Secretome Analysis of Clavibacter nebraskensis Strains Treated with Natural Xylem Sap In Vitro Predicts Involvement of Glycosyl Hydrolases and Proteases in Bacterial Aggressiveness
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Growth Conditions
2.2. Aggressiveness Assay
2.3. Hydrogen Peroxide and Superoxide Localization
2.4. Xylem Sap Collection
2.5. Bacterial Induction with Xylem Sap
2.6. Protein Extraction, Reduction, Alkylation and Digestion
2.7. LC–MS Analysis
2.8. LC–MS Data Analysis
2.9. Cellulase Activity Assay
2.10. Protease Activity Assay
2.11. Statistical Analysis
3. Results
3.1. Aggressiveness Assay
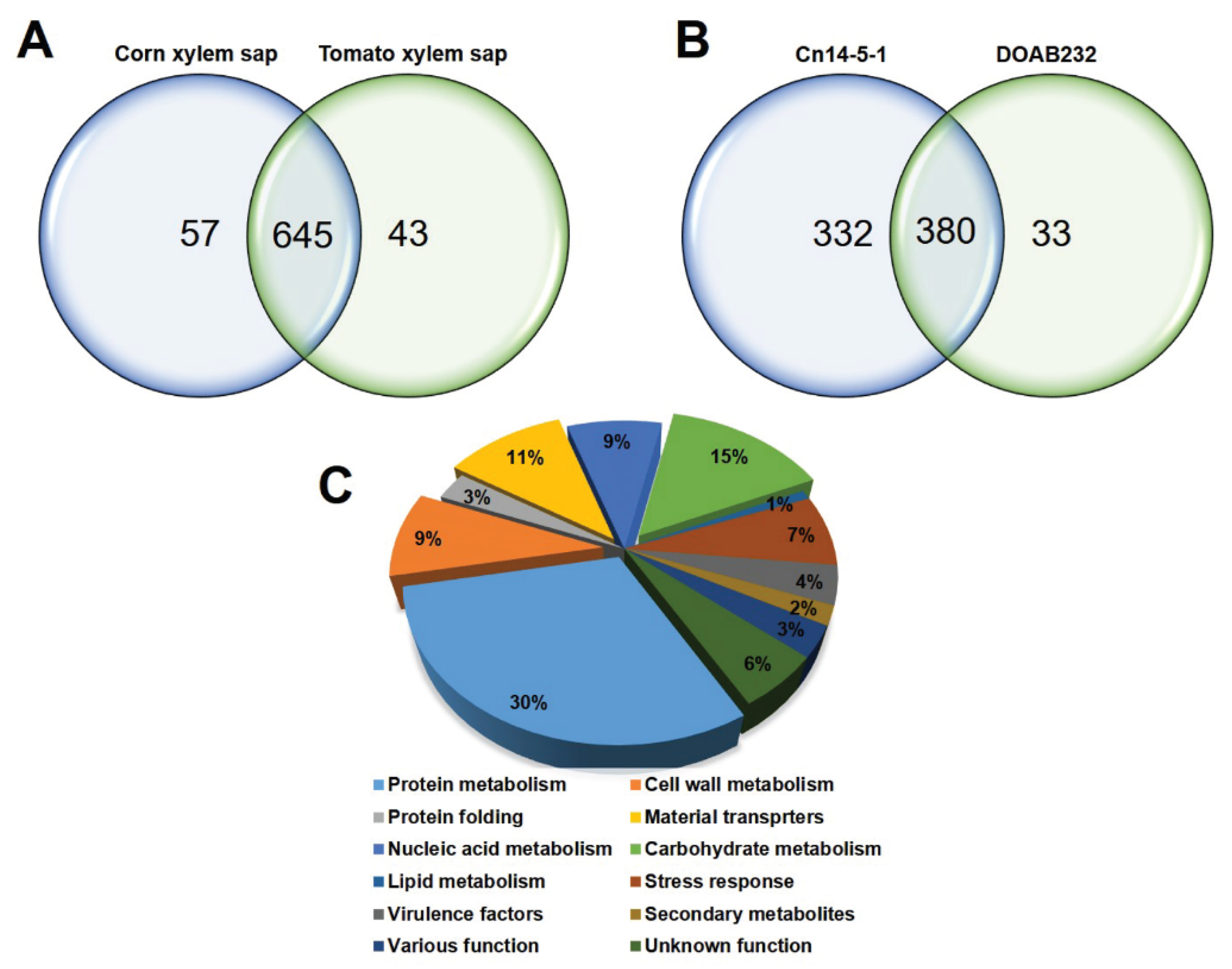
3.2. Characterisation of Cn Secretome by LC–MS Analysis
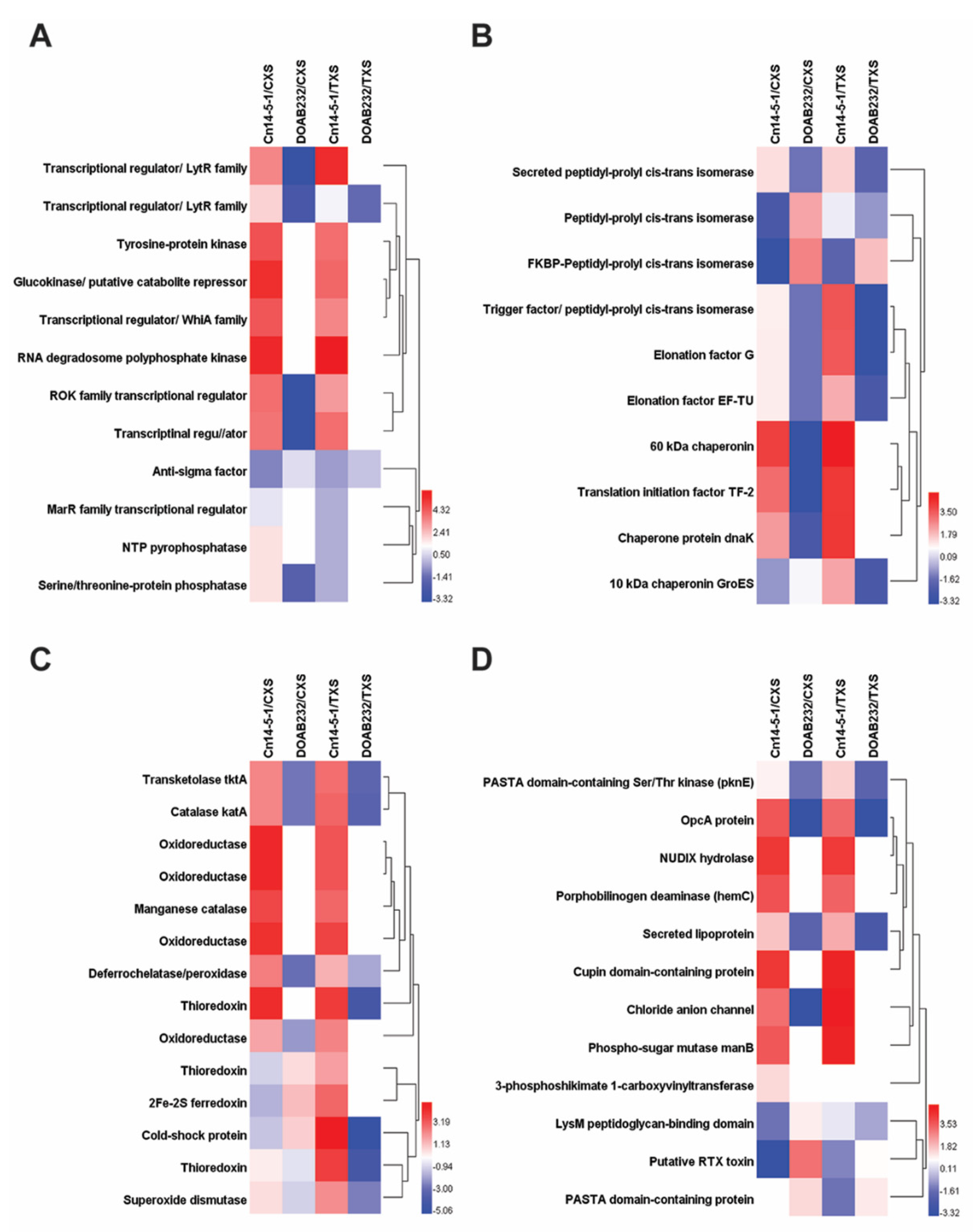
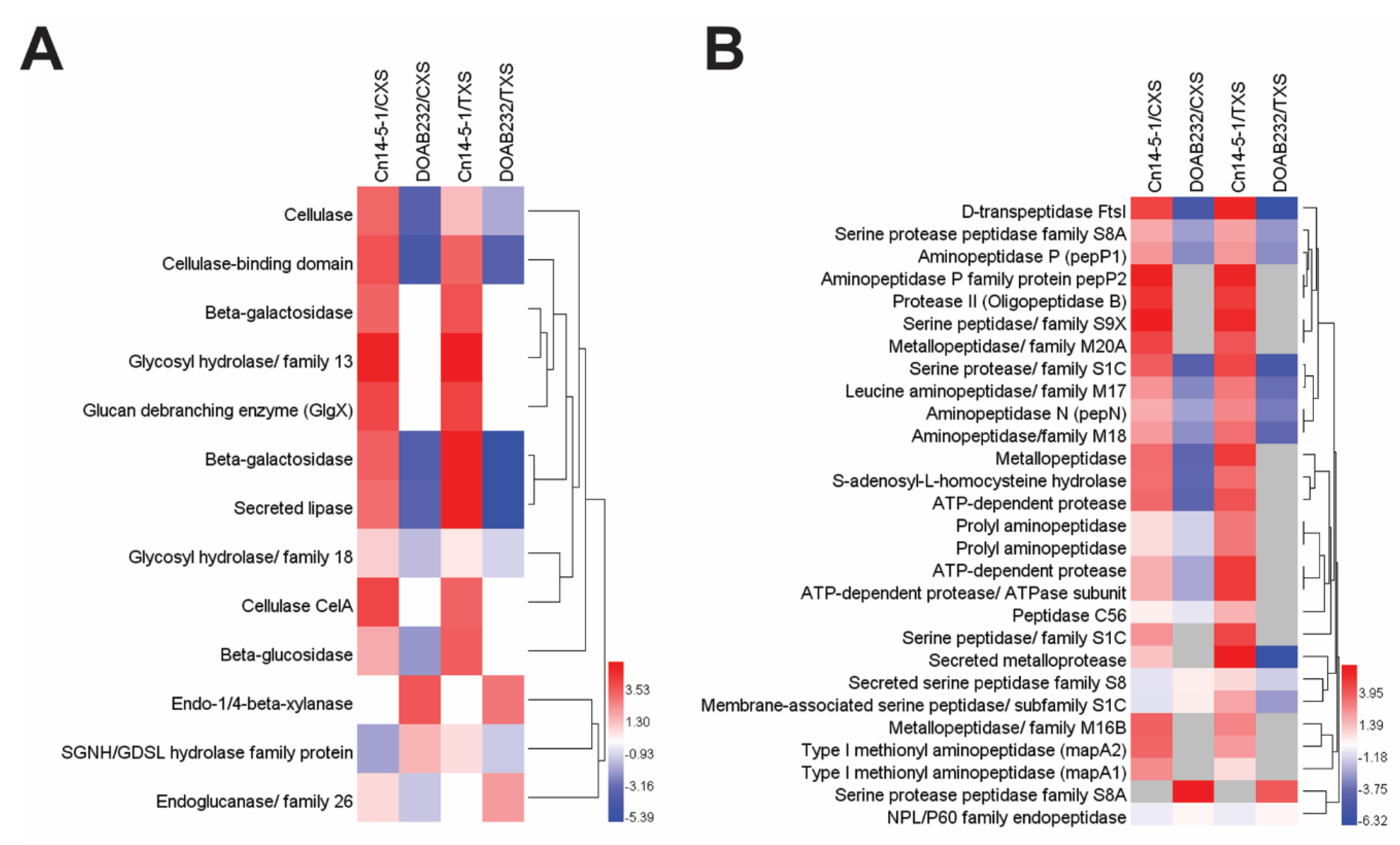
3.3. Abundance of Secreted Proteins in Xylem Sap Media
3.4. Hydrolytic Enzyme Activity Assays
3.5. Hydrogen Peroxide and Superoxide Assays
4. Discussion
5. Conclusions and Future Perspectives
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tambong, J.T. Comparative genomics of Clavibacter michiganensis subspecies, pathogens of important agricultural crops. PLoS ONE 2017, 12, e0172295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jacques, M.A.; Durand, K.; Orgeur, G.; Balidas, S.; Fricot, C.; Bonneau, S.; Quillévéré, A.; Audusseau, C.; Olivier, V.; Grimault, V.; et al. Phylogenetic analysis and polyphasic characterization of Clavibacter michiganensis strains isolated from tomato seeds reveal that nonpathogenic strains are distinct from C. michiganensis subsp. michiganensis. Appl. Environ. Microbiol. 2012, 78, 8388–8402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eichenlaub, R.; Gartemann, K.H. The Clavibacter michiganensis subspecies: Molecular investigation of Gram-positive bacterial plant pathogen. Annu. Rev. Phytopathol. 2011, 49, 445–464. [Google Scholar] [CrossRef] [PubMed]
- Vidaver, A.K.; Mandel, M. Corynebacterium nebraskense, a new orange-pigmented phytopathogenic species. Int. J. Syst. Bacteriol. 1974, 24, 482–485. [Google Scholar] [CrossRef] [Green Version]
- Schuster, M.L. Leaf Freckles and Wilt of Corn Incited by Corynebacterium nebraskense Schuster, Hoff, Mandel, Lazar, 1972; Agricultural Experiment Station, University of Nebraska: Lincoln, NE, USA, 1975. [Google Scholar]
- Li, X.; Tambong, J.; Yuan, K.X.; Chen, W.; Xu, H.; Lévesque, C.A.; De Boer, S.H. Reclassification of Clavibacter michiganensis subspecies on the basis of whole-genome and multi-locus sequence analyses. Int. J. Syst. Evol. Microbiol. 2018, 68, 234–240. [Google Scholar] [CrossRef]
- Wysong, D.S.; Doupnik, J.B.; Lane, L. Goss’s wilt and corn lethal necrosis—Can they become a major problem? In Proceedings of the Annual Corn Sorghum Research Conference; 36th American Seed Trade Association: Washington, DC, USA, 1981; pp. 104–130. [Google Scholar]
- Rocheford, T.R.; Vidaver, A.K.; Gardner, C.O.; Arbust, D.L. Effect of wind-generated sand abrasion on infection of corn (Zea mays L.) by Corynebacterium michiganense ssp. nebraskense. Phytopathology 1985, 75, 1378. [Google Scholar]
- Agarkova, I.V.; Lambrecht, P.A.; Vidaver, A.K. Genetic diversity and population structure of Clavibacter michiganensis subsp. nebraskensis. Can. J. Microbiol. 2011, 57, 366–374. [Google Scholar] [CrossRef]
- Langemeier, C.B.; Robertson, A.E.; Wang, D.; Jackson-Ziems, T.A. Factors affecting the development and severity of Goss’s bacterial wilt and leaf blight of corn, caused by Clavibacter michiganensis subsp. nebraskensis. Plant Dis. 2017, 101, 54–61. [Google Scholar] [CrossRef] [Green Version]
