Abstract
This study presents SpineCheck, a fully integrated deep-learning-based clinical decision support platform for automatic vertebra segmentation and Cobb angle (CA) measurement from scoliosis X-ray images. The system unifies end-to-end preprocessing, U-Net-based segmentation, geometry-driven angle computation, and a web-based clinical interface within a single deployable architecture. For secure clinical use, SpineCheck adopts a stateless “process-and-delete” design, ensuring that no radiographic data or Protected Health Information (PHI) are permanently stored. Five U-Net family models (U-Net, optimized U-Net-2, Attention U-Net, nnU-Net, and UNet3++) are systematically evaluated under identical conditions using Dice similarity, inference speed, GPU memory usage, and deployment stability, enabling deployment-oriented model selection. A robust CA estimation pipeline is developed by combining minimum-area rectangle analysis with Theil–Sen regression and spline-based anatomical modeling to suppress outliers and improve numerical stability. The system is validated on a large-scale dataset of 20,000 scoliosis X-ray images, demonstrating strong agreement with expert measurements based on Mean Absolute Error, Pearson correlation, and Intraclass Correlation Coefficient metrics. These findings confirm the reliability and clinical robustness of SpineCheck. By integrating large-scale validation, robust geometric modeling, secure stateless processing, and real-time deployment capabilities, SpineCheck provides a scalable and clinically reliable framework for automated scoliosis assessment.
1. Introduction
When the normal line of the spine in the frontal plane is disrupted by lateral curvatures called scoliosis, both aesthetic and functional problems occur. This distortion, measured by the Cobb angle (CA), is especially common in adolescents and can negatively affect respiratory functions as it progresses. Today, scoliosis diagnosis is still based on CA measurements made manually by experts on posteroanterior (PA) radiographs, and this method is both time-consuming and susceptible to observer-dependent errors. At the same time, repeated radiographic checks expose the patient to additional doses of radiation. Recent advances in image processing and deep learning enable faster and more accurate automatic segmentation and measurement of medical images, offering a clearer, more impartial, and repeatable workflow that reduces human-related error in the early diagnosis and follow-up of scoliosis.
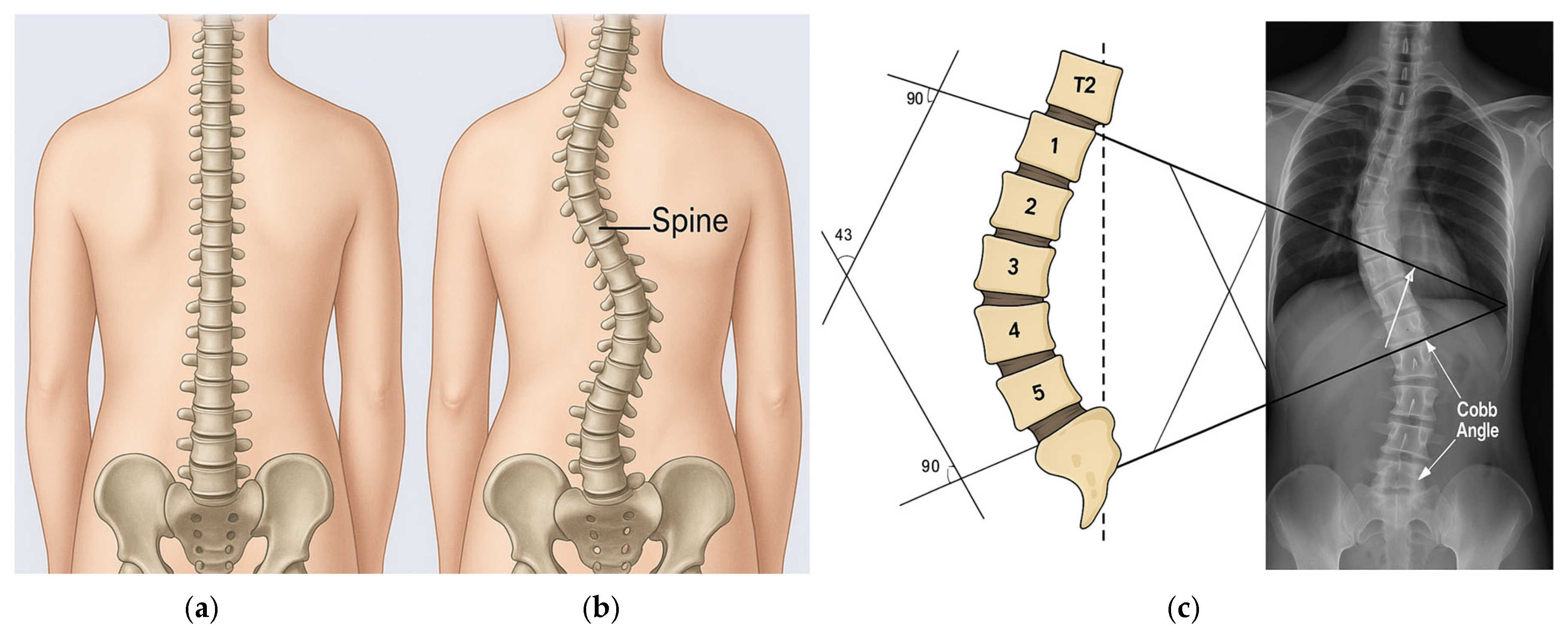
A healthy spine and a scoliosis case are shown in Figure 1. Simple manual measurement methods and classical image processing techniques have been used in scoliosis measurement for many years. The most basic method for a physician was to determine the most curved vertebrae on the X-ray and measure the CA [1] with a ruler–protractor. This simple manual angle measurement method has been accepted as the gold standard in scoliosis assessment and is still widely used in clinics [2]. An example of manual CA calculation is also given in Figure 1c. The fact that this human-based process can be slow, variable, and inaccurate has led researchers to computer-aided automated methods.
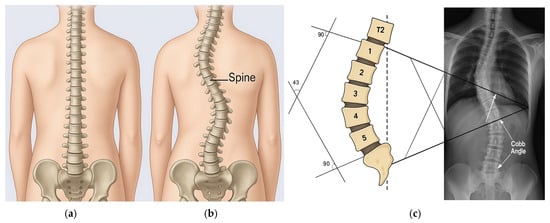
Figure 1.
(a) A healthy spine; (b) A scoliosis case; (c) Cobb Angle (CA) calculation showing vertebrae T2 and 1–5 forming the curvature, with perpendicular (90°) reference lines drawn to the superior and inferior endplates, yielding an example CA of 43°.
Table 1 shows one of the widely adopted scoliosis classifications based on CA [1] and it was used in our project. The table maps the measured spinal curvature (in degrees) to a class. As the CA increases, the scoliosis severity category progresses from normal to very severe. Ranges may vary slightly across sources.

Table 1.
CA measurements per severity.
We investigated the applicability of deep learning algorithms, particularly U-Net variants, for CA calculation and scoliosis diagnosis. To guarantee reproducible and automated measurements beyond subjective manual evaluation, the SpineCheck web-based platform was developed. Comparative evaluations of different architectures and preprocessing strategies guided the selection of the most suitable model based on accuracy, computational efficiency, and clinical applicability. A large X-ray dataset was partitioned into training, validation, and test sets to assess generalization, with augmentation and normalization employed to improve robustness against varying image quality. The trained U-Net segmentation model was integrated with an automatic end-vertebra detection and CA calculation module, whose results were statistically compared with manual measurements. Finally, a prototype web application was implemented to demonstrate integration into clinical workflows, highlighting its potential as a practical tool for both academic research and medical practice.
1.1. Related Work
Before the widespread adoption of deep learning, scoliosis assessment relied on classical image processing and rule-based methods. Early studies experimented with edge detection, Hough transform-based line fitting, and active contour models to identify vertebral endplates and measure spinal curvature angles. These approaches provided a foundation but faced significant challenges in robustness. For example, Eltanboly et al. [3] introduced a semi-automatic 3D level set-based segmentation method using statistical shape priors, achieving promising results on limited datasets. However, traditional methods generally required manual parameter tuning, were sensitive to noise, and often failed in low-contrast radiographs. As a result, they did not deliver the fully automatic and reliable measurements needed for clinical practice, though they paved the way for subsequent machine learning approaches.
The advent of convolutional neural networks (CNNs) transformed medical image analysis, and scoliosis research quickly adopted deep learning. U-Net, with its encoder–decoder architecture and skip connections, became a standard tool for biomedical segmentation [4]. Horng et al. [5] applied U-Net, Dense U-Net, and Residual U-Net for spine segmentation on scoliosis X-rays, reporting that Residual U-Net yielded the most accurate CA estimations compared with manual expert measurements. Deng et al. [6] improved accuracy by combining U-Net with BiSeNet, while Nascimento et al. [7] proposed a cascade UNet3+ architecture that achieved high Dice scores on the AASCE 2019 dataset [8]. Similarly, Wang et al. [9] enhanced vertebra localization by employing a multi-stage CNN with shape constraints. These works illustrate that segmentation-based pipelines can achieve clinically relevant precision while offering the additional advantage of visual validation of spinal curves.
Despite their success, segmentation-based models require pixel-level annotations, which are labor-intensive to prepare. To reduce labeling burden, researchers also investigated landmark-based approaches. Instead of segmenting every vertebra, these methods detect specific key points—such as vertebral corners or centers—and derive the CA from their coordinates. Wu et al. [10] demonstrated that simple CNNs, such as BoostNet, could estimate vertebral corner points and indirectly calculate curvature. Khanal et al. [2] advanced this strategy with a two-stage model combining object detection with corner regression, producing results close to expert measurements. Yi et al. [11] further refined landmark localization by first identifying vertebral centers and then corner points, improving robustness. Landmark-based methods demand less annotation and computational power, yet their accuracy depends heavily on correct keypoint placement. Consequently, many pipelines integrate post-processing heuristics or combine segmentation with landmark detection to minimize anatomical inconsistencies.
The literature also reflects an evolution of U-Net itself into numerous derivatives [12,13,14,15,16,17,18,19,20]. Dense U-Net enhances gradient flow through dense connections, while Residual U-Net facilitates deeper training via skip residual blocks. Architectures such as UNet++, UNet3+, and Attention U-Net incorporate multi-scale feature fusion and adaptive focus mechanisms to improve segmentation under challenging conditions. Xu et al. [16] combined Residual U-Net with Transformer blocks (RUnT) to capture both local detail and global context, outperforming conventional CNN-only models. In MR imaging, Kim et al. [17] introduced BSU-Net for fine-grain segmentation of intervertebral discs, and S.V.V. et al. [18] applied MU-Net with Meijering filters for lumbar spine segmentation. Nascimento et al. [7] showed that cascade UNet3+ models achieved strong performance in scoliosis-specific datasets. Collectively, these studies highlight U-Net’s versatility and establish it as the backbone of scoliosis imaging research.
Beyond radiographs, alternative imaging modalities have also been explored. CT enables detailed three-dimensional visualization of the spine, which is valuable in surgical planning. Li et al. [19] used a U-Net-based model to segment vertebrae in preoperative CT images and calculate 3D CAs, reporting high correlation with conventional 2D angles. However, CT involves high radiation exposure, limiting its routine use. MRI, while radiation-free, is primarily employed for evaluating neurological involvement and soft tissues. Deep learning models such as BSU-Net [13] and MU-Net [14] have achieved accurate segmentation of vertebrae and discs in MR scans, though angular measurements remain less common.
Ultrasound has gained attention as a safe and portable alternative for scoliosis screening. Banerjee et al. [20] proposed Skip-Inception U-Net (SIU-Net), which fused multi-scale features to improve noisy ultrasound spine segmentation. Zhou et al. [21] introduced an automatic curvature angle estimation method based on clustering of vertebral contours in ultrasound images, demonstrating agreement with expert annotations. While still experimental, these studies suggest ultrasound could support non-invasive screening, especially for adolescents, provided challenges with image variability are resolved.
Other approaches have considered surface-based or photogrammetric techniques, such as Moiré topography and back-surface scanning, to estimate spinal asymmetry. Recent works combine posture photographs or depth camera data with AI classifiers to predict scoliosis presence [22]. Smartphone-based or wearable sensor systems have also been tested for curve estimation, though they currently lack the quantitative accuracy of radiographic CA measurements.
A number of studies integrate segmentation and angle calculation into automated pipelines. Tan et al. [23] proposed an algorithm that segments vertebral endplates, fits enclosing rectangles, and calculates slope differences to estimate CAs. More recent frameworks combine segmentation with geometric or regression-based refinements, ensuring consistency with clinical conventions. Comparative evaluations (e.g., Kumar et al. [22]) show that CNN-based methods can achieve mean absolute errors below 2–3°, comparable to inter-expert variability, positioning them as credible clinical decision support tools.
Despite the substantial progress in automated scoliosis analysis, several critical limitations persist in the current literature. Most existing studies primarily focus on algorithm-level segmentation or CA estimation performance in isolated experimental settings, with validations typically conducted on small- to medium-scale datasets, often ranging from fewer than 100 images to approximately 1000–2000 radiographs. In contrast, large-scale clinical validations remain scarce. Furthermore, although robust geometric methods for CA computation have been proposed, only a limited number of studies explicitly address the combined influence of segmentation uncertainty, vertebral outliers, and anatomical consistency on angle stability under realistic clinical conditions. Additionally, while numerous U-Net variants have been individually demonstrated, systematic comparisons of multiple U-Net family architectures under identical training, inference, and deployment constraints—particularly with respect to inference speed, GPU memory usage, and operational stability—are rarely reported. Finally, the majority of existing works remain confined to offline, research-oriented pipelines, offering little insight into real-time integration within clinical workflows or into the design of full-stack, deployment-ready systems.
To address these limitations, the present study introduces SpineCheck, a fully integrated clinical decision support platform. This system unifies deep learning-based vertebra segmentation, robust geometry-driven CA computation, and a web-based user interface into a single end-to-end system. Unlike many prior works, this platform is validated on a large-scale dataset of 20,000 scoliosis X-ray images. Furthermore, deployment-oriented comparison of five U-Net family architectures, assessing them not only for segmentation accuracy but also for computational efficiency, GPU memory, and software robustness. By establishing a reproducible, full-stack architecture that incorporates robust regression and anatomical modeling, this work sets a scalable, clinically oriented framework for automated scoliosis assessment, moving beyond isolated algorithmic proofs-of-concept.
1.2. Contributions
This study presents SpineCheck, an end-to-end, deep-learning-based platform for automatic vertebra segmentation and CA measurement from scoliosis X-ray images. Unlike prior works that primarily focus on algorithmic proof-of-concept solutions, SpineCheck is designed as a clinically oriented, secure, and deployable decision support system. The main contributions of this study are summarized as follows:
- A Fully Integrated and Secure Clinical AI Platform: We present a production-ready system that integrates deep learning-based vertebra segmentation, automated CA computation, and a web-based clinical interface. Built on a modern full-stack architecture (FastAPI, React, and PyTorch 2.6.0+cu118), the platform enables real-time interaction and visualization. A stateless “process-and-delete” backend ensures that no raw images or Protected Health Information (PHI) are permanently stored, strengthening data security and regulatory compliance.
- Comprehensive Multi-Model Evaluation of U-Net Family Architectures: Five U-Net-based segmentation models (U-Net, U-Net-2, Attention U-Net, nnU-Net, and UNet3++) are systematically evaluated under identical conditions. In addition to Dice performance, models are assessed for training time, inference speed, GPU memory usage, deployment complexity, and clinical suitability, providing a practical benchmark for medical AI deployment.
- Robust Geometry-Based CA Estimation with Outlier Suppression: A reliable geometric CA computation pipeline is proposed, combining minimum-area rectangle analysis with Theil–Sen regression for slope estimation and a spline-based anatomical spine model for automated outlier detection and correction. This design improves numerical stability in challenging scoliosis cases.
- Large-Scale Clinical Validation on a 20,000-Image Dataset: The system is validated on the publicly available Spinal-AI2024 dataset containing 20,000 scoliosis X-ray images. Performance is evaluated using MAE, Pearson correlation, and ICC metrics, with stratified error analysis, demonstrating strong agreement with expert measurement.
