Deep Convolutional Neural Network for Detection and Prediction of Waxy Corn Seed Viability Using Hyperspectral Reflectance Imaging
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Preparation
2.2. Standard Germination Experiment
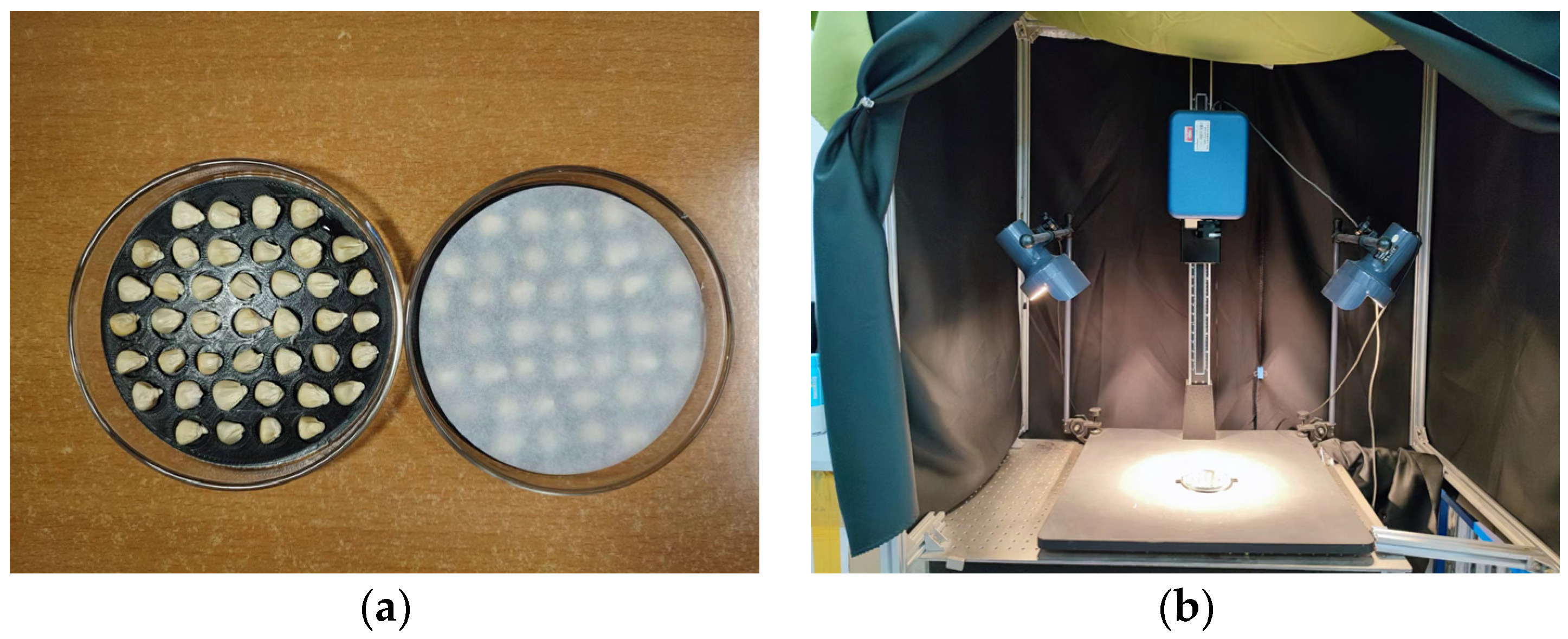
2.3. Hyperspectral Image Acquisition and Correction
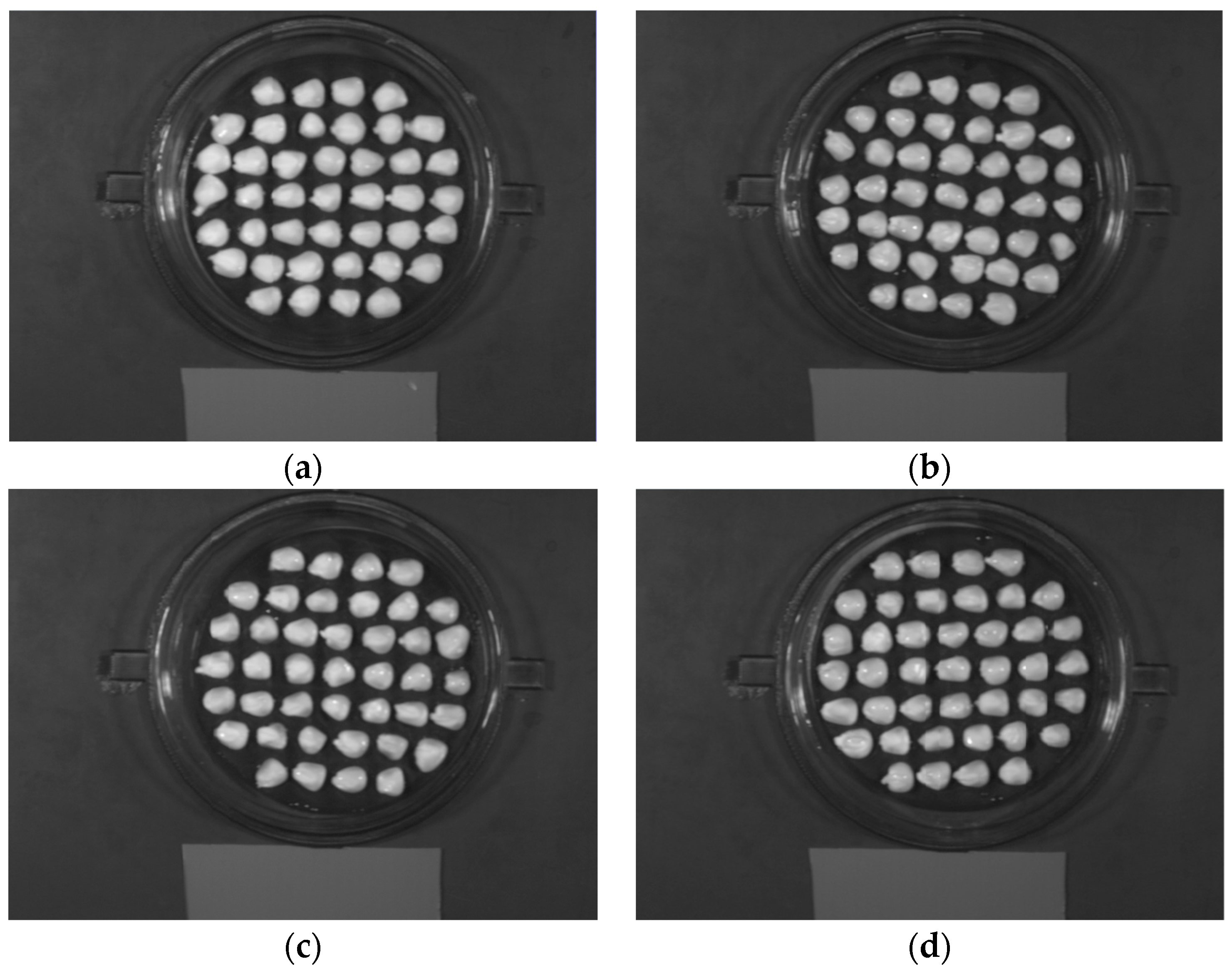
2.4. Spectra Extraction
2.5. Multivariate Data Analysis
2.6. Software
3. Results and Discussion
3.1. Overview of Waxy Corn Seed Spectra
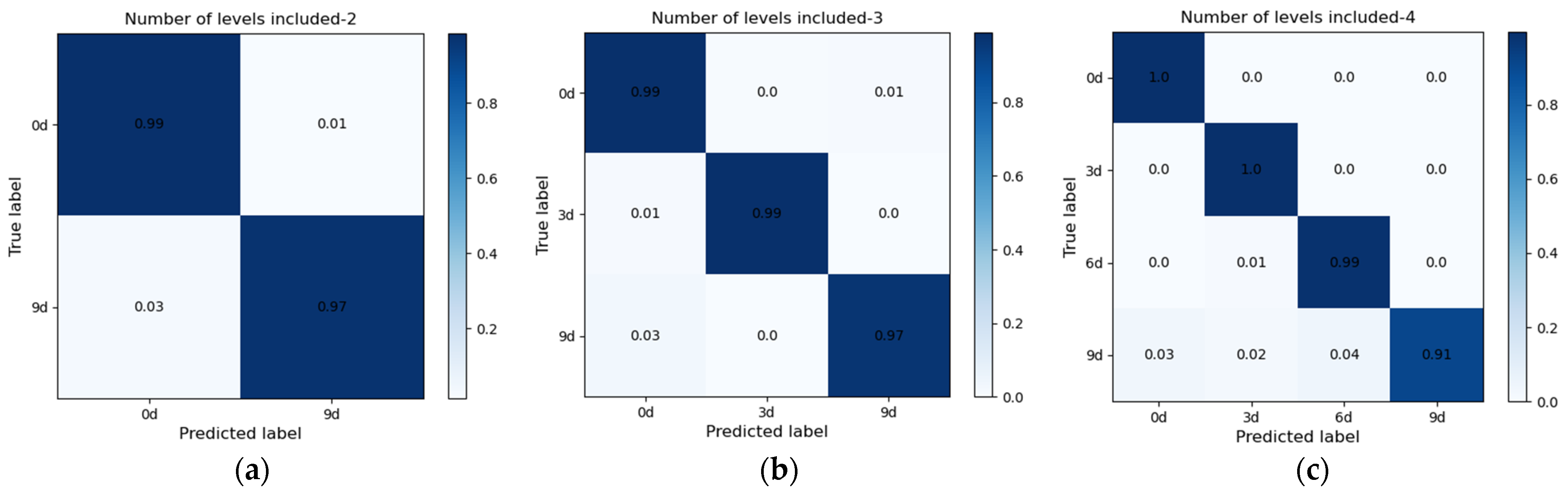
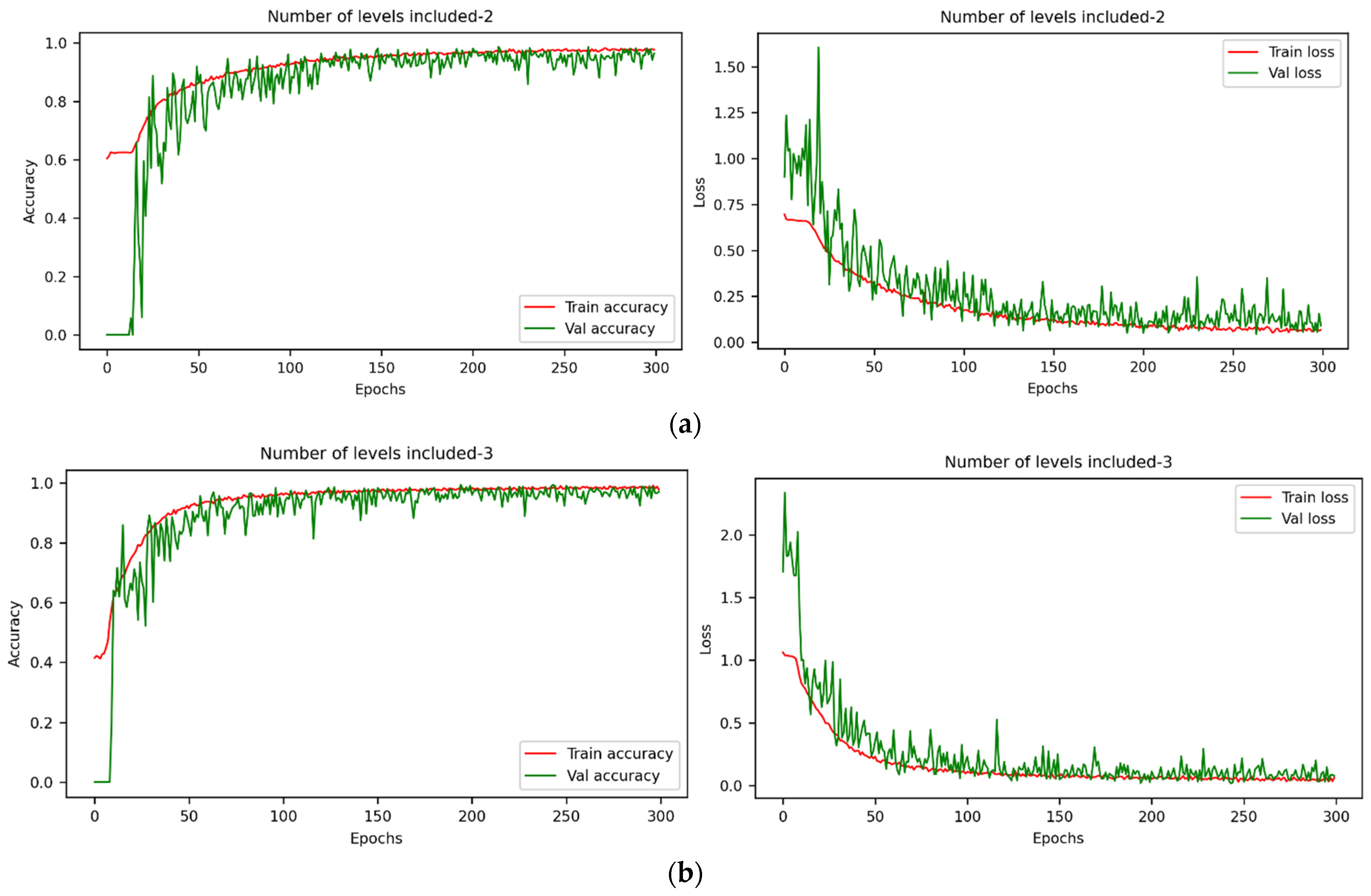
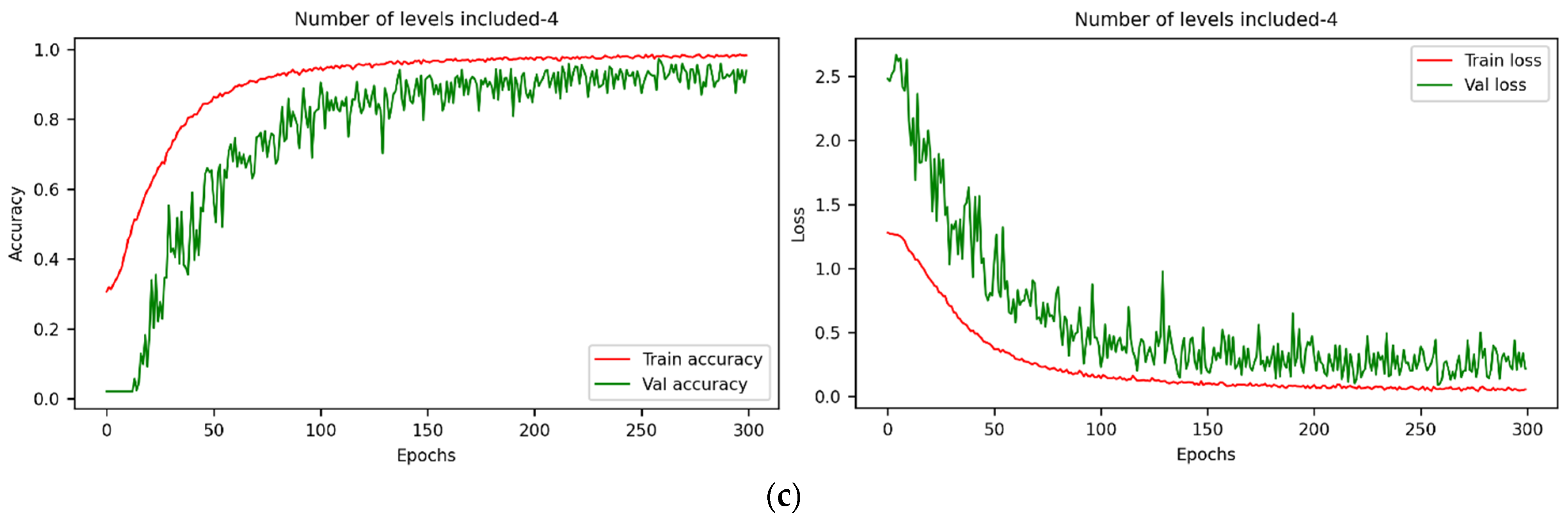
3.2. Detection Results of Seed Viability under Different Models with Full-Wavelength Data
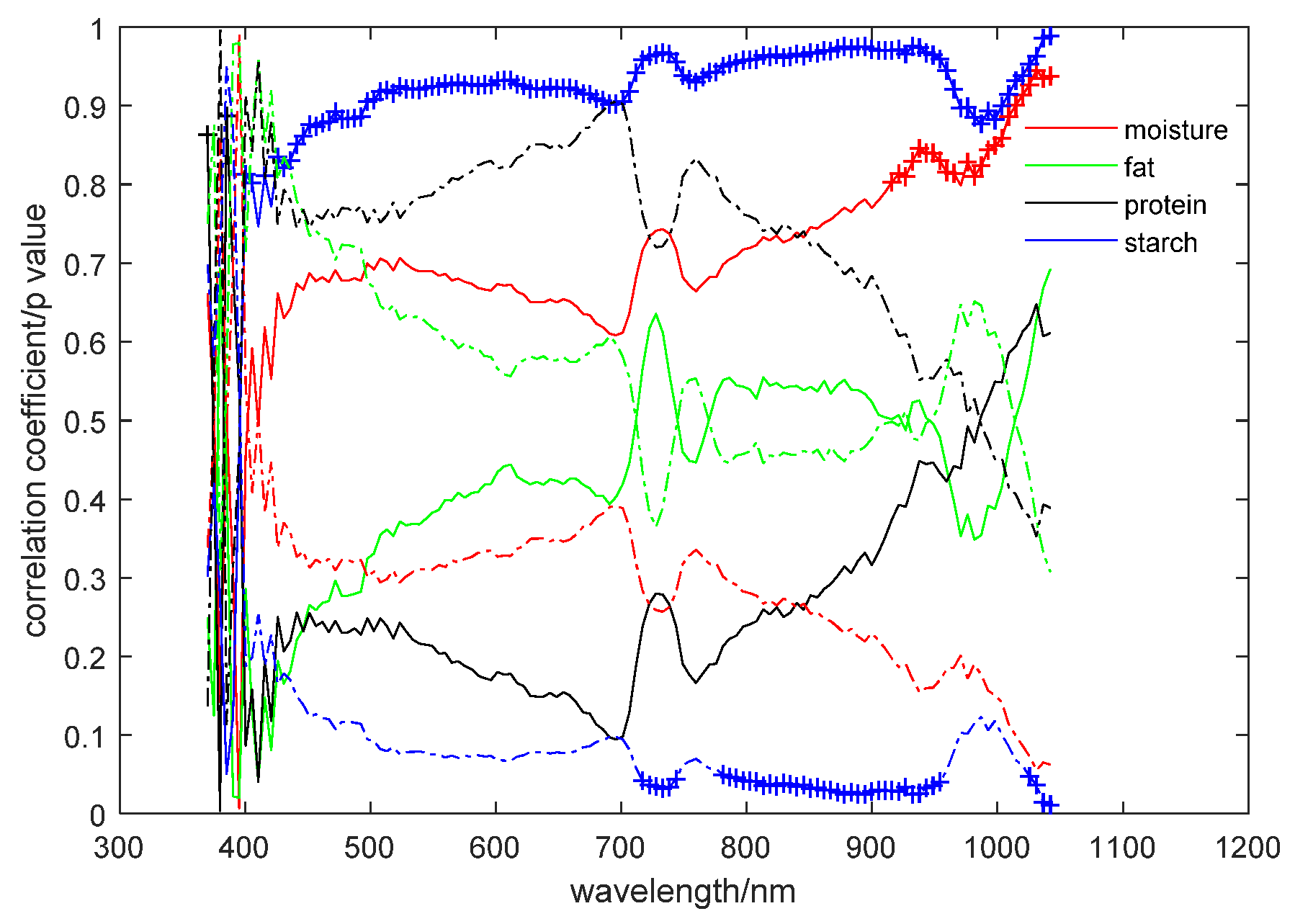
3.3. Chemical Composition Analysis and Characteristic Wavelength Selection
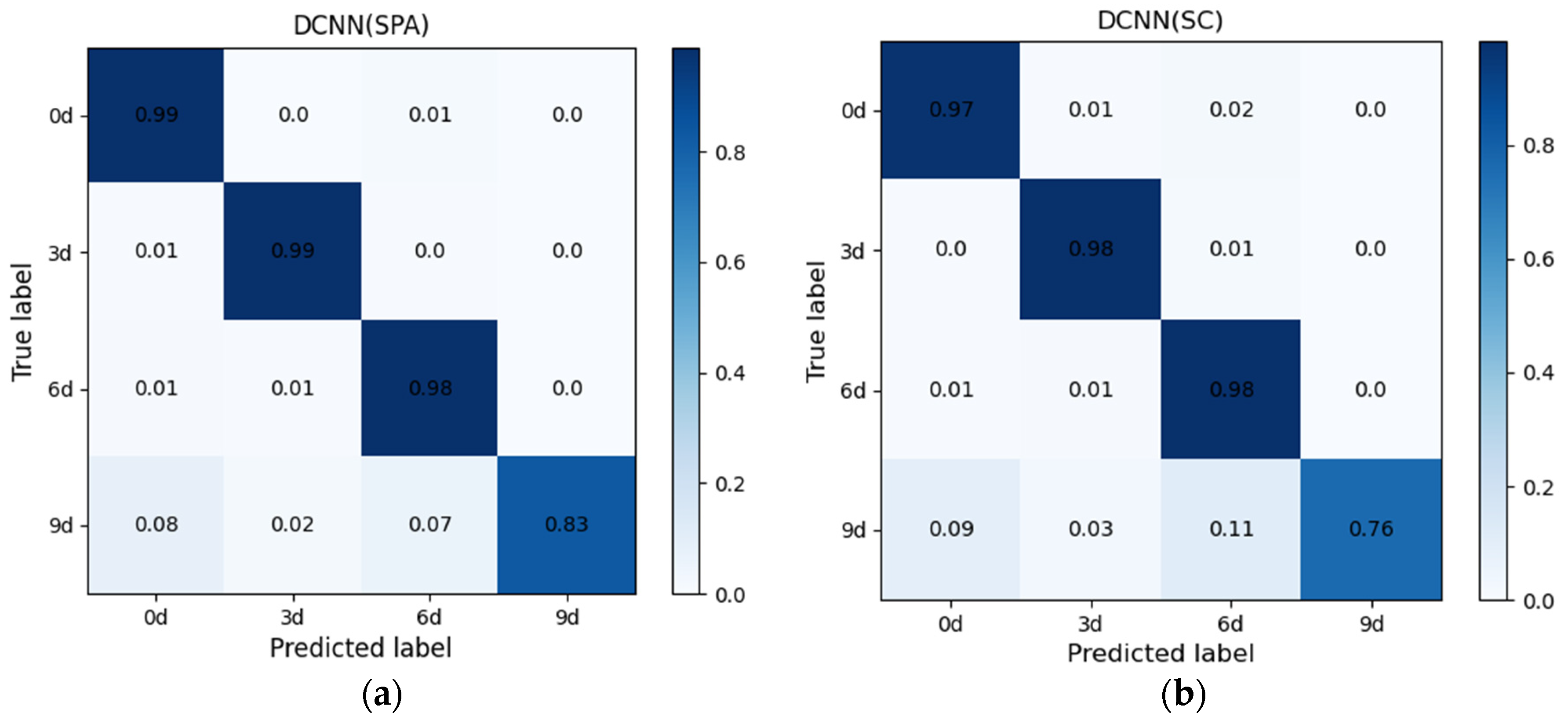
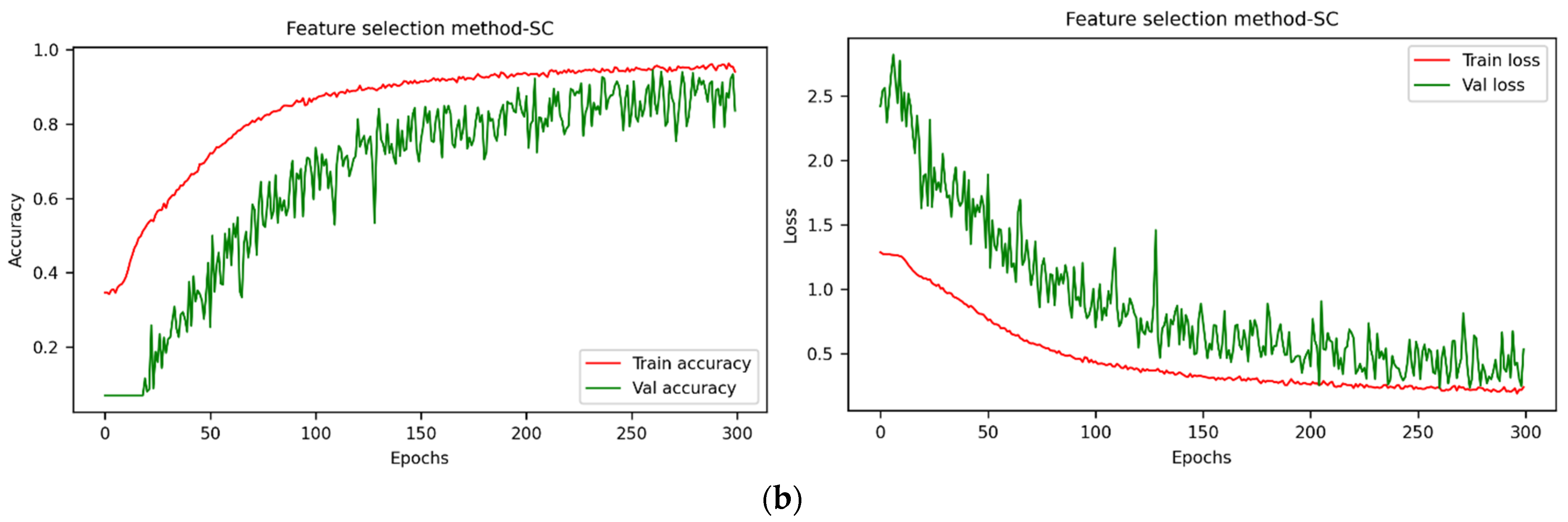
3.4. Classification Results and Analysis of the Viability Detection Model Based on Optimal Band Spectra
3.5. Seed Viability Prediction Based on the Optimal Model
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- De Bittencourt, S.R.M.; Grzybowski, C.R.D.S.; Panobianco, M.; Vieira, R.D. Metodologia alternativa para condução do teste de envelhecimento acelerado em sementes de milho. Ciênc. Rural 2012, 42, 1360–1365. [Google Scholar] [CrossRef]
