How to Tackle Underdeterminacy in Metabolic Flux Analysis? A Tutorial and Critical Review
Abstract
1. Introduction
- Dealing with the underdeterminacy—this strategy is adopted in several methods where minimal and maximal bounds on the admissible fluxes are determined. This category of methods includes Flux Pathway Analysis (FPA), where convex analysis is used to decompose the admissible flux distributions into Elementary Flux Modes (EFMs) or Extreme Pathways [3,4], Flux Variability Analysis (FVA), which is a Linear-Programming (LP)-based method determining the range of admissible fluxes [5], Flux Spectrum Approach (FSA), which is another LP-based method taking insufficient and uncertain measurements into account [6]. Random sampling of the admissible solution set allows determining the marginal probability density functions of the fluxes [7,8,9,10], and statistical methods based on the maximum entropy principle can be used to infer intracellular flux distributions [11,12].
- Reducing or eliminating the underdeterminacy—this strategy consists in adding constraints in various ways, e.g., including more measurements of the extracellular fluxes or, possibly, measurements of the intracellular fluxes using specific techniques such as tracing [13,14] and parallel labeling [15], leading to the sophisticated procedures of MFA. Alternatively, additional constraints can be introduced by formulating biological assumptions either based on prior knowledge and/or experimental observations [16,17] or systematic procedures to determine active constraints [18]. The use of thermodynamic constraints can be important in relation with reaction reversibility and the limitation of the solution space [19]. Moreover, thermodynamical constraints can prevent infeasible loops in a metabolic network as demonstrated in [20]. Underdeterminacy can also be reduced (or even eliminated) through the formulation of an optimization problem originating from the assumption of an optimal metabolic behavior of the cells. This approach corresponds to Flux Balance Analysis (FBA) [21,22], which uses an objective function expressed as a linear combination of selected fluxes. Recently, the increasing availability of metabolite profiling data obtained through gas and liquid chromatography combined with mass spectroscopy has also allowed the integration of time-course absolute quantitative metabolomics in unsteady-state (or dynamic) FBA [23,24]. In the usual situation where FBA still leads to an underdetermined system with an infinite number of flux distributions that optimize the cost function, variants of FBA have been proposed in order to define a unique solution, e.g., the geometric approach developed in [25] that searches for the minimal flux distribution satisfying the given objective. Assuming that fluxes correlate with enzyme levels, this specific flux distribution would correspond to the minimization of the amount of enzymes required to satisfy the objective defined in FBA. Ultimately, the concept of Most Accurate Fluxes [26] allows computing a unique flux distribution, hence eliminating the system underdeterminacy, with a very low computational load and without any assumption regarding an optimal biological behavior.
