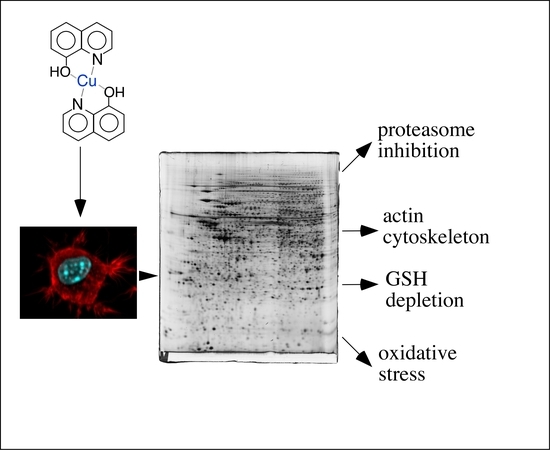
A Proteomic View of Cellular Responses to Anticancer Quinoline-Copper Complexes
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Ion Quantification by ICP-MS
2.3. Proteomics
2.3.1. Sample Preparation
2.3.2. Isoelectric Focusing
2.3.3. SDS Electrophoresis and Protein Detection
2.3.4. Image Analysis
2.3.5. Mass Spectrometry
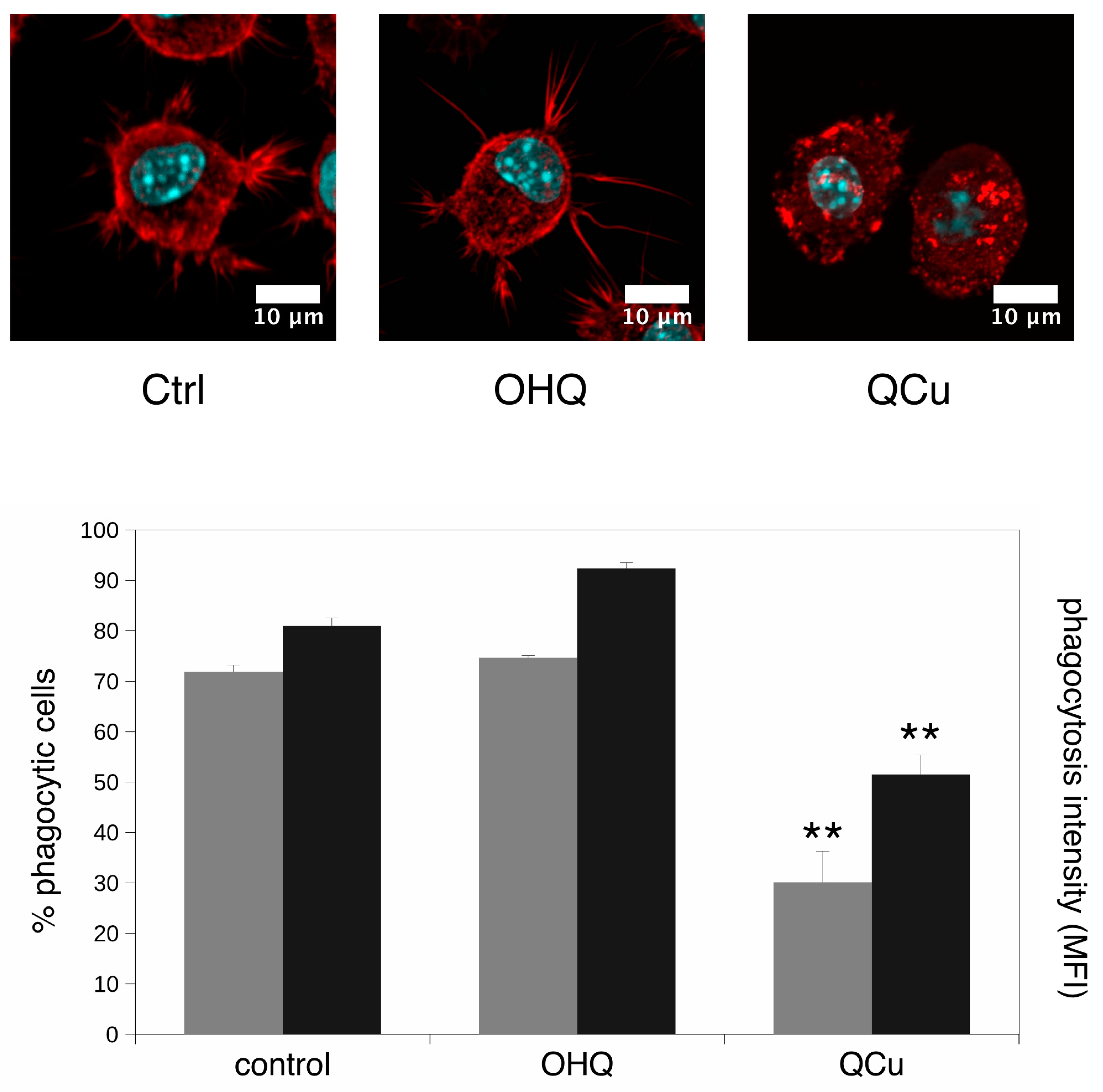
2.4. Phagocytosis and Particle Internalization Assay
2.5. F-Actin Staining
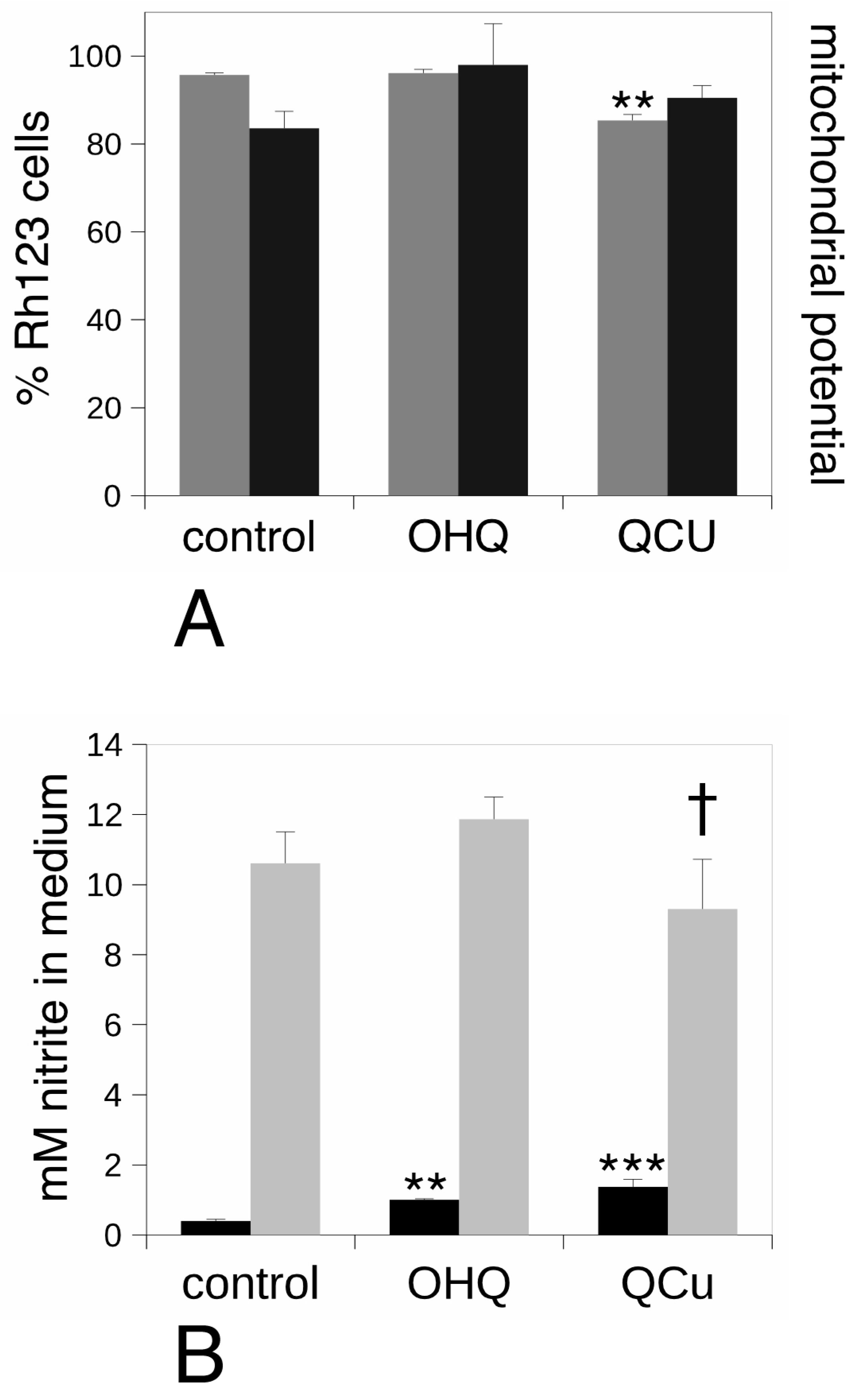
2.6. Mitochondrial Transmembrane Potential Measurement
2.7. Glutathione Assay
2.8. NO Production
3. Results
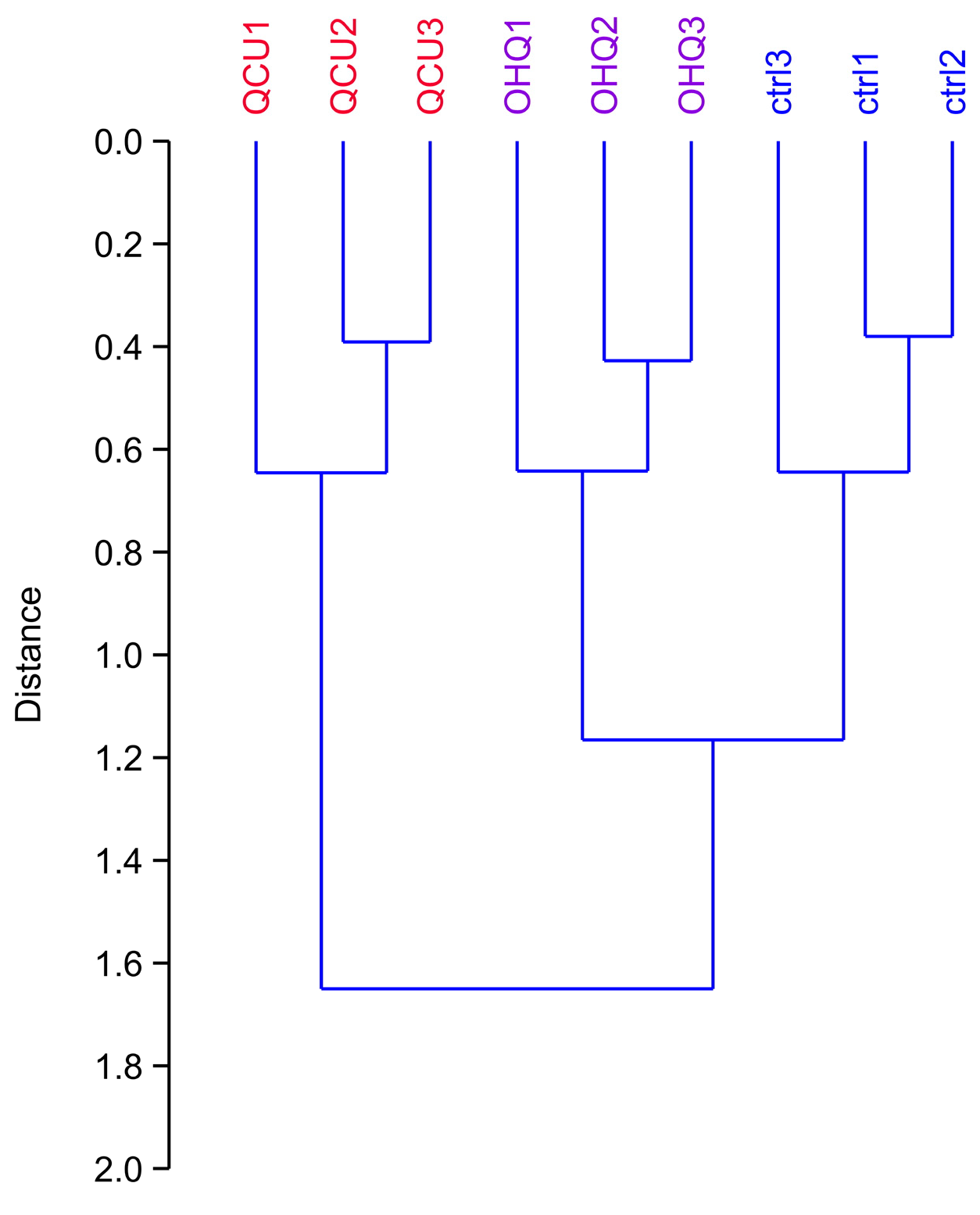
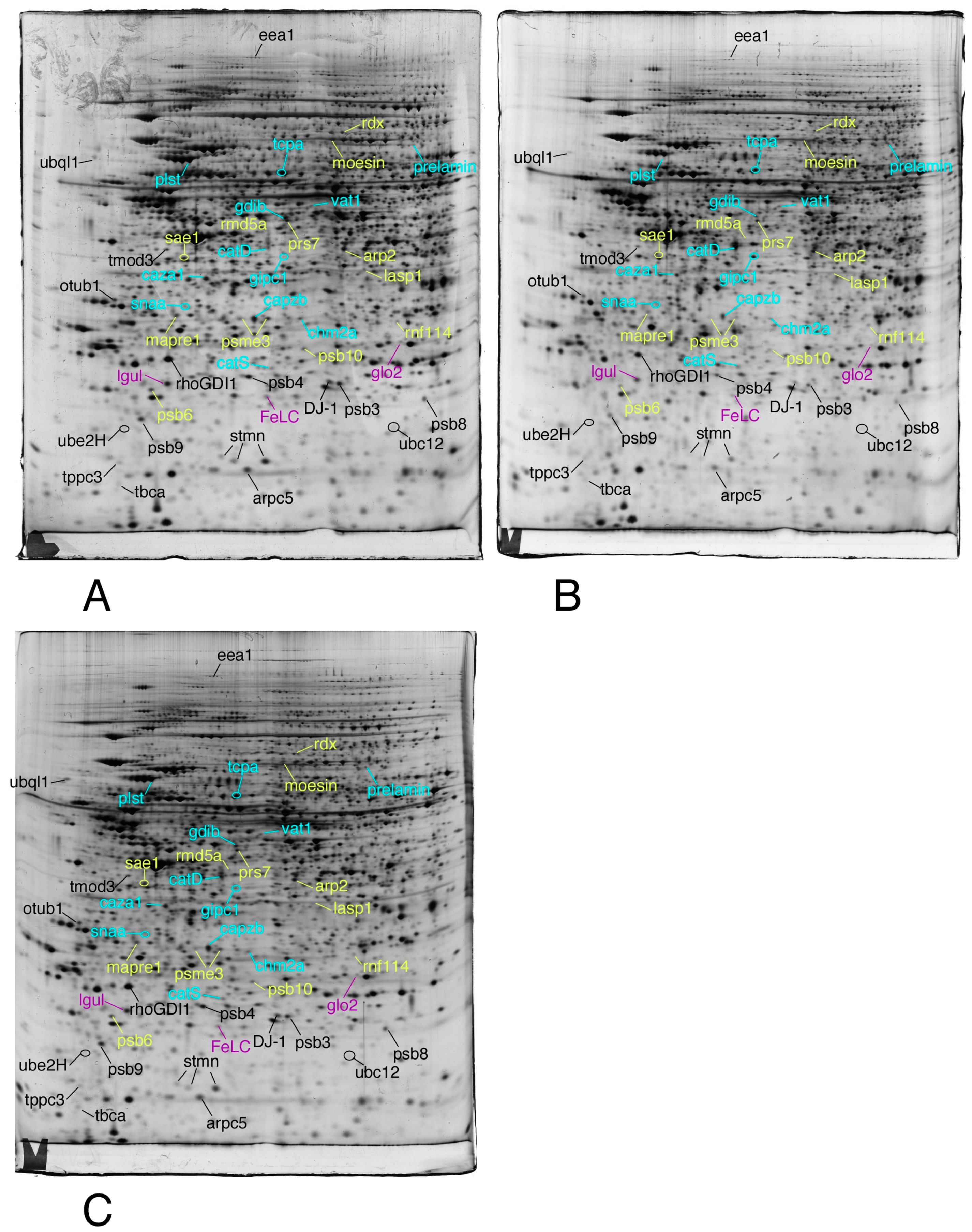
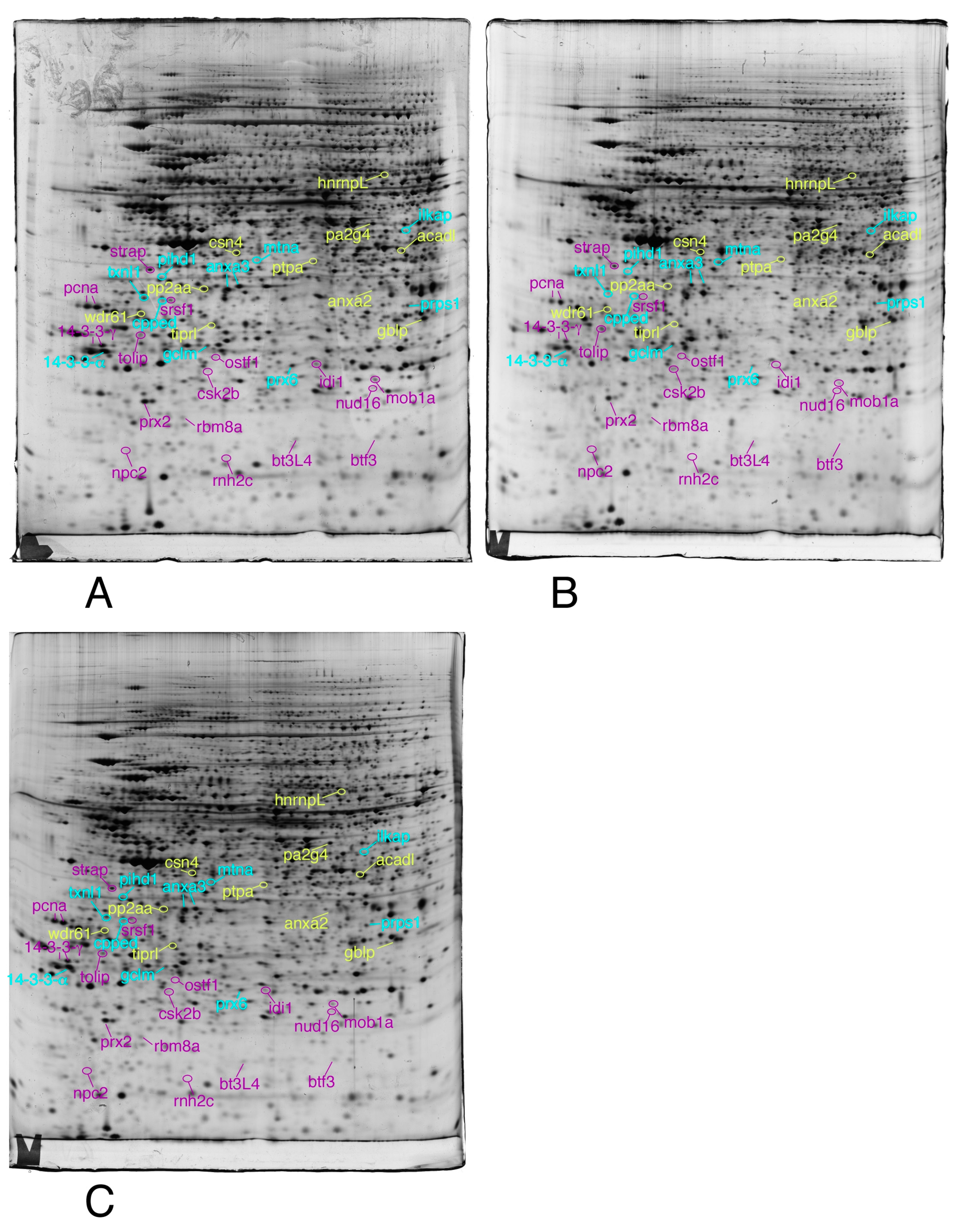
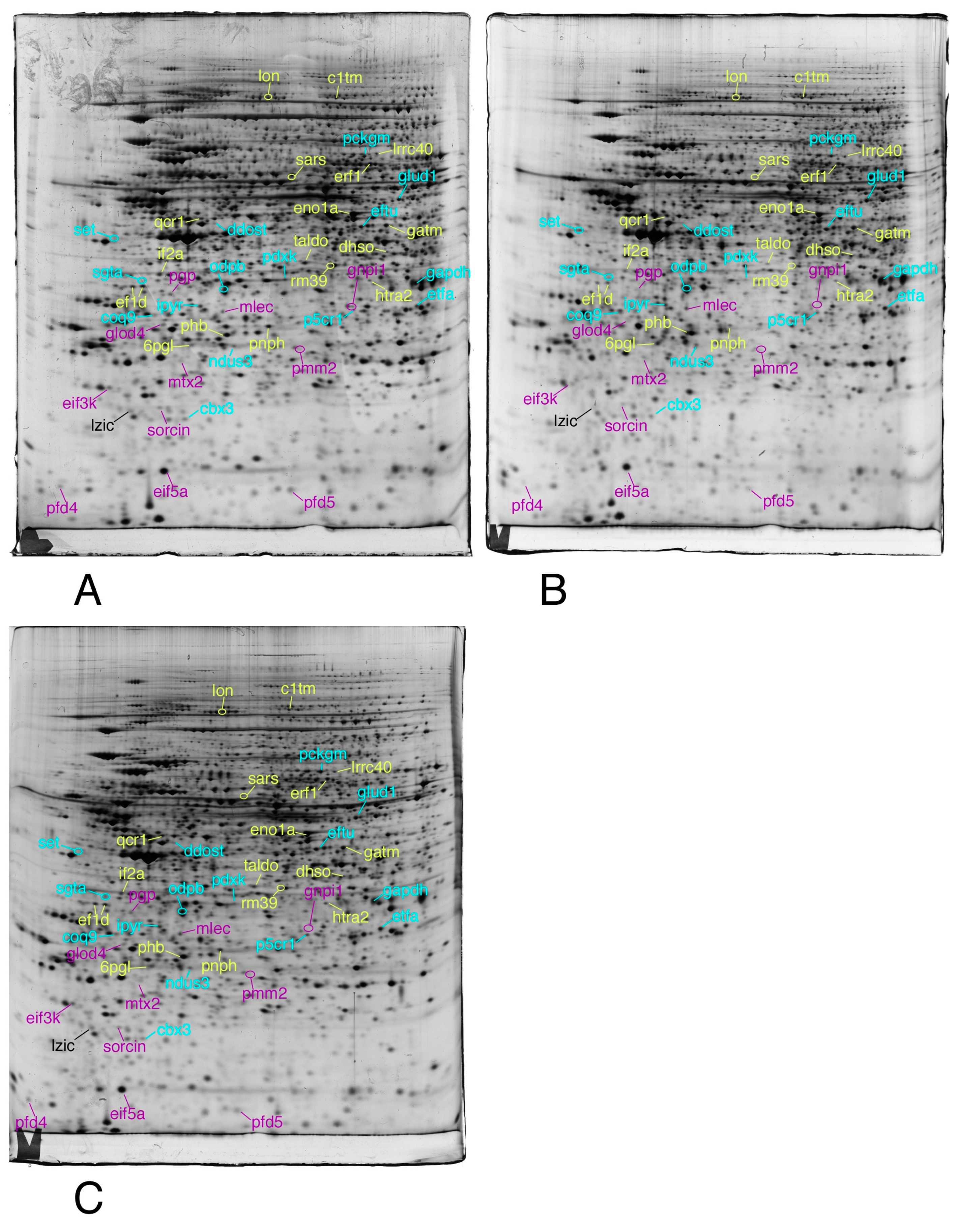
3.1. Global Analysis of the Proteomic Results
3.2. Targeted Validation Experiments
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Rosenberg, B. Platinum coordination complexes in cancer chemotherapy. Naturwissenschaften 1973, 60, 399–406. [Google Scholar] [CrossRef]
- Ho, Y.P.; Au-Yeung, S.C.; To, K.K. Platinum-based anticancer agents: Innovative design strategies and biological perspectives. Med. Res. Rev. 2003, 23, 633–655. [Google Scholar] [CrossRef]
- Dilruba, S.; Kalayda, G.V. Platinum-based drugs: Past, present and future. Cancer Chemother Pharm. 2016, 77, 1103–1124. [Google Scholar] [CrossRef]
- Milacic, V.; Chen, D.; Ronconi, L.; Landis-Piwowar, K.R.; Fregona, D.; Dou, Q.P. A novel anticancer gold(III) dithiocarbamate compound inhibits the activity of a purified 20S proteasome and 26S proteasome in human breast cancer cell cultures and xenografts. Cancer Res. 2006, 66, 10478–10486. [Google Scholar] [CrossRef]
- Arsenijevic, M.; Milovanovic, M.; Volarevic, V.; Djekovic, A.; Kanjevac, T.; Arsenijevic, N.; Dukic, S.; Bugarcic, Z.D. Cytotoxicity of gold(III) complexes on A549 human lung carcinoma epithelial cell line. Med. Chem. 2012, 8, 2–8. [Google Scholar] [CrossRef]
- Daniel, K.G.; Gupta, P.; Harbach, R.H.; Guida, W.C.; Dou, Q.P. Organic copper complexes as a new class of proteasome inhibitors and apoptosis inducers in human cancer cells. Biochem. Pharmacol. 2004, 67, 1139–1151. [Google Scholar] [CrossRef]
- Chen, D.; Dou, Q.P. New uses for old copper-binding drugs: Converting the pro-angiogenic copper to a specific cancer cell death inducer. Expert Opin. Ther. Targets 2008, 12, 739–748. [Google Scholar] [CrossRef]
