In all test cases presented here, the volume space was divided into 40 sections using Equation (21) with υ1 = 0.01 and s = 1.2. In each section the six pivots were specified (i.e., six moments were tracked). A time step of 0.01 s was adopted. All simulations in this work were carried out on a desktop computer with Intel Core i5 CPU (3.1 GHz) and 8 G memory.
4.1. Pure Aggregation
Five test cases were carried out to validate the new method for the pure aggregation process, in the first of which a constant aggregation kernel was adopted, having the form
with
C0 = 1, together with exponential initial particle size distribution
with
N0 = 1,
V0 = 1. In this test case, an analytical solution was available [
20]
with
T =
C0N0t/(2 +
C0N0t). The moments could be evaluated analytically.
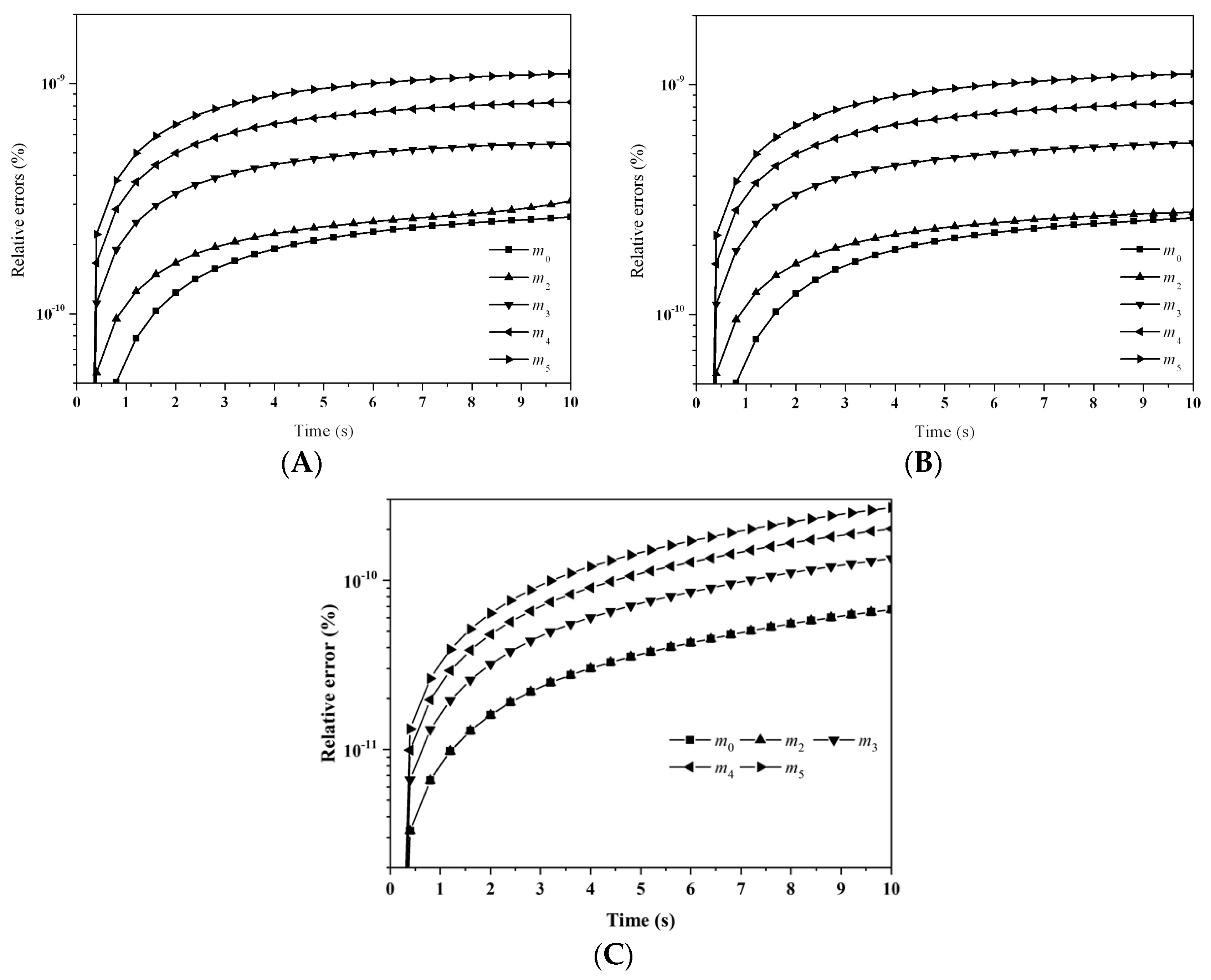
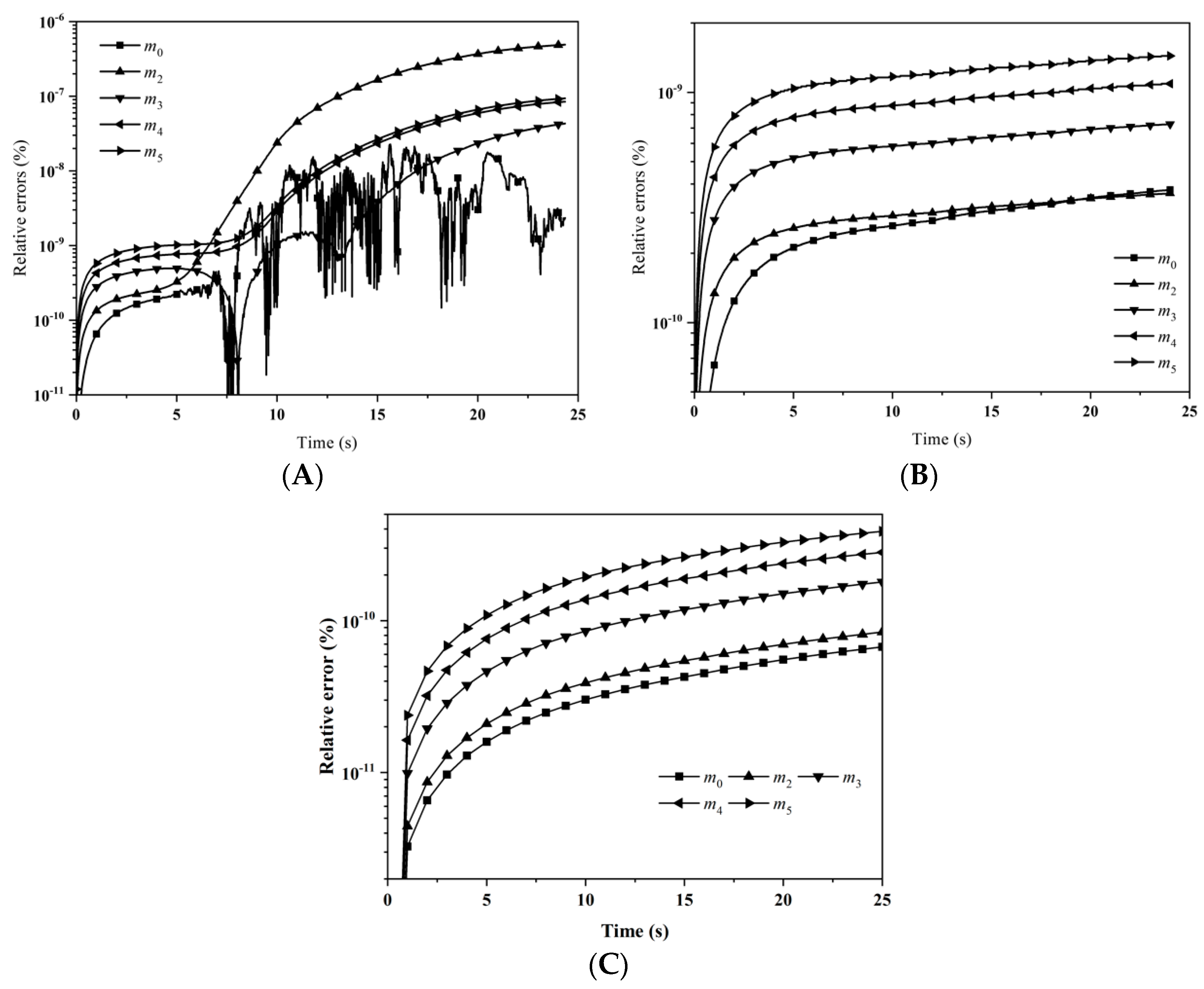
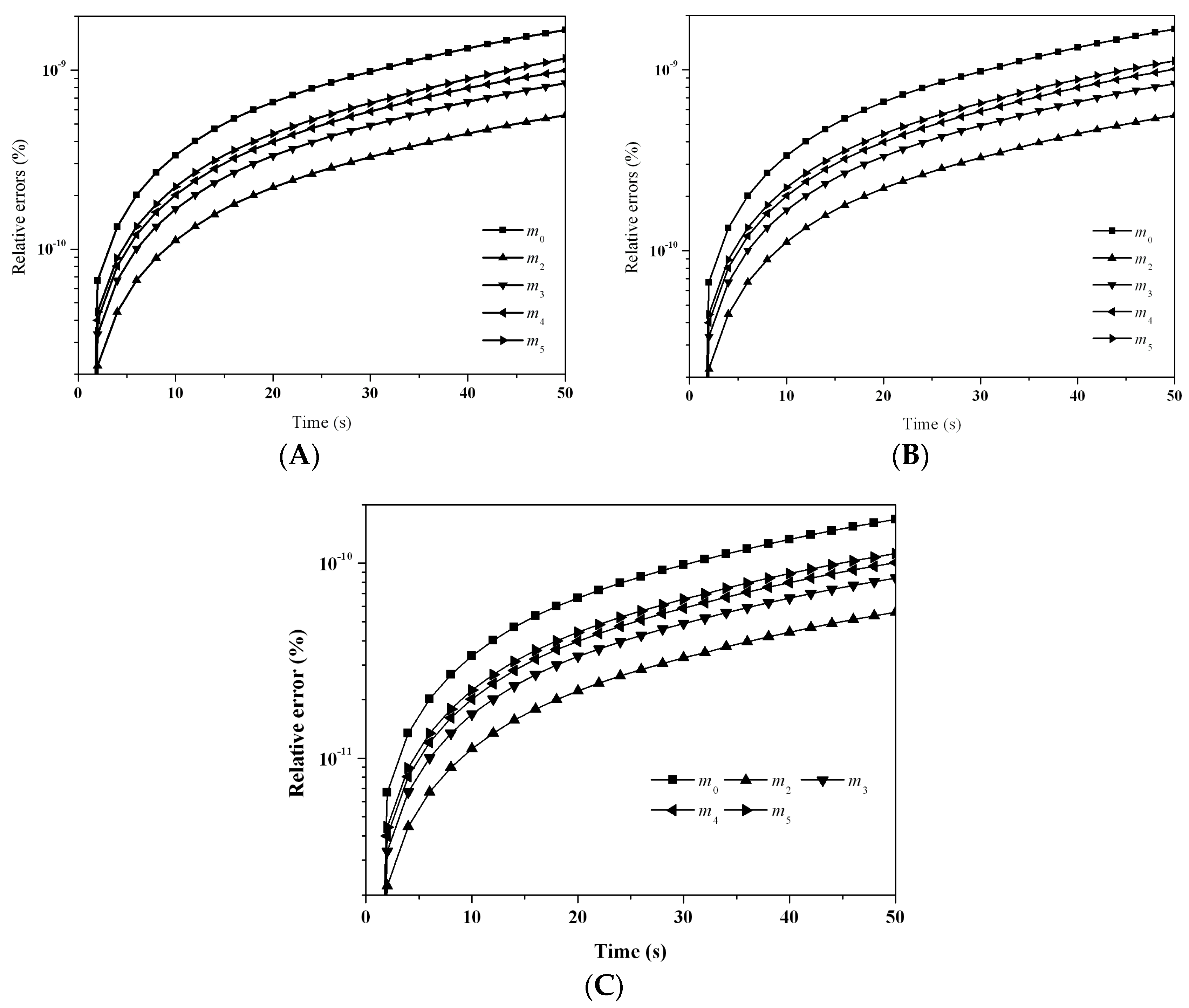
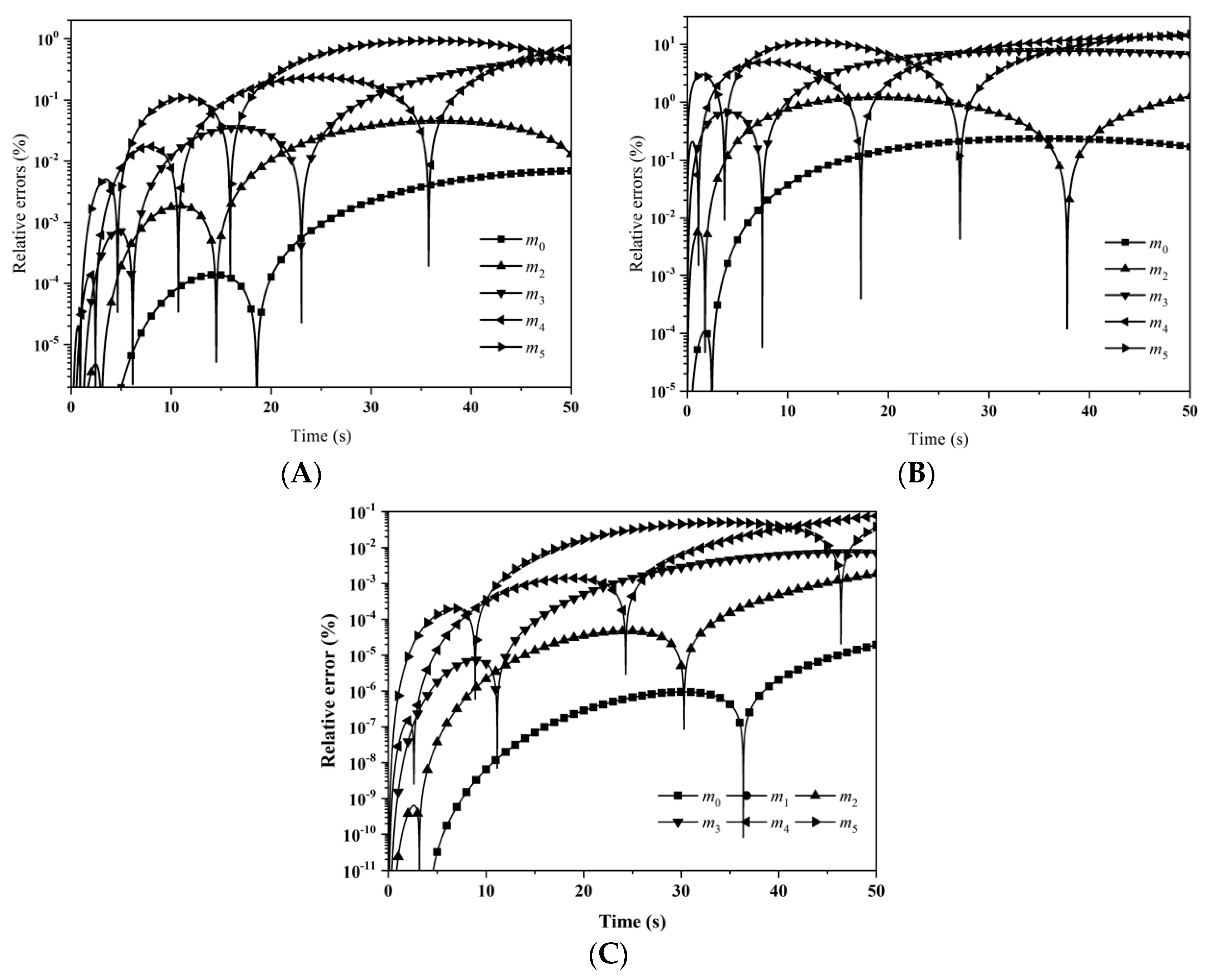
