Dynamic Optimisation of Fed-Batch Bioreactors for mAbs: Sensitivity Analysis of Feed Nutrient Manipulation Profiles
Abstract
:1. Introduction
2. Materials and Methods
2.1. Dynamic Flux Balance Model
2.2. Optimisation Software and Strategy
2.3. Sensitivty Analysis of Optimised Fed-Batch Bioreactors
2.4. Sensitivty Analysis Methodology and Case Studies
- How is performance affected with a restriction of glucose in the culture media?
- How is performance affected with an increase in glutamine in the culture media?
- How is performance affected with an increase in asparagine in the culture media?
3. Results
3.1. Glucose Fed-Batch Dynamic Optimisations
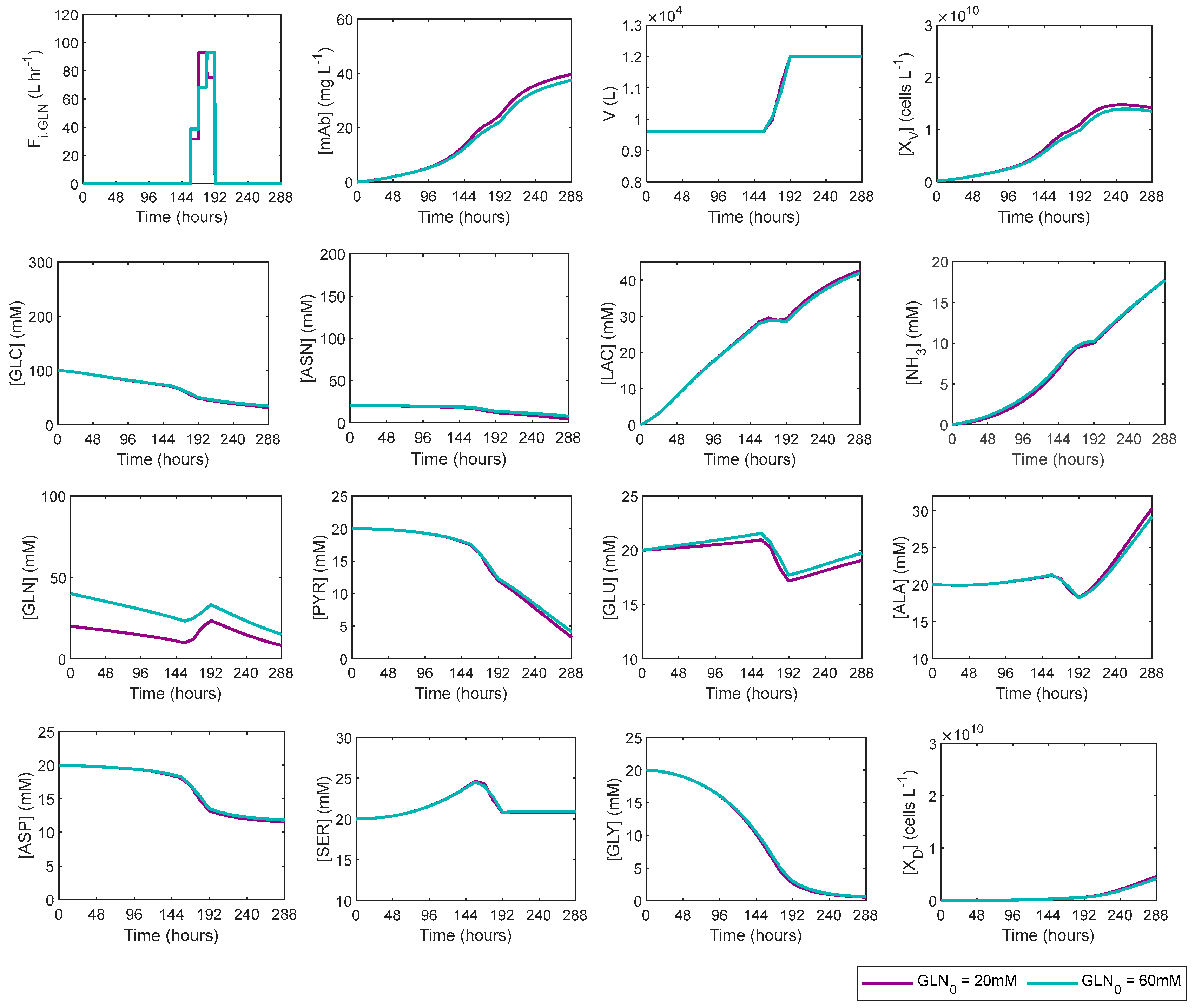
3.2. Glutamine Fed-Batch Dynamic Optimisations
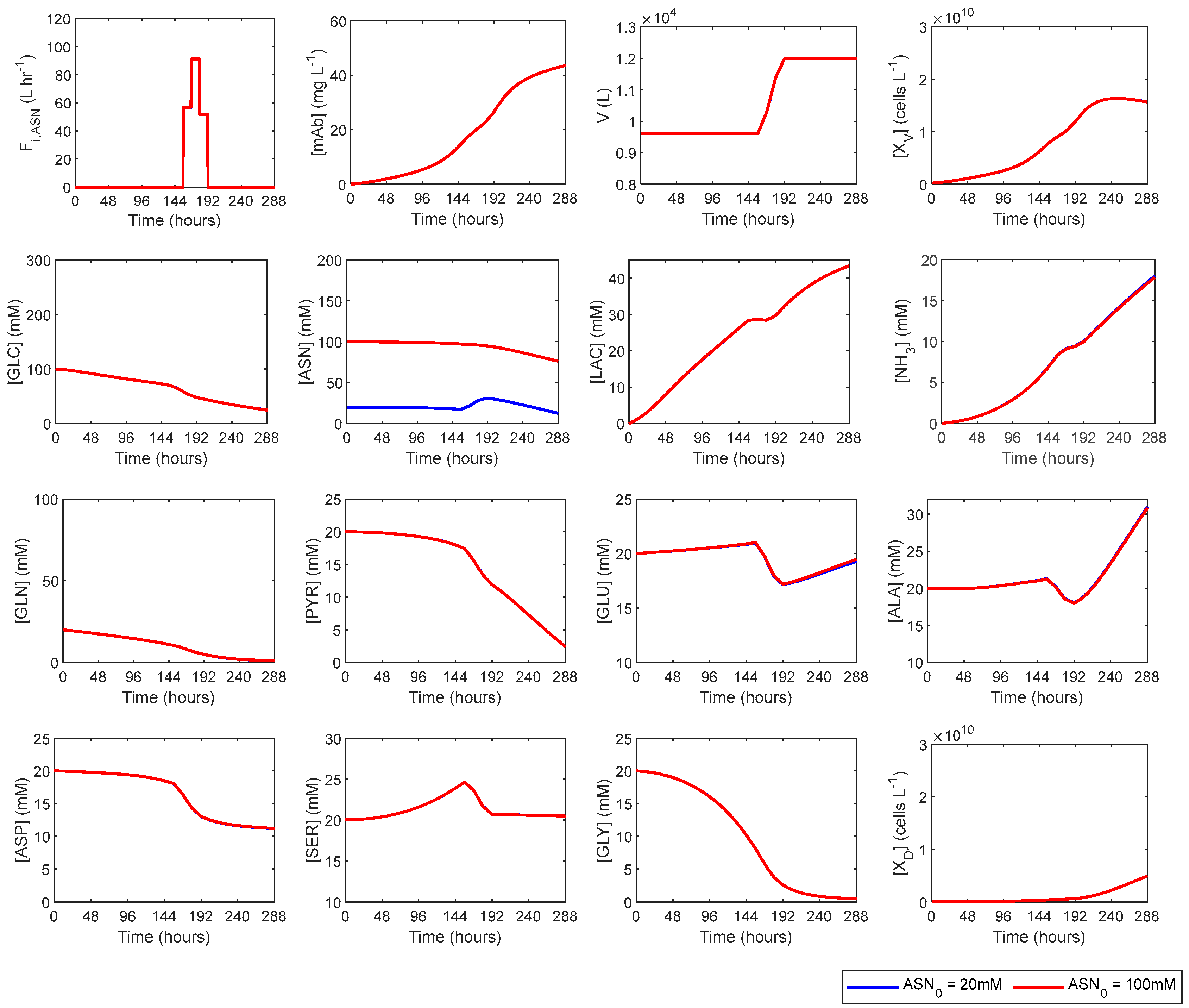
3.3. Asparagine Fed-Batch Dynamic Optimisations
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Appendix A
| (R1) | |
| (R2) | |
| (R3) | |
| (R4) | |
| (R5) | |
| (R6) | |
| (R7) | |
| (R8) | |
| (R9) | |
| (R10) | |
| (R11) | |
| (R12) | |
| (R13) | |
| (R14) | |
| (R15) | |
| (R16) | |
| (R17) | |
| (R18) | |
| (R19) | |
| (R20) | |
| (R21) | |
| (R22) | |
| (R23) |
| (E1) | |
| (E2) | |
| (E3) | |
| (E4) | |
| (E5) | |
| (E6) | |
| (E7) | |
| (E8) | |
| (E9) | |
| (E10) | |
| (E11) | |
| (E12) | |
| (E13) | |
| (E14) | |
| (E15) | |
| (E16) | |
| (E17) | |
| (E18) | |
| (E19) | |
| (E20) | |
| (E21) | |
| (E22) | |
| (E23) | |
| + | (E24) |
| (E25) | |
| (E26) | |
| (E27) | |
| (E28) | |
| (E29) | |
| (E30) | |
| (E31) | |
| (E32) | |
| (E33) | |
| (E34) | |
| (E35) | |
| (E36) | |
| (E37) | |
| (E38) | |
| (E39) | |
| (E40) | |
| (E41) | |
| (E42) | |
| (E43) | |
| (E44) | |
| (E45) | |
| (E46) | |
| (E47) | |
| (E48) | |
| (E49) | |
| (E50) | |
| (E51) | |
| (E52) | |
| (E53) | |
| (E54) | |
| (E55) | |
| (E56) | |
| (E57) | |
| (E58) | |
| (E59) | |
| (E60) | |
| (E61) | |
| (E62) | |
| (E63) | |
| (E64) | |
| (E65) |
| Parameter | Value | Unit | Parameter | Value | Unit |
|---|---|---|---|---|---|
| 8.43 × 10−12 | mmol 106 cell−1 h−1 | 0.0001875 | mM | ||
| 7.08 × 10−10 | mmol 106 cell−1 h−1 | 7 | mM | ||
| 6.63 × 10−12 | mmol 106 cell−1 h−1 | 0.324 | mM | ||
| 1.80 × 10−12 | mmol 106 cell−1 h−1 | 34.5 | mM | ||
| 9.00 × 10−14 | mmol 106 cell−1 h−1 | 5.6 | mM | ||
| 1.23 × 10−11 | mmol 106 cell−1 h−1 | 3.084 | mM | ||
