3.1. Pooling Geographically Distributed Information Enhances EDM Performance on Influenza Data
We carried out a series of numerical experiments to test the performance of EDM with and without pooling together geographically-distributed information. This subsection reports results for flu data. Each experiment was carried out for a series of conditions that we label pool, classic, and annual.
In the pool condition, we separated our influenza data series by seasons. To build forecasts for a region in a given season, the pattern library
included past and future seasons from all regions (including the one being forecast), while the data from any region and the same season was removed from
. Note, first, that including such future examples of the season being evaluated is standard in EDM [
14]. We expect that the causality between a year and the next one is fairly broken. Second, imagine that an important region would present some idiosyncratic dynamics during a season, which is later replicated in some other areas. This trend could serve as an indicator of what might happen in those adjacent regions with some delay. If we include all data from a given season, EDM could draw the inference in the opposite direction as well (using data of adjacent regions to forecast dynamics that had played out some weeks ahead). This is why, to be on the safe side, we removed all data from all regions for the season being forecast.
In condition classic, the library of patterns contained only examples from past and future seasons of a given region—which is how EDM was originally conceived [
14] and how it has been applied, e.g., to forecasting flu trends in the past [
15]. In condition annual, the library of patterns consisted of all the contemporary examples of a given region, ignoring all the examples from different years. This condition is proposed to measure the similarity between series from different regions in the same year, as the dominant flu strain will be the same and it might be possible for the effect of a region to be transmitted to a neighboring one within the same season. In
Figure 2 we represent the three conditions.
In
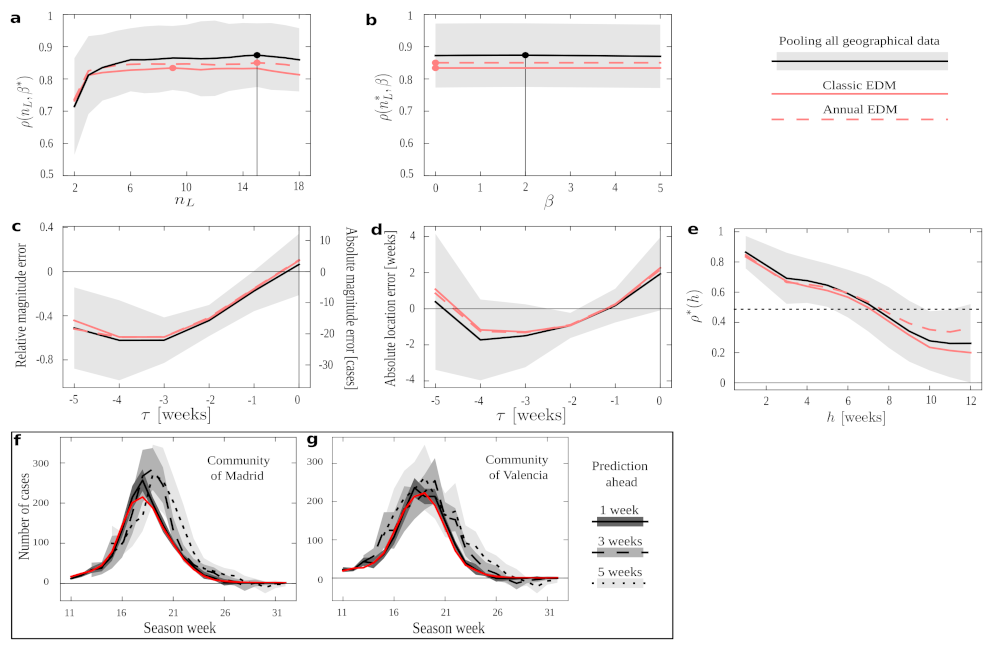
Figure 3 we show EDM performance under the different conditions in a series of numerical experiments. If we keep
fixed, we are simply trying to predict the next amount of new cases following the available data. We see the performance on this task with optimal
and varying
in
Figure 3a; and for optimal
and varying
in
Figure 3b. In both plots, condition pool outperforms all others in almost all the ranges explored and, most importantly, it does so for the optimal
and
(even for the optima derived independently for all other conditions, marked by filled red circles). Such optima would be the meta-parameters with which we should operate if we tried to forecast new time series not present in our data set—and in all cases the results suggest that we should use condition pool. Condition annual performs slightly better than the classic EDM, and both fall below pool. This demonstrates an overall advantage of pooling together epidemiological data across regions. This result might have been expected, since the pool protocol provides us with more data in our training set. However, it is not trivial that dynamics across regions (and, notably, having discarded series of a same season) would be informative to each other—each could have been affected by idiosyncratic factors such as population density, demographic structure, differences between urban and rural dynamics, etc.