- Claflin, L.E. Goss’s bacterial wilt and blight. In Compendium of Corn Diseases; White, D.H., Ed.; American Phytopathological Society: St. Paul, MN, USA, 1999. [Google Scholar]
- Mallowa, S.; Mbofung, G.; Eggenberger, S.; Den Adel, R.; Sheiding, S.R.; Robertson, A.E. Infection of maize by Clavibacter michiganensis subsp. nebraskensis does not require severe wounding. Plant Dis. 2016, 100, 724–731. [Google Scholar] [CrossRef] [Green Version]
- Langemeier, C.B.; Jackson-Ziems, T.A.; Kruger, G.R. Four common Setaria Species are alternative hosts for Clavibacter michiganensis subsp. nebraskensis, causal agent of Goss’s bacterial wilt and blight of corn. Plant Health Prog. 2014, 15, 57–60. [Google Scholar] [CrossRef] [Green Version]
- Ikley, J.T.; Wise, K.A.; Johnson, W.G. Annual ryegrass (Lolium multiflorum), Johnsongrass (Sorghum halepense), and large crabgrass (Digitaria sanguinalis) are alternative hosts for Clavibacter michiganensis subsp. nebraskensis, causal agent of Goss’s wilt of corn. Weed Sci. 2015, 63, 901–909. [Google Scholar]
- Savidor, A.; Teper, D.; Gartemann, K.-H.; Eichenlaub, R.; Chalupowicz, L.; Manulis-Sasson, S.; Barash, I.; Tews, H.; Mayer, K.; Giannone, R.; et al. The Clavibacter michiganensis subsp. michiganensis–tomato interactome reveals the perception of pathogen by the host and suggests mechanisms of infection. J. Proteome Res. 2012, 11, 736–750. [Google Scholar] [CrossRef] [PubMed]
- Hiery, E.; Poetsch, A.; Moosbauer, T.; Amin, B.; Hofmann, J.; Burkovski, A. A proteomic study of Clavibacter michiganensis subsp. michiganensis culture supernatants. Proteomes 2015, 3, 411–423. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gross, D.C.; Vidaver, A.K. A selective medium for isolation of Corynebacterium nebraskense from soil and plant parts. Phytopathology 1979, 69, 82–87. [Google Scholar] [CrossRef]
- Soliman, A.; Gulden, R.H.; Tambong, J.T.; Bajracharya, P.; Adam, L.R.; Xu, R.; Cott, M.; Daayf, F. Developed and validated inoculation and disease assessment methods for Goss’s bacterial wilt and leaf blight disease of corn. Crop Prot. 2018, 112, 59–167. [Google Scholar] [CrossRef]
- Campbell, M.T.; Proctor, C.A.; Dou, Y.; Schmitz, A.J.; Phansak, P.; Kruger, G.R.; Walia, H. Genetic and molecular characterization of submergence response identifies Subtol6 as a major submergence tolerance locus in maize. PLoS ONE 2015, 10, e0120385. [Google Scholar] [CrossRef]
- Bradford, M. A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1979, 72, 248–254. [Google Scholar] [CrossRef]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 2019, 47, D442–D450. [Google Scholar] [CrossRef]
- Keller, A.; Nesvizhskii, A.I.; Kolker, E.; Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 2002, 74, 5383–5392. [Google Scholar] [CrossRef]
- Nesvizhskii, A.I.; Keller, A.; Kolker, E.; Aebersold, R.A. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 2003, 75, 4646–4658. [Google Scholar] [CrossRef]
- Zhang, Y.H.P.; Hong, J.; Ye, X. Cellulase Assays. In Biofuels: Methods in Molecular Biology; Mielenz, J., Ed.; Humana Press: Totowa, NJ, USA, 2009; Volume 581. [Google Scholar]
- Cupp-Enyard, C. Sigma’s non-specific protease activity assay—Casein as a substrate. JoVE 2008, 19, 899. [Google Scholar] [CrossRef] [PubMed]
- Lund, R.E. Tables for an approximate test for outliers in linear models. Technometrics 1975, 17, 473–476. [Google Scholar] [CrossRef]
- Saxton, A.M. A macro for converting mean separation output to letter groupings in PROC MIXED. In Proceedings of the 23rd SAS Users Group International; SAS Institute: Cary, NC, USA, 1998; pp. 1243–1246. [Google Scholar]
