1. Introduction
Cancer is a disease driven by the accumulation of somatic mutations. Mutations on specific genes that are called cancer driver genes can affect the transcription of the genes and cause the differential expression of the genes. As many of the driver genes are signaling molecules that control the expression of downstream genes, their differential gene expressions may impinge on the cell and contribute to the hallmarks of cancer such as sustained cell proliferation and resistance to cell death [
1,
2].
In the early stages of the analysis on differential gene expression in cancer, several research works focused on comparative studies between normal and cancer cells [
1,
3]. Since the era of precision medicine or personalized medicine, however, the analysis of the differential gene expression among individual patients has become popular, as researchers have observed heterogeneity for immune responses induced by the same cancer therapy due to the diverse genetic background of individuals [
4,
5]. A recent study suggested that only around 5% of patients benefit from precision oncology [
6], which highlights the importance of improving the prediction accuracy of drug response.
As the number of molecular data from cancer patients increases, several large-scale database have been created including The Cancer Genome Atlas (TCGA) and the International Cancer Genome Consortium (ICGC) [
7,
8]. Although TCGA and the ICGC provide multi-platform genomic profiling data across cancer types, these databases do not include a large number of patient records with drug response or responses to multiple drugs as the data were collected from patients (donors). On the other hand, the Genomics of Drug Sensitivity in Cancer (GDSC) [
9] database provides large-scale drug screening results for 266 drugs on around 1000 human cancer cell lines that can be utilized to learn and predict drug responses from gene expression by computational methods.
With a large number of cell line data, various computational methods based on machine learning have been proposed to predict drug response [
10,
11]. Drug response prediction is one of the supervised learning problems. Computational models are trained to compute a drug response value (output) of cell lines (
m samples) with a genomic profile (
n input features). Depending on the type of method, learning can be performed using data of the
d entire drugs at once or for each drug separately. The genomic profile of cell lines is usually given as a matrix
, and the responses to
d drugs are given as a matrix
. The purpose of learning here is to predict the response accurately when a new cell line is given for a known drug. Various types of genomic profiles of cell lines can be provided, among which the gene expression profile is the most frequently used one [
10], owing to its representability of the cellular state and the amount of data released. Drug responses in
are often measured as the half maximal inhibitory concentration (
).
represents the amount of drug needed to inhibit a cell’s activity by 50%. Another measure is the Area Under the concentration response Curve (AUC), where a lower AUC implies lesser effectiveness of the drug.
Reference [
10] reviewed several computational methods for drug response prediction that utilized gene expression data including linear regression methods and their generalizations. The linear regression model learns a response function
with the coefficient vector
where
is a genomic profile vector of a cell line. Kernelized Ridge Regression (KRR) maps
x to a vector in a high-dimensional feature space with a kernel function and computes
w in the image vector space [
12]. Reference [
13] recently developed a method called Kernelized Ranked Learning (KRL) that recommends the
k most sensitive drugs for each cell line, rather than the response value itself [
13]. Response-Weighted Elastic Net (RWEN) is based on the linear regression model, but it incorporates additional weights to find a coefficient vector
w that results in a good prediction for cell lines with low drug responses [
14]. As shown in KRR and KRL, feature engineering such as feature selection or extraction can help improve the prediction accuracy. Recent studies on feature engineering included kernel principal component analysis integrated with bootstrapping [
15] and rough set theory [
16].
When gene expression is used as the genomic profile of a cell line, information on the relationship among genes can be incorporated into the drug response prediction process. The relationships among biological entities such as genes or proteins are usually represented as a biological network. STRING [
17] is one of the databases that provides a Protein-Protein Interaction (PPI) network that corresponds to the gene-gene relationship. As biological processes in a cell are operated by certain groups of genes with interactions like binding or regulation, we can assume that expression values of genes located close to each other in a network may affect the cellular state of a cell line together, thereby contributing to the drug response. However, the computational models for drug response prediction mentioned above do not incorporate the prior knowledge of the biological network, which may enhance the prediction accuracy.
