1. Introduction
Fermentation sits at the heart of brewing. During fermentation, fermentable sugars in brewer’s wort are converted into ethanol, carbon dioxide, and a wide range of flavour-active secondary metabolites that define the organoleptic profile of the finished beer [
1,
2,
3]. In production vessels, fermentation dynamics are shaped by yeast physiology, recipe formulation, temperature control, and fermentation-vessel geometry. The growth of the modern craft sector has further expanded this design space through the exploration of novel yeasts and recipes, and the use of mixed-culture fermentations [
4,
5,
6]. Further, many breweries now operate with continuous inline sensing and automated data logging of apparent extract (°P) and fluid temperature, creating an opportunity for digital tools that anticipate fermentation behaviour rather than simply recording it.
Mechanistic models of beer fermentation typically describe changes in yeast biomass, fermentable sugars, and ethanol through coupled ordinary differential equations (ODEs), often incorporating substrate inhibition or oxygen effects [
7,
8,
9]. Such models have been used successfully for simulation and optimisation in specific breweries. However, such models are not robust to process changes, requiring key parameters to be tuned to empirical data, meaning recipe or yeast changes require reparametrisation for the model to perform optimally [
10]. In everyday operation, many breweries primarily measure the following parameters inline: fluid temperature, apparent extract (expressed in degrees Plato), and pH (less commonly measured inline) rather than the full state vector assumed in most kinetic formulations [
8]. In particular, biomass is rarely monitored inline, leading to a discrepancy between the data collected in practice and the values forecast by conventional models.
Data-driven approaches offer an alternative route. For fermentation and bioprocessing, neural networks and related methods have been applied to soft sensing, fault detection, and the prediction of biomass or product concentrations from routinely measured variables [
11,
12,
13]. Recent work has also explored hybrid models that combine mechanistic structure with flexible empirical components [
14,
15]. More broadly, there has been increasing interest in data-driven dynamical systems and the use of digital twins within process systems engineering [
16,
17]. Recurrent neural networks, and in particular long short-term memory (LSTM) architectures [
18,
19,
20], are also well suited to this context, as they can represent non-linear temporal dependencies without requiring ODEs derived from first principles.
In brewing, most reported neural-network models have been trained on relatively small datasets, often focusing on monoculture fermentations under tightly controlled conditions [
11]. There are fewer examples of a single data-driven model trained on an extensive, heterogeneous collection of fermentations spanning multiple beer styles and mixed-culture inocula. Yet this is precisely the situation encountered in practice: breweries accumulate archives of hundreds or thousands of fermentations from a diverse product line, collected under slightly different process conditions, and an effective plant model must cope with this diversity while still providing reliable predictions for individual fermentations. Studies such as “From Data to Draught” have shown that diverse fermentation datasets can, in principle, be leveraged to develop models robust to a range of temperature and inoculum configurations [
21]. Still, a robust process model trained on a large, diverse fermentation dataset from real-world commercial breweries remains needed.
The present work addresses this need by developing a data-driven plant model for beer fermentation using an LSTM-based architecture. The model is trained on a large dataset of inline-monitored fermentations recorded using a Sennos (US) BrewIQ IoT sensor array. This dataset is primarily comprised of fermentations carried out in commercial breweries and is augmented by some pilot-scale fermentations conducted in a laboratory setting. After filtering to include only ales, India pale ales (IPAs), lagers, and mixed-culture beers, 1305 fermentations were available for use in modelling. The dataset encompasses a broad range of recipes, yeast strains, and temperature profiles, resulting in diverse apparent extract and pH trajectories.
Using this dataset, we develop a multi-output, sequence-to-sequence LSTM plant model that forecasts apparent extract and pH over short horizons, conditioned on the intended future temperature schedule; full architectural and training details are provided later in the paper.
Model performance is assessed on rolling short-horizon forecast windows and through open-loop rollouts on representative fermentations in each beverage category, providing an interpretable view of fidelity in brewer-relevant terms.
The primary objectives of this work are to describe a large, stylistically diverse beer fermentation dataset suitable for data-driven modelling, to detail the design, training, and validation of a multi-output LSTM plant model that uses inline measurements of temperature, apparent extract, and pH as key modelling parameters, and to evaluate the model’s capability to replicate typical fermentations for ales, IPAs, lagers, and mixed-culture beers. The findings demonstrate that the LSTM plant model achieves root-mean-square prediction errors for apparent extract and pH that are comparable to standard sensor tolerances, particularly for pH, indicating that LSTM-based neural networks of this type can provide a practical foundation for monitoring and short-term forecasting of beer fermentation in modern breweries.
2. Materials and Methods
