Aptamer-Based Electrochemical Sensing of Lysozyme
Abstract
:1. Introduction
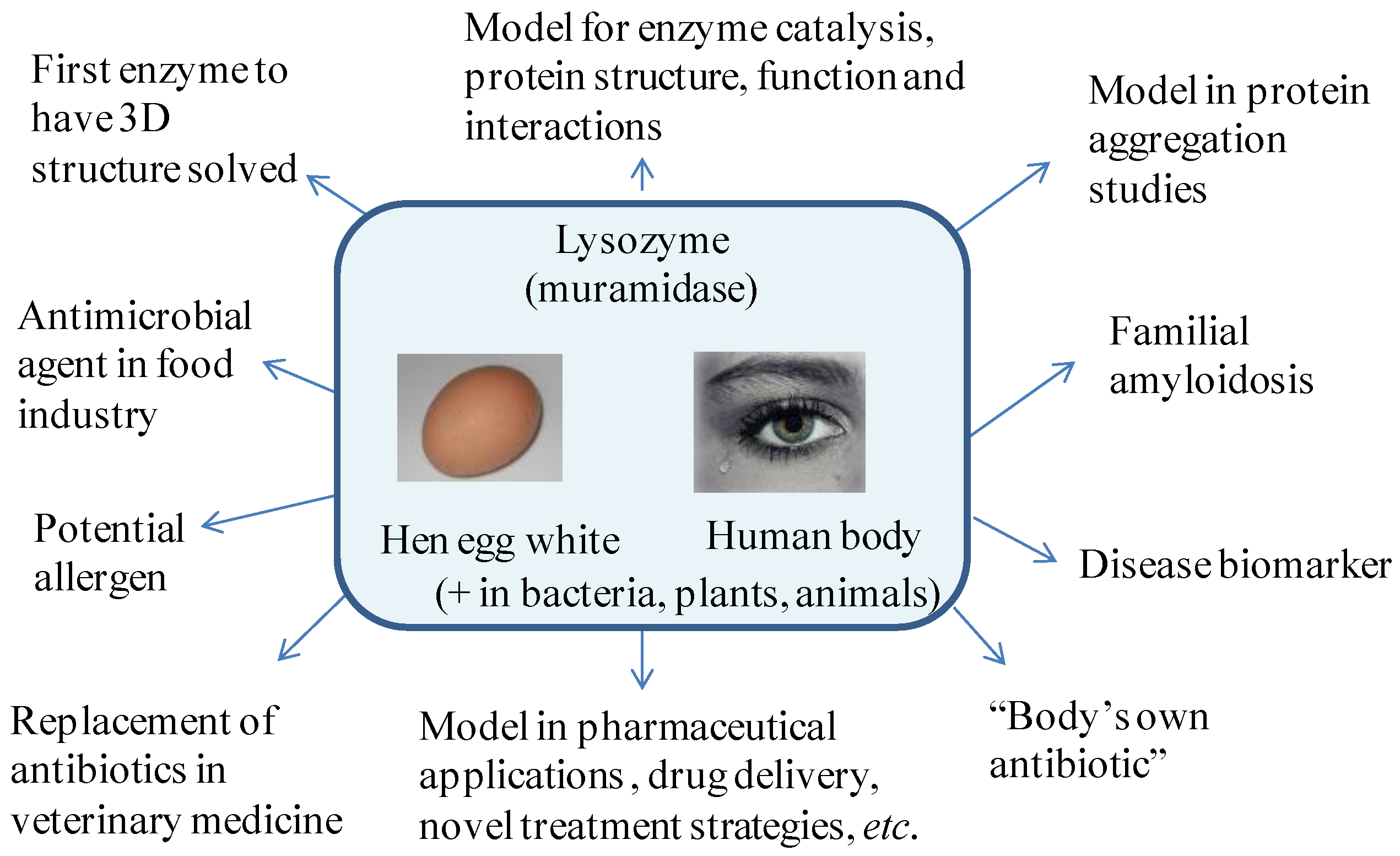
1.1. Properties of Lysozyme and Its Importance for Daily Life
1.2. Quantification Methods
2. Aptamer-Based Electrochemical Lysozyme Sensors
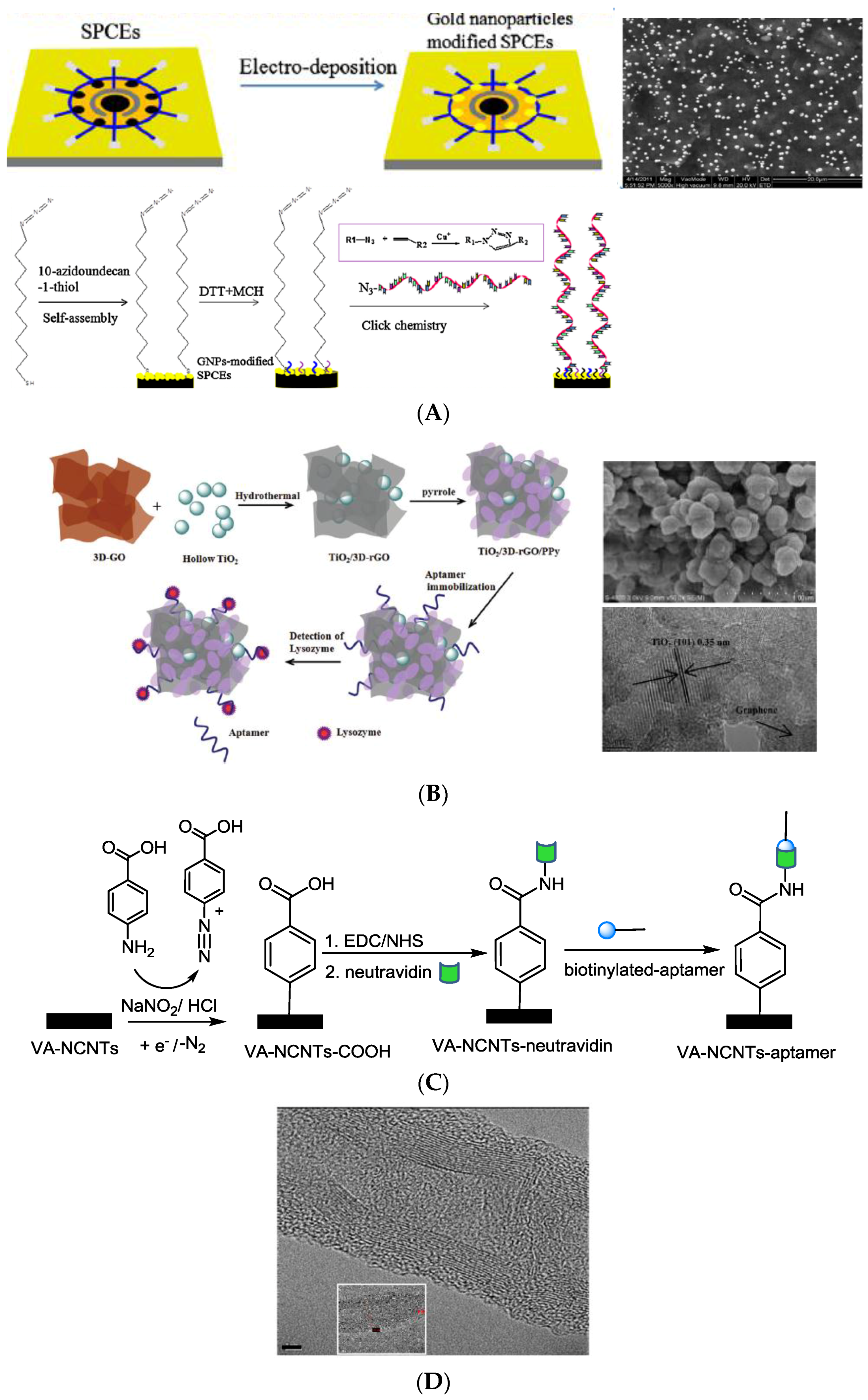
2.1. Surface Immobilization of Aptamer Ligands
- electrostatic interactions between the negatively-charged phosphate backbone of the aptamer and positively-charged materials (Figure 2E), such as polypyrrole, Fe2O3 and ferrocene-appended poly(ethyleneimine) (Fc-PEI) in a layer-by-layer approach or amine-rich films of plasma-polymerized propargylamine in a Cu2O@rGO@PpPG-modified gold electrode [74,78,84,88]
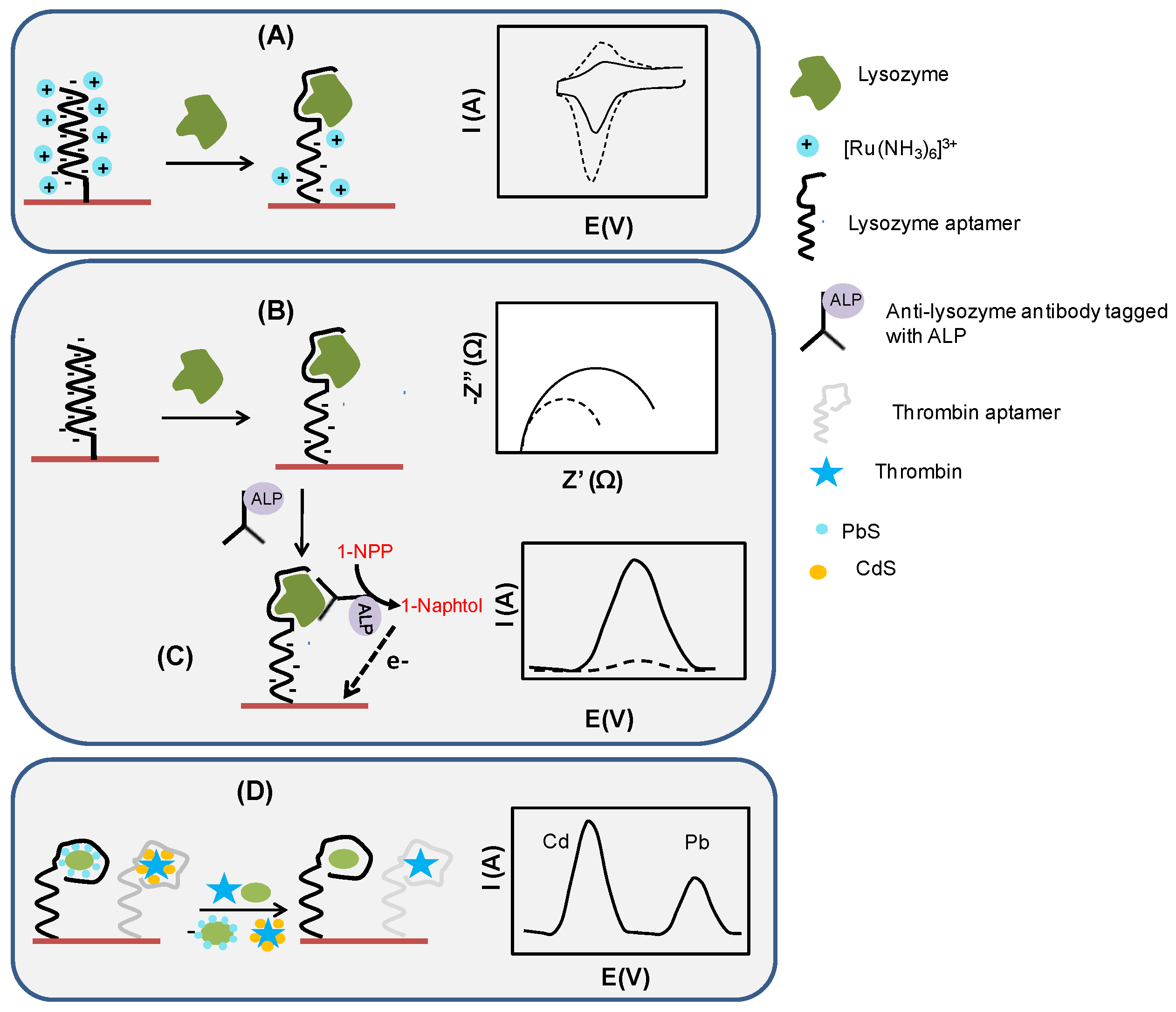
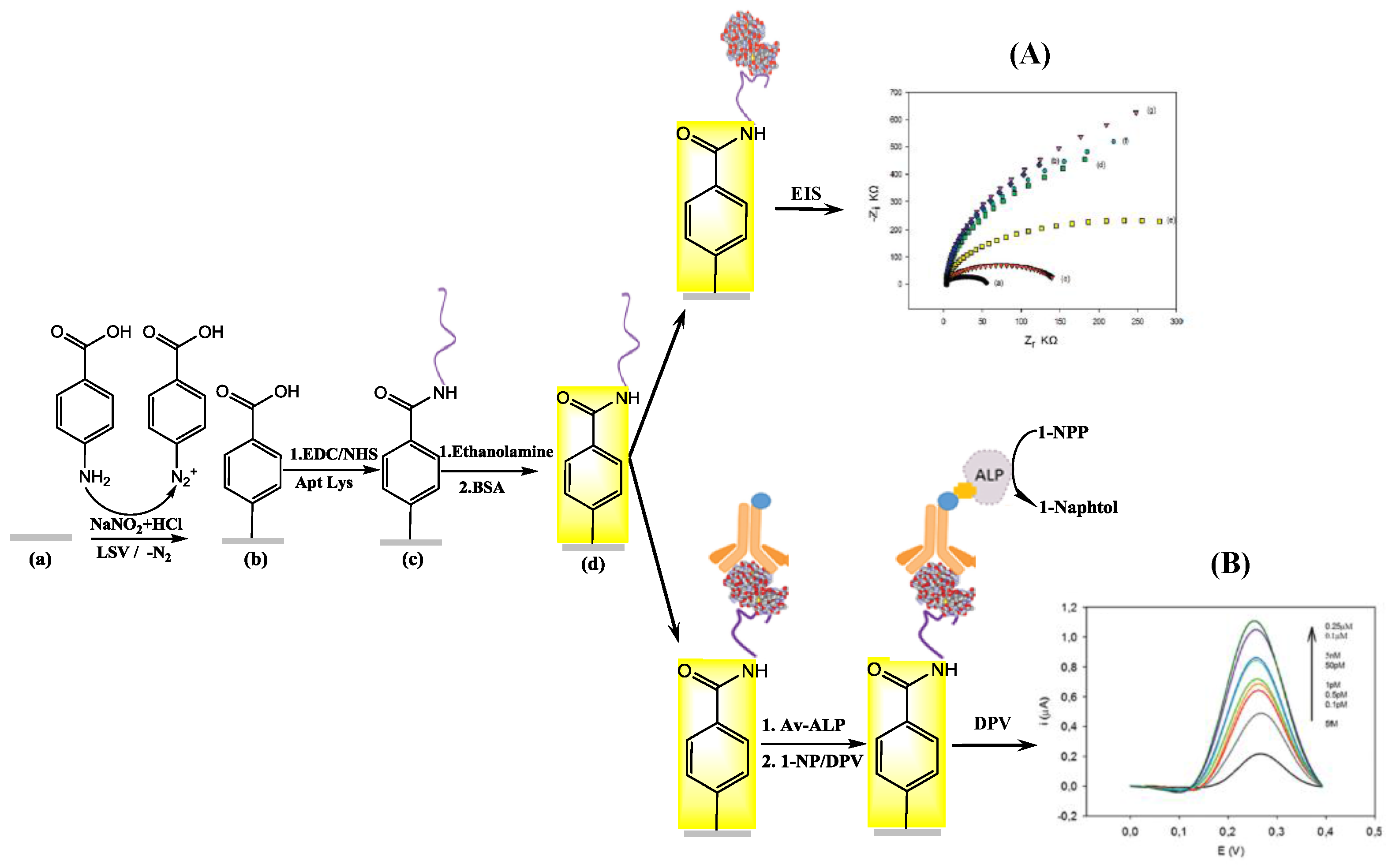
2.2. Electrochemical Assay Formats: Direct, Sandwich and Competitive Assays
- displacement of dabcyl and metallic NPs-labeled lysozyme binding aptamers forming a host-guest complex with cyclodextrin in the presence of lysozyme and subsequent release of NPs in solution [89]
- displacement of LBA from its methylene blue-tagged DNA complex in the presence of lysozyme, resulting in a conformational change of methylene blue-tagged DNA into a hairpin structure; this brings methylene blue closer to the electrode surface, leading to an increase of its signal (signal-on sensor) [90]
- electrochemical stripping of lysozyme/quantum-dots complex [94]
- desorption of lysozyme aptamer from rGO/Orange II-modified GCE, reversing the blocking effect and reestablishing efficient electron transfer from graphene-adsorbed aromatic dye Orange II [85]
2.3. Evaluation of Selectivity, Reproducibility and Storage Stability
2.4. Comparison of Electrochemical Lysozyme Sensors to Other Detection Schemes
3. Applications of Current Electrochemical Aptasensors for Lysozyme Sensing
4. Conclusions and Perspectives
- (i)
- Sensor designs and use of materials compatible with large-scale manufacturing technologies for producing commercial aptasensors. The good analytical characteristics and reproducibility of lysozyme aptasensors produced by manual, multiple step procedures is promising. Several types of electrodes modified with proteins, mediators and nanomaterials, produced by screen-printing and ink-jet printing, are already available commercially and could be used as a generic basis for lysozyme and other aptasensors;
- (ii)
- Experimental confirmation of the appropriate storage stability of the aptasensors for commercial purposes. Aptamers are inherently more stable compared to antibodies, for example; however, with the lysozyme aptasensors developed so far, storage stability beyond one month remains to be investigated;
- (iii)
- Generic approaches appropriate for high throughput, multi-analyte testing. Lysozyme analysis might prove highly beneficial in the context of the multiplexed sensing of various disease biomarkers. Going in this direction, electrochemical aptasensors have been developed for dual detection of lysozyme and interferon gamma, aiming to diagnosis acute leukemia [90]. An illustration of the potential of generic platforms was provided by an aptasensor array based on eight screen-printed electrodes modified with AuNPs, coated with azide-ended thiols, onto which three different aptamers (for lysozyme, cocaine and thrombin) were immobilized by click-chemistry [60]. Reconciling the need for a short analysis time with the simultaneous demand for a high sensitivity of detection could come from new signal amplification strategies. Among others, recent approaches based on nanomaterials, such as graphene [105] or nanoceria [106], show promising potentialities;
