Multi-Objective Optimization of Experiments Using Curvature and Fisher Information Matrix
Abstract
1. Introduction
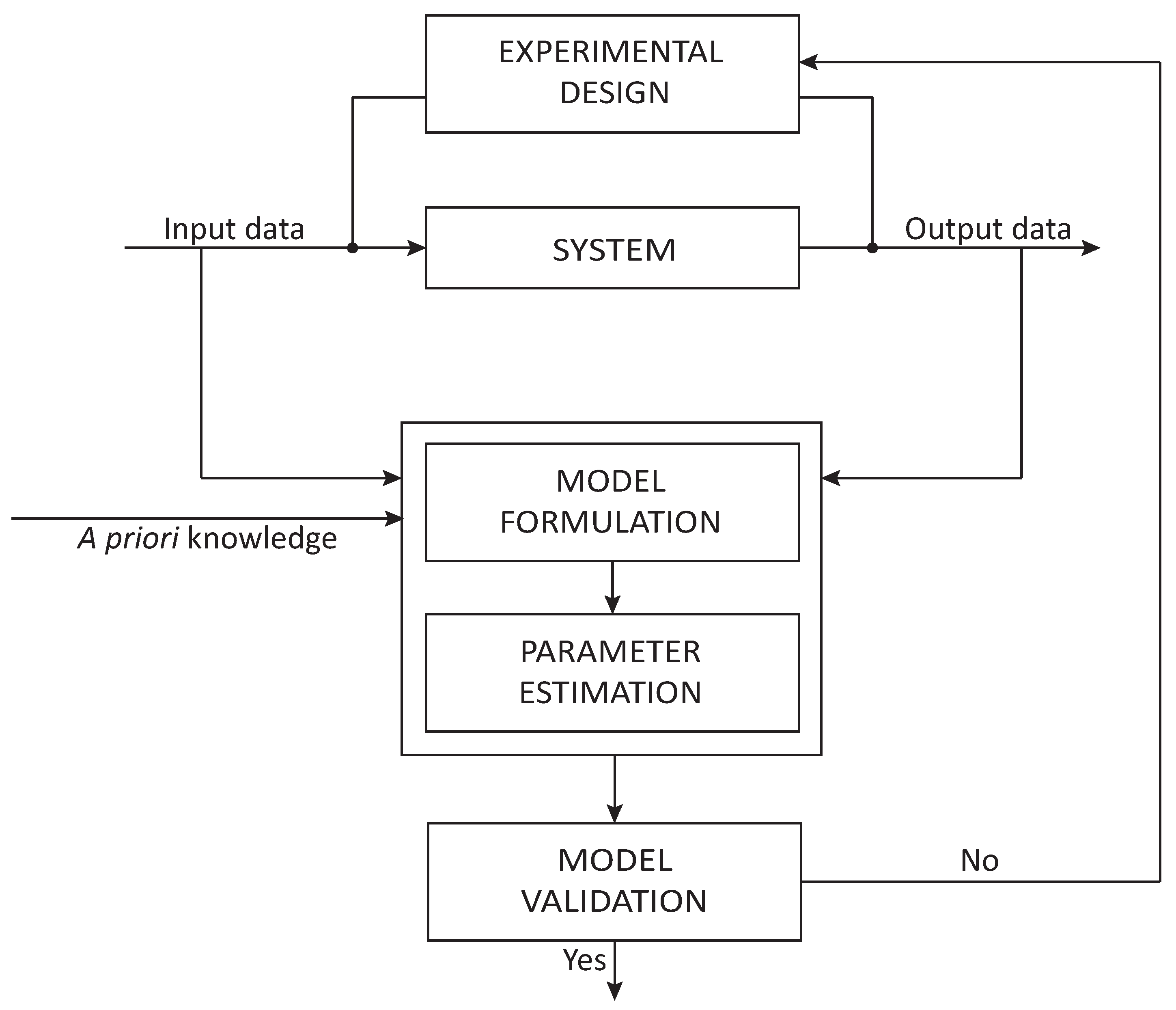
2. Model-Based Optimal Design of Experiments
2.1. Multi-Objective Design of Experiments Based on Curvatures
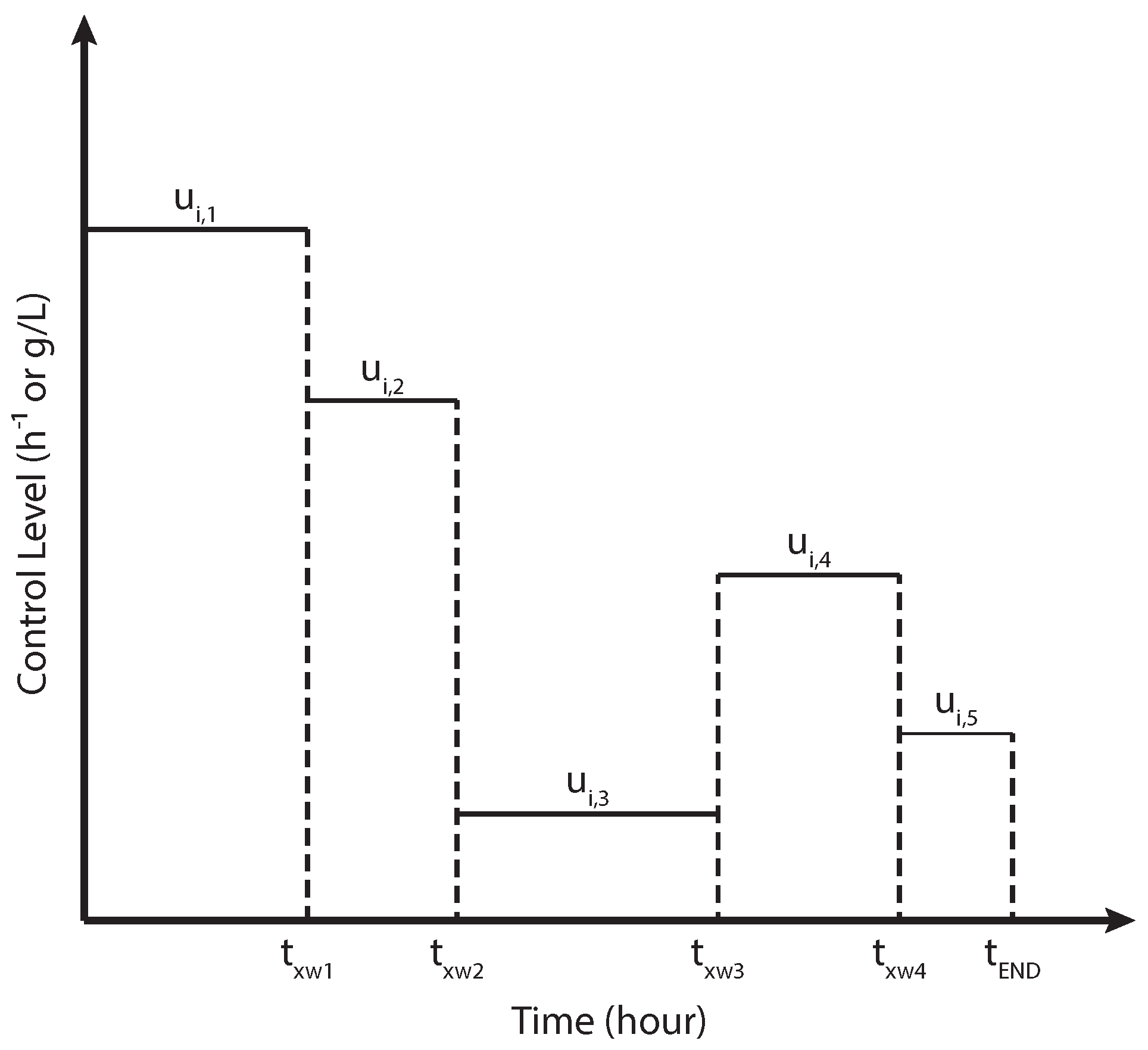
2.2. Numerical Implementation of the Curvature-Based MOO Design
3. Results
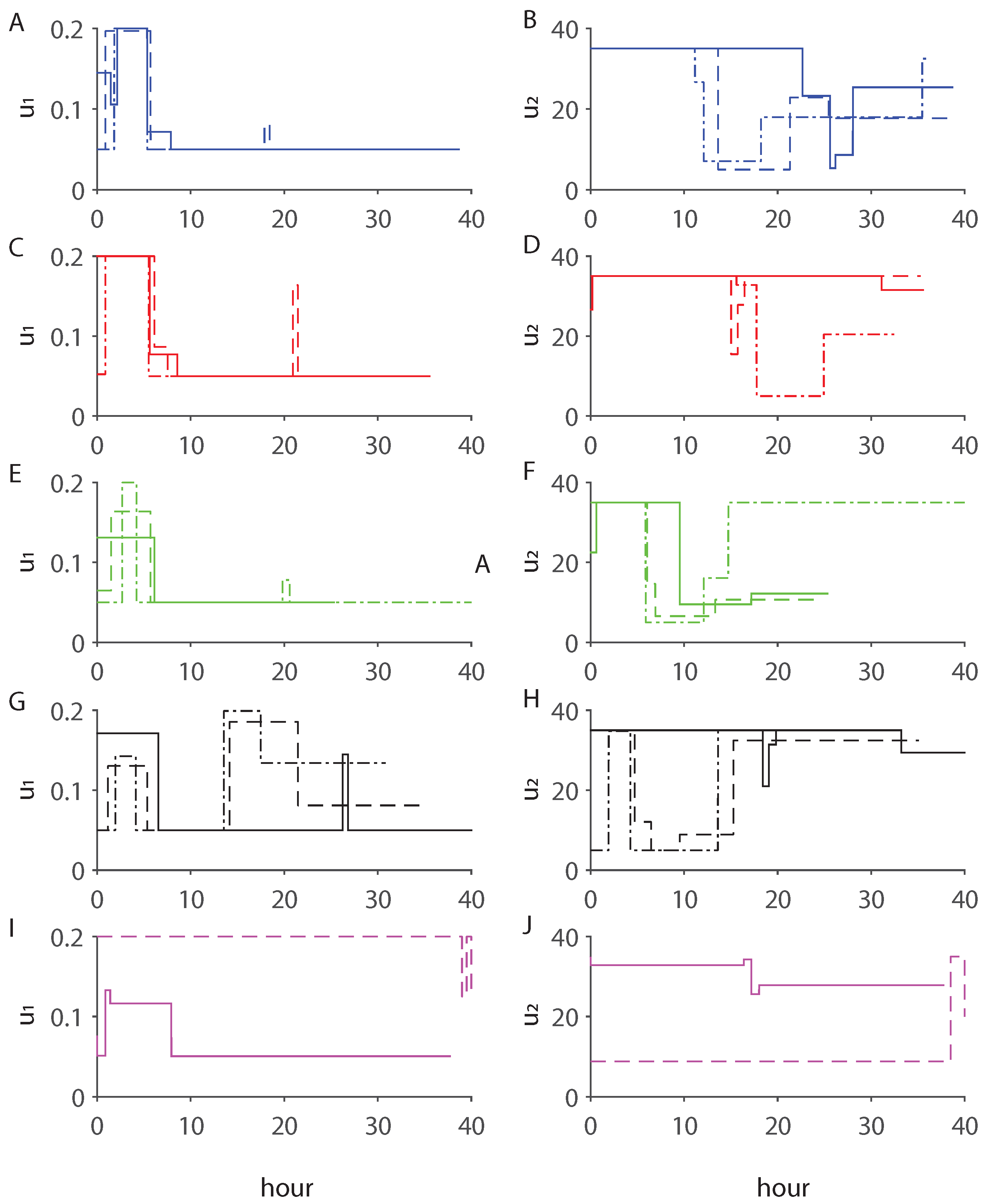
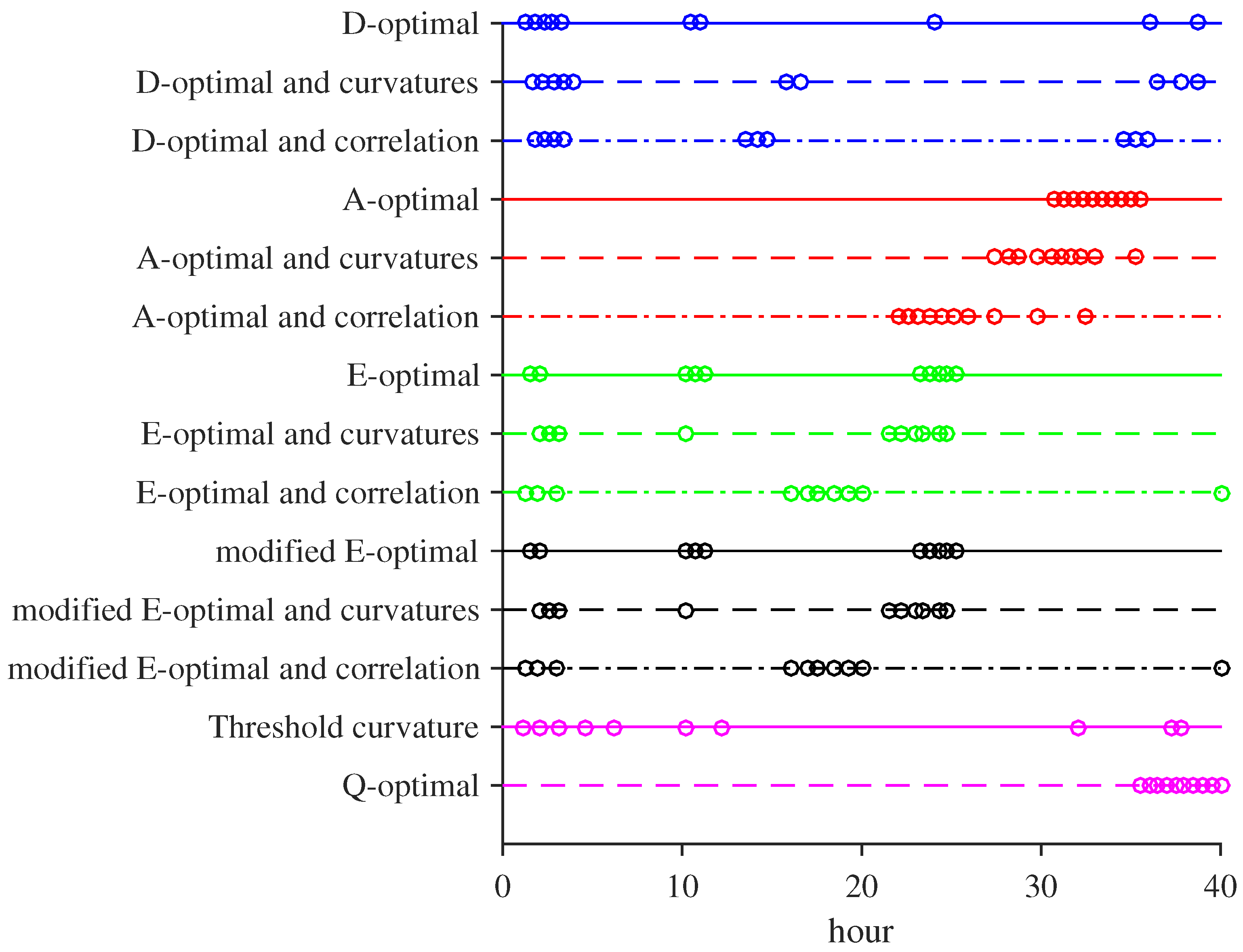
3.1. MBDOEs of Baker Yeast Fermentation Model
3.2. Performance Evaluation
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| MBDOE | Model-based design of experiments |
| FIM | Fisher information matrix |
| MOO | Multi-objective optimization |
| RMS | Root mean square |
| ODE | Ordinary differential equation |
| MLE | Maximum likelihood estimator |
| CVP | Control vector parametrization |
| nMSE | Normalized mean-square error |
References
- Srinath, S.; Gunawan, R. Parameter identifiability of power-law biochemical system models. J. Biotechnol. 2010, 149, 132–140. [Google Scholar] [CrossRef] [PubMed]
- Jia, G.; Stephanopoulos, G.; Gunawan, R. Ensemble kinetic modeling of metabolic networks from dynamic metabolic profiles. Metabolites 2012, 2, 891–912. [Google Scholar] [CrossRef] [PubMed]
- Gábor, A.; Hangos, K.; Banga, J.; Szederkànyi, G. Reaction network realizations of rational biochemical systems and their structural properties. J. Math. Chem. 2015, 53, 1657–1686. [Google Scholar] [CrossRef]
- Liu, Y.; Manesso, E.; Gunawan, R. REDEMPTION: Reduced dimension ensemble modeling and parameter estimation. Bioinformatics 2015, 31, 3387–3389. [Google Scholar] [CrossRef] [PubMed]