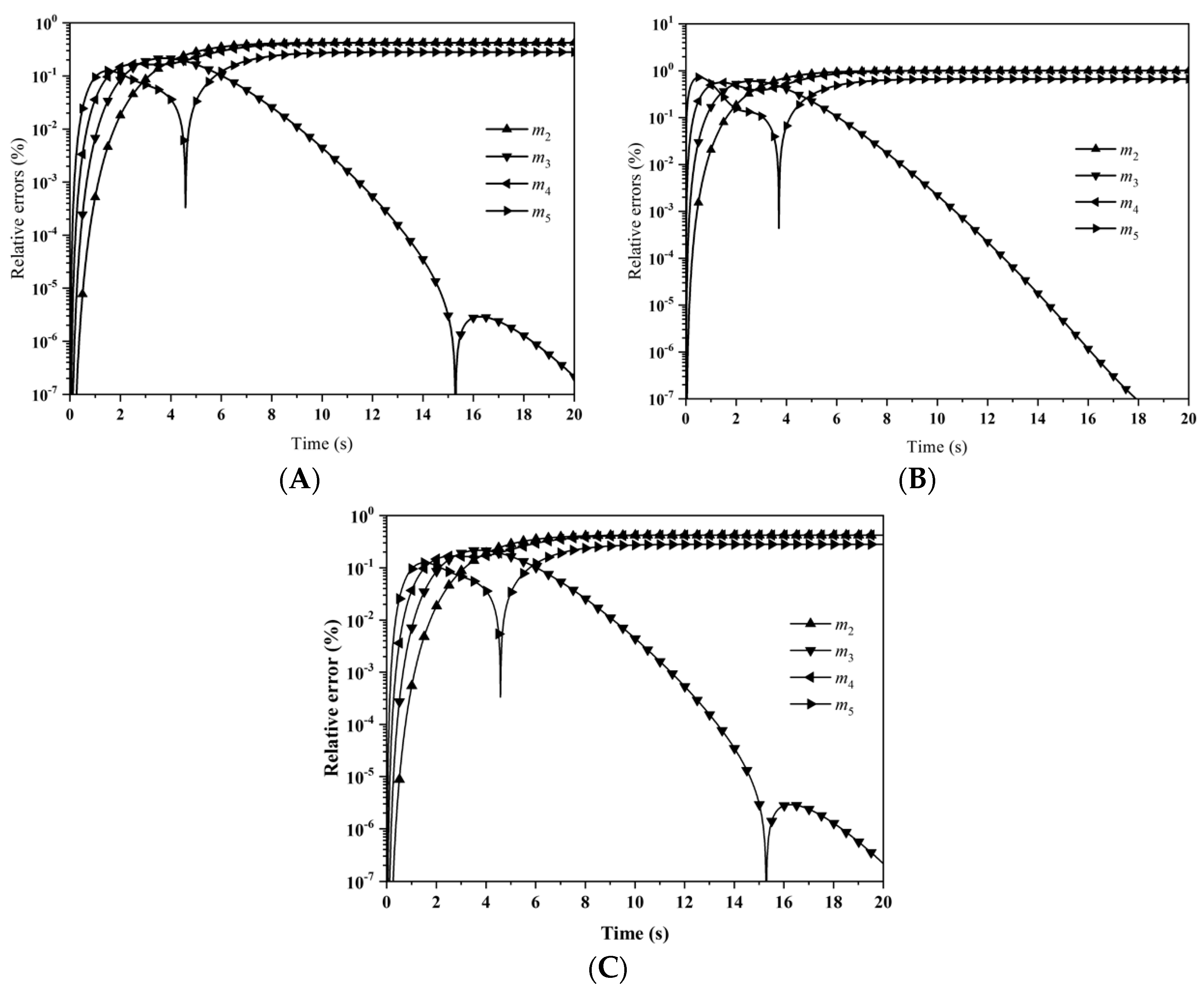
Figure 3 depicts the evolution of the percentage errors for the first six moments with: (A) LFPQMOM; (B) QMOM; and (C) FPQMOM. In the figure, the evolution for the relative errors for all the moments were similar in both methods, but a little larger than FPQMOM. Errors increased rapidly initially and leveled off after 2 s. The maximum percentage error with QMOM was 1.12 × 10
−9%, whereas with the new method, it was 1.10 × 10
−9%, a small decrease. FPQMOM had the highest accuracy. Higher rank moments had larger percentage errors with both methods. It is worth pointing out that, with LFPQMOM, the relative errors for
m1 were always zero within the simulation time, but with QMOM, they were lower than 10
−12%, but larger than zero. With the new method,
m1 could be predicted without any error. In the figure, percentage errors for
m1 were not included due to their low values.
