Breast Cancer Classification by Using Multi-Headed Convolutional Neural Network Modeling
Abstract
1. Introduction
2. Materials and Methods
2.1. Dataset
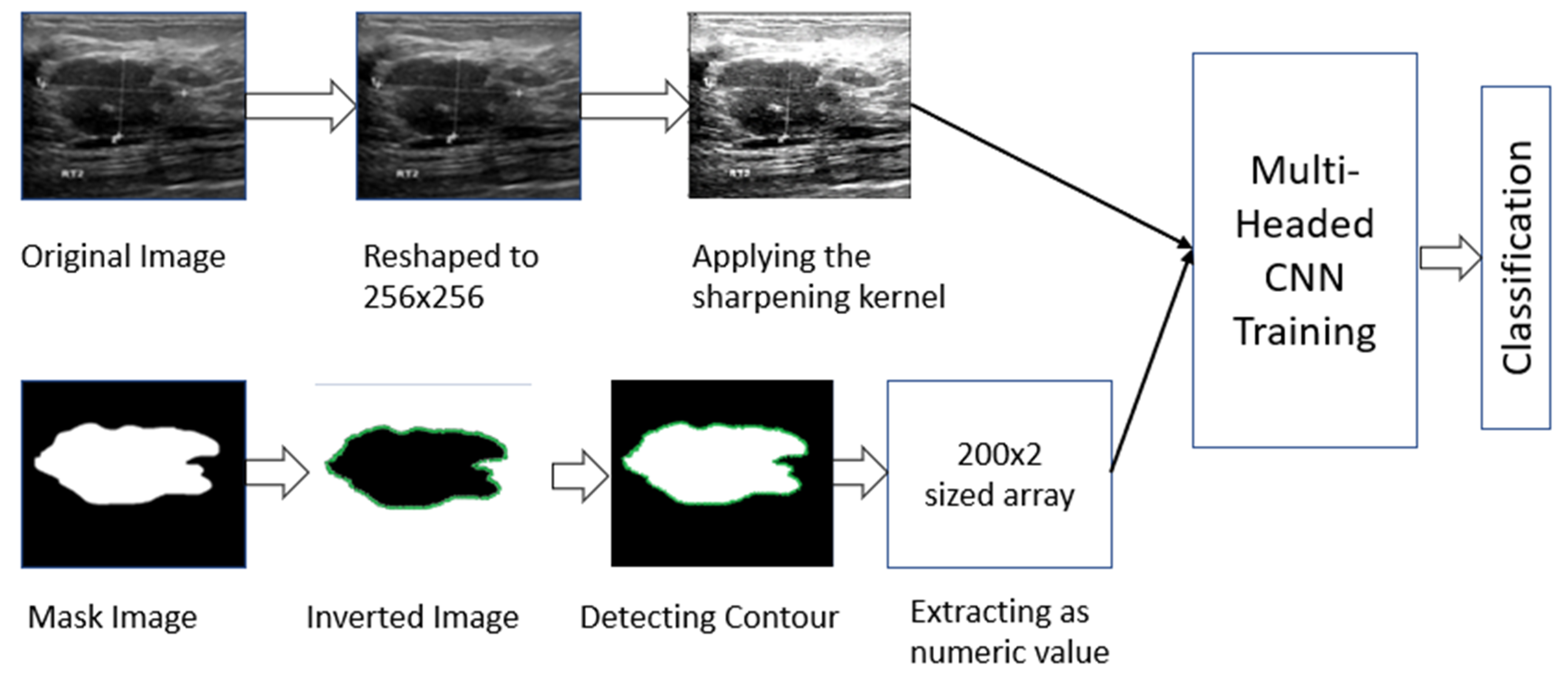
2.2. Working Procedure
2.2.1. Masked Image Processing
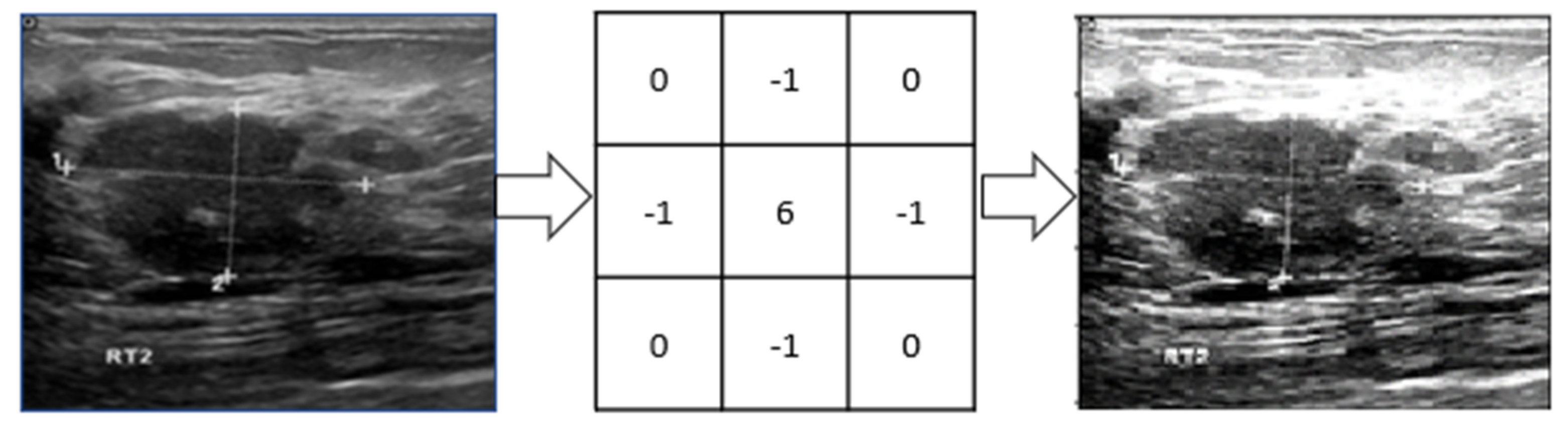
2.2.2. Raw Image Processing
2.3. Modeling and Evaluating Datasets
2.3.1. Evaluating Masked Images
2.3.2. Evaluating Original Images
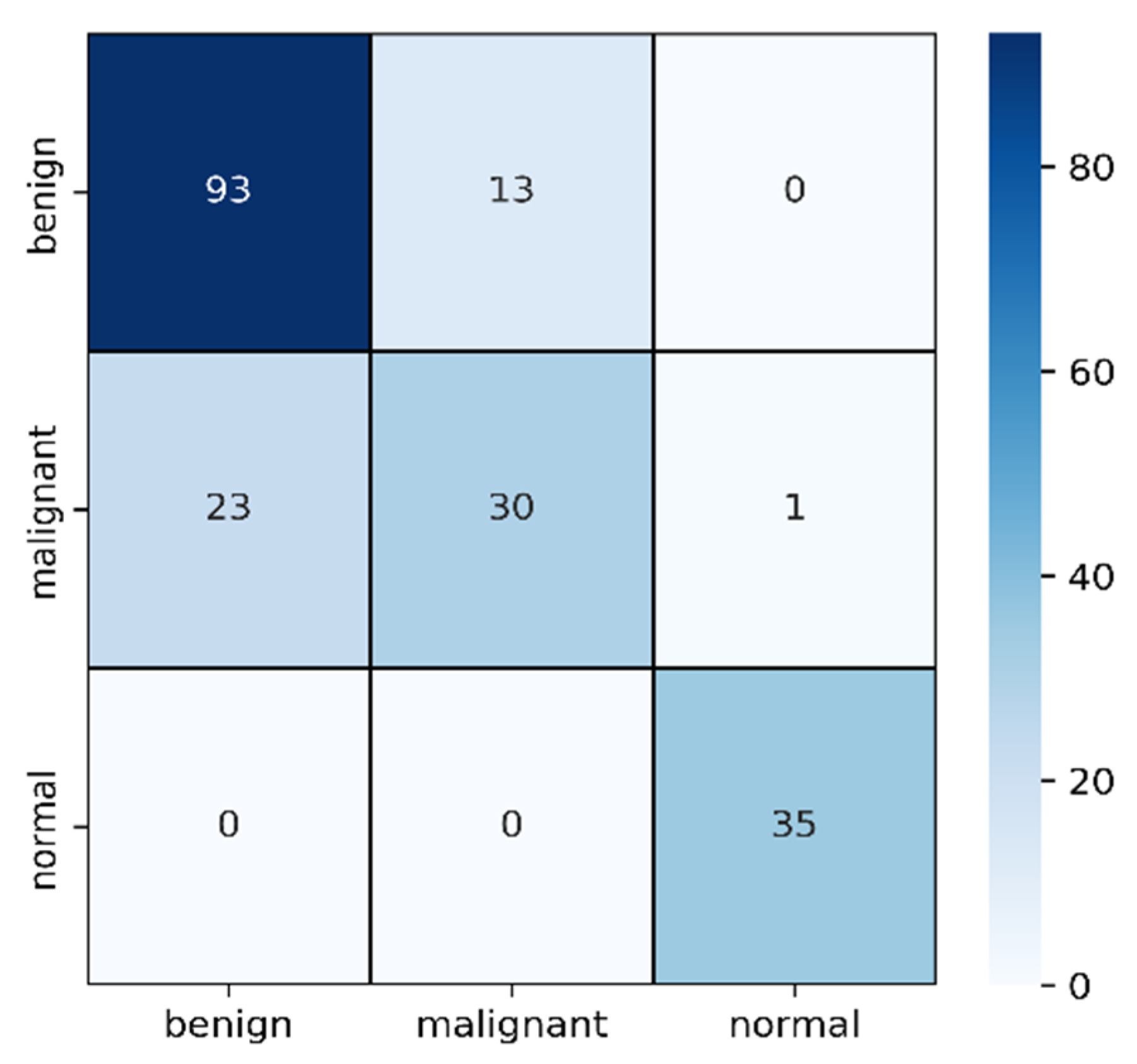
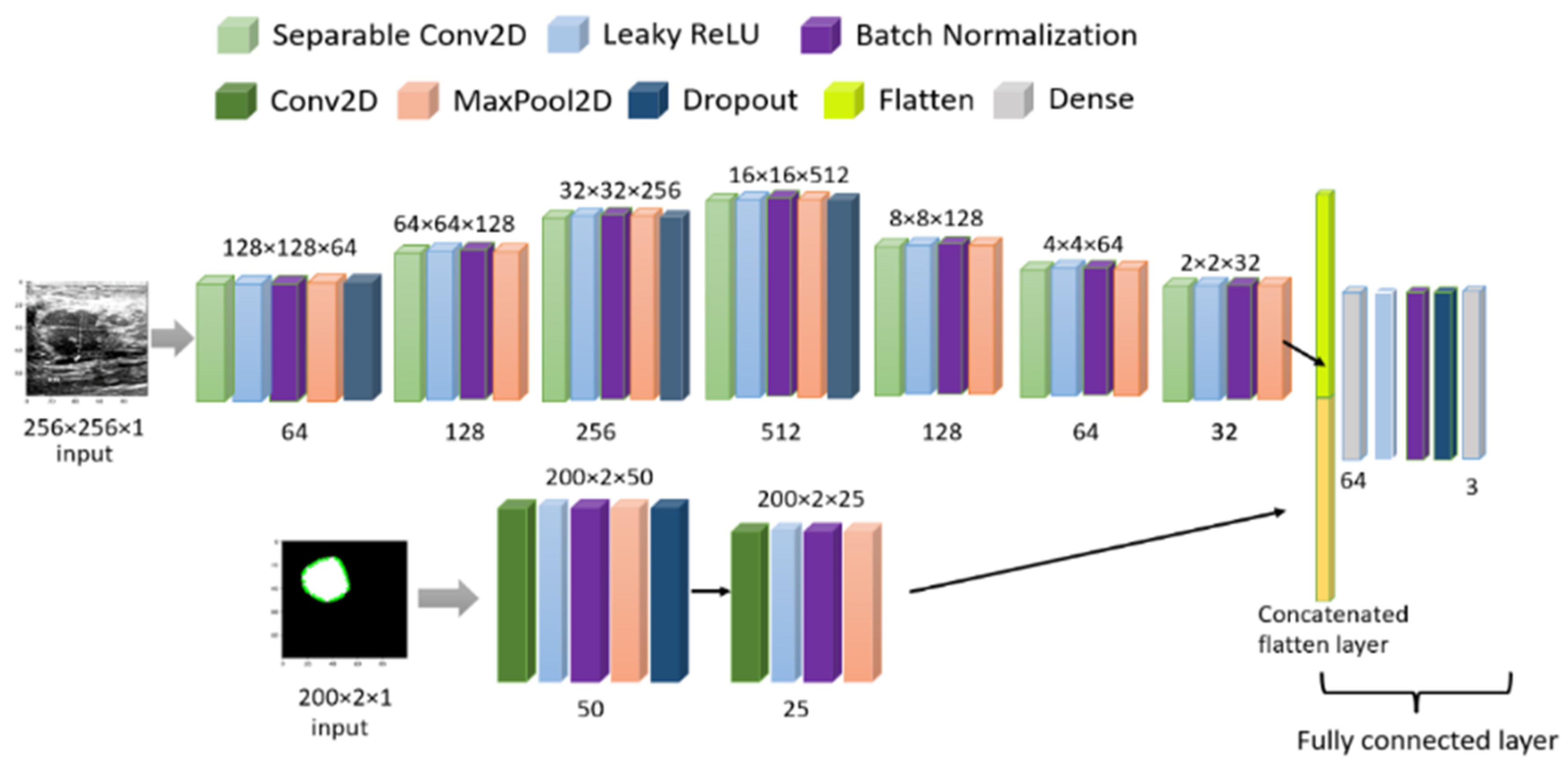
2.3.3. Validation of Multi-Headed CNN
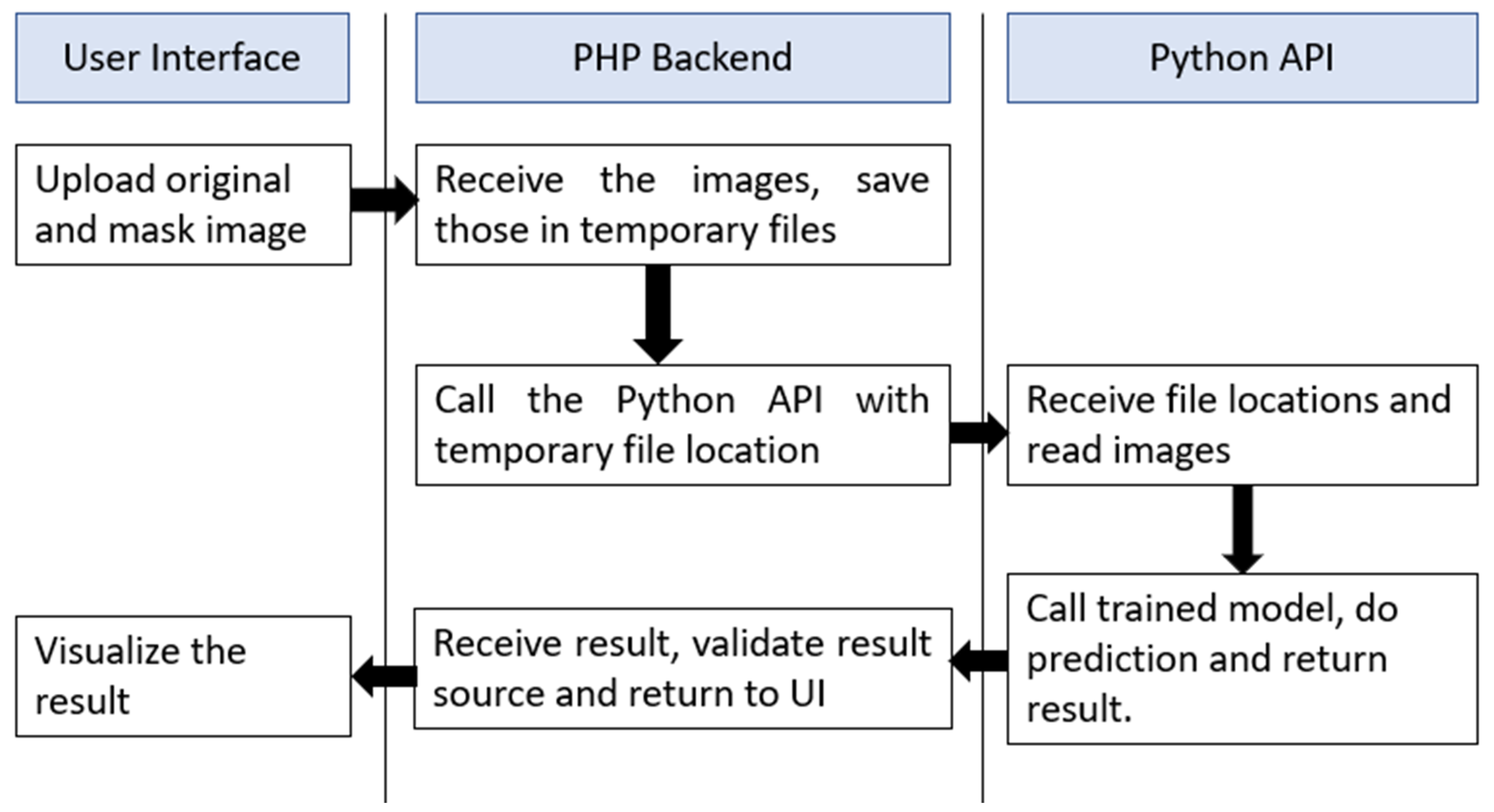
2.4. Web Application Interface
3. Discussing of Result
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Masud, M.; Rashed, A.E.E.; Hossain, M.S. Convolutional neural network-based models for diagnosis of breast cancer. Neural Comput. Appl. 2020, 34, 11383–11394. [Google Scholar] [CrossRef] [PubMed]
- Amin, M.N.; Uddin, G.; Uddin, N.; Rahaman, Z.; Siddiqui, S.A.; Hossain, S.; Islam, R.; Hasan, N.; Uddin, S.N. A hospital based survey to evaluate knowledge, awareness and perceived barriers regarding breast cancer screening among females in Bangladesh. Heliyon 2020, 6, e03753. [Google Scholar] [CrossRef] [PubMed]
- Begum, S.A.; Mahmud, T.; Rahman, T.; Zannat, J.; Khatun, F.; Nahar, K.; Towhida, M.; Joarder, M.; Harun, A.; Sharmin, F. Knowledge, Attitude and Practice of Bangladeshi Women towards Breast Cancer: A Cross Sectional Study. Mymensingh Med. J. 2019, 28, 96–104. Available online: http://europepmc.org/abstract/MED/30755557 (accessed on 19 March 2022). [PubMed]
- Islam, R.M.; Bell, R.J.; Billah, B.; Hossain, M.B.; Davis, S.R. Awareness of breast cancer and barriers to breast screening uptake in Bangladesh: A population based survey. Maturitas 2015, 84, 68–74. [Google Scholar] [CrossRef]
- Ilyas, F. Over 40,000 die of breast cancer every year in Pakistan. Dawn, 11 March 2017. [Google Scholar]
- Jemal, A.; Siegel, R.; Ward, E.; Hao, Y.; Xu, J.; Murray, T.; Thun, M.J. Cancer Statistics, 2008. CA A Cancer J. Clin. 2008, 58, 71–96. [Google Scholar] [CrossRef]
