Near-UV Circular Dichroism and Second-Derivative Fluorescence Spectroscopy as Complementary Tools for Studying Ligand–Albumin Interactions
Abstract
1. Introduction
2. Results and Discussion
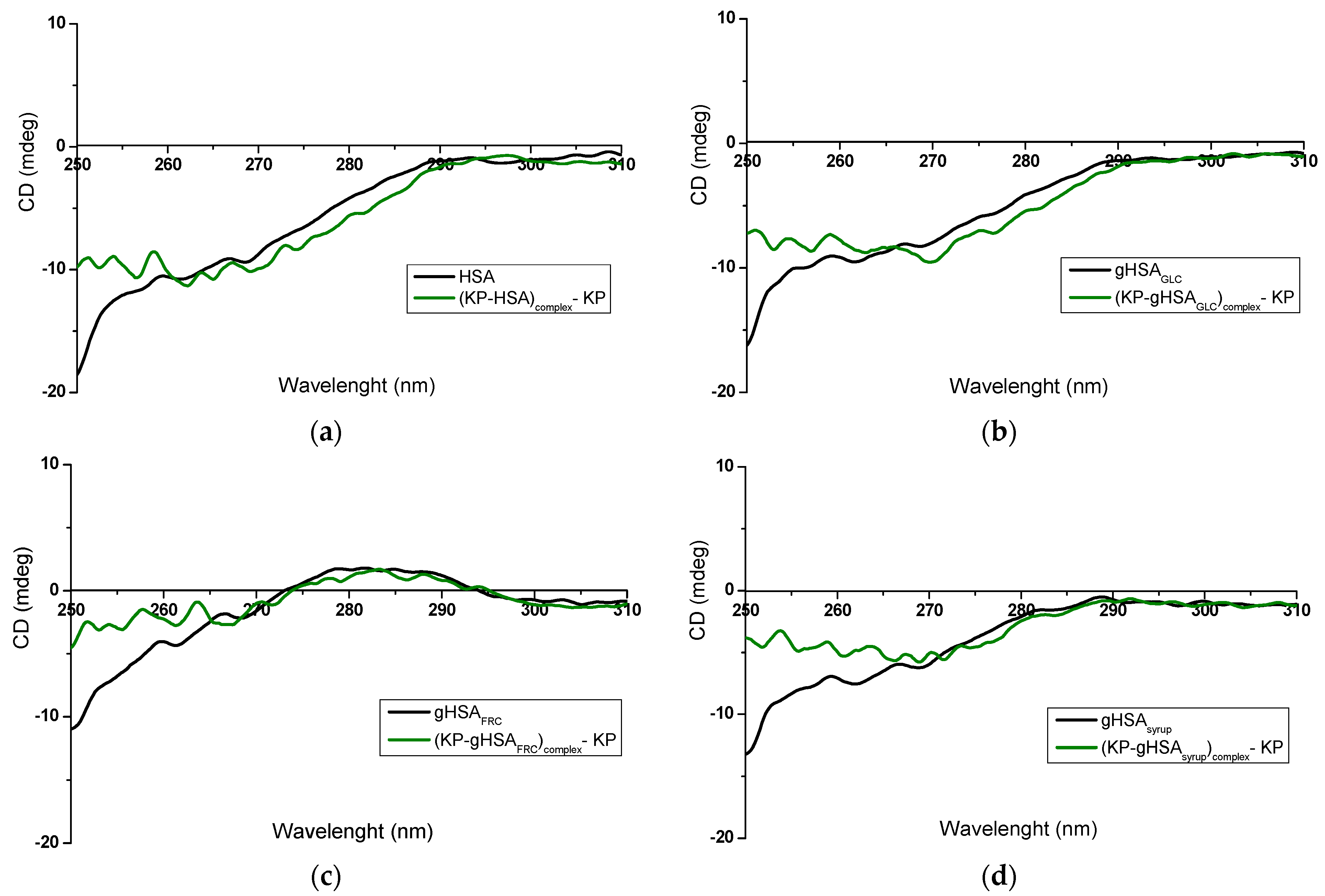
2.1. Assessment of the Tertiary Structure of Albumins by Near-UV CD Spectroscopy
2.2. Assessment of the Tertiary Structure of Albumins by Second-Derivative Fluorescence Spectra
3. Materials and Methods
3.1. Chemicals and Reagents
3.2. Methods
3.2.1. Sample Preparation
3.2.2. Circular Dichroism (CD) and Fluorescence Spectroscopy
3.3. Statistical Analysis
4. Conclusions
- (i)
- Glycation significantly alters the tertiary structure of HSA and reduces its drug-binding capacity at Sudlow’s sites I and II. Fructose-glycated HSA showed the most pronounced structural changes, confirming fructose as the most reactive glycation agent.
