Identification of Critical Genes and Signaling Pathways in Human Monocytes Following High-Intensity Exercise
Abstract
1. Introduction
2. Materials and Methods
2.1. Microarray Data
2.2. Microarray Data Pre-Processing
2.3. Screening Differentially Expressed Genes and Hierarchical Clustering Analysis
2.4. Enrichment Analyses of Differentially Expressed Genes
2.5. PPI Network and Module Analysis
2.6. TF Target Regulation Prediction
3. Results
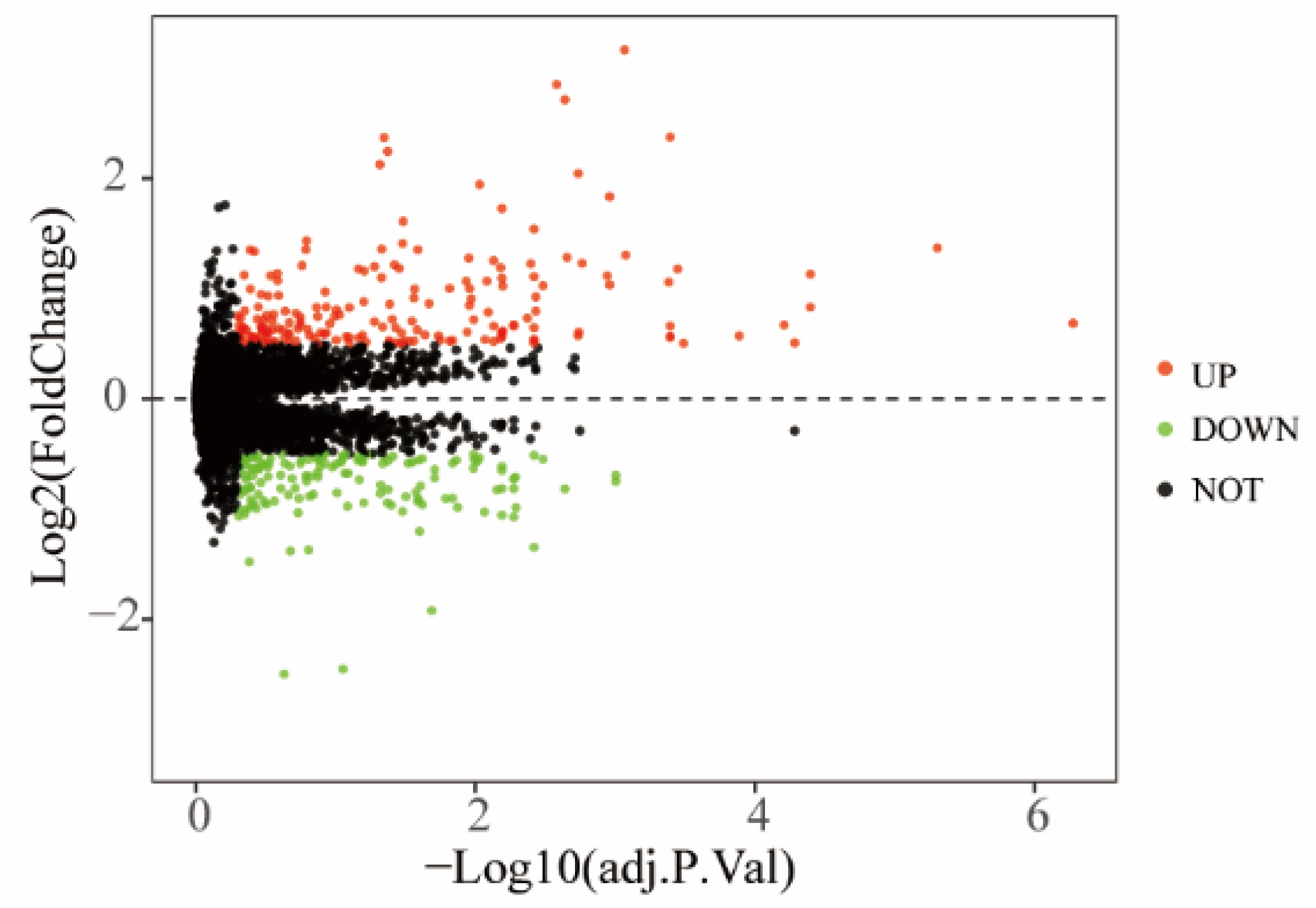
3.1. Normalization and DEGs Screening
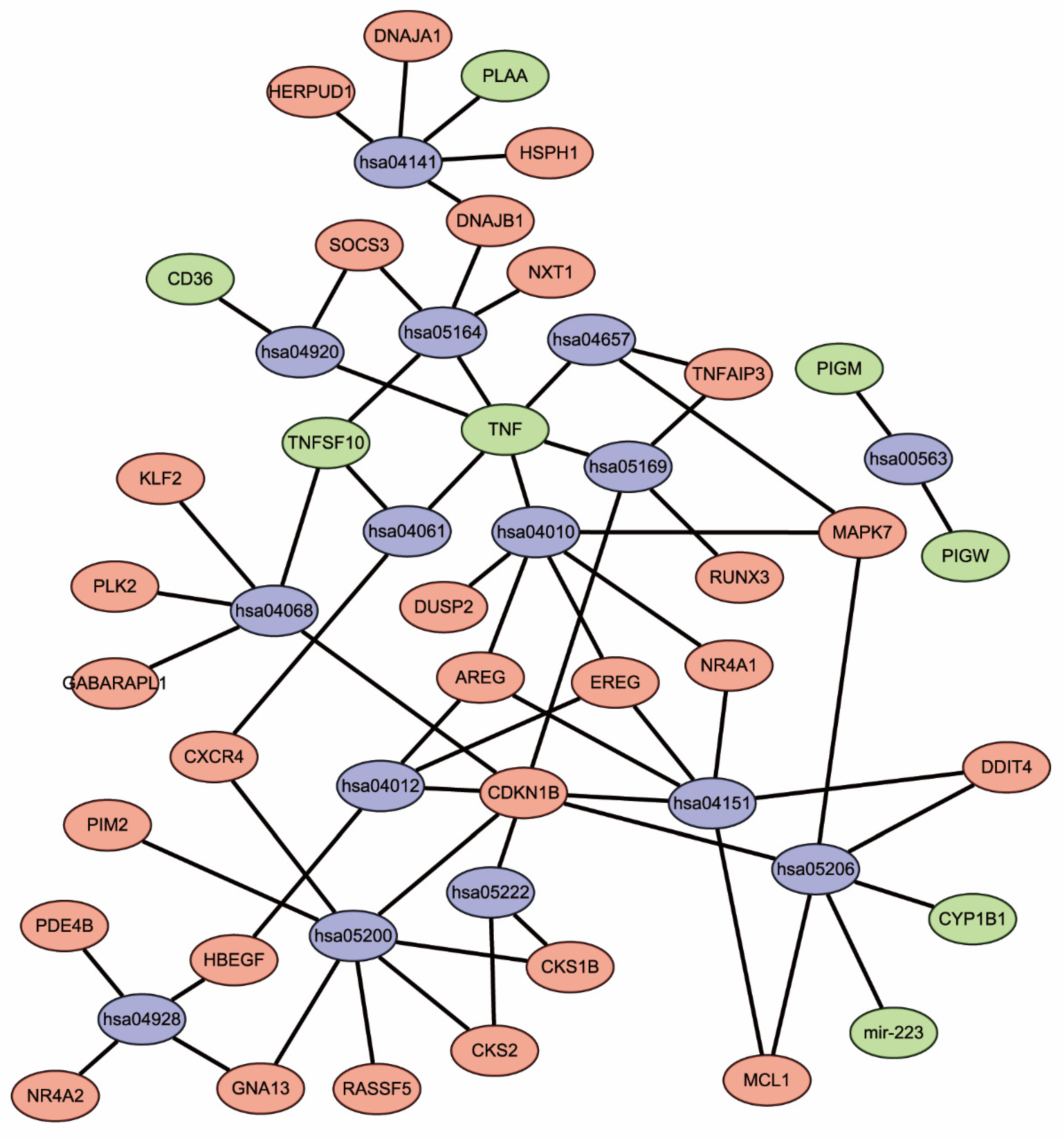
3.2. Enrichment Analyses of DEGs
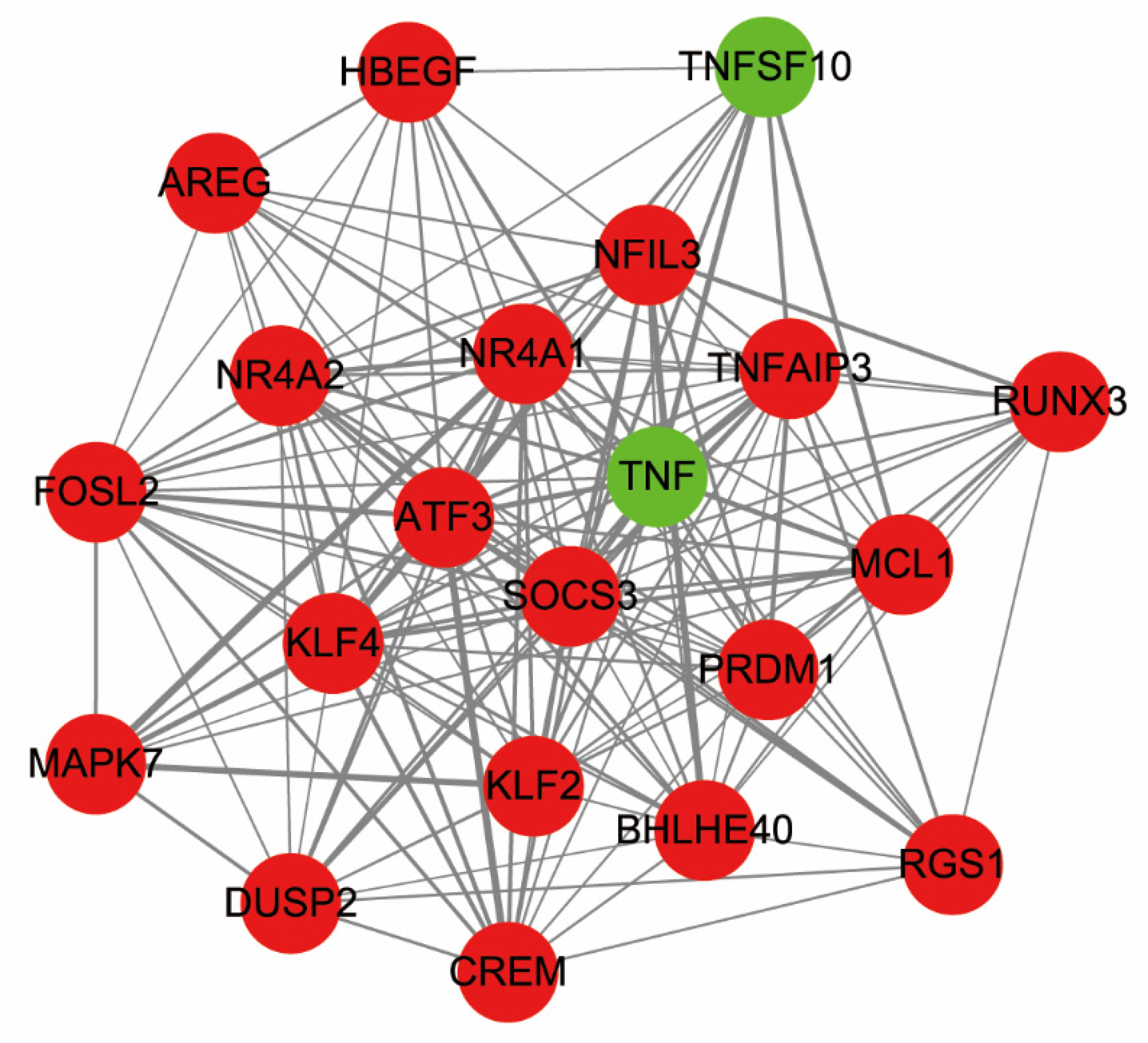
3.3. PPI Network and Module Analysis
3.4. TF Target Regulation Prediction
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kratofil, R.M.; Kubes, P.; Deniset, J.F. Monocyte Conversion During Inflammation and Injury. Arterioscler. Thromb. Vasc. Biol. 2017, 37, 35–42. [Google Scholar] [CrossRef]
- Hettinger, J.; Richards, D.M.; Hansson, J.; Barra, M.M.; Joschko, A.C.; Krijgsveld, J.; Feuerer, M. Origin of monocytes and macrophages in a committed progenitor. Nat. Immunol. 2013, 14, 821–830. [Google Scholar] [CrossRef] [PubMed]
- Guilliams, M.; Mildner, A.; Yona, S. Developmental and Functional Heterogeneity of Monocytes. Immunity 2018, 49, 595–613. [Google Scholar] [CrossRef] [PubMed]
- Wynn, T.A.; Vannella, K.M. Macrophages in Tissue Repair, Regeneration, and Fibrosis. Immunity 2016, 44, 450–462. [Google Scholar] [CrossRef] [PubMed]
- Scheffer, D.D.L.; Latini, A. Exercise-induced immune system response: Anti-inflammatory status on peripheral and central organs. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165823. [Google Scholar] [CrossRef] [PubMed]
- Simpson, R.J.; Kunz, H.; Agha, N.; Graff, R. Exercise and the Regulation of Immune Functions. Prog. Mol. Biol. Transl. Sci. 2015, 135, 355–380. [Google Scholar] [CrossRef] [PubMed]
- Khammassi, M.; Ouerghi, N.; Said, M.; Feki, M.; Khammassi, Y.; Pereira, B.; Thivel, D.; Bouassida, A. Continuous Moderate-Intensity but Not High-Intensity Interval Training Improves Immune Function Biomarkers in Healthy Young Men. J. Strength Cond. Res. 2020, 34, 249–256. [Google Scholar] [CrossRef]
- Wadley, A.J.; Chen, Y.W.; Lip, G.Y.; Fisher, J.P.; Aldred, S. Low volume-high intensity interval exercise elicits antioxidant and anti-inflammatory effects in humans. J. Sports Sci. 2016, 34, 1–9. [Google Scholar] [CrossRef]
