Ensemble Technique Coupled with Deep Transfer Learning Framework for Automatic Detection of Tuberculosis from Chest X-ray Radiographs
Abstract
1. Introduction
- A robust framework for automatic lung segmentation and TB classification based on chest X-ray images is proposed for the early detection of tuberculosis.
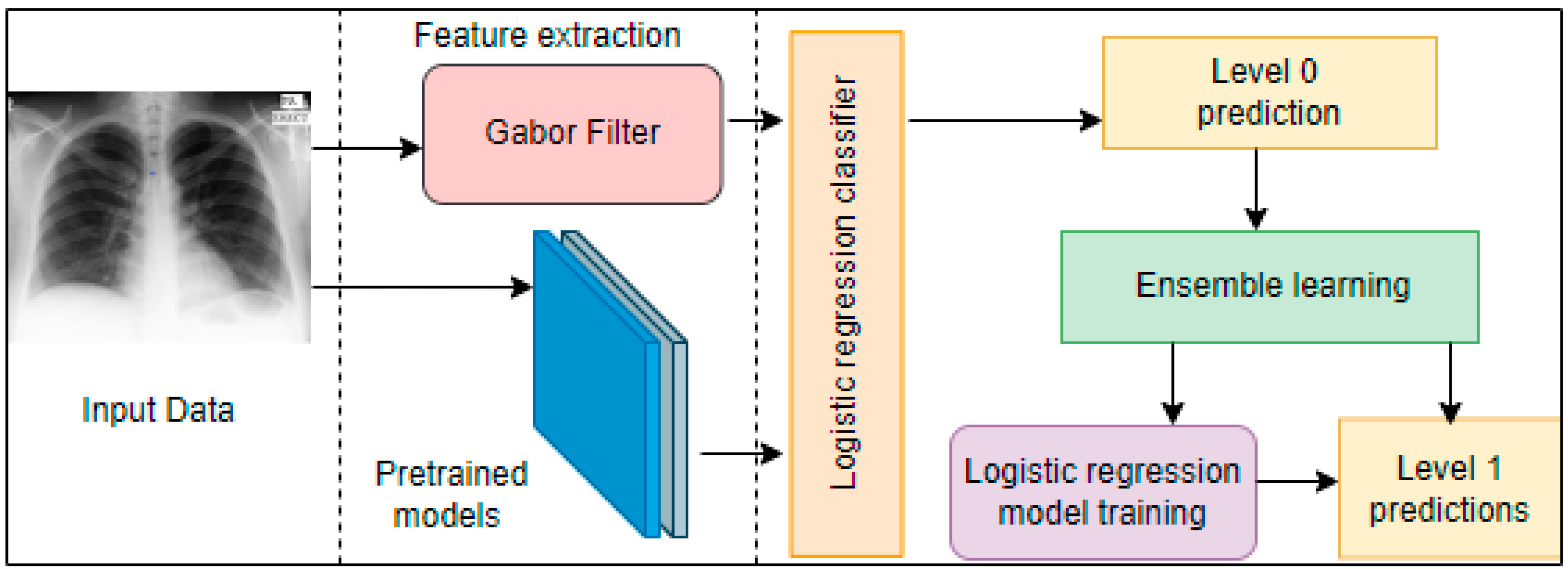
- A combined result from a customized CNN and pre-trained CNN models through stacking ensemble learning is deployed to boost classification accuracy.
- The proposed framework achieved a higher accuracy rate than other state-of-the-art TB detection models, which suggests that our model is better for mass TB screening in regions where TB is much more predominant.
1.1. Background
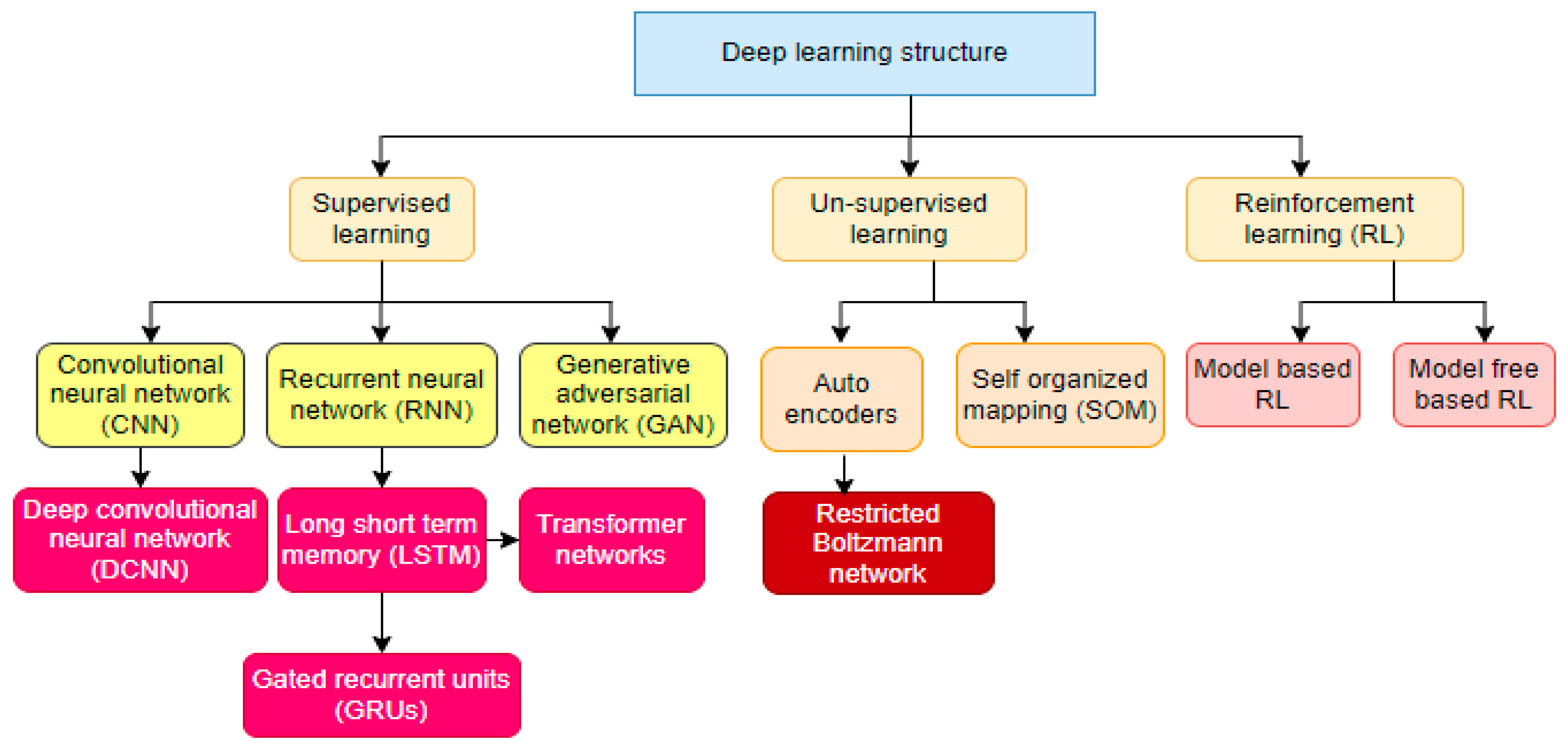
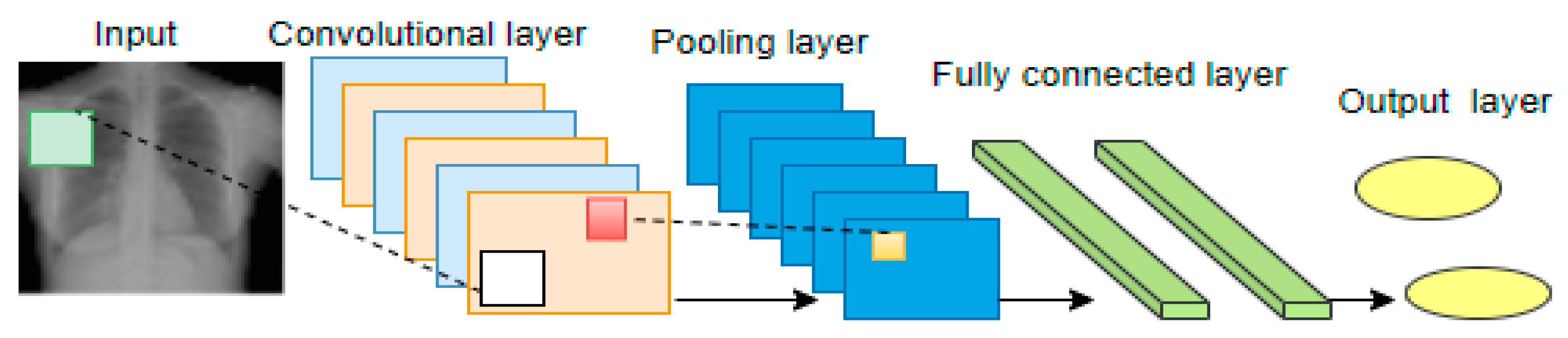
1.1.1. Deep Learning
1.1.2. Support Vector Machine
1.1.3. Deep Transfer Learning
1.1.4. U-Net
2. Related Work
3. Materials and Methods
3.1. Dataset
3.2. Image Pre-Processing
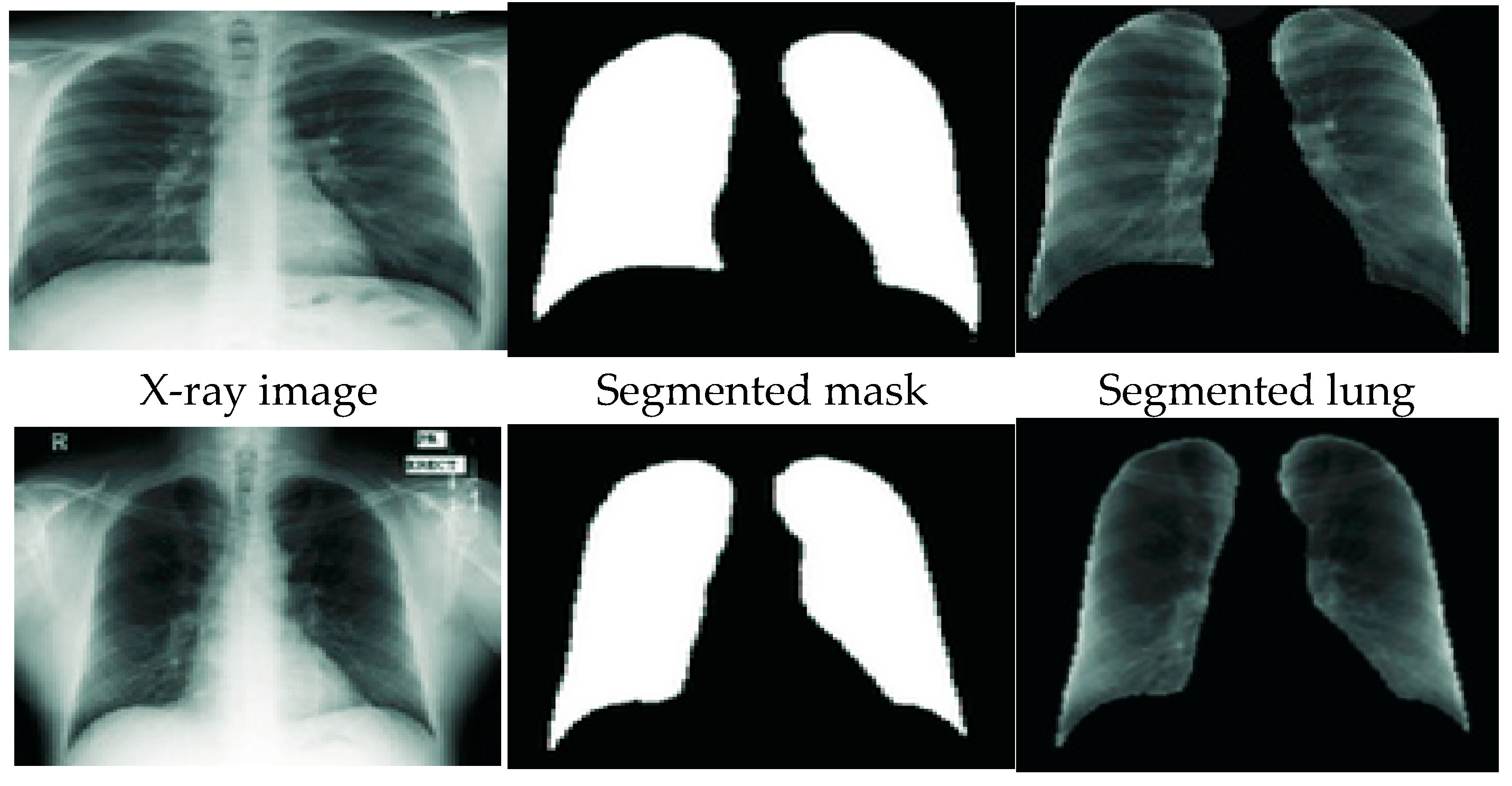
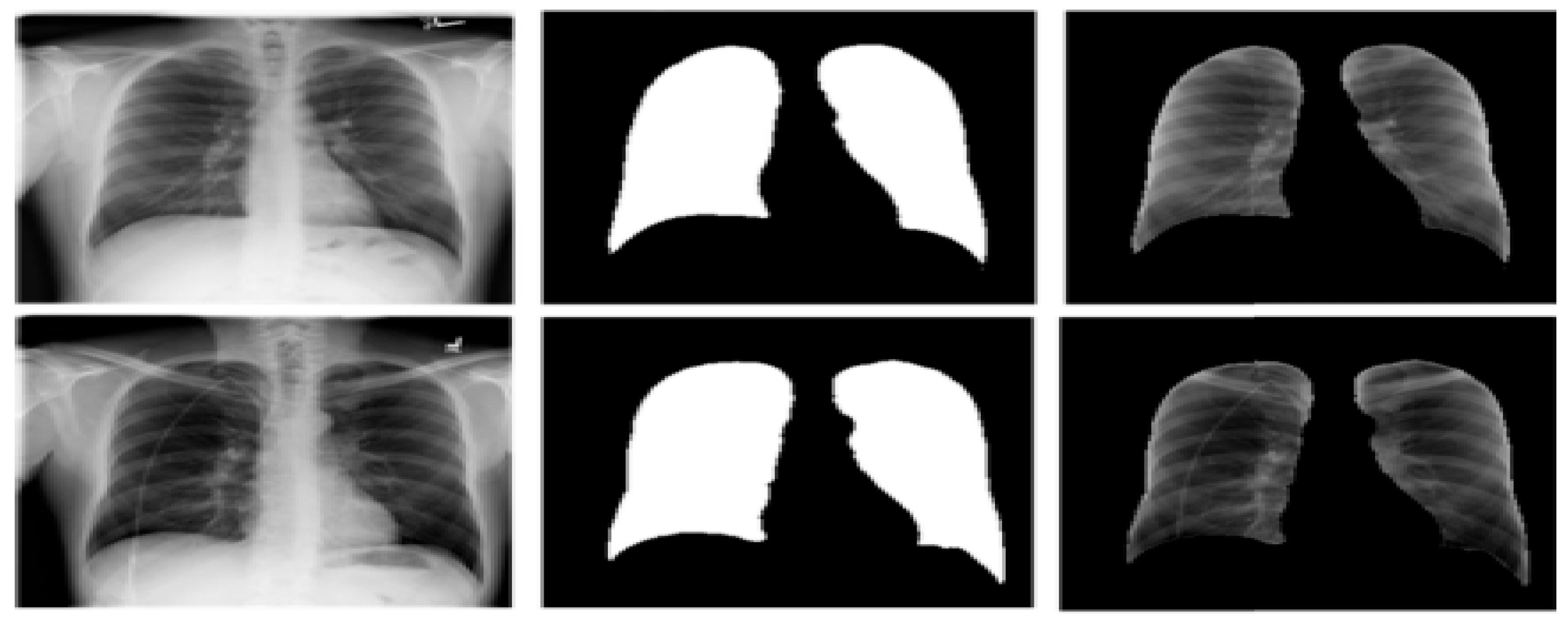
3.3. Lung Segmentation
3.4. Feature Extraction Based on CNN and Pre-Trained CNN Models
3.5. Classification
| Algorithm 1. Stack Ensemble |
| Input: Segmented images () = {(xi, yi) | xi X, yi } Output: Ensemble classifier (Ec) Step 1: Train base - models () from segmented images () For p 1 P do Train base model base on Aggregate obtained predictions from all 8 base models Step 2: Create a new dataset () from base model predictions. For n 1 to z do Create new dataset comprising {xi, yi}, where xi = {j (xi) for j = 1 − 8} Step 3: Train a second-level meta learner Learn a new classifier Ec based on newly created dataset Return Es(x) = es(es1(x), es2(x)… es8 (x)) |
4. Results and Analysis
5. Discussion
6. Conclusions
