Evaluation of Country Dietary Habits Using Machine Learning Techniques in Relation to Deaths from COVID-19
Abstract
1. Introduction
2. Methods
2.1. Principal Component Analysis (Pca)
2.2. K-Means
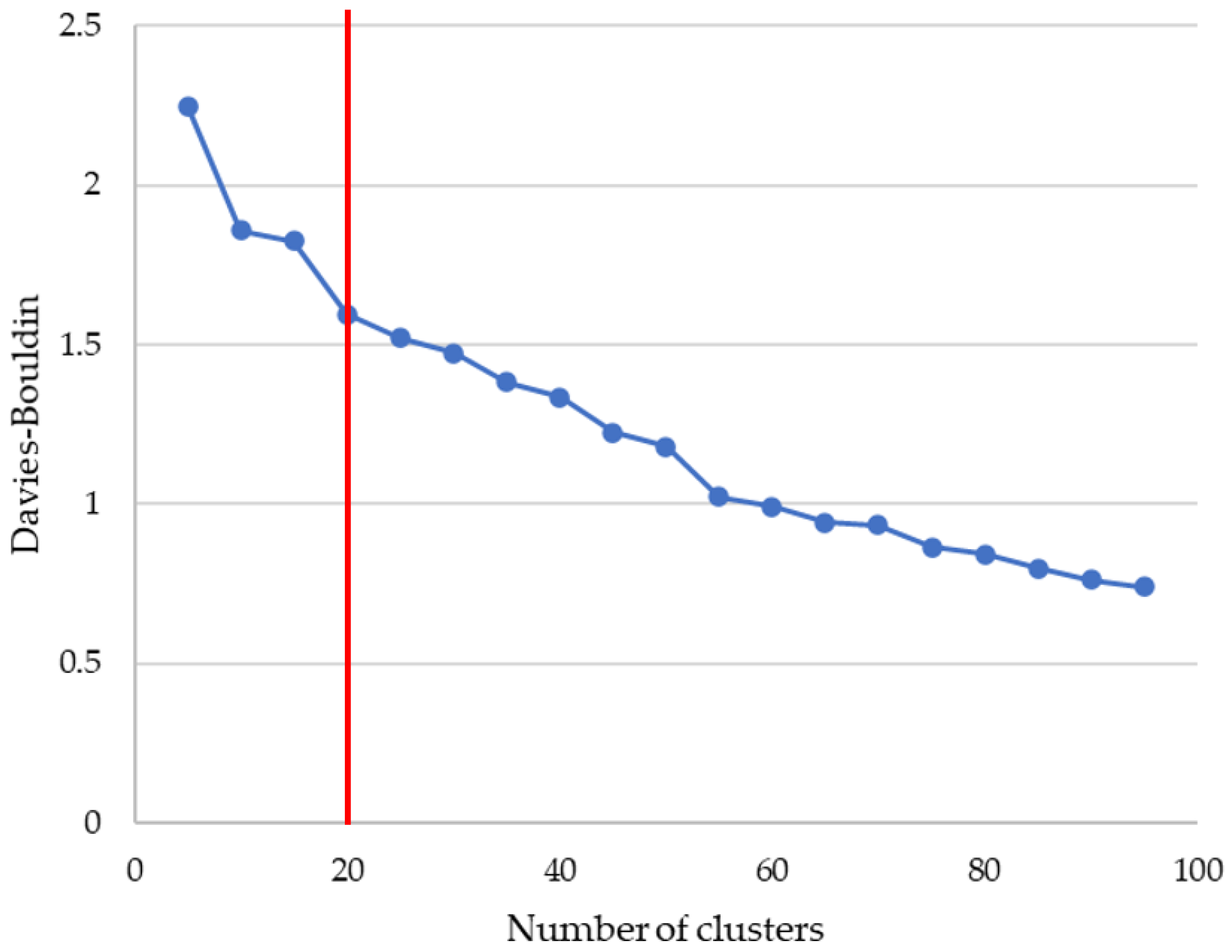
2.3. Clustering Metric: Davies–Bouldin
3. Experiments and Results
3.1. Dataset