- Brar, S.S.; Grigg, C.; Wilson, K.S.; Holder, W.D., Jr.; Dreau, D.; Austin, C.; Foster, M.; Ghio, A.J.; Whorton, A.R.; Stowell, G.W.; et al. Disulfiram inhibits activating transcription factor/cyclic AMP-responsive element binding protein and human melanoma growth in a metal-dependent manner in vitro, in mice and in a patient with metastatic disease. Mol. Cancer 2004, 3, 1049–1060. [Google Scholar]
- Sava, G.; Pacor, S.; Mestroni, G.; Alessio, E. Na[trans-RuCl4(DMSO)Im], a metal complex of ruthenium with antimetastatic properties. Clin. Exp. Metastasis 1992, 10, 273–280. [Google Scholar] [CrossRef]
- Hartinger, C.G.; Zorbas-Seifried, S.; Jakupec, M.A.; Kynast, B.; Zorbas, H.; Keppler, B.K. From bench to bedside—Preclinical and early clinical development of the anticancer agent indazolium trans-[tetrachlorobis(1H-indazole)ruthenate(III)] (KP1019 or FFC14A). J. Inorg. Biochem. 2006, 100, 891–904. [Google Scholar] [CrossRef]
- Gianferrara, T.; Bratsos, I.; Iengo, E.; Milani, B.; Ostric, A.; Spagnul, C.; Zangrando, E.; Alessio, E. Synthetic strategies towards ruthenium-porphyrin conjugates for anticancer activity. Dalton Trans 2009, 10742–10756. [Google Scholar] [CrossRef]
- Schmitt, S.M.; Frezza, M.; Dou, Q.P. New applications of old metal-binding drugs in the treatment of human cancer. Front Biosci. (Sch. Ed) 2012, 4, 375–391. [Google Scholar] [CrossRef]
- Merlino, A. Interactions between proteins and Ru compounds of medicinal interest: A structural perspective. Coord. Chem. Rev. 2016, 326, 111–134. [Google Scholar] [CrossRef]
- Ronconi, L.; Marzano, C.; Zanello, P.; Corsini, M.; Miolo, G.; Macca, C.; Trevisan, A.; Fregona, D. Gold(III) dithiocarbamate derivatives for the treatment of cancer: Solution chemistry, DNA binding, and hemolytic properties. J. Med. Chem. 2006, 49, 1648–1657. [Google Scholar] [CrossRef]
- Chiara, F.; Gambalunga, A.; Sciacovelli, M.; Nicolli, A.; Ronconi, L.; Fregona, D.; Bernardi, P.; Rasola, A.; Trevisan, A. Chemotherapeutic induction of mitochondrial oxidative stress activates GSK-3alpha/beta and Bax, leading to permeability transition pore opening and tumor cell death. Cell Death Dis. 2012, 3, e444. [Google Scholar] [CrossRef]
- Manasanch, E.E.; Orlowski, R.Z. Proteasome inhibitors in cancer therapy. Nat. Rev. Clin. Oncol. 2017, 14, 417–433. [Google Scholar] [CrossRef]