Future Work
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- World Health Organization. Global Tuberculosis Report; WHO: Geneva, Switzerland, 2020. [Google Scholar]
- World Health Organization. Global Tuberculosis Report; WHO: Geneva, Switzerland, 2021. [Google Scholar]
- Ryu, Y.J. Diagnosis of pulmonary tuberculosis: Recent advances and diagnostic algorithms. Tuberc. Respir. Dis. 2015, 78, 64–71. [Google Scholar] [CrossRef] [PubMed]
- Lo, C.-M.; Wu, Y.-H.; Li, Y.-C.; Lee, C.-C. Computer-aided bacillus detection in whole-slide pathological images using a deep convolutional neural network. Appl. Sci. 2020, 10, 4059. [Google Scholar] [CrossRef]
- Chang, R.I.; Chiu, Y.H.; Lin, J.W. Two-stage classification of tuberculosis culture diagnosis using convolutional neural network with transfer learning. J. Supercomput. 2020, 76, 8641–8656. [Google Scholar] [CrossRef]
- Swetha, K.; Sankaragomathi, B.; Thangamalar, J.B. Convolutional neural network based automated detection of mycobacterium bacillus from sputum images. In Proceedings of the 5th International Conference on Inventive Computation Technologies, ICICT 2020, Lalitpur, Nepal, 26–28 February 2020; pp. 293–300. [Google Scholar] [CrossRef]
- Verma, D.; Bose, C.; Tufchi, N.; Pant, K.; Tripathi, V.; Thapliyal, A. An efficient framework for identification of tuberculosis and pneumonia in chest x-ray images using neural network. Procedia Comput. Sci. 2020, 171, 217–224. [Google Scholar] [CrossRef]
- Msonda, P.; Uymaz, S.A.; Karaağaç, S.S. Spatial pyramid pooling in deep convolutional networks for automatic tuberculosis diagnosis. Trait. Signal 2020, 37, 1075–1084. [Google Scholar] [CrossRef]
- Singh, J.; Tripathy, A.; Garg, P.; Kumar, A. Lung tuberculosis detection using anti-aliased convolutional networks. Procedia Comput. Sci. 2020, 173, 281–290. [Google Scholar] [CrossRef]