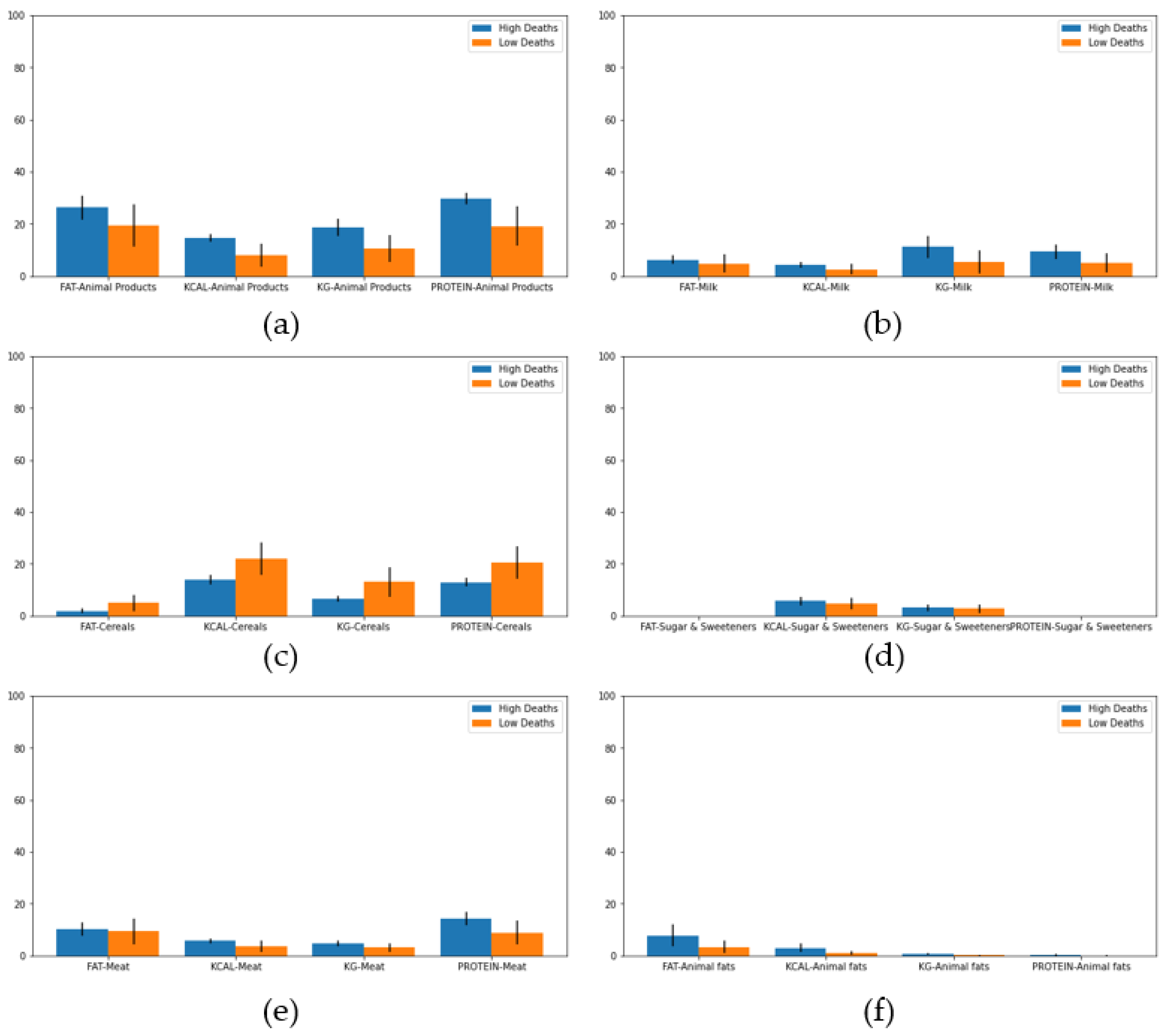
- Percentages of fat consumed from each type of food listed.
- Percentages of food supply (in kg) for each type of food listed.
- Percentages of energy (in kilocalories) consumed from each type of food listed.
- Percentages of protein consumed from each type of food listed.
