in Collagen. Appl. Opt. 2003, 42, 5209. [Google Scholar] [CrossRef] [PubMed]
- Mansfield, J.; Yu, J.; Attenburrow, D.; Moger, J.; Tirlapur, U.; Urban, J.; Cui, Z.; Winlove, P. The Elastin Network: Its Relationship with Collagen and Cells in Articular Cartilage as Visualized by Multiphoton Microscopy. J. Anat. 2009, 215, 682–691. [Google Scholar] [CrossRef] [PubMed]
- Becker, W. Fluorescence Lifetime Imaging—Techniques and Applications. J. Microsc. 2012, 247, 119–136. [Google Scholar] [CrossRef] [PubMed]
- König, K.; So, P.T.C.; Mantulin, W.W.; Tromberg, B.J.; Gratton, E. Two-Photon Excited Lifetime Imaging of Autofluorescence in Cells during UV A and NIR Photostress. J. Microsc. 1996, 183, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Yaseen, M.A.; Sakadžić, S.; Wu, W.; Becker, W.; Kasischke, K.A.; Boas, D.A. In Vivo Imaging of Cerebral Energy Metabolism with Two-Photon Fluorescence Lifetime Microscopy of NADH. Biomed. Opt. Express 2013, 4, 307–321. [Google Scholar] [CrossRef]
- Leppert, J.; Krajewski, J.; Kantelhardt, S.R.; Schlaffer, S.; Petkus, N.; Reusche, E.; Hüttmann, G.; Giese, A. Multiphoton Excitation of Autofluorescence for Microscopy of Glioma Tissue. Neurosurgery 2006, 58, 759–767. [Google Scholar] [CrossRef]
- McCamant, D.W.; Kukura, P.; Yoon, S.; Mathies, R.A. Femtosecond Broadband Stimulated Raman Spectroscopy: Apparatus and Methods. Rev. Sci. Instrum. 2004, 75, 4971–4980. [Google Scholar] [CrossRef]
- Begley, R.F.; Harvey, A.B.; Byer, R.L. Coherent Anti-Stokes Raman Spectroscopy. Appl. Phys. Lett. 2004, 25, 387–390. [Google Scholar] [CrossRef]
- Turrell, G.; Corset, J. Raman Microscopy: Developments and Applications; Academic Press: San Diego, CA, USA, 1996. [Google Scholar]
- Fu, D.; Lu, F.-K.; Zhang, X.; Freudiger, C.; Pernik, D.R.; Holtom, G.; Xie, X.S. Quantitative Chemical Imaging with Multiplex Stimulated Raman Scattering Microscopy. J. Am. Chem. Soc. 2012, 134, 3623–3626. [Google Scholar] [CrossRef]
- Fu, D.; Holtom, G.; Freudiger, C.; Zhang, X.; Xie, X.S. Hyperspectral Imaging with Stimulated Raman Scattering by Chirped Femtosecond Lasers. J. Phys. Chem. B 2013, 117, 4634–4640. [Google Scholar] [CrossRef]
- Cheng, J.-X.; Xie, X.S. Coherent Raman Scattering Microscopy; CRC Press: Boca Raton, FL, USA, 2016. [Google Scholar]
- Cheng, J.X.; Xie, X.S. Vibrational Spectroscopic Imaging of Living Systems: An Emerging Platform for Biology and Medicine. Science 2015, 350. [Google Scholar] [CrossRef] [PubMed]
- Min, W.; Freudiger, C.W.; Lu, S.; Xie, X.S. Coherent Nonlinear Optical Imaging: Beyond Fluorescence Microscopy. Annu. Rev. Phys. Chem. 2011, 62, 507–530. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.C.; Min, W.; Freudiger, C.W.; Ruvkun, G.; Xie, X.S. RNAi Screening for Fat Regulatory Genes with SRS Microscopy. Nat. Methods 2011, 8, 135. [Google Scholar] [CrossRef] [PubMed]
- Fu, D.; Yu, Y.; Folick, A.; Currie, E.; Farese, R.V., Jr.; Tsai, T.-H.; Xie, X.S.; Wang, M.C. In Vivo Metabolic Fingerprinting of Neutral Lipids with Hyperspectral Stimulated Raman Scattering Microscopy. J. Am. Chem. Soc. 2014, 136, 8820–8828. [Google Scholar] [CrossRef] [PubMed]
- Ji, M.; Orringer, D.A.; Freudiger, C.W.; Ramkissoon, S.; Liu, X.; Lau, D.; Golby, A.J.; Norton, I.; Hayashi, M.; Agar, N.Y.R.; et al. Rapid, Label-Free Detection of Brain Tumors with Stimulated Raman Scattering Microscopy. Sci. Transl. Med. 2013, 5, 201ra119. [Google Scholar] [CrossRef] [PubMed]
- Ekins, S. Pharmaceutical Applications of Raman Spectroscopy; John Wiley & Sons: Hoboken, NJ, USA, 2008. [Google Scholar]
- Staveley, L.A.K. The Characterization of Chemical Purity: Organic Compounds; Elsevier: Amsterdam, The Netherlands, 1971. [Google Scholar] [CrossRef]