Deep learning with neural networks has shown remarkable achievements compared to the traditional machine learning methods in the field of drug development such as drug-drug interaction extraction [
18], drug target prediction [
19,
20], drug side effect detection [
21], and drug discovery [
22]. For drug response prediction, a number of methods have been developed as well, each of which utilizes different input data for prediction [
11]. Multi-omics Late Integration (MOLI) [
23] is a deep learning model that uses multi-omics data including gene expression, Copy Number Variation (CNV), and somatic mutations to characterize a cell line. Three separate subnetworks of MOLI learn representations for each type of omics data, and a final network uses concatenated features and classifies the response of a cell as a responder or non-responder. Reference [
24] proposed a deep autoencoder model for representation learning of cancer cells from input data consisting of gene expression, CNV, and somatic mutations. The latent variables learned from the deep autoencoder are used to train an elastic net or support vector machine to classify the response. Those methods share two characteristics in common: the integration of multiple input data (multi-omics) and binary classification of the drug response. Although the integration of multiple types of omics data can improve the learning of the status of the cell lines, it might limit the availability of the method for testing on different cell lines or patients as the model requires additional data other than gene expression. Furthermore, a certain threshold of the
values should be set before binary classification of the drug response, which may vary depending on the experimental condition such as drug or tumor types.
The Convolutional Neural Network (CNN) is one of the neural network models adopted for drug response prediction [
11]. The CNN has been actively used for image, video, text, and sound data due to its strong ability to preserve the local structure of data and learn hierarchies of features [
25]. Twin Convolutional Neural Network for drugs in SMILES format (tCNNS) [
26] takes a one-hot encoded representation of drugs and feature vectors of cell lines as the inputs for two encoding subnetworks of a One-Dimensional (1D) CNN. One-hot encodings of drugs in tCNNS are derived from Simplified Molecular Input Line Entry System (SMILES) strings that describe the chemical structure of a drug compound. Binary feature vectors of cell lines represent 735 mutation states or CNVs of a cell. Cancer Drug Response profile scan (CDRscan) [
27] proposes an ensemble model composed of five CNNs, each of which predicts the
values from the binary encoding of the genomic signature (mutation) and the drug signature (PaDEL-descriptors [
28]). KekuleScope [