- Williams, P.; Manley, M.; Fox, G.; Geladi, P. Indirect Detection of Fusarium Verticillioides in Maize (Zea mays L.) Kernels by near Infrared Hyperspectral Imaging. J. Near Infrared Spectrosc. 2010, 18, 49–58. [Google Scholar] [CrossRef]
- Ambrose, A.; Kandpal, L.M.; Kim, M.S.; Lee, W.-H.; Cho, B.-K. High speed measurement of corn seed viability using hyperspectral imaging. Infrared Phys. Technol. 2016, 75, 173–179. [Google Scholar] [CrossRef]
- Sena, D.V.D.A.; Alves, E.U.; De Medeiros, D.S. Vigor tests to evaluate the physiological quality of corn seeds cv. ‘Sertanejo’. Cienci Rural 2017, 47, 1678. [Google Scholar] [CrossRef]
- Fessel, S.A.; Panobianco, M.; Vieira, R.D.; Da Cruz, M.C.P.; De Paula, R.C. Electrical conductivity testing of corn seeds as influenced by temperature and period of storage. Pesqui. Agropecu. Bras. 2006, 41, 1551–1559. [Google Scholar] [CrossRef]
- Song, L.; Wang, Q.; Wang, C.; Lin, Y.; Yu, D.; Xu, Z.; Huang, Q.; Wu, Y. Effect of γ-irradiation on rice seed vigor assessed by near-infrared spectroscopy. J. Stored Prod. Res. 2015, 62, 46–51. [Google Scholar] [CrossRef]
- Via, F.D.J.S.F.I.; Ivano, A.D.; Raniele, T.G.A.E.D.S.; Itamar, R.T.; Carlos, R.S. Physiological quality of quinoa seeds submitted to different storage conditions. Afr. J. Agric. Res. 2016, 11, 1299–1308. [Google Scholar] [CrossRef]