- Cheng, H.; Shi, X.; Min, R.; Hu, L.; Cai, X.; Du, H. Approaches for automated detection and classification of masses in mammograms. Pattern Recognit. 2006, 39, 646–668. [Google Scholar] [CrossRef]
- Cheng, H.; Cai, X.; Chen, X.; Hu, L.; Lou, X. Computer-aided detection and classification of microcalcifications in mammograms: A survey. Pattern Recognit. 2003, 36, 2967–2991. [Google Scholar] [CrossRef]
- Jesneck, J.L.; Lo, J.Y.; Baker, J.A. Breast Mass Lesions: Computer-aided Diagnosis Models with Mammographic and Sonographic Descriptors. Radiology 2007, 244, 390–398. [Google Scholar] [CrossRef]
- Shankar, P.M.; Piccoli, C.W.; Reid, J.M.; Forsberg, F.; Goldberg, B.B. Application of the compound probability density function for characterization of breast masses in ultrasound B scans. Phys. Med. Biol. 2005, 50, 2241–2248. [Google Scholar] [CrossRef]
- Taylor, K.J.; Merritt, C.; Piccoli, C.; Schmidt, R.; Rouse, G.; Fornage, B.; Rubin, E.; Georgian-Smith, D.; Winsberg, F.; Goldberg, B.; et al. Ultrasound as a complement to mammography and breast examination to characterize breast masses. Ultrasound Med. Biol. 2002, 28, 19–26. [Google Scholar] [CrossRef]
- Zhi, H.; Ou, B.; Luo, B.-M.; Feng, X.; Wen, Y.-L.; Yang, H.-Y. Comparison of Ultrasound Elastography, Mammography, and Sonography in the Diagnosis of Solid Breast Lesions. J. Ultrasound Med. 2007, 26, 807–815. [Google Scholar] [CrossRef] [PubMed]
- Chang, R.-F.; Wu, W.-J.; Moon, W.K.; Chen, D.-R. Improvement in breast tumor discrimination by support vector machines and speckle-emphasis texture analysis. Ultrasound Med. Biol. 2003, 29, 679–686. [Google Scholar] [CrossRef] [PubMed]
- Sahiner, B.; Chan, H.-P.; Roubidoux, M.A.; Hadjiiski, L.; Helvie, M.A.; Paramagul, C.; Bailey, J.; Nees, A.V.; Blane, C. Malignant and Benign Breast Masses on 3D US Volumetric Images: Effect of Computer-aided Diagnosis on Radiologist Accuracy. Radiology 2007, 242, 716–724. [Google Scholar] [CrossRef]
- Chen, C.-M.; Chou, Y.-H.; Han, K.-C.; Hung, G.-S.; Tiu, C.-M.; Chiou, H.-J.; Chiou, S.-Y. Breast Lesions on Sonograms: Computer-aided Diagnosis with Nearly Setting-Independent Features and Artificial Neural Networks. Radiology 2003, 226, 504–514. [Google Scholar] [CrossRef]
