1. Introduction
In general terms, gene therapy is the introduction of a specific cell function through the modification of the cellular genetic material of a patient for the treatment of hereditary or acquired genetic diseases. The effective delivery of genes to the target tissue/cells is carried out using gene delivery vehicles known as vectors, which remains a critical step in gene therapy protocols [
1,
2]. This area has seen several approved treatments based on viral vectors that vary from vector-based cancer therapies to the treatment of monogenic disorders with life-long benefits [
2,
3]. The recombinant adeno-associated virus (rAAV) is a versatile viral vector technology for gene therapy applications that may be designed for specific functional interventions. It has proven to be safe and efficient in preclinical and clinical evaluations because of its unique biological and physicochemical features, and rAAV may be employed in a wide range of therapeutic applications in various genetic disorders [
1,
2,
3,
4]. Although rAAV is one of the most effective vehicles for directly translating the genomic revolution into medicinal therapies, the manufacturing of rAAV viral vectors remains challenging [
5], limiting the generalization of AAV-based treatments.
One of the technological limitations in upstream processing in rAAV manufacturing is the low rAAV yield in large-scale production [
5]. Low titers and a high variability in product quality are often the results of an upstream procedure involving an insufficient triple-plasmid transfection of suspension-based cell culture [
5]. The situation can be improved by following the Food and Drug Administration’s initiative of process analytical technology (PAT), which requires understanding the process and a timely monitoring of critical process parameters (CPP) that affect critical quality attributes (CQA) [
6]. However, current techniques for monitoring the rAAV manufacturing in bioreactors are expensive, laborious, and time-consuming. Sample taking is usually required to measure the CPPs, such as the cell density and metabolites, and the quantification of the CQAs, such as a rAAV genome titer using a quantitative Polymerase Chain Reaction (qPCR)/droplet digital Polymerase Chain Reaction (ddPCR) or a viral capsid titer using an enzyme-linked immunosorbent assay (ELISA), takes one day to complete [
7]. Recently, in situ monitoring technologies, such as Raman spectroscopy [
8,
9] and fluorescence spectroscopy [
10,
11,
12], have been developed to estimate the cell density and metabolites in mammalian cell cultures in real-time, but have not been reported as detecting the rAAV titer. Moreover, the setup of a spectroscopy system is costly in terms of the investment and calibration effort [
13]. On the other hand, one solution is to develop fast and cost-effective real-time process monitoring technologies through mathematical models of the production [
14,
15,
16]. Mathematical modeling (MM) is an essential component of process systems engineering (PSE) [
17,
18,
19] and is helpful in monitoring through process state estimation [
14,
17,
20]. Estimation algorithms that rely on the mathematical model can estimate variables that are not directly observable and can predict meaningful process outputs and attributes that are either not measurable online or can only be measured at a low sampling frequency [
14,
17,
19].
The mathematical representations of the rAAV production for state estimation and output prediction can be made with mechanistic kinetic models [
21,
22,
23]. A mechanistic kinetic model can be classified as unstructured and structured [
14]. An unstructured model enables the macro-modeling of the functionality of the bioreactor, and it can provide an insight toward the underlying macro-scale phenomena of the upstream process. This kind of model can be used to depict the dynamics of the cell density, viability, nutrient/metabolite concentrations, and product titer [
14], which could be determined by online applications (where the data are analyzed in a continuous mode and the sensed variable must be measured more frequently than it can change in the process) and offline applications (where samples are required to be taken and analyzed in the laboratory after proper pre-treatments) [
18,
24]. Narayanan et al. [
21,
22] and Fernandes-Platzgummer et al. [
23] have used an unstructured model for monoclonal antibody (mAb) production, which is also based on mammalian cell cultures as rAAV production. It is a good starting point for designing a mechanistic model for rAAV production without considering the complexity of the triple-plasmid transfection process. On the other hand, structured mechanistic models are more complex than unstructured ones because they describe details about the intracellular environment of a homogenous cell population [
14]. The structured model of rAAV production presented by Nguyen et al. [
25] is the first proposed model and is essential for the mechanistic understanding of rAAV production pathways. However, it is not feasible to be extended as an application of soft sensors in bioreactors because it describes the kinetic behavior of transient transfection at the subcellular level. It is most appropriate for cell-line development, where genome-level characteristics of the cells are altered to achieve certain desired process behaviors.