- (ii)
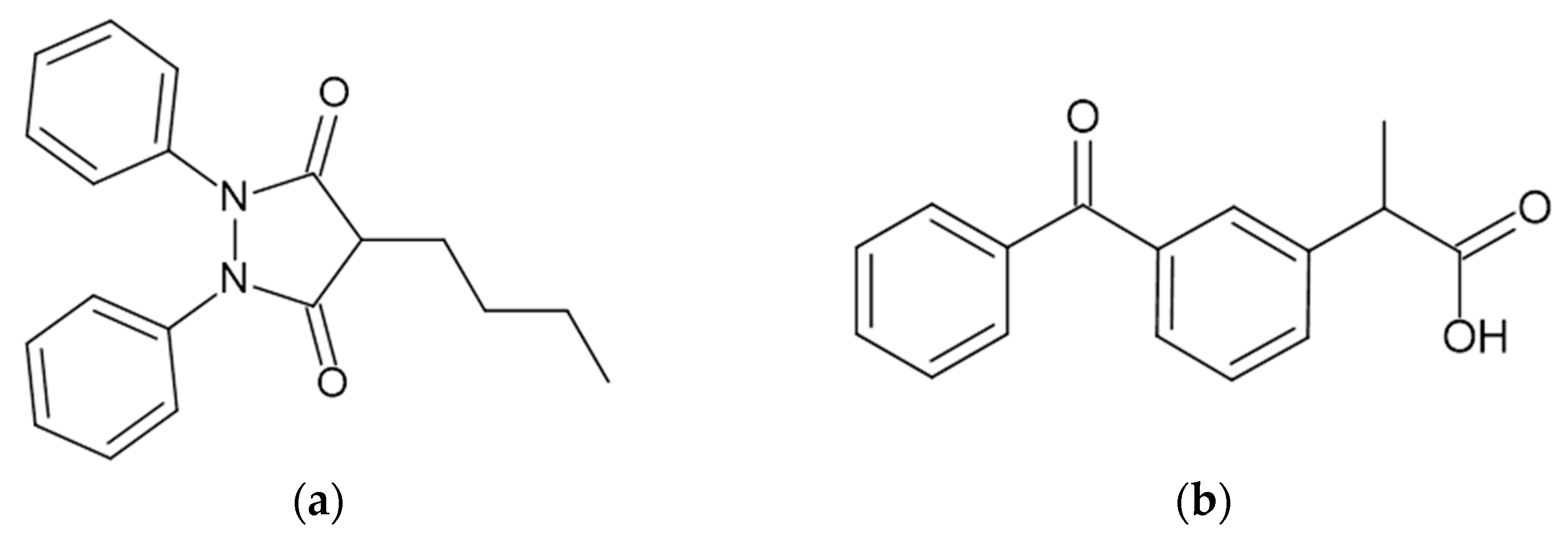
- PHB induced distinct structural rearrangements, manifested by a characteristic enhancement of the ellipticity peak at ~290 nm, which indicates perturbations in the chiral environment near Trp14 within Sudlow’s site I.
- (iii)
- KP caused weaker, site-specific conformational perturbations, primarily within hydrophobic domains enriched in Phe residues.
- (iv)
- Glycation reduces the hydrophobicity of the aromatic residue environment, making it more exposed and polar, whereas the microenvironment of Trp214 remained relatively stable.
- (v)
- Ligands modulate the conformational flexibility of glycated albumin mainly through the reorganization of Tyr-rich domains, rather than by directly affecting the surroundings of the single Trp214.
- (vi)
- The findings demonstrate that glycation heterogeneity significantly influences drug binding, a finding relevant to pharmacokinetics in diabetes and other metabolic disorders.