- Nansen, C.; Zhao, G.; Dakin, N.; Zhao, C.; Turner, S.R. Using hyperspectral imaging to determine germination of native Australian plant seeds. J. Photochem. Photobiol. B Biol. 2015, 145, 19–24. [Google Scholar] [CrossRef]
- Ropodi, A.; Panagou, E.; Nychas, G.-J. Data mining derived from food analyses using non-invasive/non-destructive analytical techniques; determination of food authenticity, quality & safety in tandem with computer science disciplines. Trends Food Sci. Technol. 2016, 50, 11–25. [Google Scholar] [CrossRef]
- Xia, J.; Cao, H.; Yang, Y.; Zhang, W.; Wan, Q.; Xu, L.; Ge, D.; Zhang, W.; Ke, Y.; Huang, B. Detection of waterlogging stress based on hyperspectral images of oilseed rape leaves (Brassica napus L.). Comput. Electron. Agric. 2019, 159, 59–68. [Google Scholar] [CrossRef]
- Díaz, J.J.V.; Aldana, A.P.S.; Zuluaga, D.V.R. Prediction of dry matter content of recently harvested ‘Hass’ avocado fruits using hyperspectral imaging. J. Sci. Food Agric. 2020, 101, 897–906. [Google Scholar] [CrossRef]
- Tian, X.; Fan, S.X.; Huang, W.Q.; Wang, Z.L.; Li, J.B. Detection of early decay on citrus using hyperspectral transmittance imaging technology coupled with principal component analysis and improved watershed segmentation algorithms. Postharvest Biol. Tech. 2020, 161, 111071–111079. [Google Scholar] [CrossRef]