- Kim, I.; Kim, C.H.; Kim, J.H.; Lee, J.; Choi, J.J.; Chen, Z.A.; Lee, M.G.; Chung, K.C.; Hsu, C.Y.; Ahn, Y.S. Pyrrolidine dithiocarbamate and zinc inhibit proteasome-dependent proteolysis. Exp. Cell Res. 2004, 298, 229–238. [Google Scholar] [CrossRef]
- Xiao, Y.; Chen, D.; Zhang, X.; Cui, Q.Z.; Fan, Y.H.; Bi, C.F.; Dou, Q.P. Molecular study on copper-mediated tumor proteasome inhibition and cell death. Int. J. Oncol. 2010, 37, 81–87. [Google Scholar]
- Verani, C.N. Metal complexes as inhibitors of the 26S proteasome in tumor cells. J. Inorg. Biochem. 2012, 106, 59–67. [Google Scholar] [CrossRef]
- Daniel, K.G.; Chen, D.; Orlu, S.; Cui, Q.C.; Miller, F.R.; Dou, Q.P. Clioquinol and pyrrolidine dithiocarbamate complex with copper to form proteasome inhibitors and apoptosis inducers in human breast cancer cells. Breast Cancer Res. 2005, 7, R897–R908. [Google Scholar] [CrossRef]
- Chen, D.; Cui, Q.C.; Yang, H.J.; Barrea, R.A.; Sarkar, F.H.; Sheng, S.J.; Yan, B.; Reddy, G.P.V.; Dou, Q.P. Clioquinol, a therapeutic agent for Alzheimer’s disease, has proteasome-inhibitory, androgen receptor-suppressing, apoptosis-inducing, and antitumor activities in human prostate cancer cells and xenografts. Cancer Res. 2007, 67, 1636–1644. [Google Scholar] [CrossRef]
- Daniel, K.G.; Chen, D.; Yan, B.; Dou, Q.P. Copper-binding compounds as proteasome inhibitors and apoptosis inducers in human cancer. Front. Biosci. -Landmark 2007, 12, 135–144. [Google Scholar]
- Milacic, V.; Jiao, P.F.; Zhang, B.; Yan, B.; Dou, Q.P. Novel 8-hydroxylquinoline analogs induce copper-dependent proteasome inhibition and cell death in human breast cancer cells. Int. J. Oncol. 2009, 35, 1481–1491. [Google Scholar]
- Zhai, S.M.; Yang, L.; Cui, Q.C.; Sun, Y.; Dou, Q.P.; Yan, B. Tumor cellular proteasome inhibition and growth suppression by 8-hydroxyquinoline and clioquinol requires their capabilities to bind copper and transport copper into cells. J. Biol. Inorg. Chem. 2010, 15, 259–269. [Google Scholar] [CrossRef]
- Schimmer, A.D. Clioquinol—A Novel Copper-Dependent and Independent Proteasome Inhibitor. Curr. Cancer Drug Targets 2011, 11, 325–331. [Google Scholar] [CrossRef]
- Oliveri, V.; Lanza, V.; Milardi, D.; Viale, M.; Maric, I.; Sgarlata, C.; Vecchio, G. Amino- and chloro-8-hydroxyquinolines and their copper complexes as proteasome inhibitors and antiproliferative agents. Metallomics 2017, 9, 1439–1446. [Google Scholar] [CrossRef]