- Peritore-Galve, F.C.; Schneider, D.J.; Yang, Y.; Thannhauser, T.W.; Smart, C.D.; Stodghill, P. Proteome profile and genome refinement of the tomato-pathogenic bacterium Clavibacter michiganensis subsp. michiganensis. Proteomics 2019, 19, e1800224. [Google Scholar] [CrossRef] [PubMed]
- Tambong, J.T.; Xu, R.; Daayf, F.; Brière, S.; Bilodeau, G.J.; Tropiano, R.; Hartke, A.; Reid, L.M.; Cott, M.; Cote, T.; et al. Genome analysis and development of a multiplex TaqMan real-time PCR for specific identification and detection of Clavibacter michiganensis subsp. nebraskensis. APS Phytopathol. 2016, 106, 1473–1485. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bellincampi, D.; Cervone, F.; Lionetti, V. Plant cell wall dynamics and wall-related susceptibility in plant–pathogen interactions. Front. Plant Sci. 2014, 28, 228. [Google Scholar] [CrossRef] [Green Version]
- Lalaoui, F.; Halama, P.; Dumortier, V.; Paul, B. Cell wall-degrading enzymes produced in vitro by isolates of Phaeosphaeria nodorum differing in aggressiveness. Plant Pathol. 2000, 49, 727–733. [Google Scholar] [CrossRef]
- Barras, F.; Gijsegem, F.V.; Chatterjee, A.K. Extracellular enzymes and pathogenesis of soft-rot Erwinia. Annu. Rev. Phytopathol. 1994, 32, 201–234. [Google Scholar] [CrossRef]
- Melotto, M.; Kunkel, B.N. Virulence strategies of plant pathogenic bacteria. In The Prokaryotes; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2013; pp. 61–82. [Google Scholar]
- Holtsmark, I.; Takle, G.; Brurberg, M. Expression of putative virulence factors in the potato pathogen Clavibacter michiganensis subsp. sepedonicus during infection. Arch. Microbiol. 2008, 189, 131–139. [Google Scholar]
- Laine, M.J.; Haapalainen, M.; Wahlroos, T.; Kankare, K.; Nissinen, R.; Kassuwi, S.; Metzler, M.C. The cellulase encoded by the native plasmid of Clavibacter michiganensis ssp. sepedonicus plays a role in virulence and contains an expansin-like domain. Phys. Mol. Plant Pathol. 2000, 57, 221–233. [Google Scholar]
- Nissinen, R.; Kassuwi, S.; Peltola, R.; Metzler, M.C. In planta—Complementation of Clavibacter michiganensis subsp. sepedonicus strains deficient in cellulase production or HR induction restores virulence. Eur. J. Plant Pathol. 2001, 107, 175–182. [Google Scholar] [CrossRef]
- Ahmad, A.; Mbofung, G.Y.; Acharya, J.; Schmidt, C.L.; Robertson, A.E. Characterization and comparison of Clavibacter michiganensis subsp. nebraskensis strains recovered from epiphytic and symptomatic infections of maize in Iowa. PLoS ONE 2015, 10, e0143553. [Google Scholar] [CrossRef] [PubMed]
- Hoston, A.; Mudgett, M.B. Cysteine proteases in phytopathogenic bacteria: Identification of plant targets and activation of innate immunity. Curr. Opin. Plant Biol. 2004, 7, 384–390. [Google Scholar]
- Figaj, D.; Ambroziak, P.; Przepiora, T.; Skorko-Glonek, J. The role of proteases in the virulence of plant pathogenic bacteria. Int. J. Mol. Sci. 2019, 20, 672. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boos, W.; Shuman, H. Maltose/maltodextrin system of Escherichia coli: Transport, metabolism, and regulation. Microbiol. Mol. Biol. Rev. 1998, 62, 204–229. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lengsfeld, C.; Schönert, S.; Dippel, R.; Boos, W. Glucose-and Glucokinase-controlled mal gene expression in Escherichia coli. J. Bacteriol. 2009, 191, 701–712. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yeh, J.I.; Kettering, R.; Saxl, R.; Bourand, A.; Darbon, E.; Joly, N.; Briozzo, P.; Deutscher, J. Structural characterizations of glycerol kinase: Unraveling phosphorylation-induced long-range activation. Biochemistry 2009, 48, 346–356. [Google Scholar] [CrossRef] [Green Version]