- Handzlik, M.K.; Shaw, A.J.; Dungey, M.; Bishop, N.C.; Gleeson, M. The influence of exercise training status on antigen-stimulated IL-10 production in whole blood culture and numbers of circulating regulatory T cells. Eur. J. Appl. Physiol. 2013, 113, 1839–1848. [Google Scholar] [CrossRef]
- Tucker, P.S.; Briskey, D.R.; Scanlan, A.T.; Coombes, J.S.; Dalbo, V.J. High intensity interval training favourably affects antioxidant and inflammation mRNA expression in early-stage chronic kidney disease. Free Radic. Biol. Med. 2015, 89, 466–472. [Google Scholar] [CrossRef]
- Aw, N.H.; Canetti, E.; Suzuki, K.; Goh, J. Monocyte Subsets in Atherosclerosis and Modification with Exercise in Humans. Antioxidants 2018, 7, 196. [Google Scholar] [CrossRef]
- Radom-Aizik, S.; Zaldivar, F.P., Jr.; Haddad, F.; Cooper, D.M. Impact of brief exercise on circulating monocyte gene and microRNA expression: Implications for atherosclerotic vascular disease. Brain Behav. Immun. 2014, 39, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Ruffino, J.S.; Davies, N.A.; Morris, K.; Ludgate, M.; Zhang, L.; Webb, R.; Thomas, A.W. Moderate-intensity exercise alters markers of alternative activation in circulating monocytes in females: A putative role for PPARgamma. Eur. J. Appl. Physiol. 2016, 116, 1671–1682. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Wang, J.; Quan, M. Novel insights into the molecular mechanisms underlying the beneficial effects of exercise training on pulmonary arterial hypertension. J. Sports Med. Phys. Fit. 2019, 59, 1584–1592. [Google Scholar] [CrossRef] [PubMed]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Xie, C.; Mao, X.; Huang, J.; Ding, Y.; Wu, J.; Dong, S.; Kong, L.; Gao, G.; Li, C.Y.; Wei, L. KOBAS 2.0: A web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011, 39, W316–W322. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Chin, C.H.; Chen, S.H.; Wu, H.H.; Ho, C.W.; Ko, M.T.; Lin, C.Y. cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 2014, 8 (Suppl. 4), S11. [Google Scholar] [CrossRef] [PubMed]
- Bader, G.D.; Hogue, C.W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 2003, 4, 2. [Google Scholar] [CrossRef]
- Janky, R.; Verfaillie, A.; Imrichova, H.; Van de Sande, B.; Standaert, L.; Christiaens, V.; Hulselmans, G.; Herten, K.; Naval Sanchez, M.; Potier, D.; et al. iRegulon: From a gene list to a gene regulatory network using large motif and track collections. PLoS Comput. Biol. 2014, 10, e1003731. [Google Scholar] [CrossRef]