- Villaverde, A.F.; Banga, J.R. Structural properties of dynamic systems biology models: Idenfiability, reachability and initial conditions. Processes 2017, 5, 29. [Google Scholar] [CrossRef]
- Franceschini, G.; Macchietto, S. Model-based design of experiments for parameter precision: State of the art. Chem. Eng. Sci. 2008, 63, 4846–4872. [Google Scholar] [CrossRef]
- Kreutz, C.; Timmer, J. Systems biology: Experimental design. FEBS J. 2009, 276, 923–942. [Google Scholar] [CrossRef] [PubMed]
- Chakrabarty, A.; Buzzard, G.T.; Rundell, A.E. Model-based design of experiments for cellular processes. WIREs Syst. Biol. Med. 2013, 5, 181–203. [Google Scholar] [CrossRef] [PubMed]
- Cover, T.M.; Thomas, J.A. Elements of Information Theory, 2nd ed.; John Wiley & Sons: Hoboken, NJ, USA, 2006. [Google Scholar]
- Faller, D.; Klingmüller, U.; Timmer, J. Simulation methods for optimal experimental design in systems biology. Simulation 2003, 79, 717–725. [Google Scholar] [CrossRef]
- Gadkar, K.G.; Gunawan, R.; Doyle, F.J., III. Iterative approach to model identification of biological networks. BMC Bioinformatics 2005, 6, 155. [Google Scholar] [CrossRef] [PubMed]
- Balsa-Canto, E.; Alonso, A.A.; Banga, J.R. Computational procedures for optimal experimental design in biological systems. IET Syst. Biol. 2008, 2, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Chung, M.; Haber, E. Experimental design for biological systems. SIAM J. Control Optim. 2012, 50, 471–489. [Google Scholar] [CrossRef]
