A Comparative Performance Evaluation of Classification Algorithms for Clinical Decision Support Systems
Abstract
1. Introduction
- RQ: What is the relative performance of classification algorithms with respect to different resampling strategies?
- RQ: Among the various families, is there a best choice in selecting classification algorithm for clinical decision support systems?
2. Related Work
- Most studies had simply reviewed previous publications by using a mapping study; thus, they could not be used as a support tool in providing a more informed choice to select the best performing classifier for disease prediction.
- When comparing multiple algorithms with multiple datasets, there existed a lack of statistical significance tests. Hence, the performance difference among the classification algorithms was still unrevealed.
3. Materials and Methods
3.1. Datasets
3.2. Classification Algorithms
- a.
- Tree-based algorithms: 5 learners
- i.
- C50 decision tree (C50)The classifier is the extension of the C4.5 algorithm presented in [41] that possesses extra improvements such as boosting, generating smaller trees, and unequal costs for different types of errors. Tree pruning is performed by a final global pruning strategy in which the costly and complex sub-trees are removed in such a way the error rate exceeds the baseline, e.g., the standard error rate of a decision tree without pruning. C50 can generate a set of rules, as well as a classification tree.
- ii.
- Credal decision tree (CDT)It takes into account imprecise probabilities and uncertainty measures for split criteria [42]. This procedure is a bit different from C4.5 or C50, where an information gain is used for a split criterion to choose the split attribute at each branching node in a tree.
- iii.
- Classification and regression tree (CART)It is trained in a recursive binary splitting manner to generate the tree. Binary splitting is a numerical process in which all the values are organized, and different split points are tested using a cost function. The split with the lowest cost is chosen [43].
- iv.
- Random tree (RT)It grows a tree using K randomly selected input features at each node without pruning. The cost function (error) is estimated during the training; thus, there is no accuracy estimation operation, i.e., cross-validation or train-test to obtain an estimate of the training error [44].
- v.
- Forest-PA (FPA)The classifier generates bootstrap samples from the training set and trains a CART classifier on the bootstrap sample using the weights of the attributes. The weights of the attributes are then updated incrementally that are presented in the latest tree. Following this step, the weights of applicable attributes are updated using their respective weight increment values that are not present in the latest tree [45].
- b.
- Ensemble methods: 7 learners
- i.
- Random forest (RF)It uses decision trees as a base classifier, while the tree is grown to a depth of one; then, the same procedure is replicated for all other nodes in the tree until the specified depth of the tree is reached [44].
- ii.
- Extra trees (XT)It works similarly to the random forest classifier, yet the features and splits are chosen at random; thus, it is also known as extremely randomized trees. Because splits are chosen at random, the computational cost (variance) of extra trees is lower than the random forest and decision tree [46].
- iii.
- Rotation forest (RoF)The classifier generates M feature subsets randomly and principal component analysis (PCA) is applied to each subset in order to restore a full feature set (e.g., using M axis rotations) for each base learner (e.g., decision tree) in the ensemble [47].
- iv.
- Gradient boosting machine (GBM)The classifier is proposed to improve the performance of the classification and regression tree (CART). It constructs an ensemble serially, where each new tree in the sequence is in charge of rectifying the prior tree’s prediction error [48].
- v.
- Extreme gradient boosting machine (XGB)The classifier is a state-of-the-art implementation of the gradient boosting algorithm. It shares a similar principle with GBM; however, less computational complexity is one of its advantages. In addition, XGB utilizes a more regularized model, making it able to reduce the complexity of the model while improving the prediction accuracy [49].
- vi.
- CForest (CF)The classifier differs from random forest in terms of the base classifier employed and the aggregation scheme implemented. It utilizes conditional inference trees as a base learner. At the same time, the aggregation scheme works by taking the average weights obtained from each tree, not by averaging predictions directly as the random forest does [50].
- vii.
- Adaboost (AB)The classifier attempts to improve the performance of a weak classifier, e.g., decision tree. The weak learner is trained sequentially on several bootstrap resamples of the learning set. Such a sequential scheme takes the results of a previous classifier into the next one to improve the final prediction by having the latter one emphasizing more on the mistakes of the earlier classifiers [51].
- c.
- Neural-based algorithms: 4 learners
- i.
- Deep learning (DL)It derives from a multilayer feed-forward neural network, which is built based on a stochastic gradient descent algorithm of back-propagation. The primary distinction from a conventional neural network is that it possesses a large number of hidden layers, e.g., greater than or equal to four layers. In addition, some fine-tuned hyper-parameters are needed to be set properly, where a grid search is an option for obtaining the best parameter settings [52].
- ii.
- Multilayer perceptron (MLP)The classifier is a fully connected feed-forward network, where the training is performed by error propagation method [53].
- iii.
- Deep neural network with a stacked autoencoder (SAEDNN)It is a deep learning classifier, where the weights are initialized by a stacked autoencoder [54]. Similar to a deep belief network, it is trained with a greedy layerwise algorithm, while reconstruction error is used as an objective function.
- iv.
- Linear support vector machine (SVM)A support vector machine works based on the principle of hyperplane that classifies the data in a higher dimensional space [55]. In this study, a linear implementation [56] is used with an -regularized and -loss descent method. This is because the linear implementation is computationally efficient as compared to LibSVM [57], for instance.
- d.
- Probability-based classifiers: 3 learners
- i.
- Naive Bayes (NB)A Naive Bayes classifier performs classification based on the conditional probability of a categorical class variable. It considers each of the variables to contribute independently to the probability [58]. In many application domains, the maximum likelihood method is prevalently considered for parameter estimation. Furthermore, it can be trained very efficiently in a supervised learning task.
- ii.
- Gaussian process (GP)A Gaussian process is defined by a mean and a covariance function. The function in any data modeling problem is considered to be a single sample in Gaussian distribution. In the classification task, the Gaussian process uses Laplace approximation for the parameter estimation [59].
- iii.
- Generalized linear model (GLM)The classifier can be used either for classification and regression tasks. In this study, a multinomial family generalization is used as we deal with multi-class response variables. It models the probability of an observation belonging to an output category given the data [60].
- e.
- Discriminant methods: 3 learners
- i.
- Linear discriminant analysis (LDA)The classifier assumes that any data model problem is Gaussian, and each attribute has the same variance. It estimates the mean and the variance for each class, while the prediction is made by estimating the probability of a test set belongs to each class. The output class is the one that gets the highest probability, in which the probability is estimated using Bayes theorem [61].
- ii.
- Mixture discriminant analysis (MDA)This classifier is an extension of linear discriminant analysis. It is used for classification based on mixture models, while the mixture of normals is employed to get a density estimation for each class [62].
- iii.
- K-nearest neighbor (K-NN)The classifier performs prediction on each row of the test set by finding the k nearest (measured by Euclidean distance) training set vectors. The classification is then made by majority voting with ties broken at random [63].
- f.
- Rule-based algorithm: 3 learners
- i.
- Repeated incremental pruning (RIP)This classifier was originally developed to improve the performance of the algorithm. The classifier constructs a rule by taking into account the following two procedures: (1) data samples are randomly divided into two subsets, i.e., a growing set and a pruning set, and (2) a rule is grown using the FOIL algorithm. After generating a rule, the rule is straightaway pruned by eliminating any final sequence of conditions from the rule [64]. In this study, we employ a Java implementation of the RIP algorithm, so-called JRIP.
- ii.
- Partial decision tree (PART)This classifier is a rule-induction procedure that avoids global optimization. It combines the two major rule generation techniques, i.e., decision tree (C4.5) and RIP. PART produces rule sets that are as accurate and of a similar size to those produced by C4.5, and more accurate than the RIP classifier [65].
- iii.
- OneR (1-R)This is a straightforward classifier that produces one rule for each predictor in the training set and selects the rule with a minimum total error as its final rule. A rule for a predictor is produced by generating a frequency table for each predictor feature against the target feature [66].
3.3. Resampling Procedures
| Algorithm 1 General resampling strategy |
Input: A dataset of n observation to , the number of subsets k, and a loss function . Process: 1. Generate k subsets of named to 2. 3. for to k do 4. 5. 6. 7. 8. end 9. Aggregate , i.e., Output: Summary of validation statistics. |
3.4. Significance Tests
- A non-parametric Friedman test [69] is exploited to inspect whether there exist significant differences between the classifiers with respect to the performance metrics as mentioned earlier [70]. The null hypothesis (): no performance differences between classifiers exist, i.e., the expected difference that is equal to zero is observed against the alternative hypothesis (): at least one group of classifiers does not have the same performance, i.e., the expected difference is not equal to zero. The statistic of the Friedman test is calculated according to Equation (1):where v denotes the number of datasets (14 in our case), w denotes the number of classifiers (25 in our case) to be compared, and the average rank of classifier algorithm is .
- Finner test [71] is an p-value adjustment in a step-down manner. Let be the ordered p-values in increasing order, so that , and are the respective hypotheses. It rejects to if w is the smallest integer. Due to its simplicity and power, Finner test is a good choice in general.
- Nemenyi test [72] works by calculating the average rank of each benchmarked algorithm and taking their differences. In case such average differences are larger than or equal to a critical difference (CD), the performances of the algorithms are significantly different. CD can be obtained using the following formula:where is the Studentized statistic divided by .
- Calculate the classifiers’ rank for each dataset using Friedman rank with respect to their area under ROC curve (AUC) metric in increasing order, from the best performer to the worst performer.
- Calculate the average rank of each classifier over all datasets. The best-performing classifier is determined by the lowest value of Friedman rank. Note that the merit is inversely proportional to numeric value.
- Calculate p-value using an omnibus test, e.g., Friedman.
- If the Friedman test demonstrates significant results (p-value < 0.05 in our case), run the Finner’s method. It is carried out based on a pairwise comparison, where the best-performing algorithm is used as a control algorithm for being compared with the remaining algorithms.
- Perform a Nemenyi test to compare the performance of classifiers by each family.
4. Results and Discussion
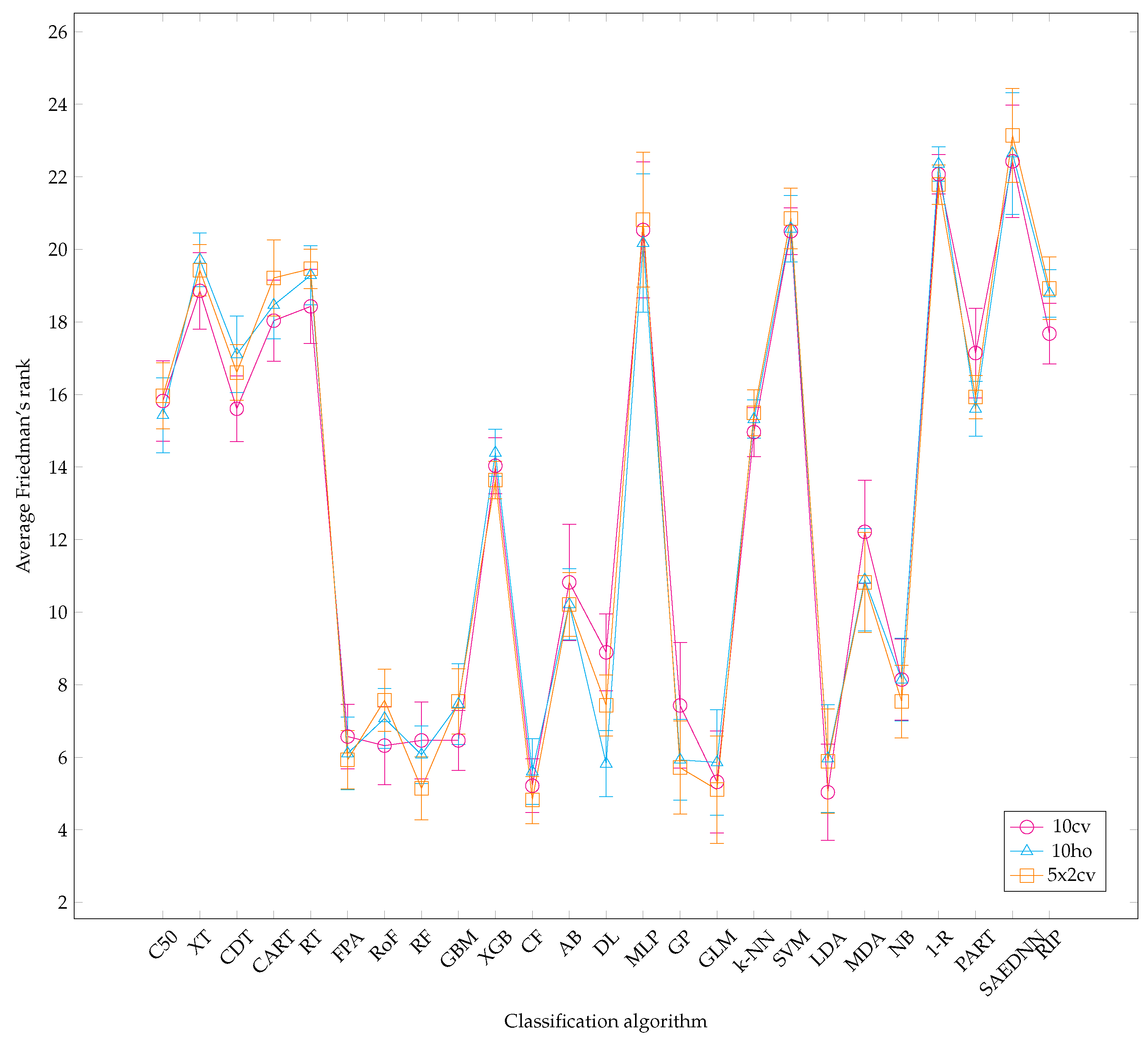
4.1. Overall Analysis
- Overall, it is worth mentioning that, over the three resampling techniques, CF have performed best with average AUC 0.857 and rank 5.16. The result is reasonably unexpected since a conditional inference tree model can outperform other gradient-based ensemble algorithms, i.e., XGB.