- Rahman, K.; Vengrenyuk, Y.; Ramsey, S.A.; Vila, N.R.; Girgis, N.M.; Liu, J.; Gusarova, V.; Gromada, J.; Weinstock, A.; Moore, K.J.; et al. Inflammatory Ly6Chi monocytes and their conversion to M2 macrophages drive atherosclerosis regression. J. Clin. Investig. 2017, 127, 2904–2915. [Google Scholar] [CrossRef]
- Avolio, E.; Spinetti, G.; Madeddu, P. Training monocytes by physical exercise: Good practice for improving collateral artery development and postischemic outcomes. Arterioscler. Thromb. Vasc. Biol. 2015, 35, 1733–1735. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Seternes, O.M.; Kidger, A.M.; Keyse, S.M. Dual-specificity MAP kinase phosphatases in health and disease. Biochim. Biophys. Acta Mol. Cell Res. 2019, 1866, 124–143. [Google Scholar] [CrossRef] [PubMed]
- Smallie, T.; Ross, E.A.; Ammit, A.J.; Cunliffe, H.E.; Tang, T.; Rosner, D.R.; Ridley, M.L.; Buckley, C.D.; Saklatvala, J.; Dean, J.L.; et al. Dual-Specificity Phosphatase 1 and Tristetraprolin Cooperate To Regulate Macrophage Responses to Lipopolysaccharide. J. Immunol. 2015, 195, 277–288. [Google Scholar] [CrossRef]
- Koga, Y.; Tsurumaki, H.; Aoki-Saito, H.; Sato, M.; Yatomi, M.; Takehara, K.; Hisada, T. Roles of Cyclic AMP Response Element Binding Activation in the ERK1/2 and p38 MAPK Signalling Pathway in Central Nervous System, Cardiovascular System, Osteoclast Differentiation and Mucin and Cytokine Production. Int. J. Mol. Sci. 2019, 20, 1346. [Google Scholar] [CrossRef]
- Li, B.; Yang, Y.; Chen, L.; Chen, S.; Zhang, J.; Tang, W. 18alpha-Glycyrrhetinic acid monoglucuronide as an anti-inflammatory agent through suppression of the NF-kappaB and MAPK signaling pathway. Medchemcomm 2017, 8, 1498–1504. [Google Scholar] [CrossRef]
- Lin, Y.; Lin, Y.; Lin, X.; Sun, X.; Luo, K. Combination of PET and CXCR4-Targeted Peptide Molecule Agents for Noninvasive Tumor Monitoring. J. Cancer 2019, 10, 3420–3426. [Google Scholar] [CrossRef]
- Doring, Y.; Pawig, L.; Weber, C.; Noels, H. The CXCL12/CXCR4 chemokine ligand/receptor axis in cardiovascular disease. Front. Physiol. 2014, 5, 212. [Google Scholar] [CrossRef]
- Page, M.; Ridge, L.; Gold Diaz, D.; Tsogbayar, T.; Scambler, P.J.; Ivins, S. Loss of CXCL12/CXCR4 signalling impacts several aspects of cardiovascular development but does not exacerbate Tbx1 haploinsufficiency. PLoS ONE 2018, 13, e0207251. [Google Scholar] [CrossRef]