- Grove, A. Regulation of metabolic pathways by MarR family transcription factors. Comput. Struct. Biotechnol. J. 2017, 15, 366–371. [Google Scholar] [CrossRef]
- Ramos, J.L.; Martínez-Bueno, M.; Molina-Henares, A.J.; Terán, W.; Watanabe, K.; Zhang, X.; Gallegos, M.T.; Brennan, R.; Tobes, R. The TetR family of transcriptional repressors. Mol. Biol. Rev. 2005, 69, 326–356. [Google Scholar] [CrossRef] [Green Version]
- Cuthbertson, L.; Nodwella, J.R. The TetR Family of regulators. Microbiol. Mol. Biol. Rev. 2013, 77, 440–475. [Google Scholar] [CrossRef] [Green Version]
- Titgemeyer, F.; Reizer, J.; Reizer, A.; Saier, M.H.J. Evolutionary relationships between sugar kinases and transcriptional repressors in bacteria. Microbiology 1994, 140, 2349–2354. [Google Scholar] [CrossRef] [Green Version]
- Kazanov, M.D.; Li, X.; Gelfand, M.S.; Osterman, A.L.; Rodionov, D.A. Functional diversification of ROK-family transcriptional regulators of sugar catabolism in the Thermotogae phylum. Nucleic Acids Res. 2012, 41, 790–803. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thapa, S.P.; Pattathil, S.; Hahn, M.G.; Jacques, M.-A.; Gilbertson, R.L.; Coaker, G. Genomic analysis of Clavibacter michiganensis reveals insight into virulence strategies and genetic diversity of a Gram-positive bacterial pathogen. Mol. Plant Microbe Interact 2017, 30, 786–802. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laluk, K.; Mengiste, T. Necrotroph attacks on plants: Wanton Destruction or Covert Extortion? Arabidopsis Book 2010, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fatima, U.; Senthil-Kumar, M. Plant and pathogen nutrient acquisition strategies. Front. Plant Sci. 2015, 6, 750. [Google Scholar] [CrossRef]
- Delmotte, N.; Knief, C.; Chaffron, S.; Innerebner, G.; Roschitzki, B.; Schlapbach, R.; von Mering, C.; Vorholt, J.A. Community proteogenomics reveals insights into the physiology of phyllosphere bacteria. Proc. Natl. Acad. Sci. USA 2009, 106, 16428–16433. [Google Scholar] [CrossRef] [Green Version]
- Yu, J.; Ge, J.; Heuveling, J.; Schneider, E.; Yang, M. Structural basis for substrate specificity of an amino acid ABC transporter. Proc. Natl. Acad. Sci. USA 2015, 112, 5243–5248. [Google Scholar] [CrossRef] [Green Version]
- Braakman, I.; Hebert, D.N. Protein folding in the endoplasmic reticulum. Cold Spring Harb. Perspect. Biol. 2013, 5, a013201. [Google Scholar] [CrossRef] [Green Version]
- Park, C.-J.; Seo, Y.-S. Heat Shock Proteins: A Review of the Molecular Chaperones for Plant Immunity. Plant Pathol. J. 2015, 31, 323–333. [Google Scholar] [CrossRef] [Green Version]
- Stoller, G.; Rucknagel, K.P.; Nierhaus, K.H.; Schmid, F.X.; Fischer, G.; Rahfeld, J.-U. A ribosome-associated peptidyl-prolyl cis/trans isomerase identified as the trigger factor. EMBO J. 1995, 20, 4939–4948. [Google Scholar] [CrossRef]
- Rospert, S.; Glick, B.S.; Jenö, P.; Schatz, G.; Todd, M.J.; Lorimer, G.H.; Viitanen, P.V. Identification and functional analysis of chaperonin 10, the groES homolog from yeast mitochondria. Proc. Natl. Acad. Sci. USA 1993, 90, 10967–10971. [Google Scholar] [CrossRef] [Green Version]
- Levy-Rimler, G.; Viitanen, P.; Weiss, C.; Sharkia, R.; Greenberg, A.; Niv, A.; Lustig, A.; Delarea, Y.; Azem, A. The effect of nucleotides and mitochondrial chaperonin 10 on the structure and chaperone activity of mitochondrial chaperonin 60. Eur. J. Biochem. 2001, 268, 3465–3472. [Google Scholar] [CrossRef] [PubMed]
