Optimization of L-Citrulline Operon in Corynebacterium glutamicum for L-Citrulline Production
Abstract
:1. Introduction
2. Materials and Methods
2.1. Microorganism and Cultivation Conditions
2.2. Analytical Methods
2.3. Transcriptional Analysis
2.4. Construction of Plasmids and Strains
3. Results and Discussion
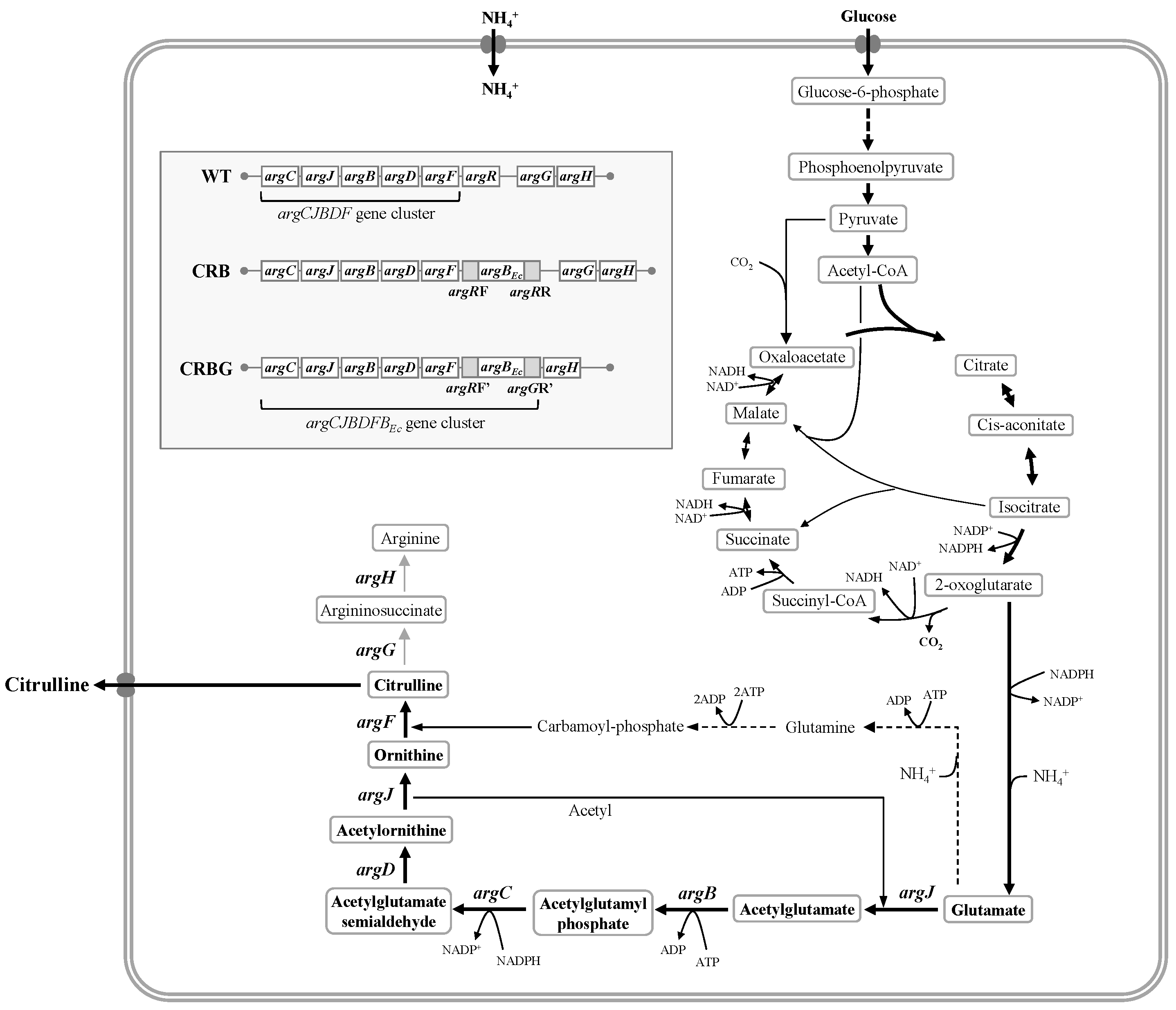
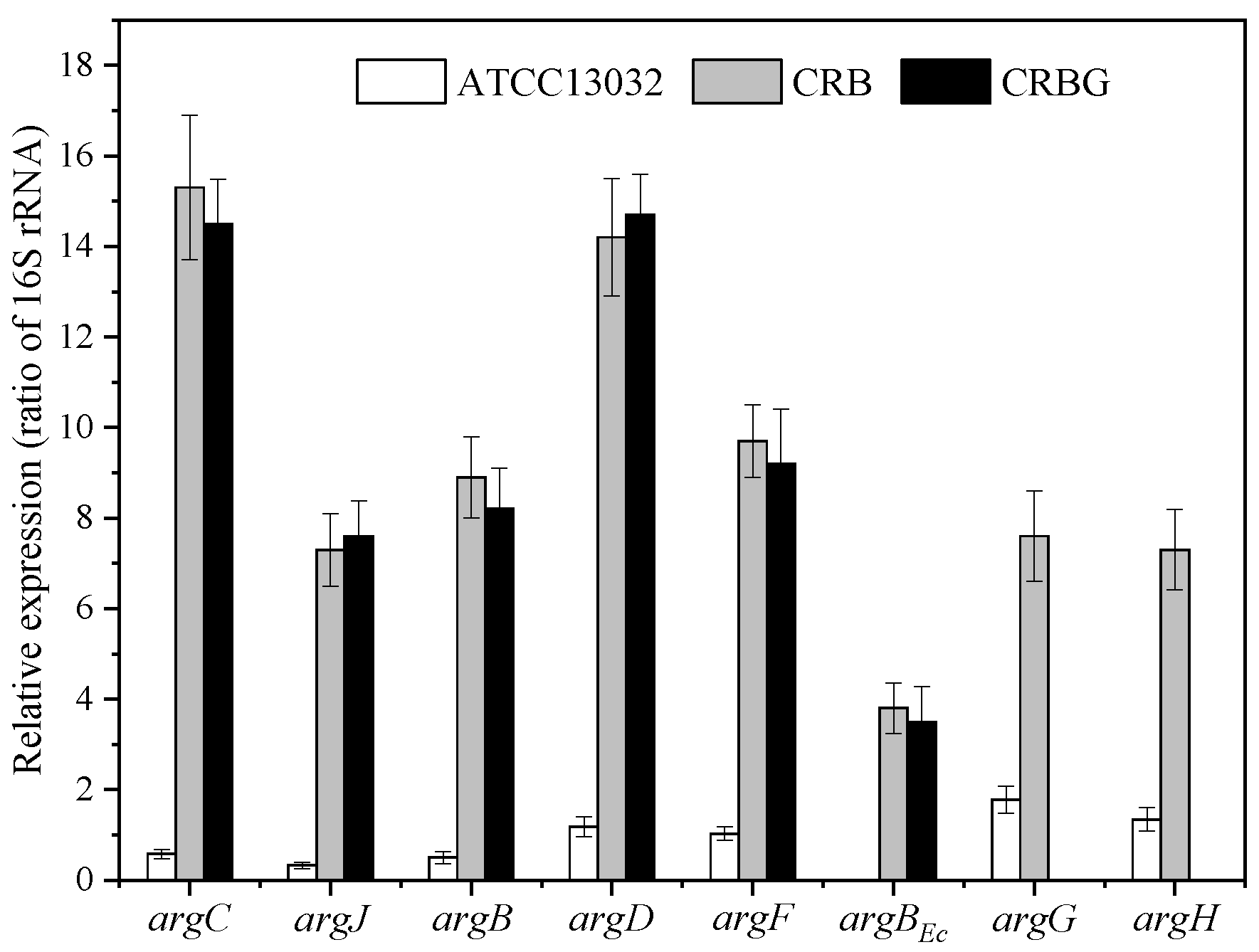
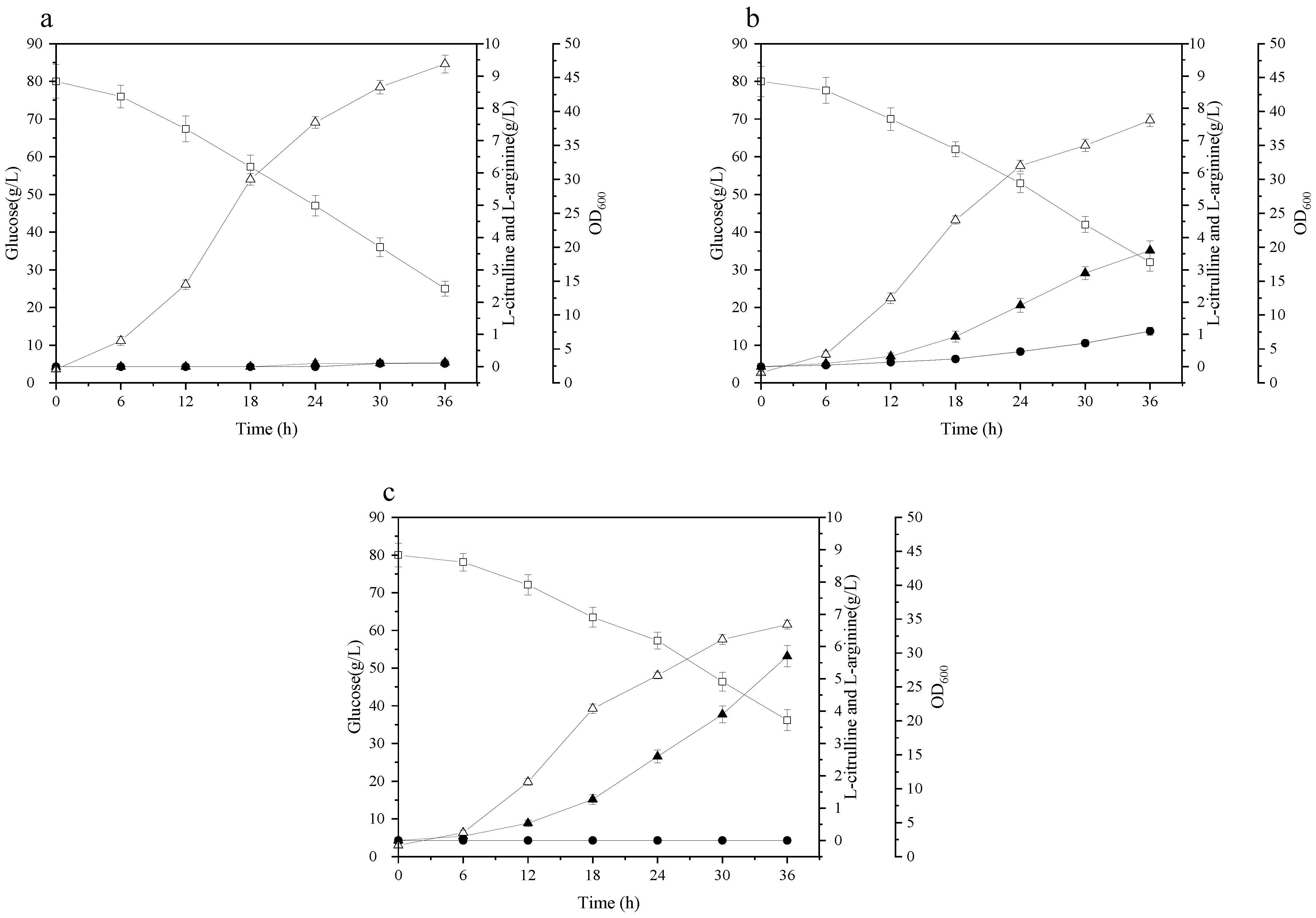
3.1. Effects of argR/argG Deletion and argBEc Expression on L-Citrulline Biosynthesis of C. glutamicum
3.2. Further Enhancing of L-Citrulline Biosynthesis by Plasmid-Based Overexpression of the Optimized L-Citrulline Operon
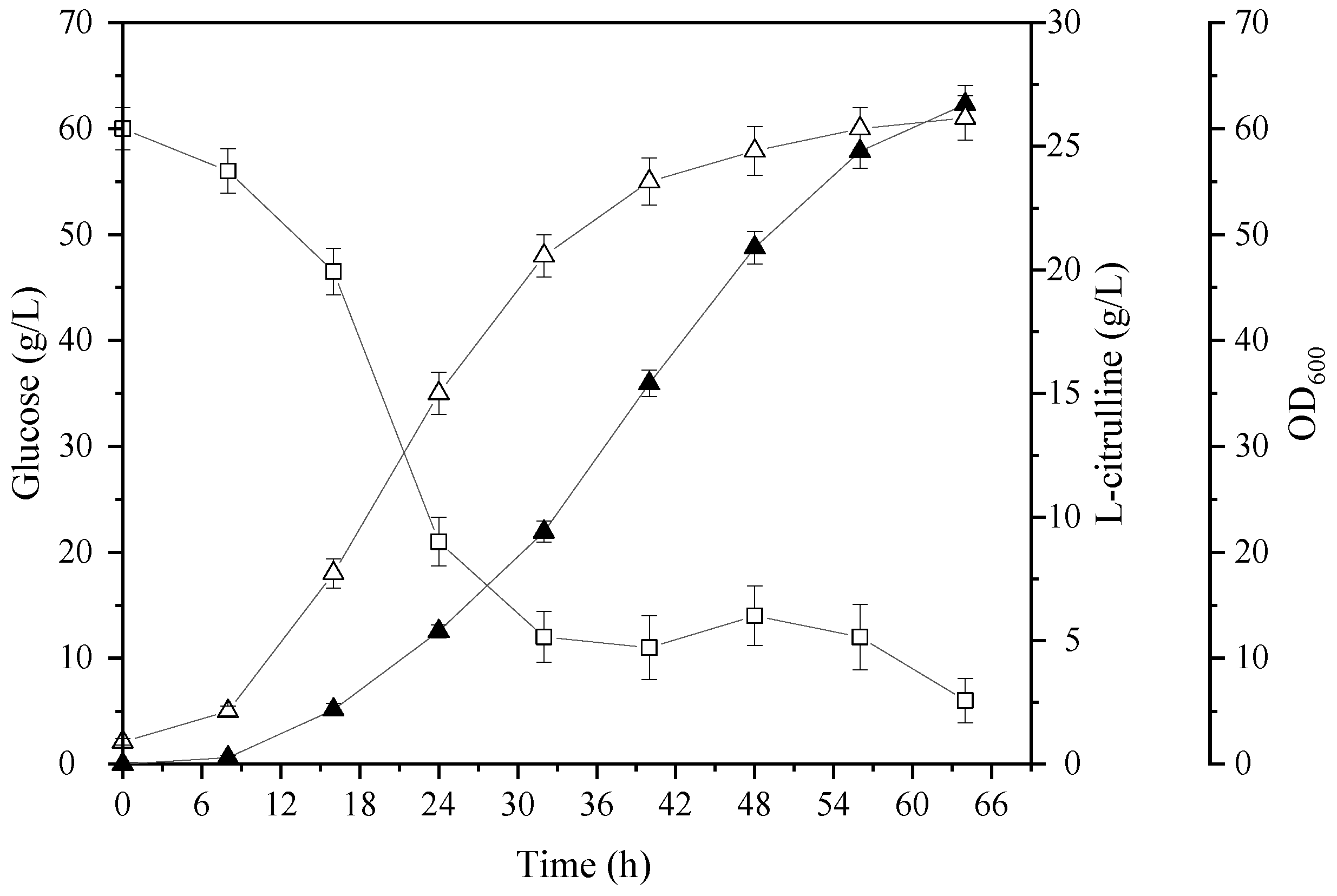
3.3. Fed-Batch Fermentation for L-Citrulline Production
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Breuillard, C.; Cynober, L.; Moinard, C. Citrulline and nitrogen homeostasis: An overview. Amino Acids 2015, 47, 685–691. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Sun, X.; Chen, X.; Liu, D.; Liu, L. Enzymatic production of L-citrulline by hydrolysis of the guanidinium group of L-arginine with recombinant arginine deiminase. J. Biotechnol. 2015, 208, 37–43. [Google Scholar] [CrossRef]
- Su, L.; Ma, Y.; Wu, J. Extracellular expression of natural cytosolic arginine deiminase from Pseudomonas putida and its application in the production of L-citrulline. Bioresour. Technol. 2015, 196, 176–183. [Google Scholar] [CrossRef] [PubMed]