Supplementary Materials
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| AGEs | Advanced glycation end-products |
| CD | Circular dichroism |
| FRC | D(−)-fructose |
| GLC | D(+)-glucose |
| GFS | Glucose–fructose syrup |
| HSA | Human serum albumin |
| gHSAFRC | Human serum albumin glycated by fructose |
| gHSAGLC | Human serum albumin glycated by glucose |
| gHSAsyrup | Human serum albumin glycated by glucose–fructose syrup |
| KP | Ketoprofen |
| NSAIDs | Nonsteroidal anti-inflammatory drugs |
| SD | Standard deviation |
| PHB | Phenylbutazone |
References
- Peters, T. All About Albumin. In Biochemistry, Genetics, and Medical Applications; Academic Press: San Diego, CA, USA, 1995; pp. 9–19,228–234. [Google Scholar]
- Fasano, M.; Curry, S.; Terreno, E.; Galliano, M.; Fanali, G.; Narciso, P.; Notari, S.; Ascenzi, P. The extraordinary ligand binding properties of human serum albumin. IUBMB Life 2005, 57, 787–796. [Google Scholar] [CrossRef]
- Yamasaki, K.; Chuang, V.T.G.; Maruyama, T.; Otagiri, M. Albumin–Drug Interaction and Its Clinical Implication. Biochim. Biophys. Acta 2013, 1830, 5435–5443. [Google Scholar] [CrossRef]
- Mishra, V.; Heath, R.J. Structural and Biochemical Features of Human Serum Albumin Essential for Eukaryotic Cell Culture. Int. J. Mol. Sci. 2021, 22, 8411. [Google Scholar] [CrossRef]
- Kragh-Hansen, U. Human Serum Albumin: A Multifunctional Protein with Multiple Binding Sites for Endogenous and Exogenous Ligands. Biol. Chem. 2016, 397, 1037–1048. [Google Scholar]
- Sitar, M.E.; Aydin, S.; Cakatay, U. Human serum albumin and its relation with oxidative stress. Clin. Lab. 2013, 59, 945–952. [Google Scholar] [CrossRef]
- Wang, Z.M.; Ho, J.X.; Ruble, J.R.; Rose, J.; Ruker, F.; Ellenburg, M.; Murphy, R.; Click, J.; Soistman, E.; Wilkerson, L.; et al. Structural studies of several clinically important oncology drugs in complex with human serum albumin. Biochim. Biophys. Acta 2013, 1830, 5356–5374. [Google Scholar] [CrossRef]
- He, X.; Carter, D. Atomic structure and chemistry of human serum albumin. Nature 1992, 358, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Rabbani, G.; Ahn, S.N. Structure, enzymatic activities, glycation and therapeutic potential of human serum albumin: A natural cargo. Int. J. Biol. Macromol. 2019, 123, 979–990. [Google Scholar] [CrossRef]
- Yazdani, F.; Shareghi, B.; Farhadian, S.; Momeni, L. Structural insights into the binding behavior of favonoids naringenin with human serum albumin. J. Mol. Liq. 2022, 349, 118431. [Google Scholar] [CrossRef]
- Sudlow, G.; Birkett, D.J.; Wade, D.N. The characterization of two specific drug binding sites on human serum albumin. Mol. Pharmacol. 1975, 11, 824–832. [Google Scholar] [CrossRef]
- Karimi, M.; Bahrami, S.; Ravari, S.B.; Zangabad, P.S.; Mirshekari, H.; Bozorgomid, M.; Shahreza, S.; Sori, M.; Hamblin, M.R. Albumin nanostructures as advanced drug delivery systems. Expert. Opin. Drug Deliv. 2016, 13, 1609–1623. [Google Scholar] [CrossRef]
- Ghuman, J.; Zunszain, P.A.; Petitpas, I.; Bhattacharya, A.A.; Otagiri, M.; Curry, S. Structural basis of the drug-binding specificity of human serum albumin. J. Mol. Biol. 2005, 353, 38–52. [Google Scholar] [CrossRef] [PubMed]