| 1.20 × 10−12 | mmol 106 cell−1 h−1 | 4.55 | mM | ||
| 2.65 × 10−14 | mmol 106 cell−1 h−1 | 21.5 | mM | ||
| 3.35 × 10−13 | mmol 106 cell−1 h−1 | 1.50 × 10−10 | mM | ||
| 1.48 × 10−11 | mmol 106 cell−1 h−1 | 1 | - | ||
| 2.35 × 10−13 | mmol 106 cell−1 h−1 | 1.87 × 10−8 | mM | ||
| 8.80 × 10−13 | mmol 106 cell−1 h−1 | 1 | - | ||
| 8.80 × 10−13 | mmol 106 cell−1 h−1 | 9.35 × 10−8 | mM | ||
| 1.40 × 10−12 | mmol 106 cell−1 h−1 | 2 | - | ||
| 3.15 × 10−13 | mmol 106 cell−1 h−1 | 7.896 × 10−8 | mM | ||
| 2.12 × 10−13 | mmol 106 cell−1 h−1 | 4 | - | ||
| 4.75 × 10−14 | mmol 106 cell−1 h−1 | 0.00468 | h−1 | ||
| 2.30 × 10−12 | mmol 106 cell−1 h−1 | 0.017 | h−1 | ||
| 2.21 × 10−11 | mmol 106 cell−1 h−1 | 0.00375 | - | ||
| 1.15 | mM | 0.00375 | - | ||
| 0.32 | mM | 0.12075 | - | ||
| 6.72 | mM | 0.1105 | - | ||
| 0.015 | mM | 4.3 × 108 | cells mmol biomass−1 | ||
| 4.97 × 10−12 | mmol 106 cell−1 h−1 | PO | 3 | - | |
| 0.105 | mM | 0.0002 | mM | ||
| 38.5 | mM | 0.02 | mM | ||
| 1.25 | mM | 0.0024 | h−1 | ||
| 0.2892 | mM | 0.01 | h−1 | ||
| 0.27 | mM | 48.5 | mM | ||
| 77.5 | mM |
References
- Chames, P.; Van Regenmortel, M.; Weiss, E.; Baty, D. Therapeutic antibodies: Successes, limitations and hopes for the future. Br. J. Pharmacol. 2009, 157, 220–233. [Google Scholar] [CrossRef]
- Mahmuda, A.; Bande, F.; Al-Zihiry, K.J.K.; Abdulhaleem, N.; Majid, R.A.; Hamat, R.A.; Abdullah, W.O.; Unyah, Z. Monoclonal antibodies: A review of therapeutic applications and future prospects. Trop. J. Pharm. Res. 2017, 16, 713–722. [Google Scholar] [CrossRef]
- Dhara, V.G.; Naik, H.M.; Majewska, N.I.; Betenbaugh, M.J. Recombinant antibody production in CHO and NS0 cells: Differences and similarities. BioDrugs 2018, 32, 571–584. [Google Scholar] [CrossRef]
- Jain, E.; Kumar, A. Upstream processes in antibody production: Evaluation of critical parameters. Biotechnol. Adv. 2008, 26, 46–72. [Google Scholar] [CrossRef]
- Liu, J.K. The history of monoclonal antibody development—Progress, remaining challenges and future innovations. Ann. Med. Surg. 2014, 3, 113–116. [Google Scholar] [CrossRef]
- Lu, R.-M.; Hwang, Y.-C.; Liu, I.-J.; Lee, C.-C.; Tsai, H.-Z.; Li, H.-J.; Wu, H.-C. Development of therapeutic antibodies for the treatment of diseases. J. Biomed. Sci. 2020, 27, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Grilo, A.L.; Mantalaris, A. The increasingly human and profitable monoclonal antibody market. Trends Biotechnol. 2019, 37, 9–16. [Google Scholar] [CrossRef]