- Open, Reproducible, and Scalable Research Infrastructure: The full pipeline—covering preprocessing, annotation handling, inference, visualization, and secure stateless processing—is designed for reproducibility and scalability. The modular architecture supports both large-scale benchmarking and real-world clinical deployment without redesign or persistent data storage.
Through these contributions, SpineCheck advances the state of the art by bridging the gap between algorithm-level scoliosis analysis methods and secure, fully deployable, clinically reliable AI systems for automated CA measurement.
2. SpineCheck Architecture
2.1. Workflow and Computation Pipeline Architecture
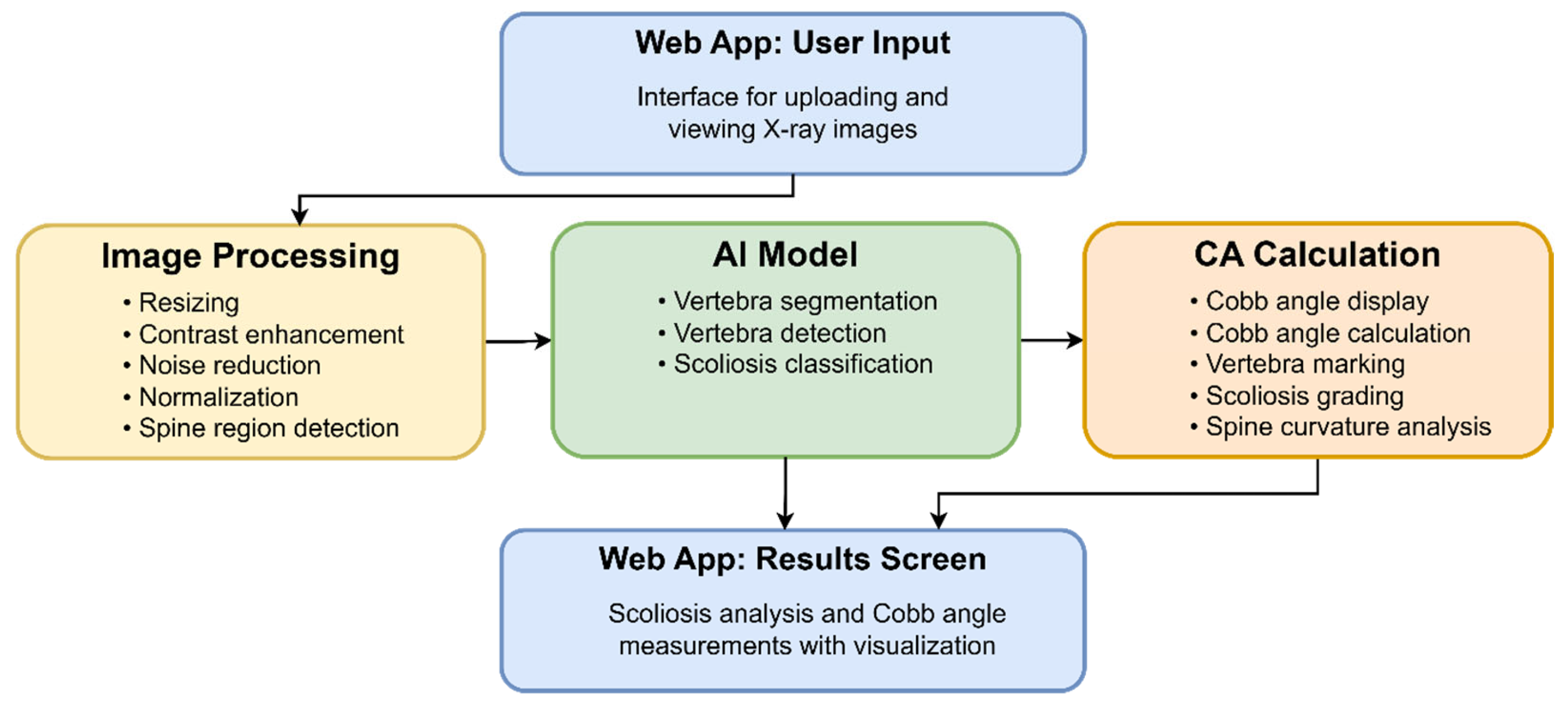
The main components of the workflow and computation pipeline architecture are shown in Figure 2. X-ray images are uploaded to the program via a web application interface. The images then go through Image Processing, AI Model, CA Calculation, Results, and Visualization modules. Finally, the visual results and CA estimations are displayed to the user. The generated images, such as an X-ray with a red curve overlay and a vertebral segmentation image, can be saved to disk.
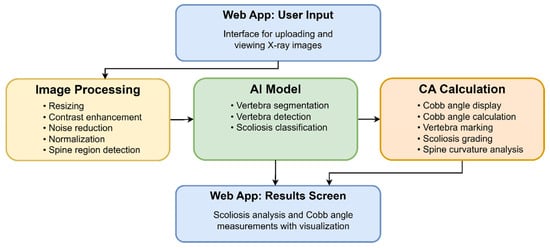
Figure 2.
The components of the workflow and computation pipeline. The web application layer represents the user interface where the clinician or technician uploads X-ray images via a browser and monitors the analysis progress. Here, patient information is entered, the image file (DICOM, PNG, JPG, etc.) is uploaded to the system, and visual notifications are received regarding the next stages of the process. This layer also handles basic user experience functions such as session management, access control, and file upload errors.
The raw X-ray image undergoes some preprocessing to make it compatible with the system. First, all images are scaled to 512 × 512 pixels. This process ensures that all images are the same scale during processing, helping the system operate more efficiently and quickly. The contrast of the image is then increased. This enhances important areas, such as the spine, and makes it easier for the system to recognize these structures. Noise and small, unwanted details are then filtered out. This step is performed to improve system performance, particularly in blurry or noisy images. In the next stage, the pixels in the image are adjusted to a specific range (usually between 0 and 1). This normalization process ensures that images of varying brightness are presented to the system uniformly, allowing the model to learn more accurately. Finally, the system automatically detects only the region containing the spine and crops this portion of the image. The model focuses solely on the spine and avoids wasting time on unnecessary areas. This step both reduces processing time and increases the accuracy of the results.
X-ray data, which has been processed through image processing steps and normalized to the spine region, is fed into a deep learning-based artificial intelligence model. A vertebra segmentation network based on the U-Net architecture was used for this task. The U-Net’s encoder–decoder model performs vertebra segmentation at the pixel level by learning both global structure and fine details. Each vertebra in the spine is clearly distinguished from its neighbors and semantically labeled.
The segmentation output is in the form of masks of vertebral boundaries, which also allow for the extraction of vertebral center locations (centroid points). This way, not only shape but also positional information is obtained. In addition to vertebral segmentation, SpineCheck reports a scoliosis severity label that is deterministically derived from the estimated CA using the thresholds given in Table 1 (<10°: normal; 10–24°: mild; 25–39°: moderate; 40–49°: severe; ≥50°: very severe). No separate classification network was trained.
Analysis data from the AI Model is processed by the CA Calculation module. The CA is calculated between the identified vertebrae, segmentation masks are overlaid onto the original X-ray image with translucent colors (“overlay drawing”), and labels are placed on the vertebrae. In the final stage, both the classified scoliosis degree and the CAs are summarized on a single screen. Clinicians, technicians, or physicians can review the angle using interactive measurement tools and review data history on this screen. This 5-stage process takes an X-ray image and processes it step-by-step through automatic segmentation, angle measurement, classification, and visual reporting, providing the end user with clinically relevant, directly interpretable outputs.
2.2. Project Components Architecture
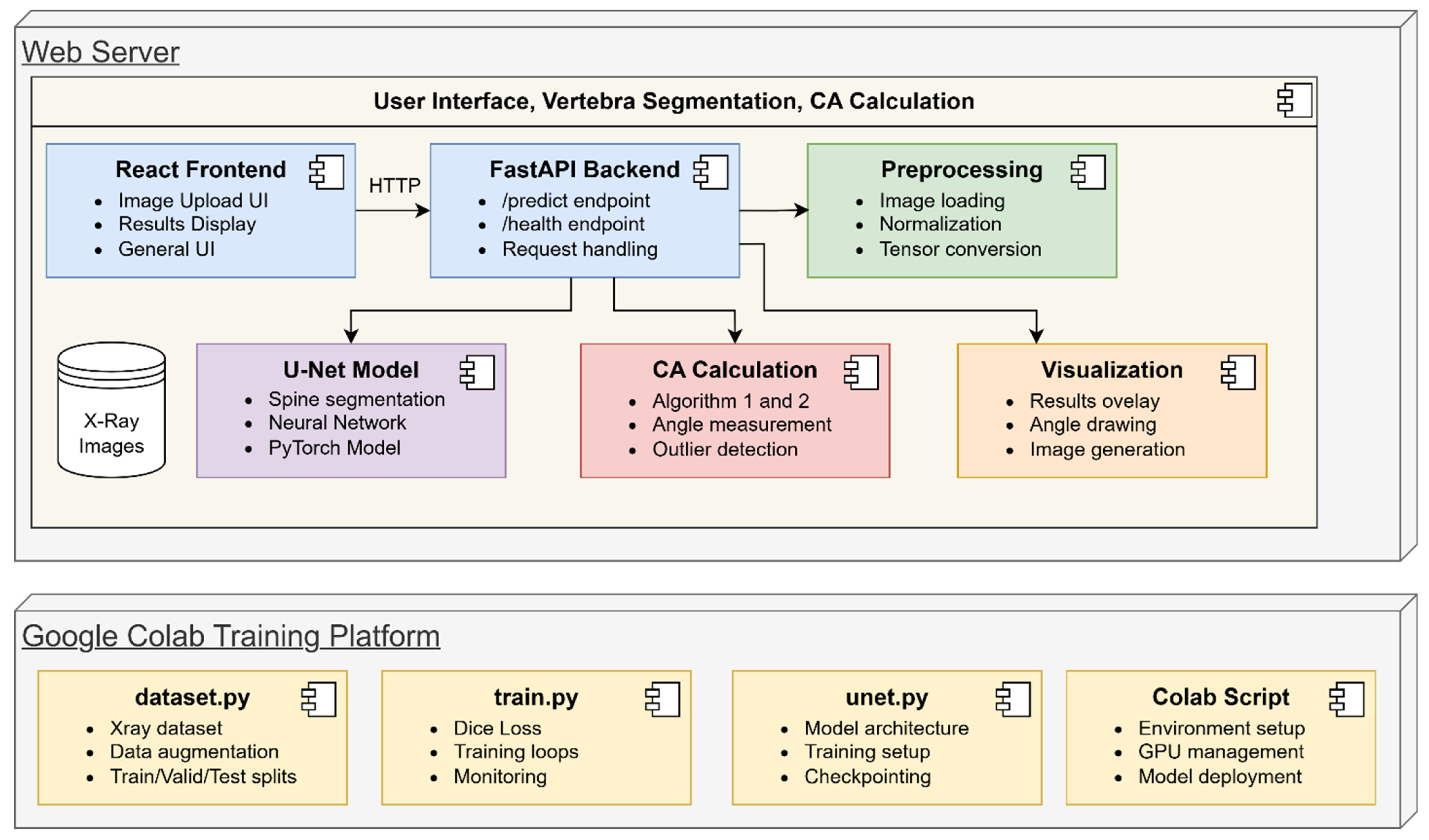
The SpineCheck project is an end-to-end system that performs automatic spine segmentation and CA measurement using spinal X-ray images. The basic components and operation of this system are explained below using the diagram shown in Figure 3. This architectural diagram illustrates the relationships between all components and the data transferred between them at each step.
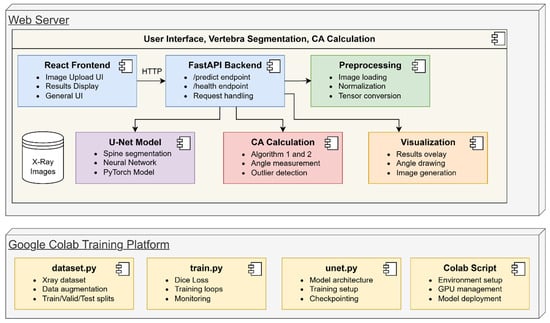
Figure 3.
The project component architecture of SpineCheck.
The user uploads the X-ray image to the React-based interface accessed from the web browser. This interface includes an area that provides the user with a preview of the image, a button that initiates the file upload process, and dynamic sections that display analysis results in graphical or tabular format. When the user clicks the “Start Analysis” button and sends the image to the FastAPI-based backend, the relevant requests are directed to the/predict endpoint via HTTP. The FastAPI backend saves the incoming file to temporary storage and initiates a workflow that triggers the preprocessing pipeline. The /health endpoint is also available to monitor the overall system status, allowing the background application to be run.
After capturing the image, the FastAPI backend first calls the preprocessing module. In this phase, the raw X-ray image is loaded into memory, the pixel values are normalized, and the data is converted to a tensor, making it compatible with PyTorch’s torch.Tensor format. This normalization process is performed to ensure consistent results across images from different sources. The converted X-ray data is then passed to the U-Net Model and CA Calculator components, respectively.
The U-Net Model component contains the neural network architecture that handles the segmentation of the spine. Both low-level spatial details and high-level meaningful features are preserved by its encoder–decoder structure. The model detects the spine regions at the pixel level and produces a binary mask. This mask is used to determine the reference points required for the next step, the CA calculation. The weights are optimized during U-Net training, and segmentation accuracy is maintained at a high level. The Dice score obtained from the test data supports these successful results.
The original image with the created spine mask is transferred to the CA Calculator module by the FastAPI application. In this stage, the positions of the endplates of the most superior and most inferior vertebrae are determined based on the mask. In a two-stage method, the coordinates of the endplate boundaries are first determined, and then the CA is obtained by calculating the slope difference between these lines. A simple outlier detection mechanism is also implemented to prevent incorrect segmentation or measurement deviations that may occur during the calculation process. This increases the reliability of the calculated angle and yields clinically meaningful results.
The CA and other intermediate data obtained with the segmentation mask are forwarded to the Visualization component. This component overlays the segmentation mask onto the original X-ray image as a translucent layer and graphically marks the detected vertebral lines. The calculated angular value is displayed in the user interface via text boxes or indicator graphics. This allows the physician or clinical user to clearly see both the segmentation accuracy and the reference lines used for the measurement.
The entire model development and training process is carried out in a separate layer called the Google Colab Training Platform. This layer is located at the bottom of the component diagram and focuses on the model preparation phase. In the first step of training, X-ray images and corresponding mask labels are loaded via the dataset.py file. This file manages data augmentation and splits the data into training, validation and test subsets.
The train.py file contains the scripts that execute the main training cycle. Here, the appropriate loss functions (e.g., Dice Loss or Combined Loss), the optimization algorithm (Adam), and the learning rate planner (ReduceLROnPlateau) are defined to optimize the model. During training, performance is measured on the validation set at the end of each epoch; the best weights, based on the specified criteria, are stored as checkpoints. This reduces the risk of overfitting and allows monitoring of the outputs at each stage of the training process. Upon completion of training, the best model weights obtained are ready for subsequent inference processes.
The unet.py file, which defines the U-Net architecture, contains the encoder–decoder structure. This file defines the model’s feedforward phase and the skip connections between layers. Thus, spine segmentation is achieved by simultaneously processing both low-level textural features and the high-level meaningful representation. Weights are updated under this architecture during the training phase, while the same structure is used to create the mask during inference.
The Colab Script component automates the training flow using the GPU resources of the Google Colab environment. This component handles the environment setup, loading the necessary libraries, downloading the dataset from Drive, and saving the training outputs and logs to Drive. This allows long training processes to be completed safely without being affected by Colab’s timeouts. Furthermore, GPU memory management and the efficient use of computational resources are regulated by the code blocks defined in these scripts. After the training process is complete, the model weights generated in the Colab layer are transferred to the FastAPI backend. When the user uploads an X-ray image on the frontend, the backend calls the trained U-Net model to perform the segmentation instantly. The CA Calculator module then steps in, processes the mask, and calculates the angle between the spinal lines.