- Bradley, W.G. History of medical imaging. Proc. Am. Philos. Soc. 2015, 152, 349–361. [Google Scholar]
- Barani, M.; Mukhtar, M.; Rahdar, A.; Sargazi, S.; Pandey, S.; Kang, M. Recent advances in nanotechnology-based diagnosis and treatments of human osteosarcoma. Biosensors 2021, 11, 55. [Google Scholar] [CrossRef]
- Douche, D.; Sert, Y.; Brandan, S.A.; Kawther, A.A. 5-((1H-imidazol-1-yl) methyl) quinolin-8-ol as potential antiviral SARS-CoV-2 candidate: Synthesis, crystal structure, Hirshfeld surface analysis, DFT and molecular docking studies. J. Mol. Struct. 2021, 1232, 130005. [Google Scholar] [CrossRef]
- Gümüş, M.; Babacan, Ş.N.; Demir, Y.; Sert, Y.; Koca, İ.; Gülçin, İ. Discovery of sulfadrug–pyrrole conjugates as carbonic anhydrase and acetylcholinesterase inhibitors. Arch. Pharm. 2022, 355, e2100242. [Google Scholar] [CrossRef]
- Murphy, K.; Habib, S.S.; Asad Zaidi, S.M.; Khowaja, S.; Khan, A.; Melendez, J.; Scholten, E.T.; Amad, F.; Schalekamp, S.; Verhagen, M.; et al. Computer aided detection of tuberculosis on chest radiographs: An evaluation of the CAD4TB v6 system. Sci. Rep. 2020, 10, 5492. [Google Scholar] [CrossRef] [PubMed]
- Dou, Q.; Chen, H.; Yu, L.; Zhao, L.; Qin, J.; Wang, D.; Mok, V.C.T.; Shi, L.; Heng, P.-A. Automatic detection of cerebral microbleeds from MR Images via 3D convolutional neural networks. IEEE Trans. Med. Imaging 2016, 35, 1182–1195. [Google Scholar] [CrossRef] [PubMed]
- Abbas, A.; Abdelsamea, M.M.; Gaber, M.M. DeTrac: Transfer learning of class decomposed medical images in convolutional neural networks. IEEE Access 2020, 8, 74901–74913. [Google Scholar] [CrossRef]