- Ren, G.; Wang, Y.; Ning, J.; Zhang, Z. Evaluation of Dianhong black tea quality using near-infrared hyperspectral imaging technology. J. Sci. Food Agric. 2020, 101, 2135–2142. [Google Scholar] [CrossRef]
- Hacisalihoglu, G.; Freeman, J.; Armstrong, P.R.; Seabourn, B.W.; Porter, L.D.; Settles, A.M.; Gustin, J.L. Protein, weight, and oil prediction by single-seed near-infrared spectroscopy for selection of seed quality and yield traits in pea (Pisum sativum). J. Sci. Food Agric. 2020, 100, 3488–3497. [Google Scholar] [CrossRef]
- Bai, X.; Zhang, C.; Xiao, Q.; He, Y.; Bao, Y. Application of near-infrared hyperspectral imaging to identify a variety of silage maize seeds and common maize seeds. RSC Adv. 2020, 10, 11707–11715. [Google Scholar] [CrossRef]
- Femenias, A.; Gatius, F.; Ramos, A.J.; Sanchis, V.; Marín, S. Standardisation of near infrared hyperspectral imaging for quantification and classification of DON contaminated wheat samples. Food Control 2019, 111, 107074. [Google Scholar] [CrossRef]
- Kandpal, L.M.; Lohumi, S.; Kim, M.S.; Kang, J.-S.; Cho, B.-K. Near-infrared hyperspectral imaging system coupled with multivariate methods to predict viability and vigor in muskmelon seeds. Sens. Actuators B Chem. 2016, 229, 534–544. [Google Scholar] [CrossRef]
- Li, Y.; Sun, J.; Wu, X.; Chen, Q.; Lu, B.; Dai, C. Detection of viability of soybean seed based on fluorescence hyperspectra and CARS-SVM-AdaBoost model. J. Food Process. Preserv. 2019, 43, e14238. [Google Scholar] [CrossRef]