2.3. Data Privacy and Security
To ensure patient data privacy and facilitate secure clinical deployment, the SpineCheck platform utilizes a stateless processing architecture designed to adhere to data minimization principles. The backend system is engineered to process radiographs without permanent storage. Upon upload, images are handled within a volatile temporary environment strictly for the duration of the preprocessing and U-Net inference cycles. Once the CA calculation is complete and the analysis results (JSON and visualization overlays) are transmitted back to the user interface, the source files are immediately and automatically deleted from the server. This “process-and-delete” mechanism ensures that no Protected Health Information (PHI) or longitudinal image history is retained in the database or file system, significantly reducing the risk of data breaches and simplifying compliance with data protection regulations.
3. SpineCheck Dataset, Model Selection, Training and Performance
3.1. X-Ray Images Dataset and Data Preprocessing Before Training
The source dataset was obtained from Roboflow [24] and initially contained 1018 spine radiographs stored as JPG files with heterogeneous resolutions. Each image is paired with a text annotation in the YOLOv5/8-seg (polygon) format—i.e., the TXT file shares the image basename and begins with the class ID followed by normalized polygon vertices as (x, y) pairs. In our corpus, vertebral bodies are annotated as quadrilateral polygons (four vertices; eight numbers). We emphasize that throughout this paper, references to “YOLO format” denote this segmentation-oriented variant rather than the bounding-box format used in object detection.
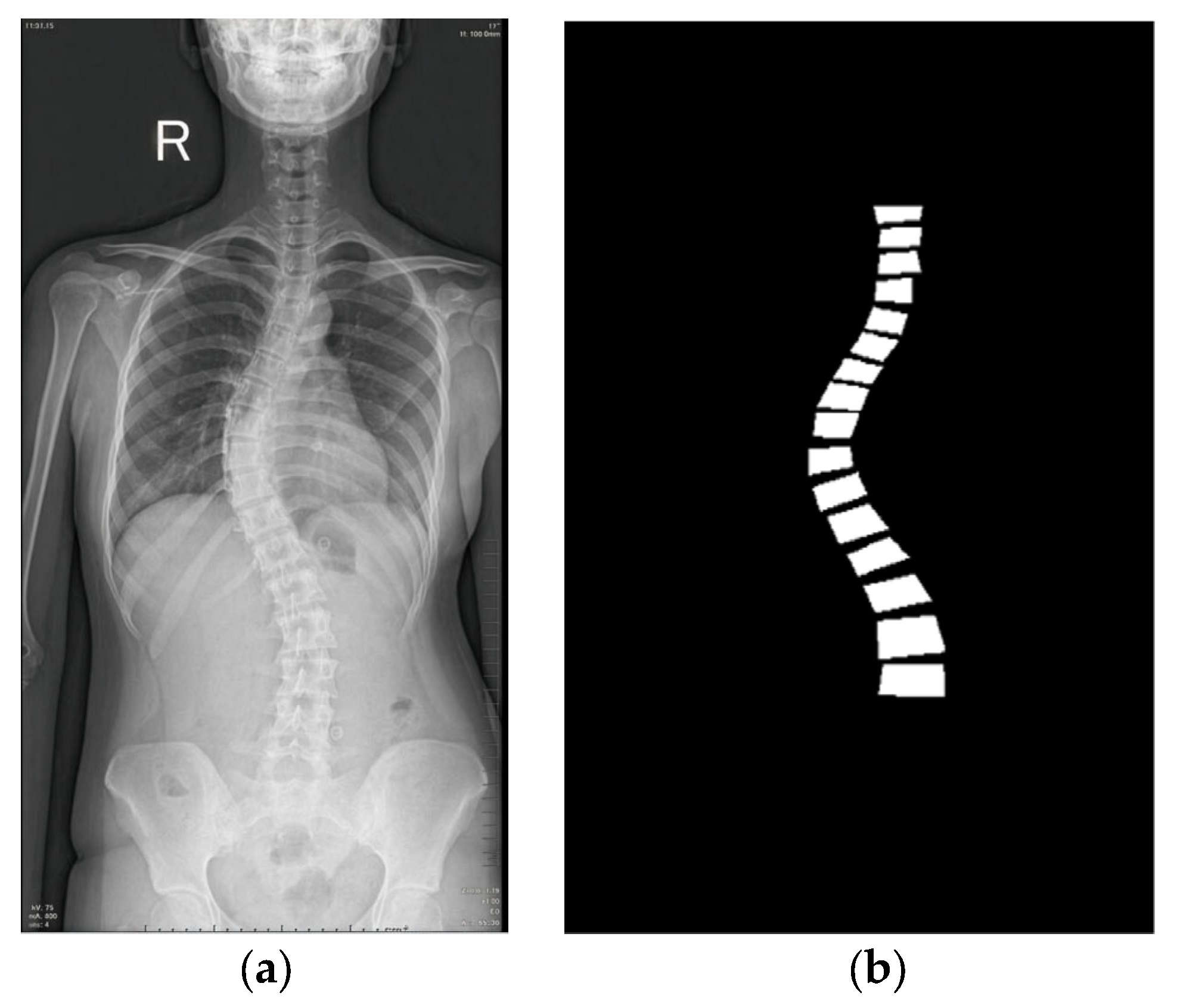
For learning-ready supervision, polygon vertices were denormalized to pixel coordinates and rasterized to produce a single-channel binary mask per image. An example radiograph and its derived mask are shown in Figure 4. Prior to dataset splitting, we conducted a manual quality-control review to remove corrupted/defective files (e.g., unreadable or truncated images) and clearly mislabeled annotations (e.g., polygons not enclosing the intended vertebra). The source contained 1018 radiographs; after this screening, 281 were excluded, yielding a final dataset of 737 radiographs with valid polygon annotations and corresponding masks.
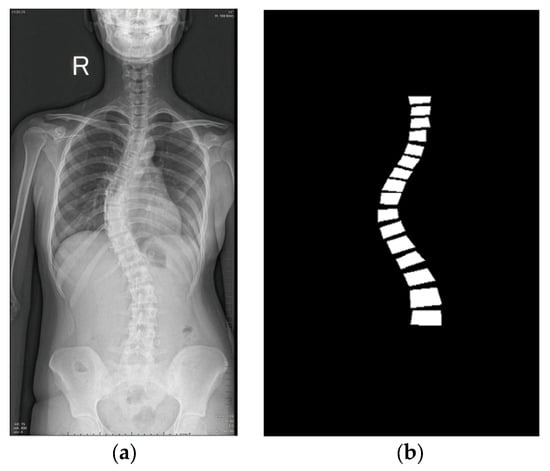
Figure 4.
An example from the dataset [20]: (a) A raw X-ray image showing the patient’s right (R) side; (b) its masked image.
Both the radiographs and masks were resampled to 512 × 512 pixels. To minimize intensity loss, images were resized with area-based interpolation, whereas masks were resized with nearest-neighbor interpolation to preserve class boundaries. In the preprocessing pipeline, each image was read from the file system, converted to RGB, and paired with its single-channel (grayscale) mask, rescaled to 512 × 512 without exception. This standardization keeps all samples at a fixed spatial size so models can be trained directly without layers specific to dynamic resolutions. The filtered dataset was then partitioned at the image level into training/validation/test subsets of 82/10/8% (604/74/59 images). This pipeline—polygon-to-mask conversion followed by standardized splits—ensures supervision is consistent across heterogeneous inputs while avoiding ambiguity between detection-style boxes and the polygonal labels actually provided by the Roboflow dataset.
We then applied a series of sequential preprocessing steps to convert the raw X-ray images and their segmentation masks into “training-ready” tensors. In the first stage of the workflow, the image is read from the file system and converted to RGB format, and the single-channel (grayscale) mask is rescaled to 512 × 512 pixels without exception. Thus, all samples are kept at a fixed spatial size; models can be trained directly with a universal input size without the need for layers specific to dynamic resolutions.
When optional augmentation is enabled, the image and mask pair are randomly flipped horizontally and vertically and rotated in the range of ±30°. In addition, slight brightness and contrast changes are added only to the image side. Since all geometric operations are applied conjugately to both channels, pixel-to-pixel alignment is not disrupted; photometric changes are used only in the image instead of the mask, preserving the consistency of class labels. These steps increase data diversity and play a protective role against overfitting.
After scaling, the content is converted to tensor format; pixel values are scaled into the range [0, 1] and standardized on a per-channel basis using common image classification statistics (μ = [0.485, 0.456, 0.406]—σ = [0.229, 0.224, 0.225]). This normalization balances brightness/contrast variability across different datasets and speeds up the network’s optimization process. Masks are not normalized; instead, they are forced into binary form (0 or 1) by applying a threshold of 0.5 on the single-channel tensor so that the loss functions can directly distinguish between pixels as “background” and “spine.” As a result, each call returns two elements to the system: the normalized 3-channel image tensor and the mask, reduced to the same-sized set of pixel values {0, 1}. This preprocessing pipeline makes the data uniform in both size and statistical distribution, ensuring reliability and reproducibility of model training.
3.2. Model Selection for Vertebra Segmentation
The segmentation framework is based on the U-Net architecture, which is widely adopted in biomedical imaging tasks for its ability to integrate fine structural details with global contextual information. There are many variants of U-Net architecture today, including classical U-Net [4], Attention U-Net [13], UNet++ [14], UNet3++ [15], and nnU-Net [16]. These architectures were used in the project, except UNet++, to evaluate the vertebra segmentation performance of different models for the SpineCheck system and to select the most suitable one. Classic U-Net is included as the first model, as it provides a reliable foundation in medical image segmentation. U-Net-2 is the optimized and more efficient variant of this architecture developed in this study. Attention U-Net is employed to examine the potential benefits of attention mechanisms in capturing fine anatomical details more accurately. nnU-Net is incorporated as a strong candidate due to its consistently high performance reported across various medical imaging studies. UNet3++, with its densely connected multi-scale design, is included for its potential to enhance deep feature integration through improved multi-scale information flow. Together, these architectures enable comprehensive comparison of classical, optimized, attention-enhanced, high-performance, and dense segmentation strategies to determine the optimal model for vertebra segmentation in the SpineCheck system.
Structural and component-level differences among U-Net models lead each architecture to exhibit distinct performance characteristics in image segmentation tasks. As presented in Table 2, these differences are primarily concentrated in critical components such as the design of core convolutional blocks, the use of residual connections, the types of skip-connection structures, the incorporation of attention mechanisms, normalization choices, activation functions, upsampling strategies, base filter sizes, and deep supervision schemes.

Table 2.
Structural and component differences between U-Net variants.
This table systematically presents how the U-Net-based architectures compared in this study differ in terms of their structural components. U-Net, U-Net-2, and Attention U-Net employ the double convolution block (Conv → BN → ReLU × 2) as their core building unit, whereas UNet3++ introduces a more complex and deeper architecture through dense blocks that enable multi-scale feature fusion. In contrast, nnU-Net distinguishes itself from the other architectures through its use of Instance Normalization and a LeakyReLU-based double convolution structure.
Residual connections are present only in UNet3++ and nnU-Net, and the skip-connection mechanisms exhibit notable variation across the architectures. U-Net and U-Net-2 adopt direct concatenation, while Attention U-Net regulates these connections through attention gates. UNet3++ achieves a richer and more comprehensive information flow via multi-scale dense concatenation, whereas nnU-Net employs direct symmetric concatenation, providing an optimized and more balanced variant of the U-Net structure.
The attention mechanism is incorporated solely in the Attention U-Net architecture and is absent in the others. In terms of normalization, most models utilize BatchNorm, whereas nnU-Net’s preference for Instance Normalization distinguishes it as a unique configuration. Regarding activation functions, four models employ ReLU, while nnU-Net adopts LeakyReLU, reflecting a different activation strategy.
In relation to upsampling methods, U-Net-2 and Attention U-Net use only the ConvTranspose2D (stride = 2) operation, whereas UNet3++ applies ConvTranspose2D in combination with its multi-scale skip-fusion mechanism. In contrast, both U-Net and nnU-Net also support bilinear interpolation, which can provide more memory-efficient or artifact-reduced upsampling depending on the task. The number of base filters in the first layer also varies across architectures: U-Net, U-Net-2, and Attention U-Net begin with 64 filters, UNet3++ with 56, and nnU-Net with 48 filters.
Finally, deep supervision is present only in the UNet3++ and nnU-Net models. This mechanism contributes to more effective optimization of multi-scale feature maps, thereby enhancing the overall learning process of these architectures.
Table 3 summarizes the system and hardware specifications used for the training and evaluation of U-Net models. The table can be examined under five main sections: operating system, hardware components, training parameters, input and batch size, and base filter numbers. In the training parameters section, the number of epochs and learning rate are indicated, while the input size and batch size reflect the model’s data processing capacity. Base filter numbers represent the representational power and learning capacity of the different models.

Table 3.
System and hardware specifications used for training and evaluation of U-Net models.
The models were developed and optimized in the Google Colab environment. The operating system used was Linux 6.6.105+-x86 64-with-glibc2.35, providing a stable training environment compatible with CUDA-based GPU support. The training device was CUDA-enabled, with an NVIDIA A100-SXM4-80GB GPU serving as the graphics processing unit; its 79.32 GB of VRAM allowed high-speed processing of large datasets. On the CPU side, a 12-core processor and 167 GB of RAM played a critical role in accelerating data transfer to the GPU and efficiently managing parallel operations. The input size of (3, 512, 512) ensured that the model received data with appropriate color channels and resolution, while a batch size of 8 balanced memory usage and training efficiency. Training parameters included 100 epochs and a learning rate of 0.0001, enabling the model to be optimized steadily over sufficient training cycles and achieve high segmentation accuracy. This combination of hardware and parameters was selected to reduce training time while enhancing the model’s performance and stability.
Table 4 summarizes the performance comparison of different U-Net models. The table presents key performance metrics, including the number of parameters, best validation loss, Dice score, training time, and inference time. These metrics allow for the comparison of the models in terms of segmentation accuracy, computational efficiency, and speed.

Table 4.
Performance comparison of U-Net-based models for vertebra segmentation.
The primary differences between the models clearly emerge through their architectural complexity as well as their training and inference efficiency. Since the parameter counts are very close to each other, the performance variations largely stem from architectural innovations and additional mechanisms integrated into the models.
When examining the Dice scores, nnU-Net, UNet3++ and Attention U-Net stand out as the best-performing group in terms of segmentation accuracy. This superior performance is attributed to nnU-Net’s well-optimized configurations proven across various medical imaging tasks, UNet3++’s multi-scale dense connections that enhance feature representation, and Attention U-Net’s attention mechanisms that more effectively highlight critical regions. U-Net-2, although slightly behind these top three models in accuracy, achieves both the shortest training time and the fastest inference speed. These characteristics make it the most suitable option for scenarios where computational efficiency is a priority. In contrast, the U-Net, due to its simpler architecture, yields the lowest Dice score and requires the longest training time. Overall, the table shows that incorporating multi-scale structures and attention mechanisms significantly improves segmentation accuracy, whereas streamlined and optimized models provide notable benefits in terms of speed and computational efficiency.
Table 5 compares the peak GPU memory usage of U-Net models during training and inference. Training memory refers to the GPU resources required during backpropagation and parameter updates, whereas inference memory reflects the lower GPU usage during the forward pass. The parameter values (M) in the table provide a quantitative indication of each model’s architectural complexity and computational load.

Table 5.
Peak GPU memory usage during training and inference.
As seen in the table, U-Net has the lowest number of parameters and also uses the least training and inference memory. Its simple and streamlined architecture makes it one of the most efficient models. nnU-Net, although it has slightly more parameters than U-Net, maintains low memory usage because it preserves the core U-Net structure while applying optimized configuration rules. Attention U-Net contains more parameters than the previous models, and its attention mechanisms require additional computation, which increases GPU memory consumption during both training and inference. U-Net-2 has a similar parameter count to nnU-Net but requires more memory due to its architectural modifications and additional processing blocks. UNet3++ has the highest number of parameters among all models and also exhibits the highest memory consumption. This is primarily due to its densely connected skip pathways and deeper multi-scale structure, which significantly increase computational load and memory requirements.