- Zhang, Z.; Wang, H.Y.; Yan, M.C.; Wang, H.N.; Zhang, C.Y. Novel copper complexes as potential proteasome inhibitors for cancer treatment. Mol. Med. Rep. 2017, 15, 3–11. [Google Scholar] [CrossRef]
- Ikotun, O.F.; Higbee, E.M.; Ouellette, W.; Doyle, R.P. Pyrophosphate-bridged complexes with picomolar toxicity. J. Inorg. Biochem. 2009, 103, 1254–1264. [Google Scholar] [CrossRef]
- Chen, H.L.; Chang, C.Y.; Lee, H.T.; Lin, H.H.; Lu, P.J.; Yang, C.N.; Shiau, C.W.; Shaw, A.Y. Synthesis and pharmacological exploitation of clioquinol-derived copper-binding apoptosis inducers triggering reactive oxygen species generation and MAPK pathway activation. Bioorg. Med. Chem. 2009, 17, 7239–7247. [Google Scholar] [CrossRef]
- Jungwirth, U.; Kowol, C.R.; Keppler, B.K.; Hartinger, C.G.; Berger, W.; Heffeter, P. Anticancer Activity of Metal Complexes: Involvement of Redox Processes. Antioxid. Redox Signal. 2011, 15, 1085–1127. [Google Scholar] [CrossRef]
- Allensworth, J.L.; Evans, M.K.; Bertucci, F.; Aldrich, A.J.; Festa, R.A.; Finetti, P.; Ueno, N.T.; Safi, R.; McDonnell, D.P.; Thiele, D.J.; et al. Disulfiram (DSF) acts as a copper ionophore to induce copper-dependent oxidative stress and mediate anti-tumor efficacy in inflammatory breast cancer. Mol. Oncol. 2015, 9, 1155–1168. [Google Scholar] [CrossRef]
- Wright, C.; Moore, R.D. Disulfiram treatment of alcoholism. Am. J. Med. 1990, 88, 647–655. [Google Scholar] [CrossRef]
- Gholz, L.M.; Arons, W.L. Prophylaxis and Therapy of Amebiasis and Shigellosis with Iodochlorhydroxyquin. Am. J. Trop Med. Hyg. 1964, 13, 396–401. [Google Scholar] [CrossRef]
- Rohde, W.; Mikelens, P.; Jackson, J.; Blackman, J.; Whitcher, J.; Levinson, W. Hydroxyquinolines inhibit ribonucleic acid-dependent deoxyribonucleic acid polymerase and inactivate Rous sarcoma virus and herpes simplex virus. Antimicrob Agents Chemother 1976, 10, 234–240. [Google Scholar] [CrossRef]
- Jaouen, G.R.; Vessières, A.; Top, S. Ferrocifen type anti cancer drugs. Chem. Soc. Rev. 2015, 44, 8802–8817. [Google Scholar] [CrossRef]
- Patra, M.; Gasser, G. The medicinal chemistry of ferrocene and its derivatives. Nat. Rev. Chem. 2017, 1, 0066. [Google Scholar] [CrossRef]