- Zhang, J.; Dai, L.; Cheng, F. Classification of Frozen Corn Seeds Using Hyperspectral VIS/NIR Reflectance Imaging. Molecules 2019, 24, 149. [Google Scholar] [CrossRef]
- Kamilaris, A.; Prenafeta-Boldú, F.X. Deep learning in agriculture: A survey. Comput. Electron. Agric. 2018, 147, 70–90. [Google Scholar] [CrossRef]
- Shen, D.; Wu, G.; Suk, H.-I. Deep Learning in Medical Image Analysis. Annu. Rev. Biomed. Eng. 2017, 19, 221–248. [Google Scholar] [CrossRef]
- Zhu, S.; Zhou, L.; Gao, P.; Bao, Y.; He, Y.; Feng, L. Near-Infrared Hyperspectral Imaging Combined with Deep Learning to Identify Cotton Seed Varieties. Molecules 2019, 24, 3268. [Google Scholar] [CrossRef]
- Feng, L.; Zhu, S.; Zhang, C.; Bao, Y.; Feng, X.; He, Y. Identification of Maize Kernel Vigor under Different Accelerated Aging Times Using Hyperspectral Imaging. Molecules 2018, 23, 3078. [Google Scholar] [CrossRef]
- Zhang, T.; Wei, W.; Zhao, B.; Wang, R.; Li, M.; Yang, L.; Wang, J.; Sun, Q. A Reliable Methodology for Determining Seed Viability by Using Hyperspectral Data from Two Sides of Wheat Seeds. Sensors 2018, 18, 813. [Google Scholar] [CrossRef]
- GB 5009.3-2016; Determination of Moisture in Food. General Administration of Quality Supervision. Inspection and Quarantine of the People’s Republic of China: Beijing, China, 2016; Volume 1, pp. 1–2.