Table 6 summarizes the functional and operational evaluation of U-Net family models. The table presents key functional metrics, including core segmentation capability, best Dice score, lowest validation loss, model complexity, inference speed, adaptability/automation, training difficulty, and deployment flexibility. These metrics allow for the comparison of the models in terms of accuracy, efficiency, and ease of use, guiding the selection of the most suitable U-Net model.

Table 6.
Functional and operational evaluation of five U-Net models.
As shown in the table, U-Net and U-Net-2 stand out with their low model complexity, ease of training, and very fast inference speeds. Because these models maintain a simple encoder–decoder structure without additional modules, they have low computational cost and run efficiently on both GPU and CPU. This minimalist architecture continues to be a strong baseline for medical image segmentation tasks.
Attention U-Net distinguishes itself by achieving a lower validation loss. This improvement stems from the attention gates, which help the model focus more selectively on relevant anatomical structures while suppressing background noise. However, these additional attention mechanisms increase architectural complexity and make the training process slightly more challenging compared to standard U-Net.
UNet3++ and nnU-Net incorporate advanced components such as higher architectural complexity, deep supervision, and Instance Normalization. While these models are more modern and capable, they require greater computational resources, have longer training times, and do not match the classical U-Net in inference speed. Their primary goal is to push segmentation performance further; indeed, nnU-Net achieved the highest Dice score in our experiments (0.8448). However, this performance comes at the cost of increased design and optimization complexity.
Overall, the results indicate that although modern U-Net variants can achieve slightly higher Dice scores, these improvements come at the cost of increased architectural complexity and a more challenging optimization process. In contrast, U-Net (classical) exhibits a more predictable, stable, and error-tolerant learning behavior due to its simple encoder–decoder structure, low parameter count, and minimal need for auxiliary modules. In the following section, the role and performance of the U-Net architecture are examined in detail, including its training process, hyperparameter choices, and its contributions to CA calculation.
3.3. U-Net Architecture
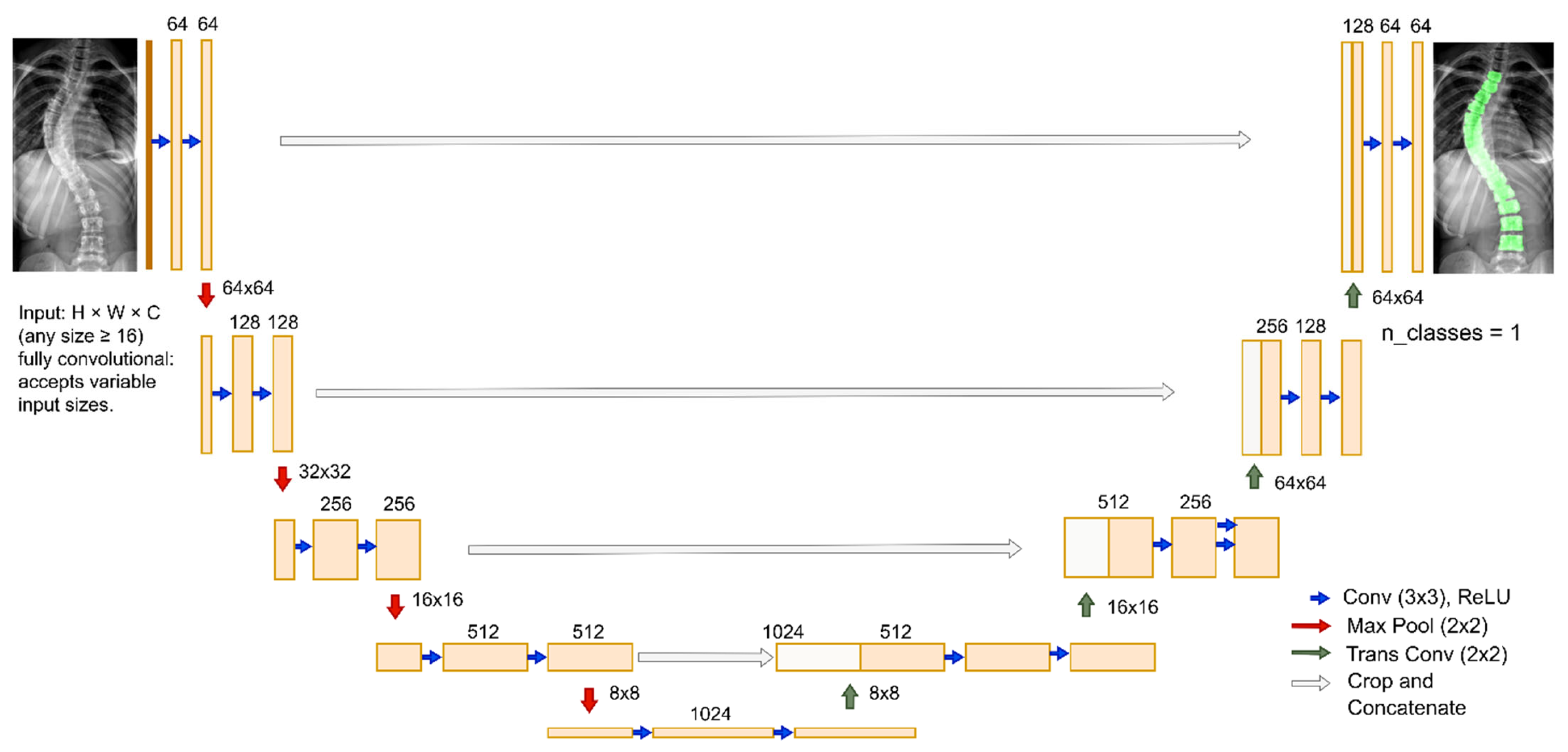
As illustrated in Figure 5, the network follows an encoder–decoder paradigm with skip connections that enable efficient pixel-wise classification even in low-contrast radiographs. The U-Net architecture is modular, as depicted in Figure 5, consisting of DoubleConv, Down, Up, and OutConv components.
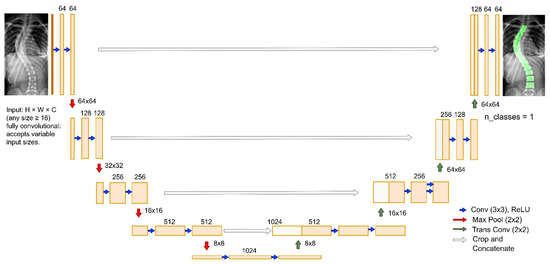
Figure 5.
U-Net architecture used in SpineCheck Light brown rectangles are convolution output feature maps with learned representations. White rectangles are skip-connection feature maps restoring fine spatial detail.
The encoder is composed of repeated DoubleConv modules (two sequential Conv–BatchNorm–ReLU layers), which progressively extract hierarchical representations. Each Down block combines max-pooling for spatial reduction with increased channel depth, allowing the network to aggregate broader contextual information. Conversely, the decoder employs Up blocks, where spatial resolution is restored either by bilinear interpolation or by transposed convolution. Bilinear upsampling provides memory efficiency and minimizes artifacts, while transposed convolution increases representational capacity at higher computational cost. Skip connections between encoder and decoder stages preserve fine-grained anatomical features such as vertebral edges, ensuring precise boundary delineation. The final OutConv layer applies a 1 × 1 convolution to generate logits for the target classes, followed by sigmoid or softmax activation to yield probability maps.
The architecture is parameterized by two primary hyperparameters. First, the features parameter defines the number of filters in the initial layer (set to 64 in this work), doubling at each encoder stage until the bottleneck is reached (512 channels with bilinear interpolation or 1024 with transposed convolution). This value governs the trade-off between representational power and computational cost. Second, the bilinear flag selects the upsampling operator: when enabled (bilinear = True), the decoder uses bilinear interpolation with scale_factor = 2 (and align_corners = True); when disabled, it uses a 2 × 2 transposed convolution (stride 2). Standard settings are retained throughout: convolutions use 3 × 3 kernels with padding 1 to preserve spatial dimensions, and downsampling is performed by 2 × 2 max-pooling with stride 2 in each Down block.
This modularity ensures flexibility for adaptation to different segmentation tasks. Furthermore, gradient checkpointing can be activated during training to alleviate GPU memory usage by recomputing intermediate activations during backpropagation, which is particularly beneficial for high-resolution X-ray images. This approach enables the model to be trained within hardware limitations without sacrificing accuracy.
This U-Net configuration provides a balanced compromise between accuracy, computational efficiency, and memory consumption. Its encoder effectively captures both localized and global spinal features, while the decoder restores high-resolution details with structural consistency. This architecture is therefore well suited for robust segmentation of spinal structures in radiographic images, providing accurate binary masks that establish a reliable foundation for subsequent CA estimation as well as the training and optimization process.
3.4. Training Process and Hyperparameters
Building upon this architecture, the U-Net model was trained on high-resolution spine radiographs. The training process and selected hyperparameters were carefully designed to ensure both computational efficiency and segmentation accuracy. Model training was conducted using the XrayDataset of image–mask pairs. The training, validation, and test subsets, with batched access, are handled by DataLoader objects. Random shuffling was applied during training, while validation followed a sequential order to ensure consistent evaluation.
The network was implemented in PyTorch with n_channels = 3 (RGB inputs) and n_classes = 1 (binary segmentation output). Training employed a combined loss function, defined as an equal-weighted sum of Binary Cross-Entropy and Dice Loss, balancing pixel-level classification with regional overlap accuracy. To stabilize optimization, a scheduler reduced the learning rate by half when validation loss plateaued. The smoothing factor in Dice Loss (1.0) prevented division errors and improved sensitivity to thin structures such as spinal edges.
Training was initially conducted for all models using a batch size of 8 and 100 epochs. However, the learning curves showed that the U-Net model fully converged between epochs 40 and 50, with no meaningful improvements beyond this point. Therefore, performance evaluations were based on the 50-epoch results, where the model produced its most stable outputs.
Using a batch size of 8 and 50 epochs provided an effective balance between gradient diversity and memory efficiency. The validation split (10%) enabled intermediate performance monitoring and learning rate adjustments. Performance was tracked using Dice score (0–1 scale, with higher values indicating better overlap) and combined loss (approaching zero for improved accuracy). These metrics guided model convergence and informed potential early stopping. Finally, the bilinear upsampling option was generally preferred to reduce computational overhead and artifacts while still preserving anatomical precision. This configuration enabled stable training on high-resolution spine radiographs and provided reliable segmentation masks for subsequent CA estimation.
3.5. Model Performance and Analysis
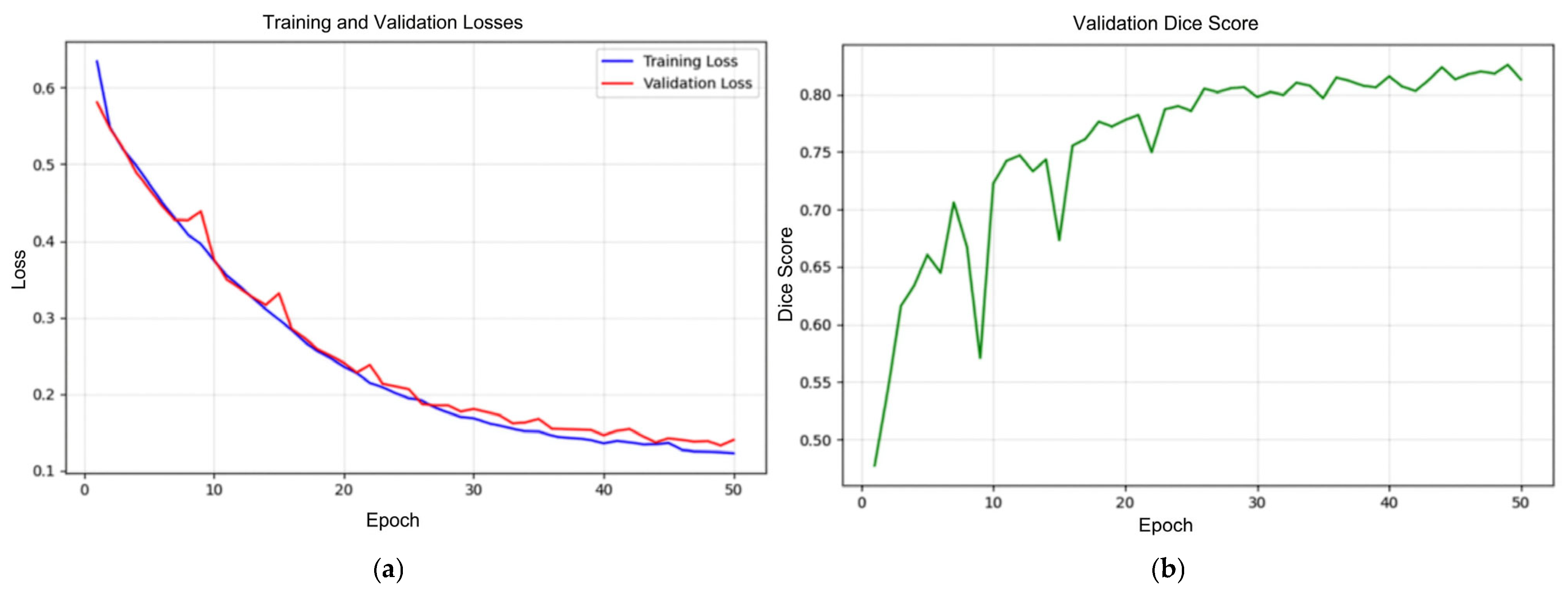
The training and validation losses, as well as the validation Dice score curves, are shown in Figure 6. Both training and validation losses consistently decrease throughout the 50 epochs, and the two curves remain closely aligned. This close alignment indicates that the model successfully learns from the training dataset while maintaining strong generalization to unseen validation samples, suggesting minimal signs of overfitting. The smooth downward trend of both curves reflects stable optimization and steady convergence.
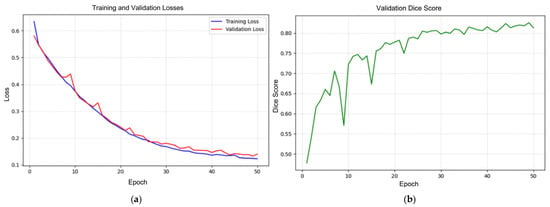
Figure 6.
Training loss (blue), validation loss (red) (a), and validation Dice score (green) (b) curves.
The validation Dice score curve demonstrates a clear upward trajectory, beginning around 0.48 and rising steadily to approximately 0.84 by the end of training. This continuous improvement indicates that the model enhances its segmentation capability over time, effectively learning vertebral boundaries and shapes. Minor fluctuations observed during training are expected due to variations within the dataset and do not disrupt the overall positive progression.
The loss reduction and Dice score improvement confirm that the U-Net model achieved effective segmentation performance, with strong convergence and generalization properties across the training process. Notably, a Dice coefficient above 0.80 is generally considered a strong performance benchmark in medical image segmentation tasks, indicating that the trained model reached a level of accuracy suitable for practical use.