2.1. Experimental Facilities and Instrumentation
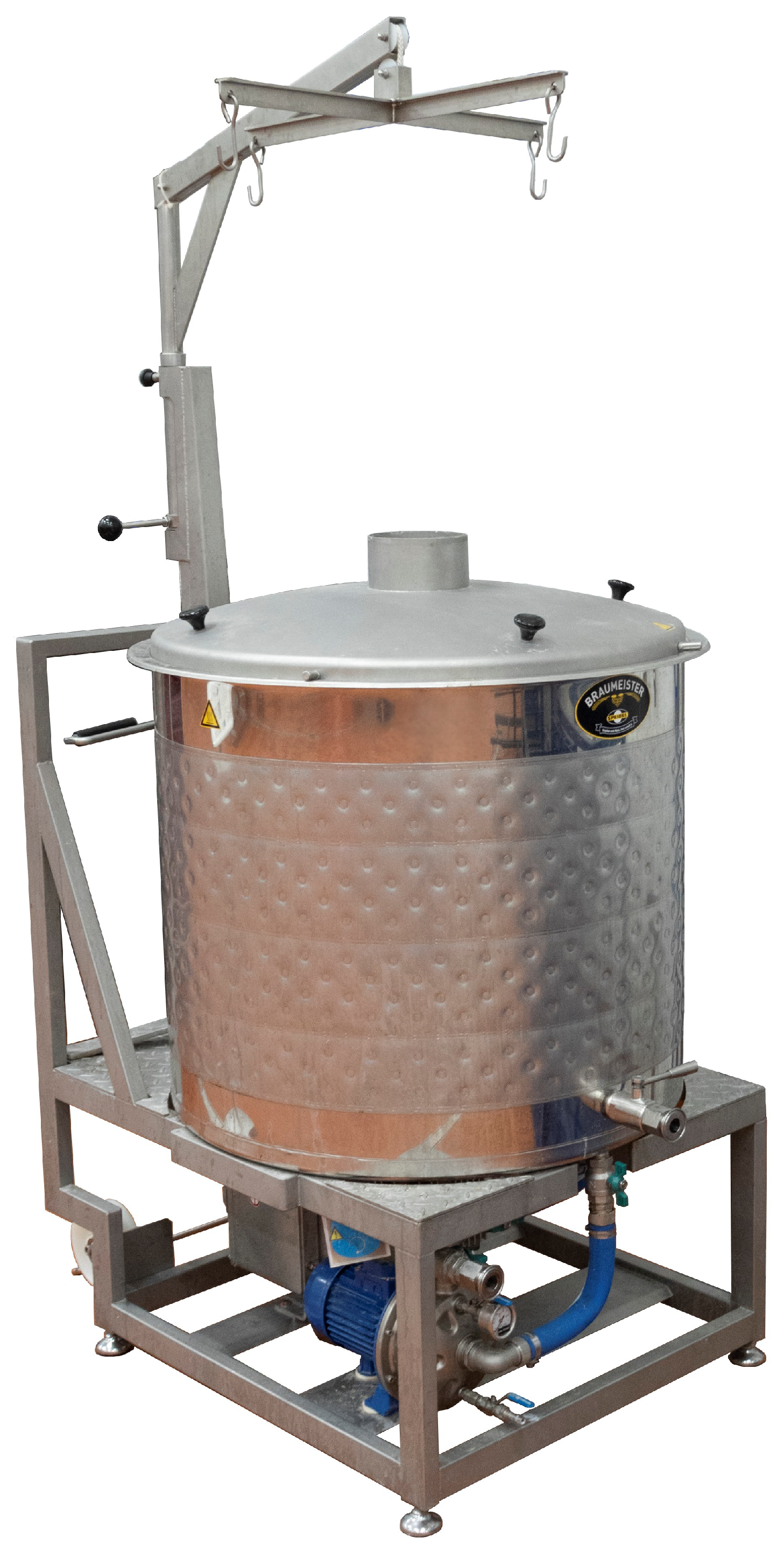
All pilot-scale fermentations were carried out at the Advanced Food Innovation Centre (AFIC) at Sheffield Hallam University (Sheffield, UK). Wort was produced using a 200 L Speidel’s Braumeister brewhouse (
Figure 1) and transferred to a 240 L fermenter (
Figure 2), which has been cleaned in place (CIP) and sanitised thoroughly prior to use (see
Figure 3). This in-house pilot brewery enabled the production of fermentations analogous to those in small craft breweries. However, the majority of the data are sourced from data collected in commercial breweries at a range of scales using a wide variety of recipes and equipment.
All inline measurements within the dataset were recorded using the Sennos (Durham, NC, USA) BrewIQ multi-sensor fermentation monitor, shown in
Figure 4. The device measures fluid temperature, specific gravity and apparent extract (°P), pH, dissolved oxygen, and conductivity, and streams data to a secure cloud platform at regular intervals. In this study, temperature, apparent extract, and pH are used directly in the modelling pipeline, while the remaining channels are omitted.
2.2. Wort Recipe, Yeast Inocula, and Operating Conditions
All pilot-scale batches were based on a single, IPA-style wort designed to provide a consistent substrate across fermentations. The grain bill comprised predominantly pale ale malt and Carapils, and hop additions chosen to produce a contemporary IPA aroma and bitterness profile (Cascade and Centennial). Using a standardised recipe isolates the influence of yeast strain, inoculum configuration, and temperature programme on the observed dynamics. This recipe was particularly important as these pilot-scale experiments constitute the overwhelming majority of data within the mixed-culture beers category.
Pilot-scale fermentations were performed by directly pitching active dry brewing yeast produced by Fermentis (Marquette-lez-Lille, France). Ale fermentations used a Saccharomyces cerevisiae strain suited to American pale ales and IPAs (Fermentis (Marquette-lez-Lille, France) SafAle US-05), while lager fermentations used Saccharomyces pastorianus (Fermentis (Marquette-lez-Lille, France) SafLager S-23). Mixed-culture beers combined ale and lager strains to create more complex flavour profiles.
Fermentations were grouped into four beverage categories: ales, IPAs, lagers, and mixed-culture beers. They were categorised based on the recorded style descriptor within the metadata of each fermentation batch.
Figure 5 illustrates three representative beers produced from the IPA-style wort using different yeast configurations at 22 °C. Even with identical wort, differences in yeast physiology and fermentation temperature yield distinct appearance and flavour profiles, reflected in the corresponding apparent extract and pH trajectories.
2.3. Dataset Construction and Preprocessing
Raw data were exported from the BrewIQ platform as comma-separated value files, each corresponding to a single fermentation. For every batch, the data included the following required attributes: the elapsed time since yeast pitch (hours_from_pitch, in hours), apparent extract (plato, in °P), fluid temperature (fluid_temp, in °C), and a style descriptor (style_name). Additional variables such as pH, dissolved oxygen, and conductivity were recorded for many fermentations, depending on the configuration used at each brewery.
The data were preprocessed and resampled before modelling. Batches lacking finite values for any of the required columns (time, Plato, temperature, style name) were discarded and two fermentations were removed on this basis. The remaining fermentations came from a mixture of pilot-scale trials and routine production in commercial breweries. The commercial data covered a wide variety of monoculture yeast inocula, temperature profiles, and recipes, including a broad range of ale and lager styles. Style names were normalised by stripping accents and diacritics, standardising capitalisation and spelling, and then mapped to beverage categories using rule-based pattern matching against curated lists. The following categories were identified: ales, IPAs, lagers, mixed-culture beers, stouts and porters, Belgian styles, sour beers, wheat beers, wines, seasonal and historical beers, flavoured and experimental beers, spirits, and uncategorised entries.
The LSTM plant model was trained on the four most populous categories: ales, IPAs, lagers, and mixed-culture beers. Restricting to these categories yielded 1305 fermentations in total. Within this modelling subset, the sampling period was 30 min, the time since pitch was capped at 300 h, and the median duration ranged from 141 h for ales to 300 h for lagers and mixed-culture fermentations. For each batch, original gravity descriptors were computed from the apparent extract (°P) time series. The initial gravity OGinit was defined as the first finite degrees Plato value at h where available. A reference original gravity, OGref, was then obtained as the median Plato over the first 2 h of the fermentation; this approach was chosen to mitigate against measurement noise.
Temperature setpoints were reconstructed from the fluid temperature time series to provide a smoothed representation of the brewer’s intended temperature programme. The recorded temperatures were first smoothed using a three-point centred moving average, then rounded to the nearest 0.5 °C. Changes smaller than 0.5 °C were treated as noise, while sequences of approximately constant temperature were treated as plateaux. The resulting piecewise-constant trajectory, denoted , captures the main steps of the temperature schedule while filtering sensor noise and short-lived perturbations.