- Drukker, K.; Giger, M.; Horsch, K.; Kupinski, M.A.; Vyborny, C.J.; Mendelson, E.B. Computerized lesion detection on breast ultrasound. Med. Phys. 2002, 29, 1438–1446. [Google Scholar] [CrossRef]
- Andre, M.P.; Galperin, M.; Olson, L.K.; Richman, K.; Payrovi, S.; Phan, P. Improving the Accuracy of Diagnostic Breast Ultrasound. Acooustical Imaging 2002, 26, 453–460. [Google Scholar] [CrossRef]
- Meraj, T.; Alosaimi, W.; Alouffi, B.; Rauf, H.T.; Kumar, S.A.; Damaševičius, R.; Alyami, H. A quantization assisted U-Net study with ICA and deep features fusion for breast cancer identification using ultrasonic data. PeerJ Comput. Sci. 2021, 7, e805. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Aggarwal, A.; Choudhury, T. Breast Cancer Detection Using Machine Learning Algorithms. In Proceedings of the 2018 International Conference on Computational Techniques, Electronics and Mechanical Systems (CTEMS), Belgaum, India, 21–22 December 2018; pp. 114–118. [Google Scholar] [CrossRef]
- Ojha, U.; Goel, S. A study on prediction of breast cancer recurrence using data mining techniques. In Proceedings of the 2017 7th International Conference on Cloud Computing, Data Science & Engineering—Confluence, Noida, India, 12–13 January 2017; pp. 527–530. [Google Scholar] [CrossRef]
- Jabeen, K.; Khan, M.A.; Alhaisoni, M.; Tariq, U.; Zhang, Y.-D.; Hamza, A.; Mickus, A.; Damaševičius, R. Breast Cancer Classification from Ultrasound Images Using Probability-Based Optimal Deep Learning Feature Fusion. Sensors 2022, 22, 807. [Google Scholar] [CrossRef]
- Irfan, R.; Almazroi, A.; Rauf, H.; Damaševičius, R.; Nasr, E.; Abdelgawad, A. Dilated Semantic Segmentation for Breast Ultrasonic Lesion Detection Using Parallel Feature Fusion. Diagnostics 2021, 11, 1212. [Google Scholar] [CrossRef] [PubMed]
- Mekha, P.; Teeyasuksaet, N. Deep Learning Algorithms for Predicting Breast Cancer Based on Tumor Cells. In Proceedings of the 2019 Joint International Conference on Digital Arts, Media and Technology with ECTI Northern Section Conference on Electrical, Electronics, Computer and Telecommunications Engineering (ECTI DAMT-NCON), Nan, Thailand, 30 January–2 February 2019; pp. 343–346. [Google Scholar] [CrossRef]