- Fones, H.; Preston, G.M. Reactive oxygen and oxidative stress tolerance in plant pathogenic Pseudomonas. FEMS Microbiol. Lett. 2012, 327, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hasset, D.J.; Cohen, M.S. Bacterial adaptation to oxidative stress: Implications for pathogenesis and interaction with phagocytic cells. FASEB J. 1989, 3, 2574–2582. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kovacs-Simon, A.; Titball, R.W.; Michell, S.L. Lipoproteins of bacterial pathogens. Infect. Immun. 2011, 79, 548–561. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zückert, W.R. Secretion of bacterial lipoproteins: Through the cytoplasmic membrane, the periplasm and beyond. Biochim. Biophys. Acta 2014, 1843, 1509–1516. [Google Scholar] [CrossRef] [Green Version]
- El Hadrami, A.; Islam, M.R.; Adam, L.R.; Fouad, D. A cupin domain-containing protein with a quercetinase activity (VdQase) regulates Verticillium dahliae’s pathogenicity and contributes to counteracting host defenses. Front. Plant Sci. 2015, 6, 440. [Google Scholar] [CrossRef] [Green Version]
- Débarbouillé, M.; Dramsi, S.; Dussurget, O.; Nahori, M.-A.; Vaganay, E.; Jouvion, G.; Cozzone, A.; Msadek, T.; Duclos, B. Characterization of a serine/threonine kinase involved in virulence of Staphylococcus aureus. J. Bacteriol. 2009, 191, 4070–4081. [Google Scholar] [CrossRef] [Green Version]

| Identified Proteins | Corn Xylem Sap (CXS) | Tomato Xylem Sap (TXS) | ||
|---|---|---|---|---|
| Cn14-5-1 | DOAB232 | Cn14-5-1 | DOAB232 | |
| Total proteins | 658 | 396 | 678 | 229 |
| Unique proteins | 269 | 40 | 459 | 10 |
| Differentially abundant | 270 | 40 | 200 | 18 |
| Accession No. | Identified Proteins | MW (KDa) | Gene/Locus | Type of Sap | Functional Group | |
|---|---|---|---|---|---|---|
| CXS | TXS | |||||
| CCE76040.1 | Serine peptidase, family S9X | 76 | CMN_02099 | + | − | Protein degradation |
| CCE75111.1 | pepP2 protein | 57 | pepP2 | + | − | Protein degradation |
| CCE76391.1 | Polyphosphate kinase | 82 | ppK | + | − | Protein phosphorylation |
| CCE75799.1 | Glucokinase | 33 | glkA | + | − | Catabolite repressor |
| CCE74648.1 | Radical SAM domain-containing protein | 46 | CMN_00689 | + | − | Stress response |
| CCE76022.1 | Protease II (oligopeptidase B) | 80 | ptrB | + | − | Protein degradation |
| CCE75620.1 | Metallopeptidase, peptidase family M20A | 47 | CMN_01675 | + | − | Protein degradation |
| CCE74827.1 | Putative tyrosine-protein kinase | 49 | CMN_00871 | + | − | Stress response |
| CCE74652.1 | Oxidoreductase | 35 | CMN_00693 | + | + | Stress response |
| CCE75671.1 | Transcriptional regulator, WhiA family | 35 | whiA | + | − | Signal transduction |
| CCE74496.1 | Thioredoxin | 21 | ccbD | + | − | Stress response |
| CCE75993.1 | Oxidoreductase | 36 | CMN_02051 | + | + | Stress response |
| CCE75476.1 | NUDIX hydrolase | 19 | CMN_01528 | + | + | Virulence |
| CCE76025.1 | Cupin_2 domain-containing protein | 32 | CMN_02083 | + | + | Virulence |
| AHN16207.1 | Cellulase | 78 | CelA | + | + | Pathogenicity |
| CCE75945.1 | Metallopeptidase, family M16B | 48 | pepR | + | − | Protein degradation |
| CCE76486.1 | Methionine aminopeptidase | 29 | mapA2 | + | − | Protein degradation |
| CAN00621.1 | Porphobilinogen deaminase | 34 | hemC | + | + | Pathogenicity |
| AJW80248.1 | Manganese catalase | 31 | DZF93_04220 | + | + | Stress response |
| CCE74906.1 | ManB protein | 61 | manB | + | + | Pathogenicity |
| CCE74104.1 | Putative glycosyl hydrolase, family 2 | 69 | CMN_00123 | + | − | Pathogenicity |
| AJW78817.1 | ATP-dependent Clp protease proteolytic subunit | 21 | clpP | + | + | Stress response |
| CCE76317.1 | Serine peptidase, family S1C | 51 | CMN_02381 | + | − | Protein degradation |