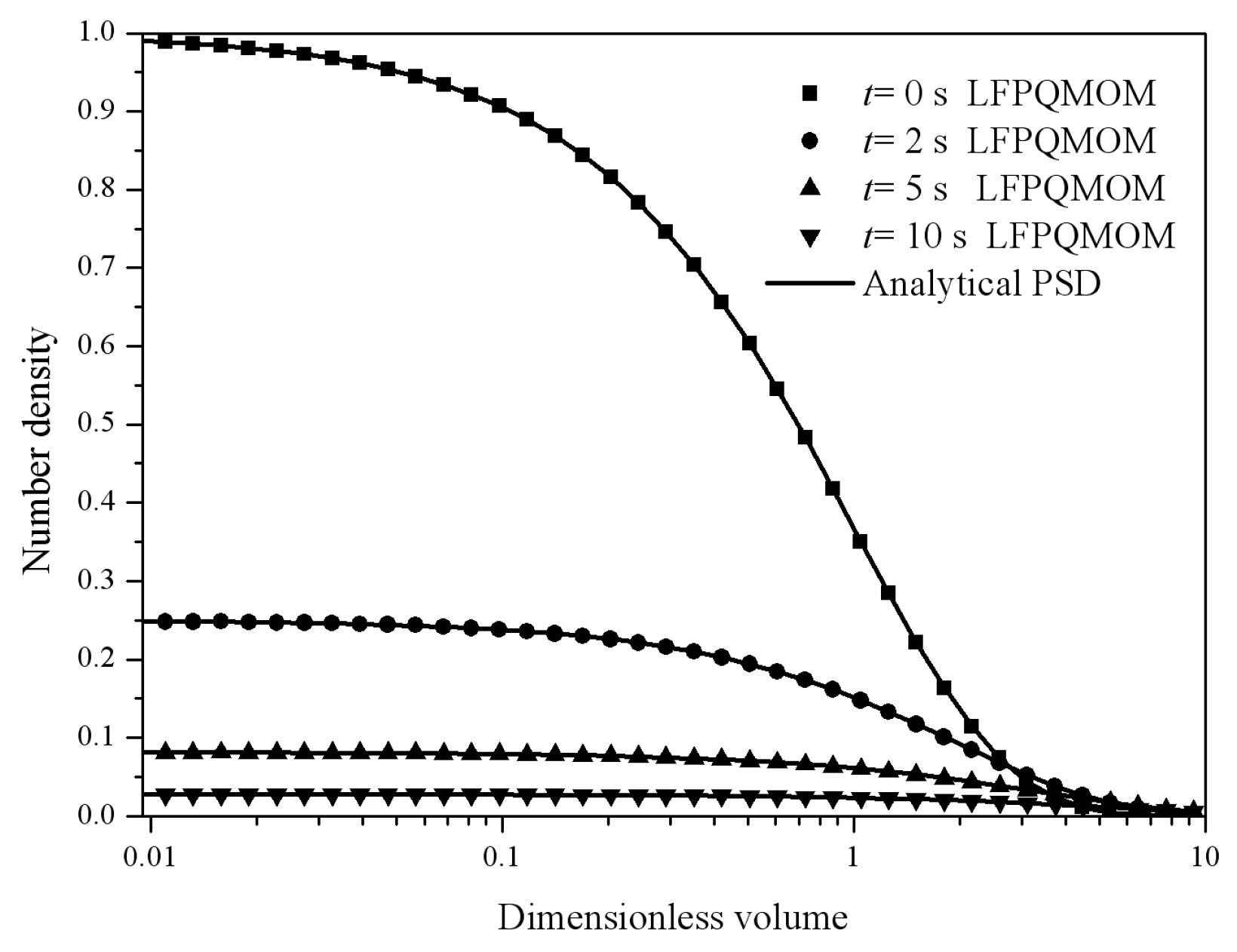
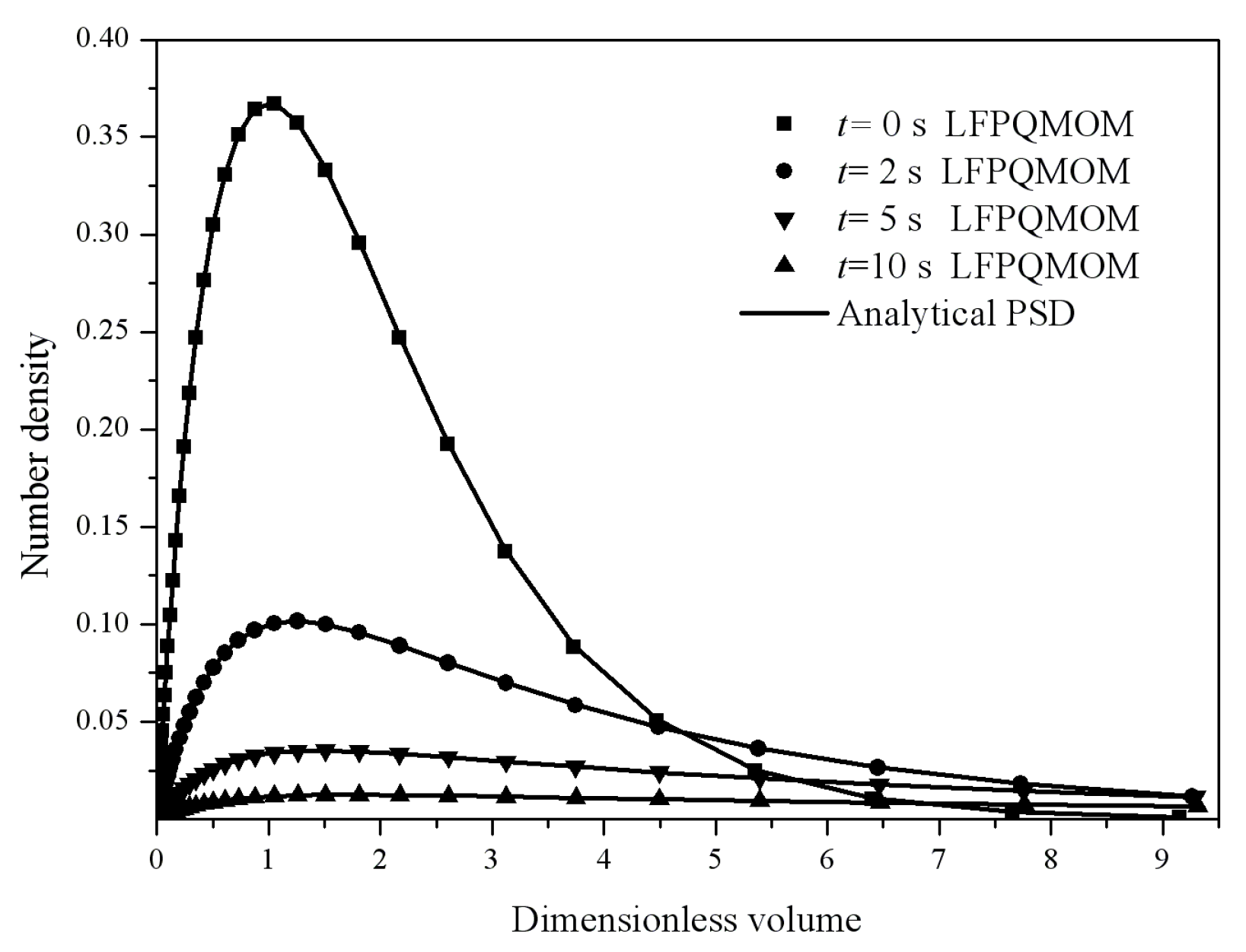
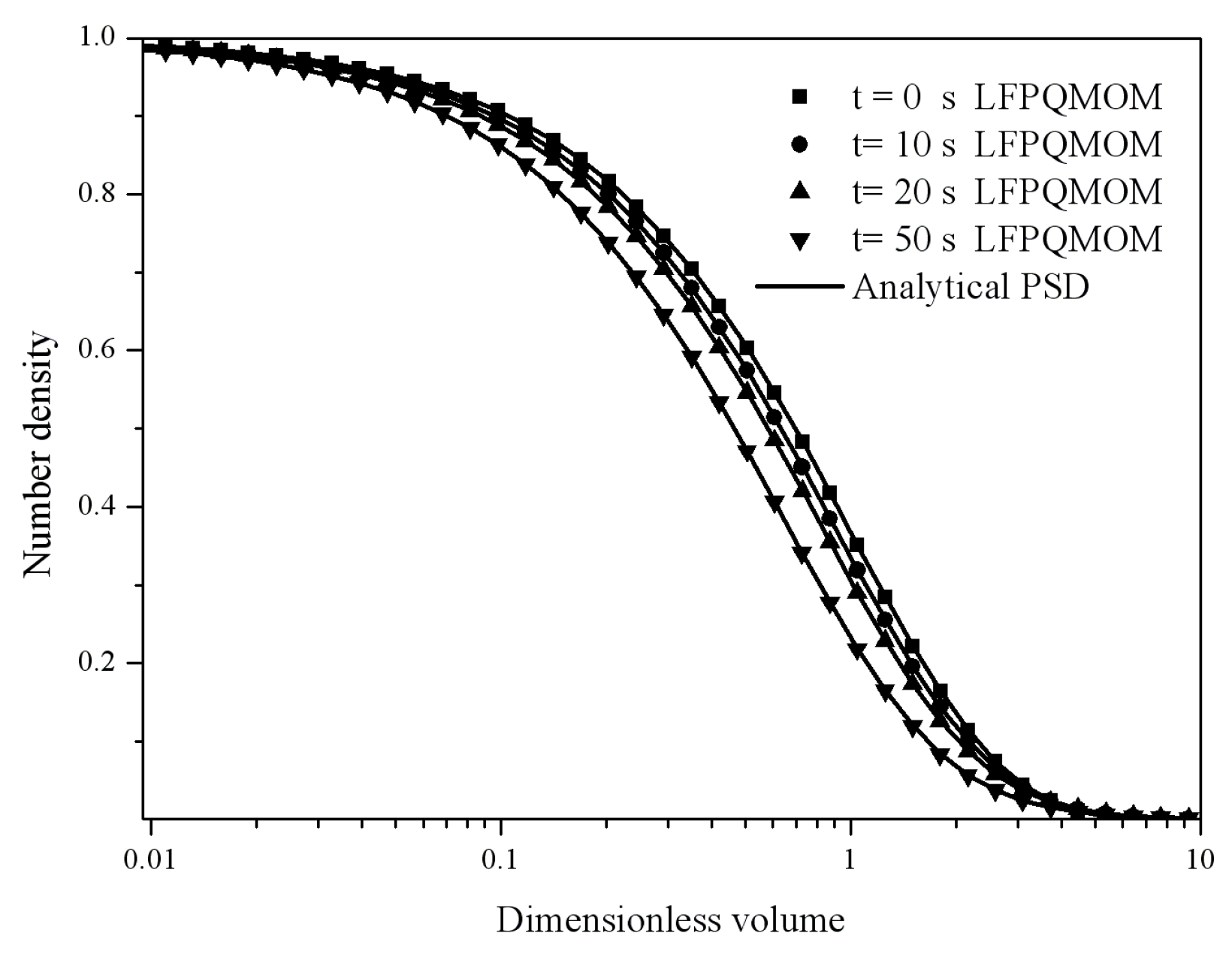
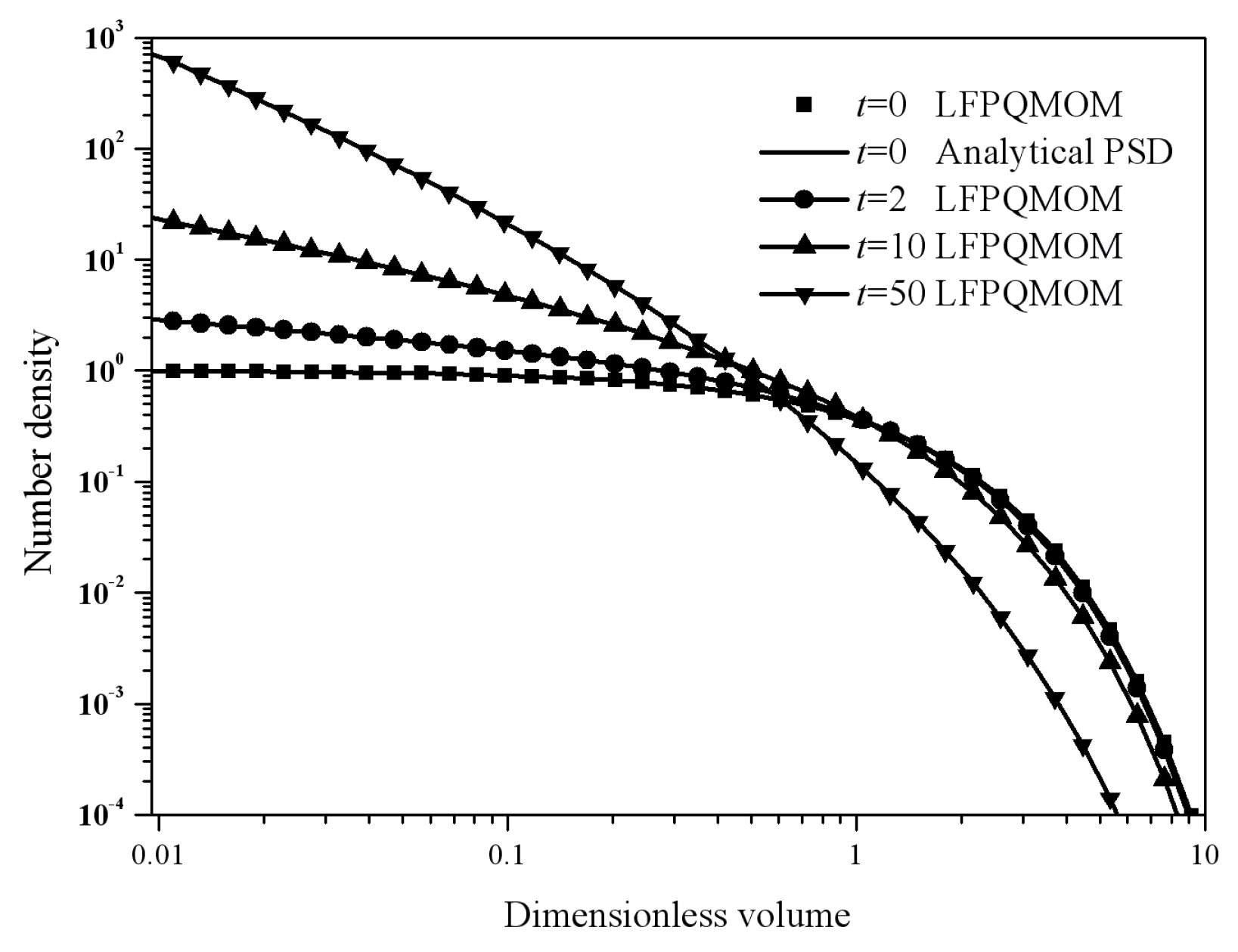
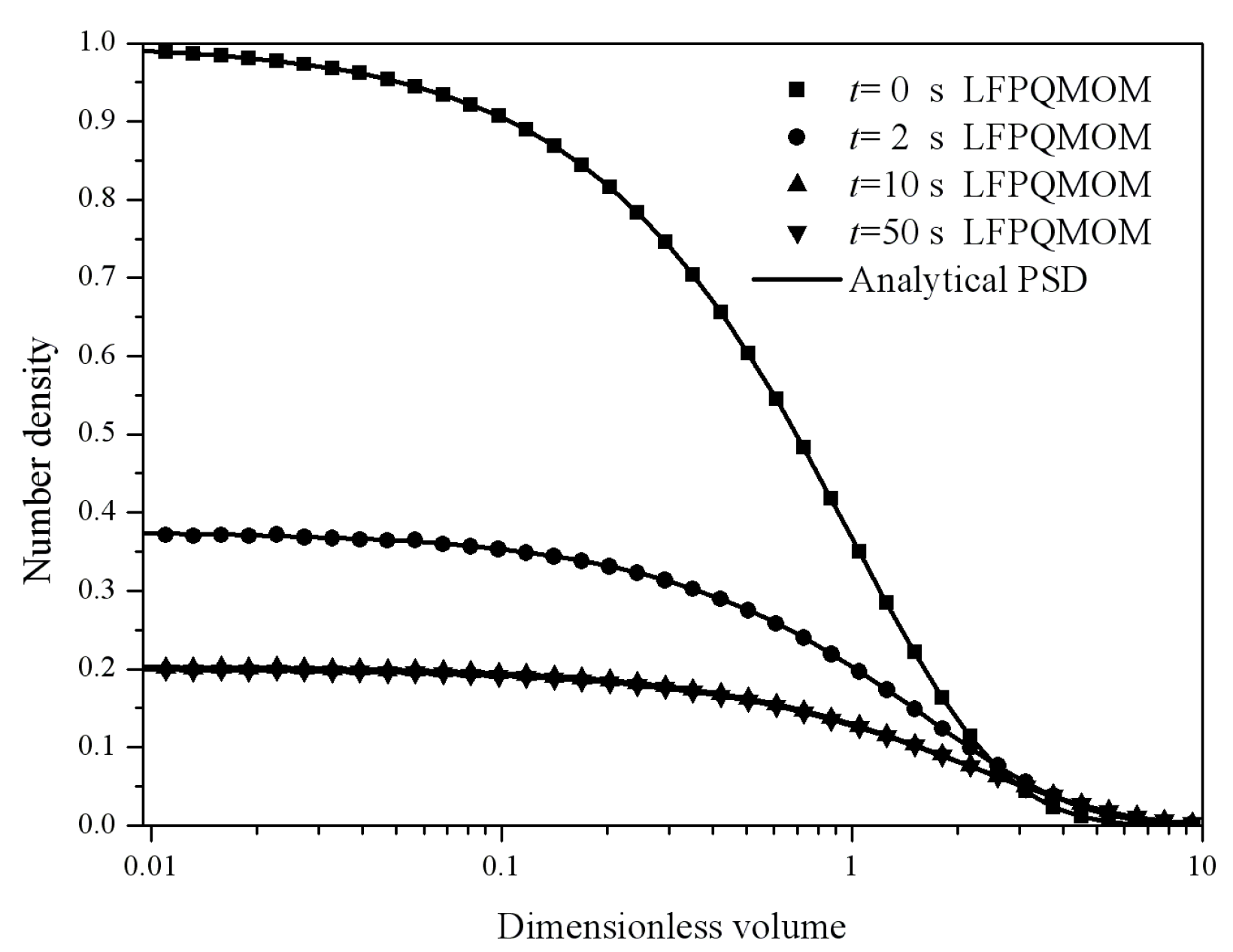
Figure 4 gives the particle size distribution predicted by LFPQMOM at 0, 2, 5, and 10 s. In the figure, the numerical PSDs agreed exactly with the analytical PSDs. The number density approached zero when the dimensionless volume was larger than 10.
The computational time was about 28,360, 1795, and 625 ms for LFPQMOM, QMOM, and FPQMOM, respectively. The computational load of LFPQMOM was much larger than QMOM and FPQMOM, since, in LFPQMOM, 40 sections were adopted in this work and, in each section, six moments were tracked. The total number of variables included in the simulation with LFPQMOM was 240, whereas, in QMOM and FPQMOM, only six variables were included. The computational time for each variable on average was 118, 299, and 104 ms for LFPQMOM, QMOM, and FPQMOM, respectively. From the view of a one-variable computational load, LFPQMOM and FPQMOM were more efficient than QMOM because of the vandermode linear system solver used.