- Chen, D.-R.; Kuo, W.-J.; Chang, R.-F.; Moon, W.K.; Lee, C.C. Use of the bootstrap technique with small training sets for computer-aided diagnosis in breast ultrasound. Ultrasound Med. Biol. 2002, 28, 897–902. [Google Scholar] [CrossRef]
- Malathi, M.; Sinthia, P.; Farzana, F.; Mary, G.A.A. Breast cancer detection using active contour and classification by deep belief network. Mater. Today Proc. 2021, 45, 2721–2724. [Google Scholar] [CrossRef]
- Abdelhafiz, D.; Bi, J.; Ammar, R.; Yang, C.; Nabavi, S. Convolutional neural network for automated mass segmentation in mammography. BMC Bioinform. 2020, 21, 192. [Google Scholar] [CrossRef]
- Hossain, M.S.; Muhammad, G.; Guizani, N. Explainable AI and Mass Surveillance System-Based Healthcare Framework to Combat COVID-I9 Like Pandemics. IEEE Netw. 2020, 34, 126–132. [Google Scholar] [CrossRef]
- Kumar, P.; Srivastava, S.; Mishra, R.K.; Sai, Y.P. End-to-end improved convolutional neural network model for breast cancer detection using mammographic data. J. Déf. Model. Simul. Appl. Methodol. Technol. 2020, 19, 375–384. [Google Scholar] [CrossRef]
- Yasmin, S.; Pathan, R.K.; Biswas, M.; Khandaker, M.U.; Faruque, M.R.I. Development of A Robust Multi-Scale Featured Local Binary Pattern for Improved Facial Expression Recognition. Sensors 2020, 20, 5391. [Google Scholar] [CrossRef]
- Masud, M.; Hossain, M.S.; Alhumyani, H.; Alshamrani, S.S.; Cheikhrouhou, O.; Ibrahim, S.; Muhammad, G.; Rashed, A.E.E.; Gupta, B.B. Pre-Trained Convolutional Neural Networks for Breast Cancer Detection Using Ultrasound Images. ACM Trans. Internet Technol. 2021, 21, 85. [Google Scholar] [CrossRef]
- Liu, J.; Li, W.; Zhao, N.; Cao, K.; Yin, Y.; Song, Q.; Chen, H.; Gong, X. Integrate Domain Knowledge in Training CNN for Ultrasonography Breast Cancer Diagnosis. Medical Image Computing and Computer Assisted Intervention—MICCAI 2018, Proceeding of the 21st International Conference, Granada, Spain 16–20 September 2018; Springer: Cham, Switzerland, 2018; pp. 868–875. [Google Scholar] [CrossRef]