- Zech, J.R.; Badgeley, M.A.; Liu, M.; Costa, A.B.; Titano, J.J.; Oermann, E.K. Variable generalization performance of a deep learning model to detect pneumonia in chest radiographs: A cross-sectional study. PLoS Med. 2018, 15, e1002683. [Google Scholar] [CrossRef] [PubMed]
- Shin, H.C.; Roberts, K.; Lu, L.; Demner-Fushman, D.; Yao, J.; Summers, R.M. Learning to read chest x-rays: Recurrent neural cascade model for automated image annotation. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 26 June–1 July 2016; pp. 2497–2506. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. Lect. Notes Comput. Sci. 2015, 9351, 234–241. [Google Scholar] [CrossRef]
- Kotei, E.; Thirunavukarasu, R. Computational techniques for the automated detection of mycobacterium tuberculosis from digitized sputum smear microscopic images: A systematic review. Prog. Biophys. Mol. Biol. 2022, 171, 4–16. [Google Scholar] [CrossRef]
- Jaeger, S.; Candemir, S.; Antani, S.; Wáng, Y.-X.J.; Lu, P.-X.; Thoma, G. Two public chest X-ray datasets for computer-aided screening of pulmonary diseases. Quant. Imaging Med. Surg. 2014, 4, 475–477. [Google Scholar]
- Stirenko, S.; Kochura, Y.; Alienin, O.; Rokovyj, O.; Gang, P.; Zeng, W.; Gordienko, Y.G. Chest x-ray analysis of tuberculosis by deep learning with segmentation and augmentation. In Proceedings of the 2018 IEEE 38th International Conference on Electronics and Nanotechnology, ELNANO 2018, Kyiv, Ukraine, 24–26 April 2018; pp. 422–428. [Google Scholar] [CrossRef]
- Firmino, M.; Angelo, G.; Morais, H.; Dantas, M.R.; Valentim, R. Computer-aided detection (CADe) and diagnosis (CADx) system for lung cancer with likelihood of malignancy. Biomed. Eng. Online 2016, 15, 2. [Google Scholar] [CrossRef]
- Brownlee, J. Machine Learning Mastery. Available online: https://machinelearningmastery.com/what-is-deep-learning/ (accessed on 10 June 2021).