In the second test case, a constant aggregation kernel was used, i.e., Equation (35) with
C0 = 1, but with Gaussian initial particle size distribution, having the form
with
N0 = 1 and
V0 = 1. In this test case, analytical particle size distribution is given by [
20]
with
T =
C0N0t/(2 +
C0N0t). The moments can be evaluated analytically.
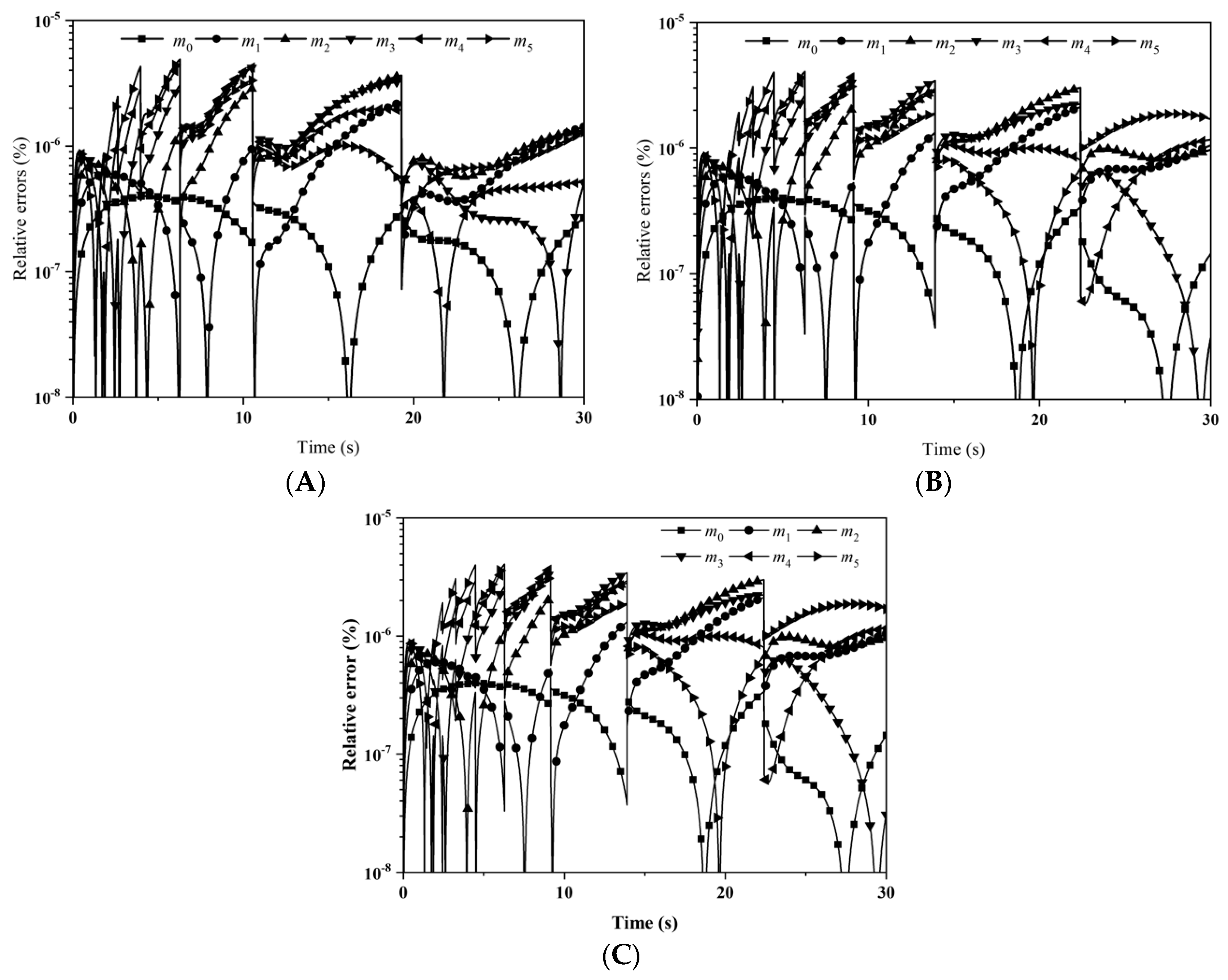
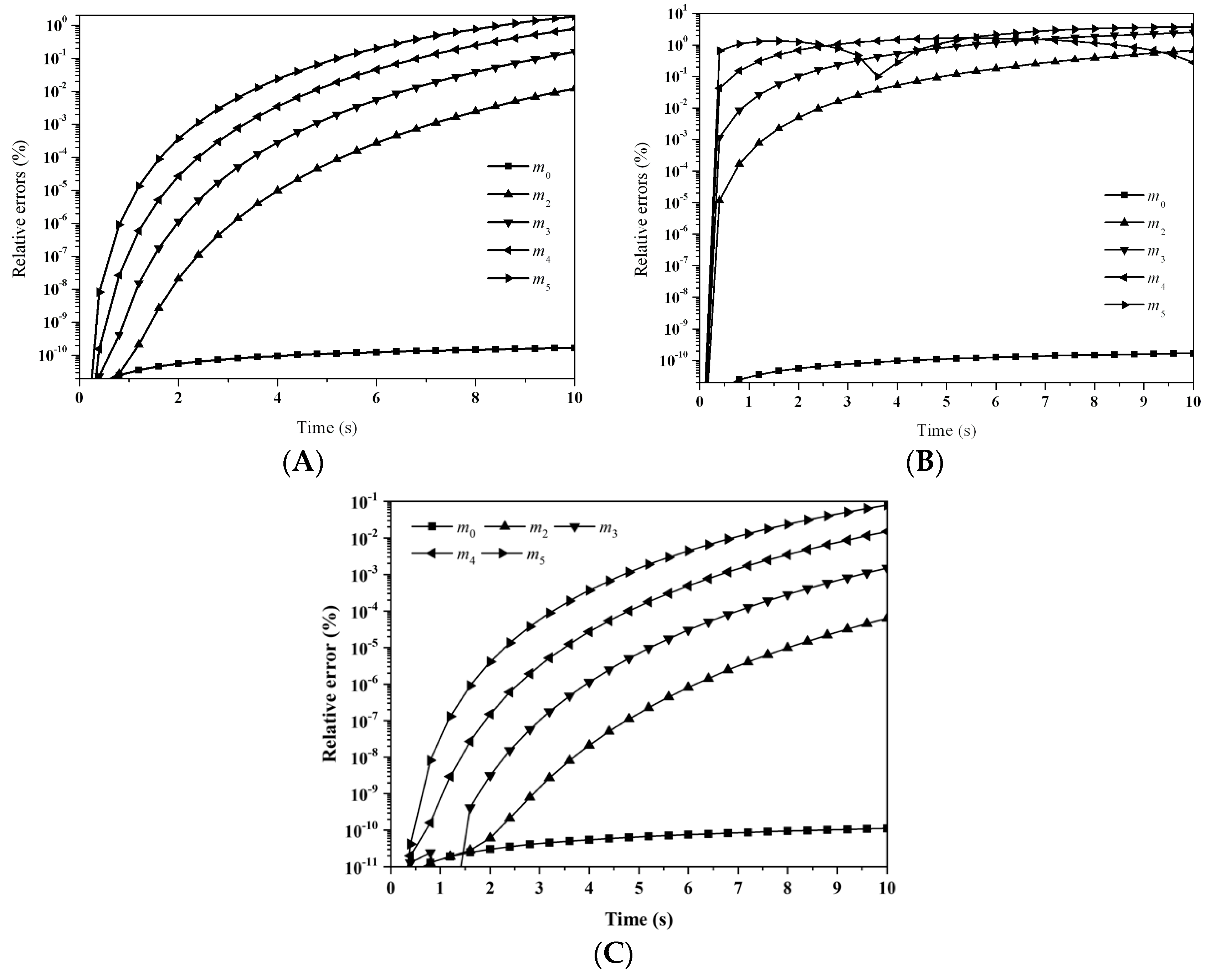
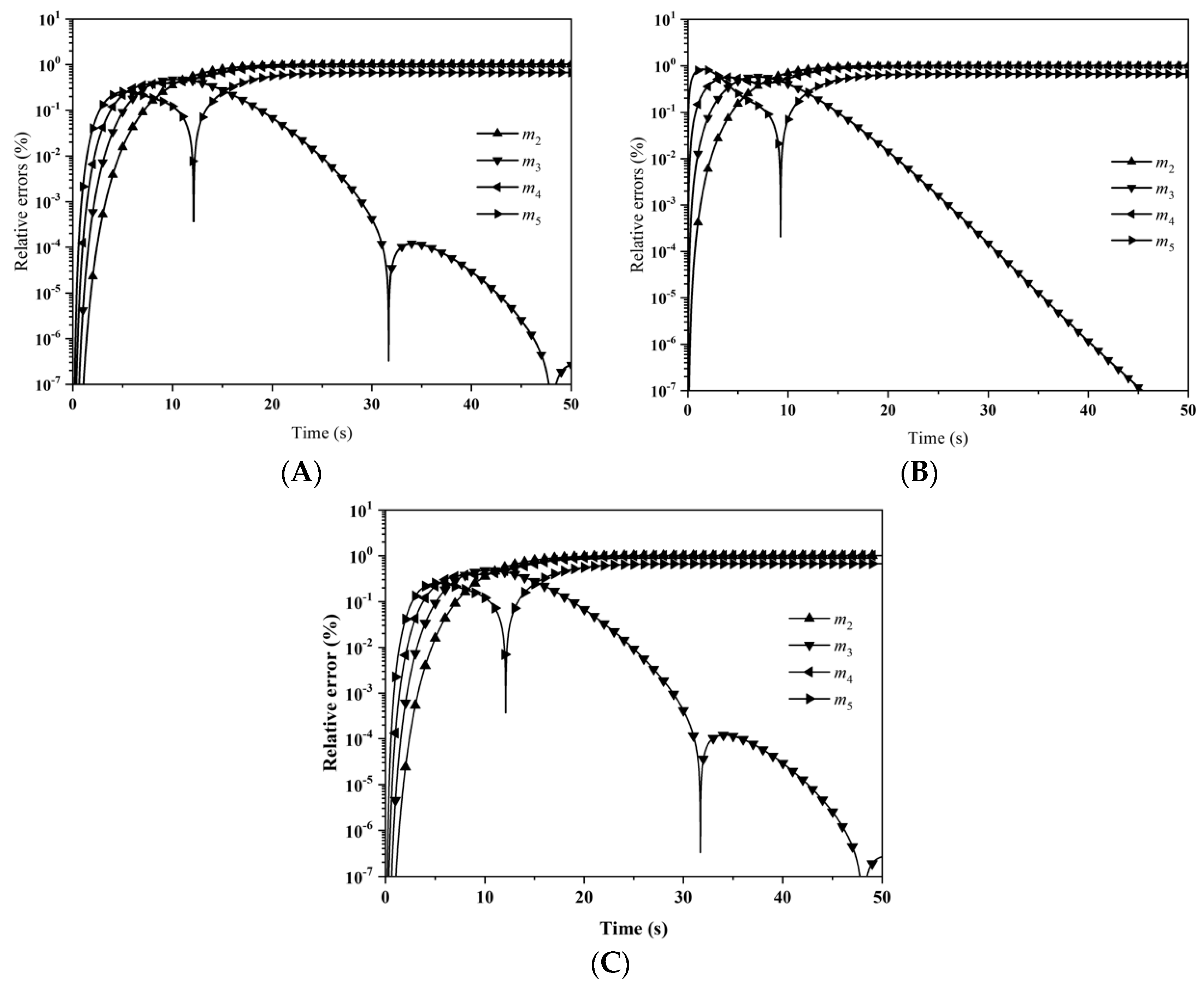
Figure 5 illustrates the relative errors for the first six moments with: (A) LFPQMOM; (B) QMOM; and (C) FPQMOM. With LFPQMOM, the relative errors rose rapidly initially, and became flat after 2 s. The relative errors for
m2 after 6 s and
m4 and
m5 after 9 s began to rise, but for
m0 and
m3 they began to fluctuate after 7 s. With QMOM, the relative errors for all the moments rose sharply initially, and leveled off after 2 s, the evolution of which was very similar to that in the previous case. With the new method, the prediction for the moments of