Table 1 summarises the modelling dataset by beverage category. Ales and IPAs dominate in terms of batch count and time points, reflecting their prevalence in contemporary production. The number of lager and mixed-culture fermentations is smaller but still sufficient to provide meaningful coverage for modelling. Almost all ale, IPA, and lager fermentations included pH measurements, whereas pH coverage was lower in mixed-culture fermentations.
2.4. LSTM Plant Model Formulation
The plant model (see
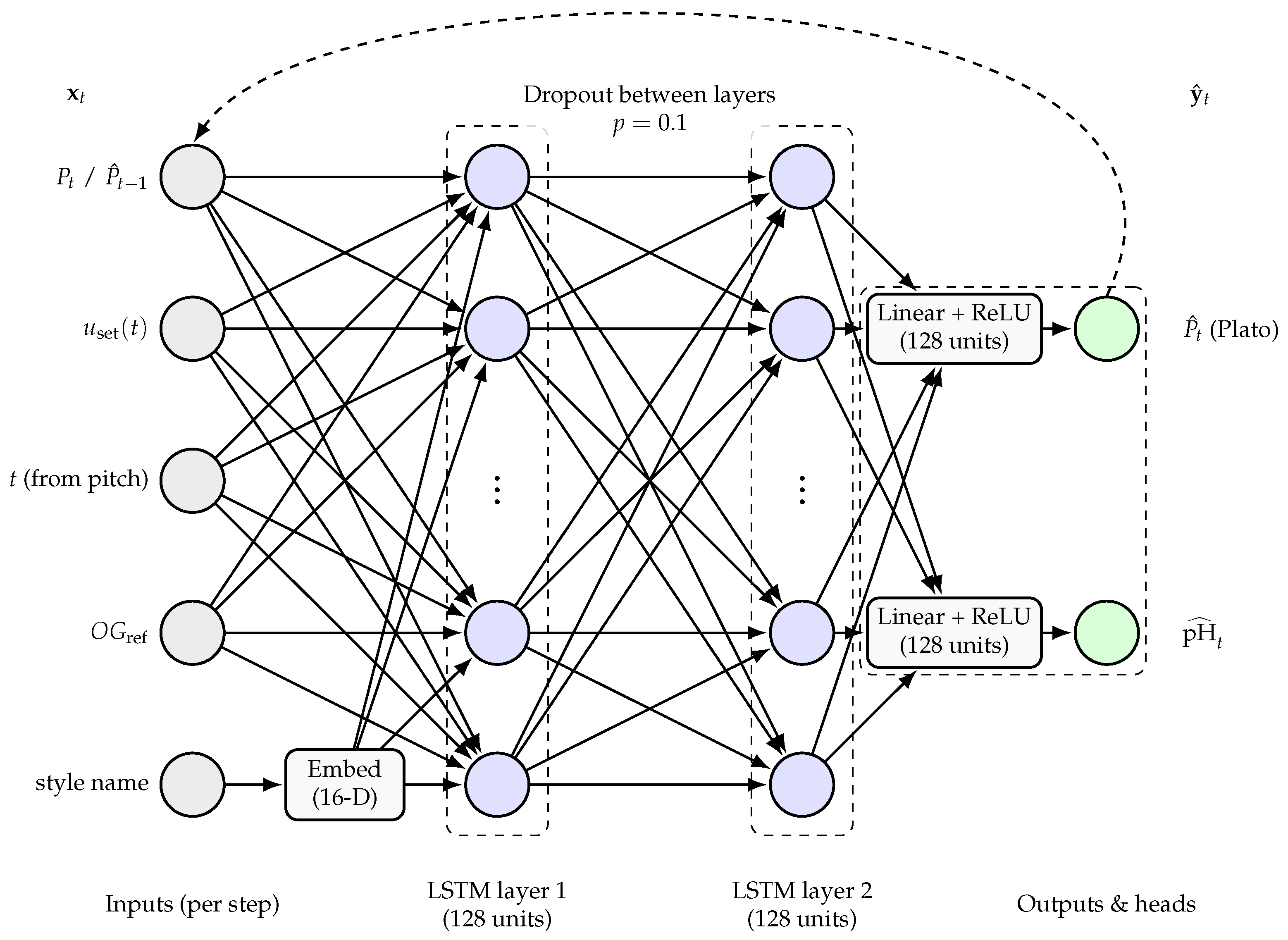
Figure 6) is formulated as a multi-output, sequence-to-sequence LSTM network implemented in PyTorch (version 2.9.1). Its purpose is to predict short-horizon trajectories of degrees Plato and pH given a recent history of process measurements and a planned future temperature schedule. This model structure was chosen as, in combination with an optimiser, it is suitable for direct integration within a model predictive control (MPC) controller.
The model operates at a fixed sampling interval of 0.5 h. For each fermentation, a history window of length h and a forecast horizon of length h are considered, yielding time steps per history and forecast segment. At each time step in the history window, the inputs to the model comprise the reconstructed temperature setpoint , the time since pitch, the original gravity descriptor (OGref), the current apparent extract value (°P), and a categorical beer style label. The style label is mapped to an integer index and embedded into a low-dimensional continuous vector through a learnable embedding layer. All continuous inputs are standardised using dataset-wide means and standard deviations, so that the network operates on inputs with approximately zero mean and unit variance per channel. The original gravity descriptor (OGref) is broadcast across time to give a constant sequence for each batch.
The recurrent backbone is a two-layer LSTM with 128 hidden units per layer. For the history window, the input sequence is formed by concatenating the scaled temperature, time, original gravity (OGref), apparent extract (°P), and style embedding at each time step, giving an input dimension of , where E is the 16-dimensional embedding vector. The LSTM processes this sequence and returns the sequence of hidden states, together with the final hidden and cell states. Two output heads, implemented as small feed-forward networks, map the hidden state at each forecast step to apparent extract and pH predictions, respectively. Both heads consist of a linear layer, a rectified linear unit activation, and a final linear layer that outputs a single scalar per time step.
Inference proceeds in two phases. First, the history window is passed through the LSTM to initialise the hidden and cell states. The last apparent extract value (°P) in the history segment is retained as the starting point for the autoregressive decoder. Second, the model rolls forward step by step over the forecast horizon. At each forecast step, the decoder forms an input vector by concatenating the scaled future temperature setpoint, future time, original gravity (OGref), the previously predicted degrees Plato value, and the style embedding. This input is passed through the same LSTM, using the hidden and cell states carried over from the previous step. The apparent extract (°P) and pH heads then produce predictions for the current forecast step, and the degrees Plato prediction is fed back as the apparent extract (°P) input for the next step. In this way, predicted apparent extract acts as a surrogate state variable in the decoder, allowing the network to propagate its own predictions forward in time.