- Hamada, K.; Fujita, K.; Smith, N.I.; Kobayashi, M.; Inouye, Y.; Kawata, S. Raman Microscopy for Dynamic Molecular Imaging of Living Cells. J. Biomed. Opt. 2008, 13, 44027. [Google Scholar] [CrossRef] [PubMed]
- Kirsch, M.; Schackert, G.; Salzer, R.; Krafft, C. Raman Spectroscopic Imaging for in Vivo Detection of Cerebral Brain Metastases. Anal. Bioanal. Chem. 2010, 398, 1707–1713. [Google Scholar] [CrossRef] [PubMed]
- Hollon, T.; Lewis, S.; Freudiger, C.W.; Xie, X.S.; Orringer, D.A. Improving the Accuracy of Brain Tumor Surgery via Raman-Based Technology. Neurosurg. Focus 2016, 40, E9. [Google Scholar] [CrossRef]
- Jermyn, M.; Mok, K.; Mercier, J.; Desroches, J.; Pichette, J.; Saint-Arnaud, K.; Bernstein, L.; Guiot, M.-C.; Petrecca, K.; Leblond, F. Intraoperative Brain Cancer Detection with Raman Spectroscopy in Humans. Sci. Transl. Med. 2015, 7, 274ra19. [Google Scholar] [CrossRef]
- Evans, C.L.; Xu, X.; Kesari, S.; Xie, X.S.; Wong, S.T.C.; Young, G.S. Chemically-Selective Imaging of Brain Structures with CARS Microscopy. Opt. Express 2007, 15, 12076. [Google Scholar] [CrossRef]
- Leahy, C.; Radhakrishnan, H.; Srinivasan, V.J. Volumetric Imaging and Quantification of Cytoarchitecture and Myeloarchitecture with Intrinsic Scattering Contrast. Biomed. Opt. Express 2013, 4, 1913–1978. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, J.; Delafontaine-Martel, P.; Pouliot, P.; Girouard, H.; Descoteaux, M. Fully Automated Dual-Resolution Serial Optical Coherence Tomography Aimed at Diffusion MRI Validation in Whole Mouse Brains. Neurophotonics 2018, 5, 045004. [Google Scholar] [CrossRef] [PubMed]
- Economo, M.N.; Clack, N.G.; Lavis, L.D.; Gerfen, C.R.; Svoboda, K.; Myers, E.W.; Chandrashekar, J. A Platform for Brain-Wide Imaging and Reconstruction of Individual Neurons. Elife 2016, 5, e10566. [Google Scholar] [CrossRef] [PubMed]
- Amato, S.P.; Pan, F.; Schwartz, J.; Ragan, T.M. Whole Brain Imaging with Serial Two-Photon Tomography. Front. Neuroanat. 2016, 10, 31. [Google Scholar] [CrossRef]
- Delafontaine-Martel, P.; Lefebvre, J.; Tardif, P.-L.; Lévy, B.I.; Pouliot, P.; Lesage, F. Whole Brain Vascular Imaging in a Mouse Model of Alzheimer’s Disease with Two-Photon Microscopy. J. Biomed. Opt. 2018, 23, 076501. [Google Scholar] [CrossRef] [PubMed]
- Magnain, C.; Augustinack, J.C.; Reuter, M.; Wachinger, C.; Frosch, M.P.; Ragan, T.; Akkin, T.; Wedeen, V.J.; Boas, D.A.; Fischl, B. Blockface Histology with Optical Coherence Tomography: A Comparison with Nissl Staining. Neuroimage 2014, 84, 524–533. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Magnain, C.; Wang, R.; Dubb, J.; Varjabedian, A.; Tirrell, L.S.; Stevens, A.; Augustinack, J.C.; Konukoglu, E.; Aganj, I.; et al. As-PSOCT: Volumetric Microscopic Imaging of Human Brain Architecture and Connectivity. Neuroimage 2018, 165, 56–68. [Google Scholar] [CrossRef] [PubMed]
- Magnain, C.; Augustinack, J.C.; Tirrell, L.; Fogarty, M.; Frosch, M.P.; Boas, D.; Fischl, B.; Rockland, K.S. Colocalization of Neurons in Optical Coherence Microscopy and Nissl-Stained Histology in Brodmann’s Area 32 and Area 21. Brain Struct. Funct. 2019, 224, 351–362. [Google Scholar] [CrossRef] [PubMed]
- Abdeladim, L.; Matho, K.S.; Clavreul, S.; Mahou, P.; Sintes, J.-M.; Solinas, X.; Arganda-Carreras, I.; Turney, S.G.; Lichtman, J.W.; Chessel, A.; et al. Multicolor Multiscale Brain Imaging with Chromatic Multiphoton Serial Microscopy. Nat. Commun. 2019, 10, 1662. [Google Scholar] [CrossRef] [PubMed]
- Hayworth, K.J.; Xu, C.S.; Lu, Z.; Knott, G.W.; Fetter, R.D.; Tapia, J.C.; Lichtman, J.W.; Hess, H.F. Ultrastructurally Smooth Thick Partitioning and Volume Stitching for Large-Scale Connectomics. Nat. Methods 2015, 12. [Google Scholar] [CrossRef] [PubMed]