- Nawaz, M.; Sewissy, A.A.; Soliman, T.H.A. Multi-Class Breast Cancer Classification using Deep Learning Convolutional Neural Network. Int. J. Adv. Comput. Sci. Appl. 2018, 9, 316–332. [Google Scholar] [CrossRef]
- Hijab, A.; Rushdi, M.A.; Gomaa, M.M.; Eldeib, A. Breast Cancer Classification in Ultrasound Images using Transfer Learning. In Proceedings of the 2019 5th International Conference on Advances in Biomedical Engineering (ICABME), Tripoli, Lebanon, 17–19 October 2019; pp. 1–4. [Google Scholar] [CrossRef]
- Sivanandan, R.; Jayakumari, J. A new CNN architecture for efficient classification of ultrasound breast tumor images with activation map clustering based prediction validation. Med. Biol. Eng. Comput. 2021, 59, 957–968. [Google Scholar] [CrossRef]
- Karthik, R.; Menaka, R.; Kathiresan, G.; Anirudh, M.; Nagharjun, M. Gaussian Dropout Based Stacked Ensemble CNN for Classification of Breast Tumor in Ultrasound Images. IRBM 2021, in press. [Google Scholar] [CrossRef]
- Wang, Y.; Choi, E.J.; Choi, Y.; Zhang, H.; Jin, G.Y.; Ko, S.-B. Breast Cancer Classification in Automated Breast Ultrasound Using Multiview Convolutional Neural Network with Transfer Learning. Ultrasound Med. Biol. 2020, 46, 1119–1132. [Google Scholar] [CrossRef]
- Casal-Guisande, M.; Comesaña-Campos, A.; Dutra, I.; Cerqueiro-Pequeño, J.; Bouza-Rodríguez, J.-B. Design and Development of an Intelligent Clinical Decision Support System Applied to the Evaluation of Breast Cancer Risk. J. Pers. Med. 2022, 12, 169. [Google Scholar] [CrossRef]
- Shirazi, A.Z.; Chabok, S.J.S.M.; Mohammadi, Z. A novel and reliable computational intelligence system for breast cancer detection. Med. Biol. Eng. Comput. 2017, 56, 721–732. [Google Scholar] [CrossRef] [PubMed]
- Kanimozhi, U.; Ganapathy, S.; Manjula, D.; Kannan, A. An Intelligent Risk Prediction System for Breast Cancer Using Fuzzy Temporal Rules. Natl. Acad. Sci. Lett. 2018, 42, 227–232. [Google Scholar] [CrossRef]
- Dua, D.; Graff, C. UCI Machine Learning Repository. 2017. Available online: http://archive.ics.uci.edu/ml (accessed on 17 January 2021).