- CF is as good as RF since CF works similar to RF [73]. Therefore, it is not surprising that both CF and RF are not significantly different.
- Other ensemble learners, i.e., RF and RoF have performed better than other ensemble models, i.e., FPA and XGB.
- Regarding a highly performance of RF, it is obvious that RF is built based on an ensemble of decision trees. The randomness of each tree split usually provides better prediction performance. In addition, RF is resilient in order to deal with imbalance datasets [74]. Note that several datasets employed in this experiment are highly imbalanced (see Table 1).
- LDA is listed in the top-5 best performing classifiers. LDA has been known as a simple but robust predictor when the dataset is linearly separable.
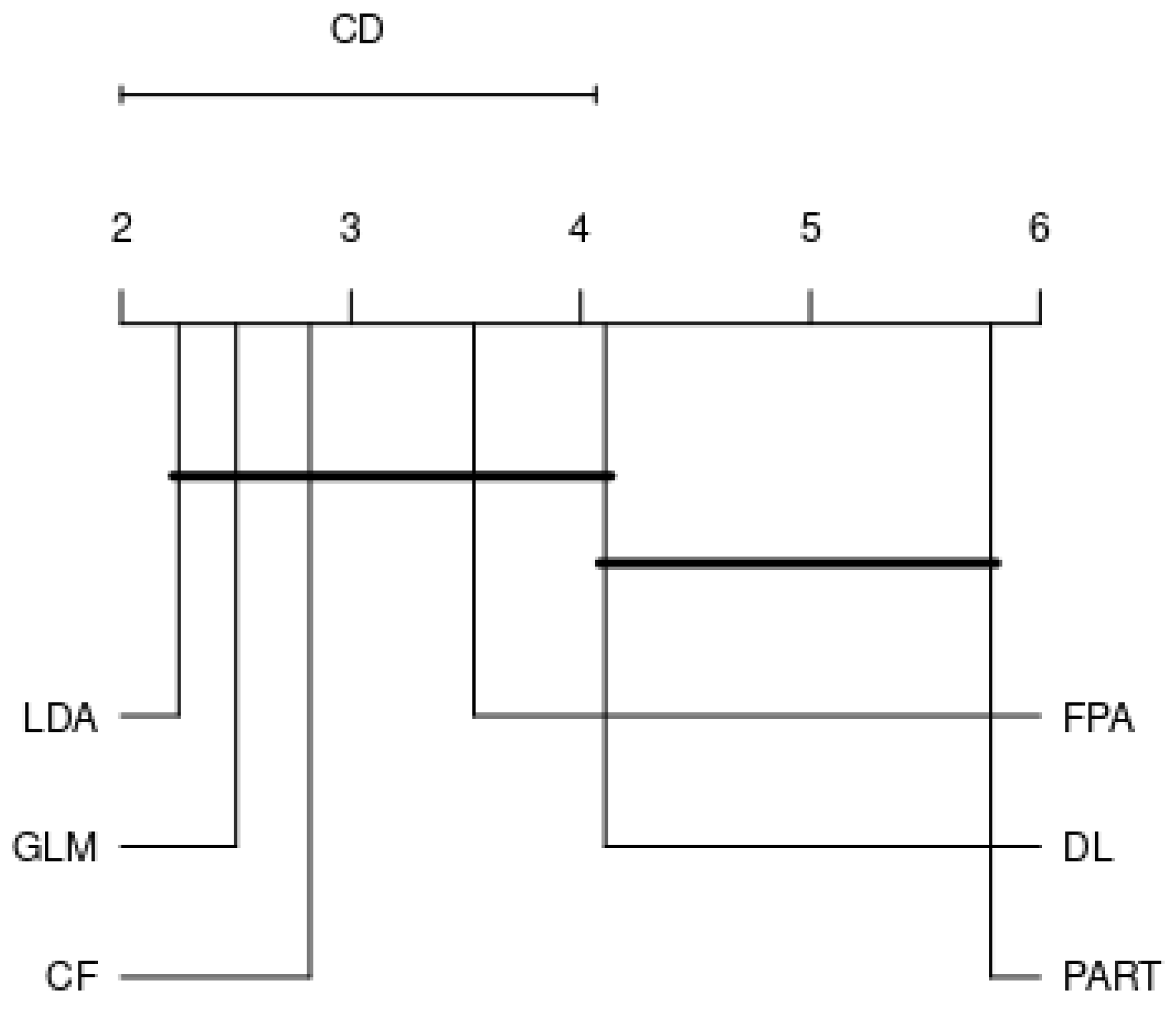
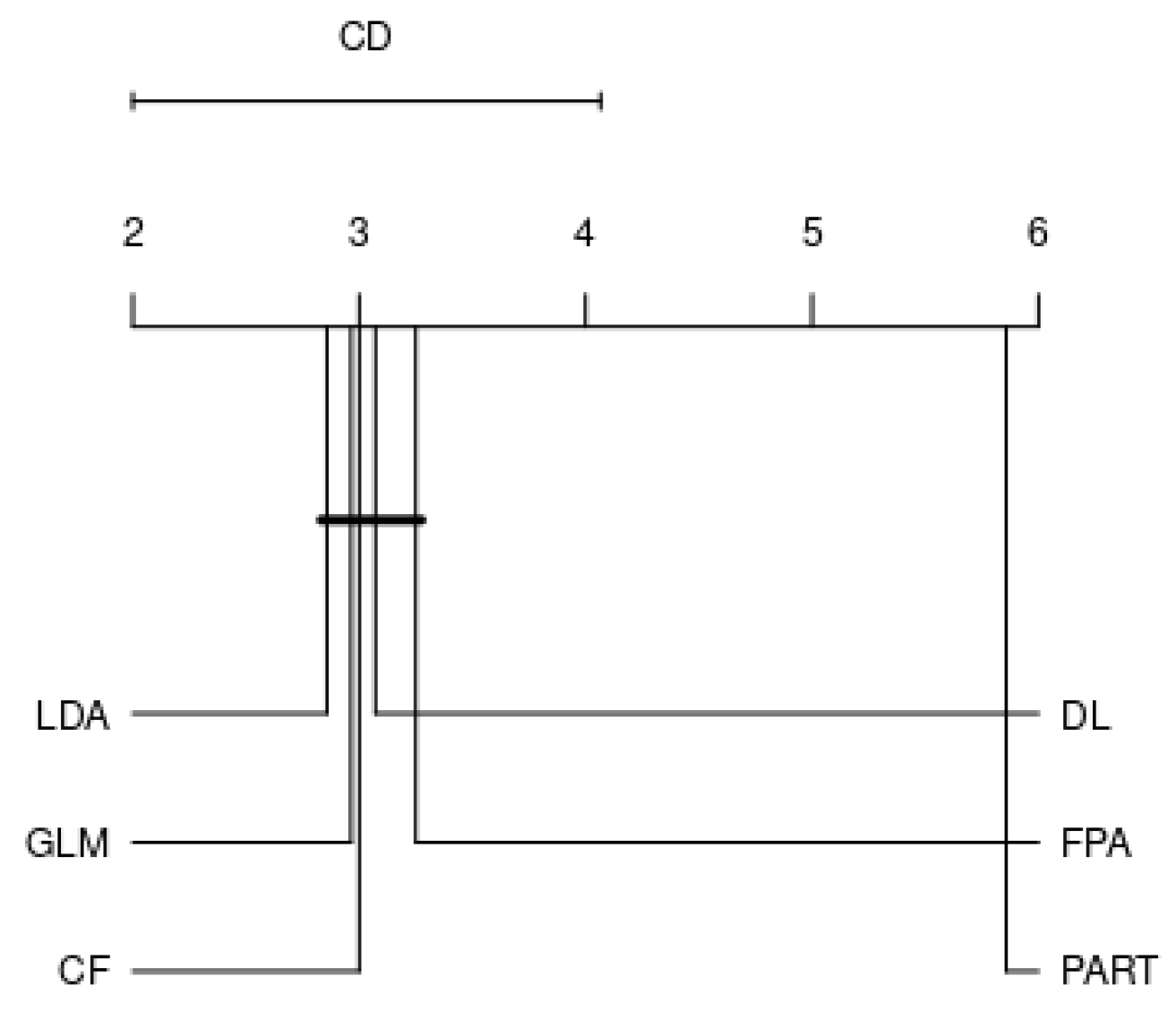
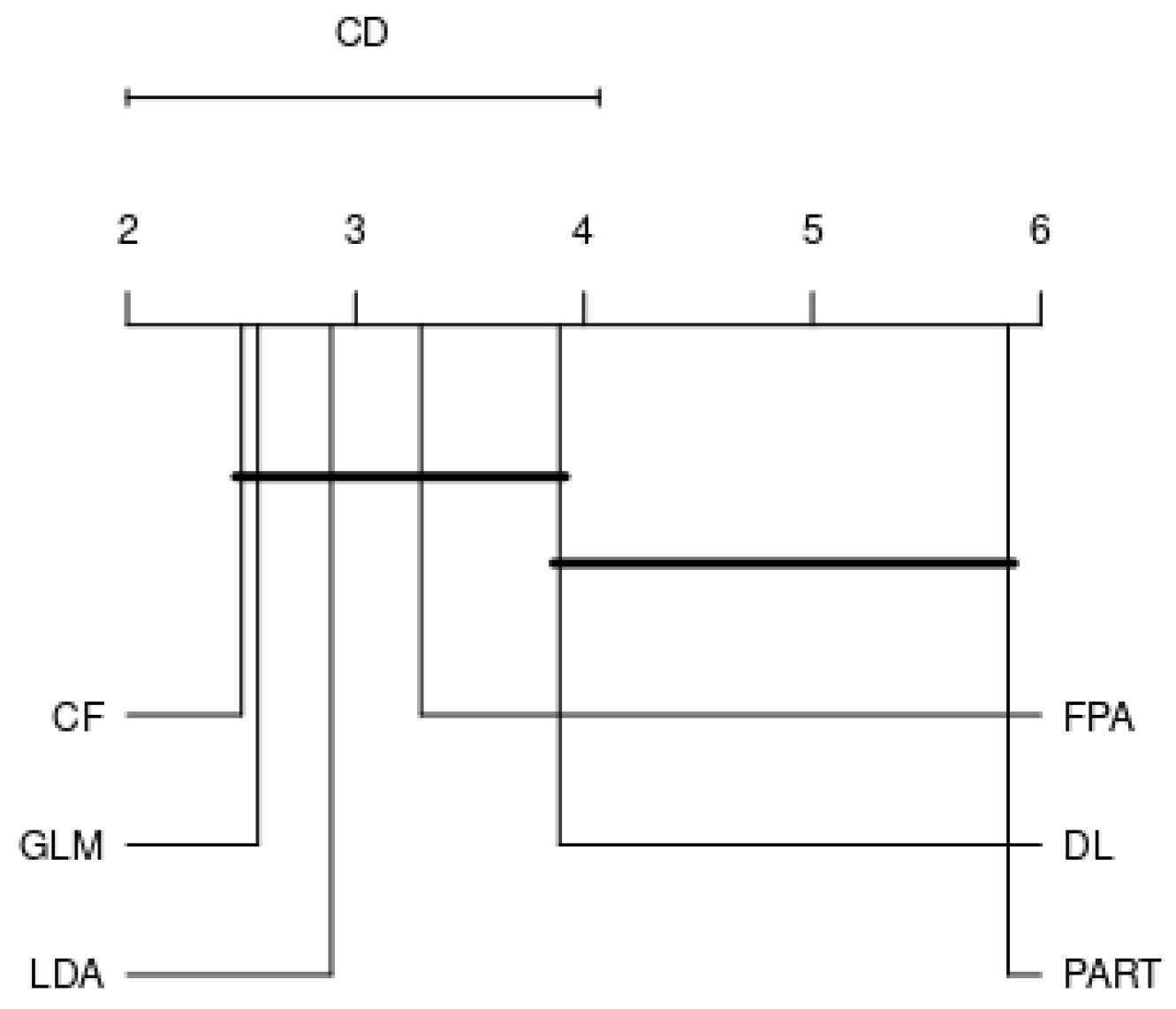
4.2. Analysis by Each Family
5. Conclusions
- RQ: What is the relative performance of classification algorithms with respect to different resampling strategies? According to the Friedman rank, different resampling techniques had no significant impact on several classifiers.
