Abnormal Hypermethylation of CpG Dinucleotides in Promoter Regions of Matrix Metalloproteinases Genes in Breast Cancer and its Relation to Epigenomic Subtypes and HER2 Overexpression
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Material
2.2. DNA Isolation and Methylation-Sensitive Restriction Enzyme Digestion PCR (MSRE-PCR)
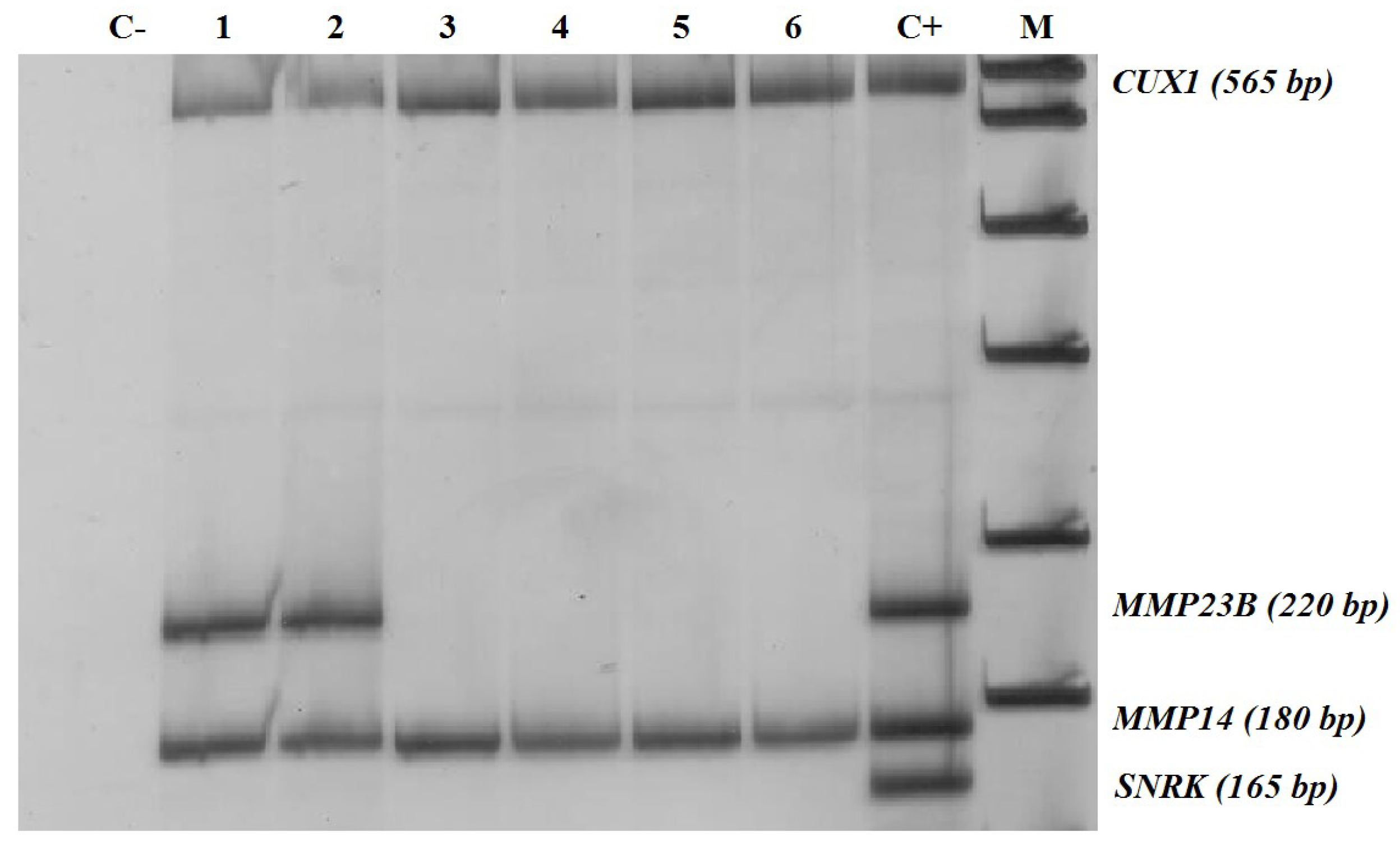
2.3. Methylation-Sensitive Restriction Enzyme Digestion PCR (MSRE-PCR) Assays
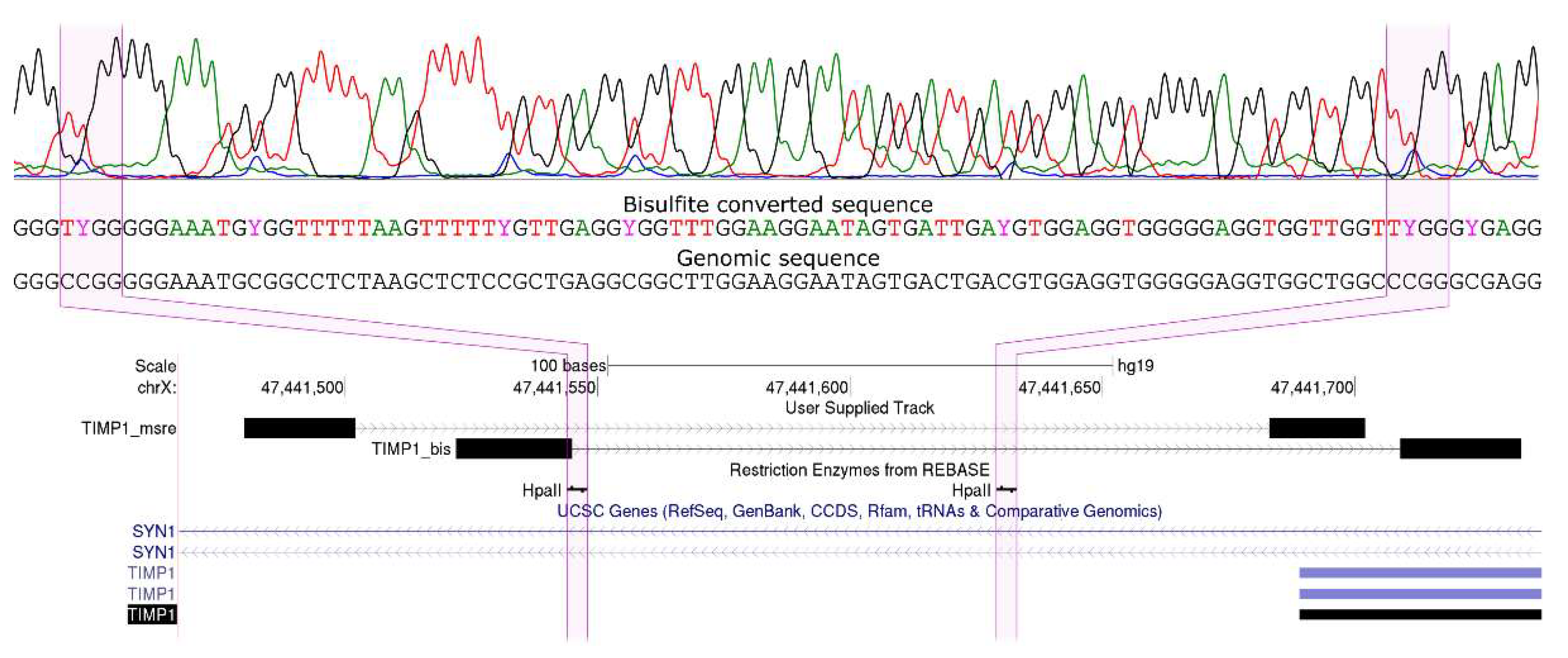
2.4. Bisulfite Sequencing by Sanger
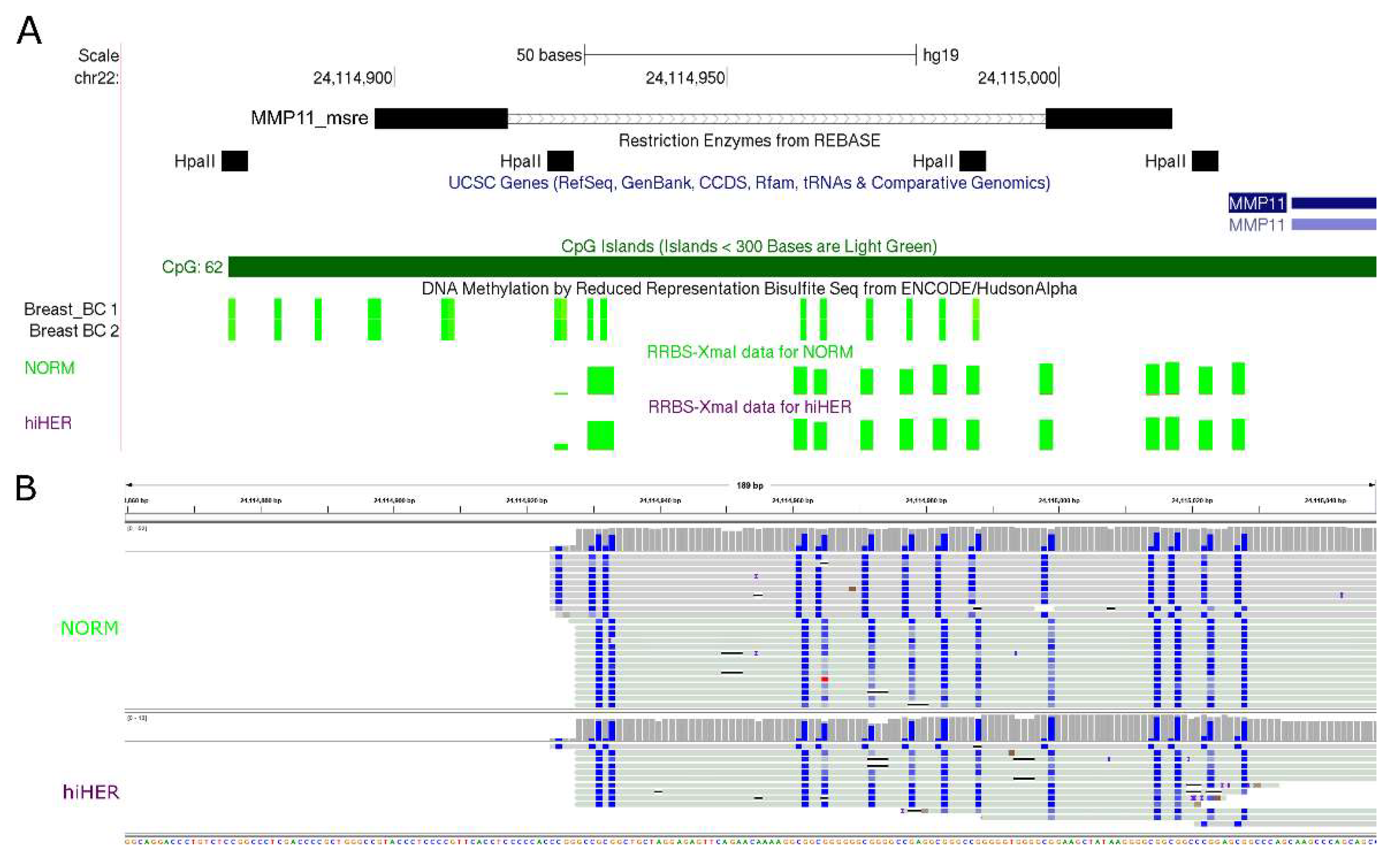
2.5. Validation of MSRE-PCR Results by RRBS
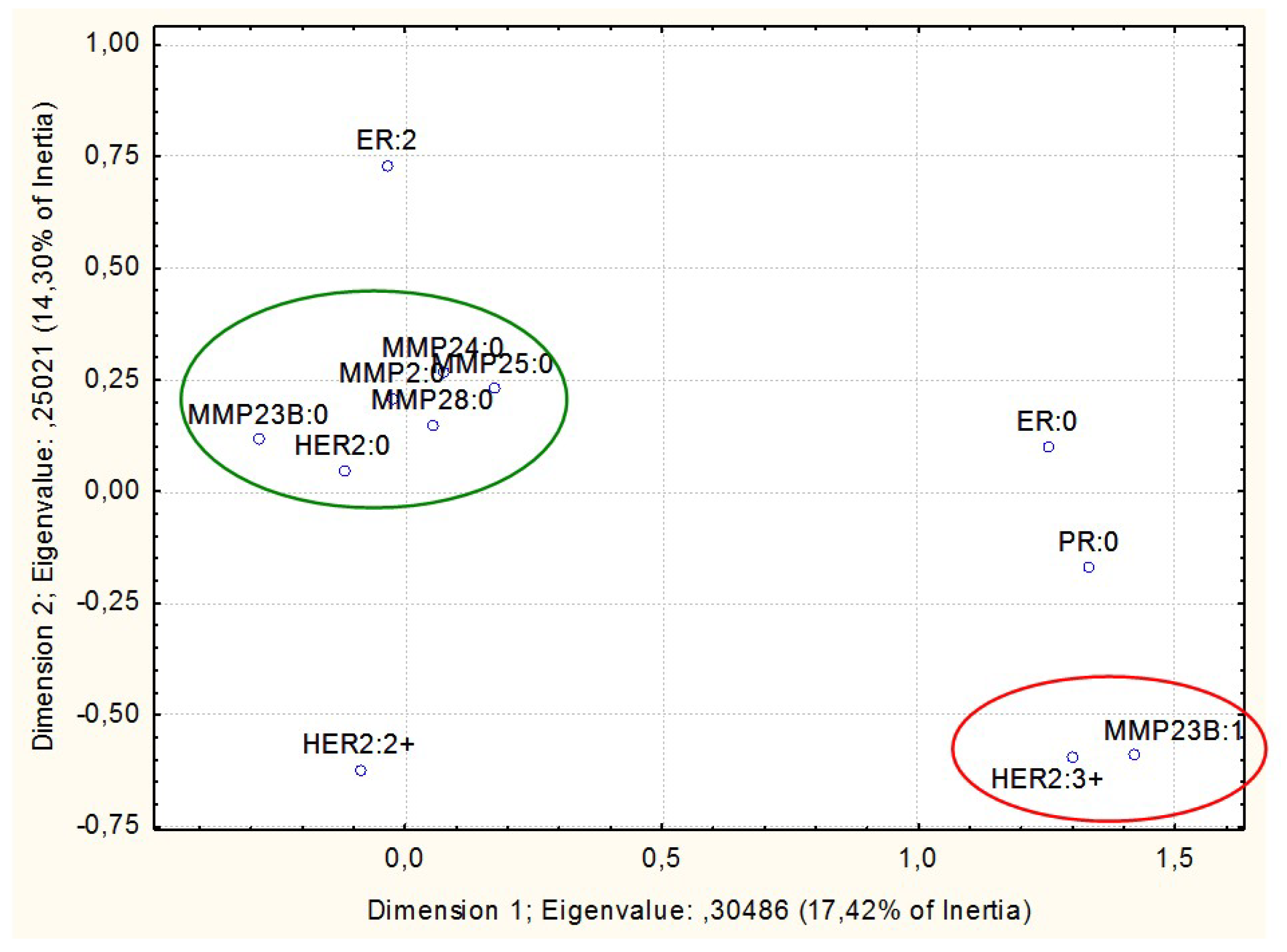
3. Results
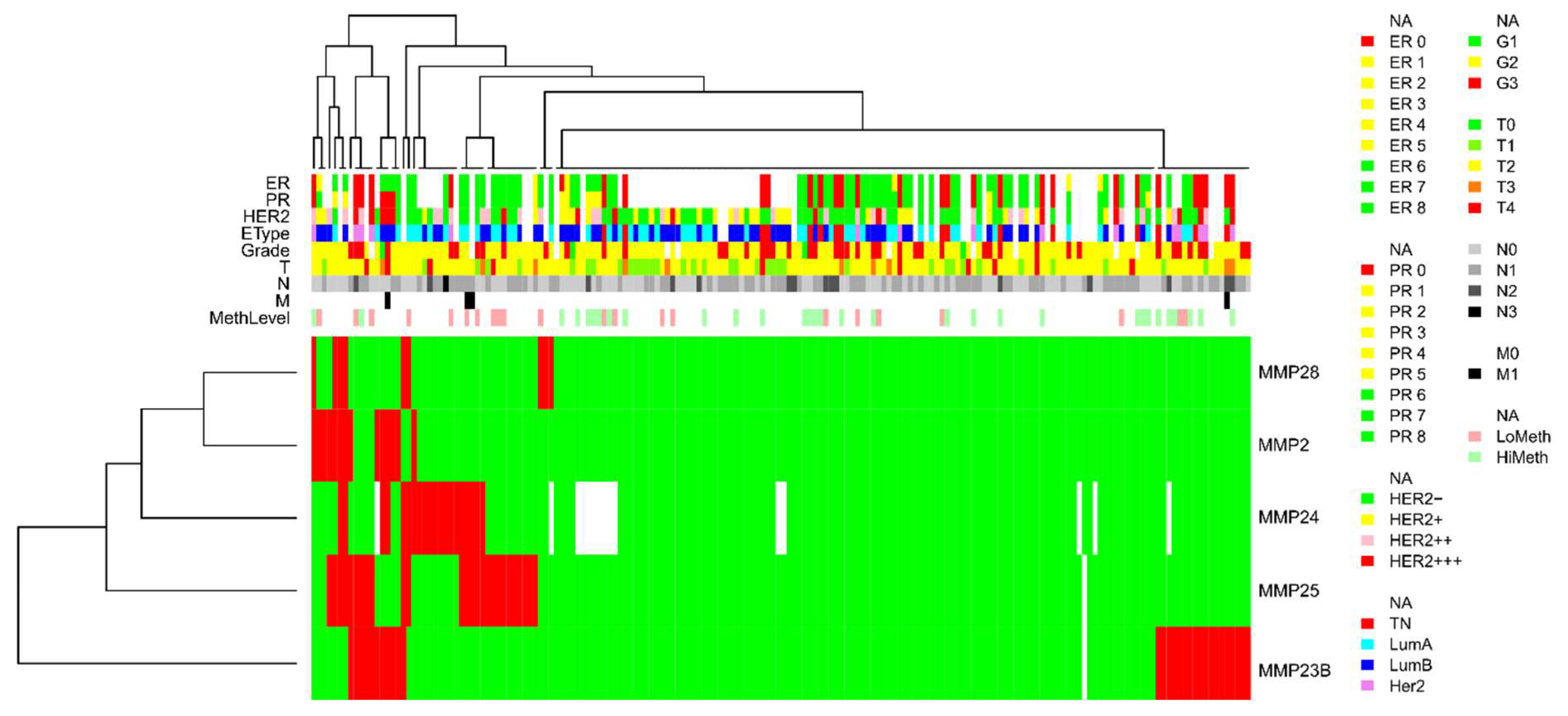
- Non-methylated in normal breast tissues but prone to abnormal hypermethylation in breast cancer (MMP2, MMP23B, MMP24, MMP25, and MMP28; with the proportion of methylated samples indicated in Table 1)
- Non-methylated in both normal breast tissues and in breast cancer (TIMP2, TIMP3, MMP11, MMP15, MMP16, and MMP17)
- Constitutively methylated in normal and cancerous breast tissues (TIMP1, TIMP4, MMP14, and MMP21)
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| BC | breast cancer |
| EMT | epithelial–mesenchymal transition |
| CpG | 5′-cytosine-phosphate-guanine-3′ |
| MMP | matrix metalloproteinase |
| MSRE-PCR | methylation-sensitive restriction enzyme digestion PCR |
| TIMPs | tissue inhibitors of matrix metalloproteinases |
| RRBS | reduced representation bisulfite sequencing |
References