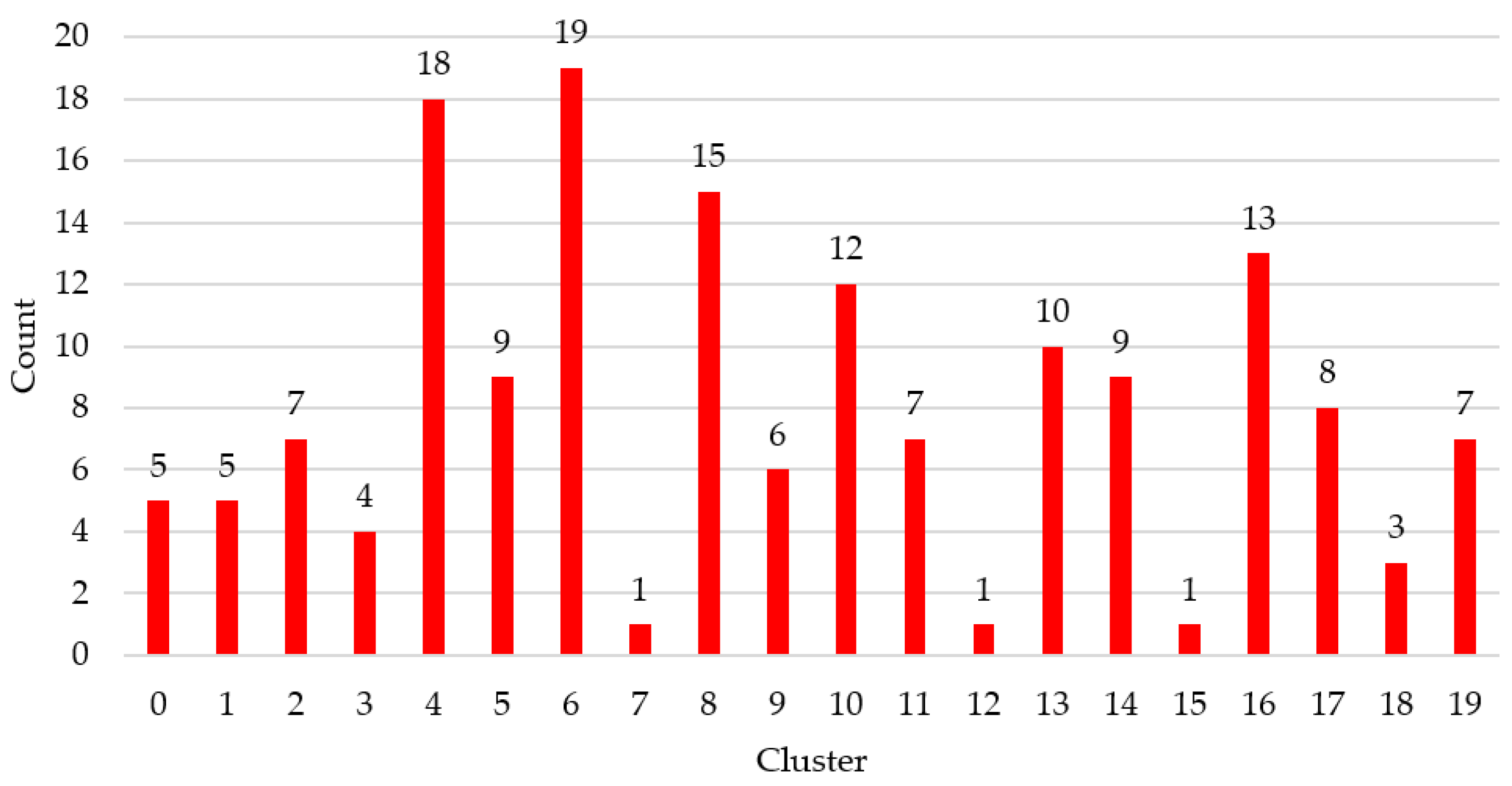
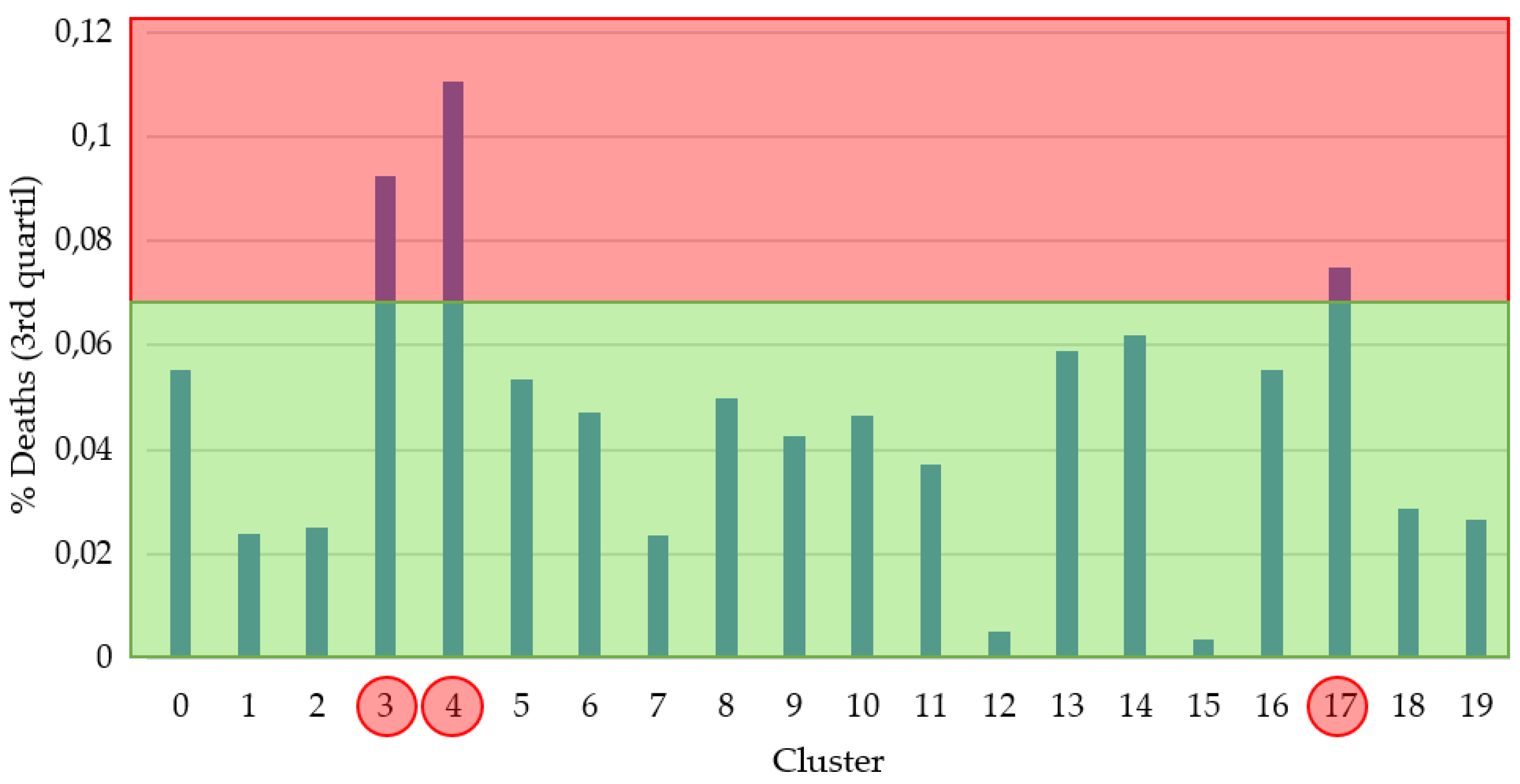
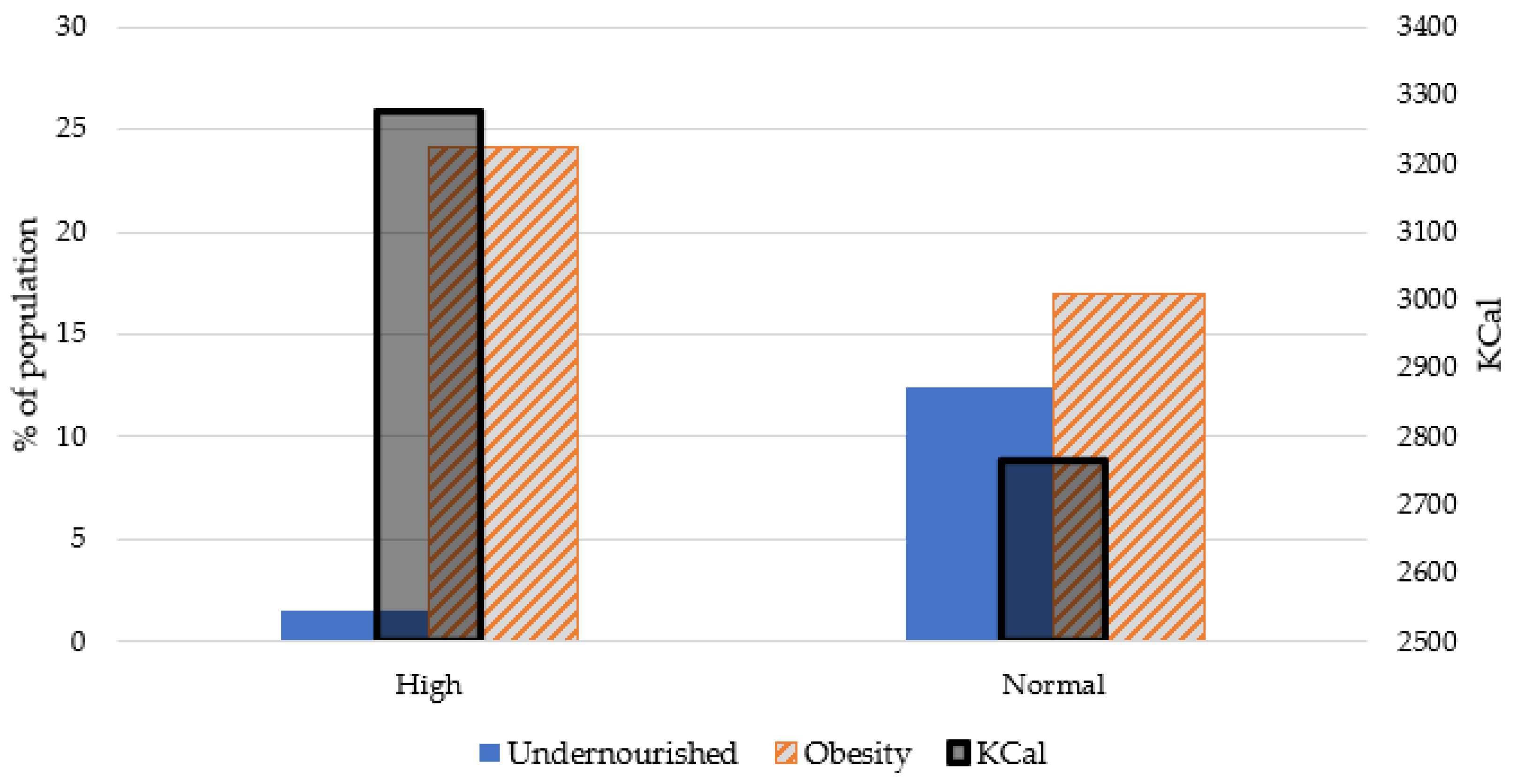
3.2. Experiments
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Chen, H.; Guo, J.; Wang, C.; Luo, F.; Yu, X.; Zhang, W.; Li, J.; Zhao, D.; Xu, D.; Gong, Q.; et al. Clinical characteristics and intrauterine vertical transmission potential of COVID-19 infection in nine pregnant women: A retrospective review of medical records. Lancet 2020, 395, 809–815. [Google Scholar] [CrossRef]
- Kampf, G.; Todt, D.; Pfaender, S.; Steinmann, E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J. Hosp. Infect. 2020, 104, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.W.; Yuan, S.; Kok, K.H.; To, K.K.W.; Chu, H.; Yang, J.; Xing, F.; Liu, J.; Yip, C.C.Y.; Poon, R.W.S.; et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: A study of a family cluster. Lancet 2020, 395, 514–523. [Google Scholar] [CrossRef]
- Otter, J.A.; Donskey, C.; Yezli, S.; Douthwaite, S.; Goldenberg, S.D.; Weber, D.J. Transmission of SARS and MERS coronaviruses and influenza virus in healthcare settings: The possible role of dry surface contamination. J. Hosp. Infect. 2016, 92, 235–250. [Google Scholar] [CrossRef]
- Bai, Y.; Yao, L.; Wei, T.; Tian, F.; Jin, D.Y.; Chen, L.; Wang, M. Presumed Asymptomatic Carrier Transmission of COVID-19. JAMA 2020, 323, 1406–1407. [Google Scholar] [CrossRef] [PubMed]