- RQ: Among the various families, is there a best choice in selecting a classification algorithm for a clinical decision support system? This study revealed that choosing the classification algorithms for disease prediction highly depended on types of practical problems, i.e., imbalanced dataset, linear or nonlinear separable, and expert knowledge regarding data and domain. Therefore, it can be concluded that CF, LDA, GLM, RF, and GP were the best choices so far in clinical decision support system field since they are resilient to imbalanced datasets.
Author Contributions
Funding
Conflicts of Interest
Appendix A. List of Learning Parameters Used in the Experiment
- C50_rConfidence factor: 0.25; smallest number of samples: 2; fuzzy threshold: No; sample: 0.
- CDT_wk-th root attribute: 1; maximum tree depth: no restriction; minimum variance proportion: 0.001; imprecise Dirichlet model: 1.0.
- CART_wMinimal cost-complexity pruning: yes; number of folds in the internal cross-validation: 5; heuristic search for binary split: yes.
- RT_wMaximum depth of the tree: unlimited; the number of randomly chosen attributes: 0; amount of data used for backfitting: 0.
- FPA_wNumber of trees to build the forest: 100; number of pruning folds: 2; minimum number of objects: 2.
- RF_rThe implementation of HO is used. Number of trees to build the forest: 100; maximum depth of the tree: unlimited; number of randomly chosen attributes: 0; number of iterations: 100.
- XT_rMinimum number of instances required at a node: 2; number of attributes to randomly choose at a node: −1.
- RoF_wBase classifier: C4.5; number of iterations: 10; projection filter: principle component analysis.
- GBM_rThe implementation of HO is used. Fold assignment: Auto; number of trees: 100; maximum depth: 5; minimum observations in a leaf: 10; number of bins: 20; number of bins at the root level: 1024; number of bins for categorical columns: 1024; learning rate: 0.1; learning rate annealing: 1; sample rate: 1; column sample rate: 1; column sample rate per tree: 1; minimum split improvement: .
- XGB_rMaximum depth: 6; learning rate: 0.3; minimum sum of instance weight: 1; subsample ratio: 1; number of trees to grow per round: 1; number of trees: 100.
- CF_rThe argument: 5; number of trees: 100, minimum criterion: 0.
- AB_wNumber of instances to process: 100; base classifier: decision stump; number of iterations: 10; resampling is used: no; weight threshold for weight pruning: 100.
- DL_rThe implementation of HO is used. L1 and L2 regularization: 0; Hidden layer dropout ratio: 0.5; input dropout ratio: 0; number of training samples for which momentum increases: 1,000,000; learning rate decay: 1; learning rate annealing: ; learning rate: 0.005; adaptive learning rate smoothing factor: ; adaptive learning rate time decay: 0.99; adaptive rate: yes; number of epochs: 100; hidden layer sizes: c(200, 200), activation function: rectifier; maximum relative size of the training data: 5.
- MLP_rMaximum iterations: 100; number of units in the hidden layer: 200; learning function: standard backpropagation, parameter for the learning function: 0.2; activation function for hidden layer: logistic, number of hidden layer: 1.
- SAEDNN_rNumber of units in the hidden layers: 200; activation function: sigmoid; learning rate for gradient descent: 0.8; momentum rate for gradient descent: 0.5; learning rate scale: 1; number of epochs: 100, function of output unit: sigmoid; drop out fraction for hidden layer: 0, number of hidden layers: 2.
- SVM_wCost parameter: 1; bias: 1; : 0.001; parameter of the insensitive loss function: 0.1; number of iterations: 100.
- NB_wUse kernel estimator: no; use supervised discretization: no.
- GP_rData transformation: normalize training data; kernel type: polynomial kernel; exponent value: 1; level of Gaussian noise: 1.
- GLM_rFamily: Gaussian; tweedie variance power: 0; tweedie link power: 1; : ; solver: auto; : 0; number of lambdas to be used in a search: 100; missing value handling: mean imputation;
- LDA_rPrior: proportion; tolerance: ; degrees of freedom: t distribution.
- MDA_rNumber of iterations: 100; number of sub-classes per class: 3; regression method used in optimal scaling: polynomial regression.
- K-NN_wNumber of neighbours to use: 2; distance weighting method: no; nearest neighbours search method: Euclidean distance.
- RIP_wThe amount of data used for pruning: 3; minimum total weight: 2; number of optimizations: 2; use pruning: yes.
- PART_rConfidence factor: 0.25; the amount of data used for reduced-error pruning: 3; reduced-error pruning: no; binary splits: no; MDL correction: yes.
- 1-R_wThe minimum bucket size used for discretizing numeric attributes: 6.
Appendix B. Performance Results of All Benchmarked Classifiers
| DATASET | C50 | XT | CDT | CART | RT | FPA | RoF | RF | GBM | XGB | CF | AB | DL | MLP | GP | GLM | k-NN | SVM | LDA | MDA | NB | 1-R | PART | SAEDNN | RIP |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Breast cancer (diagnostic) | 0.965 | 0.929 | 0.964 | 0.938 | 0.929 | 0.986 | 0.992 | 0.990 | 0.991 | 0.970 | 0.990 | 0.989 | 0.992 | 0.495 | 0.992 | 0.993 | 0.980 | 0.561 | 0.993 | 0.969 | 0.976 | 0.887 | 0.937 | 0.505 | 0.953 |
| Breast cancer (original) | 0.969 | 0.918 | 0.962 | 0.960 | 0.960 | 0.990 | 0.988 | 0.990 | 0.990 | 0.978 | 0.991 | 0.986 | 0.991 | 0.987 | 0.983 | 0.994 | 0.979 | 0.961 | 0.995 | 0.990 | 0.983 | 0.908 | 0.947 | 0.504 | 0.964 |
| Pima Indian | 0.784 | 0.647 | 0.779 | 0.727 | 0.684 | 0.823 | 0.833 | 0.817 | 0.809 | 0.783 | 0.829 | 0.801 | 0.777 | 0.497 | 0.832 | 0.832 | 0.717 | 0.596 | 0.832 | 0.796 | 0.819 | 0.649 | 0.794 | 0.499 | 0.734 |
| Statlog | 0.816 | 0.744 | 0.800 | 0.797 | 0.787 | 0.900 | 0.885 | 0.897 | 0.883 | 0.831 | 0.898 | 0.878 | 0.862 | 0.500 | 0.905 | 0.900 | 0.818 | 0.778 | 0.903 | 0.882 | 0.898 | 0.706 | 0.736 | 0.498 | 0.778 |
| Wisconsin prognostic | 0.602 | 0.486 | 0.494 | 0.475 | 0.579 | 0.629 | 0.730 | 0.617 | 0.741 | 0.682 | 0.642 | 0.695 | 0.696 | 0.497 | 0.640 | 0.739 | 0.617 | 0.504 | 0.792 | 0.722 | 0.642 | 0.504 | 0.500 | 0.487 | 0.638 |
| RSMH | 0.878 | 0.905 | 0.902 | 0.891 | 0.893 | 0.964 | 0.945 | 0.962 | 0.960 | 0.920 | 0.966 | 0.974 | 0.974 | 0.981 | 0.980 | 0.974 | 0.957 | 0.940 | 0.974 | 0.957 | 0.977 | 0.817 | 0.878 | 0.978 | 0.917 |
| Tabriz Iran | 0.558 | 0.557 | 0.545 | 0.529 | 0.569 | 0.754 | 0.712 | 0.708 | 0.720 | 0.663 | 0.720 | 0.727 | 0.680 | 0.503 | 0.724 | 0.721 | 0.545 | 0.500 | 0.721 | 0.534 | 0.753 | 0.499 | 0.613 | 0.502 | 0.528 |
| Thoracic surgery | 0.488 | 0.567 | 0.528 | 0.500 | 0.519 | 0.582 | 0.567 | 0.657 | 0.606 | 0.551 | 0.655 | 0.476 | 0.541 | 0.521 | 0.609 | 0.620 | 0.548 | 0.503 | 0.645 | 0.527 | 0.642 | 0.496 | 0.499 | 0.500 | 0.496 |
| Diabetic retinopathy | 0.683 | 0.616 | 0.665 | 0.668 | 0.616 | 0.731 | 0.821 | 0.754 | 0.770 | 0.668 | 0.770 | 0.652 | 0.786 | 0.508 | 0.764 | 0.768 | 0.648 | 0.643 | 0.796 | 0.761 | 0.682 | 0.528 | 0.694 | 0.499 | 0.637 |
| ILPD | 0.664 | 0.595 | 0.668 | 0.508 | 0.588 | 0.727 | 0.708 | 0.751 | 0.725 | 0.665 | 0.734 | 0.604 | 0.693 | 0.524 | 0.678 | 0.738 | 0.637 | 0.544 | 0.715 | 0.713 | 0.638 | 0.533 | 0.666 | 0.504 | 0.549 |
| Seizure | 0.966 | 0.903 | 0.955 | 0.942 | 0.897 | 0.990 | 0.995 | 0.995 | 0.995 | 0.890 | 0.992 | 0.898 | 0.979 | 0.505 | 0.485 | 0.521 | 0.900 | 0.609 | 0.529 | 0.528 | 0.984 | 0.746 | 0.950 | 0.505 | 0.937 |
| Chronic kidney | 0.982 | 0.973 | 0.991 | 0.995 | 0.998 | 1.000 | 1.000 | 0.999 | 1.000 | 0.969 | 0.998 | 1.000 | 0.995 | 0.687 | 0.994 | 0.998 | 0.970 | 0.689 | 0.998 | 0.966 | 1.000 | 0.921 | 0.973 | 0.500 | 0.976 |
| Cleveland | 0.806 | 0.752 | 0.809 | 0.810 | 0.710 | 0.898 | 0.900 | 0.904 | 0.892 | 0.823 | 0.906 | 0.894 | 0.882 | 0.501 | 0.904 | 0.902 | 0.841 | 0.739 | 0.904 | 0.896 | 0.894 | 0.724 | 0.773 | 0.507 | 0.810 |
| Z-Alizadeh | 0.764 | 0.748 | 0.838 | 0.799 | 0.638 | 0.914 | 0.893 | 0.919 | 0.900 | 0.823 | 0.924 | 0.898 | 0.892 | 0.506 | 0.924 | 0.927 | 0.828 | 0.526 | 0.898 | 0.867 | 0.877 | 0.677 | 0.756 | 0.497 | 0.728 |