- Wang, X.; Khalil, R.A. Matrix metalloproteinases, vascular remodeling, and vascular disease. Adv. Pharmacol. 2018, 81, 241–330. [Google Scholar] [CrossRef] [PubMed]
- Yip, C.; Foidart, P.; Noël, A.; Sounni, N.E. MT4-MMP: The GPI-Anchored membrane-type matrix metalloprotease with multiple functions in diseases. Int. J. Mol. Sci. 2019, 20, 354. [Google Scholar] [CrossRef] [PubMed]
- Tokito, A.; Jougasaki, M. Matrix metalloproteinases in non-neoplastic disorders. Int. J. Mol. Sci. 2016, 17, 1178. [Google Scholar] [CrossRef] [PubMed]
- Myasoedova, V.A.; Chistiakov, D.A.; Grechko, A.V.; Orekhov, A.N. Matrix metalloproteinases in pro-atherosclerotic arterial remodeling. J. Mol. Cell. Cardiol. 2018, 123, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Kessenbrock, K.; Plaks, V.; Werb, Z. Matrix metalloproteinases: Regulators of the tumor microenvironment. Cell 2010, 141, 52–67. [Google Scholar] [CrossRef] [PubMed]
- Quintero-Fabián, S.; Arreola, R.; Becerril-Villanueva, E.; Torres-Romero, J.C.; Arana-Argáez, V.; Lara-Riegos, J.; Ramírez-Camacho, M.A.; Alvarez-Sánchez, M.E. Role of Matrix Metalloproteinases in Angiogenesis and Cancer. Front. Oncol. 2019, 9, 1370. [Google Scholar] [CrossRef]
- Moss, L.A.S.; Jensen-Taubman, S.; Stetler-Stevenson, W.G. Matrix Metalloproteinases. Changing Roles in Tumor Progression and Metastasis. Am. J. Pathol. 2012, 181, 1895–1899. [Google Scholar] [CrossRef]
- Hadler-Olsen, E.; Winberg, J.O.; Uhlin-Hansen, L. Matrix metalloproteinases in cancer: Their value as diagnostic and prognostic markers and therapeutic targets. Tumour Biol. 2013, 34, 2041–2051. [Google Scholar] [CrossRef]