The most important features that we would like to predict in an epidemic episode are how many people will be affected and how long it will last. The maximum height and location in time of the peak are a first proxy. To study how well we can forecast this, in a second experiment we aligned the data from all seasons taking each peak as a temporal reference. Then, we looked at how good the forecast of this peak was if EDM only had data until time units (weeks in the case of flu) before.
Figure 3c shows the average error (as relative and absolute magnitudes) that EDM makes in predicting the peak’s height.
Figure 3d shows the error (in weeks) in predicting when the epidemics will reach its maximum. We appreciate that all protocols present quite similar curves. Thus, while pool produces better forecasts on average (as shown above), our results suggest that the uncertainty in predicting the magnitude and end of an epidemic process cannot be alleviated by more abundant data. This is in line with recent research [
12] that shows how behind epidemic processes lie mathematical mechanisms that make them inherently unpredictable. Unfortunately, our non-parametric method cannot circumvent such problems.
Looking at these plots with greater detail, we see how errors become smaller as we get closer to the actual peak—as might be expected (however, observe the case for COVID-19). The smallest average error in magnitude happens as the data up to the very time of the peak is considered (), while location is better predicted a week before the peak happens (). Error changes signs from negative to positive, meaning that EDM progresses (in average) from underestimating to overestimating. The forecast at (which is the furthest from the peak that we can study with the available data) is more accurate than some others for peak magnitude and than many others for peak location. This effect is noteworthy for estimating peak location: this forecast degrades notably before becoming better—perhaps because the steepest phase of the exponential dynamics happens somewhere between and . It is noteworthy, though, that this does not impact magnitude estimation as much.
With time series aligned with respect to their peaks as in the previous experiment, we also measured EDM performance (as captured by correlation between data and estimate) as a function of
. This way, we quantify how well our method works given that it is
time units before or after the peak. Again, all protocols perform quite similarly in this experiment, with pool being notably worse than others in some cases.
Figure 3e shows
, which starts and ends close to 0 (i.e., forecast is of poor quality further away from the peak). Performance raises up to
as the peak is approached, and remains at a similar level right after the peak before starting to decline gently. The dent at
(performance becomes factually nil) is explained because the slope of the data series changes around the peak. Unless both data and prediction are perfectly synchronized (which,
Figure 3d proves, is not the case), this leads to an average correlation of zero at that point.
Finally, it is relevant to establish for how long a forecast remains informative.
Figure 3e shows the EDM performance,
, as it tries to predict
h time units ahead in time. Correlation remains above
for predictions up to 7 weeks ahead of the available data, with pool being the preferred protocol in most cases. (Protocol annual becomes better around the time that correlation drops below
.) We show examples of how a relatively worse (
Figure 3f) and better (
Figure 3g) forecast degrade as we elaborate estimates with more time in advance. We see how this forecast degrades rapidly for a specific season of the Autonomous Community of Madrid, while it remains quite stable for some other seasons in the Community of València. This, together with the large deviations around most of the measures reported in
Figure 3 (gray shadings), suggests that the right protocol might depend on the region studied, and that we might rather address this in a case by case basis. Below, we make some efforts to gain some insight about this issue.