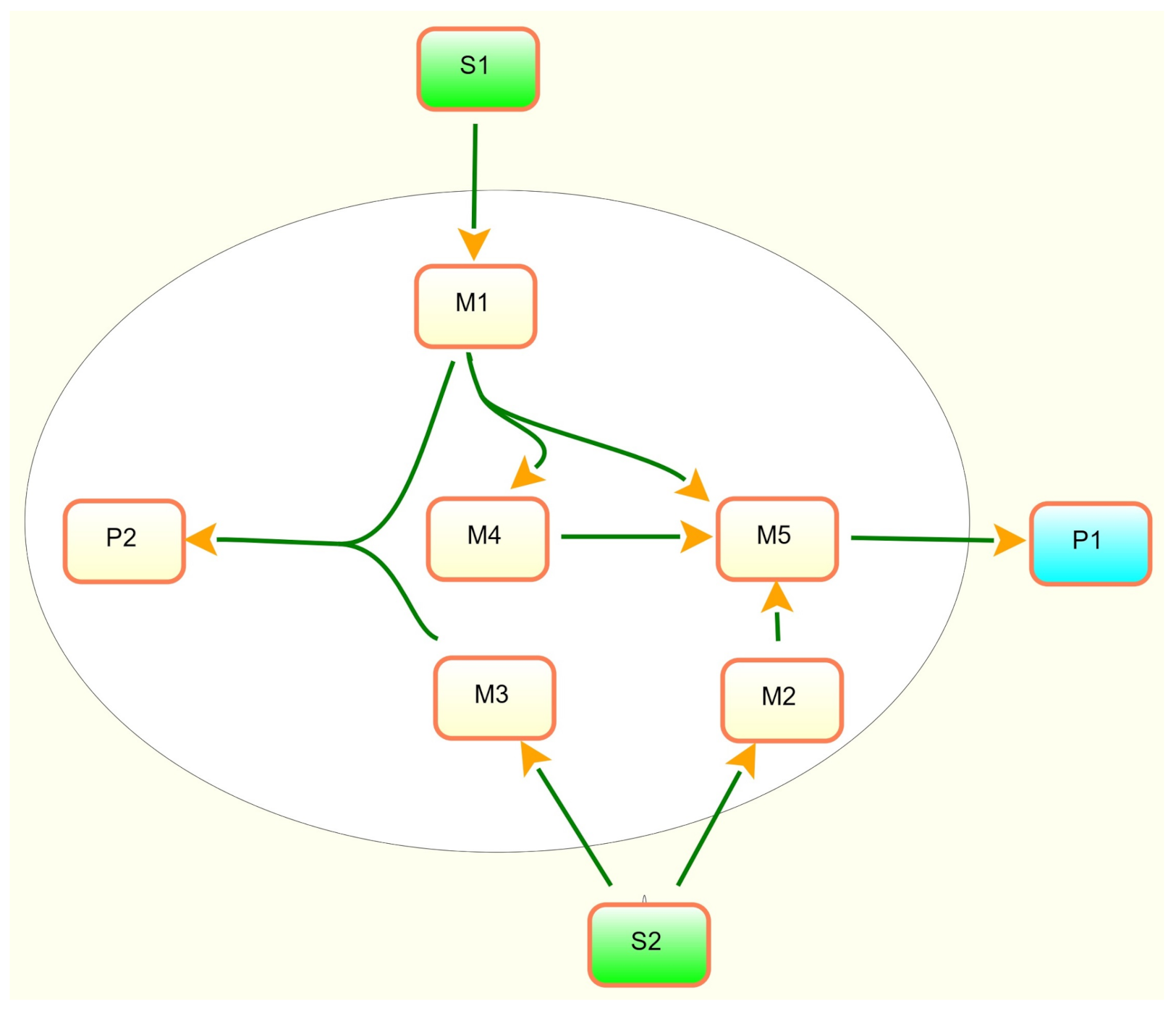
2. A Toy Example
3. Dealing with the Underdeterminacy
4. Reducing or Eliminating the Underdeterminacy
5. An Overview of Important Topics
5.1. How to Select the Size/Detail of the Metabolic Network?
5.2. Dynamic Metabolic Flux Interval Analysis
5.3. How to Represent the Accumulation of Internal Metabolites?
5.4. Model Reduction to Macroscopic Scale
5.5. How to Handle the Measurement Errors?
5.6. Some Further Perspectives on Sampling Algorithms
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Stephanopoulos, G.; Aristidou, A.A.; Nielsen, J. Metabolic Engineering: Principles and Methodologies; Elsevier Science: San Diego, CA, USA, 1998. [Google Scholar]
- Stelling, J.; Klamt, S.; Bettenbrock, K.; Schuster, S.; Gilles, E.D. Metabolic network structure determines key aspects of functionality and regulation. Nature 2002, 420, 190–193. [Google Scholar] [CrossRef]
- Schuster, S.; Hilgetag, C. On elementary flux modes in biochemical reaction systems at steady state. J. Biol. Syst. 1994, 2, 165–182. [Google Scholar] [CrossRef]
- Klamt, S.; Stelling, J. Two approaches for metabolic pathway analysis? Trends Biotechnol. 2003, 21, 64–69. [Google Scholar] [CrossRef]
- Mahadevan, R.; Schilling, C. The effects of alternate optimal solutions in constraint-based genome-scale metabolic models. Metab. Eng. 2003, 5, 264–276. [Google Scholar] [CrossRef] [PubMed]
- Llaneras, F.; Picó, J. An interval approach for dealing with flux distributions and elementary modes activity patterns. J. Theor. Biol. 2007, 246, 290–308. [Google Scholar] [CrossRef] [PubMed]
- Thiele, I.; Price, N.D.; Vo, T.D.; Palsson, B.Ø. Candidate metabolic network states in human mitochondria impact of diabetes, ischemia, and diet. J. Biol. Chem. 2005, 280, 11683–11695. [Google Scholar] [CrossRef]
- Saa, P.A.; Nielsen, L.K. ll-ACHRB: A scalable algorithm for sampling the feasible solution space of metabolic networks. Bioinformatics 2016, 32, 2330–2337. [Google Scholar] [CrossRef] [PubMed]
- Megchelenbrink, W.; Huynen, M.; Marchiori, E. optGpSampler: An improved tool for uniformly sampling the solution-space of genome-scale metabolic networks. PLoS ONE 2014, 9, e86587. [Google Scholar] [CrossRef] [PubMed]
- Haraldsdóttir, H.S.; Cousins, B.; Thiele, I.; Fleming, R.M.; Vempala, S. CHRR: Coordinate hit-and-run with rounding for uniform sampling of constraint-based models. Bioinformatics 2017, 33, 1741–1743. [Google Scholar] [CrossRef] [PubMed]
- De Martino, A.; De Martino, D. An introduction to the maximum entropy approach and its application to inference problems in biology. Heliyon 2018, 4, e00596. [Google Scholar] [CrossRef]
- De Martino, D.; Andersson, A.M.; Bergmiller, T.; Guet, C.C.; Tkačik, G. Statistical mechanics for metabolic networks during steady state growth. Nat. Commun. 2018, 9, 2988. [Google Scholar] [CrossRef] [PubMed]
- Ahn, W.S.; Antoniewicz, M.R. Metabolic flux analysis of CHO cells at growth and non-growth phases using isotopic tracers and mass spectrometry. Metab. Eng. 2011, 13, 598–609. [Google Scholar] [CrossRef] [PubMed]
- Long, C.P.; Antoniewicz, M.R. High-resolution 13 C metabolic flux analysis. Nat. Protoc. 2019, 14, 2856–2877. [Google Scholar] [CrossRef]