- Hildebrand, D.G.C.; Cicconet, M.; Torres, R.M.; Choi, W.; Quan, T.M.; Moon, J.; Wetzel, A.W.; Scott Champion, A.; Graham, B.J.; Randlett, O.; et al. Whole-Brain Serial-Section Electron Microscopy in Larval Zebrafish. Nature 2017, 545, 345–349. [Google Scholar] [CrossRef] [PubMed]
- Mayerich, D.; Abbott, L.; McCormick, B. Knife-Edge Scanning Microscopy for Imaging and Reconstruction of Three-Dimensional Anatomical Structures of the Mouse Brain. J. Microsc. 2008, 231, 134–143. [Google Scholar] [CrossRef] [PubMed]
- Choe, Y.; Mayerich, D.; Kwon, J.; Miller, D.E.; Chung, J.R.; Sung, C.; Keyser, J.; Abbott, L.C. Knife-Edge Scanning Microscopy for Connectomics Research. In Proceedings of the 2011 International Joint Conference on Neural Networks, San Jose, CA, USA, 31 July–5 August 2011; pp. 2258–2265. [Google Scholar] [CrossRef]
- Tsai, P.S.; Blinder, P.; Squier, J.A.; Kleinfeld, D. All-Optical In Situ Histology of Brain Tissue with Femtosecond Laser Pulses. Cold Spring Harb. Protoc. 2013, 2013. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, Q.T.; Tsai, P.S.; Kleinfeld, D. MPScope: A Versatile Software Suite for Multiphoton Microscopy. J. Neurosci. Methods 2006, 156, 351–359. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, Q.-T.; Driscoll, J.; Dolnick, E.M.; Kleinfeld, D. MPScope 2.0. A Computer System for Two-Photon Laser Scanning Microscopy with Concurrent Plasma-Mediated Ablation and Electrophysiology. Vivo Opt. Imaging Brain Funct. Second Ed. 2009, 1–18. [Google Scholar] [CrossRef]
- Edelstein, A.; Amodaj, N.; Hoover, K.; Vale, R.; Stuurman, N. Computer Control of Microscopes Using ΜManager. Curr. Protoc. Mol. Biol. 2010, 92, 14–20. [Google Scholar] [CrossRef]
- Edelstein, A.D.; Tsuchida, M.A.; Amodaj, N.; Pinkard, H.; Vale, R.D.; Stuurman, N. Advanced Methods of Microscope Control Using ΜManager Software. J. Biol. Methods 2014, 1, e10. [Google Scholar] [CrossRef]
- Babaloukas, G.; Tentolouris, N.; Liatis, S.; Sklavounou, A.; Perrea, D. Evaluation of Three Methods for Retrospective Correction of Vignetting on Medical Microscopy Images Utilizing Two Open Source Software Tools. J. Microsc. 2011, 244, 320–324. [Google Scholar] [CrossRef]
- Tomer, R.; Ye, L.; Hsueh, B.; Deisseroth, K. Advanced CLARITY for Rapid and High-Resolution Imaging of Intact Tissues. Nat. Protoc. 2014, 9, 1682–1697. [Google Scholar] [CrossRef]
- Berlanga, M.L.; Phan, S.; Bushong, E.A.; Wu, S.; Kwon, O.; Phung, B.S.; Lamont, S.; Terada, M.; Tasdizen, T.; Martone, M.E.; et al. Three-Dimensional Reconstruction of Serial Mouse Brain Sections: Solution for Flattening High-Resolution Large-Scale Mosaics. Front. Neuroanat. 2011, 5, 17. [Google Scholar] [CrossRef]
- Seiriki, K.; Kasai, A.; Hashimoto, T.; Schulze, W.; Niu, M.; Yamaguchi, S.; Nakazawa, T.; Inoue, K.-I.; Uezono, S.; Takada, M.; et al. High-Speed and Scalable Whole-Brain Imaging in Rodents and Primates. Neuron 2017, 94, 1085–1100.e6. [Google Scholar] [CrossRef] [PubMed]
- Peng, T.; Thorn, K.; Schroeder, T.; Wang, L.; Theis, F.J.; Marr, C.; Navab, N. A BaSiC Tool for Background and Shading Correction of Optical Microscopy Images. Nat. Commun. 2017, 8, 14836. [Google Scholar] [CrossRef] [PubMed]
- Yayon, N.; Dudai, A.; Vrieler, N.; Amsalem, O.; London, M.; Soreq, H. Intensify3D: Normalizing Signal Intensity in Large Heterogenic Image Stacks. Sci. Rep. 2018, 8, 4311. [Google Scholar] [CrossRef] [PubMed]
- Price, D.L.; Chow, S.K.; MacLean, N.A.B.; Hakozaki, H.; Peltier, S.; Martone, M.E.; Ellisman, M.H. High-Resolution Large-Scale Mosaic Imaging Using Multiphoton Microscopy to Characterize Transgenic Mouse Models of Human Neurological Disorders. Neuroinformatics 2006, 4, 65–80. [Google Scholar] [CrossRef]