- Czub, M.P.; Handing, K.B.; Venkataramany, B.S.; Cooper, D.R.; Shabalin, I.G.; Minor, W. Albumin-Based Transport of Nonsteroidal Anti-Inflammatory Drugs in Mammalian Blood Plasma. J. Med. Chem. 2020, 63, 6847–6862. [Google Scholar] [CrossRef] [PubMed]
- Dubois, N.; Lapicque, F.; Magdalou, J.; Abiteboul, M.; Netter, P. Stereoselective binding of the glucuronide of ketoprofen enantiomers to human serum albumin. Biochem. Pharmacol. 1994, 48, 1693–1699. [Google Scholar] [CrossRef]
- Bakaeean, B.; Kabiri, M.; Iranfar, H.; Saberi, M.R.; Chamani, J. Binding Effect of Common Ions to Human Serum Albumin in the Presence of Norfloxacin: Investigation with Spectroscopic and Zeta Potential Approaches. J. Solut. Chem. 2012, 41, 1777–1801. [Google Scholar] [CrossRef]
- Simard, J.R.; Zunszain, P.A.; Hamilton, J.A.; Curry, S. Locating of high and low affinity fatty acid binding sites on human serum albumin revealed by NMR drug-competition analysis. J. Mol. Biol. 2006, 361, 336–351. [Google Scholar] [CrossRef]
- Bhattacharya, A.A.; Grüne, T.; Curry, S. Crystallographic Analysis Reveals Common Modes of Binding of Medium and Long-Chain Fatty Acids to Human Serum Albumin. J. Mol. Biol. 2000, 303, 721–732. [Google Scholar] [CrossRef]
- Kragh-Hansen, U.; Brennan, S.O.; Minchiotti, L.; Galliano, M. Modified high-affinity binding of Ni2+, Ca2+ and Zn2+ to natural mutants of human serum albumin and proalbumin. Biochem. J. 1994, 301, 217–223. [Google Scholar] [CrossRef]
- Otagiri, M.; Chuang, V.T.G. Albumin in Medicine: Pathological and Clinical Applications; Springer: Singapore, 2016. [Google Scholar]
- Awasthi, S.; Saraswathi, N.T. Non-enzymatic glycation mediated structure–function changes in proteins: Case of serum albumin. RSC Adv. 2016, 6, 90739–90753. [Google Scholar] [CrossRef]
- Arif, Z.; Arfat, M.; Ahmad, J.; Zaman, A.; Islam, S.; Khan, M. Relevance of Nitroxidation of Albumin in Rheumatoid Arthritis: A Biochemical and Clinical Study. J. Clin. Cell. Immunol. 2015, 6, 1000324. [Google Scholar] [CrossRef]
- Oettl, K.; Stauber, R.E. Physiological and pathological changes in the redox state of human serum albumin critically influence its binding properties. Br. J. Pharmacol. 2007, 151, 580–590. [Google Scholar] [CrossRef] [PubMed]
- Shaklai, N.; Garlick, R.L.; Bunn, H.F. Nonenzymatic glycosylation of human serum albumin alters its conformation and function. J. Biol. Chem. 1984, 259, 3812–3817. [Google Scholar] [CrossRef] [PubMed]
- Vetter, S.W.; Indurthi, V.S. Moderate glycation of human serum albumin affects folding, stability, and ligand binding. Clin. Chim. Acta 2011, 412, 2105–2116. [Google Scholar] [CrossRef]
- Anguizola, J.; Matsuda, R.; Barnaby, O.S.; Hoy, K.S.; Wa, C.; DeBolt, E.; Koke, M.; Hage, D.S. Review: Glycation of human serum albumin. Clin. Chim. Acta 2013, 425, 64–76. [Google Scholar] [CrossRef]
- Mötzing, M.; Blüher, M.; Grunwald, T.; Hoffmann, R. Immunological Quantitation of the Glycation Site Lysine-414 in Serum Albumin in Human Plasma Samples by Indirect ELISA Using Highly Specific Monoclonal Antibodies. Chembiochem 2024, 25, e202300550. [Google Scholar] [CrossRef]