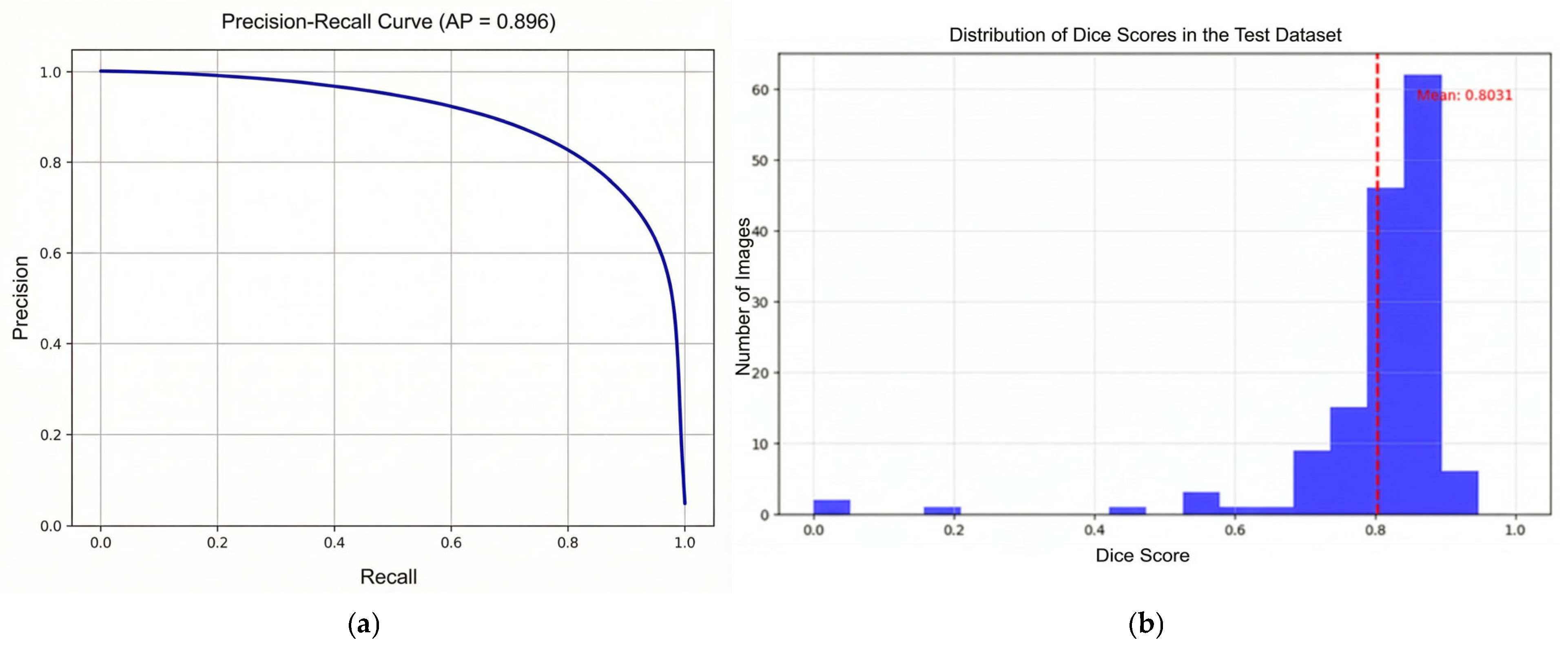
The precision–recall curve shown in Figure 7a demonstrates that the model maintains high precision across a wide range of recall values. The area under the precision–recall curve (average precision, AP = 0.896) indicates strong performance, suggesting that the model can reliably detect vertebral regions with few false positives, even as recall increases.
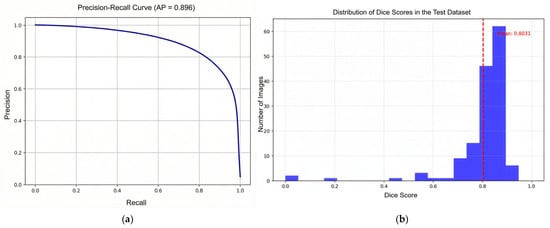
Figure 7.
Precision–Recall curve (a) and distribution of Dice scores on the test dataset curves (b). The dashed red line in the Dice score distribution plot (b) denotes the mean Dice value (0.8031) computed over all test images.
The histogram given in Figure 7b shows the distribution of Dice similarity coefficients for individual test samples. The majority of scores cluster between 0.75 and 0.90, with a mean Dice score of approximately 0.8031 (red dashed line). This confirms that the model performs consistently well across most test images, with only a few outliers exhibiting lower segmentation accuracy. The high average precision and strong mean Dice score demonstrate robust segmentation on the test set, meeting accuracy levels generally regarded as strong benchmarks in medical image segmentation tasks.
For completeness, the AP is computed pixel-wise by flattening the predicted probability map and ground-truth mask across the test set to form a binary classification problem (positive = vertebra pixel), then integrating the precision–recall curve (area-under-PR). This AP is not object-detection mAP and complements Dice by integrating precision–recall trade-offs under class imbalance typical of vertebra vs. background pixels.
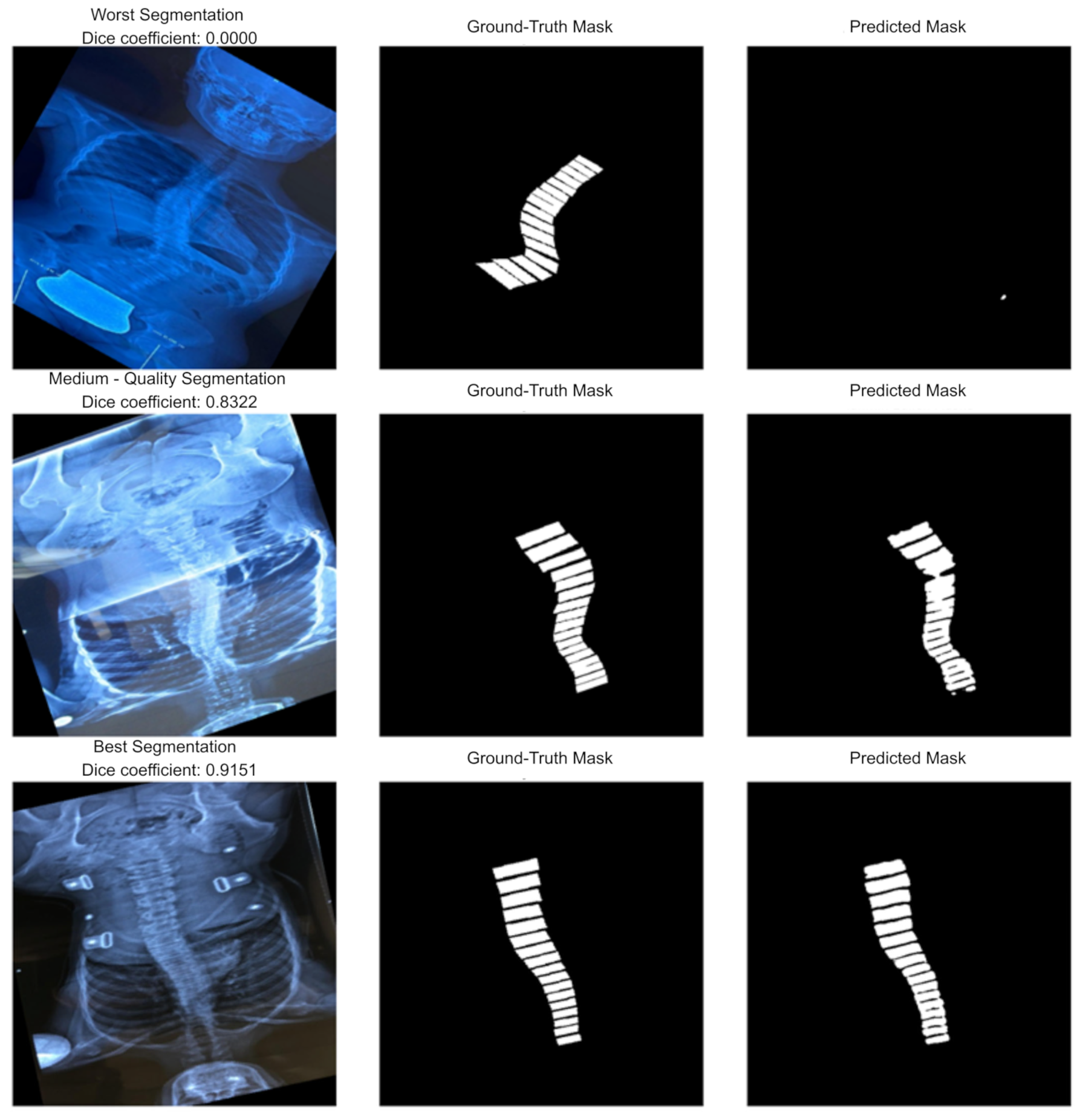
Figure 8 illustrates qualitative segmentation results of the U-Net model during testing, highlighting 3 different performance levels:
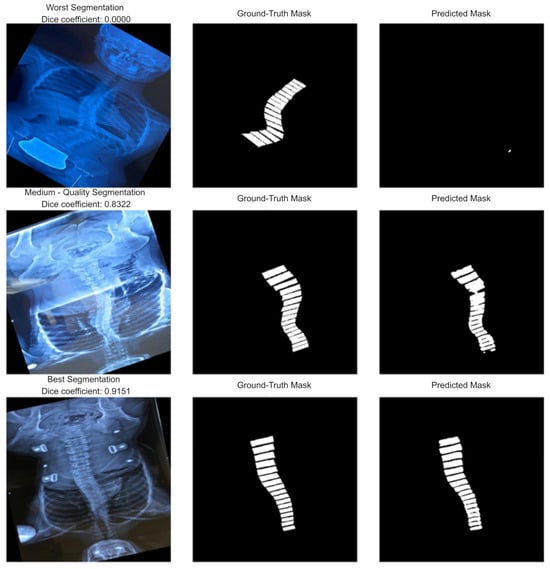
Figure 8.
Examples of worst (top row)/medium (middle row)/best (bottom row) segmentations.
Worst Segmentation (top row, Dice = 0.0000): The predicted mask fails to capture the vertebral structure entirely, resulting in no meaningful overlap with the ground truth. This case represents a complete failure of the segmentation process, possibly due to poor image quality, extreme anatomical variation, or noise.
Medium-Quality Segmentation (middle row, Dice = 0.8322): The predicted mask aligns reasonably well with the ground truth but shows minor inaccuracies in boundary definition and continuity. While the vertebral column is mostly identified, some misalignments and partial omissions reduce accuracy.
Best Segmentation (bottom row, Dice = 0.9151): The predicted mask closely matches the ground truth with minimal deviation. The vertebral boundaries are well preserved, and segmentation quality is high, reflecting strong model performance on this sample.
In addition to quantitative evaluations such as Dice score and precision–recall analysis, qualitative examples illustrate the performance variability of the U-Net model. Representative cases from the test set, corresponding to the worst, medium, and best segmentation outcomes, demonstrate how the model performs under different conditions. These visual results highlight both the strengths of the model, where the predicted masks closely align with the ground truth, and its limitations, where segmentation accuracy deteriorates due to challenging image characteristics. Such qualitative analysis provides an essential complement to numerical metrics, offering deeper insights into the practical reliability and interpretability of the model in real-world applications.
4. Calculation of the CA and Validation
4.1. Automatic Scoliosis Diagnosis and Measurement Algorithm
This section details the automated pipeline used to derive the CA and classify scoliosis severity from the segmentation masks generated by the deep learning model. The complete algorithmic procedure is formally defined in Algorithm 1, which outlines the transformation from raw pixel data to a clinical diagnosis.
| Algorithm 1. Pseudocode for automatic scoliosis diagnosis and measurement |
 |
As illustrated in Step 1 and Step 2 (lines 2–10), the process begins by normalizing the input radiograph and generating a probability map via the U-Net model. To ensure robustness against false positives—such as artifacts resembling spinal structures—the algorithm calculates a vertical projection profile and applies Gaussian smoothing to locate the primary spinal centerline. This allows for the definition of a dynamic Region of Interest (ROI), excluding non-spinal lateral regions. Subsequently, to address potential segmentation mergers where vertebrae appear connected, Step 3 (lines 11–16) applies a distance-transform-based watershed algorithm to separate adjacent vertebral bodies into distinct geometric entities.
Once the vertebrae are isolated and filtered by geometric properties (lines 19–23), the system applies a geometric modeling approach to approximate the endplates. Adopting the geometry-first methodology proposed by Tan et al. [23], Step 5 (lines 26–37) fits a minimum enclosing rectangle (rotated bounding box) to the contour of each detected vertebra. The superior and inferior edges of this rectangle serve as proxies for the vertebral endplates. To mitigate the impact of jagged segmentation boundaries on angle calculation, the system samples equidistant points along these edges and fits a line using Theil–Sen regression (lines 33–34). This robust estimator is less sensitive to outliers than standard least-squares fitting, yielding a precise slope for both the superior and inferior endplates of every vertebra, even in the presence of contour irregularities.
To further refine the measurements and enhance clinical robustness, Step 6 (lines 38–44) enforces anatomical consistency through outlier suppression. A second-degree polynomial spline is fitted through the centroids of all detected vertebrae to model the natural spinal curvature. As detailed in line 43, vertebrae whose centroids deviate significantly from this spline—specifically, those exceeding a data-driven distance threshold—are flagged as outliers and excluded from the angle calculation. This step prevents spurious segmentations or misidentified artifacts from distorting the final CA.
The final CA is determined in Step 7 (lines 45–56) by iterating through all valid pairs of vertebrae. The algorithm calculates the acute angle difference between the superior endplate of a cranial vertebra and the inferior endplate of a caudal vertebra. To replicate clinical practices, the search is constrained to pairs separated by at least two and at most ten vertebrae (line 49). The pair yielding the maximum angle is selected as the representative CA. Finally, this value is mapped to a severity category using the classification thresholds defined in Table 1, and the diagnosis is returned alongside the indices of the upper and lower end vertebrae (UEV, LEV).
4.2. The Validation of Cobb Angle Measurements
The clinical utility of the SpineCheck platform depends not only on segmentation accuracy but also on the final precision of the CA measurement. To validate the system, we assessed the end-to-end performance of the geometric algorithm using the Spinal-AI2024 dataset [25,26]. This CA measurement validation phase utilized a large-scale subset of 20,000 X-ray images to ensure statistical significance. The automated pipeline described in Section 4.1 was applied to the segmentation outputs of five different deep learning models—U-Net, U-Net-2 (our improved version), Attention U-Net, nnU-Net, and UNet3Plus—and the resulting angles were compared against the CAs provided as reference annotations in the Spinal-AI2024 dataset. For each model, we computed the mean absolute error (MAE), the Pearson correlation coefficient r, and the intraclass correlation coefficient ICC(A,1) between automatic measurements and these dataset-provided reference CAs.
Because a single global error value on the full dataset can be heavily skewed by cases where segmentation failed completely (resulting in gross outliers), we performed a stratified analysis to evaluate the geometric precision of the models under increasingly realistic conditions. First, we analyzed a high-precision “clean” subset of 828 images with an absolute error below 5°, representing cases with reliable segmentations and near-ideal behavior of the geometric estimator. The agreement metrics for this clean subset are reported in Table 7. All models achieved very low MAE values of approximately 2° and ICC(A,1) values around 0.95–0.96, indicating excellent agreement with the dataset reference angles. Within this subset, U-Net and Attention U-Net attained the smallest errors and the highest agreement statistics, confirming that, in the absence of major segmentation artifacts, the geometric CA algorithm can reproduce the reference measurements with high fidelity.

Table 7.
Agreement metrics for a clean subset with absolute Cobb-angle error below 5°.
To bridge the gap between this idealized scenario and the full cohort, we then considered a broader subset that included all cases with an absolute error below 10°, resulting in 4192 images. This intermediate analysis captures the majority of clinically acceptable predictions while still excluding the most severe failure cases. The corresponding metrics are summarized in Table 8. In this subset, the MAE values increased to roughly 4–5°, and ICC(A,1) decreased slightly but remained in the range typically interpreted as good agreement. Again, U-Net retained a small but consistent advantage in MAE over the more complex architectures, suggesting that its smoother vertebral boundaries are slightly more favorable for stable endplate regression than the sharper but noisier segmentations produced by deeper variants such as nnU-Net and UNet3Plus.

Table 8.
Agreement metrics for a clean subset with absolute Cobb-angle error below 10°.
Finally, Table 9 reports global agreement metrics on the entire 20,000-image evaluation dataset, including all radiographs irrespective of segmentation quality. As expected, the error increases markedly when all failure cases are included: MAE values rise to roughly 10–15°, and both Pearson r and ICC(A,1) fall to the “fair” agreement range. This degradation is mainly driven by a relatively small number of images with poor or incomplete segmentations, where the pipeline substantially underestimates the CA or fails to detect curvature at all. Reporting the 5° and 10° subsets alongside the full-dataset results is therefore essential to disentangle the intrinsic accuracy of the geometric Cobb estimator from the impact of segmentation robustness and to clarify the conditions under which the system attains clinically acceptable performance.

Table 9.
Global agreement metrics for automatic CA estimations on the full dataset.