- Rothe, C.; Schunk, M.; Sothmann, P.; Bretzel, G.; Froeschl, G.; Wallrauch, C.; Zimmer, T.; Thiel, V.; Janke, C.; Guggemos, W.; et al. Transmission of 2019-NCOV infection from an asymptomatic contact in Germany. N. Engl. J. Med. 2020, 382, 970–971. [Google Scholar] [CrossRef]
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: A descriptive study. Lancet 2020, 395, 507–513. [Google Scholar] [CrossRef]
- De Wit, E.; Feldmann, F.; Cronin, J.; Jordan, R.; Okumura, A.; Thomas, T.; Scott, D.; Cihlar, T.; Feldmann, H. Prophylactic and therapeutic remdesivir (GS-5734) treatment in the rhesus macaque model of MERS-CoV infection. Proc. Natl. Acad. Sci. USA 2020, 117, 6771–6776. [Google Scholar] [CrossRef]
- Liu, X.; Wang, X.J. Potential inhibitors for 2019-nCoV coronavirus M protease from clinically approved medicines. J. Genet. Genom. 2020, 47, 119–121. [Google Scholar] [CrossRef]
- Sarma, P.; Kaur, H.; Kumar, H.; Mahendru, D.; Avti, P.; Bhattacharyya, A.; Prajapat, M.; Shekhar, N.; Kumar, S.; Singh, R.; et al. Virological and clinical cure in COVID-19 patients treated with hydroxychloroquine: A systematic review and meta-analysis. J. Med. Virol. 2020, 92, 776–785. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Zhou, Q.; Li, Y.; Garner, L.V.; Watkins, S.P.; Carter, L.J.; Smoot, J.; Gregg, A.C.; Daniels, A.D.; Jervey, S.; et al. Research and Development on Therapeutic Agents and Vaccines for COVID-19 and Related Human Coronavirus Diseases. ACS Cent. Sci. 2020, 6, 315–331. [Google Scholar] [CrossRef] [PubMed]
- Gozes, O.; Frid-Adar, M.; Greenspan, H.; Browning, P.D.; Zhang, H.; Ji, W.; Bernheim, A.; Siegel, E. Rapid AI Development Cycle for the Coronavirus (COVID-19) Pandemic: Initial Results for Automated Detection & Patient Monitoring using Deep Learning CT Image Analysis. arXiv 2020, arXiv:2003.05037. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Berlin/Heidelberg, Germany, 2015; Volume 9351, pp. 234–241. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; IEEE Computer Society: Washington, DC, USA, 2016; Volume 2016-Decem, pp. 770–778. [Google Scholar] [CrossRef]
- Li, L.; Qin, L.; Xu, Z.; Yin, Y.; Wang, X.; Kong, B.; Bai, J.; Lu, Y.; Fang, Z.; Song, Q.; et al. Artificial Intelligence Distinguishes COVID-19 from Community Acquired Pneumonia on Chest CT. Radiology 2020, 296, E65–E71. [Google Scholar] [CrossRef]
- Pirouz, B.; Shaffiee Haghshenas, S.; Shaffiee Haghshenas, S.; Piro, P. Investigating a Serious Challenge in the Sustainable Development Process: Analysis of Confirmed cases of COVID-19 (New Type of Coronavirus) Through a Binary Classification Using Artificial Intelligence and Regression Analysis. Sustainability 2020, 12, 2427. [Google Scholar] [CrossRef]