- Davis, M.I.; Puhl, H.L., 3rd. Nr4a1-eGFP is a marker of striosome-matrix architecture, development and activity in the extended striatum. PLoS ONE 2011, 6, e16619. [Google Scholar] [CrossRef]
- Liu, X.; Wang, Y.; Lu, H.; Li, J.; Yan, X.; Xiao, M.; Hao, J.; Alekseev, A.; Khong, H.; Chen, T.; et al. Genome-wide analysis identifies NR4A1 as a key mediator of T cell dysfunction. Nature 2019, 567, 525–529. [Google Scholar] [CrossRef] [PubMed]
- Hanna, R.N.; Shaked, I.; Hubbeling, H.G.; Punt, J.A.; Wu, R.; Herrley, E.; Zaugg, C.; Pei, H.; Geissmann, F.; Ley, K.; et al. NR4A1 (Nur77) deletion polarizes macrophages toward an inflammatory phenotype and increases atherosclerosis. Circ. Res. 2012, 110, 416–427. [Google Scholar] [CrossRef] [PubMed]
- Freire, P.R.; Conneely, O.M. NR4A1 and NR4A3 restrict HSC proliferation via reciprocal regulation of C/EBPalpha and inflammatory signaling. Blood 2018, 131, 1081–1093. [Google Scholar] [CrossRef]
- Sabaratnam, R.; Pedersen, A.J.; Eskildsen, T.V.; Kristensen, J.M.; Wojtaszewski, J.F.P.; Hojlund, K. Exercise Induction of Key Transcriptional Regulators of Metabolic Adaptation in Muscle Is Preserved in Type 2 Diabetes. J. Clin. Endocrinol. Metab. 2019, 104, 4909–4920. [Google Scholar] [CrossRef]
- Miao, H.; Miao, C.X.; Li, N.; Han, J. FOXJ2 controls meiosis during spermatogenesis in male mice. Mol. Reprod. Dev. 2016, 83, 684–691. [Google Scholar] [CrossRef]
- Martin-de-Lara, F.; Sanchez-Aparicio, P.; de la Fuente, C.A.; Rey-Campos, J. Biological effects of FoxJ2 over-expression. Transgenic Res. 2008, 17, 1131–1141. [Google Scholar] [CrossRef][Green Version]
- Carow, B.; Rottenberg, M.E. SOCS3, a Major Regulator of Infection and Inflammation. Front. Immunol. 2014, 5, 58. [Google Scholar] [CrossRef]
- Pal, S.; Nath, P.; Biswas, S.; Mukherjee, U.; Maitra, S. Nonylphenol attenuates SOCS3 expression and M1 polarization in lipopolysaccharide-treated rat splenic macrophages. Ecotoxicol. Environ. Saf. 2019, 174, 574–583. [Google Scholar] [CrossRef]
- Chen, Y.H.; Hsieh, S.C.; Chen, W.Y.; Li, K.J.; Wu, C.H.; Wu, P.C.; Tsai, C.Y.; Yu, C.L. Spontaneous resolution of acute gouty arthritis is associated with rapid induction of the anti-inflammatory factors TGFbeta1, IL-10 and soluble TNF receptors and the intracellular cytokine negative regulators CIS and SOCS3. Ann. Rheum. Dis. 2011, 70, 1655–1663. [Google Scholar] [CrossRef] [PubMed]