- (iv)
- Validation of novel aptasensors in comparison with methods currently used in clinical and analytical laboratories, such as ELISA and HPLC. So far, only three studies reported comparative results obtained with the aptasensor and by classical methods [73,78,91]. In the particular case of lysozyme, a comparison with other methods should be made with caution, since some methods measure the amount of enzymatically-active lysozyme, while others determine the total amount of protein. Moreover, differences between results provided by methods based on very different principles, e.g., chromatographic separation and affinity, are not uncommon [107].
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| AuNPs | gold nanoparticles |
| B-AB | biotinylated antibody |
| BiDNA | bifunctional aptamer for adenosine and lysozyme, linker DNA |
| CD | cyclodextrin |
| CPE | carbon paste electrode |
| CPSA | chronopotentiometric stripping analysis |
| Cu2O@rGO@PpPG | nanocomposite of reduced graphene oxide, cuprous oxide and plasma-polymerized propargylamine |
| CV | cyclic voltammetry |
| DLAP | dabcyl-labeled aptamer modified metal nanoparticles |
| DPASV | differential pulse adsorptive stripping voltammetry |
| DPV | differential pulse voltammetry |
| DTT | dithiothreitol |
| EDC | 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide |
| EIS | electrochemical impedance spectroscopy |
| Fc | ferrocene |
| FLD | fluorescence detector |
| GCE | glassy carbon electrode |
| GO | graphene oxide |
| GR | graphene |
| IDA–Cu/AuNps/GCE | iminodiacetic acid–copper ion complex immobilized on a glassy carbon electrode modified with gold nanoparticles |
| ITO | indium tin oxide |
| IFN-γ | interferon gamma |
| LBA | lysozyme binding aptamer |
| MCH | mercaptohexanol |
| MeB-cDNA | methylene blue-tagged complementary DNA |
| MWCNTs-CS | multiwalled carbon nanotubes-chitosan nanocomposites |
| NHS | N-hydroxysuccinimide |
| O-GNs | Orange II functionalized graphene nanosheets |
| p–ATP | p-aminothiophenol |
| PABA | poly-aminobenzoic acid |
| PEI | polyethyleneimine |
| PGE | pencil graphite electrode |
| RLS | resonance light scattering |
| SA-ALP | streptavidin-conjugate of alkaline phosphatase |
| SERS | surface-enhanced Raman scattering |
| SPCE | screen-printed carbon electrode |
| SWV | square wave voltammetry |
| TBA | thrombin binding aptamer |
| TCA/AuNP/ssDNA | thiocyanuric acid (TCA)/gold nanoparticles (AuNPs) modified with ssDNA |
| (THH) Au NCs | tetrahexahedral gold nanocrystals |
| TiO2@PPAA | composite made of polyacrylic acid and hollow TiO2 spheres |
| TiO2/3D-rGO/PPy | hollow titanium dioxide nanoball, three-dimensional reduced graphene oxide and polypyrrole |
| TPA | tripropylamine |
| TWJ | three-way junction |
| VANCNT | vertically-aligned nitrogen-doped carbon nanotubes |
References