- Kelly, S.M.; Price, N.C. The Use of Circular Dichroism in the Investigation of Protein Structure and Function. Curr. Protein Pept. Sci. 2000, 1, 349–384. [Google Scholar] [CrossRef]
- Ranjbar, B.; Gill, P. Circular Dichroism Techniques: Biomolecular and Nanostructural Analyses—A Review. Chem. Biol. Drug Des. 2009, 74, 101–120. [Google Scholar] [CrossRef] [PubMed]
- Zsila, F. Circular Dichroism Spectroscopic Detection of Ligand Binding Induced Subdomain IB Specific Structural Adjustment of Human Serum Albumin. J. Phys. Chem. B 2013, 117, 10798–10806. [Google Scholar] [CrossRef]
- Carter, D.C.; Ho, J.X. Structure of Serum Albumin. Adv. Protein Chem. 1994, 45, 153–203. [Google Scholar] [CrossRef]
- Mocz, G.; Ross, J. Fluorescence Techniques in Analysis of Protein–Ligand Interactions. Methods Mol. Biol. 2013, 1008, 169–210. [Google Scholar] [CrossRef]
- Kumar, V.; Sharma, V.K.; Kalonia, D.S. Second derivative tryptophan fluorescence spectroscopy as a tool to characterize partially unfolded intermediates of proteins. Int. J. Pharm. 2005, 294, 193–199. [Google Scholar] [CrossRef]
- Mozo-Villarías, A. Second derivative fluorescence spectroscopy of tryptophan in proteins. J. Biochem. Biophys. Methods 2002, 50, 163–178. [Google Scholar] [CrossRef]
- Biswas, P. Modern Biophysical Approaches to Study Protein–Ligand Interactions. Biophys. Rev. Lett. 2018, 13, 133–155. [Google Scholar] [CrossRef]
- Hekmat, A.; Hajebrahimi, Z.; Motamedzade, A. Structural Changes of Human Serum Albumin (HSA) in Simulated Microgravity. Curr. Med. Chem. 2017, 24, 3941–3949. [Google Scholar] [CrossRef]
- Szkudlarek, A.; Sułkowska, A.; Maciążek-Jurczyk, M.; Chudzik, M.; Równicka-Zubik, J. Effects of Non-Enzymatic Glycation in Human Serum Albumin. Spectroscopic Analysis. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2016, 152, 645–653. [Google Scholar] [CrossRef]
- Szkudlarek, A.; Pentak, D.; Ploch, A.; Pożycka, J.; Maciążek-Jurczyk, M. In Vitro Investigation of the Interaction of Tolbutamide and Losartan with Human Serum Albumin in Hyperglycemia States. Molecules 2017, 22, 2249. [Google Scholar] [CrossRef] [PubMed]
- Zaman, A.; Arif, Z.; Moinuddin; Alam, K. Fructose–Human Serum Albumin Interaction Undergoes Numerous Biophysical and Biochemical Changes before Forming AGEs and Aggregates. Int. J. Biol. Macromol. 2017, 109, 896–906. [Google Scholar] [CrossRef] [PubMed]
- Muzammil, S.; Kumar, Y.; Tayyab, S. Molten Globule-Like State of Human Serum Albumin at Low pH. Eur. J. Biochem. 2001, 266, 26–32. [Google Scholar] [CrossRef]
- Kabir, M.Z.; Mukarram, A.K.; Mohamad, S.B.; Alias, Z.; Tayyab, S. Characterization of the Binding of an Anticancer Drug, Lapatinib to Human Serum Albumin. J. Photochem. Photobiol. B 2016, 160, 229–239. [Google Scholar] [CrossRef]
- Musa, K.A.; Ridzwan, N.F.W.; Mohamad, S.B.; Tayyab, S. Exploring the Combination Characteristics of Lumefantrine, an Antimalarial Drug and Human Serum Albumin through Spectroscopic and Molecular Docking Studies. J. Biomol. Struct. Dyn. 2021, 39, 691–702. [Google Scholar] [CrossRef] [PubMed]