- Shin, J.H.; Lee, S.Y. Metabolic engineering of microorganisms for the production of L-arginine and its derivatives. Microb. Cell Fact. 2014, 13, 166. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Kim, H.U.; Kim, T.Y.; Park, J.S.; Kim, S.S.; Lee, S.Y. Metabolic engineering of Corynebacterium glutamicum for L-arginine production. Nat. Commun. 2014, 5, 4618. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, J.H.; Wendisch, V.F. Production of amino acids—Genetic and metabolic engineering approaches. Bioresour. Technol. 2017, 245, 1575–1587. [Google Scholar] [CrossRef]
- Heider, S.A.E.; Wendisch, V.F. Engineering microbial cell factories: Metabolic engineering of Corynebacterium glutamicum with a focus on non-natural products. Biotechnol. J. 2015, 10, 1170–1184. [Google Scholar] [CrossRef]
- Jensen, J.V.K.; Eberhardt, D.; Wendisch, V.F. Modular pathway engineering of Corynebacterium glutamicum for production of the glutamate-derived compounds ornithine, proline, putrescine, citrulline, and arginine. J. Biotechnol. 2015, 214, 85–94. [Google Scholar] [CrossRef]
- Wendisch, V.F.; Jorge, J.M.P.; Pérez-García, F.; Sgobba, E. Updates on industrial production of amino acids using Corynebacterium glutamicum. World J. Microbiol. Biotechnol. 2016, 32, 105. [Google Scholar] [CrossRef]