- Fioravanti, D.; Giarratano, Y.; Maggio, V.; Agostinelli, C.; Chierici, M.; Jurman, G.; Furlanello, C. Phylogenetic convolutional neural networks in metagenomics. BMC Bioinform. 2018, 19 (Suppl. 2). [Google Scholar] [CrossRef]
- Grisoni, F.; Moret, M.; Lingwood, R.; Schneider, G. Bidirectional molecule generation with recurrent neural networks. J. Chem. Inf. Model. 2020, 60, 1175–1183. [Google Scholar] [CrossRef]
- Méndez-lucio, O.; Baillif, B.; Clevert, D.; Rouquié, D.; Wichard, J. De novo generation of hit-like molecules from gene expression signatures using artificial intelligence. Nat. Commun. 2020, 11, 10. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Liu, Q.; Xie, W.; Hu, P. YOLO-LOGO: A transformer-based YOLO segmentation model for breast mass detection and segmentation in digital mammograms. Comput. Methods Programs Biomed. 2022, 221, 106903. [Google Scholar] [CrossRef] [PubMed]
- Thai, L.H.; Hai, T.S.; Thuy, N.T. Image Classification using Support Vector Machine and artificial neural network. Int. J. Inf. Technol. Comput. Sci. 2012, 4, 32–38. [Google Scholar] [CrossRef]
- Pandit, M.K.; Banday, S.A.; Naaz, R.; Chishti, M.A. Automatic detection of COVID-19 from chest radiographs using deep learning. Radiogr. J. 2020, 27, 483–489. [Google Scholar] [CrossRef] [PubMed]
- Sitaula, C.; Hossain, M.B. Attention-based VGG-16 model for COVID-19 chest X-ray image classification. Appl. Intell. 2021, 51, 2850–2863. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, A.; Latief, J.; Mudasir, M. CoroNet: A deep neural network for detection and diagnosis of COVID-19 from chest x-ray images. Comput. Methods Programs Biomed. 2020, 196, 105581. [Google Scholar] [CrossRef]
- Ozturk, T.; Talo, M.; Yildirim, E.A.; Baloglu, U.B.; Yildirim, O.; Acharya, U.R. Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput. Biol. Med. 2020, 121, 103792. [Google Scholar] [CrossRef] [PubMed]
- Showkatian, E.; Salehi, M.; Ghaffari, H.; Reiazi, R.; Sadighi, N. Deep learning-based automatic detection of tuberculosis disease in chest X-ray images. Pol. J. Radiol. 2022, 87, 118–124. [Google Scholar] [CrossRef]
- Priya, E.; Srinivasan, S. Automated object and image level classification of TB images using support vector neural network classifier. Biocybern. Biomed. Eng. 2016, 36, 670–678. [Google Scholar] [CrossRef]
- Ahmad, T.; Lund, L.H.; Rao, P.; Ghosh, R.; Warier, P.; Vaccaro, B.; Dahlstrom, U.; O’Connor, C.M.; Felker, G.M.; Desai, N.R. Machine learning methods improve prognostication, identify clinically distinct phenotypes, and detect heterogeneity in response to therapy in a large cohort of heart failure patients. J. Am. Heart Assoc. 2018, 7, e008081. [Google Scholar] [CrossRef]
- Sankaran, A.; Jain, A.; Vashisth, T.; Vatsa, M.; Singh, R. Adaptive latent fingerprint segmentation using feature selection and random decision forest classification. Inf. Fusion 2017, 34, 1–15. [Google Scholar] [CrossRef]
- Hooda, R.; Sofat, S.; Kaur, S.; Mittal, A.; Meriaudeau, F. Deep-learning: A potential method for tuberculosis detection using chest radiography. In Proceedings of the 2017 IEEE International Conference on Signal and Image Processing Applications, ICSIPA 2017, Kuching, Malaysia, 12–14 September 2017; pp. 497–502. [Google Scholar] [CrossRef]
- Hwang, E.J.; Park, S.; Jin, K.-N.; Kim, J.I.; Choi, S.Y.; Lee, J.H.; Goo, J.M.; Aum, J.; Yim, J.-J.; Parl, C.M.; et al. Development and validation of a deep learning-based automatic detection algorithm for active pulmonary tuberculosis on chest radiographs. Clin. Infect. Dis. 2019, 69, 739–747. [Google Scholar] [CrossRef] [PubMed]
- Devnath, L.; Luo, S.; Summons, P.; Wang, D. Tuberculosis classification in chest radiographs using deep convolutional neural networks. Int. J. Adv. Sci. Eng. Technol. 2018. [Google Scholar]
- Cao, Y.; Liu, C.; Liu, B.; Brunette, M.J.; Zhang, N.; Sun, T.; Zhang, P.; Peinado, J.; Garavito, E.S.; Garcia, L.L.; et al. Improving tuberculosis diagnostics using deep learning and mobile health technologies among resource-poor and marginalized communities. In Proceedings of the 2016 IEEE 1st International Conference on Connected Health: Applications, Systems and Engineering Technologies, CHASE 2016, Washington, DC, USA, 27–29 June 2016; pp. 274–281. [Google Scholar] [CrossRef]
- Liu, C.; Cao, Y.; Alcantara, M.; Liu, B.; Brunette, M.J.; Peinado, J.; Curioso, W. TX-CNN: Detecting tuberculosis in chest x-ray images using convolutional neural network. In Proceedings of the International Conference on Image Processing, ICIP, Beijing, China, 17–20 September 2017; pp. 2314–2318. [Google Scholar] [CrossRef]
- Heo, S.J.; Kim, Y.; Yun, S.; Lim, S.-S.; Kim, J.; Nam, C.-M.; Park, E.-C.; Jung, I.; Yoon, J.-H. Deep learning algorithms with demographic information help to detect tuberculosis in chest radiographs in annual workers’ health examination data. Int. J. Environ. Res. Public Health 2019, 16, 250. [Google Scholar] [CrossRef] [PubMed]
- Hwang, E.J.; Park, S.; Jin, K.-N.; Kim, J.I.; Choi, S.Y.; Lee, J.H.; Goo, J.M.; Aum, J.; Yim, J.-J.; Cohen, J.G.; et al. Development and validation of a deep learning-based automated detection algorithm for major thoracic diseases on chest radiograph. JAMA Netw. Open 2019, 2, e191095. [Google Scholar] [CrossRef]