| AVERAGE | 0.780 | 0.739 | 0.779 | 0.753 | 0.741 | 0.849 | 0.855 | 0.854 | 0.856 | 0.801 | 0.858 | 0.819 | 0.839 | 0.587 | 0.815 | 0.831 | 0.785 | 0.650 | 0.835 | 0.793 | 0.840 | 0.685 | 0.765 | 0.535 | 0.760 |
| DATASET | C50 | XT | CDT | CART | RT | FPA | RoF | RF | GBM | XGB | CF | AB | DL | MLP | GP | GLM | k-NN | SVM | LDA | MDA | NB | 1-R | PART | SAEDNN | RIP |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Breast cancer (diagnostic) | 0.955 | 0.914 | 0.945 | 0.923 | 0.925 | 0.987 | 0.990 | 0.991 | 0.993 | 0.952 | 0.988 | 0.987 | 0.995 | 0.500 | 0.991 | 0.994 | 0.968 | 0.524 | 0.992 | 0.967 | 0.971 | 0.881 | 0.935 | 0.500 | 0.939 |
| Breast cancer (original) | 0.970 | 0.923 | 0.953 | 0.941 | 0.925 | 0.991 | 0.989 | 0.992 | 0.990 | 0.971 | 0.993 | 0.990 | 0.994 | 0.991 | 0.983 | 0.995 | 0.986 | 0.963 | 0.995 | 0.993 | 0.989 | 0.900 | 0.963 | 0.559 | 0.954 |
| Pima Indian | 0.768 | 0.652 | 0.737 | 0.739 | 0.667 | 0.823 | 0.826 | 0.820 | 0.824 | 0.783 | 0.830 | 0.798 | 0.833 | 0.523 | 0.836 | 0.829 | 0.747 | 0.561 | 0.829 | 0.778 | 0.809 | 0.665 | 0.782 | 0.500 | 0.709 |
| Statlog | 0.814 | 0.721 | 0.755 | 0.787 | 0.705 | 0.896 | 0.885 | 0.890 | 0.872 | 0.843 | 0.895 | 0.886 | 0.887 | 0.450 | 0.913 | 0.907 | 0.828 | 0.775 | 0.909 | 0.897 | 0.914 | 0.658 | 0.762 | 0.524 | 0.749 |
| Wisconsin prognostic | 0.628 | 0.528 | 0.604 | 0.514 | 0.557 | 0.645 | 0.734 | 0.648 | 0.759 | 0.631 | 0.671 | 0.720 | 0.742 | 0.515 | 0.652 | 0.742 | 0.610 | 0.480 | 0.831 | 0.699 | 0.636 | 0.508 | 0.592 | 0.500 | 0.613 |
| RSMH | 0.900 | 0.874 | 0.898 | 0.898 | 0.883 | 0.955 | 0.958 | 0.957 | 0.953 | 0.925 | 0.953 | 0.957 | 0.961 | 0.964 | 0.964 | 0.961 | 0.952 | 0.932 | 0.960 | 0.958 | 0.962 | 0.821 | 0.902 | 0.965 | 0.897 |
| Tabriz Iran | 0.536 | 0.608 | 0.533 | 0.500 | 0.562 | 0.753 | 0.711 | 0.699 | 0.650 | 0.631 | 0.709 | 0.709 | 0.695 | 0.518 | 0.747 | 0.717 | 0.588 | 0.500 | 0.730 | 0.576 | 0.740 | 0.499 | 0.679 | 0.492 | 0.529 |
| Thoracic surgery | 0.497 | 0.555 | 0.496 | 0.500 | 0.514 | 0.660 | 0.578 | 0.687 | 0.621 | 0.584 | 0.692 | 0.525 | 0.669 | 0.560 | 0.641 | 0.675 | 0.563 | 0.519 | 0.695 | 0.600 | 0.696 | 0.507 | 0.559 | 0.500 | 0.502 |
| Diabetic retinopathy | 0.681 | 0.605 | 0.655 | 0.647 | 0.602 | 0.714 | 0.732 | 0.747 | 0.760 | 0.670 | 0.748 | 0.652 | 0.797 | 0.498 | 0.761 | 0.767 | 0.640 | 0.676 | 0.793 | 0.762 | 0.677 | 0.484 | 0.680 | 0.500 | 0.633 |
| ILPD | 0.677 | 0.571 | 0.622 | 0.514 | 0.602 | 0.737 | 0.737 | 0.736 | 0.723 | 0.654 | 0.740 | 0.700 | 0.731 | 0.474 | 0.674 | 0.724 | 0.612 | 0.504 | 0.703 | 0.705 | 0.726 | 0.534 | 0.653 | 0.500 | 0.550 |
| Seizure | 0.965 | 0.895 | 0.949 | 0.938 | 0.900 | 0.994 | 0.994 | 0.994 | 0.995 | 0.893 | 0.992 | 0.905 | 0.992 | 0.546 | 0.995 | 0.515 | 0.893 | 0.617 | 0.530 | 0.528 | 0.953 | 0.753 | 0.945 | 0.513 | 0.931 |
| Chronic kidney | 0.971 | 0.968 | 0.991 | 0.989 | 0.992 | 1.000 | 1.000 | 1.000 | 1.000 | 0.965 | 0.999 | 0.999 | 0.998 | 0.679 | 0.998 | 0.999 | 0.973 | 0.694 | 0.998 | 0.984 | 0.996 | 0.921 | 0.978 | 0.511 | 0.954 |
| Cleveland | 0.810 | 0.740 | 0.789 | 0.807 | 0.735 | 0.897 | 0.887 | 0.899 | 0.881 | 0.847 | 0.901 | 0.892 | 0.882 | 0.493 | 0.892 | 0.892 | 0.840 | 0.683 | 0.897 | 0.885 | 0.885 | 0.725 | 0.801 | 0.499 | 0.797 |
| Z-Alizadeh | 0.811 | 0.732 | 0.880 | 0.842 | 0.714 | 0.938 | 0.925 | 0.936 | 0.923 | 0.869 | 0.936 | 0.911 | 0.910 | 0.505 | 0.932 | 0.929 | 0.837 | 0.538 | 0.899 | 0.845 | 0.879 | 0.641 | 0.780 | 0.500 | 0.788 |
| AVERAGE | 0.785 | 0.735 | 0.772 | 0.753 | 0.735 | 0.856 | 0.853 | 0.857 | 0.853 | 0.801 | 0.861 | 0.831 | 0.863 | 0.587 | 0.856 | 0.832 | 0.788 | 0.640 | 0.840 | 0.798 | 0.845 | 0.678 | 0.787 | 0.540 | 0.753 |
| DATASET | C50 | XT | CDT | CART | RT | FPA | RoF | RF | GBM | XGB | CF | AB | DL | MLP | GP | GLM | k-NN | SVM | LDA | MDA | NB | 1-R | PART | SAEDNN | RIP |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Breast cancer (diagnostic) | 0.956 | 0.912 | 0.945 | 0.923 | 0.919 | 0.986 | 0.989 | 0.988 | 0.989 | 0.959 | 0.988 | 0.987 | 0.993 | 0.495 | 0.990 | 0.993 | 0.977 | 0.501 | 0.991 | 0.975 | 0.976 | 0.887 | 0.933 | 0.500 | 0.928 |
| Breast cancer (original) | 0.967 | 0.926 | 0.959 | 0.952 | 0.938 | 0.990 | 0.989 | 0.989 | 0.989 | 0.969 | 0.990 | 0.988 | 0.992 | 0.988 | 0.984 | 0.994 | 0.984 | 0.958 | 0.994 | 0.992 | 0.987 | 0.905 | 0.956 | 0.516 | 0.951 |
| Pima Indian | 0.745 | 0.627 | 0.756 | 0.713 | 0.657 | 0.818 | 0.815 | 0.812 | 0.795 | 0.762 | 0.819 | 0.796 | 0.800 | 0.512 | 0.827 | 0.825 | 0.718 | 0.565 | 0.826 | 0.788 | 0.809 | 0.652 | 0.755 | 0.503 | 0.703 |
| Statlog | 0.817 | 0.741 | 0.757 | 0.761 | 0.726 | 0.898 | 0.888 | 0.898 | 0.881 | 0.824 | 0.898 | 0.881 | 0.875 | 0.474 | 0.904 | 0.896 | 0.820 | 0.695 | 0.895 | 0.881 | 0.898 | 0.705 | 0.780 | 0.505 | 0.749 |
| Wisconsin prognostic | 0.577 | 0.529 | 0.523 | 0.521 | 0.533 | 0.625 | 0.664 | 0.604 | 0.658 | 0.603 | 0.650 | 0.650 | 0.703 | 0.479 | 0.600 | 0.718 | 0.557 | 0.501 | 0.762 | 0.706 | 0.625 | 0.497 | 0.569 | 0.493 | 0.591 |
| RSMH | 0.875 | 0.879 | 0.892 | 0.875 | 0.886 | 0.963 | 0.952 | 0.966 | 0.957 | 0.892 | 0.963 | 0.959 | 0.969 | 0.977 | 0.975 | 0.970 | 0.954 | 0.938 | 0.970 | 0.964 | 0.976 | 0.831 | 0.890 | 0.969 | 0.858 |
| Tabriz Iran | 0.562 | 0.570 | 0.576 | 0.500 | 0.565 | 0.708 | 0.695 | 0.684 | 0.671 | 0.606 | 0.716 | 0.704 | 0.681 | 0.546 | 0.716 | 0.705 | 0.566 | 0.500 | 0.700 | 0.644 | 0.727 | 0.504 | 0.602 | 0.492 | 0.538 |
| Thoracic surgery | 0.502 | 0.531 | 0.510 | 0.500 | 0.515 | 0.622 | 0.592 | 0.665 | 0.596 | 0.580 | 0.651 | 0.565 | 0.588 | 0.515 | 0.674 | 0.642 | 0.527 | 0.515 | 0.640 | 0.569 | 0.624 | 0.518 | 0.549 | 0.500 | 0.501 |
| Diabetic retinopathy | 0.669 | 0.605 | 0.670 | 0.667 | 0.615 | 0.713 | 0.767 | 0.749 | 0.762 | 0.671 | 0.743 | 0.660 | 0.739 | 0.497 | 0.743 | 0.763 | 0.634 | 0.670 | 0.783 | 0.730 | 0.678 | 0.504 | 0.667 | 0.508 | 0.634 |
| ILPD | 0.635 | 0.602 | 0.629 | 0.504 | 0.587 | 0.727 | 0.689 | 0.726 | 0.712 | 0.654 | 0.726 | 0.695 | 0.698 | 0.518 | 0.649 | 0.737 | 0.592 | 0.519 | 0.714 | 0.715 | 0.726 | 0.546 | 0.649 | 0.508 | 0.567 |
| Seizure | 0.950 | 0.893 | 0.943 | 0.925 | 0.892 | 0.993 | 0.993 | 0.994 | 0.994 | 0.890 | 0.993 | 0.903 | 0.991 | 0.544 | 0.993 | 0.516 | 0.876 | 0.596 | 0.527 | 0.514 | 0.953 | 0.755 | 0.933 | 0.511 | 0.928 |
| Chronic kidney | 0.970 | 0.946 | 0.953 | 0.955 | 0.965 | 0.996 | 0.994 | 0.998 | 0.997 | 0.965 | 0.997 | 0.996 | 0.996 | 0.659 | 0.995 | 0.995 | 0.974 | 0.709 | 0.996 | 0.985 | 0.993 | 0.920 | 0.960 | 0.525 | 0.953 |
| Cleveland | 0.828 | 0.742 | 0.777 | 0.774 | 0.733 | 0.900 | 0.888 | 0.898 | 0.881 | 0.832 | 0.898 | 0.888 | 0.873 | 0.498 | 0.900 | 0.896 | 0.825 | 0.675 | 0.896 | 0.880 | 0.892 | 0.739 | 0.781 | 0.506 | 0.784 |
| Z-Alizadeh | 0.817 | 0.707 | 0.816 | 0.832 | 0.676 | 0.903 | 0.898 | 0.916 | 0.892 | 0.830 | 0.915 | 0.880 | 0.893 | 0.505 | 0.885 | 0.912 | 0.826 | 0.539 | 0.864 | 0.821 | 0.864 | 0.614 | 0.755 | 0.499 | 0.744 |
| AVERAGE | 0.776 | 0.729 | 0.765 | 0.743 | 0.729 | 0.846 | 0.844 | 0.849 | 0.841 | 0.788 | 0.853 | 0.825 | 0.842 | 0.586 | 0.845 | 0.826 | 0.774 | 0.634 | 0.826 | 0.797 | 0.838 | 0.684 | 0.770 | 0.538 | 0.745 |
References
- Lim, S.; Tucker, C.S.; Kumara, S. An unsupervised machine learning model for discovering latent infectious diseases using social media data. J. Biomed. Inform. 2017, 66, 82–94. [Google Scholar] [CrossRef] [PubMed]
- Esfandiari, N.; Babavalian, M.R.; Moghadam, A.M.E.; Tabar, V.K. Knowledge discovery in medicine: Current issue and future trend. Expert Syst. Appl. 2014, 41, 4434–4463. [Google Scholar] [CrossRef]
- Abdar, M.; Zomorodi-Moghadam, M.; Zhou, X.; Gururajan, R.; Tao, X.; Barua, P.D.; Gururajan, R. A new nested ensemble technique for automated diagnosis of breast cancer. Pattern Recognit. Lett. 2018, 132, 123–131. [Google Scholar] [CrossRef]