- Vermeij, E.A.; Broeren, M.G.; Bennink, M.B.; Arntz, O.J.; Gjertsson, I.; van Lent, P.L.; van den Berg, W.B.; Koenders, M.I.; van de Loo, F.A. Disease-regulated local IL-10 gene therapy diminishes synovitis and cartilage proteoglycan depletion in experimental arthritis. Ann. Rheum. Dis. 2015, 74, 2084–2091. [Google Scholar] [CrossRef]
- Link, W. Introduction to FOXO Biology. Methods Mol. Biol. 2019, 1890, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Eijkelenboom, A.; Burgering, B.M. FOXOs: Signalling integrators for homeostasis maintenance. Nat. Rev. Mol. Cell Biol. 2013, 14, 83–97. [Google Scholar] [CrossRef] [PubMed]
- Newton, R.H.; Shrestha, S.; Sullivan, J.M.; Yates, K.B.; Compeer, E.B.; Ron-Harel, N.; Blazar, B.R.; Bensinger, S.J.; Haining, W.N.; Dustin, M.L.; et al. Maintenance of CD4 T cell fitness through regulation of Foxo1. Nat. Immunol. 2018, 19, 838–848. [Google Scholar] [CrossRef] [PubMed]
- Ushmorov, A.; Wirth, T. FOXO in B-cell lymphopoiesis and B cell neoplasia. Semin. Cancer Biol. 2018, 50, 132–141. [Google Scholar] [CrossRef]
- Becker, T.; Loch, G.; Beyer, M.; Zinke, I.; Aschenbrenner, A.C.; Carrera, P.; Inhester, T.; Schultze, J.L.; Hoch, M. FOXO-dependent regulation of innate immune homeostasis. Nature 2010, 463, 369–373. [Google Scholar] [CrossRef]
- Matsuzaki, T.; Alvarez-Garcia, O.; Mokuda, S.; Nagira, K.; Olmer, M.; Gamini, R.; Miyata, K.; Akasaki, Y.; Su, A.I.; Asahara, H.; et al. FoxO transcription factors modulate autophagy and proteoglycan 4 in cartilage homeostasis and osteoarthritis. Sci. Transl. Med. 2018, 10, eaan0746. [Google Scholar] [CrossRef]
- Lu, M.; Xu, W.; Gao, B.; Xiong, S. Blunting Autoantigen-induced FOXO3a Protein Phosphorylation and Degradation Is a Novel Pathway of Glucocorticoids for the Treatment of Systemic Lupus Erythematosus. J. Biol. Chem. 2016, 291, 19900–19912. [Google Scholar] [CrossRef] [PubMed]
- Tsuchiya, K.; Westerterp, M.; Murphy, A.J.; Subramanian, V.; Ferrante, A.W., Jr.; Tall, A.R.; Accili, D. Expanded granulocyte/monocyte compartment in myeloid-specific triple FoxO knockout increases oxidative stress and accelerates atherosclerosis in mice. Circ. Res. 2013, 112, 992–1003. [Google Scholar] [CrossRef] [PubMed]