- Laschtschenko, P. Uber die keimtötende und entwicklungshemmende Wirkung von Hühnereiweiß. Z. Hyg. Infekt. Krankh. 1909, 64, 419–427. [Google Scholar] [CrossRef]
- Fleming, A. On a Remarkable Bacteriolytic Element Found in Tissues and Secretions. Proc. R. Soc. B Biol. Sci. 1922, 93, 306–317. [Google Scholar] [CrossRef]
- Jollès, P. Recent Developments in the Study of Lysozymes. Angew. Chem. Int. Ed. Engl. 1964, 3, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, F.E.; Proctor, V.A.; Goetsch, S.J. Egg-white lysozyme as a food preservative: an overview. World’s Poult. Sci. J. 1991, 47, 141–163. [Google Scholar] [CrossRef]
- Callewaert, L.; Michiels, C.W. Lysozymes in the animal kingdom. J. Biosci. 2010, 35, 127–160. [Google Scholar] [CrossRef] [PubMed]
- Jolles, P.; Jolles, J. What‘s new in lysozyme research? Always a model system, today as yesterday. Mol. Cell Biochem. 1984, 63, 165–189. [Google Scholar] [PubMed]
- Blake, C.C.F.; Koenig, D.F.; Mair, G.A.; North, A.C.T.; Phillips, D.C.; Sarma, V.R. Structure of Hen Egg-White Lysozyme: A Three-dimensional Fourier Synthesis at 2 Å Resolution. Nature 1965, 206, 757–761. [Google Scholar] [CrossRef] [PubMed]
- Blake, C.C.; Mair, G.A.; North, A.C.; Phillips, D.C.; Sarma, V.R. On the conformation of the hen egg-white lysozyme molecule. Proc. R. Soc. Lond. B Biol. Sci. 1967, 167, 365–377. [Google Scholar] [CrossRef] [PubMed]
- Miranker, A.; Radford, S.E.; Karplus, M.; Dobson, C.M. Demonstration by NMR of folding domains in lysozyme. Nature 1991, 349, 633–636. [Google Scholar] [CrossRef] [PubMed]
- Dobson, C.M.; Evans, P.A.; Radford, S.E. Understanding how proteins fold: The lysozyme story so far. Trends Biochem. Sci. 1994, 19, 31–37. [Google Scholar] [CrossRef]
- Merlini, G.; Bellotti, V. Lysozyme: a paradigmatic molecule for the investigation of protein structure, function and misfolding. Clin. Chim. Acta. 2005, 357, 168–172. [Google Scholar] [CrossRef] [PubMed]
- Goda, S.; Takano, K.; Yamagata, Y.; Nagata, R.; Akutsu, H.; Maki, S.; Namba, K.; Yutani, K. Amyloid protofilament formation of hen egg lysozyme in highly concentrated ethanol solution. Protein Sci. 2000, 9, 369–375. [Google Scholar] [CrossRef] [PubMed]
- Booth, D.R.; Sunde, M.; Bellotti, V.; Robinson, C.V.; Hutchinson, W.L.; Fraser, P.E.; Hawkins, P.N.; Dobson, C.M.; Radford, S.E.; Blake, C.C.; et al. Instability, unfolding and aggregation of human lysozyme variants underlying amyloid fibrillogenesis. Nature 1997, 385, 787–793. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, R.; Ravi, V.K.; Kumar, S.; Kumar, M.V.; Chandra, N. Lysozyme: A model protein for amyloid research. Adv. Protein Chem. Struct. Biol. 2011, 84, 63–111. [Google Scholar] [PubMed]
- Dumoulin, M.; Last, A.M.; Desmyter, A.; Decanniere, K.; Canet, D.; Larsson, G.; Spencer, A.; Archer, D.B.; Sasse, J.; Muyldermans, S.; et al. A camelid antibody fragment inhibits the formation of amyloid fibrils by human lysozyme. Nature 2003, 424, 783–788. [Google Scholar] [CrossRef] [PubMed]
- Haas, M.; Moolenaar, F.; Meijer, D.K.; de Zeeuw, D. Specific drug delivery to the kidney. Cardiovasc. Drugs Ther. 2002, 16, 489–496. [Google Scholar] [CrossRef] [PubMed]
- Dolman, M.E.; Harmsen, S.; Storm, G.; Hennink, W.E.; Kok, R.J. Drug targeting to the kidney: Advances in the active targeting of therapeutics to proximal tubular cells. Adv. Drug Deliv. Rev. 2010, 62, 1344–1357. [Google Scholar] [CrossRef] [PubMed]