- Kowalski, K. Insight into the Biological Activity of Organometallic Acetylsalicylic Acid (Aspirin) Derivatives. ChemPlusChem 2019, 84, 403–415. [Google Scholar] [CrossRef]
- Liu, W.; Gust, R. Metal N-heterocyclic carbene complexes as potential antitumor metallodrugs. Chem. Soc. Rev. 2013, 42, 755–773. [Google Scholar] [CrossRef]
- Wehbe, M.; Malhotra, A.K.; Anantha, M.; Lo, C.; Dragowska, W.H.; Dos Santos, N.; Bally, M.B. Development of a copper-clioquinol formulation suitable for intravenous use. Drug Deliv. Transl. Res. 2018, 8, 239–251. [Google Scholar] [CrossRef]
- Weniger, M.A.; Rizzatti, E.G.; Perez-Galan, P.; Liu, D.; Wang, Q.; Munson, P.J.; Raghavachari, N.; White, T.; Tweito, M.M.; Dunleavy, K.; et al. Treatment-induced oxidative stress and cellular antioxidant capacity determine response to bortezomib in mantle cell lymphoma. Clin Cancer Res 2011, 17, 5101–5112. [Google Scholar] [CrossRef]
- Zanellato, I.; Heldt, J.-M.; Vessières, A.; Jaouen, G.R.; Osella, D. Antiproliferative effect of ferrocifen drug candidates on malignant pleural mesothelioma cell lines. Inorg. Chim. Acta 2009, 362, 4037–4042. [Google Scholar] [CrossRef]
- Rabilloud, T.; Lescuyer, P. Proteomics in mechanistic toxicology: History, concepts, achievements, caveats, and potential. Proteomics 2015, 15, 1051–1074. [Google Scholar] [CrossRef] [PubMed]
- Guidi, F.; Modesti, A.; Landini, I.; Nobili, S.; Mini, E.; Bini, L.; Puglia, M.; Casini, A.; Dyson, P.J.; Gabbiani, C.; et al. The molecular mechanisms of antimetastatic ruthenium compounds explored through DIGE proteomics. J. Inorg. Biochem. 2013, 118, 94–99. [Google Scholar] [CrossRef] [PubMed]
- Gamberi, T.; Massai, L.; Magherini, F.; Landini, I.; Fiaschi, T.; Scaletti, F.; Gabbiani, C.; Bianchi, L.; Bini, L.; Nobili, S.; et al. Proteomic analysis of A2780/S ovarian cancer cell response to the cytotoxic organogold(III) compound Aubipy(c). J. Proteom. 2014, 103, 103–120. [Google Scholar] [CrossRef]
- Mao, X.; Li, X.; Sprangers, R.; Wang, X.; Venugopal, A.; Wood, T.; Zhang, Y.; Kuntz, D.A.; Coe, E.; Trudel, S.; et al. Clioquinol inhibits the proteasome and displays preclinical activity in leukemia and myeloma. Leukemia 2009, 23, 585–590. [Google Scholar] [CrossRef] [PubMed]
- Prachayasittikul, V.; Prachayasittikul, S.; Ruchirawat, S.; Prachayasittikul, V. 8-Hydroxyquinolines: A review of their metal chelating properties and medicinal applications. Drug Des. Dev. Ther. 2013, 7, 1157–1178. [Google Scholar] [CrossRef]