- Kastelic, M.; Kopač, D.; Novak, U.; Likozar, B. Dynamic metabolic network modeling of mammalian Chinese hamster ovary (CHO) cell cultures with continuous phase kinetics transitions. Biochem. Eng. J. 2019, 142, 124–134. [Google Scholar] [CrossRef]
- Kirsch, B.J.; Bennun, S.V.; Mendez, A.A.; Johnson, A.S.; Wang, H.; Qiu, H.; Li, N.; Lawrence, S.M.; Bak, H.; Betenbaugh, M.J. Metabolic analysis of the asparagine and glutamine dynamics in an industrial Chinese hamster ovary fed-batch process. Biotechnol. Bioeng. 2022, 119, 807–819. [Google Scholar] [CrossRef]
- Kontoravdi, C.; Pistikopoulos, E.N.; Mantalaris, A. Systematic development of predictive mathematical models for animal cell cultures. Comput. Chem. Eng. 2010, 34, 1192–1198. [Google Scholar] [CrossRef]
- Kiparissides, A.; Georgakis, C.; Mantalaris, A.; Pistikopoulos, E.N. Design of in silico experiments as a tool for nonlinear sensitivity analysis of knowledge-driven models. Ind. Eng. Chem. Res. 2014, 53, 7517–7525. [Google Scholar] [CrossRef]
- Kiparissides, A.; Pistikopoulos, E.N.; Mantalaris, A. On the model-based optimization of secreting mammalian cell (GS-NS0) cultures. Biotechnol. Bioeng. 2015, 112, 536–548. [Google Scholar] [CrossRef] [PubMed]
- Kelley, B.; Renshaw, T.; Kamarck, M. Process and operations strategies to enable global access to antibody therapies. Biotechnol. Prog. 2021, 37, e3139. [Google Scholar] [CrossRef] [PubMed]
- Lohmann, L.J.; Strube, J. Accelerating biologics manufacturing by modeling: Process integration of precipitation in mAb downstream processing. Processes 2020, 8, 58. [Google Scholar] [CrossRef]
- Lohmann, L.J.; Strube, J. Process Analytical Technology for precipitation process integration into biologics manufacturing towards autonomous operation: mAb case study. Processes 2021, 9, 488. [Google Scholar] [CrossRef]
- Jones, W.; Gerogiorgis, D.I. Parametric analysis of mammalian cell (GS-NS0) culture performance for advanced mAb biopharmaceutical manufacturing. Comp. Aid. Chem. Eng. 2021, 50, 1923–1928. [Google Scholar]
- Jones, W.; Gerogiorgis, D.I. Dynamic optimisation and comparative analysis of fed-batch and perfusion bioreactor performance for monoclonal antibody (mAb) manufacturing. Comp. Aid. Chem. Eng. 2022, 51, 1117–1122. [Google Scholar]