- Wasserfall, F.; Teuber, M. Action of egg white lysozyme on Clostridium tyrobutyricum. Appl. Environ. Microbiol. 1979, 38, 197–199. [Google Scholar] [PubMed]
- Masschalck, B.; Michiels, C.W. Antimicrobial properties of lysozyme in relation to foodborne vegetative bacteria. Crit. Rev. Microbiol. 2003, 29, 191–214. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Han, S.-Y.; Zhang, B.; Li, M.; Sheng, W.-J. Development of lysozyme-combined antibacterial system to reduce sulfur dioxide and to stabilize Italian Riesling ice wine during aging process. Food Sci. Nutr. 2015, 3, 453–465. [Google Scholar] [CrossRef] [PubMed]
- Peñas, E.; di Lorenzo, C.; Uberti, F.; Restani, P. Allergenic Proteins in Enology: A Review on Technological Applications and Safety Aspects. Molecules 2015, 20, 13144–13164. [Google Scholar] [CrossRef] [PubMed]
- Weber, P.; Steinhart, H.; Paschke, A. Investigation of the allergenic potential of wines fined with various proteinogenic fining agents by ELISA. J. Agric. Food Chem. 2007, 55, 3127–3133. [Google Scholar] [CrossRef] [PubMed]
- Weber, P.; Kratzin, H.; Brockow, K.; Ring, J.; Steinhart, H.; Paschke, A. Lysozyme in wine: A risk evaluation for consumers allergic to hen‘s egg. Mol. Nutr. Food Res. 2009, 53, 1469–1477. [Google Scholar] [CrossRef] [PubMed]
- Schneider, N.; Werkmeister, K.; Becker, C.M.; Pischetsrieder, M. Prevalence and stability of lysozyme in cheese. Food Chem. 2011, 128, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Silvetti, T.; Brasca, M.; Lodi, R.; Vanoni, L.; Chiolerio, F.; Groot, M.; Bravi, A. Effects of Lysozyme on the Microbiological Stability and Organoleptic Properties of Unpasteurized Beer. J. Inst. Brew. 2010, 116, 33–40. [Google Scholar] [CrossRef]
- Zimoch-Korzycka, A.; Jarmoluk, A. The use of chitosan, lysozyme, and the nano-silver as antimicrobial ingredients of edible protective hydrosols applied into the surface of meat. J. Food Sci. Technol. 2015, 52, 5996–6002. [Google Scholar] [CrossRef] [PubMed]
- Chander, R.; Lewis, N.F. Effect of lysozyme and sodium EDTA on shrimp microflora. Eur. J. Appl. Microbiol. Biotechnol. 1980, 10, 253–258. [Google Scholar] [CrossRef]
- Humphrey, B.D.; Huang, N.; Klasing, K.C. Rice expressing lactoferrin and lysozyme has antibiotic-like properties when fed to chicks. J. Nutr. 2002, 132, 1214–1218. [Google Scholar] [PubMed]
- Oliver, W.T.; Wells, J.E. Lysozyme as an alternative to antibiotics improves growth performance and small intestinal morphology in nursery pigs. J. Anim. Sci. 2013, 91, 3129–3136. [Google Scholar] [CrossRef] [PubMed]
- Proctor, V.A.; Cunningham, F.E.; Fung, D.Y.C. The chemistry of lysozyme and its use as a food preservative and a pharmaceutical. CRC Crit. Rev. Food Sci. Nutr. 1988, 26, 359–395. [Google Scholar] [CrossRef] [PubMed]
- Osserman, E.F.; Lawlor, D.P. Serum and urinary lysozyme (muramidase) in monocytic and monomyelocytic leukemia. J. Exp. Med. 1966, 124, 921–952. [Google Scholar] [CrossRef] [PubMed]
- Immundiagnostik AG. Manual-Lysozyme ELISA for the in vitro Determination of Lysozyme in Serum, Urine and Liquor. Avaiable online: www.immundiagnostik.com/fileadmin/pdf/LYSOZYM_Serum%20Urin%20Liq_K6902.pdf (accessed on 22 February 2016).