- Transtrum, M.K.; Qiu, P. Optimal experiment selection for parameter estimation in biological differential equation models. BMC Bioinform. 2012, 13, 181. [Google Scholar] [CrossRef] [PubMed]
- Maheshwari, V.; Rangaiah, G.P.; Samavedham, L. A multi-objective framework for model based design of experiments to improve parameter precision and minimize parameter correlation. Ind. Eng. Chem. Res. 2013, 52, 8289–8304. [Google Scholar] [CrossRef]
- Sinkoe, A.; Hahn, J. Optimal experimental design for parameter estimation of an IL-6 signaling model. Processes 2017, 5, 49. [Google Scholar] [CrossRef]
- Vanlier, J.; Tiemann, C.A.; Hilbers, P.A.J.; van Riel, N.A.W. A Bayesian approach to targeted experiment design. Bioinformatics 2012, 28, 1136–1142. [Google Scholar] [CrossRef] [PubMed]
- Weber, P.; Kramer, A.; Dingler, C.; Radde, N. Trajectory-oriented Bayesian experiment design versus Fisher A-optimal design: An in-depth comparison study. Bioinformatics 2012, 28, i535–i541. [Google Scholar] [CrossRef] [PubMed]
- Liepe, J.; Filippi, S.; Komorowski, M.; Stumpf, M.P.H. Maximizing the information content of experiments in systems biology. PLoS Comput. Biol. 2013, 9, e1002888. [Google Scholar] [CrossRef] [PubMed]
- Apgar, J.F.; Toettcher, J.E.; Endy, D.; White, F.M.; Tidor, B. Stimulus design for model selection and validation in cell signaling. PLoS Comput. Biol. 2008, 4, e30. [Google Scholar] [CrossRef] [PubMed]
- Daunizeau, J.; Preuschoff, K.; Friston, K.; Stephan, K. Optimizing experimental design for comparing models of brain function. PLoS Comput. Biol. 2011, 7, e1002280. [Google Scholar] [CrossRef] [PubMed]