- Al-Dhabyani, W.; Gomaa, M.; Khaled, H.; Fahmy, A. Dataset of breast ultrasound images. Data Brief 2019, 28, 104863. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Gkioxari, G.; Dollár, P.; Girshick, R. Mask r-cnn. In Proceedings of the IEEE International Conference on Computer Vision, Venice, Italy, 22–29 October 2017; pp. 2961–2969. [Google Scholar]
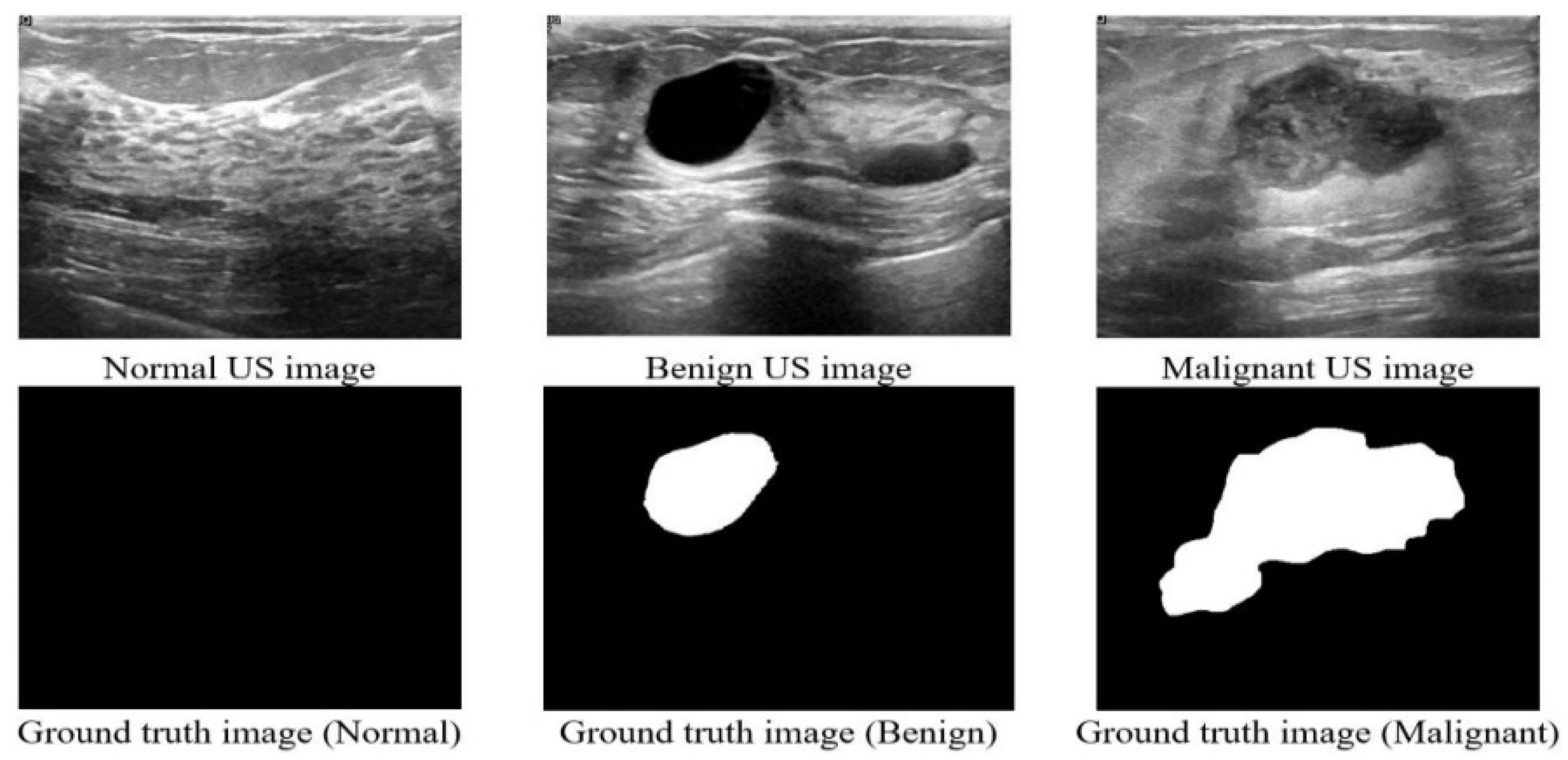







| Type | Number of Images | Shape | Color | Format |
|---|---|---|---|---|
| Benign | 487 | 500 × 500 Pixel | Grayscale | png |
| Malignant | 210 | |||
| Normal | 133 |
| Precision | Recall | F1 Score | Support | |
|---|---|---|---|---|
| Benign | 0.80 | 0.88 | 0.84 | 106 |
| Malignant | 0.70 | 0.56 | 0.62 | 54 |
| Normal | 0.97 | 1.00 | 0.99 | 35 |
| accuracy | 0.81 | 195 | ||
| macro avg | 0.82 | 0.81 | 0.81 | 195 |
| weighted avg | 0.80 | 0.81 | 0.80 | 195 |
| Precision | Recall | F1 Score | Support | |
|---|---|---|---|---|
| Benign | 0.73 | 0.98 | 0.84 | 106 |
| Malignant | 1.00 | 0.63 | 0.77 | 54 |
| Normal | 0.89 | 0.46 | 0.60 | 35 |
| accuracy | 0.79 | 195 | ||
| macro avg | 0.87 | 0.69 | 0.74 | 195 |
| weighted avg | 0.83 | 0.79 | 0.78 | 195 |
| Layer (Type) | Output Shape | Param | Connected to |
|---|---|---|---|
| “input_2 (InputLayer)” | [(None, 256, 256, 1)] | 0 | [] |
| “separable_conv2d (SeparableConv2D)” | (None, 256, 256, 64) | 137 | “[input_2[0][0]]” |
| “leaky_re_lu_5 (LeakyReLU)” | (None, 256, 256, 64) | 0 | “[separable_conv2d [0][0]]” |
| “batch_normalization_4 (BatchNormalization)” | (None, 256, 256, 64) | 256 | “[leaky_re_lu_5[0][0]]” |
| “max_pooling2d_5 (MaxPooling2D)” | (None, 128, 128, 64) | 0 | “[batch_normalization_4[0][0]]” |
| “dropout_3 (Dropout)” | (None, 128, 128, 64) | 0 | “[max_pooling2d_5[0][0]]” |
| “separable_conv2d_1 (SeparableConv2D)” | (None, 128, 128, 128) | 8896 | “[dropout_3[0][0]]” |
| “leaky_re_lu_6 (LeakyReLU)” | (None, 128, 128, 128) | 0 | “[separable_conv2d_1[0][0]]” |
| “batch_normalization_5 (BatchNormalization)” | (None, 128, 128, 128) | 512 | “[leaky_re_lu_6[0][0]]” |
| “input_1 (InputLayer)” | [(None, 200, 2, 1)] | 0 | [] |
| “max_pooling2d_6 (MaxPooling2D)” | (None, 64, 64, 128) | 0 | “[batch_normalization_5[0][0]]” |
| “conv2d_2 (Conv2D)” | (None, 200, 2, 50) | 500 | “[input_1[0][0]]” |
| “separable_conv2d_5 (SeparableConv2D)” | (None, 64, 64, 64) | 9408 | “[max_pooling2d_6[0][0]]” |