From an implementation perspective, it is important to note that SpineCheck explicitly returns a CA of 0° in cases where the segmentation module does not detect a sufficient number of vertebrae to support a reliable curvature estimate. These zero-valued outputs are also displayed as such in the user interface, making failure cases transparent to clinicians. However, when these values are compared against the non-zero reference CAs provided in the Spinal-AI2024 dataset, they exert a disproportionate influence on the global agreement statistics, systematically lowering Pearson r and ICC(A,1) on the full 20,000-image cohort. Many of these extreme discrepancies originate from low-quality or artifact-heavy radiographs in the public dataset rather than from the Cobb estimator itself. For this reason, the stratified MAE < 5° and MAE < 10° analyses are essential: they isolate the intrinsic geometric accuracy of the Cobb-angle algorithm in cases with reliable segmentations and provide a fairer basis for comparing the five segmentation backbones.
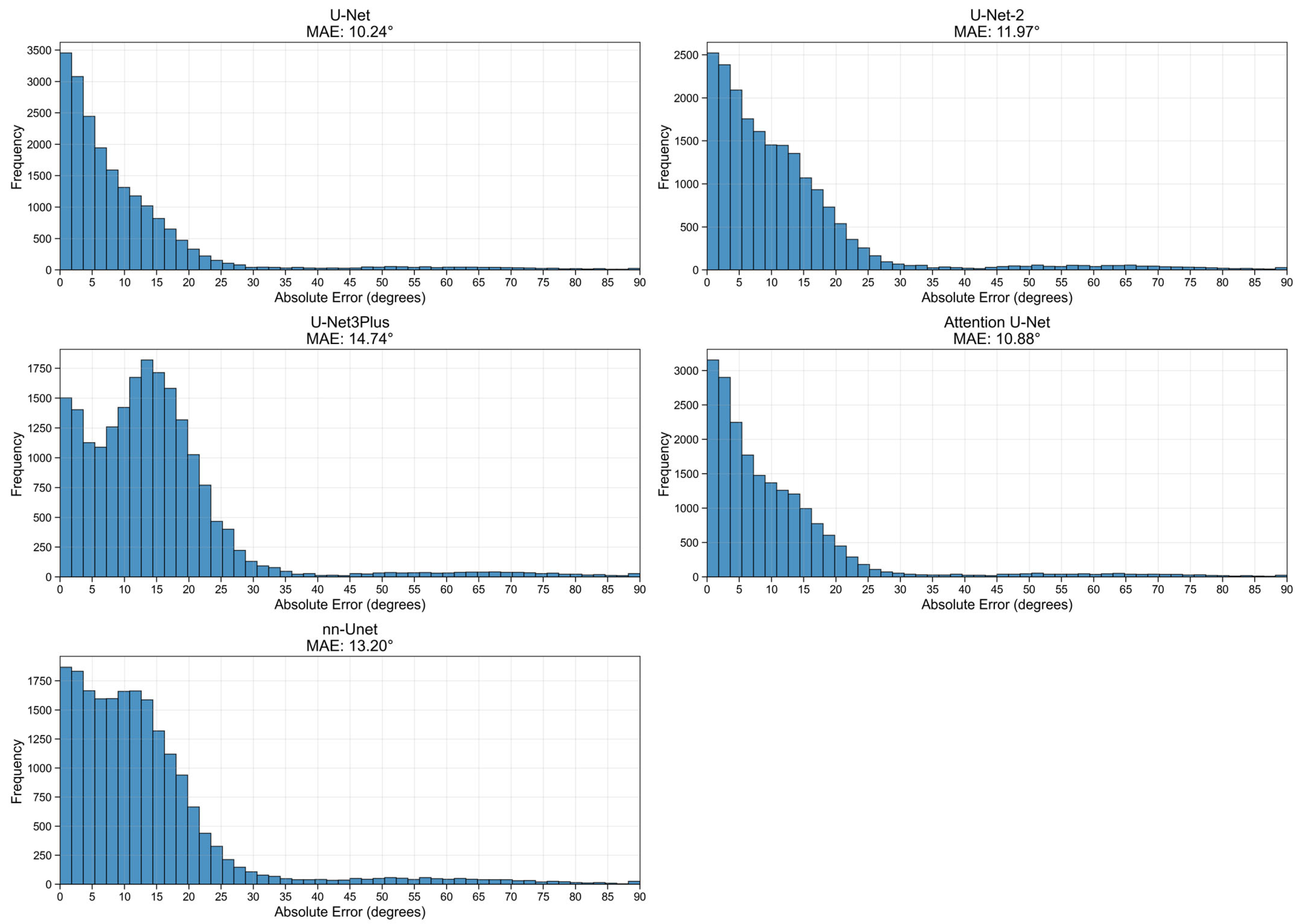
Figure 9 presents the histograms of the absolute errors for the five evaluated models on the full 20,000-image set. The distributions are right-skewed, indicating that for the majority of cases, the automated measurements remain close to the reference angles, with the highest bar heights occurring at small error values. The panel titles report the global MAE for each model, providing a compact summary of their overall performance. While the U-Net and Attention U-Net variants show a tighter concentration of errors near zero, the longer tails in the distributions for UNet3Plus and nnU-Net reflect a higher incidence of outliers and failure cases with very large errors. These findings support the choice of the optimized U-Net backbone for the final SpineCheck deployment, as it offers the most reliable balance between segmentation performance and downstream geometric accuracy.
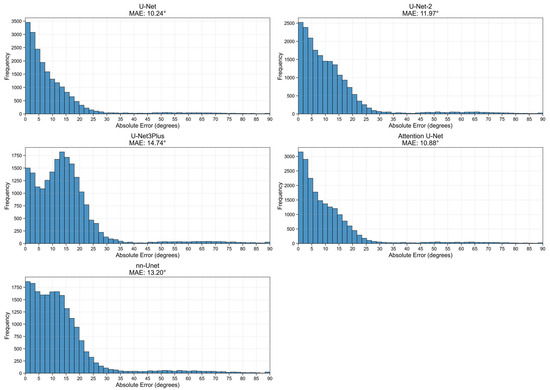
Figure 9.
Distribution of MAE values across five models evaluated on 20,000 images.
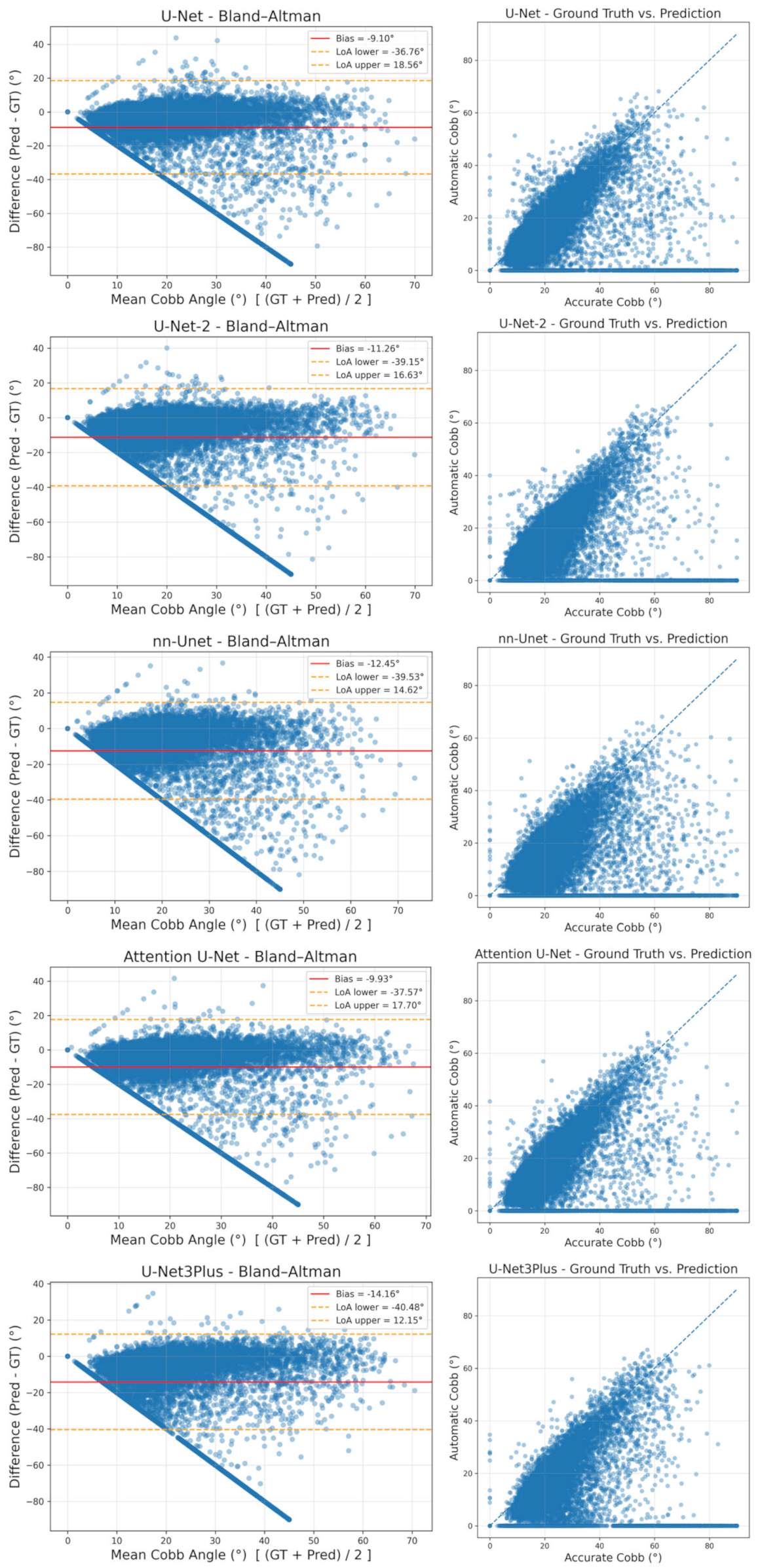
In addition to these distributional analyses, we conducted a visual agreement analysis using Bland–Altman and scatter plots for all five models, and the results are shown in Figure 10. In the Bland–Altman diagrams, the vertical axis represents the difference between automatic and reference measurements (Pred−GT), the horizontal axis shows their mean, the solid line denotes the bias (mean signed difference), and the dashed lines indicate the 95% limits of agreement defined as bias ± 1.96·SD. Across models, the bias is consistently negative, reflecting a tendency of the pipeline to underestimate the CA, while the wide limits of agreement on the full dataset are largely attributable to the same outlier cases that dominate the global MAE. The downward “fan” of points towards large negative differences corresponds to images in which the system returns 0° because no reliable curvature could be detected, whereas the dataset annotations indicate substantial scoliosis. When these failure cases are disregarded, most points cluster near the zero-difference line, and the scatter plots—where the x-axis is labeled “Accurate Cobb (°)” and represents the dataset reference angle—show dense clouds of measurements around the identity line, corroborating the strong agreement observed in the stratified analyses of Table 7 and Table 8.
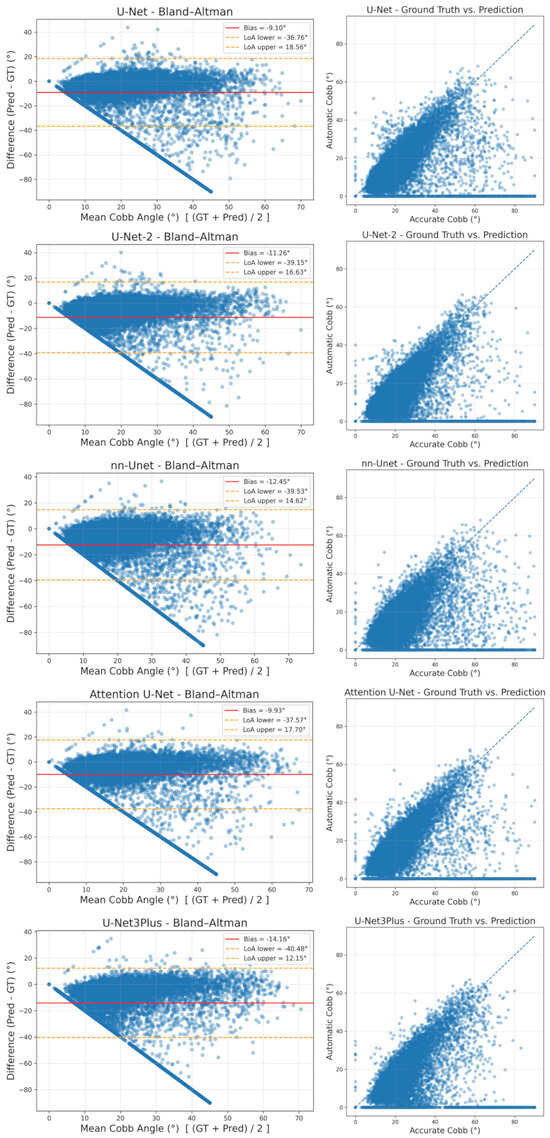
Figure 10.
Agreement analysis of automatic vs. expert CAs across five U-Net variants.
In the visual agreement plots shown in Figure 10, graphical elements follow a consistent convention. In each Bland–Altman panel (left column), blue circles correspond to individual radiographs, the solid red horizontal line summarizes the mean difference (bias) between automatic and accurate Cobb estimates, and the two yellow dashed lines delineate the 95% limits of agreement, indicating the range within which most differences are expected to fall. In the scatter plots (right column), blue circles represent paired automatic versus accurate Cobb values for each image, while the blue dashed diagonal denotes the line of identity, against which deviations directly reflect disagreement between the two measurements.
5. SpinCheck User Interface
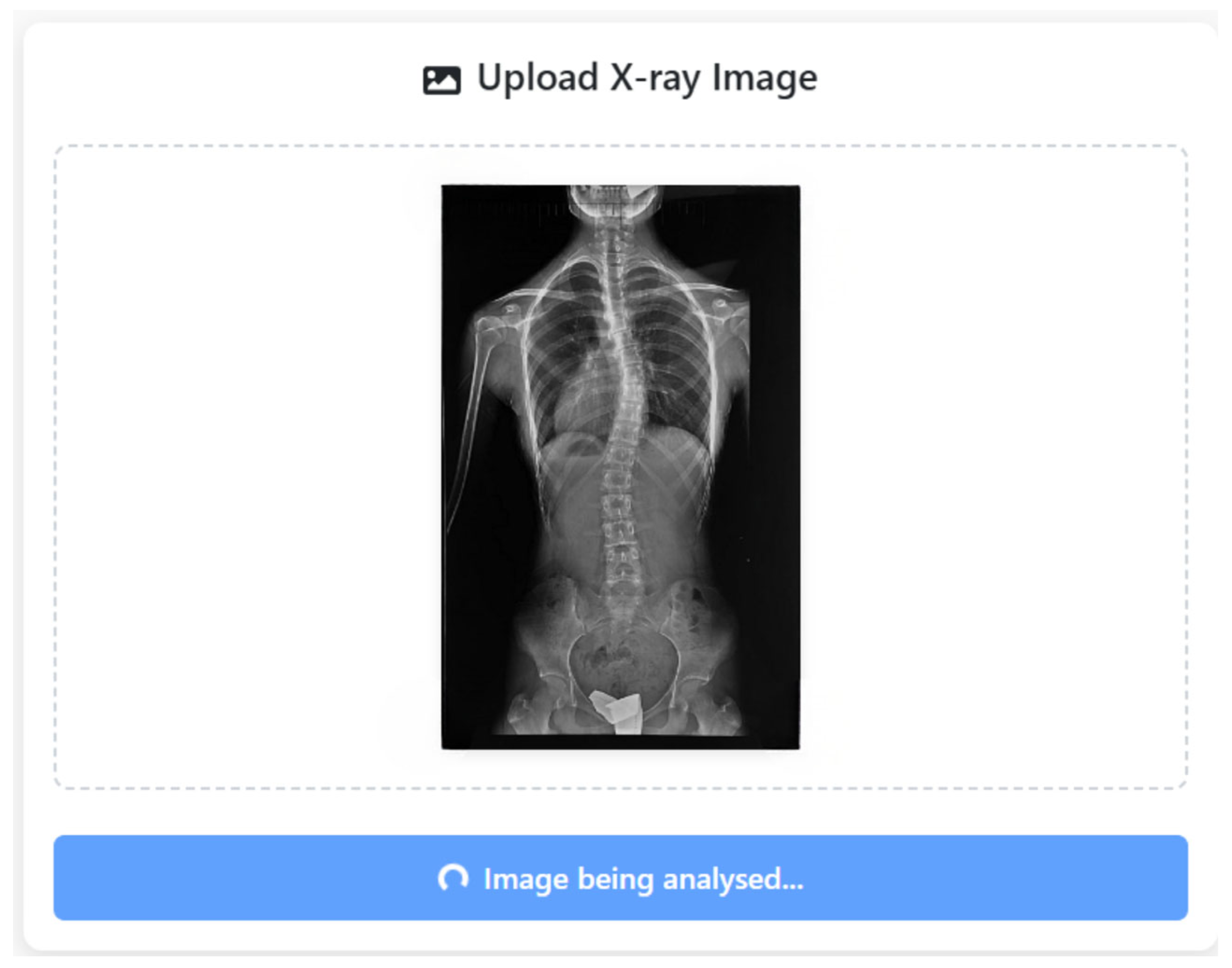
During the analysis workflow, once the X-ray is submitted to the SpineCheck user interface, the system performs vertebra segmentation and geometric curve modeling automatically, as shown in Figure 11. The CA is then determined by evaluating slope differences between all valid vertebral pairs. The maximum angular separation is selected as the defining curvature, calculated via trigonometric transformation and rounded to the nearest 0.5°. The final value is compared against the 10° diagnostic threshold for scoliosis.