- Jones, W.; Gerogiorgis, D.I. Dynamic simulation, optimisation and economic analysis of fed-batch vs. perfusion bioreactors for advanced mAb manufacturing. Comput. Chem. Eng. 2022, 165, 107855. [Google Scholar]
- Varadaraju, H.; Schneiderman, S.; Zhang, L.; Fong, H.; Menkhaus, T.J. Process and economic evaluation for monoclonal antibody purification using a membrane-only process. Biotechnol. Prog. 2011, 27, 1297–1305. [Google Scholar] [CrossRef]
- Mir-Artigues, P.; Twyman, R.M.; Alvarez, D.; Cerda Bennasser, P.; Balcells, M.; Christou, P.; Capell, T. A simplified techno-economic model for the molecular pharming of antibodies. Biotechnol. Bioeng. 2019, 116, 2526–2539. [Google Scholar] [CrossRef]
- Gupta, P.; Kateja, N.; Mishra, S.; Kaur, H.; Rathore, A.S. Economic assessment of continuous processing for manufacturing of biotherapeutics. Biotechnol. Prog. 2021, 37, e3108. [Google Scholar] [CrossRef]
- Gerogiorgis, D.I.; Barton, P.I. Steady-state optimization of a continuous pharmaceutical process. Comp. Aid. Chem. Eng. 2009, 27, 927–932. [Google Scholar]
- Rodman, A.D.; Gerogiorgis, D.I. Dynamic optimization of beer fermentation: Sensitivity analysis of attainable performance vs. product flavour constraints. Comput. Chem. Eng. 2017, 106, 582–595. [Google Scholar] [CrossRef]
- Rodman, A.D.; Fraga, E.S.; Gerogiorgis, D.I. On the application of a nature-inspired stochastic evolutionary algorithm to constrained multi-objective beer fermentation. Comput. Chem. Eng. 2018, 108, 448–459. [Google Scholar] [CrossRef]
- Shirahata, H.; Diab, S.; Sugiyama, H.; Gerogiorgis, D.I. Dynamic modelling, simulation and economic evaluation of two CHO cell-based production modes towards developing biopharmaceutical manufacturing processes. Chem. Eng. Res. Des. 2019, 150, 218–233. [Google Scholar] [CrossRef]
- Botelho Ferreira, K.; Benlegrimet, A.; Diane, G.; Pasquier, V.; Guillot, R.; De Poli, M.; Chappuis, L.; Vishwanathan, N.; Souquet, J.; Broly, H.; et al. Transfer of continuous manufacturing process principles for mAb production in a GMP environment: A step in the transition from batch to continuous. Biotechnol. Prog. 2022, 38, e3259. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, D.; Parulekar, S.J.; Cinar, A. A dynamic EFM-based model for antibody producing cell lines and model based evaluation of fed-batch processes. Biochem. Eng. J. 2020, 156, 107494. [Google Scholar] [CrossRef]