- GB 5009.6-2016; Determination of Fat in Food. General Administration of Quality Supervision. Inspection and Quaran-tine of the People’s Republic of China: Beijing, China, 2016; Volume 1, pp. 1–2.
- GB 5009.5-2016; Determination of Protein in Food. General Administration of Quality Supervision. Inspection and Quarantine of the People’s Republic of China: Beijing, China, 2016; Volume 1, pp. 1–5.
- GB 5009.9-2016; Determination of Starch in Food. General Administration of Quality Supervision. Inspection and Quarantine of the People’s Republic of China: Beijing, China, 2016; Volume 1, pp. 1–2.
- Huo, Z.J.; Pan, X.L.; Li, J.Y. Problems and countermeasures in the operation of GB/T3543-1995 rules for agricultural seed testing. Modern Agri. 2004, 1, 14–15. [Google Scholar] [CrossRef]
- ISTA. International Rules for Seed Testing, International Seed Testing Association Chapter 1:1-12; ISTA: Palmerston North, New Zealand, 2015. [Google Scholar]
- Ru, C.; Li, Z.; Tang, R. A Hyperspectral Imaging Approach for Classifying Geographical Origins of Rhizoma Atractylodis Macrocephalae Using the Fusion of Spectrum-Image in VNIR and SWIR Ranges (VNIR-SWIR-FuSI). Sensors 2019, 19, 2045. [Google Scholar] [CrossRef]
- Wu, D.; He, Y.; Feng, S.; Sun, D.-W. Study on infrared spectroscopy technique for fast measurement of protein content in milk powder based on LS-SVM. J. Food Eng. 2008, 84, 124–131. [Google Scholar] [CrossRef]
- Gerhardt, N.; Schwolow, S.; Rohn, S.; Pérez-Cacho, P.R.; Galán-Soldevilla, H.; Arce, L.; Weller, P. Quality assessment of olive oils based on temperature-ramped HS-GC-IMS and sensory evaluation: Comparison of different processing approaches by LDA, kNN, and SVM. Food Chem. 2019, 278, 720–728. [Google Scholar] [CrossRef]
- Qiu, G.; Lü, E.; Wang, N.; Lu, H.; Wang, F.; Zeng, F. Cultivar Classification of Single Sweet Corn Seed Using Fourier Transform Near-Infrared Spectroscopy Combined with Discriminant Analysis. Appl. Sci. 2019, 9, 1530. [Google Scholar] [CrossRef]
- Fraiwan, L.; Lweesy, K.; Khasawneh, N.; Wenz, H.; Dickhaus, H. Automated sleep stage identification system based on time–frequency analysis of a single EEG channel and random forest classifier. Comput. Methods Programs Biomed. 2012, 108, 10–19. [Google Scholar] [CrossRef]
- Jiao, C.; Chen, C.; McGarvey, R.G.; Bohlman, S.; Jiao, L.; Zare, A. Multiple instance hybrid estimator for hyperspectral target characterization and sub-pixel target detection. ISPRS J. Photogramm. Remote. Sens. 2018, 146, 235–250. [Google Scholar] [CrossRef]
- Chen, L.-C.; Papandreou, G.; Kokkinos, I.; Murphy, K.; Yuille, A.L. DeepLab: Semantic Image Segmentation with Deep Convolutional Nets, Atrous Convolution, and Fully Connected CRFs. IEEE Trans. Pattern Anal. Mach. Intell. 2018, 40, 834–848. [Google Scholar] [CrossRef]
- Zhong, L.; Guo, X.; Xu, Z.; Ding, M. Soil properties: Their prediction and feature extraction from the LUCAS spectral library using deep convolutional neural networks. Geoderma 2021, 402, 115366. [Google Scholar] [CrossRef]
- Li, J.; Tian, X.; Huang, W.; Zhang, B.; Fan, S. Application of long-wave near infrared hyperspectral imaging for measurement of color distribution in salmon fillet. Food Anal. Methods 2016, 9, 3087–3098. [Google Scholar] [CrossRef]
- Williams, P.J.; Kucheryavskiy, S. Classification of maize kernels using NIR hyperspectral imaging. Food Chem. 2016, 209, 131–138. [Google Scholar] [CrossRef]
- Liu, D.; Sun, D.-W.; Zeng, X.-A. Recent Advances in Wavelength Selection Techniques for Hyperspectral Image Processing in the Food Industry. Food Bioprocess Technol. 2013, 7, 307–323. [Google Scholar] [CrossRef]