A simple unstructured mechanistic kinetic model (UMKM) has a low prediction ability, and it is not enough to process state estimation because it is improbable that a single set of parameter values enables a kinetic model to satisfactorily for several data sets collected under distinct operating circumstances [
26]. Given this, UMKM is commonly implemented with the Kalman filter approach [
27] to improve the prediction accuracy and generate predictions in between sampling instances. In various data analysis methods, the Kalman filter and its non-linear extensions, such as the extended Kalman filter, are powerful tools for predicting values of the unobserved states. Although there are several applications of the extended Kalman filter for mAb production [
22,
28] and other cultivation processes [
29,
30], its application to the rAAV production process has not been reported.
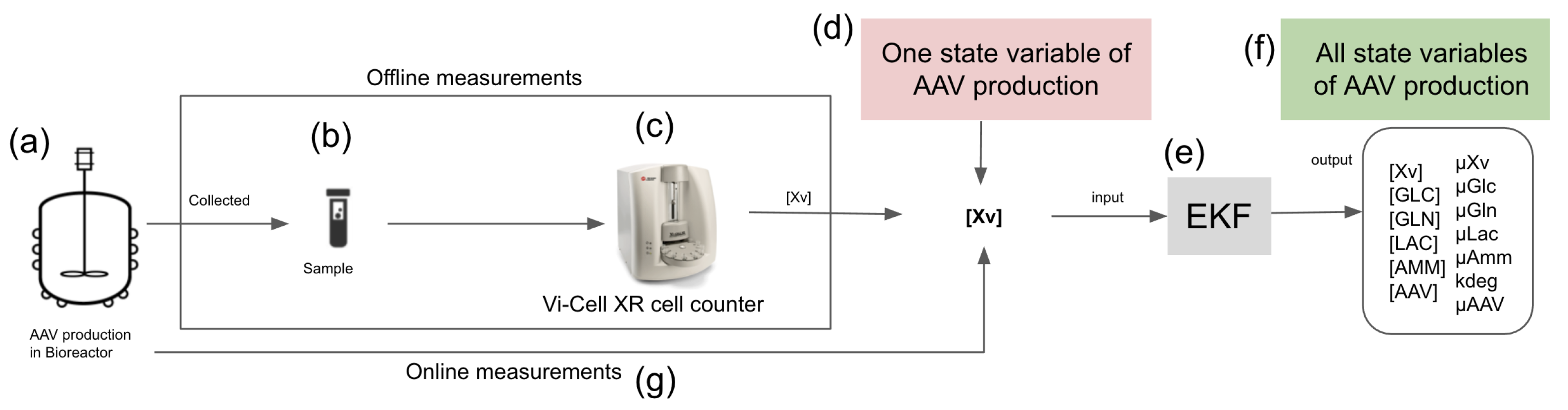
In this research, an extended Kalman filter (EKF) was proposed to supervise the rAAV production using only online viable cell density (Xv) measurements to estimate the other process state variables, including glucose (GLC) concentration, glutamine (GLN) concentration, lactate (LAC) concentration, ammonium (AMM) concentration, and rAAV viral titers that are measured at a low sampling frequency. The proposed EKF was applied to the cell expansion phase (CEP) and viral vector production phase (VVPP) of the upstream process using a UMKM based on mass balances (only dependent on Xv measurements) as a process model. Three datasets were used in the development of the proposed EKF, and the data were collected from the production of rAAV by a triple-plasmid transfection of HEK293SF-3F6 cells in three different environments: the shake-flasks dataset (offline data), bioreactor 1 dataset (offline data), and bioreactor 2 dataset (online and offline data). The parameters used in the UMKM were estimated with a neural ordinary differential equation and Bayesian inference approaches using the bioreactor 1 dataset. Furthermore, they were also estimated during the execution of EKF using the joint estimation method, and the EKF parameters were obtained from the shake-flasks and bioreactor 1 datasets. Our approach was evaluated with the bioreactor 1 and 2 datasets, and we showed that the proposed approach can only use the online Xv measurements and estimate the GLC, LAC, and rAAV viral titer effectively. The proposed approach is the first EKF approach developed to monitor rAAV production, and it uses only one device as opposed to the current approaches, which require multiple assays/devices. Our results indicate that the proposed EKF has the potential to be generalized and extended to an online soft-sensor, and to be classified as a cost-effective and rapid approach to monitoring rAAV production.