- Flassig, R.J.; Sundmacher, K. Optimal design of stimulus experiments for robust discrimination of biochemical reaction networks. Bioinformatics 2012, 28, 3089–3096. [Google Scholar] [CrossRef] [PubMed]
- Busetto, A.G.; Hauser, A.; Krummenacher, G.; Sunnaker, M.; Dimopoulos, S.; Ong, C.S.; Stelling, J.; Buhmann, J.M. Near-optimal experimental design for model selection in systems biology. Bioinformatics 2013, 29, 2625–2632. [Google Scholar] [CrossRef] [PubMed]
- Silk, D.; Kirk, P.D.W.; Barnes, C.P.; Toni, T.; Stumpf, M.P.H. Model selection in systems biology depends on experimental design. PLoS Comput. Biol. 2014, 10, e1003650. [Google Scholar] [CrossRef] [PubMed]
- Bazil, J.N.; Buzzard, G.T.; Rundell, A.E. A global parallel model based design of experiments method to minimize model output uncertainty. Bull. Math. Biol. 2012, 74, 688–716. [Google Scholar] [CrossRef] [PubMed]
- Mdluli, T.; Buzzard, G.T.; Rundell, A.E. Efficient optimization of stimuli for model-based design of experiments to resolve dynamical uncertainty. PLoS Comput. Biol. 2015, 11, e1004488. [Google Scholar] [CrossRef] [PubMed]
- Cochran, W.G. Experiments for Nonlinear Functions. J. Am. Stat. Assoc. 1973, 68, 771–781. [Google Scholar] [CrossRef]
- Bates, D.M.; Watts, D.G. Relative Curvature Measures of Nonlinearity. J. R. Stat. Soc. Ser. B 1980, 42, 1–25. [Google Scholar]
- Hamilton, D.C.; Watts, D.G. A quadratic design criterion for precise estimation in nonlinear regression models. Technometrics 1985, 27, 241–250. [Google Scholar] [CrossRef]