- Chow, S.K.; Hakozaki, H.; Price, D.L.; MacLean, N.A.B.; Deerinck, T.J.; Bouwer, J.C.; Martone, M.E.; Peltier, S.T.; Ellisman, M.H. Automated Microscopy System for Mosaic Acquisition and Processing. J. Microsc. 2006, 222, 76–84. [Google Scholar] [CrossRef]
- Preibisch, S.; Saalfeld, S.; Tomancak, P. Fast Stitching of Large 3d Biological Datasets. In Proceedings of the Image J User and Developer Conference, Luxembourg, 6–7 November 2008. [Google Scholar]
- Preibisch, S.; Saalfeld, S.; Tomancak, P. Globally Optimal Stitching of Tiled 3D Microscopic Image Acquisitions. Bioinformatics 2009, 25, 1463–1465. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An Open-Source Platform for Biological-Image Analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Bria, A.; Iannello, G. TeraStitcher—A Tool for Fast Automatic 3D-Stitching of Teravoxel-Sized Microscopy Images. BMC Bioinform. 2012, 13, 316. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Ruan, Z.; Long, F.; Simpson, J.H.; Myers, E.W. V3D Enables Real-Time 3D Visualization and Quantitative Analysis of Large-Scale Biological Image Data Sets. Nat. Biotechnol. 2010, 28, 348–353. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Bria, A.; Zhou, Z.; Iannello, G.; Long, F. Extensible Visualization and Analysis for Multidimensional Images Using Vaa3D. Nat. Protoc. 2014, 9, 193–208. [Google Scholar] [CrossRef]
- Ren, J.; Choi, H.; Chung, K.; Bouma, B.E. Label-Free Volumetric Optical Imaging of Intact Murine Brains. Sci. Rep. 2017, 7, 46306. [Google Scholar] [CrossRef] [PubMed]
- Silvestri, L.; Allegra Mascaro, A.L.; Costantini, I.; Sacconi, L.; Pavone, F.S. Correlative Two-Photon and Light Sheet Microscopy. Methods 2014, 66, 268–272. [Google Scholar] [CrossRef] [PubMed]
- Piccinini, F.; Bevilacqua, A.; Lucarelli, E. Automated Image Mosaics by Non-Automated Light Microscopes: The MicroMos Software Tool. J. Microsc. 2013, 252, 226–250. [Google Scholar] [CrossRef] [PubMed]
- Emmenlauer, M.; Ronneberger, O. XuvTools: Free, Fast and Reliable Stitching of Large 3D Datasets. J. Microsc. 2009, 233, 42–60. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.C.; Tward, D.J.; Mitra, P.P.; Miller, M.I. On Variational Solutions for Whole Brain Serial-Section Histology Using a Sobolev Prior in the Computational Anatomy Random Orbit Model. PLoS. Comput. Biol. 2018, 14, e1006610. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.C.; Lin, M.K.; Fu, Y.; Hata, J.; Miller, M.I.; Mitra, P.P. Joint Atlas-Mapping of Multiple Histological Series Combined with Multimodal MRI of Whole Marmoset Brains. arXiv 2018, arXiv:1805.04975. [Google Scholar]
- Ali, S.; Schober, M.; Schlöme, P.; Amunts, K.; Axer, M.; Rohr, K. Towards Ultra-High Resolution 3D Reconstruction of a Whole Rat Brain from 3D-PLI Data; Springer: Cham, Switzerland, 2018. [Google Scholar]
- Amunts, K.; Lepage, C.; Borgeat, L. BigBrain: An Ultrahigh-Resolution 3D Human Brain Model. Science 2013, 340, 1472–1475. [Google Scholar] [CrossRef] [PubMed]
- Susaki, E.A.; Ueda, H.R. Review Whole-Body and Whole-Organ Clearing and Imaging Techniques with Single-Cell Resolution: Toward Organism-Level Systems Biology in Mammals. Cell Chem. Biol. 2016, 23, 137–157. [Google Scholar] [CrossRef]
- Chung, K.; Deisseroth, K. CLARITY for Mapping the Nervous System. Nat. Methods 2013, 10, 508–513. [Google Scholar] [CrossRef]
- Qi, Y.; Yu, T.; Xu, J.; Wan, P.; Ma, Y.; Zhu, J.; Li, Y.; Gong, H.; Luo, Q.; Zhu, D. FDISCO: Advanced Solvent-Based Clearing Method for Imaging Whole Organs. Arch. Stud. Urbani Reg. 2019, 5, eaau8355. [Google Scholar] [CrossRef]
- Sharpe, J. Optical Projection Tomography as a Tool for 3D Microscopy and Gene Expression Studies. Science (80-) 2002, 296, 541–545. [Google Scholar] [CrossRef] [PubMed]