- Becker, J.; Wittmann, C. Bio-based production of chemicals, materials and fuels—Corynebacterium glutamicum as versatile cell factory. Curr. Opin. Biotechnol. 2012, 23, 631–640. [Google Scholar] [CrossRef]
- Man, Z.; Rao, Z.; Xu, M.; Guo, J.; Yang, T.; Zhang, X.; Xu, Z. Improvement of the intracellular environment for enhancing L-arginine production of Corynebacterium glutamicum by inactivation of H2O2-forming flavin reductases and optimization of ATP supply. Metab. Eng. 2016, 38, 310–321. [Google Scholar] [CrossRef] [PubMed]
- Man, Z.; Xu, M.; Rao, Z.; Guo, J.; Yang, T.; Zhang, X.; Xu, Z. Systems pathway engineering of Corynebacterium crenatum for improved L-arginine production. Sci. Rep. 2016, 6, 28629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ikeda, M.; Mitsuhashi, S.; Tanaka, K.; Hayashi, M. Reengineering of a Corynebacterium glutamicum L-arginine and L-citrulline producer. Appl. Environ. Microbiol. 2009, 75, 1635–1641. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, M.; Rao, Z.; Dou, W.; Yang, J.; Jin, J.; Xu, Z. Site-directed mutagenesis and feedback-resistant N-acetyl-L-glutamate kinase (NAGK) increase Corynebacterium crenatum L-arginine production. Amino Acids 2012, 43, 255–266. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.Y.; Lee, J.; Lee, S.Y. Metabolic engineering of Corynebacterium glutamicum for the production of L-ornithine. Biotechnol. Bioeng. 2015, 112, 416–421. [Google Scholar] [CrossRef] [PubMed]