2.5. Training Procedure and Loss Function
Before training, global scaling factors are estimated for each continuous variable. For every fermentation in the modelling subset, the sequences of temperature setpoint, time since pitch, original gravity (OGref), degrees Plato, and pH are concatenated, and a mean and standard deviation are computed across the combined dataset for each channel. These statistics are used to transform all inputs and targets to zero mean and unit variance. Scaling is applied identically to training and evaluation data.
Overlapping training examples are constructed by sliding a window across each fermentation. For each valid time index, a history segment of duration h and an immediately following forecast segment of duration h are extracted at 0.5 h resolution. The history contains scaled sequences of temperature setpoint, time since pitch, original gravity (OGref), degrees Plato, and the style embedding. The forecast contains scaled temperature, time, original gravity (OGref), and the corresponding degrees Plato and pH targets. Windows containing any missing values in the input series or in the Plato targets are discarded. Windows with missing pH values are retained, but the missing entries are marked in a binary pH mask. This approach allows the model to use incomplete pH information without discarding otherwise useful windows.
Noise augmentation is applied to the training inputs to improve robustness to measurement variation. During training, the reconstructed temperature setpoint and apparent extract sequences are perturbed by truncated Gaussian noise. For each of these variables x, a zero-mean Gaussian noise term is added with a standard deviation chosen so that three standard deviations match the nominal sensor tolerance (approximately °C for temperature and °P for Plato). The perturbation is clipped to lie within the corresponding tolerance band, and the result is further restricted to physically plausible limits of 0–30 °C for temperature and 0–20 °P for apparent extract. The perturbed sequences are then scaled and used as inputs, while the original, unperturbed apparent extract and pH sequences form the targets. This procedure mimics realistic sensor noise without corrupting the ground truth.