- Johnson, A.E.; Pollard, T.J.; Shen, L.; Li-Wei, H.L.; Feng, M.; Ghassemi, M.; Moody, B.; Szolovits, P.; Celi, L.A.; Mark, R.G. MIMIC-III, a freely accessible critical care database. Sci. Data 2016, 3, 160035. [Google Scholar] [CrossRef]
- Firdaus, M.A.; Nadia, R.; Tama, B.A. Detecting major disease in public hospital using ensemble techniques. In Proceedings of the 2014 International Symposium on Technology Management and Emerging Technologies, Bandung, Indonesia, 27–29 May 2014; pp. 149–152. [Google Scholar]
- Kadi, I.; Idri, A.; Fernandez-Aleman, J. Knowledge discovery in cardiology: A systematic literature review. Int. J. Med Inform. 2017, 97, 12–32. [Google Scholar] [CrossRef]
- Tama, B.A.; Rhee, K.H. In-depth analysis of neural network ensembles for early detection method of diabetes disease. Int. J. Med Eng. Inform. 2018, 10, 327–341. [Google Scholar] [CrossRef]
- Tama, B.A.; Rhee, K.H. Tree-based classifier ensembles for early detection method of diabetes: An exploratory study. Artif. Intell. Rev. 2019, 51, 355–370. [Google Scholar] [CrossRef]
- Wolpert, D.H.; Macready, W.G. No free lunch theorems for optimization. IEEE Trans. Evol. Comput. 1997, 1, 67–82. [Google Scholar] [CrossRef]
- Hosni, M.; Abnane, I.; Idri, A.; de Gea, J.M.C.; Alemán, J.L.F. Reviewing Ensemble Classification Methods in Breast Cancer. Comput. Methods Programs Biomed. 2019, 177, 89–112. [Google Scholar] [CrossRef]
- Idri, A.; Benhar, H.; Fernández-Alemán, J.; Kadi, I. A systematic map of medical data preprocessing in knowledge discovery. Comput. Methods Programs Biomed. 2018, 162, 69–85. [Google Scholar] [CrossRef]
- Idrissi, T.E.; Idri, A.; Bakkoury, Z. Systematic map and review of predictive techniques in diabetes self-management. Int. J. Inf. Manag. 2019, 46, 263–277. [Google Scholar] [CrossRef]
- Petersen, K.; Feldt, R.; Mujtaba, S.; Mattsson, M. Systematic Mapping Studies in Software Engineering. In Proceedings of the 12th International Conference on Evaluation and Assessment in Software Engineering, Bari, Italy, 26–27 June 2008; Volume 8, pp. 68–77. [Google Scholar]
- Kitchenham, B.A.; Budgen, D.; Brereton, O.P. Using mapping studies as the basis for further research—A participant-observer case study. Inf. Softw. Technol. 2011, 53, 638–651. [Google Scholar] [CrossRef]
- Arji, G.; Safdari, R.; Rezaeizadeh, H.; Abbassian, A.; Mokhtaran, M.; Ayati, M.H. A systematic literature review and classification of knowledge discovery in traditional medicine. Comput. Methods Programs Biomed. 2019, 168, 39–57. [Google Scholar] [CrossRef] [PubMed]
- Jothi, N.; Husain, W. Data mining in healthcare—A review. Procedia Comput. Sci. 2015, 72, 306–313. [Google Scholar] [CrossRef]
- Garciarena, U.; Santana, R. An extensive analysis of the interaction between missing data types, imputation methods, and supervised classifiers. Expert Syst. Appl. 2017, 89, 52–65. [Google Scholar] [CrossRef]
- Abdar, M.; Zomorodi-Moghadam, M.; Das, R.; Ting, I.H. Performance analysis of classification algorithms on early detection of liver disease. Expert Syst. Appl. 2017, 67, 239–251. [Google Scholar] [CrossRef]
- Jain, D.; Singh, V. Feature selection and classification systems for chronic disease prediction: A review. Egypt. Inform. J. 2018, 19, 179–189. [Google Scholar] [CrossRef]
- Moreira, M.W.; Rodrigues, J.J.; Korotaev, V.; Al-Muhtadi, J.; Kumar, N. A comprehensive review on smart decision support systems for health care. IEEE Syst. J. 2019, 13, 3536–3545. [Google Scholar] [CrossRef]
- Sohail, M.N.; Jiadong, R.; Uba, M.M.; Irshad, M. A comprehensive looks at data mining techniques contributing to medical data growth: A survey of researcher reviews. In Recent Developments in Intelligent Computing, Communication and Devices; Springer: Berlin/Heidelberg, Germany, 2019; pp. 21–26. [Google Scholar]
- Nayar, N.; Ahuja, S.; Jain, S. Swarm intelligence and data mining: A review of literature and applications in healthcare. In Proceedings of the Third International Conference on Advanced Informatics for Computing Research, Shimla, India, 15–16 June 2019; pp. 1–7. [Google Scholar]
- Das, R. A comparison of multiple classification methods for diagnosis of Parkinson disease. Expert Syst. Appl. 2010, 37, 1568–1572. [Google Scholar] [CrossRef]
- Bashir, S.; Qamar, U.; Khan, F.H.; Naseem, L. HMV: A medical decision support framework using multi-layer classifiers for disease prediction. J. Comput. Sci. 2016, 13, 10–25. [Google Scholar] [CrossRef]
- Bashir, S.; Qamar, U.; Khan, F.H. IntelliHealth: A medical decision support application using a novel weighted multi-layer classifier ensemble framework. J. Biomed. Inform. 2016, 59, 185–200. [Google Scholar] [CrossRef] [PubMed]
- Asuncion, A.; Newman, D. UCI Machine Learning Repository. 2007. Available online: http://www.ics.uci.edu/mlearn/MLRepository.html (accessed on 16 October 2020).
- Heydari, M.; Teimouri, M.; Heshmati, Z.; Alavinia, S.M. Comparison of various classification algorithms in the diagnosis of type 2 diabetes in Iran. Int. J. Diabetes Dev. Ctries. 2016, 36, 167–173. [Google Scholar] [CrossRef]
- Alizadehsani, R.; Zangooei, M.H.; Hosseini, M.J.; Habibi, J.; Khosravi, A.; Roshanzamir, M.; Khozeimeh, F.; Sarrafzadegan, N.; Nahavandi, S. Coronary artery disease detection using computational intelligence methods. Knowl. Based Syst. 2016, 109, 187–197. [Google Scholar] [CrossRef]
- Aličković, E.; Subasi, A. Breast cancer diagnosis using GA feature selection and Rotation Forest. Neural Comput. Appl. 2017, 28, 753–763. [Google Scholar] [CrossRef]
- Zheng, B.; Yoon, S.W.; Lam, S.S. Breast cancer diagnosis based on feature extraction using a hybrid of K-means and support vector machine algorithms. Expert Syst. Appl. 2014, 41, 1476–1482. [Google Scholar] [CrossRef]
- Maglogiannis, I.; Zafiropoulos, E.; Anagnostopoulos, I. An intelligent system for automated breast cancer diagnosis and prognosis using SVM based classifiers. Appl. Intell. 2009, 30, 24–36. [Google Scholar] [CrossRef]
- Huang, Y.P.; Basanta, H.; Wang, T.H.; Kuo, H.C.; Wu, W.C. A Fuzzy Approach to Determining Critical Factors of Diabetic Retinopathy and Enhancing Data Classification Accuracy. Int. J. Fuzzy Syst. 2019, 21, 1844–1857. [Google Scholar] [CrossRef]
- Raza, K. Improving the prediction accuracy of heart disease with ensemble learning and majority voting rule. In U-Healthcare Monitoring Systems; Elsevier: Amsterdam, The Netherlands, 2019; pp. 179–196. [Google Scholar]
- Abdar, M.; Książek, W.; Acharya, U.R.; Tan, R.S.; Makarenkov, V.; Pławiak, P. A new machine learning technique for an accurate diagnosis of coronary artery disease. Comput. Methods Programs Biomed. 2019, 179, 104992. [Google Scholar] [CrossRef]
- Amin, M.S.; Chiam, Y.K.; Varathan, K.D. Identification of significant features and data mining techniques in predicting heart disease. Telemat. Inform. 2019, 36, 82–93. [Google Scholar] [CrossRef]
- Mangat, V.; Vig, R. Novel associative classifier based on dynamic adaptive PSO: Application to determining candidates for thoracic surgery. Expert Syst. Appl. 2014, 41, 8234–8244. [Google Scholar] [CrossRef]
- Andrzejak, R.G.; Lehnertz, K.; Mormann, F.; Rieke, C.; David, P.; Elger, C.E. Indications of nonlinear deterministic and finite-dimensional structures in time series of brain electrical activity: Dependence on recording region and brain state. Phys. Rev. E 2001, 64, 061907. [Google Scholar] [CrossRef] [PubMed]
- Polat, H.; Mehr, H.D.; Cetin, A. Diagnosis of chronic kidney disease based on support vector machine by feature selection methods. J. Med. Syst. 2017, 41, 55. [Google Scholar] [CrossRef] [PubMed]
- Bischl, B.; Lang, M.; Kotthoff, L.; Schiffner, J.; Richter, J.; Studerus, E.; Casalicchio, G.; Jones, Z.M. mlr: Machine Learning in R. J. Mach. Learn. Res. 2016, 17, 5938–5942. [Google Scholar]
- Hall, M.; Frank, E.; Holmes, G.; Pfahringer, B.; Reutemann, P.; Witten, I.H. The WEKA data mining software: An update. ACM SIGKDD Explor. Newsl. 2009, 11, 10–18. [Google Scholar] [CrossRef]
- Quinlan, J.R. C4.5: Programs for Machine Learning; Elsevier: Amsterdam, The Netherlands, 1992. [Google Scholar]