4. Discussion
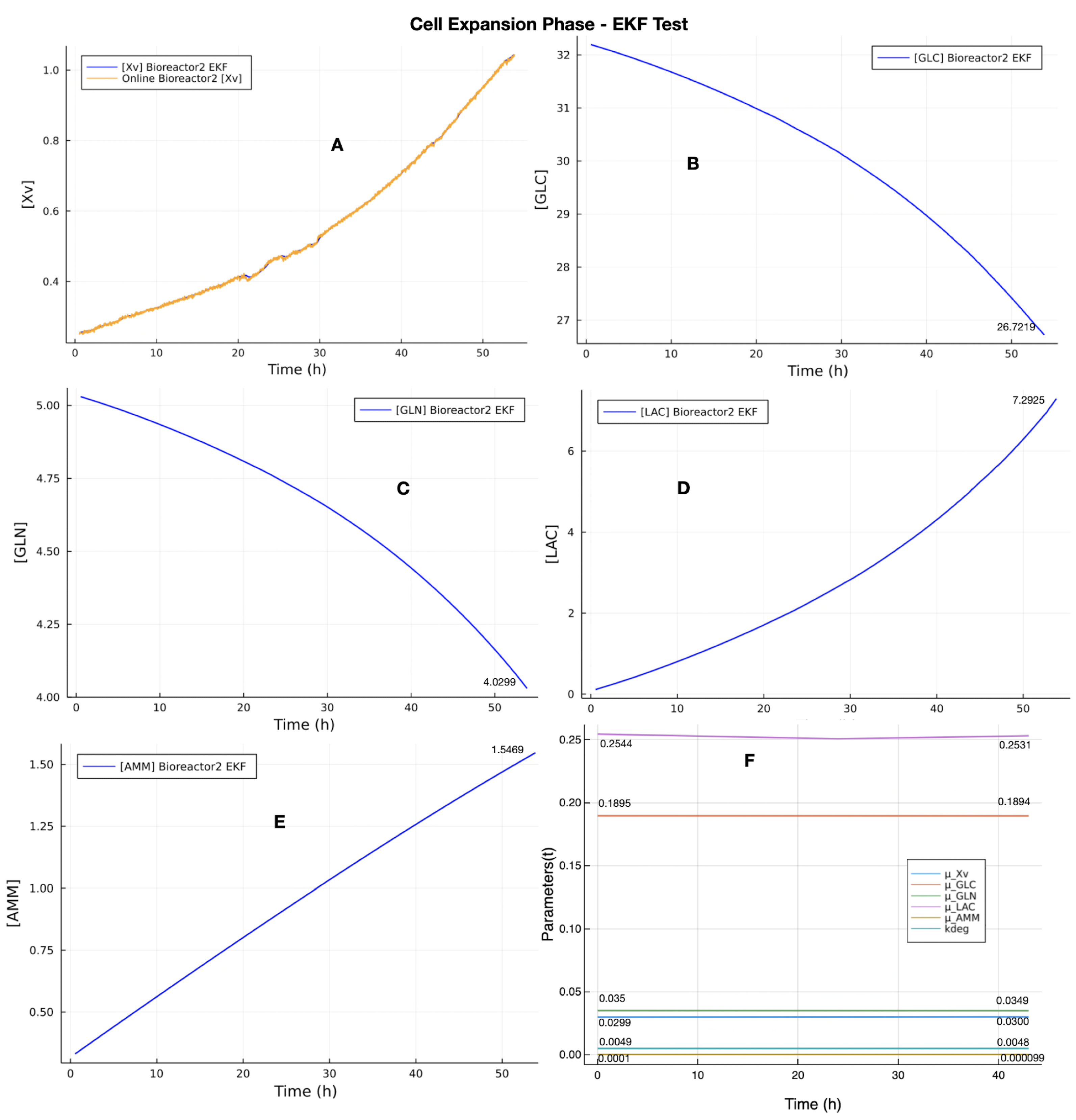
The main result of the evaluation came from the EKF test (
Section 3.3). However, the EKF test depended on the results of the UMKM parameters estimation (
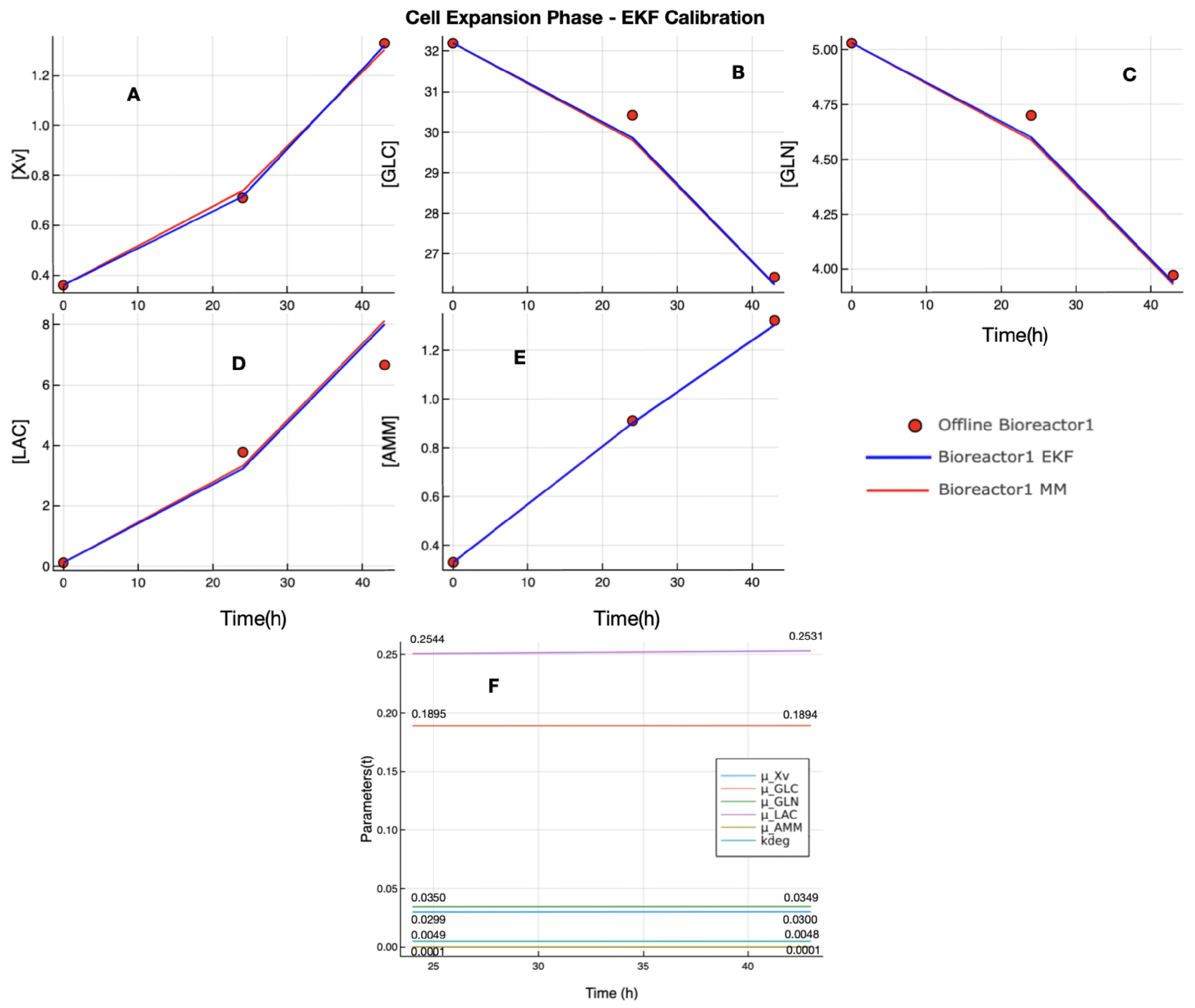
Section 3.1) and the EKF calibration (
Section 3.2) to be performed, as described in
Section 2.6. The UMKM parameters estimation performed by NODE and Bayesian inference found congruent parameters values (
Section 3.1). Besides these values being used as the initial condition in the state variables vector
, they were also used in the EKF calibration to obtain the final values of EKF parameters
and
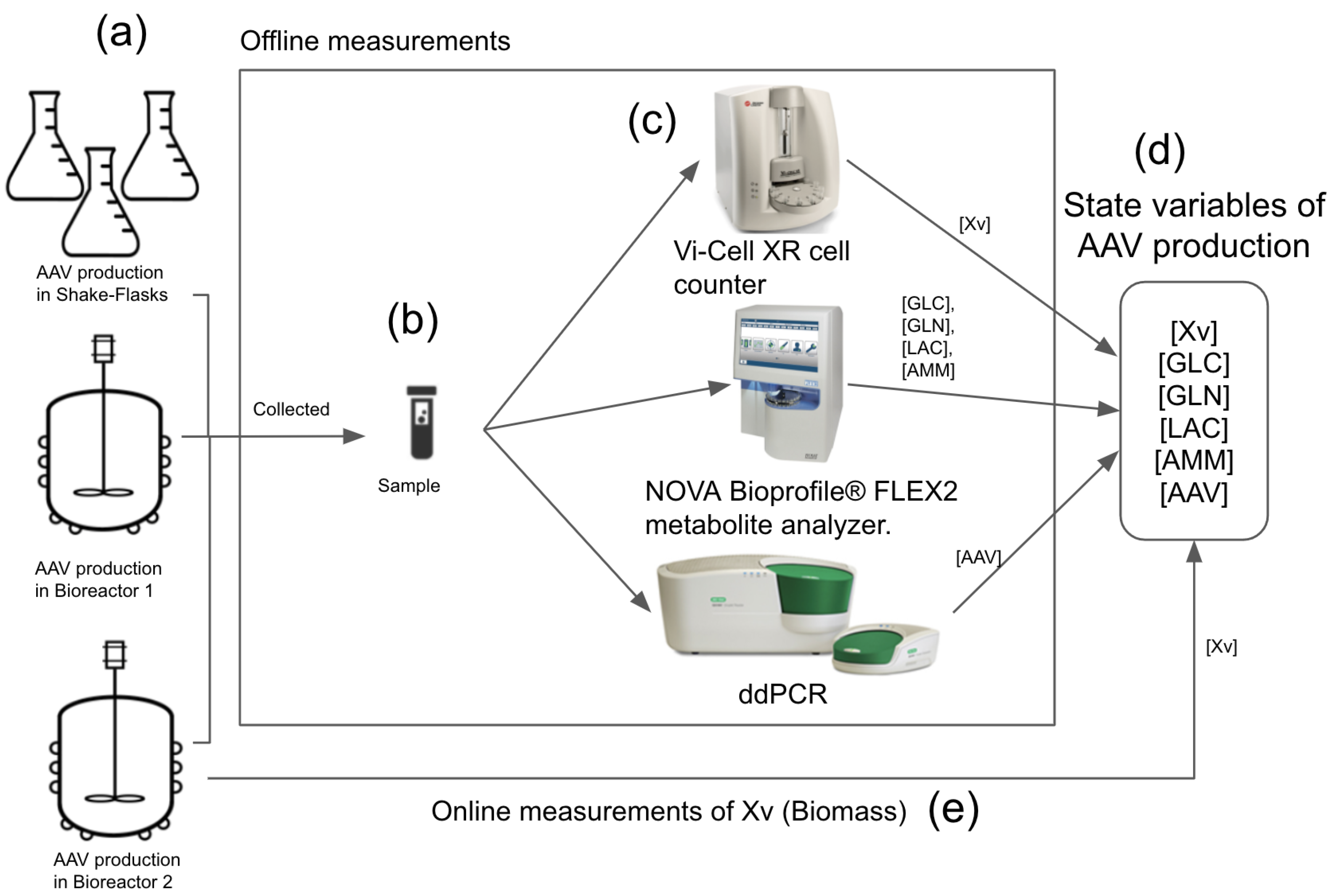
to be used in the EKF test. The results of the EKF test showed that the proposed EKF, with the process model (UMKM) depending only on the online viable cells (Xv) measurements, was able to estimate the other state variables of rAAV production, with values very close to the offline measurements. These results imply that the proposed EKF has solid potential to evolve