- Crown, S.B.; Antoniewicz, M.R. Parallel labeling experiments and metabolic flux analysis: Past, present and future methodologies. Metab. Eng. 2013, 16, 21–32. [Google Scholar] [CrossRef]
- Richelle, A.; Gziri, K.M.; Bogaerts, P. A methodology for building a macroscopic FBA-based dynamical simulator of cell cultures through flux variability analysis. Biochem. Eng. J. 2016, 114, 50–64. [Google Scholar] [CrossRef]
- Bogaerts, P.; Gziri, K.M.; Richelle, A. From MFA to FBA: Defining linear constraints accounting for overflow metabolism in a macroscopic FBA-based dynamical model of cell cultures in bioreactor. J. Process Control 2017, 60, 34–47. [Google Scholar] [CrossRef]
- Nikdel, A.; Braatz, R.D.; Budman, H.M. A systematic approach for finding the objective function and active constraints for dynamic flux balance analysis. Bioprocess Biosyst. Eng. 2018, 41, 641–655. [Google Scholar] [CrossRef] [PubMed]
- Soh, K.C.; Hatzimanikatis, V. Constraining the flux space using thermodynamics and integration of metabolomics data. In Metabolic Flux Analysis; Springer Science + Business Media: New York, NY, USA, 2014; pp. 49–63. [Google Scholar]
- De Martino, D. Thermodynamics of biochemical networks and duality theorems. Phys. Rev. E 2013, 87, 052108. [Google Scholar] [CrossRef]
- Raman, K.; Chandra, N. Flux balance analysis of biological systems: Applications and challenges. Br. Bioinform. 2009, 10, 435–449. [Google Scholar] [CrossRef]
- Orth, J.D.; Thiele, I.; Palsson, B.Ø. What is flux balance analysis? Nat. Biotechnol. 2010, 28, 245–248. [Google Scholar] [CrossRef]
- Willemsen, A.M.; Hendrickx, D.M.; Hoefsloot, H.C.; Hendriks, M.M.; Wahl, S.A.; Teusink, B.; Smilde, A.K.; van Kampen, A.H. MetDFBA: Incorporating time-resolved metabolomics measurements into dynamic flux balance analysis. Mol. BioSyst. 2015, 11, 137–145. [Google Scholar] [CrossRef] [PubMed]
- Bordbar, A.; Yurkovich, J.T.; Paglia, G.; Rolfsson, O.; Sigurjónsson, Ó.E.; Palsson, B.O. Elucidating dynamic metabolic physiology through network integration of quantitative time-course metabolomics. Sci. Rep. 2017, 7, 46249. [Google Scholar] [CrossRef] [PubMed]
- Smallbone, K.; Simeonidis, E. Flux balance analysis: A geometric perspective. J. Theor. Biol. 2009, 258, 311–315. [Google Scholar] [CrossRef] [PubMed]
- Gziri, K.M.; Bogaerts, P. Determining a unique solution to underdetermined metabolic networks via a systematic path through the Most Accurate Fluxes. IFAC-PapersOnLine 2019, 52, 352–357. [Google Scholar] [CrossRef]
- Zanghellini, J.; Ruckerbauer, D.E.; Hanscho, M.; Jungreuthmayer, C. Elementary flux modes in a nutshell: Properties, calculation and applications. Biotechnol. J. 2013, 8, 1009–1016. [Google Scholar] [CrossRef]
- Pfeiffer, T.; Sanchez-Valdenebro, I.; Nuno, J.; Montero, F.; Schuster, S. METATOOL: For studying metabolic networks. Bioinformatics 1999, 15, 251–257. [Google Scholar] [CrossRef]
- Terzer, M.; Stelling, J. Large-scale computation of elementary flux modes with bit pattern trees. Bioinformatics 2008, 24, 2229–2235. [Google Scholar] [CrossRef] [PubMed]