- Porstmann, B.; Jung, K.; Schmechta, H.; Evers, U.; Pergande, M.; Porstmann, T.; Kramm, H.J.; Krause, H. Measurement of lysozyme in human body fluids: Comparison of various enzyme immunoassay techniques and their diagnostic application. Clin. Biochem. 1989, 22, 349–355. [Google Scholar] [CrossRef]
- Lonnerdal, B. Biochemistry and physiological function of human milk proteins. Am. J. Clin. Nutr. 1985, 42, 1299–1317. [Google Scholar] [PubMed]
- Hankiewicz, J.; Swierczek, E. Lysozyme in human body fluids. Clin. Chim. Acta 1974, 57, 205–209. [Google Scholar] [CrossRef]
- Venge, P.; Foucard, T.; Henriksen, J.; Hakansson, L.; Kreuger, A. Serum-levels of lactoferrin, lysozyme and myeloperoxidase in normal, infection-prone and leukemic children. Clin. Chim. Acta 1984, 136, 121–130. [Google Scholar] [CrossRef]
- Grieco, M.H.; Reddy, M.M.; Kothari, H.B.; Lange, M.; Buimovici-Klein, E.; William, D. Elevated beta 2-microglobulin and lysozyme levels in patients with acquired immune deficiency syndrome. Clin. Immunol. Immunopathol. 1984, 32, 174–184. [Google Scholar] [CrossRef]
- Perillie, P.E.; Khan, K.; Finch, S.C. Serum lysozyme in pulmonary tuberculosis. Am. J. Med. Sci. 1973, 265, 297–302. [Google Scholar] [CrossRef] [PubMed]
- Pascual, R.S.; Gee, J.B.; Finch, S.C. Usefulness of serum lysozyme measurement in diagnosis and evaluation of sarcoidosis. N. Engl. J. Med. 1973, 289, 1074–1076. [Google Scholar] [CrossRef] [PubMed]
- Syrjanen, S.M.; Syrjanen, K.J. Lysozyme in the labial salivary glands of patients with rheumatoid arthritis. Z. Rheumatol. 1983, 42, 332–336. [Google Scholar] [PubMed]
- Falchuk, K.R.; Perrotto, J.L.; Isselbacher, K.J. Serum lysozyme in Crohn‘s disease. A useful index of disease activity. Gastroenterology 1975, 69, 893–896. [Google Scholar] [PubMed]
- Levinson, S.S.; Elin, R.J.; Yam, L. Light chain proteinuria and lysozymuria in a patient with acute monocytic leukemia. Clin. Chem. 2002, 48, 1131–1132. [Google Scholar] [PubMed]
- Guder, W.G.; Hofmann, W. Clinical role of urinary low molecular weight proteins: their diagnostic and prognostic implications. Scand. J. Clin. Lab Investig. Suppl. 2008, 241, 95–98. [Google Scholar] [CrossRef] [PubMed]
- Ocaña, C.; Hayat, A.; Mishra, R.; Vasilescu, A.; del Valle, M.; Marty, J.L. A novel electrochemical aptamer-antibody sandwich assay for lysozyme detection. Analyst 2015, 140, 4148–4153. [Google Scholar] [CrossRef] [PubMed]
- Hun, X.; Chen, H.; Wang, W. Design of Ultrasensitive Chemiluminescence Detection of Lysozyme in Cancer Cells Based on Nicking Endonuclease Signal Amplification Technology. Biosens. Bioelectron. 2010, 26, 248–254. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Li, L.; Tian, Y.; Mu, X.; Guo, L. Signal amplification architecture for electrochemical aptasensor based on network-like thiocyanuric acid/gold nanoparticle/ssDNA. Biosens. Bioelectron. 2012, 38, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Li, L.-D.; Chen, Z.-B.; Zhao, H.-Z.; Guo, L.; Mu, X. An Aptamer-Based Biosensor for the Detection of Lysozyme With Gold Nanoparticles Amplification. Sens. Actuators B Chem. 2010, 149, 110–115. [Google Scholar] [CrossRef]
- Vasilescu, A.; Gaspar, S.; Gheorghiu, M.; David, S.; Dinca, V.; Peteu, S.; Wang, Q.; Li, M.; Boukherroub, R.; Szunerits, S. Surface Plasmon Resonance based sensing of lysozyme in serum on Micrococcus lysodeikticus-modified graphene oxide surfaces. Biosens. Bioelectron. 2016, in press. [Google Scholar] [CrossRef] [PubMed]
- Weth, F.; Schroeder, T.; Buxtorf, U.P. Determination of lysozyme content in eggs and egg products using SDS-gel electrophoresis. Z. Lebensm. Unters. Forsch. 1988, 187, 541–545. [Google Scholar] [CrossRef] [PubMed]
- Galyean, R.D.; Cotterill, O.J. Ion-Exchange Chromatographic Determination of Lysozyme in Egg White. J. Food Sci. 1981, 46, 1827–1834. [Google Scholar] [CrossRef]
- Daeschel, M.A.; Musafija-Jeknic, T.; Wu, Y.; Bizzarri, D.; Villa, A. High-Performance Liquid Chromatography Analysis of Lysozyme in Wine. Am. J. Enol. Vitic. 2002, 53, 154–157. [Google Scholar]
- Francina, A.; Cloppet, H.; Guinet, R.; Rossi, M.; Guyotat, D.; Gentilhomme, O.; Richard, M. A rapid and sensitive non-competitive avidin-biotin immuno-enzymatic assay for lysozyme. J. Immunol. Methods 1986, 87, 267–272. [Google Scholar] [CrossRef]
- Vidal, M.L.; Gautron, J.; Nys, Y. Development of an ELISA for quantifying lysozyme in hen egg white. J. Agric. Food Chem. 2005, 53, 2379–2385. [Google Scholar] [CrossRef] [PubMed]
- Lacorn, M.G.C.; Haas-Lauterbach, S.; Immer, U. Sensitive lysozyme testing in red and white wine using the RIDASCREEN FAST Lysozyme ELISA. Bulletin de l'OIV 2010, 83, 507–511. [Google Scholar]
- Subramanian, P.; Lesniewski, A.; Kaminska, I.; Vlandas, A.; Vasilescu, A.; Niedziolka-Jonsson, J.; Pichonat, E.; Happy, H.; Boukherroub, R.; Szunerits, S. Lysozyme detection on aptamer functionalized graphene-coated SPR interfaces. Biosens. Bioelectron. 2013, 50, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Mascini, M.; Palchetti, I.; Tombelli, S. Nucleic acid and peptide aptamers: fundamentals and bioanalytical aspects. Angew. Chem. Int. Ed. Engl. 2012, 51, 1316–1332. [Google Scholar] [CrossRef] [PubMed]
- Sener, G.; Uzun, L.; Say, R.; Denizli, A. Use of molecular imprinted nanoparticles as biorecognition element on surface plasmon resonance sensor. Sens. Actuators B Chem. 2011, 160, 791–799. [Google Scholar] [CrossRef]