- Biegler, L.T. Solution of dynamic optimization problems by successive quadratic programming and orthogonal collocation. Comput. Chem. Eng. 1984, 8, 243–247. [Google Scholar] [CrossRef]
- Wächter, A.; Biegler, L.T. On the implementation of an interior-point filter line-search algorithm for large-scale nonlinear programming. Math. Program. 2006, 106, 25–57. [Google Scholar] [CrossRef]
- Hedengren, J.D.; Shishavan, R.A.; Powell, K.M.; Edgar, T.F. Nonlinear modeling, estimation and predictive control in APMonitor. Comput. Chem. Eng. 2014, 70, 133–148. [Google Scholar] [CrossRef]
- Fan, Y.; Jimenez Del Val, I.; Müller, C.; Sen, J.W.; Rasmussen, S.K.; Kontoravdi, C.; Weilguny, D.; Andersen, M.R. Amino acid and glucose metabolism in fed-batch CHO cell culture affects antibody production and glycosylation. Biotechnol. Bioeng. 2015, 112, 521–535. [Google Scholar] [CrossRef]
- Zhou, W.; Chen, C.C.; Buckland, B.; Aunins, J. Fed-batch culture of recombinant NS0 myeloma cells with high monoclonal antibody production. Biotechnol. Bioeng. 1997, 55, 783–792. [Google Scholar] [CrossRef]
- Mason, G.F.; Gruetter, R.; Rothman, D.L.; Behar, K.L.; Shulman, R.G.; Novotny, E.J. Simultaneous determination of the rates of the TCA cycle, glucose utilization, α-ketoglutarate/glutamate exchange, and glutamine synthesis in human brain by NMR. J. Cereb. Blood Flow Metab. 1995, 15, 12–25. [Google Scholar] [CrossRef]
- Farid, S.S. Process economics of industrial monoclonal antibody manufacture. J. Chromatogr. B 2007, 848, 8–18. [Google Scholar] [CrossRef]
- Kappatou, C.D.; Mhamdi, A.; Campano, A.Q.; Mantalaris, A.; Mitsos, A. Model-based dynamic optimization of monoclonal antibodies production in semibatch operation—Use of reformulation techniques. Ind. Eng. Chem. Res. 2018, 57, 9915–9924. [Google Scholar] [CrossRef]
- Tong, X.; Li, X.; Pratt, N.L.; Hillen, J.B.; Stanford, T.; Ward, M.; Roughead, E.E.; Lai, E.C.-C.; Shin, J.-Y.; Cheng, F.W.; et al. Monoclonal antibodies and Fc-fusion protein biologic medicines: A multinational cross-sectional investigation of accessibility and affordability in Asia Pacific regions between 2010 and 2020. Lancet Reg. Health (West. Pac.) 2022, 26, 100506. [Google Scholar] [CrossRef]