- Van Klinken, J.B.; Willems van Dijk, K. FluxModeCalculator: An efficient tool for large-scale flux mode computation. Bioinformatics 2016, 32, 1265–1266. [Google Scholar] [CrossRef][Green Version]
- Jungers, R.M.; Zamorano, F.; Blondel, V.D.; Vande Wouwer, A.; Bastin, G. Fast computation of minimal elementary decompositions of metabolic flux vectors. Automatica 2011, 47, 1255–1259. [Google Scholar] [CrossRef]
- Provost, A.; Bastin, G. Dynamic metabolic modelling under the balanced growth condition. J. Process Control 2004, 14, 717–728. [Google Scholar] [CrossRef]
- Zamorano, F.; Vande Wouwer, A.; Jungers, R.M.; Bastin, G. Dynamic metabolic models of CHO cell cultures through minimal sets of elementary flux modes. J. Biotechnol. 2013, 164, 409–422. [Google Scholar] [CrossRef] [PubMed]
- Fernandes de Sousa, S.; Bastin, G.; Jolicoeur, M.; Vande Wouwer, A. Dynamic metabolic flux analysis using a convex analysis approach: Application to hybridoma cell cultures in perfusion. Biotechnol. Bioeng. 2016, 113, 1102–1112. [Google Scholar] [CrossRef] [PubMed]
- Provost, A. Metabolic Design of Dynamic Bioreaction Models; Faculté des Sciences Appliquées, Université Catholique de Louvain: Louvain-la-Neuve, Belgium, 2006. [Google Scholar]
- Becker, S.A.; Feist, A.M.; Mo, M.L.; Hannum, G.; Palsson, B.Ø.; Herrgard, M.J. Quantitative prediction of cellular metabolism with constraint-based models: The COBRA Toolbox. Nat. Protoc. 2007, 2, 727–738. [Google Scholar] [CrossRef] [PubMed]
- Klamt, S.; Saez-Rodriguez, J.; Gilles, E.D. Structural and functional analysis of cellular networks with CellNetAnalyzer. BMC Syst. Biol. 2007, 1, 1–13. [Google Scholar] [CrossRef]
- Abbate, T.; Dewasme, L.; Vande Wouwer, A.; Bogaerts, P. Adaptive flux variability analysis of HEK cell cultures. Comput. Chem. Eng. 2020, 133, 106633. [Google Scholar] [CrossRef]
- Bogaerts, P.; Rooman, M. DISCOPOLIS: An algorithm for uniform sampling of metabolic flux distributions via iterative sequences of linear programs. IFAC-PapersOnLine 2019, 52, 269–274. [Google Scholar] [CrossRef]
- Rubinstein, R. Generating random vectors uniformly distributed inside and on the surface of different regions. Eur. J. Oper. Res. 1982, 10, 205–209. [Google Scholar] [CrossRef]
- Herrmann, H.A.; Dyson, B.C.; Vass, L.; Johnson, G.N.; Schwartz, J.M. Flux sampling is a powerful tool to study metabolism under changing environmental conditions. NPJ Syst. Biol. Appl. 2019, 5, 1–8. [Google Scholar] [CrossRef]
- Fallahi, S.; Skaug, H.J.; Alendal, G. A comparison of Monte Carlo sampling methods for metabolic network models. PLoS ONE 2020, 15, e0235393. [Google Scholar] [CrossRef]
- Smith, R.L. Efficient Monte Carlo procedures for generating points uniformly distributed over bounded regions. Oper. Res. 1984, 32, 1296–1308. [Google Scholar] [CrossRef]
- Hoops, S.; Sahle, S.; Gauges, R.; Lee, C.; Pahle, J.; Simus, N.; Singhal, M.; Xu, L.; Mendes, P.; Kummer, U. COPASI—A complex pathway simulator. Bioinformatics 2006, 22, 3067–3074. [Google Scholar] [CrossRef]
- Ma, H.W.; Zeng, A.P. The connectivity structure, giant strong component and centrality of metabolic networks. Bioinformatics 2003, 19, 1423–1430. [Google Scholar] [CrossRef] [PubMed]
- Klamt, S.; Gilles, E.D. Minimal cut sets in biochemical reaction networks. Bioinformatics 2004, 20, 226–234. [Google Scholar] [CrossRef] [PubMed]