- Dockal, M.; Carter, D.C.; Rüker, F. Conformational Transitions of the Three Recombinant Domains of Human Serum Albumin Depending on pH. J. Biol. Chem. 2000, 275, 3042–3050. [Google Scholar] [CrossRef] [PubMed]
- Yamasaki, K.; Maruyama, T.; Kragh-Hansen, U.; Otagiri, M. Characterization of Site I on Human Serum Albumin: Concept about the Structure of a Drug Binding Site. Biochim. Biophys. Acta 1996, 1295, 147–157. [Google Scholar] [CrossRef]
- Russeva, V.; Mihailova, D. Binding of Phenylbutazone to Human Serum Albumin. Characterization and Identification of Binding Sites. Arzneimittelforschung 1999, 49, 255–258. [Google Scholar] [CrossRef]
- Chignell, C.F. Optical Studies of Drug–Protein Complexes II. Interaction of Phenylbutazone and Its Analogues with Human Serum Albumin. Mol. Pharmacol. 1969, 5, 244–252. [Google Scholar] [CrossRef]
- Bertozo, L.C.; Tavares Neto, E.; Oliveira, L.C.; Ximenes, V.F. Oxidative Alteration of Trp-214 and Lys-199 in Human Serum Albumin Increases Binding Affinity with Phenylbutazone: A Combined Experimental and Computational Investigation. Int. J. Mol. Sci. 2018, 19, 2868. [Google Scholar] [CrossRef]
- Graciani, F.S.; Ximenes, V.F. Investigation of Human Albumin-Induced Circular Dichroism in Dansylglycine. PLoS ONE 2013, 8, e76849. [Google Scholar] [CrossRef] [PubMed]
- Manoharan, P.; Wong, Y.H.; Tayyab, S. Stabilization of Human Serum Albumin against Urea Denaturation by Diazepam and Ketoprofen. Protein Pept. Lett. 2015, 22, 611–617. [Google Scholar] [CrossRef]
- Rogóż, W.; Owczarzy, A.; Kulig, K.; Maciążek-Jurczyk, M. Ligand–Human Serum Albumin Analysis: The Near-UV CD and UV–Vis Spectroscopic Studies. Naunyn Schmiedebergs Arch. Pharmacol. 2025, 398, 3119–3131. [Google Scholar] [CrossRef]
- Balestrieri, C.; Colonna, G.; Giovane, A.; Irace, G.; Servillo, L. Second- Derivative Spectroscopy of Proteins. Eur. J. Biochem. 1978, 90, 433–440. [Google Scholar] [CrossRef]
- Chudzik, M.; Maciążek-Jurczyk, M.; Pawełczak, B.; Sułkowska, A. Spectroscopic Studies on the Molecular Ageing of Serum Albumin. Molecules 2017, 22, 34. [Google Scholar] [CrossRef] [PubMed]
- Szkudlarek, A. Effect of Palmitic Acid on Tertiary Structure of Glycated Human Serum Albumin. Processes 2023, 11, 2746. [Google Scholar] [CrossRef]
- Fanali, G.; di Masi, A.; Trezza, V.; Marino, M.; Fasano, M.; Ascenzi, P. Human Serum Albumin: From Bench to Bedside. Mol. Asp. Med. 2012, 33, 209–290. [Google Scholar] [CrossRef]





| Sample | H275nm mean ± SD * | λmin (nm) | λmax (nm) | H295nm mean ± SD * | λmin (nm) | λmax (nm) |
|---|---|---|---|---|---|---|
| HSA | 0.842 ± 0.040 (a) | 301 | 313 | 0.014 ± 0.003 | 383 | 389 |
| (PHB–HSA)complex—PHBnorm | 1.732 ± 0.172 (a) | 308 | 318 | 0.020 ± 0.006 | 391 | 395 |
| (KP–HSA)complex—KPnorm | 0.558 ± 0.086 (a) | 308 | 317 | 0.011 ± 0.007 | 392 | 396 |
| gHSAGLC | 0.794 ± 0.012 (b) | 301 | 314 | 0.016 ± 0.005 | 387 | 394 |
| (PHB–gHSAGLC)complex—PHBnorm | 1.442 ± 0.050 (b) | 308 | 318 | 0.020 ± 0.008 | 378 | 384 |