- Abellán, J.; Moral, S. Building classification trees using the total uncertainty criterion. Int. J. Intell. Syst. 2003, 18, 1215–1225. [Google Scholar] [CrossRef]
- Breiman, L.; Friedman, J.H.; Olshen, R.A.; Stone, C.J. Classification and Regression Trees; Chapman and Hall/CRC: Boca Raton, FL, USA, 1984. [Google Scholar]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Adnan, M.N.; Islam, M.Z. Forest PA: Constructing a decision forest by penalizing attributes used in previous trees. Expert Syst. Appl. 2017, 89, 389–403. [Google Scholar] [CrossRef]
- Geurts, P.; Ernst, D.; Wehenkel, L. Extremely randomized trees. Mach. Learn. 2006, 63, 3–42. [Google Scholar] [CrossRef]
- Rodriguez, J.J.; Kuncheva, L.I.; Alonso, C.J. Rotation forest: A new classifier ensemble method. IEEE Trans. Pattern Anal. Mach. Intell. 2006, 28, 1619–1630. [Google Scholar] [CrossRef]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. Ann. Stat. 2001, 29, 1189–1232. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. Xgboost: A scalable tree boosting system. In Proceedings of the 22nd ACM Sigkdd International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 785–794. [Google Scholar]
- Hothorn, T.; Lausen, B.; Benner, A.; Radespiel-Tröger, M. Bagging survival trees. Stat. Med. 2004, 23, 77–91. [Google Scholar] [CrossRef] [PubMed]
- Freund, Y.; Schapire, R.E. A decision-theoretic generalization of online learning and an application to boosting. J. Comput. Syst. Sci. 1997, 55, 119–139. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436. [Google Scholar] [CrossRef] [PubMed]
- Rosenblatt, F. The perceptron: A probabilistic model for information storage and organization in the brain. Psychol. Rev. 1958, 65, 386. [Google Scholar] [CrossRef]
- Bengio, Y.; Lamblin, P.; Popovici, D.; Larochelle, H. Greedy layer-wise training of deep networks. In Advances in Neural Information Processing Systems 19: Proceedings of the 2006 Conference; Mit Press: Cambridge, MA, USA, 2007; pp. 153–160. [Google Scholar]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Fan, R.E.; Chang, K.W.; Hsieh, C.J.; Wang, X.R.; Lin, C.J. LIBLINEAR: A library for large linear classification. J. Mach. Learn. Res. 2008, 9, 1871–1874. [Google Scholar]
- Chang, C.C.; Lin, C.J. LIBSVM: A library for support vector machines. ACM Trans. Intell. Syst. Technol. (TIST) 2011, 2, 27. [Google Scholar] [CrossRef]
- John, G.H.; Langley, P. Estimating continuous distributions in Bayesian classifiers. In Proceedings of the Eleventh Conference on Uncertainty in Artificial Intelligence; Morgan Kaufmann Publishers Inc.: Burlington, MA, USA, 1995; pp. 338–345. [Google Scholar]
- Williams, C.K.; Barber, D. Bayesian classification with Gaussian processes. IEEE Trans. Pattern Anal. Mach. Intell. 1998, 20, 1342–1351. [Google Scholar] [CrossRef]
- Nelder, J.A.; Wedderburn, R.W. Generalized linear models. J. R. Stat. Soc. Ser. A (Gen.) 1972, 135, 370–384. [Google Scholar] [CrossRef]
- Mika, S.; Ratsch, G.; Weston, J.; Scholkopf, B.; Mullers, K.R. Fisher discriminant analysis with kernels. In Proceedings of the Neural networks for signal processing IX: Proceedings of the 1999 IEEE Signal Processing Society Workshop (cat. no. 98th8468), Madison, WI, USA, 25 August 1999; pp. 41–48. [Google Scholar]
- Hastie, T.; Tibshirani, R. Discriminant analysis by Gaussian mixtures. J. R. Stat. Soc. Ser. B (Methodol.) 1996, 58, 155–176. [Google Scholar] [CrossRef]
- Ripley, B.D.; Hjort, N. Pattern Recognition and Neural Networks; Cambridge University Press: Cambridge, UK, 1996. [Google Scholar]
- Cohen, W.W. Fast effective rule induction. In Machine Learning Proceedings; Elsevier: Amsterdam, The Netherlands, 1995; pp. 115–123. [Google Scholar]
- Frank, E.; Witten, I.H. Generating accurate rule sets without global optimization. In Proceedings of the Fifteenth International Conference on Machine Learning (ICML), Morgan Kaufmann, Madison, WI, USA, 24–27 July 1998; pp. 144–151. [Google Scholar]
- Holte, R.C. Very simple classification rules perform well on most commonly used datasets. Mach. Learn. 1993, 11, 63–90. [Google Scholar] [CrossRef]
- Bischl, B.; Mersmann, O.; Trautmann, H.; Weihs, C. Resampling methods for meta-model validation with recommendations for evolutionary computation. Evol. Comput. 2012, 20, 249–275. [Google Scholar] [CrossRef] [PubMed]
- García, S.; Fernández, A.; Luengo, J.; Herrera, F. Advanced nonparametric tests for multiple comparisons in the design of experiments in computational intelligence and data mining: Experimental analysis of power. Inf. Sci. 2010, 180, 2044–2064. [Google Scholar] [CrossRef]
- Friedman, M. A comparison of alternative tests of significance for the problem of m rankings. Ann. Math. Stat. 1940, 11, 86–92. [Google Scholar] [CrossRef]
- Demšar, J. Statistical comparisons of classifiers over multiple data sets. J. Mach. Learn. Res. 2006, 7, 1–30. [Google Scholar]
- Finner, H. On a monotonicity problem in step-down multiple test procedures. J. Am. Stat. Assoc. 1993, 88, 920–923. [Google Scholar] [CrossRef]
- Nemenyi, P. Distribution-free multiple comparisons. Biometrics 1962, 18, 263. [Google Scholar]
- Mogensen, U.B.; Ishwaran, H.; Gerds, T.A. Evaluating random forests for survival analysis using prediction error curves. J. Stat. Softw. 2012, 50, 1. [Google Scholar] [CrossRef]
- Khoshgoftaar, T.M.; Golawala, M.; Van Hulse, J. An empirical study of learning from imbalanced data using random forest. In Proceedings of the 19th IEEE International Conference on Tools with Artificial Intelligence (ICTAI 2007), Patras, Greece, 29–31 October 2007; Volume 2, pp. 310–317. [Google Scholar]

| ID | Dataset | Disease | #Instances | #Input Variables | #Class Label | %Majority Class | Publication |
|---|---|---|---|---|---|---|---|
| 1 | Breast cancer (diagnostic) | Breast cancer | 569 | 31 | 2 | 62.70 | Aličković and Subasi [29] |
| 2 | Breast cancer (original) | Breast cancer | 699 | 10 | 2 | 65.50 | Zheng et al. [30] |
| 3 | Breast cancer (prognostic) | Breast cancer | 198 | 34 | 2 | 76.26 | Maglogiannis et al. [31] |
| 4 | Pima Indian | Diabetes | 768 | 8 | 2 | 65.10 | Tama and Rhee [8] |
| 5 | Tabriz Iran | Diabetes | 2536 | 13 | 3 | 64.87 | Heydari et al. [27] |
| 6 | RSMH | Diabetes | 435 | 11 | 2 | 79.31 | Tama and Rhee [8] |
| 7 | Diabetic Retinopathy | Diabetes | 1151 | 18 | 2 | 53.08 | Huang et al. [32] |
| 8 | Statlog | Heart disease | 261 | 13 | 2 | 56.32 | Raza [33] |
| 9 | Z-Alizadeh Sani | Heart disease | 303 | 54 | 4 | 86.14 | Abdar et al. [34] |
| 10 | Cleveland | Heart disease | 303 | 13 | 2 | 54.13 | Amin et al. [35] |
| 11 | Thoracic Surgery | Lung cancer | 470 | 16 | 2 | 85.11 | Mangat and Vig [36] |
| 12 | Epileptic Seizure Recognition | Seizure disease | 11,500 | 178 | 5 | 80.0 | Andrzejak et al. [37] |
| 13 | ILPD | Liver disease | 583 | 9 | 2 | 71.18 | Abdar et al. [3] |
| 14 | Chronic Kidney | Chronic kidney | 400 | 24 | 2 | 62.5 | Polat et al. [38] |
| C50 | XT | CDT | CART | RT | FPA | RoF | RF | GBM | XGB | CF | AB | DL | MLP | GP | GLM | k-NN | SVM | LDA | MDA | NB | 1-R | PART | SAEDNN | RIP | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| C50 | n/a | 0.363 | 0.943 | 0.512 | 0.438 | 0.002 | 0.002 | 0.002 | 0.002 | 0.594 | 0.001 | 0.123 | 0.027 | 0.146 | 0.006 | 0.001 | 0.784 | 0.149 | 0.000 | 0.279 | 0.013 | 0.047 | 0.684 | 0.035 | 0.581 |
| XT | 0.363 | n/a | 0.330 | 0.791 | 0.890 | 0.000 | 0.000 | 0.000 | 0.000 | 0.137 | 0.000 | 0.009 | 0.001 | 0.617 | 0.000 | 0.000 | 0.239 | 0.621 | 0.000 | 0.034 | 0.000 | 0.336 | 0.610 | 0.283 | 0.713 |
| CDT | 0.943 | 0.330 | n/a | 0.471 | 0.402 | 0.003 | 0.002 | 0.003 | 0.003 | 0.637 | 0.001 | 0.140 | 0.032 | 0.129 | 0.008 | 0.001 | 0.834 | 0.131 | 0.001 | 0.311 | 0.016 | 0.040 | 0.642 | 0.029 | 0.538 |
| CART | 0.512 | 0.791 | 0.471 | n/a | 0.898 | 0.000 | 0.000 | 0.000 | 0.000 | 0.225 | 0.000 | 0.021 | 0.003 | 0.459 | 0.001 | 0.000 | 0.357 | 0.465 | 0.000 | 0.068 | 0.001 | 0.221 | 0.775 | 0.180 | 0.907 |