- Bouman, A.; Schipper, M.; Heineman, M.J.; Faas, M.M. Gender difference in the non-specific and specific immune response in humans. Am. J. Reprod. Immunol. 2004, 52, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Ben-Hur, H.; Mor, G.; Insler, V.; Blickstein, I.; Amir-Zaltsman, Y.; Sharp, A.; Globerson, A.; Kohen, F. Menopause is associated with a significant increase in blood monocyte number and a relative decrease in the expression of estrogen receptors in human peripheral monocytes. Am. J. Reprod. Immunol. 1995, 34, 363–369. [Google Scholar] [CrossRef] [PubMed]
- Young, N.A.; Wu, L.C.; Burd, C.J.; Friedman, A.K.; Kaffenberger, B.H.; Rajaram, M.V.; Schlesinger, L.S.; James, H.; Shupnik, M.A.; Jarjour, W.N. Estrogen modulation of endosome-associated toll-like receptor 8: An IFNalpha-independent mechanism of sex-bias in systemic lupus erythematosus. Clin. Immunol. 2014, 151, 66–77. [Google Scholar] [CrossRef] [PubMed]
- Rauf, S.; Soejono, S.K.; Partadiredja, G. Effects of treadmill exercise training on cerebellar estrogen and estrogen receptors, serum estrogen, and motor coordination performance of ovariectomized rats. Iran. J. Basic Med. Sci. 2015, 18, 587–592. [Google Scholar]
- Michishita, R.; Shono, N.; Inoue, T.; Tsuruta, T.; Node, K. Effect of exercise therapy on monocyte and neutrophil counts in overweight women. Am. J. Med. Sci. 2010, 339, 152–156. [Google Scholar] [CrossRef]





| GO-ID | Term | Gene Counts | p-Value |
|---|---|---|---|
| GO-BP terms | |||
| GO:0070301 | cellular response to hydrogen peroxide | 6 | 2.81 × 10−5 |
| GO:0006986 | response to unfolded protein | 5 | 1.35 × 10−4 |
| GO:0008285 | negative regulation of cell proliferation | 10 | 8.43 × 10−4 |
| GO:0071499 | cellular response to laminar fluid shear stress | 3 | 0.001058 |
| GO:0051247 | positive regulation of protein metabolic process | 3 | 0.001058 |
| GO-CC terms | |||
| GO:0005634 | Nucleus | 52 | 6.68 × 10−5 |
| GO:0043227 | membrane-bounded organelle | 67 | 0.00014 |
| GO:0043231 | intracellular membrane-bounded organelle | 62 | 0.0009 |
| GO:0005737 | Cytoplasm | 45 | 0.003735 |
| GO:0005654 | Nucleoplasm | 26 | 0.017733 |
| GO-MF terms | |||
| GO:0051087 | chaperone binding | 6 | 1.70 × 10−4 |
| GO:0005515 | protein binding | 75 | 3.58 × 10−4 |
| GO:0003700 | transcription factor activity sequence-specific DNA binding | 16 | 0.001167 |
| GO:0030544 | Hsp70 protein binding | 4 | 0.001208 |
| GO:0043130 | ubiquitin binding | 5 | 0.001571 |
| Pathway | ID | Gene Counts | p-Value | Genes |