- Sharpe, J. Optical Projection Tomography. Annu. Rev. Biomed. Eng. 2004, 6, 209–228. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, D.; Marchand, P.J.; Planchette, A.L.; Nilsson, J.; Sison, M.; Extermann, J.; Lopez, A.; Sylwestrzak, M.; Sordet-Dessimoz, J.; Schmidt-Christensen, A.; et al. Optical Projection Tomography for Rapid Whole Mouse Brain Imaging. Biomed. Opt. Express 2017, 8, 5637. [Google Scholar] [CrossRef] [PubMed]
- Dodt, H.U.; Leischner, U.; Schierloh, A.; Jährling, N.; Mauch, C.P.; Deininger, K.; Deussing, J.M.; Eder, M.; Zieglgänsberger, W.; Becker, K. Ultramicroscopy: Three-Dimensional Visualization of Neuronal Networks in the Whole Mouse Brain. Nat. Methods 2007, 4, 331–336. [Google Scholar] [CrossRef] [PubMed]
- Silvestri, L.; Bria, A.; Sacconi, L.; Iannello, G.; Pavone, F.S. Confocal Light Sheet Microscopy: Micron-Scale Neuroanatomy of the Entire Mouse Brain. Opt. Express 2012, 20, 20582. [Google Scholar] [CrossRef]
- Silvestri, L.; Bria, A.; Costantini, I.; Sacconi, L.; Peng, H.; Iannello, G.; Pavone, F.S. Micron-Scale Resolution Optical Tomography of Entire Mouse Brains with Confocal Light Sheet Microscopy. J. Vis. Exp. 2013, e50696. [Google Scholar] [CrossRef]
- Ahrens, M.B.; Orger, M.B.; Robson, D.N.; Li, J.M.; Keller, P.J. Whole-Brain Functional Imaging at Cellular Resolution Using Light-Sheet Microscopy. Nat. Methods 2013, 10, 413–420. [Google Scholar] [CrossRef]
- Keller, P.J.; Ahrens, M.B. Visualizing Whole-Brain Activity and Development at the Single-Cell Level Using Light-Sheet Microscopy. Neuron 2015, 85, 462–483. [Google Scholar] [CrossRef]
- Pende, M.; Becker, K.; Wanis, M.; Saghafi, S.; Kaur, R.; Hahn, C.; Pende, N.; Foroughipour, M.; Hummel, T.; Dodt, H.U. High-Resolution Ultramicroscopy of the Developing and Adult Nervous System in Optically Cleared Drosophila Melanogaster. Nat. Commun. 2018, 9, 4731. [Google Scholar] [CrossRef]
- Gao, R.; Asano, S.M.; Upadhyayula, S.; Pisarev, I.; Milkie, D.E.; Liu, T.L.; Singh, V.; Graves, A.; Huynh, G.H.; Zhao, Y.; et al. Cortical Column and Whole-Brain Imaging with Molecular Contrast and Nanoscale Resolution. Science 2019, 363. [Google Scholar] [CrossRef]
- Richardson, D.S.; Lichtman, J.W. Clarifying Tissue Clearing. Cell 2015, 246–257. [Google Scholar] [CrossRef] [PubMed]
- Silvestri, L.; Costantini, I.; Sacconi, L.; Pavone, F.S. Clearing of Fixed Tissue: A Review from a Microscopist’s Perspective. J. Biomed. Opt. 2016, 21, 081205. [Google Scholar] [CrossRef] [PubMed]
- Pinskiy, V.; Jones, J.; Tolpygo, A.S.; Franciotti, N.; Weber, K.; Mitra, P.P. High-Throughput Method of Whole-Brain Sectioning, Using the Tape-Transfer Technique. PLoS ONE 2015, 10, e0102363. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.K.; Takahashi, Y.S.; Huo, B.-X.; Hanada, M.; Nagashima, J.; Hata, J.; Tolpygo, A.S.; Ram, K.; Lee, B.C.; Miller, M.I.; et al. A High-Throughput Neurohistological Pipeline for Brain-Wide Mesoscale Connectivity Mapping of the Common Marmoset. Elife 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Vandenberghe, M.E.; Hérard, A.-S.; Souedet, N.; Sadouni, E.; Santin, M.D.; Briet, D.; Carré, D.; Schulz, J.; Hantraye, P.; Chabrier, P.-E.; et al. High-Throughput 3D Whole-Brain Quantitative Histopathology in Rodents. Sci. Rep. 2016, 6, 20958. [Google Scholar] [CrossRef]
- Liu, C.J.; Williams, K.E.; Orr, H.T.; Akkin, T. Visualizing and Mapping the Cerebellum with Serial Optical Coherence Scanner. Neurophotonics 2016, 4, 011006. [Google Scholar] [CrossRef]
- Wang, H.; Lenglet, C.; Akkin, T. Structure Tensor Analysis of Serial Optical Coherence Scanner Images for Mapping Fiber Orientations and Tractography in the Brain. J. Biomed. Opt. 2015, 20, 036003. [Google Scholar] [CrossRef]