- Peng, Y.K.; Zhao, F.; Li, L.; Xing, Y.Y.; Fang, X.Q. Discrimination of heat-damaged tomato seeds based on near infrared spectroscopy and PCA-SVM method. Trans. Chin. Soc. Agric. Eng. 2018, 34, 159–165. [Google Scholar] [CrossRef]
- Wu, N.; Zhang, Y.; Na, R.; Mi, C.; Zhu, S.; He, Y.; Zhang, C. Variety identification of oat seeds using hyperspectral imaging: Investigating the representation ability of deep convolutional neural network. RSC Adv. 2019, 9, 12635–12644. [Google Scholar] [CrossRef]
- Zhu, S.; Zhou, L.; Zhang, C.; Bao, Y.; Wu, B.; Chu, H.; Yu, Y.; He, Y.; Feng, L. Identification of Soybean Varieties Using Hyperspectral Imaging Coupled with Convolutional Neural Network. Sensors 2019, 19, 4065. [Google Scholar] [CrossRef]
- Fu, J.R. Seed Physiology; Science Press: Beijing, China, 1985. [Google Scholar]
- Taiz, L.; Jones, R.L. Gibberellic acid, beta-1,3-glucanase and the cell walls of barley aleurone layers. Planta 1970, 92, 73–84. [Google Scholar] [CrossRef]
- Hara-Nishimura, I.; Nishimura, M.; Daussant, J. Conversion of free β-amylase to bound β-amylase on starch granules in the barley endosperm during desiccation phase of seed development. Protoplasma 1986, 134, 149–153. [Google Scholar] [CrossRef]
- Yong-Qiang, M.A.; Han, C.R. Changes of Starch in Corn during Germination; Grain Processing: Muscatine County, IA, USA, 2007. [Google Scholar]
- Chavan, J.K.; Kadam, S.S.; Salunkhe, D.K. Changes in Tannin, Free Amino Acids, Reducing Sugars, and Starch During Seed Germination of Low and High Tannin Cultivars of Sorghum. J. Food Sci. 1981, 46, 638–639. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, Y.; Song, J.; Chen, X.; Lu, Y.; Wang, W. Pearson correlation coefficient of current derivatives based pilot protection scheme for long-distance LCC-HVDC transmission lines. Int. J. Electr. Power Energy Syst. 2020, 116, 105526. [Google Scholar] [CrossRef]
- Yang, G.; Wang, Q.; Liu, C.; Wang, X.; Fan, S.; Huang, W. Rapid and visual detection of the main chemical compositions in maize seeds based on Raman hyperspectral imaging. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2018, 200, 186–194. [Google Scholar] [CrossRef]
- Feng, X.; Peng, C.; Chen, Y.; Liu, X.; Feng, X.; He, Y. Discrimination of CRISPR/Cas9-induced mutants of rice seeds using near-infrared hyperspectral imaging. Sci. Rep. 2017, 7, 15934–15943. [Google Scholar] [CrossRef]



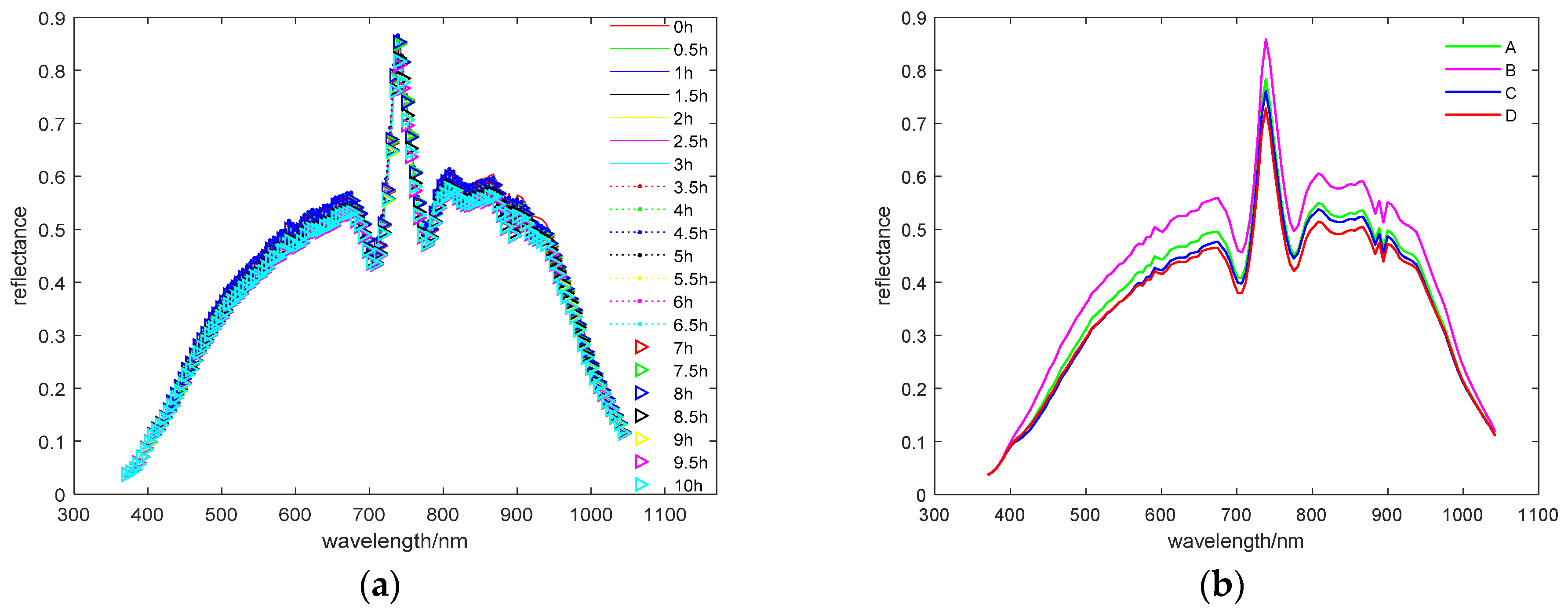



| Aging Time (d) | Vitality Level | Number of Samples | Viable Seeds | GP (%) | s (cm) | GI | VI | AGD (d) |
|---|---|---|---|---|---|---|---|---|
| 0 | A | 192 | 172 | 89.6 | 2.59 | 41.46 | 107.38 | 4.7 |
| 3 | B | 192 | 180 | 93.8 | 2.63 | 49.99 | 131.47 | 3.9 |
| 6 | C | 192 | 114 | 59.4 | 1.10 | 26.03 | 28.63 | 5 |
| 9 | D | 192 | 33 | 17.2 | 0.21 | 6.45 | 1.35 | 5.5 |
| Total | 768 | 499 | 65 | 1.63 | 30.98 | 67.21 | 4.8 |
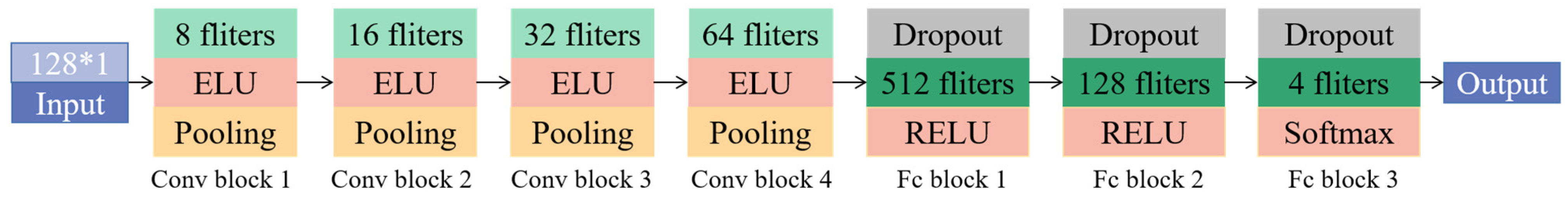
| Layer (Type) | Output Shape | Param |
|---|---|---|
| Input | (128, 1) | 0 |
| Conv1D_1 | (128, 8) | 32 |
| MaxPooling1D_1 | (64, 8) | 0 |
| Conv1D_2 | (64, 16) | 400 |
| MaxPooling1D_2 | (32, 16) | 0 |
| Conv1D_3 | (32, 32) | 1568 |
| MaxPooling1D_3 | (16, 32) | 0 |
| Conv1D_4 | (16, 64) | 6208 |
| MaxPooling1D_4 | (8, 64) | 0 |
| Flatten | (512) | 0 |
| Dropout_1 | (512) | 0 |
| Dense_1 | (512) | 262,656 |
| Dropout_2 | (512) | 0 |
| Dense_2 | (128) | 65,664 |
| Ddropout_2 | (128) | 0 |
| Dense_3 | (4) | 516 |
| Total params | / | 337,044 |
| Number of Levels Included | Discriminant Model | Main Parameters | Training Set Accuracy | Test Set Accuracy | Time (s) |
|---|---|---|---|---|---|