- Ballerstein, K.; von Kamp, A.; Klamt, S.; Haus, U.U. Minimal cut sets in a metabolic network are elementary modes in a dual network. Bioinformatics 2012, 28, 381–387. [Google Scholar] [CrossRef]
- Takiguchi, N.; Shimizu, H.; Shioya, S. An on-line physiological state recognition system for the lysine fermentation process based on a metabolic reaction model. Biotechnol. Bioeng. 1997, 55, 170–181. [Google Scholar] [CrossRef]
- Henry, O.; Kamen, A.; Perrier, M. Monitoring the physiological state of mammalian cell perfusion processes by on-line estimation of intracellular fluxes. J. Process Control 2007, 17, 241–251. [Google Scholar] [CrossRef]
- Lequeux, G.; Beauprez, J.; Maertens, J.; Van Horen, E.; Soetaert, W.; Vandamme, E.; Vanrolleghem, P.A. Dynamic metabolic flux analysis demonstrated on cultures where the limiting substrate is changed from carbon to nitrogen and vice versa. J. Biomed. Biotechnol. 2010, 2010, 621645. [Google Scholar] [CrossRef] [PubMed]
- Niklas, J.; Schräder, E.; Sandig, V.; Noll, T.; Heinzle, E. Quantitative characterization of metabolism and metabolic shifts during growth of the new human cell line AGE1. HN using time resolved metabolic flux analysis. Bioprocess Biosyst. Eng. 2011, 34, 533–545. [Google Scholar] [CrossRef]
- Varma, A.; Palsson, B.O. Stoichiometric flux balance models quantitatively predict growth and metabolic by-product secretion in wild-type Escherichia coli W3110. Appl. Environ. Microbiol. 1994, 60, 3724–3731. [Google Scholar] [CrossRef]
- Mahadevan, R.; Edwards, J.S.; Doyle III, F.J. Dynamic flux balance analysis of diauxic growth in Escherichia coli. Biophys. J. 2002, 83, 1331–1340. [Google Scholar] [CrossRef]
- Hjersted, J.L.; Henson, M.A. Optimization of fed-batch Saccharomyces cerevisiae fermentation using dynamic flux balance models. Biotechnol. Prog. 2006, 22, 1239–1248. [Google Scholar] [CrossRef] [PubMed]
- Meadows, A.L.; Karnik, R.; Lam, H.; Forestell, S.; Snedecor, B. Application of dynamic flux balance analysis to an industrial Escherichia coli fermentation. Metab. Eng. 2010, 12, 150–160. [Google Scholar] [CrossRef]
- Hanly, T.J.; Henson, M.A. Dynamic flux balance modeling of microbial co-cultures for efficient batch fermentation of glucose and xylose mixtures. Biotechnol. Bioeng. 2011, 108, 376–385. [Google Scholar] [CrossRef]
- Grafahrend-Belau, E.; Junker, A.; Eschenröder, A.; Müller, J.; Schreiber, F.; Junker, B.H. Multiscale metabolic modeling: Dynamic flux balance analysis on a whole-plant scale. Plant Physiol. 2013, 163, 637–647. [Google Scholar] [CrossRef]
- Emenike, V.N.; Schenkendorf, R.; Krewer, U. Model-based optimization of biopharmaceutical manufacturing in Pichia pastoris based on dynamic flux balance analysis. Comput. Chem. Eng. 2018, 118, 1–13. [Google Scholar] [CrossRef]
- Gomez, J.A.; Höffner, K.; Barton, P.I. DFBAlab: A fast and reliable MATLAB code for dynamic flux balance analysis. BMC Bioinform. 2014, 15, 1–10. [Google Scholar] [CrossRef]
- Llaneras, F.; Picó, J. A procedure for the estimation over time of metabolic fluxes in scenarios where measurements are uncertain and/or insufficient. BMC Bioinform. 2007, 8, 1–25. [Google Scholar] [CrossRef]