- Benabbas, L.; Asprey, S.P.; Macchietto, S. Curvature-based methods for designing optimally informative experiments in multiresponse nonlinear dynamic situations. Ind. Eng. Chem. Res. 2005, 44, 7120–7131. [Google Scholar] [CrossRef]
- Asprey, S.P.; Macchietto, S. Statistical tools for optimal dynamic model building. Comput. Chem. Eng. 2000, 24, 1261–1267. [Google Scholar] [CrossRef]
- Seber, G.A.F.; Wild, C.J. Nonlinear Regression; John Wiley & Sons: Hoboken, NJ, USA, 2003. [Google Scholar]
- Merlé, Y.; Tod, M. Impact of pharmacokinetic-pharmacodynamic model linearization on the accuracy of population information matrix and optimal design. J. Pharmacokinet. Pharmacodyn. 2001, 28, 363–388. [Google Scholar] [CrossRef] [PubMed]
- Bogacka, B.; Wright, F. Comparison of two design optimality criteria applied to a nonlinear model. J. Biopharm. Stat. 2004, 14, 909–930. [Google Scholar] [CrossRef] [PubMed]
- Rangaiah, G.P. Multi-Objective Optimization: Techniques and Applications in Chemical Engineering; World Scientific: Singapore, 2008; Volume 1. [Google Scholar]
- Varma, A.; Morbidelli, M.; Wu, H. Parametric Sensitivity in Chemical Systems; Cambridge University Press: Cambridge, UK, 1999. [Google Scholar]
- Zhang, T.; Golub, G.H. Rank-One Approximation to High Order Tensors. SIAM J. Matrix Anal. Appl. 2001, 23, 534–550. [Google Scholar] [CrossRef]
- Deb, K.; Pratap, A.; Agarwal, S.; Meyarivan, T. A fast and elitist multi-objective genetic algorithm: NSGA-II. IEEE Trans. Evol. Comput. 2002, 6, 182–197. [Google Scholar] [CrossRef]