- Hao, N.; Mu, J.; Hu, N.; Xu, S.; Yan, M.; Li, Y.; Guo, K.; Xu, L. Improvement of L-citrulline production in Corynebacterium glutamicum by ornithine acetyltransferase. J. Ind. Microbiol. Biotechnol. 2015, 42, 307–313. [Google Scholar] [CrossRef]
- Eberhardt, D.; Jensen, J.V.K.; Wendisch, V.F. L-citrulline production by metabolically engineered Corynebacterium glutamicum from glucose and alternative carbon sources. AMB Express 2014, 4, 85. [Google Scholar] [CrossRef] [Green Version]
- Schäfer, A.; Tauch, A.; Jäger, W.; Kalinowski, J.; Thierbach, G.; Pühler, A. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: Selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 1994, 145, 69–73. [Google Scholar] [CrossRef]
- Xu, D.; Tan, Y.; Shi, F.; Wang, X. An improved shuttle vector constructed for metabolic engineering research in Corynebacterium glutamicum. Plasmid 2010, 64, 85–91. [Google Scholar] [CrossRef]
- Xu, H.; Dou, W.; Xu, H.; Zhang, X.; Rao, Z.; Shi, Z.; Xu, Z. A two-stage oxygen supply strategy for enhanced L-arginine production by Corynebacterium crenatum based on metabolic fluxes analysis. Biochem. Eng. J. 2009, 43, 41–51. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, R.; Bao, T.; Yang, T.; Xu, M.; Li, H.; Xu, Z.; Rao, Z. Moderate expression of the transcriptional regulator ALsR enhances acetoin production by Bacillus subtilis. J. Ind. Microbiol. Biotechnol. 