m2,
m4, and
m5 was poor relative to QMOM, but was lower 10
−6% within the simulation time, which could still satisfy industrial needs. With FPQMOM, the numerical prediction was the best. In this test case,
m3 could be predicted exactly without any error within the simulation time with the new method. The percentage errors for
m3 were not included in the figure due to their low values with both methods.
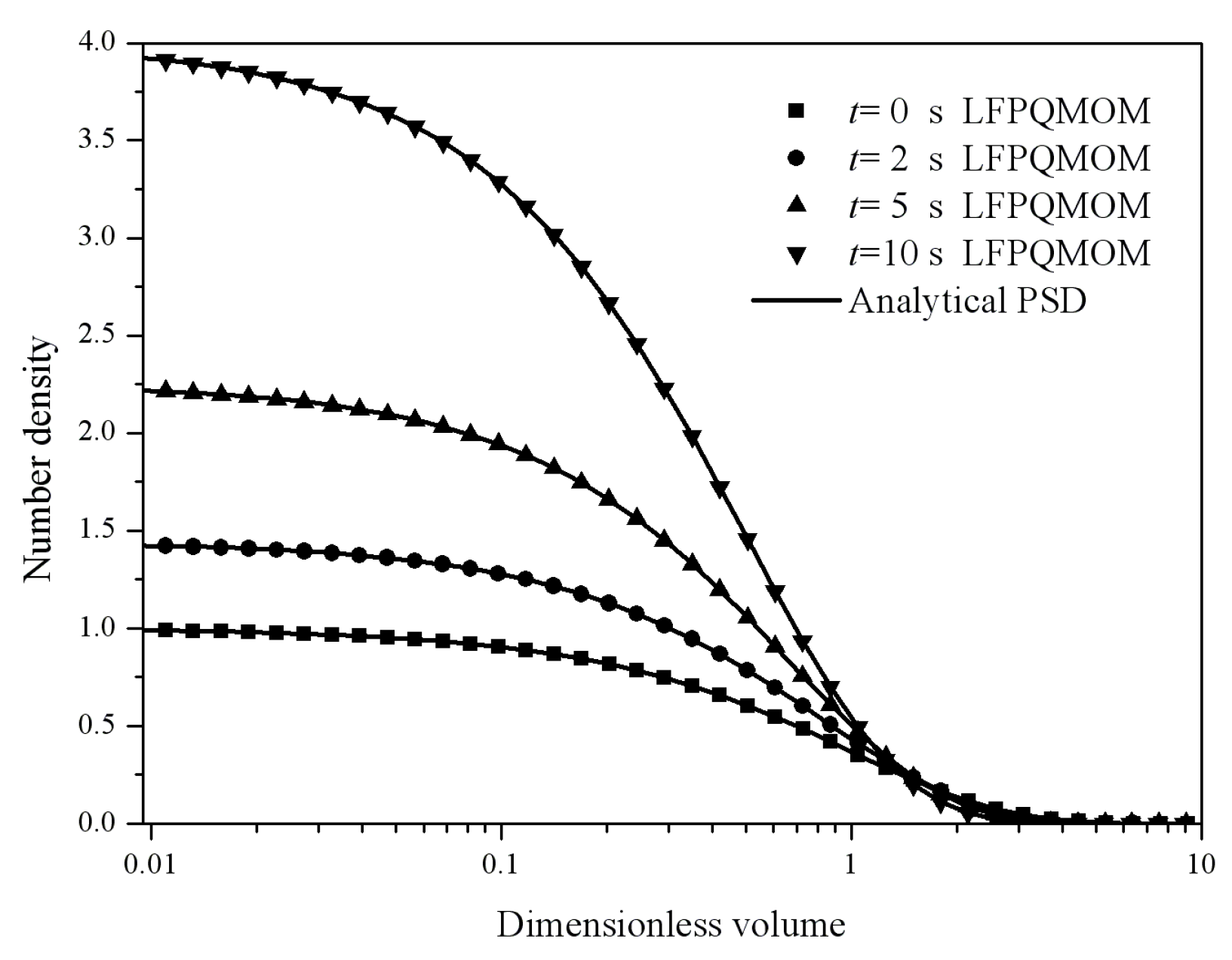
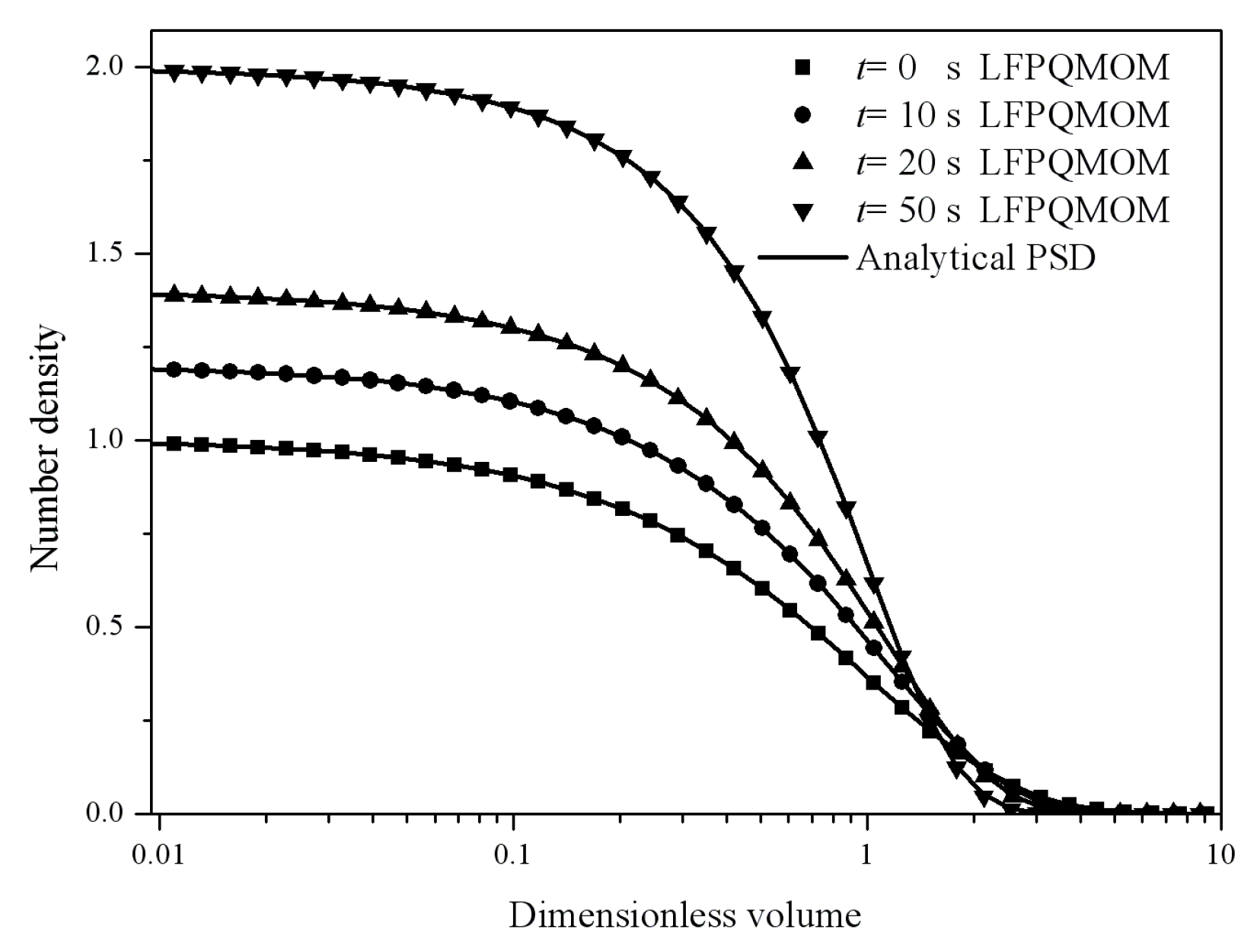
Figure 6 gives the particle size distribution predicted by LFPQMOM at 0, 2, 5, and 10 s, along with analytical solutions for comparison. In the figure, the numerical PSDs with our new method agreed well with the analytical PSDs. Again, the new method made an excellent prediction for both moments and PSD. The computational time was about 71,766, 5024, and 1709 for LFPQMOM, QMOM, and FPQMOM, respectively, in this test.