To construct training and validation sets, all admissible windows are first enumerated without augmentation. A random subset comprising 10% of these windows is assigned to the validation set, and the remaining 90% form the training window pool. For validation, the windows are used in their unaugmented form. For training, windows are regenerated with noise augmentation, using independent random draws at each epoch. Because the split is performed at the window level rather than the batch level, windows from the same fermentation can appear in both the training and validation sets. The resulting training and validation losses therefore quantify generalisation across overlapping windows drawn from the same population of fermentations, rather than strictly held-out fermentations.
The model is trained using the Adam optimiser with a learning rate of
, mini-batch size of 32, and 20 epochs. Gradients are clipped to a global norm of 1.0 to improve stability, and a dropout rate of 0.1 is applied between the two LSTM layers. These hyperparameters were chosen to balance model capacity, training stability, and computational cost, and are consistent with standard practice for recurrent neural networks in process modelling [
17,
22].
The training objective is a masked Huber loss. For a set of predictions
, targets
z, and a binary mask
m, we denote each prediction–target pair by
. With a threshold parameter
, the per-element Huber loss is defined as
and the masked Huber loss is the mean of
over all entries where
. In this work,
is used for both apparent extract and pH in scaled units. During training, the total loss for a mini-batch is
where
denotes the masked Huber loss with
,
is a mask of ones for the degrees Plato targets, and
is the pH mask. Apparent extract (degrees Plato), therefore, acts as the primary objective, with pH contributing a secondary but non-negligible term.
To encourage robustness to missing pH measurements, an additional dropout mechanism is applied to the pH mask during training. At each batch, a random binary matrix
D is sampled with entries drawn independently from a Bernoulli distribution with parameter
. The effective training mask for pH is then
where ⊙ denotes element-wise multiplication. This mechanism effectively turns off a proportion of pH targets during training, even where measurements are available, and encourages the network to infer pH dynamics from context rather than relying solely on densely labelled trajectories. At the end of each epoch, mean training and validation losses are recorded, and the model with the lowest validation loss is retained for subsequent analysis.
2.6. Medoid Selection and Full-Run Evaluation
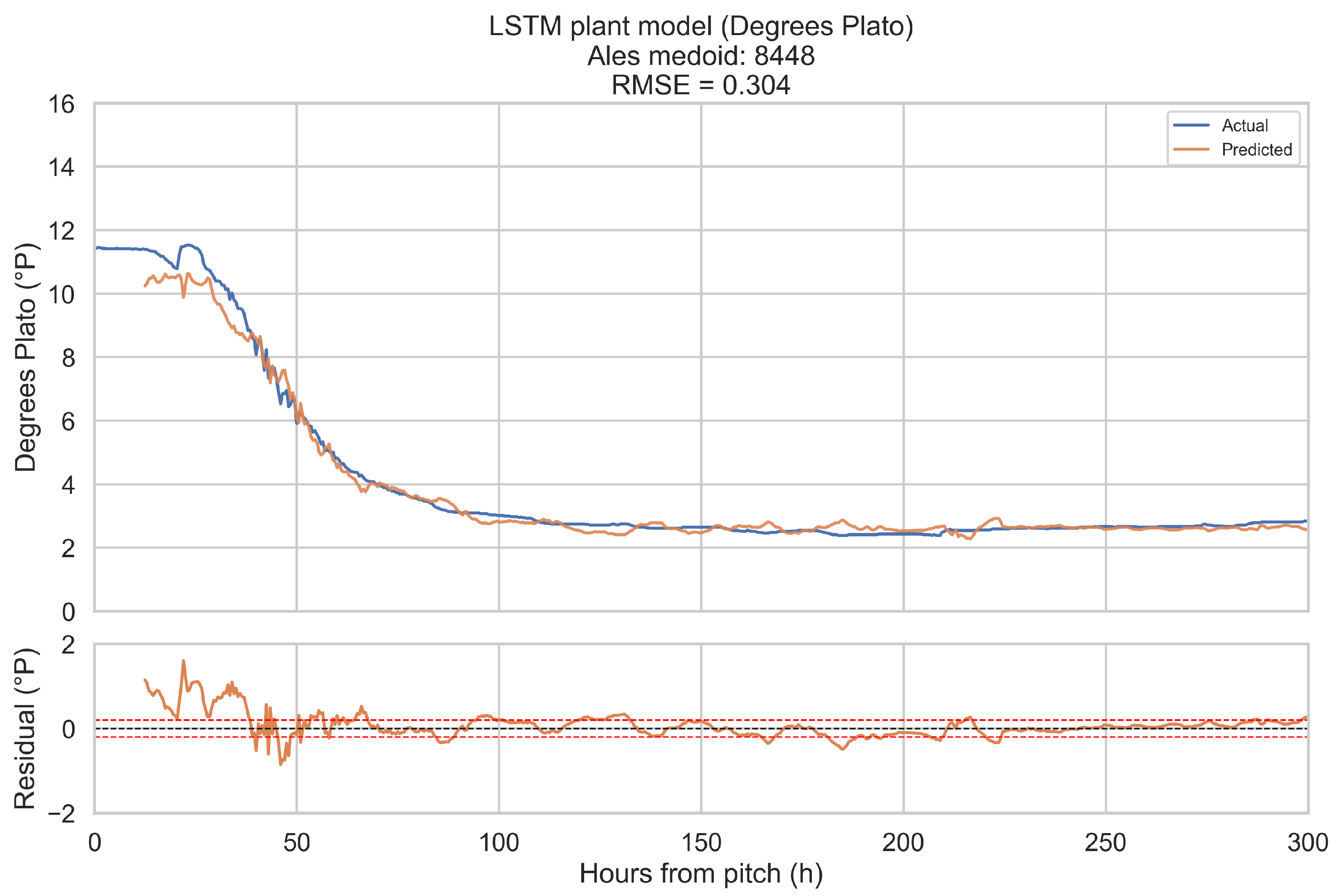
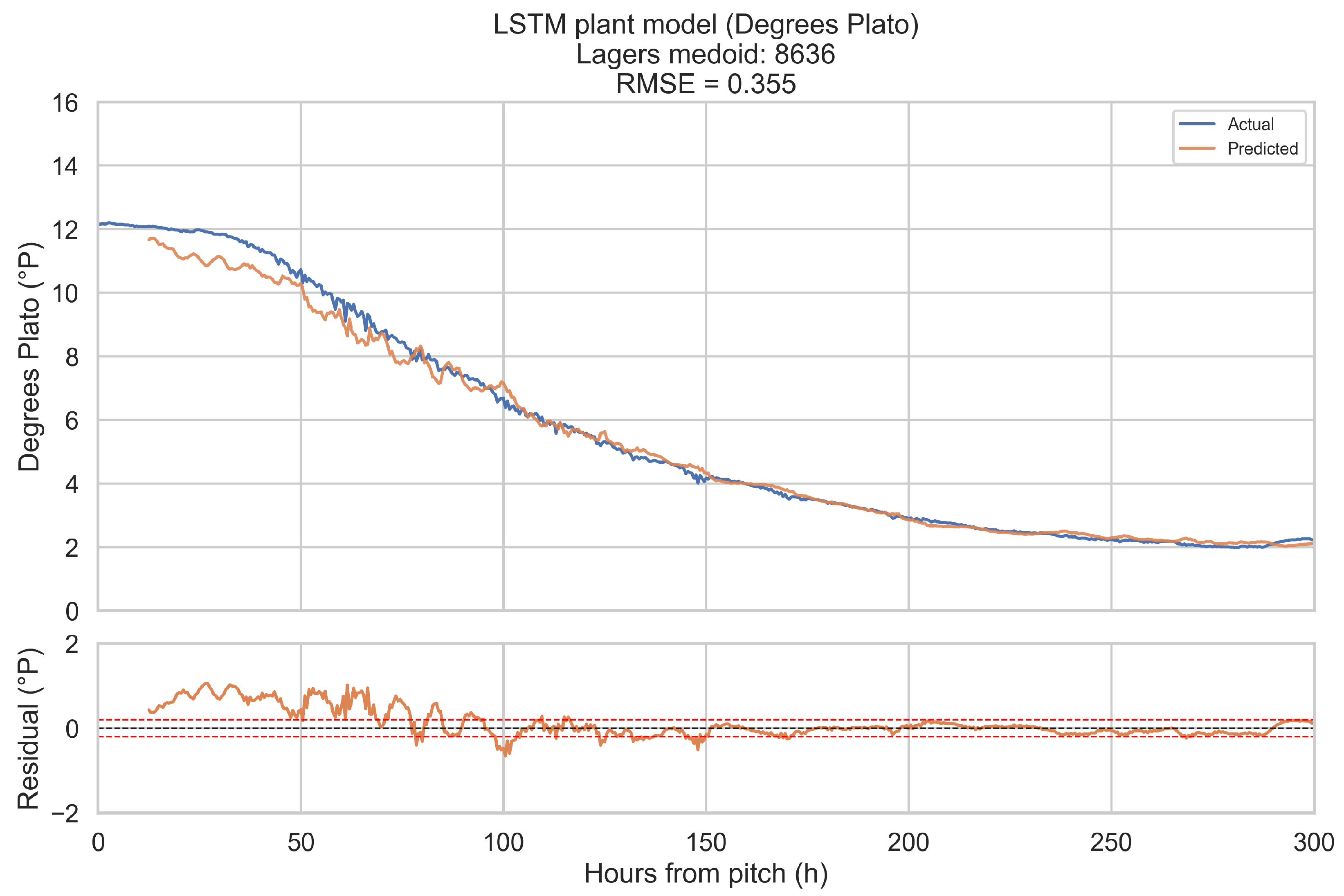
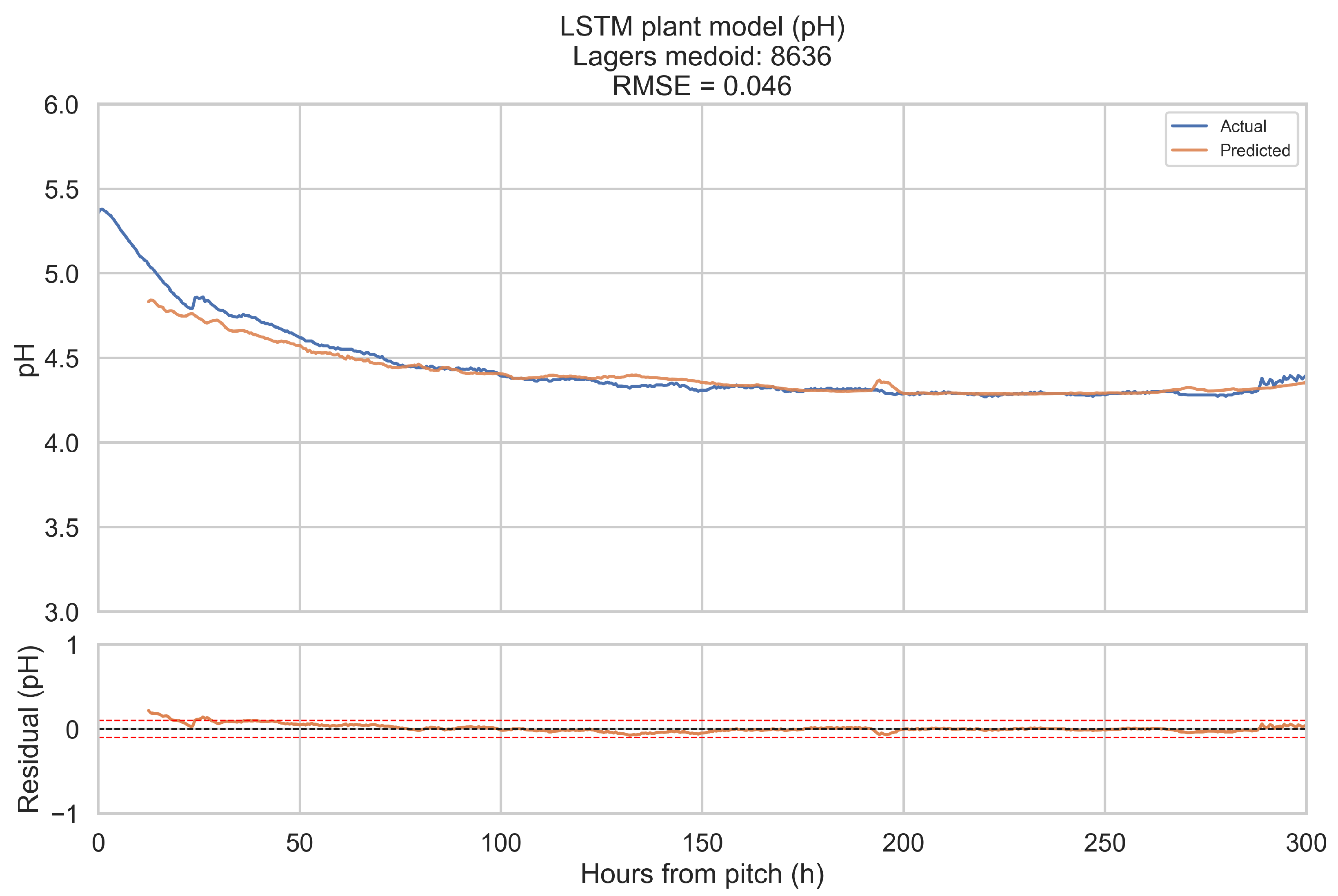