2013, 40, 1067–1076. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Rao, Z.; Dou, W.; Xu, Z. The role of ARGR repressor regulation on L-arginine production in Corynebacterium crenatum. Appl. Biochem. Biotechnol. 2013, 170, 587–597. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Rao, Z.; Yang, J.; Xia, H.; Dou, W.; Jin, J.; Xu, Z. Heterologous and homologous expression of the arginine biosynthetic argC~H cluster from Corynebacterium crenatum for improvement of L-arginine production. J. Ind. Microbiol. Biotechnol. 2012, 39, 495–502. [Google Scholar] [CrossRef]
- Baumgart, M.; Unthan, S.; Rückert, C.; Sivalingam, J.; Grünberger, A.; Kalinowski, J.; Bott, M.; Noack, S.; Frunzke, J. Construction of a prophage-free variant of Corynebacterium glutamicum ATCC 13032 for use as a platform strain for basic research and industrial biotechnology. Appl. Environ. Microbiol. 2013, 79, 6006. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, J.; Man, Z.; Rao, Z.; Xu, M.; Yang, T.; Zhang, X.; Xu, Z. Improvement of the ammonia assimilation for enhancing L-arginine production of Corynebacterium crenatum. J. Ind. Microbiol. Biotechnol. 2017, 44, 443–451. [Google Scholar] [CrossRef]
- Xu, M.; Li, J.; Shu, Q.; Tang, M.; Zhang, X.; Yang, T.; Xu, Z.; Rao, Z. Enhancement of L-arginine production by increasing ammonium uptake in an AmtR-deficient Corynebacterium crenatum mutant. J. Ind. Microbiol. Biotechnol. 2019, 46, 1155–1166. [Google Scholar] [CrossRef]