One should note that, with LFPQMOM, we did not track the moments of the PSD over the whole domain, but the moments in each section. As a result, to compute relative error with LFPQMOM, we had to resort to Equation (32) to first compute the moments of the PSD. Thus, the numerical errors of each moment presented in the figure for LFPQMOM are the compound errors, but not for one variable. For instance, to calculate errors of
m3, we calculated the
m3 with
m3i of the whole PSD using Equation (32) first, and then compared it with its analytical counterpart where
m3i is the tracked variable not
m3. In each section, a fixed pivot quadrature method of moment (FPQMOM) was used. FPQMOM was more accurate than QMOM, as can be seen in the figure. As a result, the relative errors of the moments with FPQMOM do not deviate from QMOM or LFPQMOM too much. If the relative error for a tracked variable is too small, the values of the relative errors can be random. The compound relative errors can also be random. The fluctuations in the errors of
m0 in
Figure 5A may be related to this.
In the third test case, a sum aggregation kernel was used
with
C0 = 0.01, together with the exponential initial distribution, i.e., Equation (36) with
N0 = 1 and
V0 = 1. Analytical solution was given by [
20]
with
x =
V/
V0 and
τ = 1 − exp(−
C0N0V0t).
I1(
x) is the modified Bessel function of the first kind. Moments can be evaluated using the numerical integration method.
Figure 7 shows the evolution of percentage errors for the first six moments with: (A) LFPQMOM; (B) QMOM; and (C) FPQMOM. In the figure, within the simulation time, the relative errors for all the moments were lower than 10
−5%. It should be noted that the relative errors for
m1 were not lower than 10
−12%, even though the conservation of the mass was considered in the new method. This is because no analytical solutions for the moments were available. The numerical errors for
m1 were not caused by the new method or QMOM, but came from the evaluation of the moments through numerical integration. The sharp drops in the relative errors were related with the numerical errors of the evaluation of the moments.
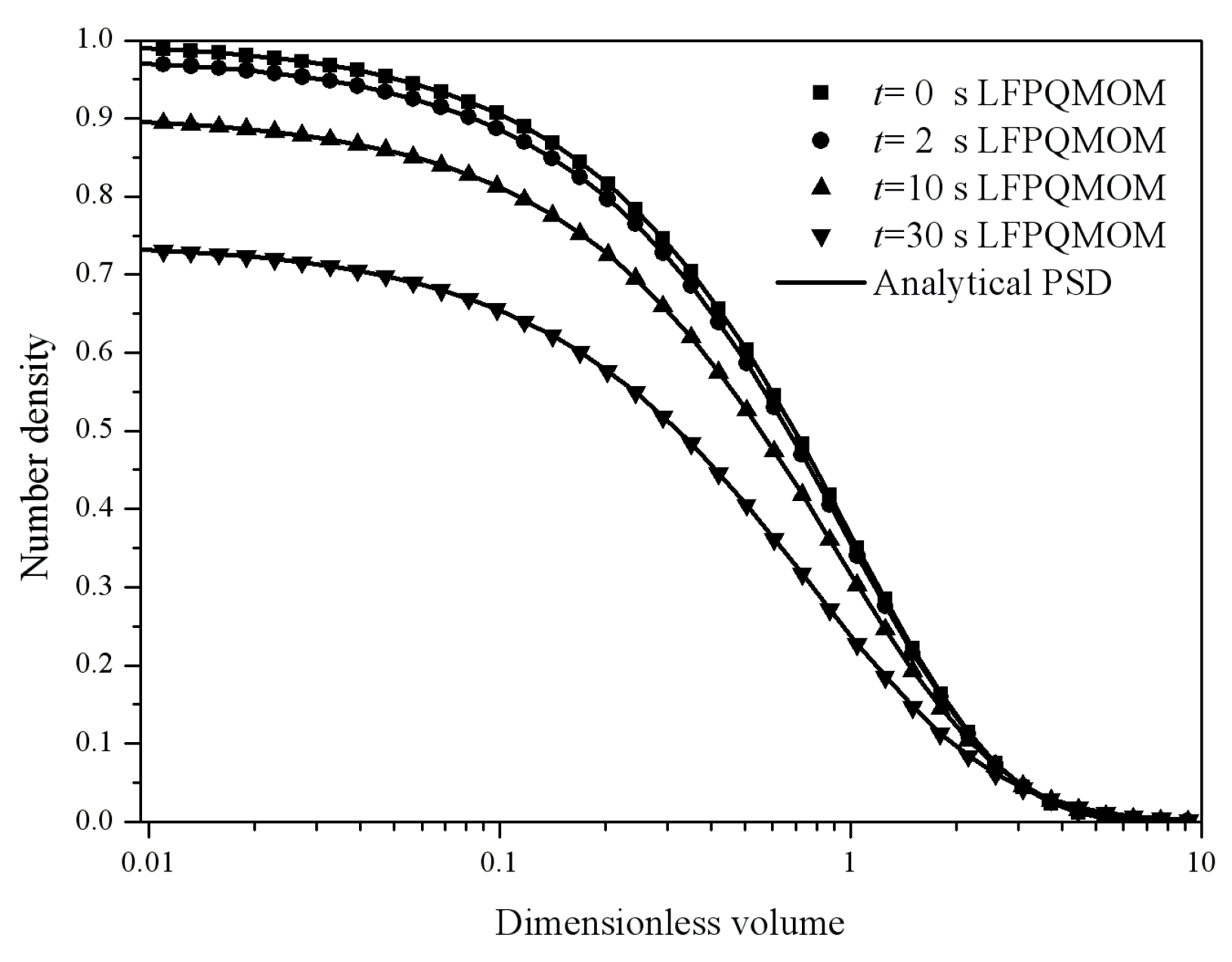
Figure 8 illustrates the particle size distribution at 0, 2, 10, and 30 s. It can be observed that the numerical predictions with the new method were in excellent agreement with the analytical counterparts. Number density with dimensionless volume larger than 10 approached zero. The computational time was about 86,314, 5570, and 1948 ms for LFPQMOM, QMOM, and FPQMOM, respectively.
In the fourth test case, a product aggregation kernel was adopted, having the form
with
C0 = 0.01, together with the exponential initial PSD, i.e., Equation (36) with
N0 = 1 and
V0 = 1. Analytical solution for this test case was given by [
20]
where
,
x =
V/
V0 and Γ (
x) is the gamma function.
Note that the product kernel is a gelling kernel, and the second and higher moments will become infinite within finite time [
21]. Most existing moment methods cannot predict the moments exactly, especially near the gelling point. The analytical solutions for moments
m0,
m1, and
m2 are given in
Table 1. Based on the solution for
m2, the actual value for
tgel can be calculated
The gelling time
tgel is 50 s in this test case. To predict the moments as exactly as possible, especially near the gelling point, two nodes were adopted for QMOM and three pivots in each section for LFPQMOM. The percentage errors of the first three moments are shown in
Figure 9A–C for LFPQMOM, QMOM, and FPQMOM, respectively. An interesting feature with the new method is that the simulation can be carried out for as long as possible before the gelling point. The maximum time that can be used as the simulation time is
tgel −
Δt. Δ
t is the time step. In this test case, a time step of 0.01 s was adopted, thus any time of
t ≤ 49.99 s could be adopted as the simulation time. With QMOM, only an even number of moments could be tracked, four moments in this test case. We could only carry out the simulation before the time of 49.6 s when
m3 diverged. After that time, QMOM was not appropriate, and the new method could also make a good prediction, with a relative error smaller than 10
−5%, as can be seen in
Figure 9A. Relative errors with FPQMOM were similar to those with LFPQMOM. The relative errors for
m1 with all methods here were lower than 10
−13%, hence were not included in the figures.
Figure 10 shows a good agreement of numerical PSD predicted by the new method at 0, 10, 20, and 50 s with the analytical solution. Number density approached zero with a dimensionless volume larger than 10. The computational time was about 143,962, 8957, and 3199 ms for LFPQMOM, QMOM, and FPQMOM, respectively, in this test.
In the final test case for pure aggregation, a product kernel was used, i.e., Equation (42) with
C0 = 0.01, but with a Gaussian initial PSD, i.e., Equation (38) with
N0 = 1 and
V0 = 1. Analytical solution for PSD is given by [
21]
where
,
x =
V/
V0 and Γ (
x) is the gamma function. The analytical moments are listed in
Table 1. The gelling time
tgel was 16.67 s. Similar to the previous case, two nodes were adopted with QMOM and three pivots in each section with LFPQMOM to track the moments exactly.