into an online soft-sensor application and to be viewed as a low-cost and fast solution for monitoring rAAV production throughout the upstream process at the macroscale. This is because the offline/online measurement process of the state variables (viable cell density, metabolites, and rAAV viral titer ) used to generate the datasets required the use of multiple assays/devices to perform the measurements of all state variables (as described in
Section 2.4.5 and
Figure 2), whereas the proposed EKF requires only one (as described in
Section 3.3 and
Figure 1). The reason for this is that the Xv measurement (viable cells) will be an input to the EKF, which is then used to estimate all state variables of rAAV production. It can consequently reduce the costs of frequent sampling. Furthermore, the fact that EKF relies only on Xv measurements to estimate all state variables is a desirable step forward for online soft sensors since they are used to estimate state variables over time that are difficult to measure directly, or that can only be measured at a low sampling rate [
18]. However, despite significant results achieved by the proposed EKF, it is important to point out that it has limitations and needs more tests and further improvements. The proposed approach cannot contribute to the understanding of the rAAV production mechanistic model and should be considered as a limitation. Furthermore, more tests and improvements should be considered to extend the proposed EKF to a stable soft-sensor application that is ready to be used in the industry. Three future research directions might be considered. The first direction is related to increasing the complexity of the mechanistic model. The UMKM and EKF had the same performance in estimating AMM, but they did not perform a prediction near the observed data properly. The main reason for this discrepancy is that the conversion between NH4+, NH3(aq), and NH3(g) is not considered in the model. The sparging of oxygen and constant air overlay flow would remove the NH3(g) so that the reaction equilibrium shifts to the direction of converting NH4+ to NH3(g), hence decreasing NH4+ at the end of process. This could be solved by introducing an AMM removal term to Equation (
5) [
35,
52]. It is noteworthy that the trend of Xv during VVPP is not exponential. This may be because of transfection, nutrition limits (GLC and GLN), and toxic compound accumulation (LAC and AMM). Second, an additional improvement is estimating the parameters with other methods to confirm the convergence obtained. An option includes calibrating parameters outside the EKF calculation with an outer optimization routine [
29]. Third, the proposed EKF needs validation with different datasets containing offline and online measurements of rAAV productions. The datasets used in this initial study allow us to test the proposed EKF, aiming to have a preliminary idea about its potential as an approach to monitoring the rAAV production using only the Xv measurements to estimate the state variables of rAAV production, but the datasets limit the final validation because of their small size and missing online and offline measurements.