- Wang, L.; Zhu, C.; Han, L.; Jin, L.; Zhou, M.; Dong, S. Label-free, regenerative and sensitive surface plasmon resonance and electrochemical aptasensors based on graphene. Chem. Commun. 2011, 47, 7794–7796. [Google Scholar] [CrossRef] [PubMed]
- Mihai, I.; Vezeanu, A.; Polonschii, C.; Albu, C.; Radu, G.-L.; Vasilescu, A. Label-free detection of lysozyme in wines using an aptamer based biosensor and SPR detection. Sens. Actuators B Chem. 2015, 206, 198–204. [Google Scholar] [CrossRef]
- Xie, D.; Li, C.; Shangguan, L.; Qi, H.; Xue, D.; Gao, Q.; Zhang, C. Click chemistry-assisted self-assembly of DNA aptamer on gold nanoparticles-modified screen-printed carbon electrodes for label-free electrochemical aptasensor. Sens. Actuators B Chem. 2014, 192, 558–564. [Google Scholar] [CrossRef]
- Erdem, A.; Eksin, E.; Muti, M. Chitosan-graphene oxide based aptasensor for the impedimetric detection of lysozyme. Colloids Surf. B Biointerfaces 2014, 115, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Zhang, D.; Li, Y.; Qi, H.; Gao, Q.; Zhang, C. Label-free and sensitive faradic impedance aptasensor for the determination of lysozyme based on target-induced aptamer displacement. Biosens. Bioelectron. 2009, 25, 94–99. [Google Scholar] [CrossRef] [PubMed]
- Ocaña, C.; Hayat, A.; Mishra, R.K.; Vasilescu, A.; Del Valle, M.; Marty, J.L. Label free aptasensor for Lysozyme detection: A comparison of the analytical performance of two aptamers. Bioelectrochemistry 2015, 105, 72–77. [Google Scholar] [CrossRef] [PubMed]
- Rajendran, M.; Ellington, A.D. Selection of fluorescent aptamer beacons that light up in the presence of zinc. Anal. Bioanal. Chem. 2008, 390, 1067–1075. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Li, Y.; Tang, Z.; Cao, Z.C.; Chen, H.W.; Mallikaratchy, P.; Sefah, K.; Yang, C.J.; Tan, W. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc. Natl. Acad. Sci. USA 2006, 103, 11838–11843. [Google Scholar] [CrossRef] [PubMed]
- Bunka, D.H.; Stockley, P.G. Aptamers come of age at last. Nat. Rev. Microbiol. 2006, 4, 588–596. [Google Scholar] [CrossRef] [PubMed]
- Iliuk, A.B.; Hu, L.; Tao, W.A. Aptamer in bioanalytical applications. Anal. Chem. 2011, 83, 4440–4452. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.C.; Ellington, A.D. Automated selection of anti-protein aptamers. Bioorg. Med. Chem. 2001, 9, 2525–2531. [Google Scholar] [CrossRef]
- Cox, J.C.; Hayhurst, A.; Hesselberth, J.; Bayer, T.S.; Georgiou, G.; Ellington, A.D. Automated selection of aptamers against protein targets translated in vitro: From gene to aptamer. Nucleic. Acids Res. 2002, 30, e108. [Google Scholar] [CrossRef] [PubMed]
- Kirby, R.; Cho, E.J.; Gehrke, B.; Bayer, T.; Park, Y.S.; Neikirk, D.P.; McDevitt, J.T.; Ellington, A.D. Aptamer-based sensor arrays for the detection and quantitation of proteins. Anal. Chem. 2004, 76, 4066–4075. [Google Scholar] [CrossRef] [PubMed]
- Tran, D.T.; Janssen, K.P.; Pollet, J.; Lammertyn, E.; Anne, J.; Van Schepdael, A.; Lammertyn, J. Selection and characterization of DNA aptamers for egg white lysozyme. Molecules 2010, 15, 1127–1140. [Google Scholar] [CrossRef] [PubMed]
- Schoukroun-Barnes, L.R.; Glaser, E.P.; White, R.J. Heterogeneous Electrochemical Aptamer-Based Sensor Surfaces for Controlled Sensor Response. Langmuir 2015, 31, 6563–6569. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Subramanian, P.; Schlechter, A.; Teblum, E.; Yemini, R.; Nessim, G.D.; Vasilescu, A.; Li, M.; Boukherroub, R.; Szunerits, S. Vertically aligned nitrogen-doped carbon nanotube carpet electrodes: Highly sensitive interfaces for the analysis of serum from patients with inflammatory bowel disease. ACS Appl. Mater. Interfaces 2016, 8, 9600–9609. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Zhai, S.; Ye, Z.; He, L.; Peng, D.; Feng, X.; Yang, Y.; Fang, S.; Zhang, H.; Zhang, Z. An electrochemical aptasensor based on a TiO2/three-dimensional reduced graphene oxide/PPy nanocomposite for the sensitive detection of lysozyme. Dalton Trans. 2015, 44, 6473–6479. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhang, S.; He, L.; Peng, D.; Yan, F.; Wang, M.; Zhao, J.; Zhang, H.; Fang, S. Feasible electrochemical biosensor based on plasma polymerization-assisted composite of polyacrylic acid and hollow TiO2 spheres for sensitively detecting lysozyme. Biosens. Bioelectron. 2015, 74, 384–390. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Li, L.; Zhao, H.; Guo, L.; Mu, X. Electrochemical impedance spectroscopy detection of lysozyme based on electrodeposited gold nanoparticles. Talanta 2011, 83, 1501–1506. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Wang, Y.; Wu, M.; Ma, X.; Yang, X. Graphene-based lysozyme binding aptamer nanocomposite for label-free and sensitive lysozyme sensing. J. Electroanal. Chem. 2013, 702, 49–55. [Google Scholar] [CrossRef]
- Fang, S.; Dong, X.; Ji, H.; Liu, S.; Yan, F.; Peng, D.; He, L.; Wang, M.; Zhang, Z. Electrochemical aptasensor for lysozyme based on a gold electrode modified with a nanocomposite consisting of reduced graphene oxide, cuprous xide, and plasma-polymerized propargylamine. Microchim. Acta 2016, 183, 633–642. [Google Scholar] [CrossRef]
- Chen, Z.; Guo, J.; Li, J.; Guo, L. Tetrahexahedral Au nanocrystals/aptamer based ultrasensitive electrochemical biosensor. RSC Adv. 2013, 3, 14385–14389. [Google Scholar] [CrossRef]