| RT | 0.438 | 0.890 | 0.402 | 0.898 | n/a | 0.000 | 0.000 | 0.000 | 0.000 | 0.180 | 0.000 | 0.014 | 0.002 | 0.532 | 0.000 | 0.000 | 0.299 | 0.538 | 0.000 | 0.049 | 0.001 | 0.275 | 0.692 | 0.225 | 0.809 |
| FPA | 0.002 | 0.000 | 0.003 | 0.000 | 0.000 | n/a | 0.934 | 0.971 | 0.971 | 0.016 | 0.677 | 0.194 | 0.489 | 0.000 | 0.784 | 0.698 | 0.006 | 0.000 | 0.642 | 0.077 | 0.637 | 0.000 | 0.001 | 0.000 | 0.000 |
| RoF | 0.002 | 0.000 | 0.002 | 0.000 | 0.000 | 0.934 | n/a | 0.962 | 0.962 | 0.013 | 0.727 | 0.168 | 0.444 | 0.000 | 0.727 | 0.752 | 0.005 | 0.000 | 0.692 | 0.064 | 0.588 | 0.000 | 0.000 | 0.000 | 0.000 |
| RF | 0.002 | 0.000 | 0.003 | 0.000 | 0.000 | 0.971 | 0.962 | n/a | 1.000 | 0.015 | 0.698 | 0.182 | 0.471 | 0.000 | 0.760 | 0.721 | 0.006 | 0.000 | 0.663 | 0.071 | 0.617 | 0.000 | 0.000 | 0.000 | 0.000 |
| GBM | 0.002 | 0.000 | 0.003 | 0.000 | 0.000 | 0.971 | 0.962 | 1.000 | n/a | 0.015 | 0.698 | 0.182 | 0.471 | 0.000 | 0.760 | 0.721 | 0.006 | 0.000 | 0.663 | 0.071 | 0.617 | 0.000 | 0.000 | 0.000 | 0.000 |
| XGB | 0.594 | 0.137 | 0.637 | 0.225 | 0.180 | 0.016 | 0.013 | 0.015 | 0.015 | n/a | 0.004 | 0.336 | 0.111 | 0.039 | 0.035 | 0.005 | 0.767 | 0.040 | 0.003 | 0.588 | 0.064 | 0.009 | 0.353 | 0.006 | 0.275 |
| CF | 0.001 | 0.000 | 0.001 | 0.000 | 0.000 | 0.677 | 0.727 | 0.698 | 0.698 | 0.004 | n/a | 0.079 | 0.270 | 0.000 | 0.512 | 0.971 | 0.001 | 0.000 | 0.952 | 0.025 | 0.383 | 0.000 | 0.000 | 0.000 | 0.000 |
| AB | 0.123 | 0.009 | 0.140 | 0.021 | 0.014 | 0.194 | 0.168 | 0.182 | 0.182 | 0.336 | 0.079 | n/a | 0.569 | 0.001 | 0.311 | 0.085 | 0.208 | 0.002 | 0.069 | 0.669 | 0.424 | 0.000 | 0.045 | 0.000 | 0.029 |
| DL | 0.027 | 0.001 | 0.032 | 0.003 | 0.002 | 0.489 | 0.444 | 0.471 | 0.471 | 0.111 | 0.270 | 0.569 | n/a | 0.000 | 0.655 | 0.283 | 0.055 | 0.000 | 0.243 | 0.318 | 0.809 | 0.000 | 0.007 | 0.000 | 0.004 |
| MLP | 0.146 | 0.617 | 0.129 | 0.459 | 0.532 | 0.000 | 0.000 | 0.000 | 0.000 | 0.039 | 0.000 | 0.001 | 0.000 | n/a | 0.000 | 0.000 | 0.081 | 0.990 | 0.000 | 0.007 | 0.000 | 0.642 | 0.311 | 0.574 | 0.396 |
| GP | 0.006 | 0.000 | 0.008 | 0.001 | 0.000 | 0.784 | 0.727 | 0.760 | 0.760 | 0.035 | 0.512 | 0.311 | 0.655 | 0.000 | n/a | 0.532 | 0.015 | 0.000 | 0.475 | 0.140 | 0.816 | 0.000 | 0.001 | 0.000 | 0.001 |
| GLM | 0.001 | 0.000 | 0.001 | 0.000 | 0.000 | 0.698 | 0.752 | 0.721 | 0.721 | 0.005 | 0.971 | 0.085 | 0.283 | 0.000 | 0.532 | n/a | 0.002 | 0.000 | 0.925 | 0.028 | 0.402 | 0.000 | 0.000 | 0.000 | 0.000 |
| k-NN | 0.784 | 0.239 | 0.834 | 0.357 | 0.299 | 0.006 | 0.005 | 0.006 | 0.006 | 0.767 | 0.001 | 0.208 | 0.055 | 0.081 | 0.015 | 0.002 | n/a | 0.083 | 0.001 | 0.412 | 0.029 | 0.023 | 0.517 | 0.016 | 0.418 |
| SVM | 0.149 | 0.621 | 0.131 | 0.465 | 0.538 | 0.000 | 0.000 | 0.000 | 0.000 | 0.040 | 0.000 | 0.002 | 0.000 | 0.990 | 0.000 | 0.000 | 0.083 | n/a | 0.000 | 0.007 | 0.000 | 0.637 | 0.313 | 0.569 | 0.402 |
| LDA | 0.000 | 0.000 | 0.001 | 0.000 | 0.000 | 0.642 | 0.692 | 0.663 | 0.663 | 0.003 | 0.952 | 0.069 | 0.243 | 0.000 | 0.475 | 0.925 | 0.001 | 0.000 | n/a | 0.021 | 0.353 | 0.000 | 0.000 | 0.000 | 0.000 |
| MDA | 0.279 | 0.034 | 0.311 | 0.068 | 0.049 | 0.077 | 0.064 | 0.071 | 0.071 | 0.588 | 0.025 | 0.669 | 0.318 | 0.007 | 0.140 | 0.028 | 0.412 | 0.007 | 0.021 | n/a | 0.217 | 0.001 | 0.129 | 0.001 | 0.087 |
| NB | 0.013 | 0.000 | 0.016 | 0.001 | 0.001 | 0.637 | 0.588 | 0.617 | 0.617 | 0.064 | 0.383 | 0.424 | 0.809 | 0.000 | 0.816 | 0.402 | 0.029 | 0.000 | 0.353 | 0.217 | n/a | 0.000 | 0.003 | 0.000 | 0.002 |
| 1-R | 0.047 | 0.336 | 0.040 | 0.221 | 0.275 | 0.000 | 0.000 | 0.000 | 0.000 | 0.009 | 0.000 | 0.000 | 0.000 | 0.642 | 0.000 | 0.000 | 0.023 | 0.637 | 0.000 | 0.001 | 0.000 | n/a | 0.129 | 0.907 | 0.180 |
| PART | 0.684 | 0.610 | 0.642 | 0.775 | 0.692 | 0.001 | 0.000 | 0.000 | 0.000 | 0.353 | 0.000 | 0.045 | 0.007 | 0.311 | 0.001 | 0.000 | 0.517 | 0.313 | 0.000 | 0.129 | 0.003 | 0.129 | n/a | 0.100 | 0.862 |
| SAEDNN | 0.035 | 0.283 | 0.029 | 0.180 | 0.225 | 0.000 | 0.000 | 0.000 | 0.000 | 0.006 | 0.000 | 0.000 | 0.000 | 0.574 | 0.000 | 0.000 | 0.016 | 0.569 | 0.000 | 0.001 | 0.000 | 0.907 | 0.100 | n/a | 0.143 |
| RIP | 0.581 | 0.713 | 0.538 | 0.907 | 0.809 | 0.000 | 0.000 | 0.000 | 0.000 | 0.275 | 0.000 | 0.029 | 0.004 | 0.396 | 0.001 | 0.000 | 0.418 | 0.402 | 0.000 | 0.087 | 0.002 | 0.180 | 0.862 | 0.143 | n/a |
| C50 | XT | CDT | CART | RT | FPA | RoF | RF | GBM | XGB | CF | AB | DL | MLP | GP | GLM | k-NN | SVM | LDA | MDA | NB | 1-R | PART | SAEDNN | RIP | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| C50 | n/a | 0.190 | 0.628 | 0.363 | 0.240 | 0.002 | 0.006 | 0.002 | 0.009 | 0.760 | 0.001 | 0.108 | 0.002 | 0.144 | 0.002 | 0.002 | 0.972 | 0.113 | 0.002 | 0.165 | 0.018 | 0.025 | 0.955 | 0.019 | 0.312 |
| XT | 0.190 | n/a | 0.441 | 0.715 | 0.900 | 0.000 | 0.000 | 0.000 | 0.000 | 0.101 | 0.000 | 0.002 | 0.000 | 0.892 | 0.000 | 0.000 | 0.180 | 0.798 | 0.000 | 0.004 | 0.000 | 0.435 | 0.209 | 0.383 | 0.783 |
| CDT | 0.628 | 0.441 | n/a | 0.692 | 0.528 | 0.000 | 0.001 | 0.000 | 0.002 | 0.421 | 0.000 | 0.026 | 0.000 | 0.359 | 0.000 | 0.000 | 0.608 | 0.297 | 0.000 | 0.048 | 0.003 | 0.106 | 0.664 | 0.086 | 0.628 |
| CART | 0.363 | 0.715 | 0.692 | n/a | 0.806 | 0.000 | 0.000 | 0.000 | 0.000 | 0.212 | 0.000 | 0.007 | 0.000 | 0.620 | 0.000 | 0.000 | 0.348 | 0.540 | 0.000 | 0.014 | 0.001 | 0.236 | 0.396 | 0.202 | 0.924 |
| RT | 0.240 | 0.900 | 0.528 | 0.806 | n/a | 0.000 | 0.000 | 0.000 | 0.000 | 0.132 | 0.000 | 0.003 | 0.000 | 0.790 | 0.000 | 0.000 | 0.226 | 0.708 | 0.000 | 0.006 | 0.000 | 0.359 | 0.265 | 0.312 | 0.885 |
| FPA | 0.002 | 0.000 | 0.000 | 0.000 | 0.000 | n/a | 0.776 | 0.990 | 0.692 | 0.007 | 0.885 | 0.209 | 0.932 | 0.000 | 0.955 | 0.939 | 0.002 | 0.000 | 0.963 | 0.141 | 0.551 | 0.000 | 0.002 | 0.000 | 0.000 |
| RoF | 0.006 | 0.000 | 0.001 | 0.000 | 0.000 | 0.776 | n/a | 0.768 | 0.908 | 0.018 | 0.670 | 0.348 | 0.715 | 0.000 | 0.737 | 0.721 | 0.007 | 0.000 | 0.744 | 0.243 | 0.752 | 0.000 | 0.005 | 0.000 | 0.000 |
| RF | 0.002 | 0.000 | 0.000 | 0.000 | 0.000 | 0.990 | 0.768 | n/a | 0.686 | 0.006 | 0.892 | 0.205 | 0.939 | 0.000 | 0.963 | 0.946 | 0.002 | 0.000 | 0.972 | 0.138 | 0.547 | 0.000 | 0.002 | 0.000 | 0.000 |
| GBM | 0.009 | 0.000 | 0.002 | 0.000 | 0.000 | 0.692 | 0.908 | 0.686 | n/a | 0.025 | 0.592 | 0.417 | 0.633 | 0.000 | 0.657 | 0.641 | 0.010 | 0.000 | 0.664 | 0.301 | 0.842 | 0.000 | 0.007 | 0.000 | 0.000 |
| XGB | 0.760 | 0.101 | 0.421 | 0.212 | 0.132 | 0.007 | 0.018 | 0.006 | 0.025 | n/a | 0.004 | 0.202 | 0.005 | 0.070 | 0.005 | 0.005 | 0.783 | 0.050 | 0.006 | 0.292 | 0.047 | 0.009 | 0.721 | 0.007 | 0.180 |
| CF | 0.001 | 0.000 | 0.000 | 0.000 | 0.000 | 0.885 | 0.670 | 0.892 | 0.592 | 0.004 | n/a | 0.158 | 0.946 | 0.000 | 0.924 | 0.939 | 0.001 | 0.000 | 0.915 | 0.104 | 0.455 | 0.000 | 0.001 | 0.000 | 0.000 |
| AB | 0.108 | 0.002 | 0.026 | 0.007 | 0.003 | 0.209 | 0.348 | 0.205 | 0.417 | 0.202 | 0.158 | n/a | 0.180 | 0.001 | 0.190 | 0.182 | 0.116 | 0.001 | 0.193 | 0.842 | 0.547 | 0.000 | 0.096 | 0.000 | 0.005 |
| DL | 0.002 | 0.000 | 0.000 | 0.000 | 0.000 | 0.932 | 0.715 | 0.939 | 0.633 | 0.005 | 0.946 | 0.180 | n/a | 0.000 | 0.972 | 0.990 | 0.002 | 0.000 | 0.963 | 0.119 | 0.500 | 0.000 | 0.001 | 0.000 | 0.000 |
| MLP | 0.144 | 0.892 | 0.359 | 0.620 | 0.790 | 0.000 | 0.000 | 0.000 | 0.000 | 0.070 | 0.000 | 0.001 | 0.000 | n/a | 0.000 | 0.000 | 0.135 | 0.908 | 0.000 | 0.002 | 0.000 | 0.528 | 0.162 | 0.469 | 0.686 |
| GP | 0.002 | 0.000 | 0.000 | 0.000 | 0.000 | 0.955 | 0.737 | 0.963 | 0.657 | 0.005 | 0.924 | 0.190 | 0.972 | 0.000 | n/a | 0.980 | 0.002 | 0.000 | 0.990 | 0.127 | 0.521 | 0.000 | 0.002 | 0.000 | 0.000 |
| GLM | 0.002 | 0.000 | 0.000 | 0.000 | 0.000 | 0.939 | 0.721 | 0.946 | 0.641 | 0.005 | 0.939 | 0.182 | 0.990 | 0.000 | 0.980 | n/a | 0.002 | 0.000 | 0.972 | 0.121 | 0.507 | 0.000 | 0.001 | 0.000 | 0.000 |
| k-NN | 0.972 | 0.180 | 0.608 | 0.348 | 0.226 | 0.002 | 0.007 | 0.002 | 0.010 | 0.783 | 0.001 | 0.116 | 0.002 | 0.135 | 0.002 | 0.002 | n/a | 0.106 | 0.002 | 0.177 | 0.020 | 0.023 | 0.932 | 0.018 | 0.297 |