- Castonguay, A.; Lefebvre, J.; Lesage, F.; Pouliot, P. Comparing Three-Dimensional Serial Optical Coherence Tomography Histology to MRI Imaging in the Entire Mouse Brain. J. Biomed. Opt. 2018, 23, 016008. [Google Scholar] [CrossRef]
- Bégin, S.; Dupont-Therrien, O.; Bélanger, E.; Daradich, A.; Laffray, S.; De Koninck, Y.; Côté, D.C. Automated Method for the Segmentation and Morphometry of Nerve Fibers in Large-Scale CARS Images of Spinal Cord Tissue. Biomed. Opt. Express 2014, 5, 4145–4161. [Google Scholar] [CrossRef]
- Zaimi, A.; Duval, T.; Gasecka, A.; Côté, D.; Stikov, N.; Cohen-Adad, J. AxonSeg: Open Source Software for Axon and Myelin Segmentation and Morphometric Analysis. Front. Neuroinform. 2016, 10, 37. [Google Scholar] [CrossRef]
- Zaimi, A.; Wabartha, M.; Herman, V.; Antonsanti, P.-L.; Perone, C.S.; Cohen-Adad, J. AxonDeepSeg: Automatic Axon and Myelin Segmentation from Microscopy Data Using Convolutional Neural Networks. Sci. Rep. 2018, 8, 3816. [Google Scholar] [CrossRef] [PubMed]
- Stikov, N.; Campbell, J.S.W.; Stroh, T.; Lavelée, M.; Frey, S.; Novek, J.; Nuara, S.; Ho, M.-K.; Bedell, B.J.; Dougherty, R.F.; et al. Quantitative Analysis of the Myelin G-Ratio from Electron Microscopy Images of the Macaque Corpus Callosum. Data Br. 2015, 4, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Stikov, N.; Campbell, J.S.W.; Stroh, T.; Lavelée, M.; Frey, S.; Novek, J.; Nuara, S.; Ho, M.-K.; Bedell, B.J.; Dougherty, R.F.; et al. In Vivo Histology of the Myelin G-Ratio with Magnetic Resonance Imaging. Neuroimage 2015, 118, 397–405. [Google Scholar] [CrossRef] [PubMed]
- Cohen-Adad, J. Microstructural Imaging in the Spinal Cord and Validation Strategies. Neuroimage 2018, 182, 169–183. [Google Scholar] [CrossRef] [PubMed]
- MacKenzie-Graham, A.; Lee, E.-F.; Dinov, I.D.; Bota, M.; Shattuck, D.W.; Ruffins, S.; Yuan, H.; Konstantinidis, F.; Pitiot, A.; Ding, Y.; et al. A Multimodal, Multidimensional Atlas of the C57BL/6J Mouse Brain. J. Anat. 2004, 204, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Johnson, G.A.; Badea, A.; Brandenburg, J.; Cofer, G.; Fubara, B.; Liu, S.; Nissanov, J. Waxholm Space: An Image-Based Reference for Coordinating Mouse Brain Research. Neuroimage 2010, 53, 365–372. [Google Scholar] [CrossRef] [PubMed]
- Majka, P.; Chlodzinska, N.; Turlejski, K.; Banasik, T.; Djavadian, R.L.; Węglarz, W.P.; Wójcik, D.K. A Three-Dimensional Stereotaxic Atlas of the Gray Short-Tailed Opossum (Monodelphis Domestica) Brain. Brain Struct. Funct. 2017. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Xu, D.; Yuan, J.; Li, X.; Guo, C.; Peng, J.; Li, Y.; Schwarz, L.A.; Li, A.; Hu, B.; et al. High-Throughput Dual-Colour Precision Imaging for Brain-Wide Connectome with Cytoarchitectonic Landmarks at the Cellular Level. Nat. Commun. 2016, 7, 12142. [Google Scholar] [CrossRef]
- Lu, H.; Scholl, C.A.; Zuo, Y.; Demny, S.; Rea, W.; Stein, E.A.; Yang, Y. Registering and Analyzing Rat FMRI Data in the Stereotaxic Framework by Exploiting Intrinsic Anatomical Features. Magn. Reson. Imaging 2010, 28, 146–152. [Google Scholar] [CrossRef]
- Li, X.; Aggarwal, M.; Hsu, J.; Jiang, H.; Mori, S. AtlasGuide: Software for Stereotaxic Guidance Using 3D CT/MRI Hybrid Atlases of Developing Mouse Brains. J. Neurosci. Methods 2013, 220, 75–84. [Google Scholar] [CrossRef]
- Rangarajan, J.R.; Vande Velde, G.; van Gent, F.; De Vloo, P.; Dresselaers, T.; Depypere, M.; van Kuyck, K.; Nuttin, B.; Himmelreich, U.; Maes, F. Image-Based in Vivo Assessment of Targeting Accuracy of Stereotactic Brain Surgery in Experimental Rodent Models. Sci. Rep. 2016, 6, 38058. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, C.; Chen, S.C. Sectioning Soft Materials with an Oscillating Blade. Precis. Eng. 2019, 56, 96–100. [Google Scholar] [CrossRef]