- Gayen, K.; Venkatesh, K.V. Analysis of optimal phenotypic space using elementary modes as applied to Corynebacterium glutamicum. BMC Bioinform. 2006, 7, 1–13. [Google Scholar] [CrossRef][Green Version]
- Zamorano, F.; Vande Wouwer, A.; Bastin, G. A detailed metabolic flux analysis of an underdetermined network of CHO cells. J. Biotechnol. 2010, 150, 497–508. [Google Scholar] [CrossRef]
- Richelle, A.; Fickers, P.; Bogaerts, P. Macroscopic modelling of baker’s yeast production in fed-batch cultures and its link with trehalose production. Comput. Chem. Eng. 2014, 61, 220–233. [Google Scholar] [CrossRef]
- Baroukh, C.; Muñoz-Tamayo, R.; Steyer, J.P.; Bernard, O. DRUM: A new framework for metabolic modeling under non-balanced growth. Application to the carbon metabolism of unicellular microalgae. PLoS ONE 2014, 9, e104499. [Google Scholar]
- López Zazueta, C.; Bernard, O.; Gouzé, J.L. Dynamical reduction of linearized metabolic networks through quasi steady state approximation. AIChE J. 2019, 65, 18–31. [Google Scholar] [CrossRef]
- Plaza, J.; Bogaerts, P. FBA-based simulator of Saccharomyces cerevisiae fed-batch cultures involving an internal unbalanced metabolite. IFAC-PapersOnLine 2019, 52, 169–174. [Google Scholar] [CrossRef]
- Haag, J.E.; Vande Wouwer, A.; Bogaerts, P. Systematic procedure for the reduction of complex biological reaction pathways and the generation of macroscopic equivalents. Chem. Eng. Sci. 2005, 60, 459–465. [Google Scholar] [CrossRef]
- Haag, J.E.; Vande Wouwer, A.; Bogaerts, P. Dynamic modeling of complex biological systems: A link between metabolic and macroscopic description. Math. Biosci. 2005, 193, 25–49. [Google Scholar] [CrossRef]
- Niu, H.; Amribt, Z.; Fickers, P.; Tan, W.; Bogaerts, P. Metabolic pathway analysis and reduction for mammalian cell cultures—Towards macroscopic modeling. Chem. Eng. Sci. 2013, 102, 461–473. [Google Scholar] [CrossRef]
- Machado, D.; Soons, Z.; Patil, K.R.; Ferreira, E.C.; Rocha, I. Random sampling of elementary flux modes in large-scale metabolic networks. Bioinformatics 2012, 28, i515–i521. [Google Scholar] [CrossRef]
- Song, H.S.; Ramkrishna, D. Reduction of a set of elementary modes using yield analysis. Biotechnol. Bioeng. 2009, 102, 554–568. [Google Scholar] [CrossRef] [PubMed]
- Soons, Z.I.; Ferreira, E.C.; Rocha, I. Selection of elementary modes for bioprocess control. IFAC Proc. Vol. 2010, 43, 156–161. [Google Scholar] [CrossRef]
- Soons, Z.I.; Ferreira, E.C.; Rocha, I. Identification of minimal metabolic pathway models consistent with phenotypic data. J. Process Control 2011, 21, 1483–1492. [Google Scholar] [CrossRef][Green Version]
- Song, H.S.; Ramkrishna, D.; Pinchuk, G.E.; Beliaev, A.S.; Konopka, A.E.; Fredrickson, J.K. Dynamic modeling of aerobic growth of Shewanella oneidensis. Predicting triauxic growth, flux distributions, and energy requirement for growth. Metab. Eng. 2013, 15, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Oddsdóttir, H.Æ.; Hagrot, E.; Chotteau, V.; Forsgren, A. On dynamically generating relevant elementary flux modes in a metabolic network using optimization. J. Math. Biol. 2015, 71, 903–920. [Google Scholar] [CrossRef]