- Radisky, E.S.; Radisky, D.C. Matrix metalloproteinases as breast cancer drivers and therapeutic targets. Front. Biosci. (Landmark Ed.) 2015, 20, 1144–1163. [Google Scholar] [CrossRef]
- Cierna, Z.; Mego, M.; Janega, P.; Karaba, M.; Minarik, G.; Benca, J.; Sedlácková, T.; Cingelova, S.; Gronesova, P.; Manasova, D.; et al. Matrix metalloproteinase 1 and circulating tumor cells in early breast cancer. BMC Cancer. 2014, 14, 472. [Google Scholar] [CrossRef]
- Mehner, G.; Hockla, A.; Miller, E.; Ran, S.; Radisky, D.C.; Radisky, E.S. Tumor cell-produced matrix metalloproteinase 9 (MMP-9) drives malignant progression and metastasis of basal-like triple negative breast cancer. Oncotarget 2014, 5, 2736–2749. [Google Scholar] [CrossRef] [PubMed]
- Radisky, E.S.; Raeeszadeh-Sarmazdeh, M.; Radisky, D.C. Therapeutic potential of matrix metalloproteinase inhibition in breast cancer. J. Cell Biochem. 2017, 118, 3531–3548. [Google Scholar] [CrossRef] [PubMed]
- Simonova, O.A.; Kuznetsova, E.B.; Poddubskaya, E.V.; Kekeeva, T.V.; Kerimov, R.A.; Trotsenko, I.D.; Tanas, A.S.; Rudenko, V.V.; Alekseeva, E.A.; Zaletayev, D.V.; et al. DNA methylation in the promoter regions of the laminin family genes in normal and breast carcinoma tissues. Mol. Biol. 2015, 49, 598–607. [Google Scholar] [CrossRef]
- Wang, H.; Maurano, M.T.; Qu, H.; Varley, K.E.; Gertz, J.; Pauli, F.; Lee, K.; Canfield, T.; Weaver, M.; Sandstrom, R.; et al. Widespread plasticity in CTCF occupancy linked to DNA methylation. Genome Res. 2012, 22, 1680–1688. [Google Scholar] [CrossRef]