- Gutiérrez, R.; Gómez, F.; Roa-Peña, L.; Romero, E. A Supervised Visual Model for Finding Regions of Interest in Basal Cell Carcinoma Images. Diagn. Pathol. 2011, 6, 26. [Google Scholar] [CrossRef] [PubMed]
- Pavillon, N.; Smith, N.I. Compressed Sensing Laser Scanning Microscopy. Opt. Express 2016, 24, 30038. [Google Scholar] [CrossRef] [PubMed]
- Woringer, M.; Darzacq, X.; Zimmer, C.; Mir, M. A Versatile Compressed Sensing Scheme for Faster and Less Phototoxic 3D Fluorescence Microscopy. bioRxiv 2017, 125815. [Google Scholar] [CrossRef]
- Ouyang, W.; Aristov, A.; Lelek, M.; Hao, X.; Zimmer, C. Deep Learning Massively Accelerates Super-Resolution Localization Microscopy. Nat. Biotechnol. 2018, 36, 460–468. [Google Scholar] [CrossRef]
- Lehtinen, J.; Munkberg, J.; Hasselgren, J.; Laine, S.; Karras, T.; Aittala, M.; Aila, T. Noise2Noise: Learning Image Restoration without Clean Data. In Proceedings of the 35th International Conference on Machine Learning, Stockholm, Sweden, 10–15 July 2018. [Google Scholar]
- Zhang, H.; Fang, C.; Xie, X.; Yang, Y.; Mei, W.; Jin, D.; Fei, P. High-Throughput, High-Resolution Deep Learning Microscopy Based on Registration-Free Generative Adversarial Network. Biomed. Opt. Express 2019, 10, 1044. [Google Scholar] [CrossRef]
- Xiong, J.; Ren, J.; Luo, L.; Horowitz, M. Mapping Histological Slice Sequences to the Allen Mouse Brain Atlas Without 3D Reconstruction. Front. Neuroinform. 2018, 12, 93. [Google Scholar] [CrossRef]
- Waithe, D.; Brown, J.; Reglinski, K.; Diez-Sevilla, I.; Roberts, D.; Eggeling, C. Object Detection Networks and Augmented Reality for Cellular Detection in Fluorescence Microscopy Acquisition and Analysis. bioRxiv 2019, 544833. [Google Scholar] [CrossRef]
- Chen, P.-H.C.; Gadepalli, K.; MacDonald, R.; Liu, Y.; Nagpal, K.; Kohlberger, T.; Dean, J.; Corrado, G.S.; Hipp, J.D.; Stumpe, M.C. Microscope 2.0: An Augmented Reality Microscope with Real-Time Artificial Intelligence Integration. arXiv 2018, arXiv:1812.00825. [Google Scholar]
- Durand, A.; Wiesner, T.; Gardner, M.A.; Robitaille, L.É.; Bilodeau, A.; Gagné, C.; De Koninck, P.; Lavoie-Cardinal, F. A Machine Learning Approach for Online Automated Optimization of Super-Resolution Optical Microscopy. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Magliaro, C.; Tirella, A.; Mattei, G.; Pirone, A.; Ahluwalia, A. HisTOOLogy: An Open-Source Tool for Quantitative Analysis of Histological Sections. J. Microsc. 2015, 260, 260–267. [Google Scholar] [CrossRef] [PubMed]
- Janowczyk, A.; Madabhushi, A. Deep Learning for Digital Pathology Image Analysis: A Comprehensive Tutorial with Selected Use Cases. J. Pathol. Inform. 2016, 7, 29. [Google Scholar] [CrossRef] [PubMed]
- Komura, D.; Ishikawa, S. Machine Learning Methods for Histopathological Image Analysis. Comput. Struct. Biotechnol. J. 2018, 16, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; McElvain, L.E.; Tolpygo, A.S.; Ferrante, D.; Friedman, B.; Mitra, P.P.; Karten, H.J.; Freund, Y.; Kleinfeld, D. An Active Texture-Based Digital Atlas Enables Automated Mapping of Structures and Markers across Brains. Nat. Methods 2019, 16. [Google Scholar] [CrossRef] [PubMed]
- Pinto, M.; Zorn, K.C.; Tremblay, J.-P.; Desroches, J.; Dallaire, F.; Aubertin, K.; Marple, E.; Kent, C.; Leblond, F.; Trudel, D.; et al. Integration of a Raman Spectroscopy System to a Robotic-Assisted Surgical System for Real-Time Tissue Characterization during Radical Prostatectomy Procedures. J. Biomed. Opt. 2019, 24, 025001. [Google Scholar] [CrossRef] [PubMed]
- Girkin, J.M.; Carvalho, M.T. The Light-Sheet Microscopy Revolution. J. Opt. 2018, 20, 053002. [Google Scholar] [CrossRef]
- Pitrone, P.G.; Schindelin, J.; Stuyvenberg, L.; Preibisch, S.; Weber, M.; Eliceiri, K.W.; Huisken, J.; Tomancak, P. OpenSPIM: An Open-Access Light-Sheet Microscopy Platform. Nat. Methods 2013, 10, 598–599. [Google Scholar] [CrossRef] [PubMed]