| “leaky_re_lu_3 (LeakyReLU)” | (None, 200, 2, 50) | 0 | “[conv2d_2[0][0]]” |
| “leaky_re_lu_10 (LeakyReLU)” | (None, 64, 64, 64) | 0 | “[separable_conv2d_5[0][0]]” |
| “batch_normalization_2 (BatchNormalization)” | (None, 200, 2, 50) | 200 | “[leaky_re_lu_3[0][0]]” |
| “batch_normalization_9 (BatchNormalization)” | (None, 64, 64, 64) | 256 | “[leaky_re_lu_10[0][0]]” |
| “max_pooling2d_3 (MaxPooling2D)” | (None, 200, 2, 50) | 0 | “[batch_normalization_2[0][0]]” |
| “max_pooling2d_10 (MaxPooling2D)” | (None, 32, 32, 64) | 0 | “[batch_normalization_9[0][0]]” |
| “dropout_2 (Dropout)” | (None, 200, 2, 50) | 0 | “[max_pooling2d_3[0][0]]” |
| “separable_conv2d_6 (SeparableConv2D)” | (None, 32, 32, 32) | 2656 | “[max_pooling2d_10[0][0]]” |
| “conv2d_3 (Conv2D)” | (None, 200, 2, 25) | 11275 | “[dropout_2[0][0]]” |
| “leaky_re_lu_11 (LeakyReLU)” | (None, 32, 32, 32) | 0 | “[separable_conv2d_6[0][0]]” |
| “leaky_re_lu_4 (LeakyReLU)” | (None, 200, 2, 25) | 0 | “[conv2d_3[0][0]]” |
| “batch_normalization_10 (BatchNormalization)” | (None, 32, 32, 32) | 128 | “[leaky_re_lu_11[0][0]]” |
| “batch_normalization_3 (BatchNormalization)” | (None, 200, 2, 25) | 100 | “[leaky_re_lu_4[0][0]]” |
| “max_pooling2d_11 (MaxPooling2D)” | (None, 16, 16, 32) | 0 | “[batch_normalization_10[0][0]]” |
| “max_pooling2d_4 (MaxPooling2D)” | (None, 200, 2, 25) | 0 | “[batch_normalization_3[0][0]]” |
| “dropout_6 (Dropout)” | (None, 16, 16, 32) | 0 | “[max_pooling2d_11[0][0]]” |
| “flatten_1 (Flatten)” | (None, 10000) | 0 | “[max_pooling2d_4[0][0]]” |
| “flatten_2 (Flatten)” | (None, 8192) | 0 | “[dropout_6[0][0]]” |
| “concatenate (Concatenate)” | (None, 18192) | 0 | “[flatten_1[0][0],flatten_2[0][0]]” |
| “dense_2 (Dense)” | (None, 64) | 1164352 | “[concatenate[0][0]]” |
| “leaky_re_lu_12 (LeakyReLU)” | (None, 64) | 0 | “[dense_2[0][0]]” |
| “batch_normalization_11 (BatchNormalization)” | (None, 64) | 256 | “[leaky_re_lu_12[0][0]]” |
| “dropout_7 (Dropout)” | (None, 64) | 0 | “[batch_normalization_11[0][0]]” |
| “dense_3 (Dense)” | (None, 3) | 195 | “[dropout_7[0][0]]” |
| Total params: 1,199,127 Trainable params: 1,198,273 Non-trainable params: 854 | |||
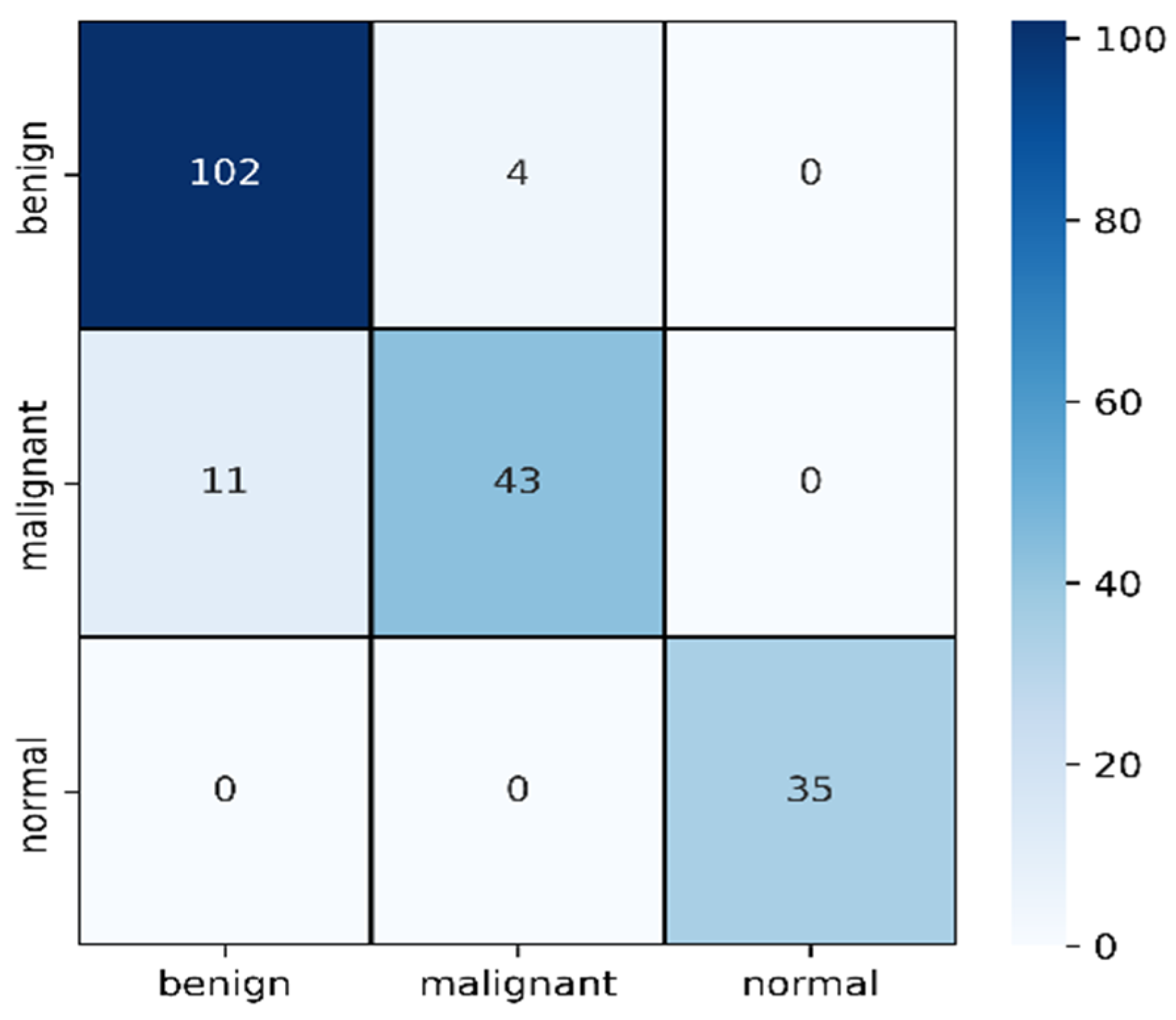
| Precision | Recall | F1 Score | Support | |
|---|---|---|---|---|
| Benign | 0.90 | 0.96 | 0.93 | 106 |
| Malignant | 0.91 | 0.80 | 0.85 | 54 |
| Normal | 1.00 | 1.00 | 1.00 | 35 |
| accuracy | 0.92 | 195 | ||
| macro avg | 0.94 | 0.92 | 0.93 | 195 |