- Tanas, A.S.; Sigin, V.O.; Kalinkin, A.I.; Litviakov, N.V.; Slonimskaya, E.M.; Ibragimova, M.K.; Ignatova, E.O.; Simonova, O.A.; Kuznetsova, E.B.; Kekeeva, T.V.; et al. Genome-wide methylotyping resolves breast cancer epigenetic heterogeneity and suggests novel therapeutic perspectives. Epigenomics 2019, 11, 605–617. [Google Scholar] [CrossRef]
- Tanas, A.S.; Borisova, M.E.; Kuznetsova, E.B.; Rudenko, V.V.; Karandasheva, K.O.; Nemtsova, M.V.; Izhevskaya, V.L.; Simonova, O.A.; Larin, S.S.; Zaletaev, D.V.; et al. Rapid and affordable genome-wide bisulfite DNA sequencing by XmaI-reduced representation bisulfite sequencing. Epigenomics 2017, 9, 833–847. [Google Scholar] [CrossRef]
- Köhrmann, A.; Kammerer, U.; Kapp, M.; Dietl, J.; Anacker, J. Expression of matrix metalloproteinases (MMPs) in primary human breast cancer and breast cancer cell lines: New findings and review of the literature. BMC Cancer 2009, 9, 188. [Google Scholar] [CrossRef]
- Mondal, S.; Adhikari, N.; Banerjee, S.; Amin, S.A.; Jha, T. Matrix metalloproteinase-9 (MMP-9) and its inhibitors in cancer: A minireview. Eur. J. Med. Chem. 2020, 194, 112260. [Google Scholar] [CrossRef]
- Melnikov, A.A.; Gartenhaus, R.B.; Levenson, A.S.; Motchoulskaia, N.A.; Levenson, V.V. MSRE-PCR for analysis of gene-specific DNA methylation. Nucleic Acids Res. 2005, 33, e93. [Google Scholar] [CrossRef]
- Díez-Villanueva, A.; Mallona, I.; Peinado, M.A. Wanderer, an interactive viewer to explore DNA methylation and gene expression data in human cancer. Epigenetics Chromatin 2015, 8, 22. [Google Scholar] [CrossRef]