- Rodriguez, M.C.; Kawde, A.-N.; Wang, J. Aptamer biosensor for label-free impedance spectroscopy detection of proteins based on recognition-induced switching of the surface charge. Chem. Commun. 2005, 34, 4267–4269. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, M.C.; Rivas, G.A. Label-free electrochemical aptasensor for the detection of lysozyme. Talanta 2009, 78, 212–216. [Google Scholar] [CrossRef] [PubMed]
- Rohrbach, F.; Karadeniz, H.; Erdem, A.; Famulok, M.; Mayer, G. Label-free impedimetric aptasensor for lysozyme detection based on carbon nanotube-modified screen-printed electrodes. Anal. Biochem. 2012, 421, 454–459. [Google Scholar] [CrossRef] [PubMed]
- Du, M.; Yang, T.; Zhao, C.; Jiao, K. Electrochemical logic aptasensor based on graphene. Sens. Actuators B Chem. 2012, 169, 255–260. [Google Scholar] [CrossRef]
- Du, M.; Yang, T.; Guo, X.; Zhong, L.; Jiao, K. Electrochemical synthesis of Fe2O3 on graphene matrix for indicator-free impedimetric aptasensing. Talanta 2013, 105, 229–234. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Han, Y.; Guo, Y.; Dong, C. Graphene-Orange II composite nanosheets with electroactive functions as label-free aptasensing platform for "signal-on" detection of protein. Biosens. Bioelectron. 2013, 45, 95–101. [Google Scholar] [CrossRef] [PubMed]
- Kawde, A.-N.; Rodriguez, M.C.; Lee, T.M.H.; Wang, J. Label-free bioelectronic detection of aptamer–protein interactions. Electrochem. Commun. 2005, 7, 537–540. [Google Scholar] [CrossRef]
- Cheng, A.K.; Ge, B.; Yu, H.Z. Aptamer-based biosensors for label-free voltammetric detection of lysozyme. Anal. Chem. 2007, 79, 5158–5164. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Chen, C.; Li, B.; Zhou, M.; Wang, E.; Dong, S. Layer-by-layer electrochemical biosensor with aptamer-appended active polyelectrolyte multilayer for sensitive protein determination. Biosens. Bioelectron. 2010, 25, 1902–1907. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Zhang, J.; Lin, Y.; Wang, Q.; Zhang, X.; Ding, Y.; Cui, H.; Fan, H. An electrochemical molecular recognition-based aptasensor for multiple protein detection. Anal. Biochem. 2015, 491, 31–36. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Song, D.; Wang, Z.; Zhang, F.; Yang, M.; Gui, R.; Xia, L.; Bi, S.; Xia, Y.; Li, Y.; et al. Single electrode biosensor for simultaneous determination of interferon gamma and lysozyme. Biosens. Bioelectron. 2015, 68, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Guo, J. A reagentless signal-off architecture for electrochemical aptasensor for the detection of lysozyme. Electrochim. Acta 2013, 111, 916–920. [Google Scholar] [CrossRef]
- Liu, D.Y.; Zhao, Y.; He, X.W.; Yin, X.B. Electrochemical aptasensor using the tripropylamine oxidation to probe intramolecular displacement between target and complementary nucleotide for protein array. Biosens. Bioelectron. 2011, 26, 2905–2910. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Fang, C.; Zhang, S. Ultrasensitive electrochemical analysis of two analytes by using an autonomous DNA machine that works in a two-cycle mode. Chemistry 2011, 17, 7531–7537. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J.A.; Wang, J.; Kawde, A.N.; Xiang, Y.; Gothelf, K.V.; Collins, G. Quantum-dot/aptamer-based ultrasensitive multi-analyte electrochemical biosensor. J. Am. Chem. Soc. 2006, 128, 2228–2229. [Google Scholar] [CrossRef] [PubMed]
- Deng, C.; Chen, J.; Nie, L.; Nie, Z.; Yao, S. Sensitive bifunctional aptamer-based electrochemical biosensor for small molecules and protein. Anal. Chem. 2009, 81, 9972–9978. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Gan, S.; Xu, Q.; Qiu, X.; Gao, P.; Huang, S. A three-way junction aptasensor for lysozyme detection. Biosens. Bioelectron. 2013, 39, 250–254. [Google Scholar] [CrossRef] [PubMed]
- Vasilescu, A.; Marty, J.-L. Electrochemical aptasensors for the assessment of food quality and safety. Trends Anal. Chem. 2016, 79, 60–70. [Google Scholar] [CrossRef]
- Arabzadeh, A.; Salimi, A. Novel voltammetric and impedimetric sensor for femtomolar determination of lysozyme based on metal-chelate affinity immobilized onto gold nanoparticles. Biosens. Bioelectron. 2015, 74, 270–276. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhu, S.; Xiong, Y.; Deng, C.; Zhang, X. Development of a MALDI-TOF MS strategy for the high-throughput analysis of biomarkers: On-target aptamer immobilization and laser-accelerated proteolysis. Angew. Chem. Int. Ed. Engl. 2013, 52, 6055–6058. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Cai, C.; Chen, X.; Chen, C. “Click on the bidirectional switch”: The aptasensor for simultaneous detection of lysozyme and ATP with high sensitivity and high selectivity. Sci. Rep. 2016, 6, 18814. [Google Scholar] [CrossRef] [PubMed]
- He, P.; Zhang, Y.; Liu, L.; Qiao, W.; Zhang, S. Ultrasensitive SERS Detection of Lysozyme by a Target-Triggering Multiple Cycle Amplification Strategy Based on a Gold Substrate. Chem. Eur. J. 2013, 19, 7452–7460. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.H.; Brown, M.B.; Martin, G.P. Turbidimetric and HPLC assays for the determination of formulated lysozyme activity. J. Pharm. Pharmacol. 2001, 53, 549–554. [Google Scholar] [CrossRef] [PubMed]
- Determination of Lysozyme in Wine Using High-Performance Liquid Chromatography. Available online: http://www.oiv.int/public/medias/2553/oiv-ma-as315–25-en.pdf (accessed on 22 March 2016).