- Egea, J.A.; Martí, R.; Banga, J.R. An evolutionary method for complex-process optimization. Comput. Oper. Res. 2010, 37, 315–324. [Google Scholar] [CrossRef]
- Egea, J.A.; Rodriguez-Fernandez, M.; Banga, J.R.; Martí, R. Scatter search for chemical and bioprocess optimization. J. Glob. Optim. 2007, 37, 481–503. [Google Scholar] [CrossRef]
- Rodriguez-Fernandez, M.; Egea, J.A.; Banga, J.R. Novel metaheuristic for parameter estimation in nonlinear dynamic biological systems. BMC Bioinform. 2006, 7, 483. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Edgar, T.F. PCA combined model-based design of experiments (DOE) criteria for differential and algebraic system parameter estimation. Ind. Eng. Chem. Res. 2008, 47, 7772–7783. [Google Scholar] [CrossRef]
- Chou, I.C.; Voit, E. Recent developments in parameter estimation and structure identification of biochemical and genomic systems. Math. Biosci. 2009, 219, 57–83. [Google Scholar] [CrossRef] [PubMed]
- Voit, E. Biochemical systems theory: A review. ISRN Biomath. 2013, 2013, 897658. [Google Scholar] [CrossRef]
- White, A.; Tolman, M.; Thames, H.D.; Withers, H.R.; Mason, K.A.; Transtrum, M.K. The limitations of model-based experimental design and parameter estimation in sloppy systems. PLoS Comput. Biol. 2016, 12, e1005227. [Google Scholar] [CrossRef] [PubMed]
- Transturm, M.K.; Qiu, P. Model reduction by manifold boundaries. Phys. Rev. Lett. 2014, 113, 098701. [Google Scholar] [CrossRef] [PubMed]
- Transturm, M.K.; Qiu, P. Bridging mechanistic and phenomenological models of complex biological systems. PLoS Comput. Biol. 2016, 12, e1004915. [Google Scholar] [CrossRef] [PubMed]