- Terada, K.; Okochi-Takada, E.; Akashi-Tanaka, S.; Miyamoto, K.; Taniyama, K.; Tsuda, H.; Asada, K.; Kaminishi, M.; Ushijima, T. Association between frequent CpG island methylation and HER2 amplification in human breast cancers. Carcinogenesis 2009, 30, 466–471. [Google Scholar] [CrossRef] [PubMed]
- Marmor, M.D.; Skaria, K.B.; Yarden, Y. Signal transduction and oncogenesis by ErbB/HER receptors. Int. J. Radiat Oncol. Biol. Phys. 2004, 58, 903–913. [Google Scholar] [CrossRef] [PubMed]
- Subramaniam, M.M.; Chan, J.Y.; Soong, R.; Ito, K.; Ito, Y.; Yeoh, K.G.; Salto-Tellez, M.; Putti, T.C. RUNX3 inactivation by frequent promoter hypermethylation and protein mislocalization constitute an early event in breast cancer progression. Breast Cancer Res. Treat. 2009, 113, 113–121. [Google Scholar] [CrossRef] [PubMed]
- Lindqvist, B.M.; Wingren, S.; Motlagh, P.B.; Nilsson, T.K. Whole genome DNA methylation signature of HER2-positive breast cancer. Epigenetics 2014, 9, 1149–1162. [Google Scholar] [CrossRef]
- Bao, W.; Fu, H.J.; Jia, L.T.; Zhang, Y.; Li, W.; Jin, B.Q.; Yao, L.B.; Chen, S.Y.; Yang, A.G. HER2-mediated upregulation of MMP-1 is involved in gastric cancer cell invasion. Arch. Biochem. Biophys. 2010, 499, 49–55. [Google Scholar] [CrossRef]
- Hegedüs, L.; Cho, H.; Xie, X.; Eliceiri, G.L. Additional MDA-MB-231 breast cancer cell matrix metalloproteinases promote invasiveness. J. Cell. Physiol. 2008, 216, 480–485. [Google Scholar] [CrossRef]