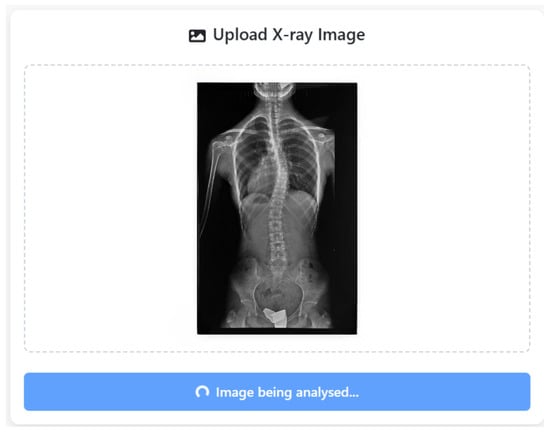
Figure 11.
SpineCheck user interface during automated analysis of an uploaded X-ray image.
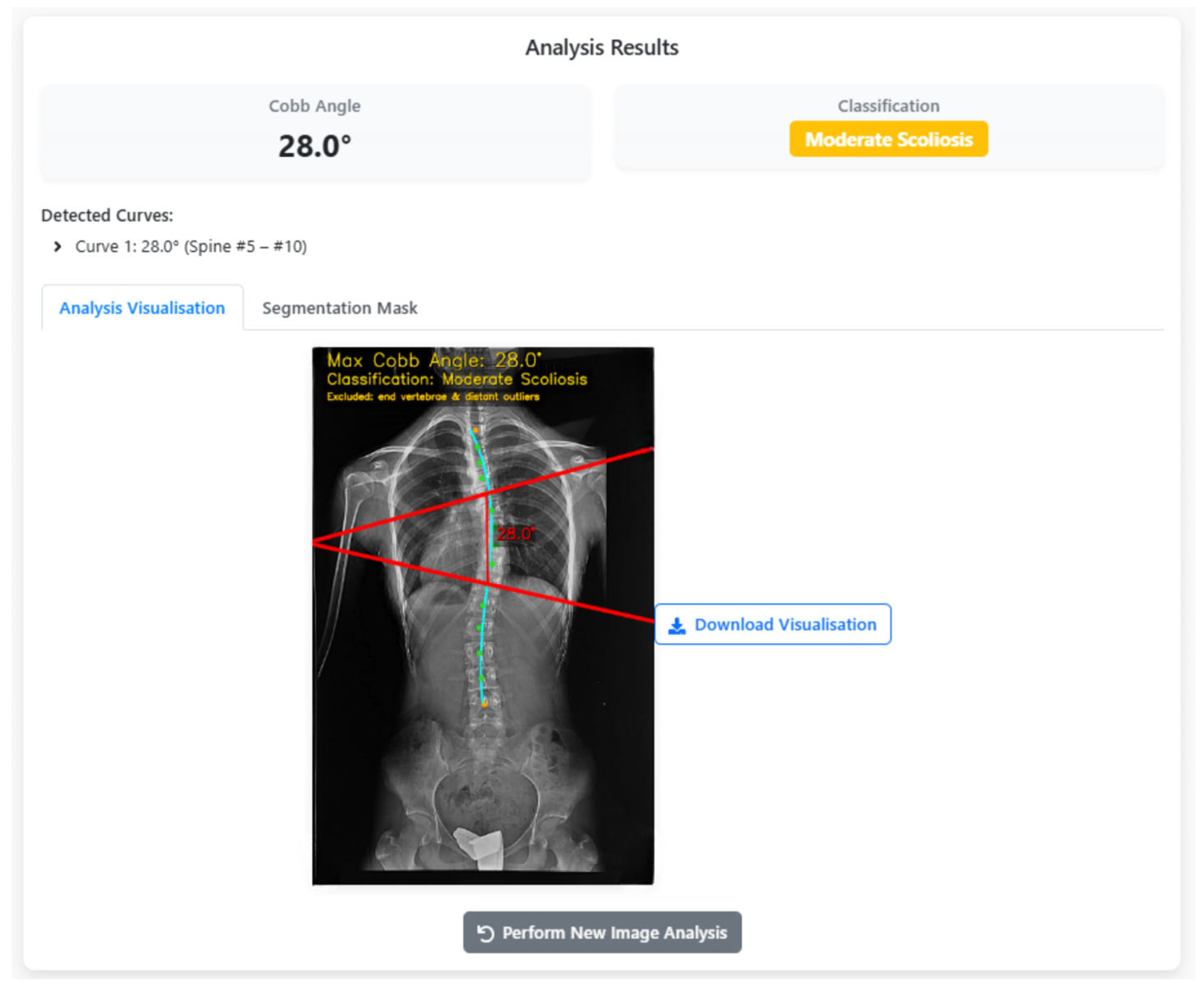
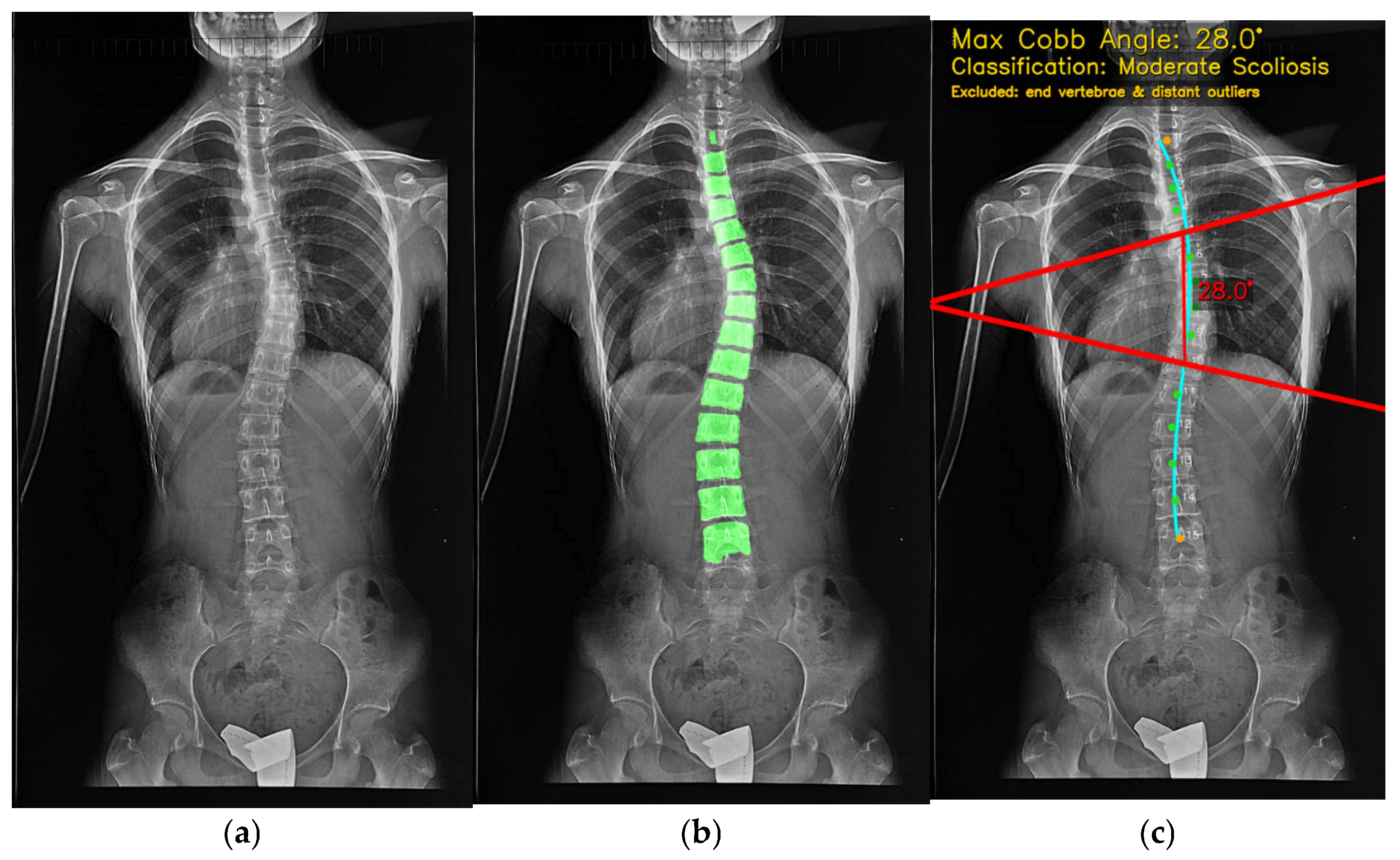
The results are presented to the user through a comprehensive visualization interface, as shown in Figure 12. Here, the maximum CA, scoliosis classification, and anatomical overlays are superimposed on the original radiograph, highlighting the analyzed vertebrae, endplate lines, and curvature trajectory. This graphical representation ensures interpretability, reproducibility, and clinical trust. When the analysis finishes, the “Maximum CA” (28.0°) is shown with key details for clinical decisions. The bottom row lists the measured vertebral range (Spine #5–#10). Red lines and blue reference points on the X-ray illustrate how the angle is computed. “Download Visualization” lets users export the figure, and “Perform New Image Analysis” restarts the workflow. Switching to the segmentation tab overlays a transparent green vertebral mask on the X-ray (Figure 13), showing extracted vertebrae and boundaries. The translucency lets users assess tissue and contours together. “Download Segmentation” saves/exports the mask for further analysis or reporting.
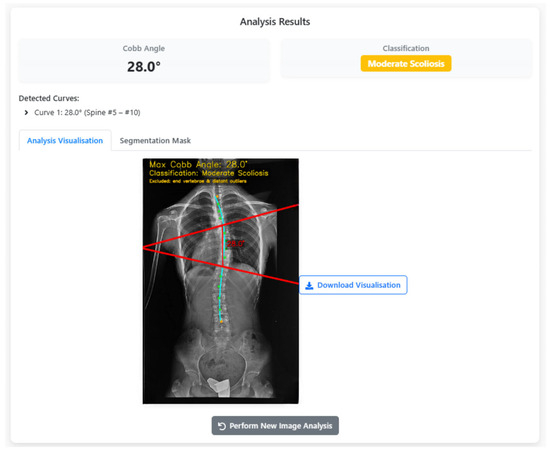
Figure 12.
CA calculation results: Visual overlay of vertebral endplates and classification.
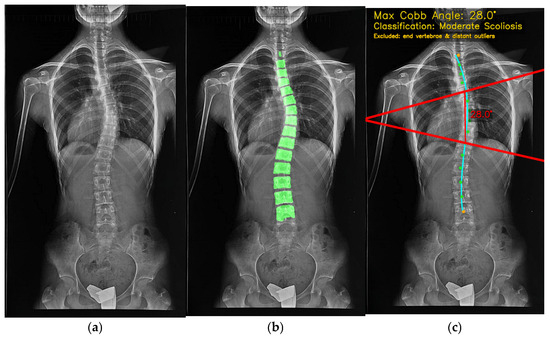
Figure 13.
The input X-ray image (a), vertebra segmentation (b) and final CA calculation output (c).
In Figure 12 and Figure 13, all colours and symbols follow a consistent convention. The bright green regions indicate the vertebral segmentation mask predicted by the U-Net model. The cyan spinal curve with small markers represents the fitted midline obtained from the centroids of the segmented vertebrae and is used for automatic curve detection. The solid red lines correspond to the upper and lower endplate tangents defining the maximum Cobb angle, and the red numeric label (e.g., “28.0°”) reports this Cobb angle in degrees. The yellow text at the top of the panel summarises the maximum Cobb angle and the corresponding scoliosis category (e.g., “Moderate Scoliosis”). The subfigure labels (a), (b) and (c) refer, respectively, to the original X-ray image, the vertebra segmentation overlay, and the final Cobb-angle visualisation generated by SpineCheck-AI. Interface text such as “Curve 1: 28.0° (Spine #5–#10)” denotes the first detected curve and the index range of vertebrae along the automatically identified spinal sequence.
6. Discussion
As summarized in Table 10, most recent systems converge on CNN backbones—typically U-Net variants—for vertebral parsing because they preserve endplate detail while remaining stable across heterogeneous acquisitions. Across datasets, scale and diversity remain decisive for generalizability: multi-center or PACS-sourced cohorts (e.g., the 2150-case series of Ha et al. [27]) broaden acquisition conditions but amplify label heterogeneity, whereas curated cohorts can yield smaller errors in mild-curve distributions (e.g., Horng et al., MAD ≈ 2.5° on curves < 20° [5]). In contrast to many prior scoliosis studies that focus primarily on algorithmic accuracy under controlled experimental conditions, our work emphasizes system-level deployability, large-scale validation, and clinical workflow integration.

Table 10.
A comparison of deep learning-based automated scoliosis measurement systems.
Within this methodological landscape, two families of Cobb-angle computation dominate. First, geometry-first pipelines—including Tan et al. and SpineCheck—fit endplate primitives (e.g., minimum-area rectangles), stabilize orientations using regression, and select the maximal inter-line angle [23]. Tan et al. explicitly combine minimum-area rectangles with least-squares line fitting to recover endplate orientations, reporting a small mean deviation (1.7 ± 1.2°) against an expert reader, exemplifying a geometry-first yet interpretable design [19]. Second, detector/centerline-guided pipelines generate a spinal trajectory and compute local slopes: in Ha et al., a Faster R-CNN (ResNet-101) proposes vertebral bodies, and a controller derives a smoothed centroid spline with endplate slope estimation; large-cohort validation showed strong correlation with clinical reports (Spearman ρ = 0.89) but higher mean differences when compared directly to report values (7.34°, 95% CI 5.90–8.78°). Agreement improved when human-drawn versus CNN-derived boxes were compared on the same images (MAD = 5.46°, ρ = 0.95) [27].
A third, practically important variant is cascaded segmentation with body-tilt angles, as in Wong et al., where vertebral-body bounding-box orientations—rather than explicit endplate lines—are used to derive CAs. In a 200-image validation, they report MAD = 2.8 ± 2.8°, ICC ≈ 0.92, and 91% of measurements within ±5°, with region- and severity-stratified analyses demonstrating stable performance across subgroups [29]. Methodologically, the body-tilt strategy is efficient but may diverge from manual endplate practice in severely wedged vertebrae—an acknowledged limitation that motivates explicit endplate modeling when wedging is prominent.
Consistent with this broader landscape, SpineCheck employs a U-Net-based vertebral segmentation pipeline. The achieved mean Dice score (~0.803 on 737 PA radiographs) is sufficient to preserve endplate geometry for downstream angle estimation and broadly aligns with prior CNN-based reports. Rather than optimizing segmentation accuracy in isolation, SpineCheck couples this performance with a geometry-driven Cobb computation stage designed for interpretability and robustness under clinical noise.