- Tardito, S.; Barilli, A.; Bassanetti, I.; Tegoni, M.; Bussolati, O.; Franchi-Gazzola, R.; Mucchino, C.; Marchio, L. Copper-Dependent Cytotoxicity of 8-Hydroxyquinoline Derivatives Correlates with Their Hydrophobicity and Does Not Require Caspase Activation. J. Med. Chem. 2012, 55, 10448–10459. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Luo, C.; Shan, C.; You, Q.; Lu, J.; Elf, S.; Zhou, Y.; Wen, Y.; Vinkenborg, J.L.; Fan, J.; et al. Inhibition of human copper trafficking by a small molecule significantly attenuates cancer cell proliferation. Nat. Chem. 2015, 7, 968. [Google Scholar] [CrossRef] [PubMed]
- Barresi, V.; Spampinato, G.; Musso, N.; Trovato Salinaro, A.; Rizzarelli, E.; Condorelli, D.F. ATOX1 gene silencing increases susceptibility to anticancer therapy based on copper ionophores or chelating drugs. J. Inorg. Biochem. 2016, 156, 145–152. [Google Scholar] [CrossRef]
- Repetto, G.; del Peso, A.; Zurita, J.L. Neutral red uptake assay for the estimation of cell viability/cytotoxicity. Nat. Protoc. 2008, 3, 1125–1131. [Google Scholar] [CrossRef]
- Rabilloud, T. Optimization of the cydex blue assay: A one-step colorimetric protein assay using cyclodextrins and compatible with detergents and reducers. PLoS ONE 2018, 13, e0195755. [Google Scholar] [CrossRef] [PubMed]
- Dalzon, B.; Aude-Garcia, C.; Collin-Faure, V.; Diemer, H.; Beal, D.; Dussert, F.; Fenel, D.; Schoehn, G.; Cianferani, S.; Carriere, M.; et al. Differential proteomics highlights macrophage-specific responses to amorphous silica nanoparticles. Nanoscale 2017, 9, 9641–9658. [Google Scholar] [CrossRef] [PubMed]
- Gianazza, E.; Celentano, F.; Magenes, S.; Ettori, C.; Righetti, P.G. Formulations for immobilized pH gradients including pH extremes. Electrophoresis 1989, 10, 806–808. [Google Scholar] [CrossRef] [PubMed]
- Rabilloud, T.; Valette, C.; Lawrence, J.J. Sample application by in-gel rehydration improves the resolution of two-dimensional electrophoresis with immobilized pH gradients in the first dimension. Electrophoresis 1994, 15, 1552–1558. [Google Scholar] [CrossRef] [PubMed]
- Luche, S.; Diemer, H.; Tastet, C.; Chevallet, M.; Van Dorsselaer, A.; Leize-Wagner, E.; Rabilloud, T. About thiol derivatization and resolution of basic proteins in two-dimensional electrophoresis. Proteomics 2004, 4, 551–561. [Google Scholar] [CrossRef] [PubMed]