|---|---|---|---|---|
| FoxO signaling pathway | hsa04068 | 5 | 1.69 × 10−5 | PLK2, CDKN1B, TNFSF10, GABARAPL1, KLF2 |
| Protein processing in endoplasmic reticulum | hsa04141 | 5 | 4.75 × 10−5 | DNAJB1, PLAA, DNAJA1, HERPUD1, HSPH1 |
| Influenza A | hsa05164 | 5 | 5.02 × 10−5 | DNAJB1, TNFSF10, NXT1, SOCS3, TNF |
| ErbB signaling pathway | hsa04012 | 4 | 5.46 × 10−5 | CDKN1B, EREG, HBEGF, AREG |
| MAPK signaling pathway | hsa04010 | 6 | 7.17 × 10−5 | AREG, NR4A1, TNF, MAPK7, DUSP2, EREG |
| MicroRNAs in cancer | hsa05206 | 6 | 7.71 × 10−5 | DDIT4, CDKN1B, MAPK7, CYP1B1, mir-223, MCL1 |
| Parathyroid hormone synthesis, secretion and action | hsa04928 | 4 | 0.000124 | NR4A2, HBEGF, PDE4B, GNA13 |
| PI3K-Akt signaling pathway | hsa04151 | 6 | 0.00019 | AREG, DDIT4, CDKN1B, NR4A1, MCL1, EREG |
| Pathways in cancer | hsa05200 | 7 | 0.000249 | PIM2, CDKN1B, CXCR4, RASSF5, CKS1B, GNA13, CKS2 |
| Adipocytokine signaling pathway | hsa04920 | 3 | 0.000619 | TNF, SOCS3, CD36 |
| Epstein–Barr virus infection | hsa05169 | 4 | 0.001303 | CDKN1B, TNF, TNFAIP3, RUNX3 |
| IL-17 signaling pathway | hsa04657 | 3 | 0.001425 | TNF, MAPK7, TNFAIP3 |
| Small-cell lung cancer | hsa05222 | 3 | 0.001425 | CDKN1B, CKS1B, CKS2 |
| Viral protein interaction with cytokine and cytokine receptor | hsa04061 | 3 | 0.001744 | TNFSF10, TNF, CXCR4 |
| Glycosylphosphatidylinositol (GPI)-anchor biosynthesis | hsa00563 | 2 | 0.001751 | PIGM, PIGW |
| Gene Symbol | Gene Description | Degree |
|---|---|---|
| TNF | Tumor necrosis factor | 56 |
| DUSP1 | Dual-specificity protein phosphatase 1 | 47 |
| ATF3 | Cyclic AMP-dependent transcription factor ATF-3 | 47 |
| CXCR4 | C-X-C chemokine receptor type 4 | 40 |
| NR4A1 | Nuclear receptor subfamily 4 group A member 1 | 37 |
| BHLHE40 | Class E basic helix-loop-helix protein 40 | 35 |
| CDKN1B | Cyclin-dependent kinase inhibitor 1B | 34 |
| SOCS3 | Suppressor of cytokine signaling 3 | 34 |
| TNFAIP3 | Tumor necrosis factor alpha-induced protein 3 | 31 |
| MCL1 | Induced myeloid leukemia cell differentiation protein Mcl-1 | 31 |
| GO-ID | Term | Gene Counts | p-Value |
|---|---|---|---|
| GO-BP terms | |||
| GO:0048523 | negative regulation of cellular process | 20 | 3.59 × 10−9 |
| GO:0031324 | negative regulation of cellular metabolic process | 16 | 2.26 × 10−8 |
| GO:0010604 | positive regulation of macromolecule metabolic process | 17 | 2.77 × 10−8 |