29] adopts transfer learning, which uses a pre-trained CNN on ImageNet data. The pre-trained CNN is trained with images of drug compounds represented as Kekulé structures to predict the drug response. Recently, several algorithms have been proposed to extend CNNs for data on irregular or non-Euclidean domains represented as graphs [
30,
31,
32]. Reference [
33] proposed a method to predict drug response called GraphDRP, which integrates two subnetworks for drug and cell line features, similar to tCNNS [
26]. Instead of one-hot encoding, GraphDRP uses a molecular graph to represent the drug structure converted from the SMILES string, and the Graph Convolutional Network (GCN) model from [
32] is used to learn the features of drugs. Along with GraphDRP, there have been a number of approaches to use graphs to represent the structural properties of drug compounds for drug development and discovery [
34].
Although the aforementioned CNN models incorporate a number of features in the input data, they do not include gene expression values in the genomic features because gene expression cannot be described as 1D binary sequences [
26,
27] or images [
29] used in those CNN models. However, gene expression is known to be the most informative data type for drug response prediction [
35,
36], whereas mutation and the CNV profiles of cell lines added little to the performance in a comparative study [
10]. Furthermore, most of the regression-based methods that utilize gene expression data in the prediction do not consider interactions between genes [
12,
13,
14]. Recent studies successfully introduced a GCN model to use gene expression data for subtype classification of cancer [
37,
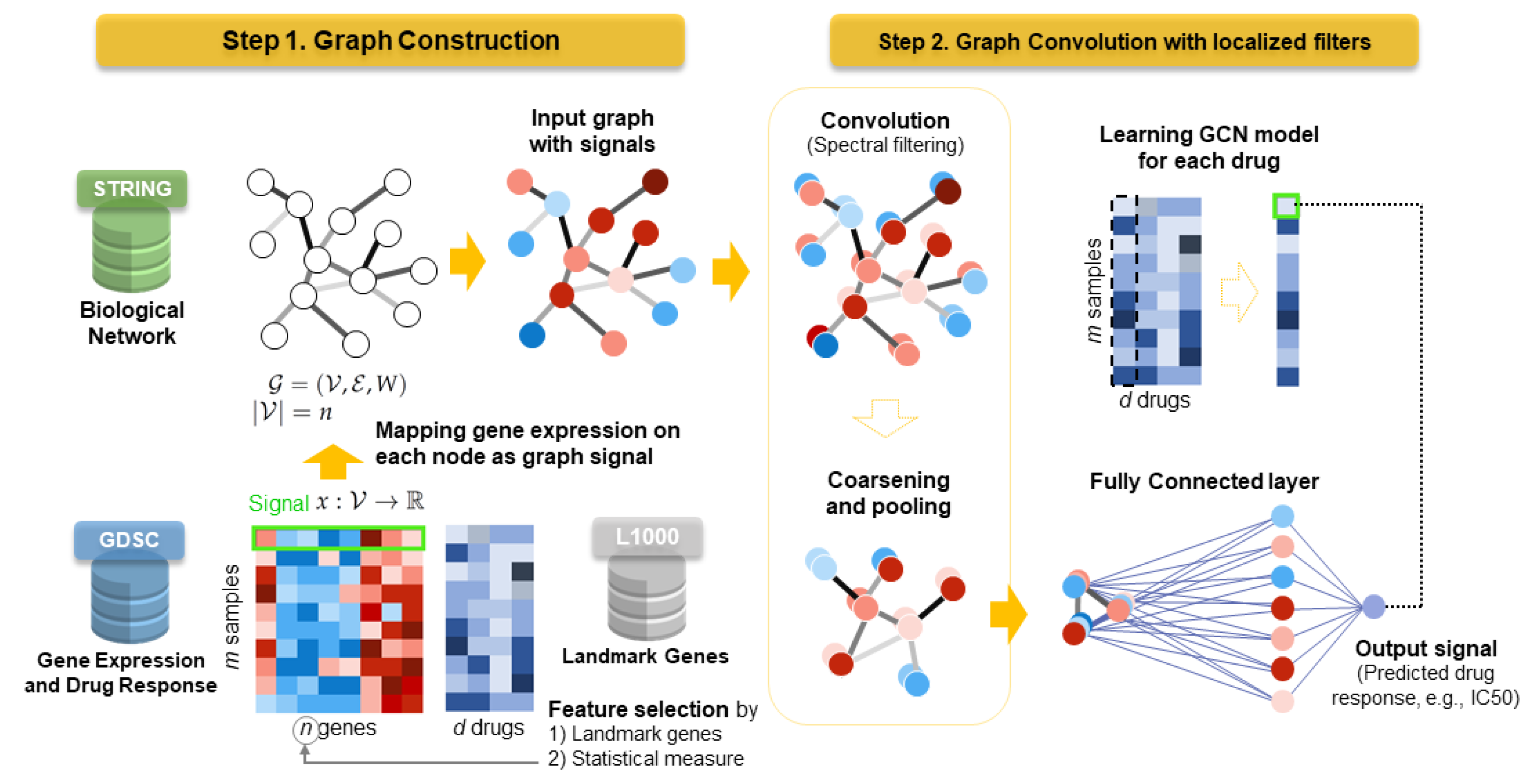
38], and a similar model can be transferred into the problem of drug response prediction. Thus, we propose an analysis framework, DrugGCN, for drug response prediction that can leverage gene expression data and network information using a GCN. DrugGCN constructs an undirected graph from a PPI network and maps gene expression values to each vertex (gene) as graph signals, which will be learned by the GCN to predict the drug response such as the
or AUC. In addition, DrugGCN incorporates the feature selection process to use genes that can possibly improve the prediction accuracy using prior knowledge. The main contributions of DrugGCN are as follows:
We propose a novel framework for drug response prediction that uses a GCN model learning genomic features of cell lines with a graph structure, which is the first approach to our knowledge.
DrugGCN generates a gene graph suitable for drug response prediction by the integration of a PPI network and gene expression data and the feature selection process of genes with high predictive power.
DrugGCN with localized filters can detect the local features in a biological network such as subnetworks of genes that contribute together to the drug response, and its learning complexity is suitable for biological networks with a huge number of vertices and edges.