| SVM | 0.113 | 0.798 | 0.297 | 0.540 | 0.708 | 0.000 | 0.000 | 0.000 | 0.000 | 0.050 | 0.000 | 0.001 | 0.000 | 0.908 | 0.000 | 0.000 | 0.106 | n/a | 0.000 | 0.002 | 0.000 | 0.608 | 0.127 | 0.547 | 0.608 |
| LDA | 0.002 | 0.000 | 0.000 | 0.000 | 0.000 | 0.963 | 0.744 | 0.972 | 0.664 | 0.006 | 0.915 | 0.193 | 0.963 | 0.000 | 0.990 | 0.972 | 0.002 | 0.000 | n/a | 0.129 | 0.528 | 0.000 | 0.002 | 0.000 | 0.000 |
| MDA | 0.165 | 0.004 | 0.048 | 0.014 | 0.006 | 0.141 | 0.243 | 0.138 | 0.301 | 0.292 | 0.104 | 0.842 | 0.119 | 0.002 | 0.127 | 0.121 | 0.177 | 0.002 | 0.129 | n/a | 0.417 | 0.000 | 0.147 | 0.000 | 0.010 |
| NB | 0.018 | 0.000 | 0.003 | 0.001 | 0.000 | 0.551 | 0.752 | 0.547 | 0.842 | 0.047 | 0.455 | 0.547 | 0.500 | 0.000 | 0.521 | 0.507 | 0.020 | 0.000 | 0.528 | 0.417 | n/a | 0.000 | 0.015 | 0.000 | 0.000 |
| 1-R | 0.025 | 0.435 | 0.106 | 0.236 | 0.359 | 0.000 | 0.000 | 0.000 | 0.000 | 0.009 | 0.000 | 0.000 | 0.000 | 0.528 | 0.000 | 0.000 | 0.023 | 0.608 | 0.000 | 0.000 | 0.000 | n/a | 0.029 | 0.932 | 0.281 |
| PART | 0.955 | 0.209 | 0.664 | 0.396 | 0.265 | 0.002 | 0.005 | 0.002 | 0.007 | 0.721 | 0.001 | 0.096 | 0.001 | 0.162 | 0.002 | 0.001 | 0.932 | 0.127 | 0.002 | 0.147 | 0.015 | 0.029 | n/a | 0.023 | 0.342 |
| SAEDNN | 0.019 | 0.383 | 0.086 | 0.202 | 0.312 | 0.000 | 0.000 | 0.000 | 0.000 | 0.007 | 0.000 | 0.000 | 0.000 | 0.469 | 0.000 | 0.000 | 0.018 | 0.547 | 0.000 | 0.000 | 0.000 | 0.932 | 0.023 | n/a | 0.240 |
| RIP | 0.312 | 0.783 | 0.628 | 0.924 | 0.885 | 0.000 | 0.000 | 0.000 | 0.000 | 0.180 | 0.000 | 0.005 | 0.000 | 0.686 | 0.000 | 0.000 | 0.297 | 0.608 | 0.000 | 0.010 | 0.000 | 0.281 | 0.342 | 0.240 | n/a |
| C50 | XT | CDT | CART | RT | FPA | RoF | RF | GBM | XGB | CF | AB | DL | MLP | GP | GLM | k-NN | SVM | LDA | MDA | NB | 1-R | PART | SAEDNN | RIP | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| C50 | n/a | 0.301 | 0.843 | 0.330 | 0.297 | 0.001 | 0.006 | 0.000 | 0.006 | 0.485 | 0.000 | 0.070 | 0.005 | 0.128 | 0.001 | 0.000 | 0.884 | 0.127 | 0.001 | 0.109 | 0.006 | 0.068 | 0.991 | 0.020 | 0.381 |
| XT | 0.301 | n/a | 0.405 | 0.946 | 0.991 | 0.000 | 0.000 | 0.000 | 0.000 | 0.069 | 0.000 | 0.002 | 0.000 | 0.669 | 0.000 | 0.000 | 0.234 | 0.662 | 0.000 | 0.005 | 0.000 | 0.478 | 0.297 | 0.266 | 0.876 |
| CDT | 0.843 | 0.405 | n/a | 0.437 | 0.400 | 0.000 | 0.003 | 0.000 | 0.003 | 0.381 | 0.000 | 0.042 | 0.003 | 0.197 | 0.000 | 0.000 | 0.736 | 0.193 | 0.000 | 0.069 | 0.003 | 0.106 | 0.835 | 0.037 | 0.485 |
| CART | 0.330 | 0.946 | 0.437 | n/a | 0.937 | 0.000 | 0.000 | 0.000 | 0.000 | 0.080 | 0.000 | 0.003 | 0.000 | 0.624 | 0.000 | 0.000 | 0.266 | 0.621 | 0.000 | 0.006 | 0.000 | 0.442 | 0.328 | 0.234 | 0.929 |
| RT | 0.297 | 0.991 | 0.400 | 0.937 | n/a | 0.000 | 0.000 | 0.000 | 0.000 | 0.068 | 0.000 | 0.002 | 0.000 | 0.675 | 0.000 | 0.000 | 0.230 | 0.669 | 0.000 | 0.005 | 0.000 | 0.485 | 0.292 | 0.269 | 0.867 |
| FPA | 0.001 | 0.000 | 0.000 | 0.000 | 0.000 | n/a | 0.621 | 0.811 | 0.624 | 0.012 | 0.736 | 0.190 | 0.645 | 0.000 | 0.946 | 0.803 | 0.002 | 0.000 | 0.991 | 0.127 | 0.624 | 0.000 | 0.001 | 0.000 | 0.000 |
| RoF | 0.006 | 0.000 | 0.003 | 0.000 | 0.000 | 0.621 | n/a | 0.470 | 0.991 | 0.055 | 0.415 | 0.431 | 0.963 | 0.000 | 0.578 | 0.463 | 0.009 | 0.000 | 0.614 | 0.330 | 0.991 | 0.000 | 0.006 | 0.000 | 0.000 |
| RF | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.811 | 0.470 | n/a | 0.473 | 0.005 | 0.920 | 0.112 | 0.486 | 0.000 | 0.859 | 0.991 | 0.001 | 0.000 | 0.818 | 0.073 | 0.473 | 0.000 | 0.000 | 0.000 | 0.000 |
| GBM | 0.006 | 0.000 | 0.003 | 0.000 | 0.000 | 0.624 | 0.991 | 0.473 | n/a | 0.054 | 0.421 | 0.426 | 0.972 | 0.000 | 0.584 | 0.470 | 0.009 | 0.000 | 0.621 | 0.328 | 1.000 | 0.000 | 0.006 | 0.000 | 0.000 |
| XGB | 0.485 | 0.069 | 0.381 | 0.080 | 0.068 | 0.012 | 0.055 | 0.005 | 0.054 | n/a | 0.004 | 0.306 | 0.049 | 0.020 | 0.009 | 0.005 | 0.578 | 0.019 | 0.011 | 0.405 | 0.054 | 0.008 | 0.486 | 0.002 | 0.099 |
| CF | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.736 | 0.415 | 0.920 | 0.421 | 0.004 | n/a | 0.092 | 0.437 | 0.000 | 0.785 | 0.929 | 0.000 | 0.000 | 0.743 | 0.058 | 0.421 | 0.000 | 0.000 | 0.000 | 0.000 |
| AB | 0.070 | 0.002 | 0.042 | 0.003 | 0.002 | 0.190 | 0.431 | 0.112 | 0.426 | 0.306 | 0.092 | n/a | 0.409 | 0.000 | 0.165 | 0.111 | 0.099 | 0.000 | 0.186 | 0.852 | 0.426 | 0.000 | 0.072 | 0.000 | 0.004 |
| DL | 0.005 | 0.000 | 0.003 | 0.000 | 0.000 | 0.645 | 0.963 | 0.486 | 0.972 | 0.049 | 0.437 | 0.409 | n/a | 0.000 | 0.607 | 0.485 | 0.008 | 0.000 | 0.638 | 0.309 | 0.972 | 0.000 | 0.005 | 0.000 | 0.000 |
| MLP | 0.128 | 0.669 | 0.197 | 0.624 | 0.675 | 0.000 | 0.000 | 0.000 | 0.000 | 0.020 | 0.000 | 0.000 | 0.000 | n/a | 0.000 | 0.000 | 0.096 | 0.991 | 0.000 | 0.001 | 0.000 | 0.769 | 0.127 | 0.485 | 0.571 |
| GP | 0.001 | 0.000 | 0.000 | 0.000 | 0.000 | 0.946 | 0.578 | 0.859 | 0.584 | 0.009 | 0.785 | 0.165 | 0.607 | 0.000 | n/a | 0.852 | 0.001 | 0.000 | 0.954 | 0.111 | 0.584 | 0.000 | 0.001 | 0.000 | 0.000 |
| GLM | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.803 | 0.463 | 0.991 | 0.470 | 0.005 | 0.929 | 0.111 | 0.485 | 0.000 | 0.852 | n/a | 0.001 | 0.000 | 0.811 | 0.072 | 0.470 | 0.000 | 0.000 | 0.000 | 0.000 |
| k-NN | 0.884 | 0.234 | 0.736 | 0.266 | 0.230 | 0.002 | 0.009 | 0.001 | 0.009 | 0.578 | 0.000 | 0.099 | 0.008 | 0.096 | 0.001 | 0.001 | n/a | 0.094 | 0.001 | 0.146 | 0.009 | 0.047 | 0.893 | 0.012 | 0.306 |
| SVM | 0.127 | 0.662 | 0.193 | 0.621 | 0.669 | 0.000 | 0.000 | 0.000 | 0.000 | 0.019 | 0.000 | 0.000 | 0.000 | 0.991 | 0.000 | 0.000 | 0.094 | n/a | 0.000 | 0.001 | 0.000 | 0.777 | 0.125 | 0.486 | 0.564 |
| LDA | 0.001 | 0.000 | 0.000 | 0.000 | 0.000 | 0.991 | 0.614 | 0.818 | 0.621 | 0.011 | 0.743 | 0.186 | 0.638 | 0.000 | 0.954 | 0.811 | 0.001 | 0.000 | n/a | 0.125 | 0.621 | 0.000 | 0.001 | 0.000 | 0.000 |
| MDA | 0.109 | 0.005 | 0.069 | 0.006 | 0.005 | 0.127 | 0.330 | 0.073 | 0.328 | 0.405 | 0.058 | 0.852 | 0.309 | 0.001 | 0.111 | 0.072 | 0.146 | 0.001 | 0.125 | n/a | 0.328 | 0.000 | 0.111 | 0.000 | 0.008 |
| NB | 0.006 | 0.000 | 0.003 | 0.000 | 0.000 | 0.624 | 0.991 | 0.473 | 1.000 | 0.054 | 0.421 | 0.426 | 0.972 | 0.000 | 0.584 | 0.470 | 0.009 | 0.000 | 0.621 | 0.328 | n/a | 0.000 | 0.006 | 0.000 | 0.000 |
| 1-R | 0.068 | 0.478 | 0.106 | 0.442 | 0.485 | 0.000 | 0.000 | 0.000 | 0.000 | 0.008 | 0.000 | 0.000 | 0.000 | 0.769 | 0.000 | 0.000 | 0.047 | 0.777 | 0.000 | 0.000 | 0.000 | n/a | 0.066 | 0.675 | 0.400 |
| PART | 0.991 | 0.297 | 0.835 | 0.328 | 0.292 | 0.001 | 0.006 | 0.000 | 0.006 | 0.486 | 0.000 | 0.072 | 0.005 | 0.127 | 0.001 | 0.000 | 0.893 | 0.125 | 0.001 | 0.111 | 0.006 | 0.066 | n/a | 0.019 | 0.376 |
| SAEDNN | 0.020 | 0.266 | 0.037 | 0.234 | 0.269 | 0.000 | 0.000 | 0.000 | 0.000 | 0.002 | 0.000 | 0.000 | 0.000 | 0.485 | 0.000 | 0.000 | 0.012 | 0.486 | 0.000 | 0.000 | 0.000 | 0.675 | 0.019 | n/a | 0.197 |
| RIP | 0.381 | 0.876 | 0.485 | 0.929 | 0.867 | 0.000 | 0.000 | 0.000 | 0.000 | 0.099 | 0.000 | 0.004 | 0.000 | 0.571 | 0.000 | 0.000 | 0.306 | 0.564 | 0.000 | 0.008 | 0.000 | 0.400 | 0.376 | 0.197 | n/a |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tama, B.A.; Lim, S. A Comparative Performance Evaluation of Classification Algorithms for Clinical Decision Support Systems. Mathematics 2020, 8, 1814. https://doi.org/10.3390/math8101814
Tama BA, Lim S. A Comparative Performance Evaluation of Classification Algorithms for Clinical Decision Support Systems. Mathematics. 2020; 8(10):1814. https://doi.org/10.3390/math8101814
Chicago/Turabian StyleTama, Bayu Adhi, and Sunghoon Lim. 2020. "A Comparative Performance Evaluation of Classification Algorithms for Clinical Decision Support Systems" Mathematics 8, no. 10: 1814. https://doi.org/10.3390/math8101814
APA StyleTama, B. A., & Lim, S. (2020). A Comparative Performance Evaluation of Classification Algorithms for Clinical Decision Support Systems. Mathematics, 8(10), 1814. https://doi.org/10.3390/math8101814