| Strain or Plasmid | Description | Reference or Source |
|---|---|---|
| Strains | ||
| E. coli JM109 | General cloning host | TaKaRa |
| C. glutamicum ATCC13032 | Wild-type strain | ATCC |
| CRB | C. glutamicum ATCC13032 derivative with insertion of argBEc into argR gene | This work |
| CRBG | C. glutamicum ATCC13032 derivative with insertion of argBEc between argR and argG genes | This work |
| CRBGC | CRBG derivative harboring pDXW-10-argCJBDF | This work |
| CRBGCo | CRBG derivative harboring pDXW-10-argCJBDFBEc | |
| plasmids | ||
| pK18mobsacB | Kanr; vector for in-frame deletions | Lab stock [18] |
| pK18RB | Derived from pK18mobsacB, harboring argRF-argBEc-argRR fragment | This work |
| pK18RBG | Derived from pK18mobsacB, harboring argRF’-argBEc-argGR’ fragment | This work |
| pDXW-10 | Kanr; shuttle vector between E. coli and C. glutamicum | Lab stock [19] |
| pDXW-10-argCJBDF | Derived from pDXW-10, for constitutive expression of argCJBDF gene cluster | This work |
| pDXW-10-argCJBDFBEc | Derived from pDXW-10, for constitutive expression of the optimized L-citrulline operon argCJBDFBEc | This work |
| Names | Sequences (5′→3′) | Restriction Sites |
|---|---|---|
| RB-RF1 | CCGGAATTCTAACGCTAACAAGCTCTCATTTTGCAGATT | EcoRI |
| RB-RR1 | TTACATTTAGCGGCGCAGCTTCTCCGACGGAC | |