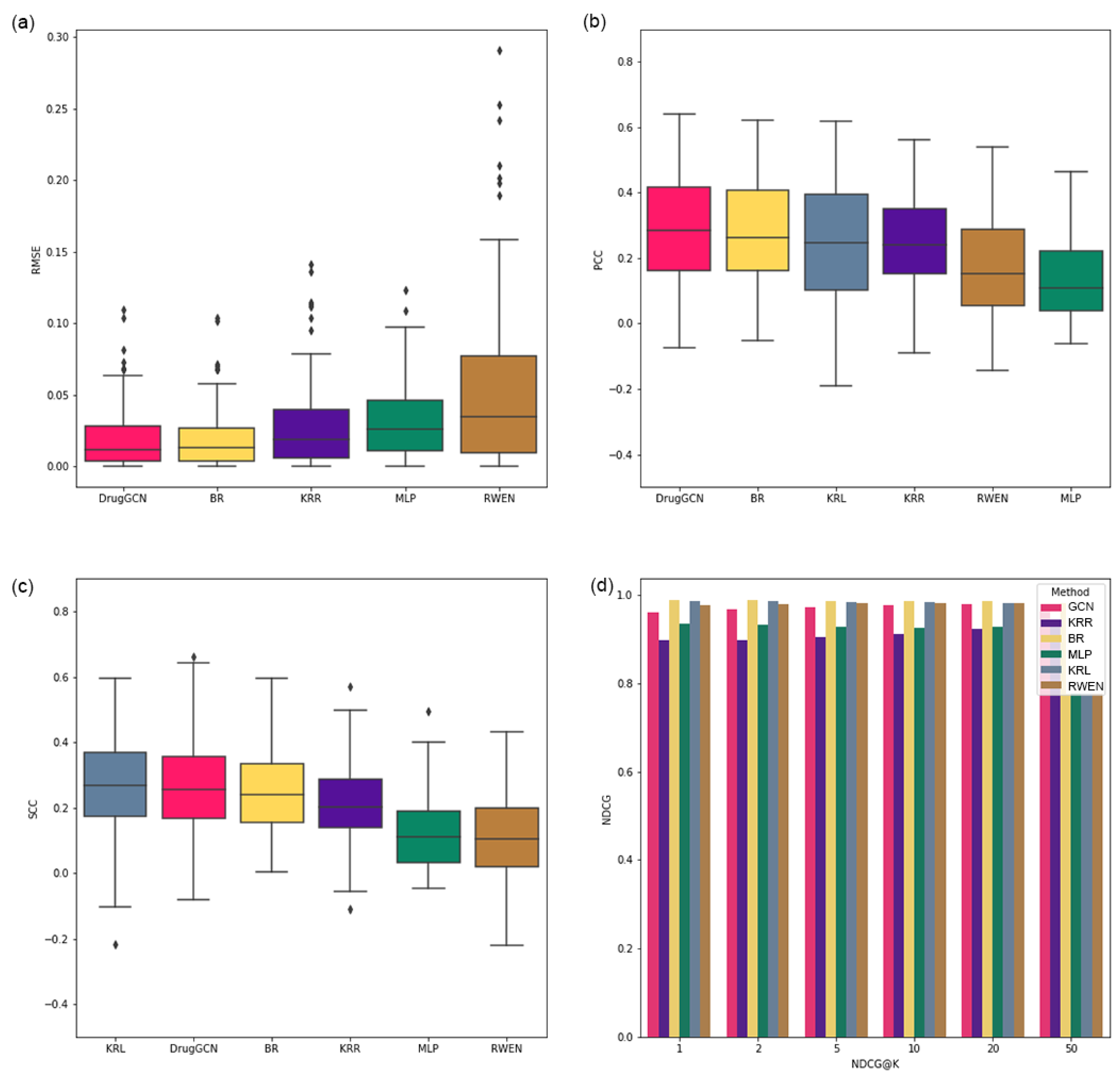
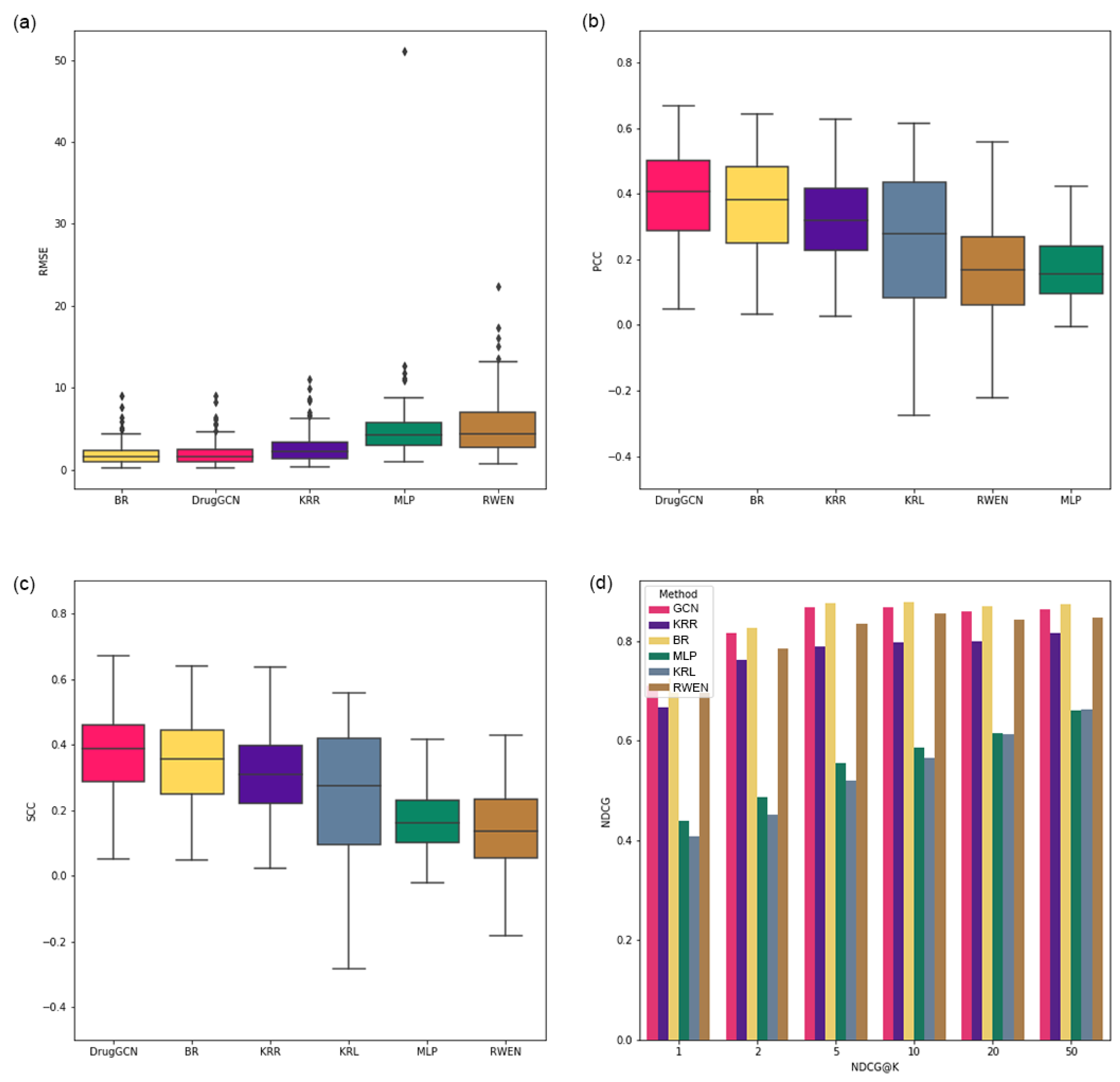
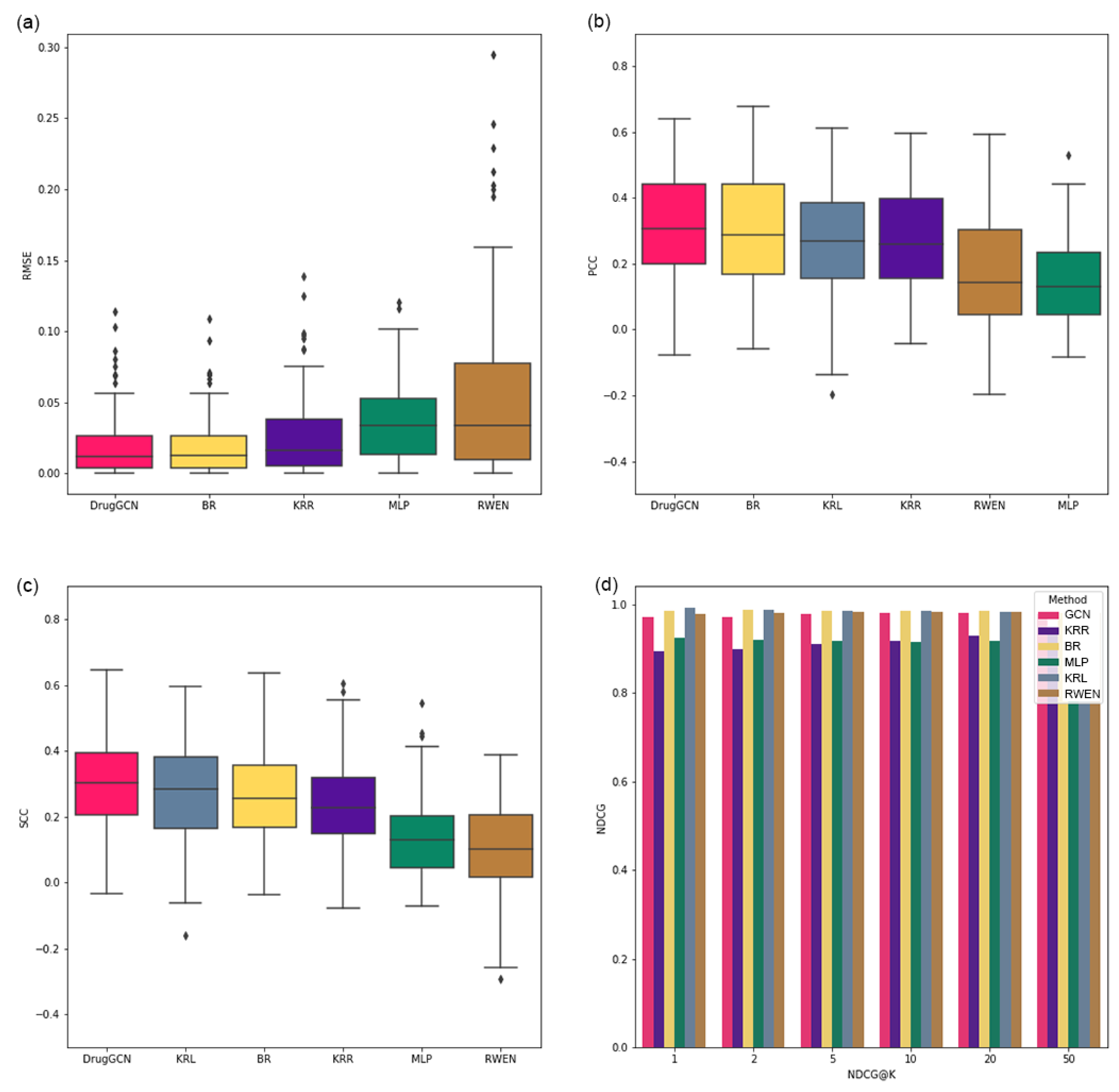
The performance of the proposed approach is demonstrated by a GDSC cell line dataset, and DrugGCN shows high prediction accuracy among the competing methods.
4. Conclusions
In this study, DrugGCN, a computational framework for drug response prediction, was proposed. DrugGCN incorporated PPI network and gene expression data into the GCN model to detect the local features in graphs by localized filtering. The effectiveness of DrugGCN was tested with four GDSC datasets, and it showed high prediction accuracy in terms of the RSME, PCC, and SCC. In the case study of ERK MAPK signaling-related drugs, we discovered supporting evidence of the hypothesis that the high accuracy of DrugGCN was due to the genes forming a subnetwork in the PPI network that provided much information to predict cellular states and consequent drug responses.
The prediction accuracy of DrugGCN can be further improved in terms of the current limitations pertaining to the model structure and input features as described below. Among the competing methods of DrugGCN, the bagging regressor showed high performance with the support of the ensemble model. An ensemble model consisting of multiple GCN was proposed in [
38] for the cancer subtype classification problem where the original graph of the biological network was divided into smaller subnetworks with hundreds of genes using prior knowledge on which genes cooperated with each other for a certain biological process, such as biological pathways from the KEGG database [
48]. The prediction accuracy of the DrugGCN model can be improved with the aforesaid ensemble model, as we showed the predictive power of subnetworks in the case study of ERK MAPK signaling.
The DrugGCN model also can be extended with additional genomic features from different omics data or drug features such as the chemical structures of drug compounds. In particular, the structural properties of drug compounds have been used for drug development and discovery as the form of a graph [
34], which can be easily integrated into the GCN model. As in the similar models [
23,
26], learning from multiple types of features can be implemented with multiple GCN models, the learned representations of which are then concatenated and put into fully connected layers.