| FIM-Based MBDOE | Criterion |
|---|---|
| D-optimal | max |
| A-optimal | max |
| E-optimal | max |
| Modified E-optimal | max |
| Design Criterion | |
|---|---|
| D-optimal | 10.0 |
| MOO D-optimal and curvatures | 10.0 |
| MOO D-optimal and correlation | 10.0 |
| A-optimal | 10.0 |
| MOO A-optimal and curvatures | 9.9 |
| MOO A-optimal and correlation | 10.0 |
| E-optimal | 10.0 |
| MOO E-optimal and curvatures | 10.0 |
| MOO E-optimal and correlation | 10.0 |
| Modified E-optimal | 10.0 |
| MOO modified E-optimal and curvatures | 10.0 |
| MOO modified E-optimal and correlation | 10.0 |
| Threshold curvature | 8.2 |
| Q-optimal | 5.5 |
| Design Criterion | ± | ± | ± | ± | |
|---|---|---|---|---|---|
| D-optimal | 7.06 × 10 | 0.3107 ± 0.0102 | 0.1831 ± 0.0276 | 0.5505 ± 0.0125 | 0.0502 ± 0.0026 |
| MOO D-optimal and curvatures | 4.71 × 10 | 0.3099 ± 0.0056 | 0.1825 ± 0.0233 | 0.5496 ± 0.0099 | 0.0499 ± 0.0018 |
| MOO D-optimal and correlation | 5.36 × 10 | 0.3117 ± 0.0134 | 0.1781 ± 0.0151 | 0.5543 ± 0.0270 | 0.0508 ± 0.0049 |
| A-optimal | 2.35 × 10 | 0.3294 ± 0.0659 | 0.2399 ± 0.1387 | 0.5841 ± 0.1083 | 0.0558 ± 0.0181 |
| MOO A-optimal and curvatures | 1.42 | 0.3669 ± 0.0947 | 0.5267 ± 0.2230 | 0.5548 ± 0.1333 | 0.0510 ± 0.0244 |
| MOO A-optimal and correlation | 4.82 | 0.0863 ± 0.0499 | 0.8927 ± 0.2555 | 0.2879 ± 0.1928 | 0.0177 ± 0.0263 |
| E-optimal | 8.01 × 10 | 0.3180 ± 0.0420 | 0.2026 ± 0.0956 | 0.5473 ± 0.0159 | 0.0496 ± 0.0026 |
| MOO E-optimal and curvatures | 3.33 × 10 | 0.3083 ± 0.0095 | 0.1829 ± 0.0164 | 0.5502 ± 0.0183 | 0.0500 ± 0.0026 |