- Gozes, O.; Greenspan, H. Deep feature learning from a hospital-scale chest x-ray dataset with application to TB detection on a small-scale dataset. In Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, EMBS, Berlin, Germany, 23–27 July 2019; pp. 4076–4079. [Google Scholar] [CrossRef]
- Pasa, F.; Golkov, V.; Pfeiffer, F.; Cremers, D.; Pfeiffer, D. Efficient deep network architectures for fast chest x-ray tuberculosis screening and visualization. Sci. Rep. 2019, 9, 2–10. [Google Scholar] [CrossRef]
- Rohilla, A.; Hooda, R.; Mittal, A. TB detection in chest radiograph using deep learning architecture. Int. J. Adv. Res. Sci. Eng. 2017, 6, 1073–1084. [Google Scholar]
- Yadav, O.; Passi, K.; Jain, C.K. Using Deep learning to classify x-ray Images of potential tuberculosis patients. In Proceedings of the 2018 IEEE International Conference on Bioinformatics and Biomedicine, BIBM 2018, San Diego, CA, USA, 18–21 November 2019; pp. 2368–2375. [Google Scholar] [CrossRef]
- Akbar, S.; GhaniHaider, N.; Tariq, H. Tuberculosis diagnosis using x-ray images. Int. J. Adv. Res. 2019, 7, 689–696. [Google Scholar] [CrossRef]
- Devnath, L.; Luo, S.; Summons, P.; Wang, D. Automated detection of pneumoconiosis with multilevel deep features learned from chest X-ray radiographs. Comput. Biol. Med. 2021, 129, 104125. [Google Scholar] [CrossRef] [PubMed]
- Rajpurkar, P.; Irvin, J.; Zhu, K.; Yang, B.; Mehta, H.; Duan, T.; Ding, D.; Bagul, A.; Langlotz, C.; Shpanskaya, K.; et al. CheXNet: Radiologist-Level Pneumonia Detection on Chest X-rays with Deep Learning. arXiv 2017, arXiv:1711.05225. [Google Scholar]
- CDC (Center for Diseases Control and Prevention). Available online: https://www.cdc.gov/niosh/topics/cwhsp/cwhsp-xray.html (accessed on 10 March 2022).
- Rahman, T.; Khandakar, A.; Abdul Kadir, M.; Islam, K.R.; Islam, K.F.; Mazhar, R.; Hamid, T.; Islam, M.T.; Kashem, S.; Mahbub, Z.B.; et al. Reliable tuberculosis detection using chest x-ray with deep learning, segmentation and visualization. IEEE Access 2020, 8, 191586–191601. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; Volume 2016, pp. 770–778. [Google Scholar] [CrossRef]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the inception architecture for computer vision. Proc. IEEE Comput. Soc. Conf. Comput. Vis. Pattern Recognit. 2016, 2016, 2818–2826. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. In Proceedings of the 3rd International Conference on Learning Representations, ICLR 2015—Conference Track Proceedings, San Diego, CA, USA, 7–9 May 2015; pp. 1–14. [Google Scholar]
- Huang, G.; Liu, Z.; van der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the 30th IEEE Computer Vision and Pattern Recognition, CVPR, Honolulu, HI, USA, 21–26 July 2017; Volume 2017, pp. 2261–2269. [Google Scholar] [CrossRef]
- Iandola, F.N.; Han, S.; Moskewicz, M.W.; Ashraf, K.; Dally, W.J.; Keutzer, K. SqueezeNet: AlexNet-level accuracy with 50× fewer parameters and <0.5 MB model size. arXiv 2016, arXiv:1602.07360v4. [Google Scholar]
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. MobileNets: Efficient Convolutional Neural Networks for Mobile Vision Applications. arXiv 2017, arXiv:1704.04861. [Google Scholar]
- Guo, R.; Passi, K.; Jain, C.K. Tuberculosis diagnostics and localization in chest x-rays via deep learning models. Front. Artif. Intell. 2020, 3, 583427. [Google Scholar] [CrossRef] [PubMed]
- Abideen, Z.U.; Ghafoor, M.; Munir, K.; Saqib, M.; Ullah, A.; Zia, T.; Tariq, S.A.; Ahmed, G.; Zahra, A. Uncertainty assisted robust tuberculosis identification with bayesian convolutional neural networks. IEEE Access 2020, 8, 22812–22825. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhou, Y.; Du, P.; Lang, G.; Xu, M.; Wu, W. A deep learning system that generates quantitative CT reports for diagnosing pulmonary Tuberculosis. Appl. Intell. 2020, 51, 4082–4093. [Google Scholar] [CrossRef]
- Sahlol, A.T.; Elaziz, M.A.; Jamal, A.T.; Damaševičius, R.; Hassan, O.F. A novel method for detection of tuberculosis in chest radiographs using artificial ecosystem-based optimisation of deep neural network features. Symmetry 2020, 12, 1146. [Google Scholar] [CrossRef]
- Munadi, K.; Muchtar, K.; Maulina, N.; Pradhan, B. Image enhancement for tuberculosis detection using deep learning. IEEE Access 2020, 8, 217897–217907. [Google Scholar] [CrossRef]
- Rajaraman, S.; Candemir, S.; Xue, Z.; Alderson, P.O.; Kohli, M.; Abuya, J.; Thoma, G.R.; Antani, S. A novel stacked generalization of models for improved TB detection in chest radiographs. In Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, EMBS, Honolulu, HI, USA, 17–21 July 2018; Volume 2018, pp. 718–721. [Google Scholar] [CrossRef]
- Lopes, U.K.; Valiati, J.F. Pre-trained convolutional neural networks as feature extractors for tuberculosis detection. Comput. Biol. Med. 2017, 89, 135–143. [Google Scholar] [CrossRef]
- Ayaz, M.; Shaukat, F.; Raja, G. Ensemble learning based automatic detection of tuberculosis in chest X-ray images using hybrid feature descriptors. Phys. Eng. Sci. Med. 2021, 44, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Lu, P.X. Chest X-ray Masks and Label. Kaggle. Available online: https://www.kaggle.com/nikhilpandey360/chest-xray-masks-and-labels (accessed on 20 April 2022).