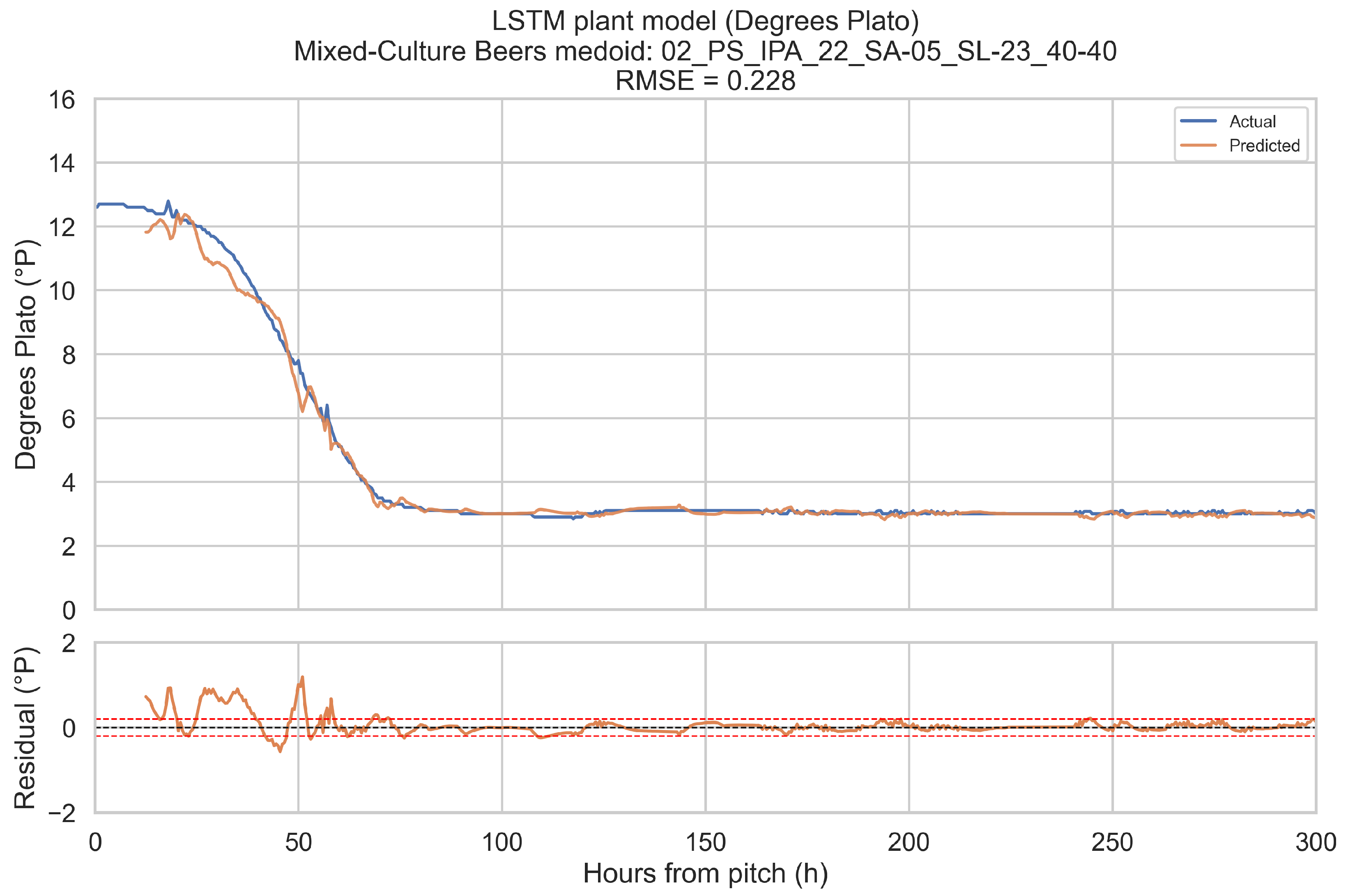
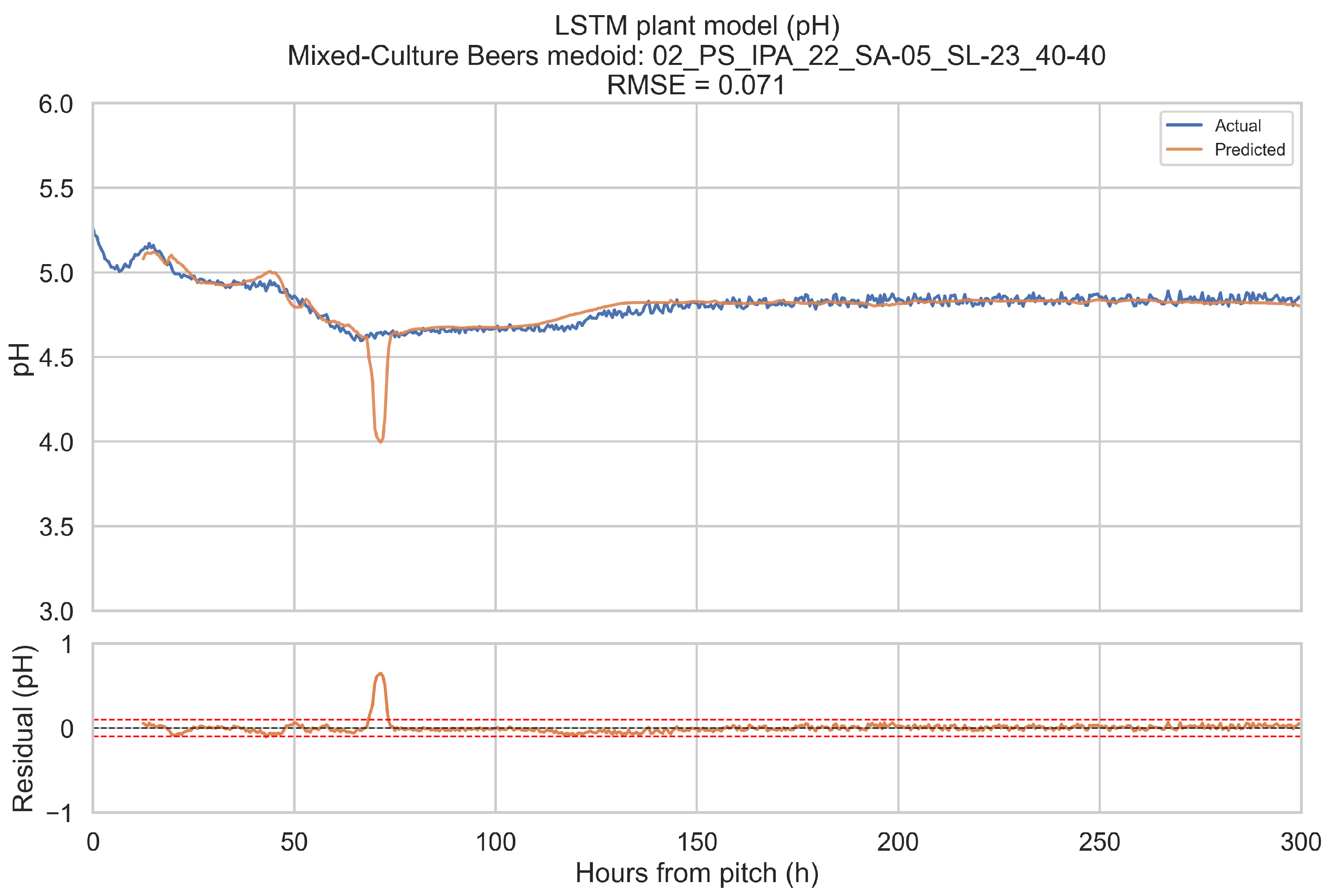
The training objective is defined on short forecast windows. To assess how well the LSTM plant model reproduces complete fermentations, it is evaluated in a one-step-ahead, open-loop configuration over the full 0–300 h horizon for a representative fermentation in each beverage category.
Representative fermentations are selected using a medoid approach. For each of the four categories (ales, IPAs, lagers, mixed-culture beers), only fermentations with BrewIQ data covering the entire 0–300 h interval are considered. Their degrees Plato trajectories are interpolated onto a uniform 1 h grid, and pairwise distances are computed between these interpolated trajectories as root-mean-square differences over time. For each candidate trajectory, the mean distance to all others in the same category is calculated. The medoid is defined as the fermentation whose mean distance is minimal. Where possible, the candidate pool is restricted to fermentations with pH measurements throughout most of the horizon, so that both degrees Plato and pH can be assessed.
For a chosen medoid, the trained LSTM plant model is rolled forward as follows: at each time step t in 0.5 h increments, a 12 h history window ending at t is extracted, scaled, and passed through the encoder to update the hidden state. The decoder then receives the next planned temperature setpoint and time index, together with original gravity (OGref), the style embedding, and the most recent degrees Plato value, and returns a one-step-ahead forecast for apparent extract (°P) and pH at time h. Predictions are transformed back to physical units using the stored scalers. This procedure is repeated across the entire 0–300 h horizon. Root-mean-square error (RMSE) and mean absolute error (MAE) are computed for each medoid, using only those time points for which the relevant ground truth measurements are available. These medoid trajectories and metrics provide an intuitive picture of model performance across complete fermentations, complementing the window-based training and validation losses.
4. Discussion
The results indicate that a single LSTM plant model can represent beer fermentation dynamics across ales, IPAs, lagers, and mixed-culture beers with an accuracy that is broadly compatible with inline measurement tolerances, especially for pH. This is achieved without separate parameter sets for each category and without explicit mechanistic equations for biomass, sugar, or ethanol concentrations. Instead, the model operates directly on temperature, degrees Plato, pH, original gravity descriptors, and beer style labels, learning the collective behaviour of 1305 fermentations. Several aspects of the formulation contribute to this performance.