| MOO E-optimal and correlation | 8.19 × 10 | 0.3108 ± 0.0164 | 0.1824 ± 0.0213 | 0.5552 ± 0.0304 | 0.0509 ± 0.0055 |
| Modified E-optimal | 6.99 × 10 | 0.3137 ± 0.0165 | 0.1986 ± 0.0920 | 0.5498 ± 0.0144 | 0.0502 ± 0.0033 |
| MOO modified E-optimal and curvatures | 3.44 × 10 | 0.3095 ± 0.0036 | 0.1789 ± 0.0034 | 0.5491 ± 0.0073 | 0.0500 ± 0.0013 |
| MOO modified E-optimal and correlation | 2.27 × 10 | 0.3088 ± 0.0048 | 0.1820 ± 0.0160 | 0.5486 ± 0.0047 | 0.0496 ± 0.0013 |
| Threshold curvature | 1.29 × 10 | 0.3144 ± 0.0307 | 0.1857 ± 0.0339 | 0.5500 ± 0.0155 | 0.0502 ± 0.0032 |
| Q-optimal | 1.91 × 10 | 0.3085 ± 0.0178 | 0.1757 ± 0.0216 | 0.5514 ± 0.0236 | 0.0504 ± 0.0119 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manesso, E.; Sridharan, S.; Gunawan, R. Multi-Objective Optimization of Experiments Using Curvature and Fisher Information Matrix. Processes 2017, 5, 63. https://doi.org/10.3390/pr5040063
Manesso E, Sridharan S, Gunawan R. Multi-Objective Optimization of Experiments Using Curvature and Fisher Information Matrix. Processes. 2017; 5(4):63. https://doi.org/10.3390/pr5040063
Chicago/Turabian StyleManesso, Erica, Srinath Sridharan, and Rudiyanto Gunawan. 2017. "Multi-Objective Optimization of Experiments Using Curvature and Fisher Information Matrix" Processes 5, no. 4: 63. https://doi.org/10.3390/pr5040063
APA StyleManesso, E., Sridharan, S., & Gunawan, R. (2017). Multi-Objective Optimization of Experiments Using Curvature and Fisher Information Matrix. Processes, 5(4), 63. https://doi.org/10.3390/pr5040063