| RB-RF2 | TAGCTTAAGTTTTGTTGGCCGGAGGCGCAGTGATGAACTGCTGGTTTCTACAGATC | |
| RB-RR2 | CGCGGATCCGCGCCCGCTGAGTAATTCACCTA | BamHI |
| RB-BF | AGCTGCGCCGCTAAATGTAACGAAGACAACTTACTGAAAG | |
| RB-BR | GGCCAACAAAACTTAAGCTA | |
| RBG-RF | CCGGAATTCGCACGCCAAGCTCTCATTTTGCAGATTTTG | EcoRI |
| RBG-RR | GGTGCGCAGCATCGCGATGTTGCCGG | |
| RBG-GF | GGCGCAGCTTTCCGGCATTGAATTTCGCTACAAGCGCGGCGTTGACGCACG | |
| RBG-GR | CGCGGATCCCTAAATACTTGAAGTTCTAGTCTGG | BamHI |
| RBG-BF | CCGGCAACATCGCGATGCTGCGCACCCCAATATTCGTTTCGGCTATGCGGAAAC | |
| RBG-BR | GAAATTCAATGCCGGAAAGCTGCGCC | |
| CF | AAGGAAAAAAGCGGCCGCAAATTCATGCTTTTACCCACTTGCAG | NotI |
| CR | GGAAGATCTGCAAAATGAGAGCTTGGCGTGCAGTGCG | BglII |
| CoR | AAGGAAAAAAGCGGCCGCGAAATTCAATGCCGGAAAGCTGCGCC | NotI |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Man, Z.; Li, J.; Cui, H.; Cai, Z.; Guo, J. Optimization of L-Citrulline Operon in Corynebacterium glutamicum for L-Citrulline Production. Processes 2022, 10, 2153. https://doi.org/10.3390/pr10102153
Man Z, Li J, Cui H, Cai Z, Guo J. Optimization of L-Citrulline Operon in Corynebacterium glutamicum for L-Citrulline Production. Processes. 2022; 10(10):2153. https://doi.org/10.3390/pr10102153
Chicago/Turabian StyleMan, Zaiwei, Jin Li, Huihui Cui, Zhiqiang Cai, and Jing Guo. 2022. "Optimization of L-Citrulline Operon in Corynebacterium glutamicum for L-Citrulline Production" Processes 10, no. 10: 2153. https://doi.org/10.3390/pr10102153