- Nikita, S.; Thakur, G.; Jesubalan, N.G.; Kulkarni, A.; Yezhuvath, V.B.; Rathore, A.S. AI-ML applications in bioprocessing: ML as an enabler of real time quality prediction in continuous manufacturing of mAbs. Comput. Chem. Eng. 2022, 164, 107896. [Google Scholar] [CrossRef]
- Ellis, M.; Durand, H.; Christofides, P.D. A tutorial review of economic model predictive control methods. J. Proc. Control 2014, 8, 1156–1178. [Google Scholar] [CrossRef]




| Run | Code | GLC (mM) | GLN (mM) | ASN (mM) | All Other Substrates (mM) |
|---|---|---|---|---|---|
| 1 | GLC_100mM | 100 | 20 | 20 | 20 |
| 2 | GLC_30mM | 30 | 20 | 20 | 20 |
| 3 | GLN_20mM | 100 | 20 | 20 | 20 |
| 4 | GLN_60mM | 100 | 60 | 20 | 20 |
| 5 | ASN_20mM | 100 | 20 | 20 | 20 |
| 6 | ASN_100mM | 100 | 20 | 100 | 20 |
| Objective function: | |
| s.t: | |
| The process model: | |
| The set of ineq. constraints: | |
| The control vector: | |
| The set of initial conditions: |
| Run | Initial Condition | (Cells mL–1) | (%) | (mg L–1) | (%) |
|---|---|---|---|---|---|
| 1 | GLC_100mM | 1.329 × 1010 | - | 35.73 | - |
| 2 | GLC_30mM | 1.905 × 1010 | 43.34 | 52.70 | 47.50 |
| 3 | GLN_20mM | 1.419 × 1010 | 6.77 | 39.68 | 11.06 |
| 4 | GLN_60mM | 1.353 × 1010 | 1.81 | 37.44 | 4.79 |
| 5 | ASN_20mM | 1.566 × 1010 | 17.83 | 43.58 | 21.97 |
| 6 | ASN_100mM | 1.566 × 1010 | 17.83 | 43.57 | 21.94 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jones, W.; Gerogiorgis, D.I. Dynamic Optimisation of Fed-Batch Bioreactors for mAbs: Sensitivity Analysis of Feed Nutrient Manipulation Profiles. Processes 2023, 11, 3065. https://doi.org/10.3390/pr11113065
Jones W, Gerogiorgis DI. Dynamic Optimisation of Fed-Batch Bioreactors for mAbs: Sensitivity Analysis of Feed Nutrient Manipulation Profiles. Processes. 2023; 11(11):3065. https://doi.org/10.3390/pr11113065
Chicago/Turabian StyleJones, Wil, and Dimitrios I. Gerogiorgis. 2023. "Dynamic Optimisation of Fed-Batch Bioreactors for mAbs: Sensitivity Analysis of Feed Nutrient Manipulation Profiles" Processes 11, no. 11: 3065. https://doi.org/10.3390/pr11113065
APA StyleJones, W., & Gerogiorgis, D. I. (2023). Dynamic Optimisation of Fed-Batch Bioreactors for mAbs: Sensitivity Analysis of Feed Nutrient Manipulation Profiles. Processes, 11(11), 3065. https://doi.org/10.3390/pr11113065