| CCE75886.1 | NTP pyrophosphatase | 22 | CMN_01943 | + | − | Signal transduction |
| CCE74756.1 | Putative secreted metalloprotease | 46 | CMN_00799 | + | − | Protein degradation |
| CCE76463.1 | Alanine racemase/kinase fusion protein | 59 | alr2 | + | − | Signal transduction |
| AJW79024.1 | HAD family phosphatase | 25 | VO01_07690 | + | − | Signal transduction |
| AJW80208.1 | MarR family transcriptional regulator | 17 | DZF93_09435 | + | − | Signal transduction |
| CCE74692.1 | Chloride anion channel | 25 | CMN_00734 | − | + | Pathogenicity |
| CCE74774.1 | ATP-dependent protease, ATPase subunit | 92 | clpC | − | + | Protein degradation |
| CCE74648.1 | Radical SAM domain-containing protein | 46 | CMN_00689 | − | + | Stress response |
| CCE76686.1 | Alpha-glucosidase, glycosyl hydrolase family 13 | 64 | aglC | − | + | Pathogenicity |
| CCE75391.1 | ATP-dependent protease, proteolytic subunit | 25 | clpP2 | − | + | Stress response |
| CCE76068.1 | 1-Deoxy-D-xylulose 5-phosphate reductoisomerase | 38 | dxr | − | + | Pathogenicity |
| CCE75358.1 | Glucan debranching enzyme | 82 | treX | − | + | Pathogenicity |
| CCE74827.1 | Putative tyrosine-protein kinase | 49 | CMN_00871 | − | + | Stress response |
| CCE76839.1 | 2-Keto acid dehydrogenase | 41 | CMN_02915 | − | + | Energy production |
| CCE74969.1 | Lipoprotein | 59 | lpqB | − | + | Pathogenicity |
| CCE74801.1 | Beta-glycosidase | 54 | bglJ | − | + | Pathogenicity |
| CCE76286.1 | Alpha glycosidase | 81 | CMN_02350 | − | + | Pathogenicity |
| AJW79410.1 | Sugar kinase | 32 | VO01_09940 | − | + | Signal transduction |
| CCE74250.1 | Alpha-L-arabinofuranosidase | 55 | abfA1 | − | + | Pathogenicity |
| AJW78269.1 | Organic hydroperoxide resistance protein | 14 | Ohr | − | + | Stress response |
| Accession No. | Identified Proteins | MW (KDa) | Gene/Locus | Fisher’s Exact Test | Type of Sap | Fold Change | Functional Group | |
|---|---|---|---|---|---|---|---|---|
| p-Value (p < 0.05) | CXS | TXS | ||||||
| CCE76069.1 | Secreted peptidyl-prolyl cis-trans isomerase | 34 | CMN_02128 | 0.00013 | − | + | 3.2 | Protein folding |
| ALD12817.1 | Cold-shock protein | 7 | AES38_07750 | 0.00018 | − | + | 38 | Stress response |
| CCE74756.1 | Putative secreted metalloprotease | 46 | CMN_00799 | 0.00029 | − | + | 80 | Protein degradation |
| CCE75860.1 | Secreted lipase | 30 | CMN_01917 | 0.00047 | + | − | 10 | Pathogenicity |
| CCE76016.1 | Beta-galactosidase, lactase | 113 | lacZ | 0.001 | − | + | 42 | Pathogenicity |
| CCE75767.1 | Metallopeptidase | 49 | CMN_01822 | 0.0016 | + | − | 14 | Protein degradation |
| CCE74047.1 | Chaperone protein dnaK | 67 | dnaK | 0.0017 | + | − | 6.1 | Protein folding |
| CCE74654.1 | Catalase | 57 | katA | 0.0018 | + | − | 5.8 | Stress response |
| CCE75796.1 | Non-specific serine/threonine protein kinase | 69 | pknE | 0.0051 | − | + | 3.3 | Virulence |
| CCE75569.1 | Leucine aminopeptidase, family M17 | 52 | pepA | 0.006 | + | − | 6.9 | Protein degradation |
| CCE74438.1 | Transcriptional regulator, LytR family | 43 | CMN_00472 | 0.009 | + | − | 13 | Virulence |
| CCE74438.1 | Transcriptional regulator, LytR family | 43 | CMN_00472 | 0.013 | − | + | 48 | Virulence |
| CCE75659.1 | Transketolase | 75 | tktA | 0.015 | + | − | 5.8 | Stress response |
| CCE74692.1 | Chloride anion channel | 25 | CMN_00734 | 0.038 | + | − | 11 | Virulence |
| CCE75389.1 | Peptidyl-prolyl cis-trans isomerase | 52 | tig | 0.043 | − | + | 14 | Pathogenicity |
| CCE74969.1 | Lipoprotein | 59 | lpqB | 0.044 | + | − | 22 | Virulence |
| CCE74654.1 | Catalase | 57 | katA | 0.071 | − | + | 9.2 | Stress response |