3.2. Exploring EDM on COVID-19 Data
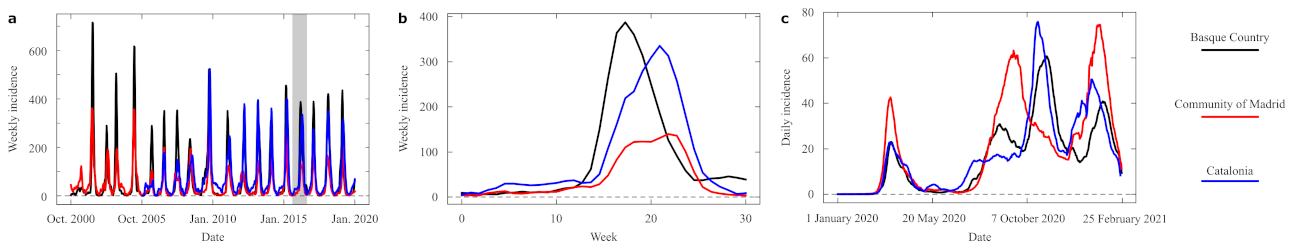
Data of the COVID-19 epidemic dynamics are affected by the various sources of unpredictability discussed above—some related to the unanticipated emergency caused by the pandemics, some others related to intrinsic properties of this malady and our social interplay with it. We have attempted to use EDM, pooling distributed geographic information from various sources, to forecast the dynamic unfolding of this crisis. Our success differed between more global (incorporating data from countries around the world) and local (as in our example from Spanish regions) attempts, and it changed over time as the pandemic changed as well. In this section, we report a brief example based on the same regions as above, now studying only conditions pool and classic. While far from successful, this attempt at forecasting allows us to quantify some aspects that reveal how the new virus unfolds with dynamics very different from those of seasonal influenza. Our data series in this case give us new infections per day, instead of weeks, so some results do not translate as readily.
Figure 4a shows the average EDM performance as a function of
with fixed optimum
. We see an optimum
(days), which is much smaller than the
(weeks) found in the case of flu. This reveals how much more changing are the dynamics for COVID-19, and how informative patterns degrade more promptly as we attempt to compare them during longer stretches of time. This is indicative of a higher number of causal factors taking turns in dominating the dynamics—resulting in a more difficult forecast. Moreover, the correlation between estimates and prediction does not reach values comparable to those achieved with influenza data.
Figure 4b shows EDM performance as a function of
with fixed, optimal
. Again, we see how the pool condition renders better results.
We repeated the experiments to estimate the quality of peak forecast, but in this case taking into account that COVID-19 ‘waves’ are much more vaguely defined than seasonal peaks. Moreover, in some cases, EDM did not forecast the existence of a peak (suggesting, in turn, that the epidemics might grow unstopped within the time-window that we looked ahead). We report only results for cases in which a peak was predicted, and the comparison of its magnitude and location with that of an ongoing wave was possible.
Figure 4c shows that the error in magnitude becomes fairly small around 5 days before the epidemic peaks. By that time, EDM can produce an accurate first proxy of what the number of affected people will be. However,
Figure 4d shows that the error in location of this peak only grows as we become closer to it. This is opposed to the results for flu, for which both magnitude and location estimates improved as the peak was approached. This indicates that EDM is consistently forecasting maxima that lie further away each time in the future of the approaching target and, in other words, suggests that COVID-19 waves do not show tell-tale signs that they are turning—thus aggravating the unpredictability of these kinds of dynamics.
The window of acceptable prediction capabilities is also much smaller for COVID-19 than for flu.
Figure 4e shows how the correlation between estimate and data has already dropped below
if we attempt to predict 5 days ahead. This is an insignificant forecasting window compared to the acceptable 7 weeks that we could look ahead with a similar accuracy in the case of flu. This points, once again, to the dynamical challenge posed by the SARS-CoV-2 pandemics.
Examples of forecast for the Community of Madrid (
Figure 4f) and Community of València (
Figure 4g) show very small deviations from their respective averages. This is due to the very scarce data available, which at the same time reveals a poverty of dynamical patterns to draw estimates from.