Firstly, the input feature set is deliberately close to what brewers observe in practice. The reconstructed temperature setpoint captures the main steps of the thermal control strategy, while the original gravity (OGref) summarises wort strength in a form that is robust to measurement noise. The use of time since pitch as an explicit input allows the LSTM to locate each forecast within the broader fermentation timeline, a key consideration given that fermentation is a time-variant process. Combined with the recent degrees Plato history, these features provide a compact yet informative representation of the present state of the process.
Secondly, the multi-output design, with a shared recurrent backbone and separate degrees Plato and pH output heads, allows the model to exploit relationships between attenuation and acidification. In many fermentations, pH dynamics are strongly coupled to yeast metabolism, and changes in pH can provide indirect information about the underlying state of the culture [
27,
28]. By jointly predicting apparent extract and pH, the network can use pH measurements when available to refine its internal representation, even though pH is not used as an input. The masked Huber loss, together with pH-specific dropout, prevents windows with sparse pH measurements from dominating training while still contributing useful information for soft-sensing.
Moreover, the choice of an LSTM architecture is well aligned with the data’s characteristics. Beer fermentations display long-range temporal dependencies, for example, decisions made in the first 24 h can influence attenuation, pH, and flavour many days later. LSTMs are designed precisely to handle such long-range time dependencies [
18,
19]. The present model uses a relatively modest architecture by modern standards, with only two layers of 128 units and a 16-dimensional style embedding. Yet, the training curves and medoid results indicate that this capacity is sufficient for the process modelling challenge. In this respect, the LSTM plant model extends previous data-driven work in brewing [
11,
29] by operating on a larger, more varied dataset and by modelling both apparent extract and pH directly, without reliance on offline biomass measurements.