Unlike standard least-squares fitting approaches commonly adopted in prior studies, SpineCheck incorporates Theil–Sen robust regression and spline-based anatomical modeling to explicitly suppress outliers and compensate for vertebral misalignment. This design choice is particularly critical for real-world radiographs affected by noise, overlap, rotation, and partial occlusions, where small segmentation deviations can otherwise propagate into large angular errors.
Within this context, SpineCheck offers 3 practical strengths. (i) It is an end-to-end web platform (FastAPI + React) that returns downloadable overlays/masks and annotated angles, moving beyond algorithm-only reports. This directly addresses a persistent translational gap, even relative to works that demonstrate a prototype web app or planned PACS integration [21]. (ii) Its geometric Cobb calculator augments classic minimum-bounding-rectangle and slope routines with anatomical outlier filtering via a third-degree centroid spline, stabilizing endplate selection when spurious vertebrae perturb local orientation—an interpretable refinement inspired by centerline-guided methods yet retaining explicit endplate semantics [25]. (iii) A transparent training and evaluation pipeline built on a five-model comparative framework (U-Net, U-Net-2, Attention U-Net, UNet3++, nnU-Net) on a 737-image PA corpus yields a verified mean Dice of 0.8031, complemented by detailed worst-, median-, and best-case analyses and qualitative failure-mode inspection.
Although nnU-Net and UNet3++ achieved marginally higher Dice coefficients under controlled experimental conditions, their substantially increased training complexity, memory consumption, and deployment overhead limited their suitability for real-time or resource-constrained clinical use. In contrast, the optimized U-Net-2 model offered a superior balance between segmentation accuracy, inference speed, computational efficiency, and software robustness, reinforcing that clinical deployment requires a fundamentally different optimization objective than offline benchmark performance alone.
As summarized in Table 10, most existing systems were evaluated on relatively limited datasets and implemented as standalone algorithmic pipelines. In contrast, SpineCheck uniquely combines large-scale clinical validation, robust geometric modeling, and a fully integrated web-based deployment architecture. This distinction is central to translation from research prototypes to routine clinical use, where scalability, reproducibility, interoperability, and usability constraints become dominant performance factors.
Looking ahead, two validation priorities would most strengthen clinical readiness. First, a multi-site external reader study with blinded assessment—reporting MAE/MAD, ICC, and SEM with region- and severity-stratified analyses—would benchmark agreement under domain shift and communicate reliability to clinicians, mirroring the reporting discipline in Suri et al. (hardware-invariant, ICC = 0.96; <2° error across internal and external tests) and Wong et al. (ICC ≈ 0.92; 91% within ±5°) [28,29]. Second, targeted ablation studies isolating the spline-based outlier filter (with and without filtering) would directly quantify its contribution to endplate stability relative to pure minimum-bounding-rectangle or body-tilt baselines. In parallel, pragmatic extensions—such as automated quality-control flags for missing vertebrae and exportable, overlay-rich reports—would further facilitate clinical integration and prospective evaluation [27].
Overall, the consolidated evidence indicates that a transparent, geometry-driven workflow, grounded in multi-model, high-quality vertebral segmentation, and enriched with lightweight anatomical filtering, remains a strong, interpretable baseline for automated Cobb measurement. With rigorous multi-site validation and standardized agreement statistics, SpineCheck is positioned for reproducible deployment, although prospective multi-center reader studies remain essential to fully quantify inter-observer variability and long-term clinical reliability.
7. Conclusions
This study presented SpineCheck, a fully integrated deep-learning-based clinical platform for automated vertebra segmentation and CA measurement from scoliosis X-ray images. The system unifies end-to-end medical image processing, robust geometric angle estimation, and a web-based clinical interface within a single deployable architecture designed for real-world use.
A systematic evaluation of five U-Net-based segmentation models demonstrated that model selection for clinical deployment must consider not only segmentation accuracy but also inference speed, memory efficiency, and software stability. The optimized U-Net-2 architecture was selected as the operational backbone of SpineCheck due to its favorable trade-off between performance and computational efficiency.
The proposed CA computation pipeline, combining robust regression with spline-based anatomical modeling, enables reliable angle estimation even in challenging scoliosis cases. Large-scale validation on 20,000 X-ray images confirmed strong statistical agreement with expert measurements using MAE, Pearson correlation, and ICC metrics, highlighting the clinical reliability of the platform.
By bridging deep learning, geometric modeling, and full-stack system design, SpineCheck advances beyond algorithm-level solutions and offers a scalable, reproducible, and clinically oriented decision support system for scoliosis assessment. The platform has strong potential to reduce clinician workload, improve measurement consistency, and enable large-scale scoliosis screening.
Future work will extend beyond clinical validation and system robustness toward a broader strategic research direction. The SpineCheck project was originally initiated in response to a real clinical need articulated by a physician at the Atlas University Hospital and further developed as an undergraduate senior design project. Through intensive experimentation, multi-architecture benchmarking, and iterative refinement, SpineCheck has now reached a stage where it is technically mature and ready for pilot deployment in clinical workflows.
Building upon this foundation, a new research phase has begun: the development of SpineCheck-LLM, an advanced, agent-enhanced scoliosis decision support platform. This new system aims to incorporate the Large Language Model (LLM)-based clinical agents into the SpineCheck pipeline. These agents will interpret CA measurements, synthesize diagnostic explanations, generate structured radiology-style reports, and provide patient-specific recommendations. The long-term goal is to transform SpineCheck from a measurement-focused tool into a comprehensive clinical assistant capable of supporting diagnostic reasoning, reporting quality control, and longitudinal disease monitoring. This new phase is also strategically aligned with the aim of securing institutional research funding and expanding collaborations with Atlas University Hospital, potentially opening the door to additional AI-augmented medical imaging projects.
Author Contributions
Writing, review and editing, İ.H.İ., H.G. and F.T.; software and models, İ.H.İ. and F.T.; methodology, H.G.; supervision and validation H.G. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by Atlas University through the Scientific Research Projects (BAP) program under Project No. 2022/3, entitled “Design and Development of Hardware and Software Frameworks of a Deep Computer for Artificial Intelligence and Data Science Fundamental and Research Laboratories.”
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
İlkhan, İ.H., Gümüşkaya, H., and Turgut, F. Vertebrae for Scoliosis (Dataset). Version 1, Mendeley Data, 25 September 2025. DOI:10.17632/4kby36n3ng.2 Available online: https://data.mendeley.com/datasets/4kby36n3ng/2 (accessed on 25 November 2025).
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Scoliosis Degrees of Curvature Chart. Scoliosis Reduction Center. Available online: https://www.scoliosisreductioncenter.com/blog/scoliosis-degrees-of-curvature-chart/ (accessed on 30 August 2025).
- Khanal, B.; Dahal, L.; Adhikari, P.; Khanal, B. Automatic Cobb Angle Detection Using Vertebra Detector and Vertebra Corners Regression. In International Workshop and Challenge on Computational Methods and Clinical Applications for Spine Imaging; Springer: Cham, Switzerland, 2019; pp. 81–87. [Google Scholar]
- Eltanboly, A.; Ghazal, M.; Hajjdiab, H.; Shalaby, A.; Switala, A.; Mahmoud, A.; Sahoo, P.; El-Azab, M.; El-Baz, A. Level Sets-Based Image Segmentation Approach Using Statistical Shape Priors. Appl. Math. Comput. 2019, 340, 164–179. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. arXiv 2015, arXiv:1505.04597. [Google Scholar] [CrossRef]
- Horng, M.-H.; Kuok, C.-P.; Fu, M.-J.; Lin, C.-J.; Sun, Y.-N. Cobb Angle Measurement of Spine from X-Ray Images Using Convolutional Neural Network. Comput. Math. Methods Med. 2019, 2019, 6357171. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Gu, F.; Zeng, D.; Lu, J.; Liu, H.; Hou, Y.; Zhang, Q. An Effective U-Net and BiSeNet Complementary Network for Spine Segmentation. Biomed. Signal Process. Control 2024, 89, 105682. [Google Scholar] [CrossRef]
- Nascimento, E.M.; Silva, A.; Almeida, J.D.S.; Pessoa, A.C.P.; Figueredo, W.K.R.; Silva, I.; Junior, G.B.; Paiva, A.; Cunha, A. A Cascade Method Based on UNet3+ for Automated Vertebrae Segmentation in X-Ray Images. Procedia Comput. Sci. 2025, 256, 789–796. [Google Scholar] [CrossRef]
- AASCE—Grand Challenge. Available online: https://aasce19.grand-challenge.org/ (accessed on 8 September 2025).
- Wang, H.; Song, S.; Lu, T.; Wang, S.; Chen, M.; Du, Y.; Zhang, J.; Xiao, R. Landmark Detection Network of Adolescent Idiopathic Scoliosis by Integrating the Image Enhancement Module and Vertebral Center Guidance. In Proceedings of the 2024 5th International Seminar on Artificial Intelligence, Networking and Information Technology (AINIT), Nanjing, China, 29–31 March 2024; IEEE: New York, NY, USA, 2024; pp. 945–949. [Google Scholar]
- Wu, H.; Bailey, C.; Rasoulinejad, P.; Li, S. Automatic Landmark Estimation for Adolescent Idiopathic Scoliosis Assessment Using BoostNet. In Proceedings of the Medical Image Computing and Computer Assisted Intervention—MICCAI 2017, Quebec City, QC, Canada, 11–13 September 2017; Descoteaux, M., Maier-Hein, L., Franz, A., Jannin, P., Collins, D.L., Duchesne, S., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 127–135. [Google Scholar]
- Yi, J.; Wu, P.; Huang, Q.; Qu, H.; Metaxas, D.N. Vertebra-focused landmark detection for scoliosis assessment. In Proceedings of the IEEE 17th International Symposium on Biomedical Imaging (ISBI), Iowa City, IA, USA, 3–7 April 2020; pp. 736–740. [Google Scholar]
- Oktay, O.; Schlemper, J.; Folgoc, L.L.; Lee, M.J.; Heinrich, M.; Misawa, K.; Mori, K.; McDonagh, S.G.; Hammerla, N.Y.; Kainz, B. Attention U-Net: Learning Where to Look for the Pancreas. arXiv 2018, arXiv:1804.03999. [Google Scholar] [CrossRef]
- Zhou, Z.; Siddiquee, M.M.R.; Tajbakhsh, N.; Liang, J. UNet++: A Nested U-Net Architecture for Medical Image Segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support; Springer: Cham, Switzerland, 2018; pp. 3–11. [Google Scholar] [CrossRef]
- Huang, H.; Lin, L.; Tong, R.; Hu, H.; Zhang, Q.; Iwamoto, Y.; Han, X.; Chen, Y.-W.; Wu, J. UNet 3+: A Full-Scale Connected UNet for Medical Image Segmentation. In Proceedings of the ICASSP 2020—IEEE International Conference on Acoustics, Speech and Signal Processing, Barcelona, Spain, 4–8 May 2020; pp. 1055–1059. [Google Scholar] [CrossRef]
- Xu, H.; Cui, X.; Li, C.; Tian, Z.; Liu, J.; Yang, J. RUnT: A Network Combining Residual U-Net and Transformer for Vertebral Edge Feature Fusion Constrained Spine CT Image Segmentation. IEEE Access 2023, 11, 55692–55705. [Google Scholar] [CrossRef]
- Isensee, F.; Jaeger, P.F.; Kohl, S.A.A.; Petersen, J.; Maier-Hein, K.H. nnU-Net: Self-Configuring Method for Deep Learning–Based Biomedical Image Segmentation. Nat. Methods 2021, 18, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Bae, W.; Masuda, K.; Chung, C.; Hwang, D. Fine-Grain Segmentation of the Intervertebral Discs from MR Spine Images Using Deep Convolutional Neural Networks: BSU-Net. Appl. Sci. 2018, 8, 1656. [Google Scholar] [CrossRef] [PubMed]
- Lakshmi, S.V.V.; Darmanayagam, S.E.; Cyril, S.R.R. Semantic Segmentation of Lumbar Vertebrae Using Meijering U-Net (MU-Net) on Spine Magnetic Resonance Images. Comput. Model. Eng. Sci. 2025, 142, 733–757. [Google Scholar] [CrossRef]
- Li, L.; Zhang, T.; Lin, F.; Li, Y.; Wong, M.-S. Automated 3D Cobb Angle Measurement Using U-Net in CT Images of Preoperative Scoliosis Patients. J. Imaging Inform. Med. 2025, 38, 309–317. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, S.; Lyu, J.; Huang, Z.; Leung, F.H.F.; Lee, T.; Yang, D.; Su, S.; Zheng, Y.; Ling, S.H. Ultrasound Spine Image Segmentation Using Multi-Scale Feature Fusion Skip-Inception U-Net (SIU-Net). Biocybern. Biomed. Eng. 2022, 42, 341–361. [Google Scholar] [CrossRef]
- Zhou, Y.; Lee, T.T.-Y.; Lai, K.K.-L.; Wu, C.; Lau, H.T.; Yang, D.; Song, Z.; Chan, C.-Y.; Chu, W.C.-W.; Cheng, J.C.-Y.; et al. Automatic Ultrasound Curve Angle Measurement via Affinity Clustering for Adolescent Idiopathic Scoliosis Evaluation. Expert Syst. Appl. 2025, 269, 126410. [Google Scholar] [CrossRef]
- Kumar, R.; Gupta, M.; Abraham, A. A Critical Analysis on Vertebra Identification and Cobb Angle Estimation Using Deep Learning for Scoliosis Detection. IEEE Access 2024, 12, 11170–11184. [Google Scholar] [CrossRef]
- Tan, Z.; Yang, K.; Sun, Y.; Wu, B.; Tao, H.; Hu, Y.; Zhang, J. An Automatic Scoliosis Diagnosis and Measurement System Based on Deep Learning. In Proceedings of the 2018 IEEE International Conference on Robotics and Bio-mimetics (ROBIO), Kuala Lumpur, Malaysia, 12–15 December 2018; IEEE: New York, NY, USA, 2018; pp. 439–443. [Google Scholar]
- Spine_Det Dataset > Overview. Available online: https://universe.roboflow.com/test42-sjp0k/spine_det (accessed on 6 May 2025).
- Ernestchenchen. Spinal-AI2024: The Project Is About the Spinal-AI2024 Dataset of Spinal Scoliosis X-Ray Images. Available online: https://github.com/Ernestchenchen/Spinal-AI2024/tree/main (accessed on 25 November 2025).
- Shao, Z.; Yuan, Y.; Ma, L.; Zhu, X. CurvNet: Latent Contour Representation and Iterative Data Engine for Curvature Angle Estimation. arxiv 2024, arXiv:2411.12604. [Google Scholar] [CrossRef]
- Ha, A.Y.; Do, B.H.; Bartret, A.L.; Fang, C.X.; Hsiao, A.; Lutz, A.M.; Banerjee, I.; Riley, G.M.; Rubin, D.L.; Stevens, K.J.; et al. Automating Scoliosis Measurements in Radiographic Studies with Machine Learning: Comparing Artificial Intelligence and Clinical Reports. J. Digit. Imaging 2022, 35, 524–533. [Google Scholar] [CrossRef] [PubMed]
- Suri, A.; Tang, S.; Kargilis, D.; Taratuta, E.; Kneeland, B.J.; Choi, G.; Agarwal, A.; Anabaraonye, N.; Xu, W.; Parente, J.B.; et al. Conquering the Cobb Angle: A Deep Learning Algorithm for Automated, Hardware-Invariant Measurement of Cobb Angle on Radiographs in Patients with Scoliosis. Radiol. Artif. Intell. 2023, 5, e220158. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.C.; Reformat, M.Z.; Parent, E.C.; Stampe, K.P.; Southon Hryniuk, S.C.; Lou, E.H. Validation of an Artificial Intelligence-Based Method to Automate Cobb Angle Measurement on Spinal Radiographs of Children with Adolescent Idiopathic Scoliosis. Eur. J. Phys. Rehabil. Med. 2023, 59, 535–542. [Google Scholar] [CrossRef] [PubMed]
- Maaliw, R.R. SCOLIONET: An Automated Scoliosis Cobb Angle Quantification Using Enhanced X-Ray Images and Deep Learning Models. J. Imaging 2023, 9, 265. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).