- Gorg, A.; Postel, W.; Weser, J.; Gunther, S.; Strahler, J.R.; Hanash, S.M.; Somerlot, L. Elimination of Point Streaking on Silver Stained Two-Dimensional Gels by Addition of Iodoacetamide to the Equilibration Buffer. Electrophoresis 1987, 8, 122–124. [Google Scholar] [CrossRef]
- Tastet, C.; Lescuyer, P.; Diemer, H.; Luche, S.; van Dorsselaer, A.; Rabilloud, T. A versatile electrophoresis system for the analysis of high- and low-molecular-weight proteins. Electrophoresis 2003, 24, 1787–1794. [Google Scholar] [CrossRef] [PubMed]
- Sinha, P.; Poland, J.; Schnolzer, M.; Rabilloud, T. A new silver staining apparatus and procedure for matrix-assisted laser desorption/ionization-time of flight analysis of proteins after two-dimensional electrophoresis. Proteomics 2001, 1, 835–840. [Google Scholar] [CrossRef]
- Yekutieli, D.; Benjamini, Y. Resampling-based false discovery rate controlling multiple test procedures for correlated test statistics. J. Stat. Plan. Inference 1999, 82, 171–196. [Google Scholar] [CrossRef]
- Diz, A.P.; Carvajal-Rodriguez, A.; Skibinski, D.O. Multiple hypothesis testing in proteomics: A strategy for experimental work. Mol. Cell Proteom. 2011, 10, M110 004374. [Google Scholar]
- Carvajal-Rodriguez, A.; de Una-Alvarez, J. Assessing significance in high-throughput experiments by sequential goodness of fit and q-value estimation. PLoS ONE 2011, 6, e24700. [Google Scholar] [CrossRef]
- Hammer, O.; Harper, D.A.T.; Ryan, P.D. Paleontological statistics software package for education and data analysis. Palaeontol. Electron. 2001, 4, 9. [Google Scholar]
- Gharahdaghi, F.; Weinberg, C.R.; Meagher, D.A.; Imai, B.S.; Mische, S.M. Mass spectrometric identification of proteins from silver-stained polyacrylamide gel: A method for the removal of silver ions to enhance sensitivity. Electrophoresis 1999, 20, 601–605. [Google Scholar] [CrossRef]
- Zhan, X.; Yang, H.; Peng, F.; Li, J.; Mu, Y.; Long, Y.; Cheng, T.; Huang, Y.; Li, Z.; Lu, M.; et al. How many proteins can be identified in a 2DE gel spot within an analysis of a complex human cancer tissue proteome? Electrophoresis 2018, 39, 965–980. [Google Scholar] [CrossRef] [PubMed]