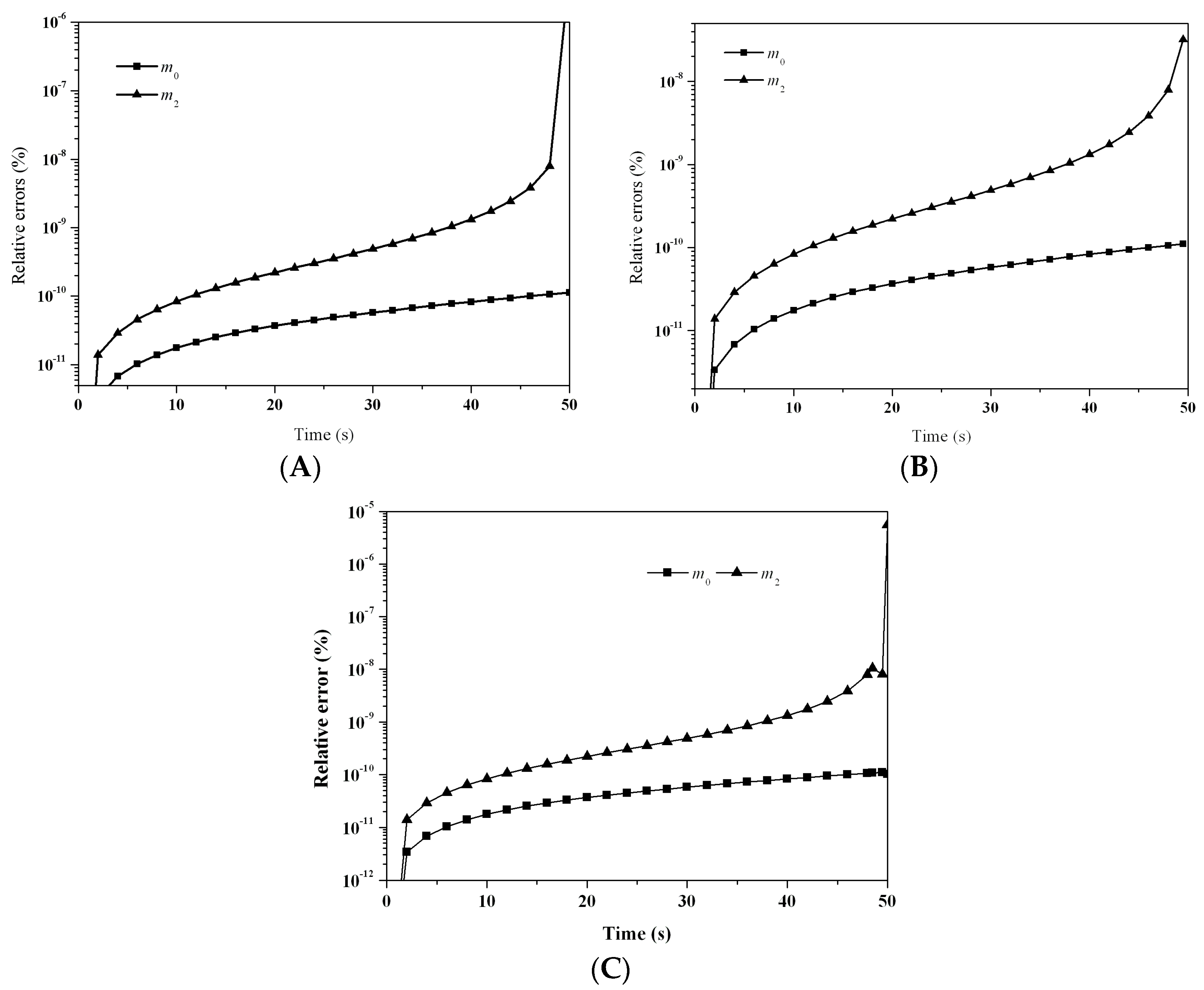
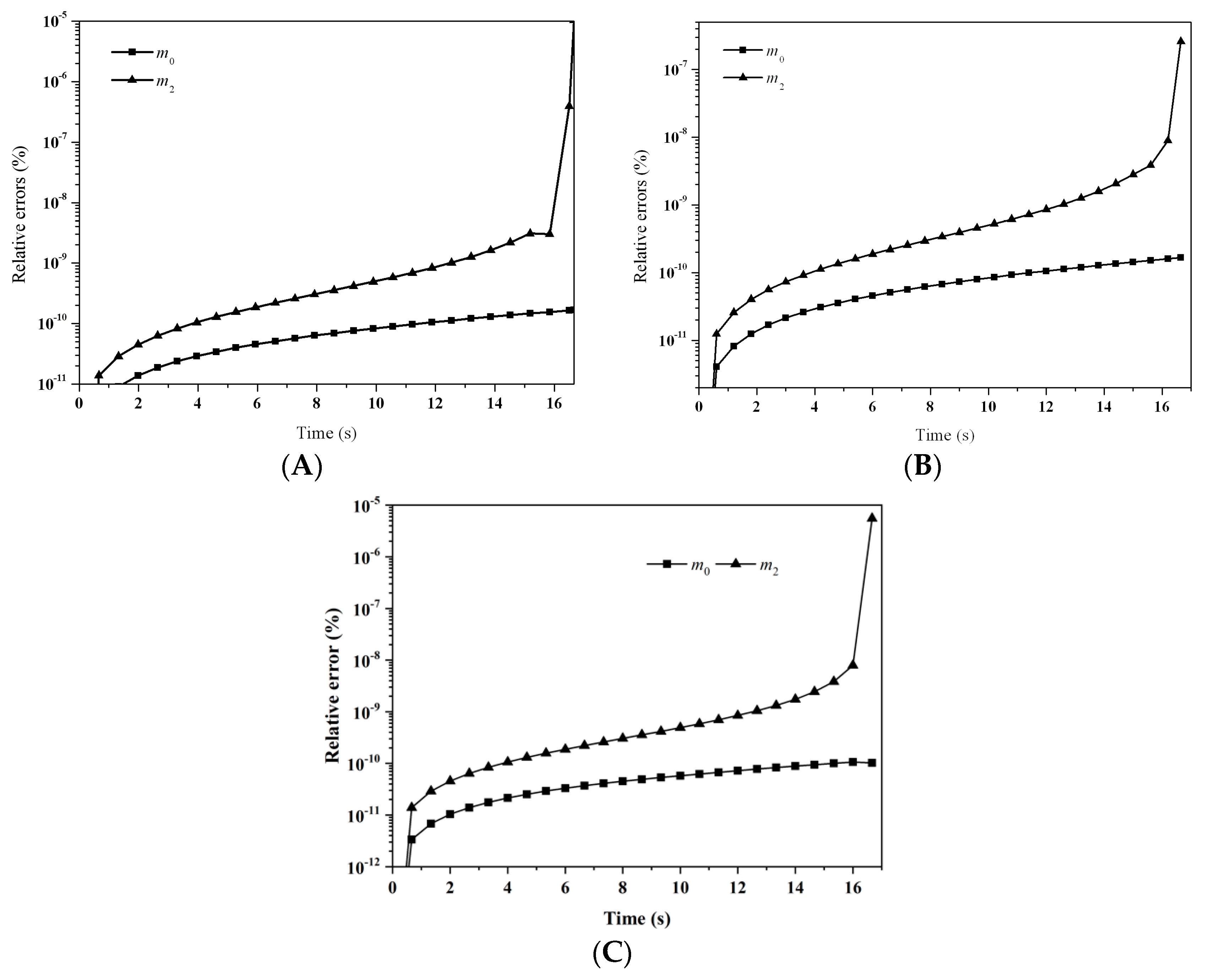
Figure 11 depicts the evolution of the relative errors for the first six moments predicted by: (A) LFPQMOM; (B) QMOM; and (C) FPQMOM. The gelling time was 16.67 and a simulation time of 16.67 − Δ
t (=16.66) could be carried out with the new method, when
m3 diverged. A maximum simulation time of 16.65 s could only be adopted with QMOM. Within a simulation of 16.65 s, the percentage errors with both methods were comparable, which were lower than 10
−6%, but, after that time, QMOM was not appropriate for this test case. Similar to the previous case, the relative errors for
m2 near the gelling point rose sharply. The relative errors for
m1 were not included in the figure for their low values. Again, numerical errors with FPQMOM were similar to LFPQMOM.
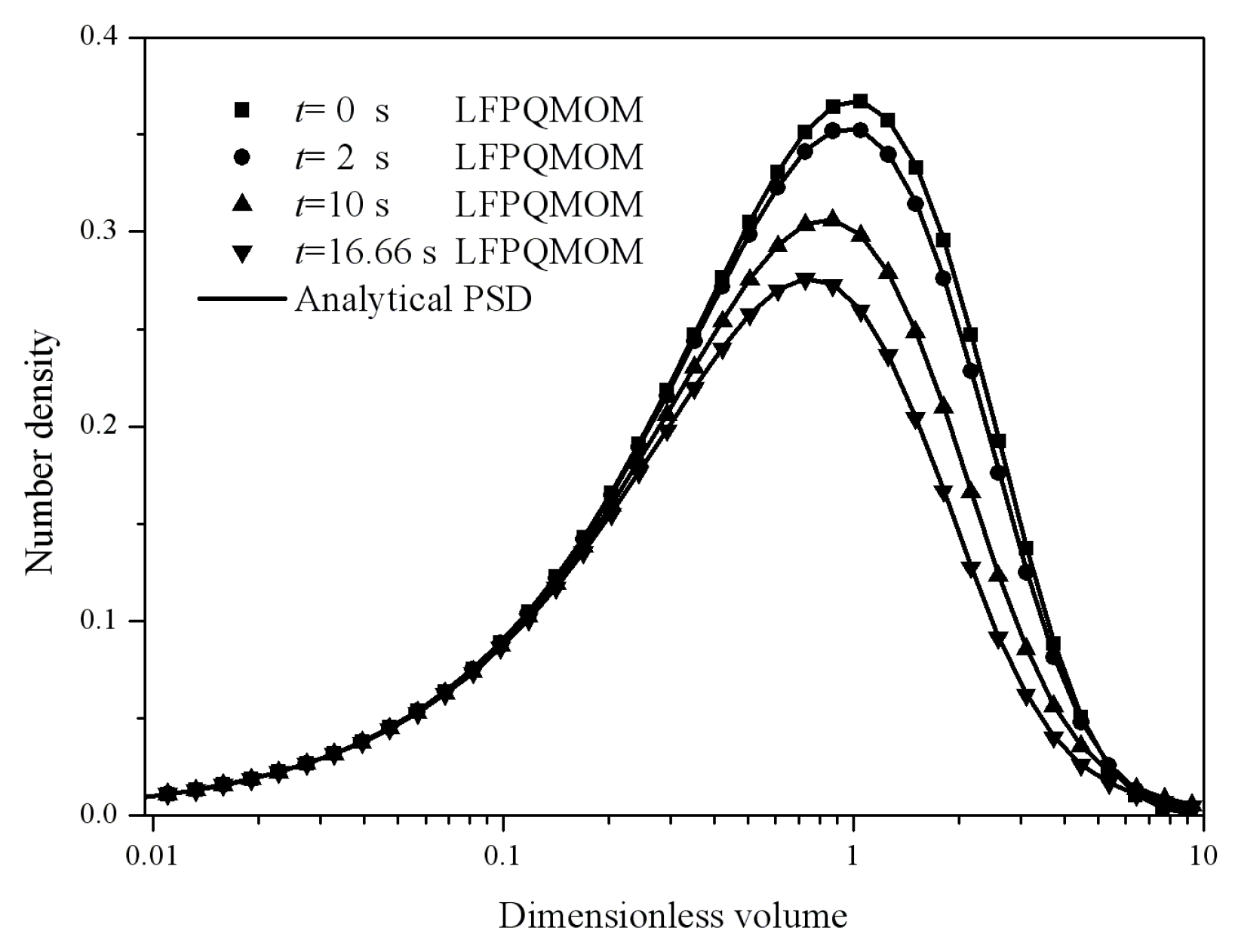
Figure 12 gives the particle size distributions at 0, 2, 10, and 16.66 s predicted by LFPQMOM along with their analytical counterparts for comparison. The numerical PSDs were in excellent agreement with the analytical PSDs on all the time points sampled. The computational time was about 46,042, 2818, and 996 ms for LFPQMOM, QMOM, and FPQMOM, respectively, in this test.
4.2. Pure Breakage
Three test cases were conducted for the pure breakage process. In the first test case, constant breakage kernel was adopted, i.e.,
with
C0 = 0.1. Uniform distribution was used as the daughter distribution
Using Equation (36) with
N0 = 1 and
V0 = 1 as the initial distribution, the analytical solution for the moments was given by [
22]
Unfortunately, analytical PSD was not available in this test case.