- Benson, C.S.; Babu, S.D.; Radhakrishna, S.; Selvamurugan, N.; Sankar, B.R. Expression of matrix metalloproteinases in human breast cancer tissues. Dis. Markers 2013, 34, 395–405. [Google Scholar] [CrossRef]
- Luo, Y.P.; Zhong, M.; Wang, L.P.; Sun, G.Q.; Li, J. Inhibitory effects of RNA interference on MMP-24 expression and invasiveness of ovarian cancer SKOV(3) cells. J. South. Med. Univ. 2009, 29, 781–784. [Google Scholar]
- Chen, Y.; Sumardika, I.W.; Tomonobu, N.; Ruma, I.M.W.; Kinoshita, R.; Kondo, E.; Inoue, Y.; Sato, H.; Yamauchi, A.; Murata, H.; et al. Melanoma cell adhesion molecule is the driving force behind the dissemination of melanoma upon S100A8/A9 binding in the original skin lesion. Cancer Lett. 2019, 452, 178–190. [Google Scholar] [CrossRef]
- Sun, Q.; Weber, C.R.; Sohail, A.; Bernardo, M.M.; Toth, M.; Zhao, H.; Turner, J.R.; Fridman, R. MMP25 (MT6-MMP) is highly expressed in human colon cancer, promotes tumor growth, and exhibits unique biochemical properties. J. Biol. Chem. 2007, 282, 21998–22010. [Google Scholar] [CrossRef]
| Gene | Primers | PCR Product Size, bp | Number of HpaII Sites within MSRE-PCR Fragment |
|---|---|---|---|
| MMP2 | F: TACAAAGGGATTGCCAGGAC R: CATTAGCGCCTCCATCGTAG | 239 | 3 |
| MMP11 | F: GTACCCTCCCCGTTCACCTC R: GCCGCCCCTTATAGCTTCC | 120 | 2 |
| MMP14 | F: GCCGACAGCGGTCTAGGAAT R: CAGGGGGAGCAGGAGACAAC | 180 | 3 |
| MMP15 | F: CTCCTCGGGCTTGGGAATTT R: CCAGCTCGGAACACTGCAC | 155 | 4 |
| MMP16 | F: CGAGAGGCAGCGGCGAAG R: CGGAACCGCCGGTGAACTTA | 100 | 3 |
| MMP17 | F: CCGGCCTCGTTAGCATACAT R: CCCTCCGCTTCGCGTTCC | 125 | 3 |