- Shi, J.; Shao, X.; Guo, X.; Fang, W.; Wu, X.; Teng, Y.; Zhang, L.; Li, Z.; Liu, Y. Dietary habits and breast cancer risk: A hospital-based case-control study in Chinese women. Clin. Breast Cancer 2020, 20, e540–e550. [Google Scholar] [CrossRef]
- Powell, H.S.; Greenberg, D.L. Screening for unhealthy diet and exercise habits: The electronic health record and a healthier population. Prev. Med. Rep. 2019, 14, 100816. [Google Scholar] [CrossRef]
- Lockhart, S.M.; O’Rahilly, S. When Two Pandemics Meet: Why Is Obesity Associated with Increased COVID-19 Mortality? Med 2020. in Press. [Google Scholar] [CrossRef]
- Yadav, R.; Aggarwal, S.; Singh, A. SARS-CoV-2-host dynamics: Increased risk of adverse outcomes of COVID-19 in obesity. Diabetes Metab. Syndr. Clin. Res. Rev. 2020, 14, 1355–1360. [Google Scholar] [CrossRef]
- Pearson, K. LIII. On lines and planes of closest fit to systems of points in space. Lond. Edinb. Dublin Philos. Mag. J. Sci. 1901, 2, 559–572. [Google Scholar] [CrossRef]
- Davies, D.L.; Bouldin, D.W. A Cluster Separation Measure. IEEE Trans. Pattern Anal. Mach. Intell. 1979, PAMI-1, 224–227. [Google Scholar] [CrossRef]
- COVID-19 Healthy Diet Dataset | Kaggle. María Ren. Available online: https://www.kaggle.com/mariaren/covid19-healthy-diet-dataset (accessed on 28 September 2020).