3.3. EDM as a Tool to Characterize the Epidemic Unfolding
Non-parametric forecasting methods are mainly results-oriented. They are often used as black-boxes—foregoing a deeper understanding of the dynamic process as long as forecasting works. This is the opposite, e.g., to compartmental modeling, in which causal relationships and meaningful parameters are inferred. With this later approach, insights can be gained about the relevant factors in the unfolding of an epidemic. However, we can turn EDM on its head, using its methods not as a predictor, but as a tool for correlating and clustering the dynamics across regions and years. Then: What regions are more informative to each-other? Can we reveal a spatial structure of how the flu or COVID-19 evolved in Spain? Are there idiosyncratic regions in which the dynamics play out rather differently? How do successive influenza seasons resemble each-other?
There is a question remaining, which is related to the fact that we are introducing data from several regions to predict another one: can we use EDM as a clustering tool?
To answer these questions we scored how often each region was within the nearest networks of each other region. Specifically, we count how many times a region takes examples from the time series of the rest of the regions to make the predictions, assuming that the dynamics of this region is related with the others. We also count how many times it takes examples from its own time series, which remarks that the dynamics of this region is different from the others. Thus, each element of the adjacency matrix characterises the number of examples taken by region i from region j.
This helps us to generate a weighted and directed graph for all regions, which may be useful to study ongoing dynamics where there are not enough past data to make good predictions using other regions’ data. We applied this technique to both diseases—influenza and COVID-19.
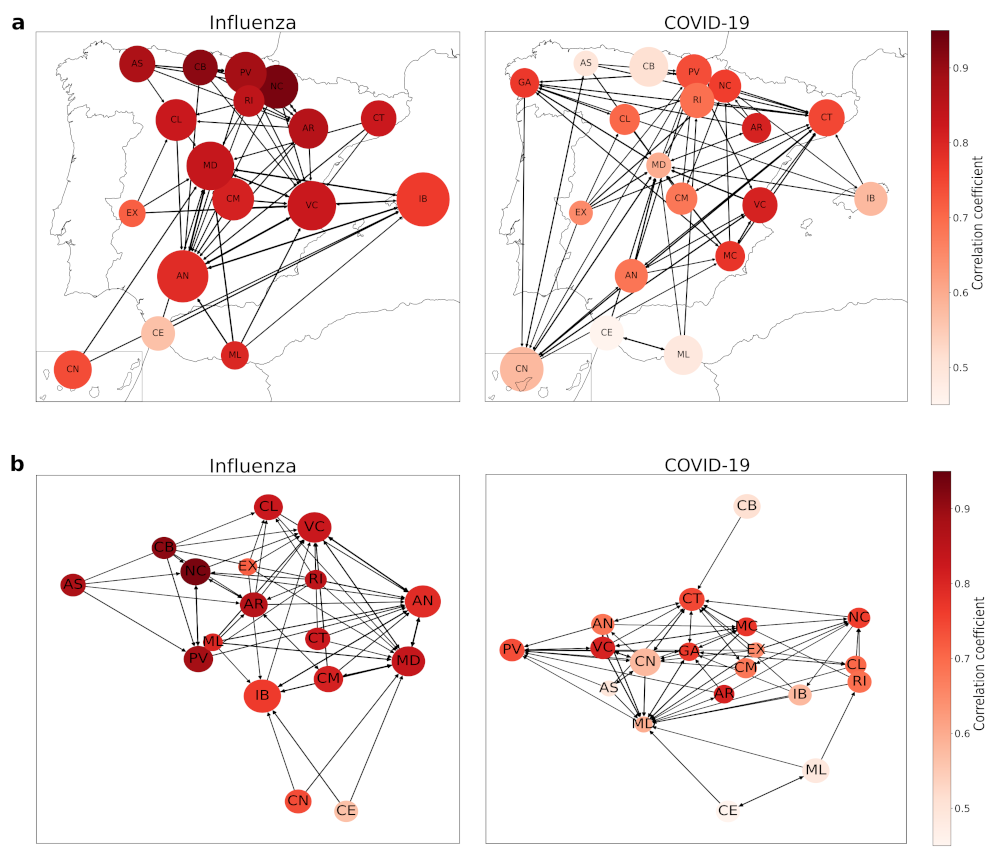
As it can be observed in
Figure 5a, northern and central autonomous regions influenza dynamics are pretty well described with
, while the southern ones are one step behind not so far with
around
Andalusia (AN), Canary Islands (CN), Extremadura (EX), Melilla (ML) and Balearic Islands (IB) as well. Northern regions are also well connected between them and also with Andalusia, which even being at the south keeps a good relationship with other northern autonomous regions. This can be observed at
Figure 5b, where we plot both influenza and COVID-19 networks with a random display, in order to visualize that well-described regions cluster together.