| GO:0009968 | negative regulation of signal transduction | 12 | 7.14 × 10−8 |
| GO:0051172 | negative regulation of nitrogen compound metabolic process | 15 | 7.14 × 10−8 |
| GO-CC terms | |||
| GO:0005634 | Nucleus | 16 | 0.0185 |
| GO:0044428 | nuclear part | 12 | 0.0337 |
| GO-MF terms | |||
| GO:0000977 | RNA polymerase II regulatory region sequence-specific DNA binding | 9 | 7.56 × 10−7 |
| GO:0001012 | RNA polymerase II regulatory region DNA binding | 9 | 7.56 × 10−7 |
| GO:0044212 | transcription regulatory region DNA binding | 10 | 7.56 × 10−7 |
| GO:0043565 | sequence-specific DNA binding | 10 | 9.79 × 10−7 |
| GO:0003677 | DNA binding | 13 | 2.79 × 10−6 |
| Pathway | ID | Gene Counts | p-Value | Genes |
|---|---|---|---|---|
| MAPK signaling pathway | hsa04010 | 5 | 7.84 × 10−8 | AREG, NR4A1, MAPK7, TNF, DUSP2 |
| IL-17 signaling pathway | hsa04657 | 3 | 6.23 × 10−6 | TNF, MAPK7, TNFAIP3 |
| TNF signaling pathway | hsa04668 | 3 | 1.07 × 10−5 | TNF, TNFAIP3, SOCS3 |
| Osteoclast differentiation | hsa04380 | 3 | 1.58 × 10−5 | TNF, SOCS3, FOSL2 |
| Apoptosis | hsa04210 | 3 | 1.89 × 10−5 | TNFSF10, MCL1, TNF |
| Fluid shear stress and atherosclerosis | hsa05418 | 3 | 2.01 × 10−5 | TNF, KLF2, MAPK7 |
| Necroptosis | hsa04217 | 3 | 3.15 × 10−5 | TNFSF10, TNF, TNFAIP3 |
| Influenza A | hsa05164 | 3 | 3.45 × 10−5 | TNFSF10, TNF, SOCS3 |
| Epstein–Barr virus infection | hsa05169 | 3 | 5.92 × 10−5 | RUNX3, TNF, TNFAIP3 |
| Type 2 diabetes mellitus | hsa04930 | 2 | 0.000148 | TNF, SOCS3 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, P.; Luo, L. Identification of Critical Genes and Signaling Pathways in Human Monocytes Following High-Intensity Exercise. Healthcare 2021, 9, 618. https://doi.org/10.3390/healthcare9060618
Li P, Luo L. Identification of Critical Genes and Signaling Pathways in Human Monocytes Following High-Intensity Exercise. Healthcare. 2021; 9(6):618. https://doi.org/10.3390/healthcare9060618
Chicago/Turabian StyleLi, Pengda, and Li Luo. 2021. "Identification of Critical Genes and Signaling Pathways in Human Monocytes Following High-Intensity Exercise" Healthcare 9, no. 6: 618. https://doi.org/10.3390/healthcare9060618
APA StyleLi, P., & Luo, L. (2021). Identification of Critical Genes and Signaling Pathways in Human Monocytes Following High-Intensity Exercise. Healthcare, 9(6), 618. https://doi.org/10.3390/healthcare9060618