- Leighty, R.W.; Antoniewicz, M.R. Dynamic metabolic flux analysis (DMFA): A framework for determining fluxes at metabolic non-steady state. Metab. Eng. 2011, 13, 745–755. [Google Scholar] [CrossRef]
- Hebing, L.; Neymann, T.; Thüte, T.; Jockwer, A.; Engell, S. Efficient generation of models of fed-batch fermentations for process design and control. IFAC-PapersOnLine 2016, 49, 621–626. [Google Scholar] [CrossRef]
- Hebing, L.; Neymann, T.; Engell, S. Application of dynamic metabolic flux analysis for process modeling: Robust flux estimation with regularization, confidence bounds, and selection of elementary modes. Biotechnol. Bioeng. 2020, 117, 2058–2073. [Google Scholar] [CrossRef]
- Abbate, T.; de Sousa, S.F.; Dewasme, L.; Bastin, G.; Vande Wouwer, A. Inference of dynamic macroscopic models of cell metabolism based on elementary flux modes analysis. Biochem. Eng. J. 2019, 151, 107325. [Google Scholar] [CrossRef]
- Maton, M.; Bogaerts, P.; Vande Wouwer, A. Selection of a Minimal Suboptimal Set of EFMs for Dynamic Metabolic Modelling. In Proceedings of the IFAC PapersOnLine 11th IFAC Symposium on Advanced Control of Chemical Processes, Venice, Italy, 13–16 June 2021. [Google Scholar]
- Haag, J.; Vande Wouwer, A.; Remy, M. A general model of reaction kinetics in biological systems. Bioprocess Biosyst. Eng. 2005, 27, 303–309. [Google Scholar] [CrossRef]
- Richelle, A.; Bogaerts, P. Systematic methodology for bioprocess model identification based on generalized kinetic functions. Biochem. Eng. J. 2015, 100, 41–49. [Google Scholar] [CrossRef]
- Kaufman, D.E.; Smith, R.L. Direction choice for accelerated convergence in hit-and-run sampling. Oper. Res. 1998, 46, 84–95. [Google Scholar] [CrossRef]
- De Martino, D.; Mori, M.; Parisi, V. Uniform sampling of steady states in metabolic networks: Heterogeneous scales and rounding. PLoS ONE 2015, 10, e0122670. [Google Scholar] [CrossRef] [PubMed]
- Bogaerts, P.; Rooman, M. DISCOPOLIS 2.0: A new recursive version of the algorithm for uniform sampling of metabolic flux distributions with linear programming. In Proceedings of the IFAC PapersOnLine 11th IFAC Symposium on Advanced Control of Chemical Processes, Venice, Italy, 13–16 June 2021. [Google Scholar]
- Yasemi, M.; Jolicoeur, M. Modelling Cell Metabolism: A Review on Constraint-Based Steady-State and Kinetic Approaches. Processes 2021, 9, 322. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bogaerts, P.; Vande Wouwer, A. How to Tackle Underdeterminacy in Metabolic Flux Analysis? A Tutorial and Critical Review. Processes 2021, 9, 1577. https://doi.org/10.3390/pr9091577
Bogaerts P, Vande Wouwer A. How to Tackle Underdeterminacy in Metabolic Flux Analysis? A Tutorial and Critical Review. Processes. 2021; 9(9):1577. https://doi.org/10.3390/pr9091577
Chicago/Turabian StyleBogaerts, Philippe, and Alain Vande Wouwer. 2021. "How to Tackle Underdeterminacy in Metabolic Flux Analysis? A Tutorial and Critical Review" Processes 9, no. 9: 1577. https://doi.org/10.3390/pr9091577
APA StyleBogaerts, P., & Vande Wouwer, A. (2021). How to Tackle Underdeterminacy in Metabolic Flux Analysis? A Tutorial and Critical Review. Processes, 9(9), 1577. https://doi.org/10.3390/pr9091577