However, the main fact is that none of the considered regions takes more examples from themselves than from the other ones. If we have a look at the proportional number of neighbors chosen by the EDM for one region respective to the others, it goes from to of the total examples, with a mean of and a standard deviation of (the maximum is in the confidence interval of two standard deviations, so we can assume it is just a statistical fluctuation). This means these regions’ dynamics may be similar and EDM does not notice any region to be considered “special” from the others, as all of them take examples from other autonomous regions.
The final goal of this work was to develop a non-parametric prediction method capable of estimating new dynamics when there is no historical data available, such as in the case of the COVID-19 pandemic—as it is a new disease with very little information at its start. We tried to apply this method to COVID-19 data for several regions at a worldwide scale—country incidences—and at a Spanish autonomous regions’ scale, but the results were not as good as the ones obtained in influenza cases. This bad performance can be explained mainly in the lack of historical data, but also because the incidence over all territories have not been the same and not even comparable, as data cannot be scaled from one region to another and examples taken by EDM might not be true to reality. In addition, the quality of the data acquired from governments have not been the best, as at the pandemic start they were running out of tests and the infected people reports were not accurate enough [
16].
This led us to compare how Spanish autonomous regions interacted with each other in this EDM approach considering COVID-19 dynamics from the first wave, from March 2020 to June 2020; the second wave, from June 2020 to December 2020; and part of the third wave, from December 2020 to February 2021, which ensures we have both ascending and descending trends, so EDM will be able to choose which one is better in each analysed case. For this reason, we repeated the clustering experiment for these data, and what we found out was there were many differences from the influenza epidemic network.
Having a look at the proportional number of neighbors chosen by the EDM—as we did in the section before—for one region respective to the others, goes from to of the total examples, with a mean of and a standard deviation of .
There are some differences between the influenza and COVID-19 networks, but the most remarkable one is the fact that some regions take a large number of examples of themselves—in particular, Community of Madrid (MD), Valencian Community (VC), and Andalusia (AN). Correlation coefficients also reflect the bad performance of EDM predicting COVID-19 dynamics, as there are less well-described regions—-with over 0.85.
In terms of connections, we have a more dispersed network, where there is no clear clustering as we had in the influenza network. The northern regions are now more connected with the southern ones, so we could think of it as an insight of a different relationship of similarity than in the influenza case—which could be related to the geographical locations and similar weather, leading to comparable incidences due to the way of life people develop. Now we can observe that dynamics differ from the previous case studied, influenza, probably related to the differences in autonomous regions pandemic management, as they were mainly independent from the central government and they carried out different measures to stop the propagation of this disease, while influenza has been fought for many years and this leads to more homogeneous actions. Despite this dispersion, we can observe in
Figure 5b the regions that are best described are centralized, as they are in the influenza network, denoting the potential application of this clustering method.
In summary, and taking all of this into account, there are several reasons why EDM is not able to perform well with COVID-19 pandemic data, but we can sum them up with two of them: lack of historical data and inhomogeneous disease incidences, which make the regions’ dynamics unpredictable from one to another.