- Kaggle. RSNA Pneumonia Detection Challenge. [Online]. Available online: https://www.kaggle.com/c/rsna-pneumonia-detection-challenge/data (accessed on 9 June 2021).
- Kaggle. Tuberculosis (TB) Chest X-ray Database. Available online: https://www.kaggle.com/datasets/tawsifurrahman/tuberculosis-tb-chest-xray-dataset (accessed on 14 February 2022).






| Reference | Module | Dataset | Classification | Highlights | Limitations |
|---|---|---|---|---|---|
| Msonda et al. [8] | CNN, AlexNet, GoogLeNet ResNet50 | Konya Education and Research Hospital, Turkey dataset (Private). Montgomery and Shenzhen dataset [21] | SVM CNN classifier | Presents a methodology that utilizes the DCNN in classifying TB-affected patients using Chest X-Rays (CXR) | Less training dataset which affects model performance |
| Akbar et al. [49] | CNN | Montgomery and Shenzhen [21] | CNN classifier | TB classification from chest X-ray images based on the CNN model | Less training dataset, and with no data augmentation |
| Abideen et al. [61] | AlexNet, VGG16, VGG19, CNN | Montgomery and Shenzhen datasets [21] | SVM | Bayesian-based CNN for uncertain cases with low discernibility among TB and non-TB manifested CXRs. | Evaluation performance is less due to less training dataset. |
| Hooda et al. [38] | CNN | Montgomery and Shenzhen datasets [21] | CNN classifier | Pure CNN model for TB classification | Less training dataset and further verification through a clinical study |
| Hwang et al. [39] | CNN | Datasets from Seoul National University (Private) Hospital. Montgomery and Shenzhen datasets | CNN classifier | DL–based algorithm for active pulmonary TB detection with lesion-wise localization and image-wise information for detection. | There is uncertainty about the model’s ability to identify different TB manifestations and other pulmonary abnormalities. |
| Rahman et al. [53] | ResNet, ChexNet, InceptionV3, DenseNet201, Vgg19 SqueezeNet, MobileNet | Kaggle Chest X-ray images [68], RSNA CXR dataset [69]. Montgomery and Shenzhen datasets [21] | CNN classifier | DL model for TB classification and visualization of learning features based on class activation map (CAM) score. | The CAM score shows that the model sometimes learns from the wrong parts of the image, which affects prediction accuracy. |
| Guo et al. [60] | CNN | Shenzhen datasets [21] | Linear average-based ensemble classifier | Proposed CNN models for TB classification and localization from CXR images | The model can bias due to uneven distribution in class datasets. |
| Rajaraman et al. [65] | AlexNet, GoogLeNet, VGG-16 and ResNet-50 | Indiana School of Medicine and Academic Model Providing Access to Healthcare (AMPATH) dataset. Montgomery and Shenzhen datasets [21] | SVM ensemble classifier | Developed a CNN model through ensemble learning that classifies hand-engineered features for TB detection from X-ray images. | The proposed model is not computationally efficient but rather memory intensive. |
| Ayaz et al. [67] | Gabor filter, Inceptionv3, VGG-Net MobileNet, ResNet50, Xception | Montgomery and Shenzhen datasets [21] | Linear regression ensemble classifier | TB detection technique that combines hand-crafted features with DCNN through ensemble learning. | The training dataset is less, and the model needs to be evaluated on a larger dataset. |
| Munadi et al. [64] | ResNet and EfficientNet | Shenzhen datasets [21] | CNN classifier | Presents three image enhancement algorithms for TB detection based on pre-trained CNN models. | The model’s accuracy can be improved since some earlier with a larger dataset. |
| Li et al. [62] | CNN, 3D DENSEVOXNET-RPN, 3DUNET-RPN | Private dataset | CNN classifier | Developed DL model to diagnose pulmonary tuberculosis (PTB) | Only five types of pulmonary lesions were considered for the study while ignoring the other signs. |
| Sahol et al. [63] | MobileNet | Montgomery and Shenzhen datasets [21] | K-Nearest Neighbor (KNN) | Proposed a hybrid method based on MobileNet and artificial ecosystem-based optimization (AEO) algorithm for TB classification. | A larger dataset is required for the evaluation of the proposed model |
| Devnath et al. [50] | CNN | NIOSH dataset [41] | CNN classifier | CNN model for TB classification | A larger dataset is required for further evaluation. |
| Pretrained Model | Input Size | No. of Parameters | Trainable Parameters | Non-Trainable Parameters |
|---|---|---|---|---|
| VGG 16 | 224 × 224 | 14,764,866 | 50,178 | 14,714,688 |
| VGG19 | 224 × 224 | 20,074,562 | 50,178 | 20,024,384 |
| InceptionResnetv2 | 299 × 299 | 54,413,538 | 76,802 | 54,336,736 |
| MobileNet | 224 × 224 | 3,090,434 | 94,082 | 2,996,352 |
| Xception | 299 × 299 | 21,271,082 | 409,602 | 20,861,480 |