| (KP–gHSAGLC)complex—KPnorm | 0.214 ± 0.003 (b) | 309 | 316 | 0.027 ± 0.001 | 383 | 391 |
| gHSAFRC | 0.172 ± 0.013 (c) | 299 | 305 | 0.009 ± 0.005 | 393 | 372 |
| (PHB–gHSAFRC)complex—PHBnorm | 0.352 ± 0.039 (c) | 308 | 317 | 0.025 ± 0.011 | 394 | 386 |
| (KP–gHSAFRC)complex—KPnorm | 0.369 ± 0.017 (c) | 311 | 302 | 0.010 ± 0.004 | 382 | 387 |
| gHSAsyrup | 0.219 ± 0.033 (d) | 300 | 307 | 0.003 ± 0.002 (e) | 381 | 386 |
| (PHB–gHSAsyrup)complex—PHBnorm | 0.641 ± 0.051 (d) | 306 | 317 | 0.002 ± 0.002 | 383 | 387 |
| (KP–gHSAsyrup)complex—KPnorm | 0.479 ± 0.010 (d) | 311 | 302 | 0.027 ± 0.009 (e) | 391 | 383 |
| Sample | H275nm | H295nm |
|---|---|---|
| Mean ± SD * | ||
| (PHB–HSA)complex—PHBnorm | 1.732 ± 0.172 | 0.020 ± 0.006 |
| (PHB–gHSAGLC)complex—PHBnorm | 1.442 ± 0.050 | 0.020 ± 0.008 |
| (PHB–gHSAFRC)complex—PHBnorm | 0.352 ± 0.039 | 0.025 ± 0.011 |
| (PHB–gHSAsyrup)complex—PHBnorm | 0.641 ± 0.051 | 0.002 ± 0.002 |
| One–way ANOVA (H275nm) Kruskal–Wallis ANOVA (H295nm) | F = 140.417 p < 0.001 | H = 7.269 p = 0.064 statistically non-significant differences (p > 0.05) |
| Tukey’s HSD test, n = 3 | all differences statistically significant (p < 0.05) | n.a. ** |
| (KP–HSA)complex—KPnorm | 0.558 ± 0.086 | 0.011 ± 0.007 |
| (KP–gHSAGLC)complex—KPnorm | 0.214 ± 0.003 | 0.020 ± 0.008 |
| (KP–gHSAFRC)complex—KPnorm | 0.369 ± 0.017 | 0.010 ± 0.004 |
| (KP–gHSAsyrup)complex—KPnorm | 0.479 ± 0.010 | 0.027 ± 0.009 |
| Kruskal–Wallis ANOVA | H = 10.385 | H = 7.989 |
| p = 0.016 | p = 0.046 | |
| MCT, n = 3 | statistically significant differences: (KP–HSA)complex—KPnorm vs. (KP–gHSAGLC)complex—KPnorm (p = 0.013) | statistically non-significant differences (p > 0.05) |
| HSA | 0.842 ± 0.040 | 0.014 ± 0.003 |
| gHSAGLC | 0.794 ± 0.012 | 0.016 ± 0.005 |
| gHSAFRC | 0.172 ± 0.013 | 0.009 ± 0.005 |
| gHSAsyrup | 0.219 ± 0.033 | 0.003 ± 0.002 |
| One-way ANOVA | F = 518.875 | F = 6.478 |
| p < 0.001 | p = 0.016 | |
| Tukey’s HSD test, n = 3 | significant differences: HSA vs. gHSAFRC, HSA vs. gHSAsyrup, gHSAGLC vs. gHSAFRC, gHSAGLC vs. gHSAsyrup (all p < 0.05) | significant differences: HSA vs. gHSAsyrup, gHSAGLC vs. gHSAsyrup (all p < 0.05) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Szkudlarek, A. Near-UV Circular Dichroism and Second-Derivative Fluorescence Spectroscopy as Complementary Tools for Studying Ligand–Albumin Interactions. Processes 2025, 13, 4000. https://doi.org/10.3390/pr13124000
Szkudlarek A. Near-UV Circular Dichroism and Second-Derivative Fluorescence Spectroscopy as Complementary Tools for Studying Ligand–Albumin Interactions. Processes. 2025; 13(12):4000. https://doi.org/10.3390/pr13124000
Chicago/Turabian StyleSzkudlarek, Agnieszka. 2025. "Near-UV Circular Dichroism and Second-Derivative Fluorescence Spectroscopy as Complementary Tools for Studying Ligand–Albumin Interactions" Processes 13, no. 12: 4000. https://doi.org/10.3390/pr13124000
APA StyleSzkudlarek, A. (2025). Near-UV Circular Dichroism and Second-Derivative Fluorescence Spectroscopy as Complementary Tools for Studying Ligand–Albumin Interactions. Processes, 13(12), 4000. https://doi.org/10.3390/pr13124000