Figure 13 shows the evolution of the relative errors for the first six moments predicted by: (A) LFPQMOM; (B) QMOM; and (C) FPQMOM. It was observed that the evolutions of the percentage errors of LFPQMOM and QMOM were very similar. The relative errors rose sharply initially and leveled off after 15 s. Lower rank moments had bigger numerical errors and within the simulation time, the relative errors for all the moments were lower than 2 × 10
−9%. The percentage errors for
m3 were not included in the figure due to their low values. Similar to the previous cases,
m3 could be predicted without any error with the new method. Numerical predictions with FPQMOM were the best in this test.
Figure 14 depicts the particle size distribution at 0, 2, 10, and 50 s. Due to lack of analytical PSD, a comparison of the numerical PSDs with their analytical counterparts was impossible, and thus, in this test case, only numerical PSDs are given at 2, 10, and 50 s. The computational time was about 14,850, 1037, and 345 ms for LFPQMOM, QMOM, and FPQMOM, respectively, in this test.
In the second test case for breakage process, linear breakage kernel was used, i.e.,
with
a0 = 0.1, together with a uniform daughter distribution, i.e., Equation (47), exponential initial distribution, i.e., Equation (36) with
N0 = 1 and
V0 = 1. Analytical solution was given by [
22]
Moments can be evaluated by numerical integration or direct integration analytically. Percentage errors for the first six moments—predicted by: (A) LFPQMOM; (B) QMOM; and (C) FPQMOM—are shown in
Figure 15. In
Figure 15B, the relative errors for the moments predicted by QMOM rose sharply at incipient time, especially for larger rank moments,
m4 and
m5 for instance. However, with the new method, the relative errors rose relatively slowly at incipient time, as can be seen in
Figure 15A. With the simulation time, the maximum error for all the moments with FPQMOM was 1.88% whereas with QMOM, the maximum was 3.66%. With the new method, the numerical accuracy was improved relative to QMOM in this test case. The relative errors for
m0 were comparable. Again, with the new method,
m1 was predicted without any error, but not with QMOM. Relative errors for
m1 were not included in the figure. FPQMOM was again the best in numerical prediction, and the relative errors for all the moments were less than 0.1% within the simulation time.
Figure 16 gives the particle size distribution at 0, 2, 5, and 10 s, from which we observed that the numerical predictions with the new method agreed very well with the analytical PSDs at the time sampled. The computational time was about 3020, 190, and 66 ms for LFPQMOM, QMOM, and FPQMOM, respectively, in this test.
In the third test case for pure breakage, the power kernel was used as the breakage kernel having the form
with
C0 = 0.01. Uniform distribution was used for the daughter distribution, i.e., Equation (47), together with the exponential initial distribution, i.e., Equation (36) with
N0 = 1 and
V0 = 1. Analytical solution was given by [
22]
The numerical method was used to evaluate the moments. Evolutions of the relative errors for the first six moments are shown in
Figure 17A,B for LFPQMOM and QMOM, respectively. In the figure, it was observed that, with the new method, all the relative errors were lower than 0.3% before the time of 20 s, and then fluctuated for
m4 and
m5, and rose for
m0–
m3. Within the simulation time, all the relative errors were lower than 2%. However, with QMOM, all the relative errors for the moments except
m0 and
m1 rose up to 0.1 within 5 s initially. Within the simulation time, the maximum percentage error for all the moments was 13.8%. With the new method, the numerical accuracy was improved in this test case. Again, LFPQMOM made an exact prediction for
m1, similar to the previous cases. The relative errors for
m1 were not included in the figure. Again, LFPQMOM had the smallest errors. Similar to the third case for the aggregation process, the sharp drops in the relative errors were related to the numerical evaluation of the moments.
Figure 18 depicts the particle size distribution predicted by LFPQMOM at 0, 10, 20, and 50 s along with analytical counterparts for comparison. The numerical predictions exactly agreed with the analytical PSDs at different time points. The computational time was about 15,050, 876, and 338 ms for LFPQMOM, QMOM, and FPQMOM, respectively, in this test.
4.3. Aggregation and Breakage
Two test cases were conducted to validate the new method for the aggregation and breakage combined processes. The resulting source for Equation (6) is the summation of Equations (28) and (31). In the first test case, a constant kernel for aggregation, i.e., Equation (35) with
C0 = 1, a linear kernel for breakage, i.e., Equation (49) with
a0 = 0.1, and uniform daughter distribution, i.e., Equation (47), along with the following initial particle size distribution were adopted
with
N0 = 1,
V0 = 1. In this test case, analytical PSD was available [
23]
with Φ(
τ) = Φ(∞)[1 + Φ(∞)tanh(Φ(∞)
τ/2)]/[Φ(∞) + tanh(Φ(∞)
τ/2)], Φ(∞) = (2
a0V0/
C0)
1/2/
N0, and
τ =
C0N0t. Moments could be evaluated analytically. Evolutions of percentage errors for the moments are shown in
Figure 19 with: (A) LFPQMOM; (B) QMOM; and (C) FPQMOM. In the figure, the evolutions for the relative errors were very similar with the three methods. Initially, all the relative errors rose sharply and leveled off after the time of 20 s, except the errors for
m3, which had a continuous drop as the time advanced. With the new method and FPQMOM, there were two fluctuations in the drop of relative errors for
m3, but not with QMOM. Within the simulation time, the maximum relative error with both methods were comparable. In the figure, the relative errors for
m0 and
m1 were not included due to their low values.
Figure 20 gives the particle size distributions at 0, 2, 10, and 50 s. It was observed that the numerical PSDs agreed very well with the analytical PSDs. Note that the PSD at 10 s coincided with the PSD at 50 s, proving that a steady state had been reached by the time of 10 s. Note that, with different physical parameters, the steady state can be different. The computational time was about 497,950, 32,275, and 10,374 ms for LFPQMOM, QMOM, and FPQMOM, respectively, in this test.
In the second test case for the aggregation and breakage combined process, a constant aggregation kernel was used, i.e., Equation (35) with
C0 = 1, a linear kernel for breakage, i.e., Equation (49) with
a0 = 0.25, and uniform daughter distribution, i.e., Equation (47), along with the following initial particle size distribution were adopted
with
N0 = 1 and
V0 = 2. When the following relation was satisfied, the analytical solution was available [
23]
The analytical solution for particle size distribution was given by [
23]
with
K1(
τ) = 7 +
τ + exp(−
τ),
K2(
τ) = 2 − 2exp(−
τ),
L2(
τ) = 9 +
τ − exp(−
τ),
p1,2 = 0.25(e
−t−τ − 9 ± (
d(
τ))
0.5)
, d(
τ) =
τ2 + (10
τ − 2
τe
−τ) + 25 − 26e
−τ + e
−2τ,
τ = C0N0t. The moments can be evaluated analytically.
Figure 21 depicts the evolutions of percentage errors for
m2–
m5 with: (A) LFPQMOM; (B) QMOM; and (C) FPQMOM. The figure is very similar to the previous case. Initially, the errors rose sharply and leveled off for
m2,
m4, and
m5, but decreased for
m3 continuously. There was one difference from the previous case: the maximum percentage error was 0.42% with the new method, and 1.00% with QMOM. The numerical accuracy was improved with the new method relative to QMOM, which was similar to FPQMOM. Relative errors for
m0 and
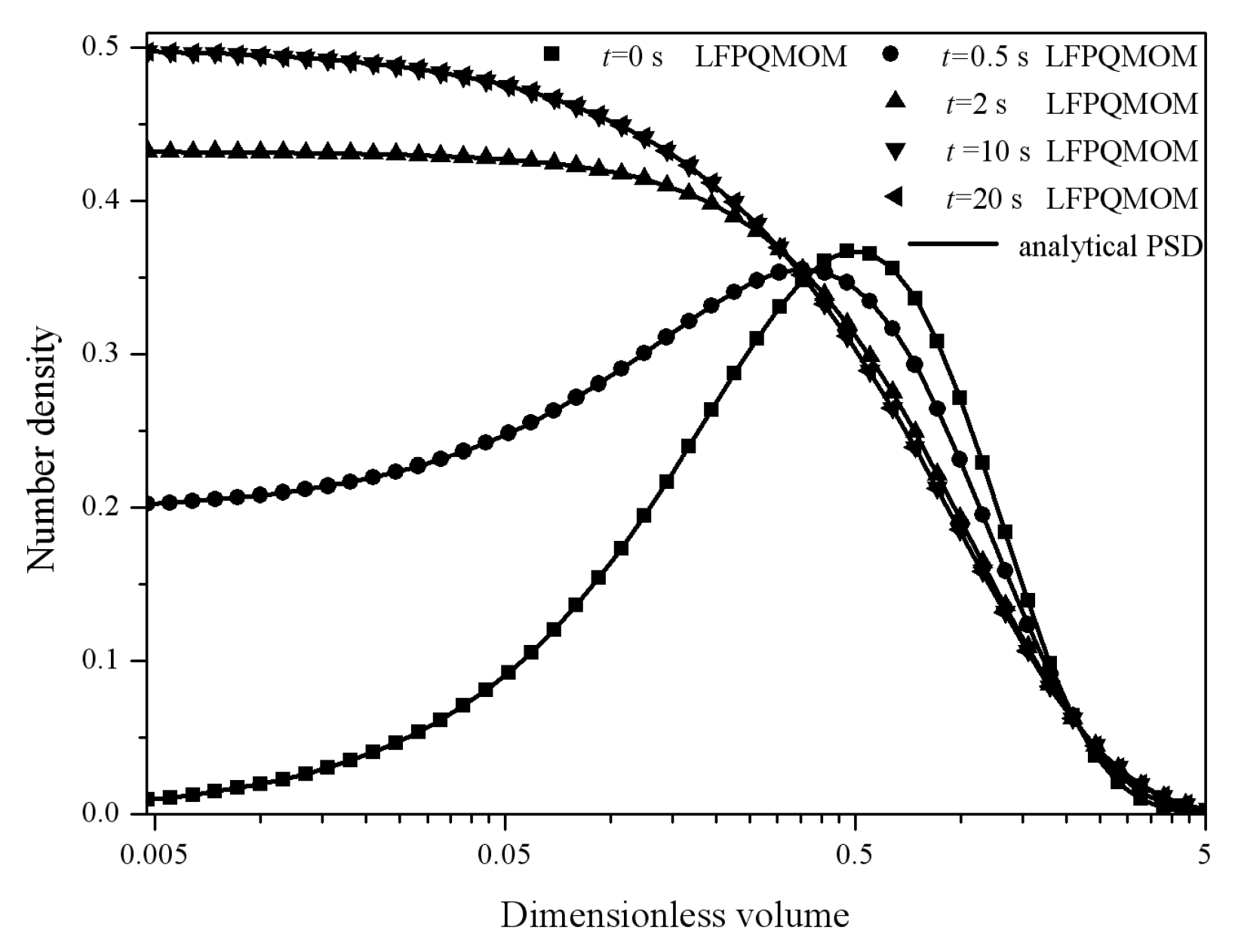
m1 were not included in the figure given their low values. The numerical and the analytical particle size distributions at 0, 0.5, 2, 10, and 20 s are shown in
Figure 22. The numerical PSDs agreed well with the analytical PSDs. The PSDs at 10 and 20 s coincided with each other, demonstrating that, by the time of 10 s, a steady state had been reached. Initially, the PSD was a Gaussian distribution, but, as the time advanced, the distributions became exponential, as can be seen in the figure. With the other parameters, the particle size distributions and the steady state could be different. The computational time was about 198,260, 12,048, and 4198 ms for LFPQMOM, QMOM, and FPQMOM, respectively, in this test.