| DenseNet201 | 224 × 224 | 18,510,146 | 188,162 | 18,321,984 |
| InceptionV3 | 299 × 299 | 21,905,186 | 102,402 | 21,802,784 |
| EfficientNetB1 | 224 × 224 | 6,700,681 | 125,442 | 6,575,239 |
| ResNet50 | 224 × 224 | 23,788,418 | 200,706 | 23,587,712 |
| Training (Chest X-ray Images for TB Dataset) | Total No. of Images | Normal Images | TB Infected |
| 1500 | 800 | 700 | |
| Validation (Montgomery dataset) | 138 | 80 | 58 |
| Testing (Shenzhen dataset) | 662 | 326 | 336 |
| Model | Accuracy | Sensitivity | Specificity |
|---|---|---|---|
| Customized CNN | 90.04 | 91.03 | 90.01 |
| VGG 16 | 92.38 | 91.42 | 92.57 |
| VGG19 | 92.86 | 92.86 | 92.7 |
| InceptionResnetv2 | 92.62 | 90.02 | 93.14 |
| MobileNet | 92.3 | 90 | 92.71 |
| Xception | 91.04 | 92.14 | 90 |
| DenseNet | 92.38 | 92.57 | 91.42 |
| InceptionV3 | 91.67 | 92.86 | 91.43 |
| EfficientNetB1 | 91.93 | 91.98 | 91.67 |
| ResNet50 | 91.58 | 91 | 91.03 |
| Model | Accuracy | Sensitivity | Specificity |
|---|---|---|---|
| Customized CNN | 93.78 | 91.2 | 92.24 |
| VGG 16 | 96.43 | 95.96 | 96.71 |
| VGG19 | 97.02 | 97.14 | 97.14 |
| InceptionResnetV2 | 96.55 | 96.43 | 96.57 |
| MobileNet | 95.36 | 95.57 | 93.01 |
| Xception | 95.95 | 95 | 96.61 |
| Densenet | 96.43 | 95.71 | 96.57 |
| EfficientNetB1 | 96.3 | 95.21 | 95.2 |
| ResNet50 | 95.3 | 95.5 | 95.3 |
| InceptionV3 | 95.76 | 95.14 | 92.81 |
| Ensemble Technique | 98.38 | 98.89 | 98.7 |
| Reference | Feature Extraction | Training and Evaluation Data | Classifier | Evaluation Metrics (%) |
|---|---|---|---|---|
| Msonda et al. [8] | AlexNet, GoogLeNet ResNet50, CNN | Konya Education and Research Hospital, Turkey dataset. Montgomery and Shenzhen dataset [21] | SVM CNN classifier | Acc = 98, Sens = 97 Spec = 99 |
| Akbar et al. [49] | CNN | Montgomery and Shenzhen [21] | CNN classifier | Acc = 92 |
| Abideen et al. [61] | AlexNet, VGG16, VGG19, CNN | Montgomery and Shenzhen datasets [21] | SVM | Acc = 96.42 |
| Hooda et al. [38] | CNN | Montgomery and Shenzhen datasets | CNN classifier | Acc = 94.73 |
| Hwang et al. [39] | CNN | Datasets from Seoul National University Hospital. Montgomery and Shenzhen datasets | CNN classifier | Acc = 97.7, Sen = 98 AUC = 98.8 |
| Rahman et al. [53] | ResNet, ChexNet, InceptionV3, DenseNet201, Vgg19 SqueezeNet, MobileNet | Kaggle Chest X-ray images [68], RSNA CXR dataset [69]. Montgomery and Shenzhen datasets [21] | CNN classifier | Acc = 98.6 Sen = 98.56 Spec = 98.54 |
| Guo et al. [60] | CNN | Shenzhen datasets [21] | Linear average ensemble classifier | Acc = 98.46 Sens = 98.76 AUC = 99 |
| Rajaraman et al. [65] | AlexNet, GoogLeNet, VGG-16 and ResNet-50 | Dataset from Indiana School of Medicine and Academic Model Providing Access to Healthcare (AMPATH). Montgomery and Shenzhen datasets | SVM ensemble classifier | Acc = 96.0 AUC = 96.5 |
| Ayaz et al., [67] | Gabor filter, Inceptionv3, VGG-Net MobileNet, ResNet50, Xception | Montgomery and Shenzhen datasets | Linear regression ensemble classifier | AUC = 99 |
| Munadi et al. [64] | ResNet and EfficientNet | Shenzhen datasets | CNN classifier | Acc = 89.92 AUC = 94.8 |
| Li et al. [62] | CNN, 3D DENSEVOXNET-RPN, 3DUNET-RPN | Private dataset | CNN classifier | Recall = 98.7 Prec = 93.7 |
| Sahol et al. [63] | MobileNet | Montgomery and Shenzhen datasets | K-nearest neighbor (KNN) | Acc = 94.1 |
| Devnath et al. [50] | CNN | NIOSH dataset [41] | CNN classifier | Acc = 87.29 |
| Prposed model | CNN, VGG, InceptionResnetV2, MobileNet, Xception, DenseNet, InceptionV3 | Chest X-ray images for tuberculosis [70] Kaggle dataset [68] Montgomery and Shenzhen datasets [21] | Stacked ensemble classifier | Acc = 98.38 Sen = 98.89 Spec = 98.70 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kotei, E.; Thirunavukarasu, R. Ensemble Technique Coupled with Deep Transfer Learning Framework for Automatic Detection of Tuberculosis from Chest X-ray Radiographs. Healthcare 2022, 10, 2335. https://doi.org/10.3390/healthcare10112335
Kotei E, Thirunavukarasu R. Ensemble Technique Coupled with Deep Transfer Learning Framework for Automatic Detection of Tuberculosis from Chest X-ray Radiographs. Healthcare. 2022; 10(11):2335. https://doi.org/10.3390/healthcare10112335
Chicago/Turabian StyleKotei, Evans, and Ramkumar Thirunavukarasu. 2022. "Ensemble Technique Coupled with Deep Transfer Learning Framework for Automatic Detection of Tuberculosis from Chest X-ray Radiographs" Healthcare 10, no. 11: 2335. https://doi.org/10.3390/healthcare10112335
APA StyleKotei, E., & Thirunavukarasu, R. (2022). Ensemble Technique Coupled with Deep Transfer Learning Framework for Automatic Detection of Tuberculosis from Chest X-ray Radiographs. Healthcare, 10(11), 2335. https://doi.org/10.3390/healthcare10112335