- Yehia, B.R.; Winegar, A.; Fogel, R.; Fakih, M.; Ottenbacher, A.; Jesser, C.; Bufalino, A.; Huang, R.H.; Cacchione, J. Association of Race With Mortality Among Patients Hospitalized With Coronavirus Disease 2019 (COVID-19) at 92 US Hospitals. JAMA Netw. Open 2020, 3, e2018039. [Google Scholar] [CrossRef] [PubMed]
- Booker, S.; Cousin, L.; Buck, H.G. Surviving Multiple Pandemics-COVID-19 and Racism for African American Older Adults: A Call to Gerontological Nursing for Social Justice. J. Gerontol. Nurs. 2020, 46, 4–6. [Google Scholar] [CrossRef] [PubMed]
- FAOSTAT. Available online: http://www.fao.org/faostat/en/#home (accessed on 28 September 2020).
- Lionetti, V.; Tuana, B.; Casieri, V.; Parikh, M.; Pierce, G. Importance of functional food compounds in cardioprotection through action on the epigenome. Eur. Heart J. 2019, 40, 575–582. [Google Scholar] [CrossRef]
- Hämäläinen, J.; Jauhiainen, S.; Kärkkäinen, T. Comparison of internal clustering validation indices for prototype-based clustering. Algorithms 2017, 10, 105. [Google Scholar] [CrossRef]
- Zheng, K.I.; Gao, F.; Wang, X.B.; Sun, Q.F.; Pan, K.H.; Wang, T.Y.; Ma, H.L.; Liu, W.Y.; George, J.; Zheng, M.H. Obesity as a risk factor for greater severity of COVID-19 in patients with metabolic associated fatty liver disease. Metab. Clin. Exp. 2020, 108, 154244. [Google Scholar] [CrossRef] [PubMed]
| Australia | Austria | Bahamas | Barbados | Belgium |
| Canada | Cyprus | Czechia | Denmark | France |
| Germany | Greece | Hungary | Ireland | Israel |
| Italy | Kazakhstan | Latvia | Lithuania | Netherlands |
| New Zealand | Norway | Poland | Portugal | Slovakia |
| Slovenia | Spain | Sweden | Switzerland | USA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
García-Ordás, M.T.; Arias, N.; Benavides, C.; García-Olalla, O.; Benítez-Andrades, J.A. Evaluation of Country Dietary Habits Using Machine Learning Techniques in Relation to Deaths from COVID-19. Healthcare 2020, 8, 371. https://doi.org/10.3390/healthcare8040371
García-Ordás MT, Arias N, Benavides C, García-Olalla O, Benítez-Andrades JA. Evaluation of Country Dietary Habits Using Machine Learning Techniques in Relation to Deaths from COVID-19. Healthcare. 2020; 8(4):371. https://doi.org/10.3390/healthcare8040371
Chicago/Turabian StyleGarcía-Ordás, María Teresa, Natalia Arias, Carmen Benavides, Oscar García-Olalla, and José Alberto Benítez-Andrades. 2020. "Evaluation of Country Dietary Habits Using Machine Learning Techniques in Relation to Deaths from COVID-19" Healthcare 8, no. 4: 371. https://doi.org/10.3390/healthcare8040371
APA StyleGarcía-Ordás, M. T., Arias, N., Benavides, C., García-Olalla, O., & Benítez-Andrades, J. A. (2020). Evaluation of Country Dietary Habits Using Machine Learning Techniques in Relation to Deaths from COVID-19. Healthcare, 8(4), 371. https://doi.org/10.3390/healthcare8040371