| MMP21 | F: GCCACTCCTCCCTCTCAGC R: CCACCCAGCCCGAGAGTC | 250 | 2 |
| MMP23B | F: ACCACACCGGGCTGTAACC R: AGGAGGCACAGGGCGACCA | 220 | 5 |
| MMP24 | F: CAGAGCCGCTCCTCAGTCTC R: AGGAGGGGGAAGAGGCTAAA | 174 | 2 |
| MMP25 | F: CTCCCGCGCCCTCTCAAC R: GAAGTGCGCGGTGGAGTC | 101 | 2 |
| MMP28 | F: CGTGCCTGTGTGGTTCCAG R: CCTGTCAGAACTCGGCAGTC | 150 | 2 |
| TIMP1 | F: TGAGTCATAGGGAGCTTGGGGG R: CGGGCCGACGAAAGGAGAT | 223 | 2 |
| TIMP2 | F: AAGCAGCGTCGCCAGCAG R: CCCCCGAGACAAAGAGGAGA | 246 | 3 |
| TIMP3 | F: CCCTCACCTGTGGAAGCGGT R: CAGACCAATGGCAGAGCCGCA | 318 | 4 |
| TIMP4 | F: ACCCCCTGCTGTGGACCTC R: CAAGCTGGGTGCTGTTGCTG | 150 | 2 |
| CUX1 | F: GCCCCCGAGGACGCCGCTACC R: AGGCGGTCCAGGGGTCCAGGC | 565 | 6 |
| SNRK | F: GCTGGGTGCGGGGTTTCGGCG R: CGGAGGCTACTGAGGCGGCGG | 165 | 3 |
| Gene | Methylated in Breast Cancer | Methylated in Normal Breast Tissues | Presence (+) or Absence (–) of Methylation in Breast Cancer Cell Lines | ||||
|---|---|---|---|---|---|---|---|
| ZR751 | MCF7 | T47D | BT474 | HS578T | |||
| MMP2 | 7.7% (14/183) | 0 | + | + | + | − | − |
| MMP23B | 17% (31/182) | 0 | + | + | + | − | + |
| MMP24 | 11.9% (20/168) | 0 | − | − | - | + | - |
| MMP25 | 15.4% (28/182) | 0 | − | + | + | − | − |
| MMP28 | 4.9% (9/183) | 0 | + | + | + | + | − |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Simonova, O.A.; Kuznetsova, E.B.; Tanas, A.S.; Rudenko, V.V.; Poddubskaya, E.V.; Kekeeva, T.V.; Trotsenko, I.D.; Larin, S.S.; Kutsev, S.I.; Zaletaev, D.V.; et al. Abnormal Hypermethylation of CpG Dinucleotides in Promoter Regions of Matrix Metalloproteinases Genes in Breast Cancer and its Relation to Epigenomic Subtypes and HER2 Overexpression. Biomedicines 2020, 8, 116. https://doi.org/10.3390/biomedicines8050116