- Voigt, F.F.; Kirschenbaum, D.; Platonova, E.; Pagès, S.; Campbell, R.A.A.; Kästli, R.; Schaettin, M.; Egolf, L.; van der Bourg, A.; Bethge, P.; et al. The MesoSPIM Initiative: Open-Source Light-Sheet Mesoscopes for Imaging in Cleared Tissue. bioRxiv 2019, 577122. [Google Scholar] [CrossRef]
- Renier, N.; Wu, Z.; Simon, D.J.; Yang, J.; Ariel, P.; Tessier-Lavigne, M. IDISCO: A Simple, Rapid Method to Immunolabel Large Tissue Samples for Volume Imaging. Cell 2014, 159, 896–910. [Google Scholar] [CrossRef]
- Renier, N.; Adams, E.L.; Kirst, C.; Wu, Z.; Azevedo, R.; Kohl, J.; Autry, A.E.; Kadiri, L.; Umadevi Venkataraju, K.; Zhou, Y.; et al. Mapping of Brain Activity by Automated Volume Analysis of Immediate Early Genes. Cell 2016, 165, 1789–1802. [Google Scholar] [CrossRef] [PubMed]
- Bakker, R.; Tiesinga, P.; Kötter, R. The Scalable Brain Atlas: Instant Web-Based Access to Public Brain Atlases and Related Content. Neuroinformatics 2015, 13, 353–366. [Google Scholar] [CrossRef] [PubMed]
| Imaging Technique | Sensitivity | Spatial Resolution (Voxel Size) | Speed | Wavelength |
|---|---|---|---|---|
| OCT/PS-OCT/SDOCT | 90 dB–110 dB [21] | 6 × 6 × 3.5 µm3 [19] 25 × 25 × 25 µm3 [16] | Fast (4 cm3/h) [19] | NIR-IR |
| Autofluorescence TPEFm | Highly variable [22] | 1 × 1 × 2.5 µm3 [12] | Slow (190 mm3/h) [12] | NIR |
| PAM | ~65 dB [23] | 1.5 mm × 1.5 mm [24] | Fast (18 cm3/h) [24] | Visible-NIR |
| SHG/ THG | >20 dB [25] | 0.6 × 0.6 × 2 µm3 [26] | Slow (125 µm3/h) [26] | NIR-IR |
| SRS/CARS | >20 dB [27] | 0.3 × 0.3 × 1.5 µm3 [28] | Slow (4000 µm3/h) [28] | NIR-IR |
| Components | Parts |
|---|---|
| Tissue preparation | Animal sacrifice protocol Brain extraction protocol Tissue storage Tissue clearing protocol Agarose embedding Pre-Acquisition with other modalities (e.g., MRI) |
| Microscope and Optical Design | Collection optics Beam scanning system Microscope objective swapping Modality swapping (e.g., PS-OCT to confocal) Various microscope modalities |
| Mechanical | Motorized sample stage Tissue sectioning system Slice collection Immersion container Sample holder |
| Acquisition control | Calibration procedures Stage motion Tissue detection Mosaic path planning Image acquisition Tissue slicing Focus depth optimization Data management system In-situ reconstruction method Visualization and control interface Acquisition cards for communication with computer |
| Reconstruction and Analysis | Preprocessing methods (illumination inhomogeneity compensation, denoising, artifacts removal, attenuation compensation, and tissue segmentation) Tile registration Tile stitching Registration to a template Brain parcellation and atlasing |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lefebvre, J.; Delafontaine-Martel, P.; Lesage, F. A Review of Intrinsic Optical Imaging Serial Blockface Histology (ICI-SBH) for Whole Rodent Brain Imaging. Photonics 2019, 6, 66. https://doi.org/10.3390/photonics6020066
Lefebvre J, Delafontaine-Martel P, Lesage F. A Review of Intrinsic Optical Imaging Serial Blockface Histology (ICI-SBH) for Whole Rodent Brain Imaging. Photonics. 2019; 6(2):66. https://doi.org/10.3390/photonics6020066
Chicago/Turabian StyleLefebvre, Joël, Patrick Delafontaine-Martel, and Frédéric Lesage. 2019. "A Review of Intrinsic Optical Imaging Serial Blockface Histology (ICI-SBH) for Whole Rodent Brain Imaging" Photonics 6, no. 2: 66. https://doi.org/10.3390/photonics6020066
APA StyleLefebvre, J., Delafontaine-Martel, P., & Lesage, F. (2019). A Review of Intrinsic Optical Imaging Serial Blockface Histology (ICI-SBH) for Whole Rodent Brain Imaging. Photonics, 6(2), 66. https://doi.org/10.3390/photonics6020066