- Guzzo, F.; Cappello, M.S.; Azzolini, M.; Tosi, E.; Zapparoli, G. The inhibitory effects of wine phenolics on lysozyme activity against lactic acid bacteria. Int. J. Food Microbiol. 2011, 148, 184–190. [Google Scholar] [CrossRef] [PubMed]
- Loo, A.H.; Bonanni, A.; Pumera, M. Mycotoxin aptasensing amplification by using inherently electroactive graphene-oxide nanoplatelet labels. ChemElectroChem 2015, 2, 743–747. [Google Scholar] [CrossRef]
- Bulbul, G.; Hayat, A.; Andreescu, S. A generic amplification strategy for electrochemical aptasensors using a non-enzymatic nanoceria tag. Nanoscale 2015, 7, 13230–13238. [Google Scholar] [CrossRef] [PubMed]
- Barbara Kerkaert, B.; Mestdagh, F.; De Meulenaer, B. Detection of hen‘s egg white lysozyme in food: Comparison between a sensitive HPLC and a commercial ELISA method. Food Chem. 2010, 120, 580–584. [Google Scholar] [CrossRef]

| DNA Sequence | Kd/nM | Reference |
|---|---|---|
| 5′-ATCAGGGCTAAAGAGTGCAGAGTTACTTAG-3′ | 31 | [68] |
| 5′-GGGAATGGATCCACATCTACGAATTCATCAGGGCTAAAGAG TGCAGAGTTACTTAGTTCACTGCAGACTTGACGAAGCTT-3′ | 29 ± 5 | [70] |
| 5′-GCAGCTAAGCAGGCGGCTCACAAAACCATTCGCATGCGGC-3′ | 2.8 ± 0.3 | [71] |
| Sample | Material | Method | LoD * | Linear Range | Comments | Reference |
|---|---|---|---|---|---|---|
| Direct Assays | ||||||
| Serum | VANCNT/NA/LBA | DPV | 100 fM | 0.1–7 pM | 2.5% decrease in signal after 2 weeks at 4 °C in buffer; RSD: 2.3% | [73] |
| Egg white | Au/TiO2/3D-rGO/PPy/LBA | DPV | 5.5 pM | 0.007–3.5 nM | 90% of initial signal after 1 month; RSD: 5.45% | [74] |
| Egg white | Au/TiO2@PPAA/LBA | EIS | 1.04 pM | 3.5 pM–7 nM | - | [75] |
| Egg white | SPCE/AuNPs/LBA | SWV | 21 fM | 0.07–3.4 pM | RSD: 4.2% | [60] |
| Egg white | Au/AuNPs/LBA | EIS | 0.01 pM | 0.1–500 pM | 84% of the original signal after 1 month in buffer at 4 °C; RSD: 2.11% (n = 3) | [76] |
| Wine | SPCE/LBA1 and LBA2 | EIS | 25 nM | 0.025–0.8 µM | Stable several days stored dry at 4 °C; RSD: <3.8% | [63] |
| Chicken egg + saliva | GCE/chitosan-GR/LBA | EIS | 6 fM | 0.01–0.5 pM | - | [77] |
| Saliva + urine + plasma | Au/Cu2O@rGO@PpPG | DPV | pM | 0.1–200 nM | 96.5% of initial activity after 15 days in buffer; RSD: 4.8% | [78] |
| Egg white + serum | GCE/THH Au NCs/APT | SWV | 0.1 pM | 0.1 pM–10 nM | 7.7% decrease in signal after storage in buffer at 4 °C for 23 days | [79] |
| N/A | PGE/chitosan–GO/LBA [1] | EIS | 28.53 nM | - | Stable 1 week at 4 °C; RSD% = 9.6% | [61] |
| N/A | ITO/PABA/SA/LBA | EIS | 14 nM | - | - | [80] |
| N/A | CPE/LBA | SWV | 18 nM (adenine) 36 nM (guanine) | 0.06–1.4 µM (adenine) (0.11–1.4 µM (guanine) | RSD: 5.1% (guanine) and 6.8% (adenine) | [81] |
| N/A | MWCNT–SPE/LBA | EIS | 862 nM | - | - | [82] |
| N/A | GR-GCE/LBA [2] | DPV | 0.08 nM | 0.2 nM–1040 nM | 4.55% decrease in signal after storage at 4 °C for 10 days; RSD: 4.23% | [83] |
| N/A | Fe2O3-GR-GCE/LBA | EIS | 11.1 pM | 35 pM–350 nM | 4.48% decrease in signal after storage at 4 °C for 10 days; RSD: 4.23% | [84] |
| N/A | GCE/O-GNs/LBA | DPV | 1 pM | 5.0 pM–0.7 nM | - | [85] |
| N/A | MB/LBA ** | CPSA | 7 nM | - | - | [86] |
| N/A | Au/LBA | CV | - | 35 nM–3.5 µM | - | [87] |
| N/A | ITO/(Fc-PEI/CNTs/Fc-PEI/LBA)3 | DPV | 11.8 pM | 13.9 pM–116 nM | 7.5% decrease after 24 days at room temperature in air; 2.25% increase after 21 days in distilled water at 4 °C | [88] |
| Sandwich Assay | ||||||
| Wine | SPCE/LBA/Lysozyme/B-AB/SA-ALP | DPV | 4.3 fM | 5 fM–5 nM | Stable 2 weeks at 4 °C; RSD: 5.5% | [44] |
| Competitive Assays | ||||||
| Serum | Au/CD/DLAP1 + DLAP2 | DPV | 64 pM | 100–1000 pM | - | [89] |
| Serum | Au/MeB-cDNA/LBA | SWV | 16.4 pM | 0.1–100 nM | Stable 3 weeks at 4 °C; RSD: <5% | [90] |
| Urine | Au/LBA-(DNA-Fc) | SWV | 0.45 nM | 7–30 nM | 7.7% decrease after storage in buffer at 4 °C; for 23 days | [91] |
| Egg white | Au/LBA/TCA/AuNPs/ cDNA | CV | 0.1 pM | 5 pM–1 nM | 84% of the original signal after one month at 4 °C; RSD: <4.3% (n = 5) | [46] |
| Egg white | Au/cDNA/LBA | LSW | 1 pM | 1.0 pM–1.1 nM | RSD < 4.2% (n = 5) | [92] |
| Ramos cancer cells | DNA machine, CdS NP–DNA/LBA ** | DPASV | 0.52 pM | 1 pM–80 nM | RSD < 6.1% (n = 3) | [93] |
| N/A | Au/TBA and LBA/(PbS-Lys and CdS-Thr) | SWV | - | 75% signal decrease for 0.07 nM | - | [94] |
| N/A | Au/DNA1/BiDNA/DNA3-AuNPs | CV | 0.7 nM | - | Stable for 2 weeks in distilled water at 4 °C RSD: 4.6% | [95] |
| N/A | Au/p-ATP-AuNPs/(LBA/Fc-cDNA) | SWV | 0.1 pM | 0.1 pM–1 nM | 15% decrease in original signal after 1-month in buffer solution | [47] |
| N/A | Au/cDNA/LBA | EIS | 70 pM | 0.2–4.0 nM | RSD: 3.7% | [62] |
| N/A | GCE/Au/(Fc-cDNA/LBA TWJ) | SWV | 0.2 nM | 0.2–100 nM | Ion and Ioff decreased by 7.9% and 18.5% after 2 weeks | [96] |
| Detection Method | Ligand | Limit of Detection | Reference |
|---|---|---|---|
| SPR | Aptamer | 0.5 nM | [55] |
| DPV | IDA–Cu complex | 60 fM | [98] |
| MALDI-TOF MS | Aptamer | 1 nM | [99] |
| RLS | Aptamer | 1 pM | [100] |
| ELISA | Antibody | 0.1 nM | [32] |
| SERS | Aptamer | 1 aM | [101] |
| Turbidimetry | Micrococcus lysodeikticus | 0.13 nM | [102] |
| HPLC-FLD | - | 10 nM | [103] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vasilescu, A.; Wang, Q.; Li, M.; Boukherroub, R.; Szunerits, S. Aptamer-Based Electrochemical Sensing of Lysozyme. Chemosensors 2016, 4, 10. https://doi.org/10.3390/chemosensors4020010
Vasilescu A, Wang Q, Li M, Boukherroub R, Szunerits S. Aptamer-Based Electrochemical Sensing of Lysozyme. Chemosensors. 2016; 4(2):10. https://doi.org/10.3390/chemosensors4020010
Chicago/Turabian StyleVasilescu, Alina, Qian Wang, Musen Li, Rabah Boukherroub, and Sabine Szunerits. 2016. "Aptamer-Based Electrochemical Sensing of Lysozyme" Chemosensors 4, no. 2: 10. https://doi.org/10.3390/chemosensors4020010
APA StyleVasilescu, A., Wang, Q., Li, M., Boukherroub, R., & Szunerits, S. (2016). Aptamer-Based Electrochemical Sensing of Lysozyme. Chemosensors, 4(2), 10. https://doi.org/10.3390/chemosensors4020010