| weighted avg | 0.92 | 0.92 | 0.92 | 195 |
| References | Method | Datasets | Findings |
|---|---|---|---|
| [20] | Random Forest, KNN and Naïve Bayes. | Wisconsin Breast Cancer dataset from UCI Repository | KNN was an excellent classifier in terms of accuracy of 94%. |
| [19] | Data Mining techniques | Wisconsin breast cancer dataset. | Detect hidden cancer-associated for classification SVM accuracy 81%. |
| [23] | Machine learning techniques | Mammogram images. | It achieved 96% accuracy by using DNN |
| [32] | Deep learning | Wisconsin breast cancer dataset. | The author compared machine learning techniques and deep learning. It achieved 96.99% accuracy with DL. |
| [24] | CAD system | Custom ultrasound images | Accuracy with 10 samples and the bootstrap method is 87.07% |
| [25] | CAD system | Mammogram images. | Random forest gives the highest accuracy of 97.51% |
| [31] | Multi-task CNN | Custom ultrasound images | With a baseline of 83.3% |
| [32] | Multi-Class CNN | BreaKHis dataset | Image level accuracy 95.4% |
| [33] | VGG16 and transfer learning | BUSI | Fined tuned model accuracy is 97% |
| Proposed Method | Multi-headed CNN | BUSI | Multiple types of datasets could generate more significant results than solo datasets. Accuracy is 92.31% (±2) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pathan, R.K.; Alam, F.I.; Yasmin, S.; Hamd, Z.Y.; Aljuaid, H.; Khandaker, M.U.; Lau, S.L. Breast Cancer Classification by Using Multi-Headed Convolutional Neural Network Modeling. Healthcare 2022, 10, 2367. https://doi.org/10.3390/healthcare10122367
Pathan RK, Alam FI, Yasmin S, Hamd ZY, Aljuaid H, Khandaker MU, Lau SL. Breast Cancer Classification by Using Multi-Headed Convolutional Neural Network Modeling. Healthcare. 2022; 10(12):2367. https://doi.org/10.3390/healthcare10122367
Chicago/Turabian StylePathan, Refat Khan, Fahim Irfan Alam, Suraiya Yasmin, Zuhal Y. Hamd, Hanan Aljuaid, Mayeen Uddin Khandaker, and Sian Lun Lau. 2022. "Breast Cancer Classification by Using Multi-Headed Convolutional Neural Network Modeling" Healthcare 10, no. 12: 2367. https://doi.org/10.3390/healthcare10122367
APA StylePathan, R. K., Alam, F. I., Yasmin, S., Hamd, Z. Y., Aljuaid, H., Khandaker, M. U., & Lau, S. L. (2022). Breast Cancer Classification by Using Multi-Headed Convolutional Neural Network Modeling. Healthcare, 10(12), 2367. https://doi.org/10.3390/healthcare10122367