| 2 | SVM | c = 2.0, g = 4.1 | 100% | 96.24% | 8.42 |
| KNN | K = 3 | 99.52% | 93.5% | 7.55 | |
| RF | mtry = 11 | 100% | 93.3% | 7.81 | |
| DCNN | lr = 10−4, batchsize = 64 | 100% | 98.20% | 369.81 | |
| 3 | SVM | c = 3.0, g = 3.5 | 97.4% | 95.30% | 75.88 |
| KNN | K = 5 | 98.37% | 92.77% | 9.96 | |
| RF | mtry = 11 | 100% | 89.13% | 13.16 | |
| DCNN | lr = 10−4, batchsize = 64 | 100% | 98.35% | 554.98 | |
| 4 | SVM | c = 3.1, g = 4.7 | 94.99% | 92.34% | 191.48 |
| KNN | K = 7 | 95.38% | 86.95% | 15.07 | |
| RF | mtry = 11 | 100% | 85.46% | 17.41 | |
| DCNN | lr = 10−4, batchsize = 64 | 99.98% | 97.47% | 608.61 |
| Sample Vitality Level | Content of Main Ingredients (g/100 g) | |||
|---|---|---|---|---|
| Moisture | Fat | Protein | Starch | |
| Normal | 9.70 ± 0.11 a | 4.20 ± 0.03 c | 11.1 ± 0.26 a | 65.6 ± 1.25 a |
| Aging for 3 days | 8.68 ± 0.07 b | 4.30 ± 0.06 b | 10.3 ± 0.10 c | 36.0 ± 0.81 c |
| Aging for 6 days | 8.10 ± 0.10 c | 4.60 ± 0.11 a | 10.6 ± 0.10 b | 34.7 ± 0.70 c |
| Aging for 9 days | 7.77 ± 0.24 c | 4.50 ± 0.08 a | 10.0 ± 0.03 d | 43.3 ± 2.95 b |
| Method of Choosing | Number of Bands | Wavelength (nm) |
|---|---|---|
| SC | 43 | 717.2, 722.5, 727.8, 733.2, 738.5, 743.8, 781.2, 786.5, 791.9, 797.2, 802.6, 807.9, 813.3, 818.7, 824.1, 829.5, 834.8, 840.2, 845.6, 851.0, 856.4, 861.8, 867.2, 872.6, 878.1, 883.5, 888.9, 894.3, 899.8, 905.2, 910.6, 916.1, 921.5, 926.9, 932.4, 937.9, 943.3, 948.8, 954.3, 1025.7, 1031.2, 1036.8, 1042.3 |
| SPA | 51 | 410.8, 451.5, 487.4, 492.6, 502.9, 508.0, 513.2, 528.7, 533.8, 539.0, 544.2, 549.4, 554.6, 575.4, 580.6, 585.8, 596.2, 606.6, 638.1, 643.3, 653.8, 659.1, 664.4, 685.5, 706.6, 711.9, 733.2, 743.8, 754.5, 759.8, 765.1, 775.8, 781.2, 786.5, 791.9, 797.2, 807.9, 813.3, 818.7, 834.8, 856.4, 888.9, 890.9, 905.2, 910.6, 932.4, 937.9, 943.3, 948.8, 970.7, 976.2 |
| Discriminant Model | Feature Selection Method (Number) | Main Parameters | Training Set Accuracy | Test Set Accuracy | Time (s) |
|---|---|---|---|---|---|
| SVM | SPA (51) | c = 1.7, g = 3.5 | 93.4% | 86.56% | 122.44 |
| SC (43) | c = 1.3, g = 1.8 | 82.92% | 79.41% | 144.51 | |
| KNN | SPA (51) | K = 4 | 95.34% | 86.31% | 5.22 |
| SC (43) | K = 4 | 89.07% | 77.34% | 5.16 | |
| RF | SPA (51) | mtry = 7 | 100% | 86.6% | 8.51 |
| SC (43) | mtry = 6 | 100% | 83.28% | 7.55 | |
| DCNN | SPA (51) | lr = 10−4, batchsize = 32 | 99.74% | 94.25% | 509.32 |
| SC (43) | lr = 10−4, batchsize = 32 | 99.57% | 91.16% | 466.34 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, X.; Pang, L.; Wang, L.; Men, S.; Yan, L. Deep Convolutional Neural Network for Detection and Prediction of Waxy Corn Seed Viability Using Hyperspectral Reflectance Imaging. Math. Comput. Appl. 2022, 27, 109. https://doi.org/10.3390/mca27060109
Zhao X, Pang L, Wang L, Men S, Yan L. Deep Convolutional Neural Network for Detection and Prediction of Waxy Corn Seed Viability Using Hyperspectral Reflectance Imaging. Mathematical and Computational Applications. 2022; 27(6):109. https://doi.org/10.3390/mca27060109
Chicago/Turabian StyleZhao, Xiaoqing, Lei Pang, Lianming Wang, Sen Men, and Lei Yan. 2022. "Deep Convolutional Neural Network for Detection and Prediction of Waxy Corn Seed Viability Using Hyperspectral Reflectance Imaging" Mathematical and Computational Applications 27, no. 6: 109. https://doi.org/10.3390/mca27060109
APA StyleZhao, X., Pang, L., Wang, L., Men, S., & Yan, L. (2022). Deep Convolutional Neural Network for Detection and Prediction of Waxy Corn Seed Viability Using Hyperspectral Reflectance Imaging. Mathematical and Computational Applications, 27(6), 109. https://doi.org/10.3390/mca27060109