From an operational perspective, prediction error magnitudes in excess of 0.3–0.5 °P in apparent extract and 0.05–0.10 pH units relative to ground truth are meaningful. For many breweries, such errors are comparable to the variability observed between replicate fermentations under nominally identical conditions. A plant model with this level of fidelity can support several tasks. It can provide short-horizon forecasts of apparent extract and pH, helping brewers to anticipate when a fermentation will reach target attenuation or pH limits. It can act as a soft sensor for pH, providing consistency checks against noisy measurements or suggesting likely trajectories when pH measurements are temporarily unavailable. It can also serve as a basis for anomaly detection by identifying fermentations whose trajectories deviate significantly from the behaviour encoded in the training data.
At the same time, there are important limitations. The LSTM plant model, at present, is trained predominantly on fermentations drawn from a variety of commercial breweries, encompassing a broad mix of monoculture yeasts, recipes, and process equipment, with the mixed-culture beers representing the category in which laboratory-scale datasets contribute most strongly. Although this provides a richer basis than a single-wort, single-site study, it still covers only a subset of the diversity found in the wider brewing industry. Generalising the model to breweries with substantially different vessel designs, control philosophies, or product portfolios may therefore require additional data volumes and attributes. Similarly, while the style embedding allows the network to adapt to different beer styles within the dataset, it does not have explicit access to formulation variables such as hop additions, adjunct use, or yeast generation, which are important factors of fermentation behaviour in other contexts.
Another limitation is that the model is purely data-driven. It does not explicitly enforce physical constraints such as non-negative extract, limits on achievable attenuation, or monotonic relationships between certain variables. Within the range of the training data, these constraints are respected implicitly, but there is no guarantee that extrapolations will remain physically realistic. Physics-informed or hybrid approaches, which embed approximate kinetic relationships or monotonic behaviours into the network [
15,
30], could be explored to improve extrapolation and provide stronger guarantees on model behaviour in sparse or novel operating regimes.
The way in which training and validation sets are constructed also warrants careful interpretation. Windows are sampled at 0.5 h intervals with substantial overlap, and the training and validation split is performed at the window level, not at the batch level. This means that windows from the same fermentation can be used for both training and validation. The excellent agreement between training and validation losses, therefore, indicates consistency across overlapping windows from the same population of fermentations, but may overestimate performance on entirely unseen fermentations. A stricter evaluation, in which complete fermentations are held out at the batch level, would provide a more conservative measure of generalisation. Similarly, the medoid-based analysis, while intuitive and informative, reports full-run errors for only one representative fermentation per category. It does not capture the full distribution of errors across all batches, nor does it quantify worst-case performance. This, however, is to be treated strictly as an exemplar of “typical” model performance, as a complement to the reported error metrics.
The term “digital twin” in the wider literature encompasses a broad spectrum of models and couplings [
17]. In the context of this research, the LSTM plant model is best viewed as a data-driven surrogate of the fermentation dynamics that could be embedded within a digital twin or MPC architecture, rather than as a complete digital twin in its own right. The present study focuses on establishing that a compact, recurrent model can reproduce the core dynamics of typical fermentations with useful accuracy.
Finally, there is scope to refine the representation of batch-level information and to optimise hyperparameters. In the current formulation, original gravity (OGref) and the style embedding are the only batch-level descriptors. Future models could incorporate additional descriptors, such as yeast generation, fermentation volume, or key recipe parameters, or could use learned embeddings of entire recipes or process histories. However, accessing or producing such a dataset presents a significant research challenge. Hyperparameters such as the number of LSTM layers, the hidden size, and the style embedding dimension were chosen to balance model capacity and training stability based on “best practices” and the scale of the dataset, rather than through an exhaustive search. Systematic hyperparameter optimisation, possibly using Bayesian optimisation or more sophisticated methods, could further improve performance.
Overall, the present results suggest that LSTM networks provide a practical route to building data-driven plant models of beer fermentations from routinely collected brewery data. They complement rather than replace mechanistic models and brewing expertise, and they provide a flexible foundation for developing more advanced monitoring and control strategies.
5. Conclusions
This paper has presented a long short-term memory (LSTM) plant model for beer fermentation, trained on a large collection of inline-monitored fermentations using Sennos’ BrewIQ sensor array (see
Figure 4). The model is built upon reconstructed temperature setpoints, time since pitch, original gravity descriptors, apparent extract, and beer style labels, and predicts short-horizon trajectories of apparent extract and pH. The training dataset comprises 1305 fermentations drawn from ales, IPAs, lagers, and mixed-culture beers.
The LSTM plant model uses a two-layer architecture with a style embedding and a multi-output head structure for apparent extract and pH. Training is carried out on overlapping 12 h history and 12 h forecast windows, using a masked Huber loss to accommodate incomplete pH data and modest noise augmentation on temperature and degrees Plato inputs.
When evaluated on representative medoid fermentations in each beverage category, the model reproduces degrees Plato and pH trajectories over the full 0–300 h horizon with errors that are on the same order as the underlying sensor tolerances. Across the four medoids, the mean apparent extract RMSE and MAE are 0.339 and 0.207 °P, and the mean pH RMSE and MAE are 0.074 and 0.044. For pH, these values are close to a 0.1 pH unit tolerance and for apparent extract, they are modestly larger than a 0.2 °P sensor tolerance, but remain within a range that is likely to be useful for monitoring and forecasting.
The research underscores the importance of careful data handling in industrial machine-learning applications. Uniform resampling at 0.5 h, robust estimation of original gravity (OGref), reconstruction of temperature setpoints, and explicit encoding of beer style all contribute to the success of the model. The masked Huber loss and pH-aware dropout provide a straightforward way to learn from incomplete pH measurements while retaining the benefits of joint apparent extract and pH modelling.
From an application standpoint, the LSTM plant model can support short-horizon forecasting of attenuation and pH (via soft sensing), and anomaly detection during fermentation. With further development, it could also be embedded within a model predictive control framework, allowing breweries to explore alternative temperature schedules and assess their likely impact on fermentation trajectories “in silico” before committing to process changes.
Key limitations are the reliance on the variables available from inline sensing (excluding biomass and other biochemical states), incomplete pH coverage, and the limited representation of mixed-culture fermentations in the training data. For future work and deployment, priorities include external validation across additional breweries and vessel geometries, uncertainty quantification and drift monitoring, and incorporation of richer operational metadata (e.g., yeast strain, pitching rate, oxygenation) where available.
In conclusion, the LSTM-based plant model introduced in this work delivers a data-driven yet computationally efficient model of beer fermentation behaviour. It demonstrates how recurrent neural networks trained on production data can serve as practical building blocks for brewery digital twins and decision-support workflows.