| CCE75600.1 | Endo-1,4-beta-xylanase | 71 | xysB | 0.084 | − | + | 2.5 | Pathogenicity |
| AJW79410.1 | Sugar kinase | 32 | VO01_09940 | 0.089 | + | − | 18 | Signal transduction |
| CCE76365.1 | 60 KDa chaperonin | 57 | groEL | <0.00010 | + | − | 19 | Protein folding |
| CCE74122.1 | Cellulose-bindingand an expansin domain | 37 | CMN_00144 | <0.00010 | + | − | 16 | Pathogenicity |
| CCE76326.1 | Secreted cellulase | 58 | cel | <0.00010 | + | − | 11 | Pathogenicity |
| CCE75788.1 | FtsI protein | 62 | ftsI | <0.00010 | + | − | 30 | Pathogenicity |
| CCE75200.1 | Serine protease, family S1C | 50 | CMN_01248 | <0.00010 | + | − | 18 | Protein degradation |
| CCE76016.1 | Beta-galactosidase, lactase | 113 | lacZ | <0.00010 | + | − | 13 | Pathogenicity |
| CCE75860.1 | Secreted lipase | 30 | CMN_01917 | <0.00010 | − | + | 41 | Pathogenicity |
| CCE76365.1 | 60 KDa chaperonin | 57 | groEL | <0.00010 | − | + | 37 | Protein folding |
| CCE75200.1 | Serine protease, family S1C | 50 | CMN_01248 | <0.00010 | − | + | 29 | Protein degradation |
| CCE74047.1 | Chaperone protein dnaK | 67 | dnaK | <0.00010 | − | + | 22 | Protein folding |
| CCE74122.1 | Cellulose-binding and an expansin domain | 37 | CMN_00144 | <0.00010 | − | + | 12 | Pathogenicity |
| CCE76326.1 | Endoglucanase | 58 | cel | <0.00010 | − | + | 3 | Pathogenicity |
| CCE76173.1 | Secreted serine peptidase family S8 | 43 | CMN_02235 | <0.00010 | − | + | 2.1 | Protein degradation |
| CAQ02078.1 | Glucose-6-phosphate 1-dehydrogenase | 58 | Zwf | 0.025 | + | − | 4.9 | Carb. Metabolism |
| CCE75669.1 | Glyceraldehyde 3-phosphate dehydrogenase | 36 | gapA | <0.00010 | + | − | 7.9 | Carb. Metabolism |
| CCE74538.1 | Putative levansucrase | 65 | sacB | 0.00051 | − | + | 4.3 | Carb. Metabolism |
| AJW79067.1 | OpcA protein | 35 | DZF93_01230 | 0.013 | + | − | 14 | Virulence |
| CCE75796.1 | PASTA domain containing Ser/Thr kinase | 69 | pknE | 0.025 | + | − | 2.3 | Virulence |
| CCE74059.1 | Glycosyl hydrolase, (chitinase) family 18 | 39 | CMN_00077 | 0.0026 | + | − | 1.6 | Pathogenicity |
| Accession No. | Identified Proteins | MW (KDa) | Gene/Locus | Type of Sap | Functional Group | |
|---|---|---|---|---|---|---|
| CXS | TXS | |||||
| CCE75047.1 | Putative secreted 5’-nucleotidase | 73 | CMN_01095 | + | + | Nucleotide catabolism |
| CAQ00484.1 | Putative solute-binding lipoprotein | 46 | CMS0363 | + | + | Virulence |
| AJW80270.1 | Integration host factor | 10 | VO01_15120 | + | − | Stress response |
| CCE74134.1 | Rhodanese domain-containing protein | 10 | CMN_00157 | + | − | Stress response |
| CCE74199.1 | Sugar ABC transporter | 43 | CMN_00224 | + | − | Cell surface |
| CCE75732.1 | Rhodanese domain-containing protein | 12 | CMN_01787 | + | − | Stress response |
| AJW79484.1 | Exodeoxyribonuclease VII small subunit | 9 | xseB | + | − | DNA catabolic |
| AJW78093.1 | General stress protein CsbD | 6 | DZF93_05670 | + | − | Stress response |
| CCE74980.1 | Alkyl hydroperoxide reductase | 17 | bcp | + | − | Stress response |
| AJW79401.1 | Antibiotic biosynthesis monooxygenase | 11 | DZF93_04600 | + | − | Stress response |
| CCE74048.1 | Heat shock chaperone GrpE | 24 | grpE | + | − | Protein folding |
| CCE76425.1 | Secreted serine protease, peptidase family S8A | 121 | sbtB | + | + | Protein degradation |
| CCE75599.1 | Endo-1,4-beta-xylanase | 45 | xysA | + | + | Pathogenicity |
| CCE76583.1 | Endoglucanase, glycosyl hydrolase family 26 | 47 | CMN_02651 | − | + | Pathogenicity |
| CCE74634.1 | Esterase | 25 | CMN_00675 | − | + | Virulence |
| Accession No. | Identified Proteins | MW (KDa) | Gene/Locus | Fisher’s Exact Test | Type of Sap | Fold Change | Functional Group | |