Simonova OA, Kuznetsova EB, Tanas AS, Rudenko VV, Poddubskaya EV, Kekeeva TV, Trotsenko ID, Larin SS, Kutsev SI, Zaletaev DV, et al. Abnormal Hypermethylation of CpG Dinucleotides in Promoter Regions of Matrix Metalloproteinases Genes in Breast Cancer and its Relation to Epigenomic Subtypes and HER2 Overexpression. Biomedicines. 2020; 8(5):116. https://doi.org/10.3390/biomedicines8050116
Chicago/Turabian StyleSimonova, Olga A., Ekaterina B. Kuznetsova, Alexander S. Tanas, Viktoria V. Rudenko, Elena V. Poddubskaya, Tatiana V. Kekeeva, Ivan D. Trotsenko, Sergey S. Larin, Sergei I. Kutsev, Dmitry V. Zaletaev, and et al. 2020. "Abnormal Hypermethylation of CpG Dinucleotides in Promoter Regions of Matrix Metalloproteinases Genes in Breast Cancer and its Relation to Epigenomic Subtypes and HER2 Overexpression" Biomedicines 8, no. 5: 116. https://doi.org/10.3390/biomedicines8050116
APA StyleSimonova, O. A., Kuznetsova, E. B., Tanas, A. S., Rudenko, V. V., Poddubskaya, E. V., Kekeeva, T. V., Trotsenko, I. D., Larin, S. S., Kutsev, S. I., Zaletaev, D. V., Nemtsova, M. V., & Strelnikov, V. V. (2020). Abnormal Hypermethylation of CpG Dinucleotides in Promoter Regions of Matrix Metalloproteinases Genes in Breast Cancer and its Relation to Epigenomic Subtypes and HER2 Overexpression. Biomedicines, 8(5), 116. https://doi.org/10.3390/biomedicines8050116