|---|---|---|---|---|---|---|---|---|
| p-Value (p < 0.05) | CXS | TXS | ||||||
| CCE74709.1 | Putative extracellular nuclease | 74 | CMN_00751 | <0.00010 | + | − | 6.0 | Nucleotide degradation |
| A5CU64.1 | 10 kDa chaperonin | 11 | groES | <0.00010 | + | − | 1.8 | Protein folding |
| ALD12817.1 | Cold-shock protein | 7 | AES38_07750 | <0.00010 | + | − | 2.1 | Stress response |
| CCE76393.1 | Phosphate-binding protein PstS | 37 | pstS | <0.00010 | + | − | 1.2 | Transport protein |
| CCE76173.1 | Secreted serine peptidase family S8 | 43 | CMN_02235 | <0.00010 | + | − | 1.4 | Protein degradation |
| CCE74340.1 | Levan fructotransferase | 57 | CMN_00371 | <0.00010 | + | − | 4.1 | Carb. metabolism |
| CCE76401.1 | Putative hydrolase | 22 | CMN_02466 | <0.00010 | + | − | 3.4 | Protein degradation |
| CCE76077.1 | Putative RTX toxin | 204 | CMN_02136 | <0.00010 | + | − | 10.0 | Virulence |
| CCE76590.1 | Sugar ABC transporter | 48 | CMN_02658 | <0.00010 | + | − | 7.5 | Transport protein |
| CCE75600.1 | Endo-1,4-beta-xylanase | 71 | xysB | <0.00010 | + | − | 6.0 | Pathogenicity |
| CCE75697.1 | FKBP-type peptidyl-prolyl cis-trans isomerase | 13 | CMN_01752 | <0.00010 | + | − | 8.0 | Protein folding |
| CCE76397.1 | Anti-sigma factor | 29 | CMN_02462 | 0.0017 | + | − | 1.8 | Stress response |
| CCE75288.1 | Membrane-associated serine peptidase | 40 | CMN_01337 | 0.0057 | + | − | 1.5 | Protein degradation |
| CCE74709.1 | Putative extracellular nuclease | 74 | CMN_00751 | <0.00010 | − | + | 8.4 | Pathogenicity |
| CCE75697.1 | FKBP-type peptidyl-prolyl cis-trans isomerase | 13 | CMN_01752 | 0.001 | − | + | 4.0 | Protein folding |
| CCE76165.1 | Ferritin-like domain-containing protein | 32 | CMN_02227 | <0.00010 | − | + | 3.5 | Element acquisition |
| CCE74340.1 | Levan fructotransferase | 57 | CMN_00371 | <0.00010 | − | + | 2.7 | Carb. metabolism |
| CCE75152.1 | Agglutinin receptor precursor | 48 | CMN_01200 | <0.00010 | − | + | 2.5 | Signal transduction |
| CCE76077.1 | Putative RTX toxin | 204 | CMN_02136 | <0.00010 | − | + | 2.0 | Pathogenicity |
| CCE76397.1 | Anti-sigma factor | 29 | CMN_02462 | <0.00010 | − | + | 1.3 | Stress response |
| CCE74757.1 | NPL/P60 family endopeptidase | 45 | CMN_00800 | <0.00010 | − | + | 1.3 | Protein degradation |
| AJW79074.1 | Superoxide dismutase | 23 | SOD | 0.027 | − | + | 5.2 | Stress response |
| CCE76077.1 | Putative RTX toxin | 204 | CMN_02136 | <0.00010 | + | − | 10 | Virulence |
| CCE75795.1 | Secreted LysM peptidoglycan-binding protein | 43 | CMN_01850 | <0.00010 | + | − | 2.4 | Virulence |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Soliman, A.; Rampitsch, C.; Tambong, J.T.; Daayf, F. Secretome Analysis of Clavibacter nebraskensis Strains Treated with Natural Xylem Sap In Vitro Predicts Involvement of Glycosyl Hydrolases and Proteases in Bacterial Aggressiveness. Proteomes 2021, 9, 1. https://doi.org/10.3390/proteomes9010001
Soliman A, Rampitsch C, Tambong JT, Daayf F. Secretome Analysis of Clavibacter nebraskensis Strains Treated with Natural Xylem Sap In Vitro Predicts Involvement of Glycosyl Hydrolases and Proteases in Bacterial Aggressiveness. Proteomes. 2021; 9(1):1. https://doi.org/10.3390/proteomes9010001
Chicago/Turabian StyleSoliman, Atta, Christof Rampitsch, James T. Tambong, and Fouad Daayf. 2021. "Secretome Analysis of Clavibacter nebraskensis Strains Treated with Natural Xylem Sap In Vitro Predicts Involvement of Glycosyl Hydrolases and Proteases in Bacterial Aggressiveness" Proteomes 9, no. 1: 1. https://doi.org/10.3390/proteomes9010001