- Triboulet, S.; Aude-Garcia, C.; Carriere, M.; Diemer, H.; Proamer, F.; Habert, A.; Chevallet, M.; Collin-Faure, V.; Strub, J.M.; Hanau, D.; et al. Molecular responses of mouse macrophages to copper and copper oxide nanoparticles inferred from proteomic analyses. Mol. Cell Proteom. 2013, 12, 3108–3122. [Google Scholar] [CrossRef] [PubMed]
- Aude-Garcia, C.; Dalzon, B.; Ravanat, J.L.; Collin-Faure, V.; Diemer, H.; Strub, J.M.; Cianferani, S.; Van Dorsselaer, A.; Carriere, M.; Rabilloud, T. A combined proteomic and targeted analysis unravels new toxic mechanisms for zinc oxide nanoparticles in macrophages. J. Proteom. 2016, 134, 174–185. [Google Scholar] [CrossRef] [PubMed]
- Perry, S.W.; Norman, J.P.; Barbieri, J.; Brown, E.B.; Gelbard, H.A. Mitochondrial membrane potential probes and the proton gradient: A practical usage guide. Biotechniques 2011, 50, 98–115. [Google Scholar] [CrossRef] [PubMed]
- Warholm, M.; Guthenberg, C.; von Bahr, C.; Mannervik, B. Glutathione transferases from human liver. Methods Enzym. 1985, 113, 499–504. [Google Scholar]
- Arrick, B.A.; Nathan, C.F.; Griffith, O.W.; Cohn, Z.A. Glutathione depletion sensitizes tumor cells to oxidative cytolysis. J. Biol. Chem. 1982, 257, 1231–1237. [Google Scholar] [PubMed]
- Bruggeman, I.M.; Spenkelink, A.; Temmink, J.H.; van Bladeren, P.J. Differential effects of raising and lowering intracellular glutathione levels on the cytotoxicity of allyl isothiocyanate, tert-butylhydroperoxide and chlorodinitrobenzene. Toxicol In Vitro 1988, 2, 31–35. [Google Scholar] [CrossRef]
- Ding, W.Q.; Liu, B.L.; Vaught, J.L.; Yamauchi, H.; Lind, S.E. Anticancer activity of the antibiotic clioquinol. Cancer Res. 2005, 65, 3389–3395. [Google Scholar] [CrossRef]
- Yu, H.J.; Zhou, Y.F.; Lind, S.E.; Ding, W.Q. Clioquinol targets zinc to lysosomes in human cancer cells. Biochem. J. 2009, 417, 133–139. [Google Scholar] [CrossRef]
- Ding, W.Q.; Lind, S.E. Metal Ionophores—An Emerging Class of Anticancer Drugs. Iubmb Life 2009, 61, 1013–1018. [Google Scholar] [CrossRef] [PubMed]
- Herrmann, A.G.; Searcy, J.L.; Le Bihan, T.; McCulloch, J.; Deighton, R.F. Total variance should drive data handling strategies in third generation proteomic studies. Proteomics 2013, 13, 3251–3255. [Google Scholar] [CrossRef] [PubMed]
- Campostrini, N.; Areces, L.B.; Rappsilber, J.; Pietrogrande, M.C.; Dondi, F.; Pastorino, F.; Ponzoni, M.; Righetti, P.G. Spot overlapping in two-dimensional maps: A serious problem ignored for much too long. Proteomics 2005, 5, 2385–2395. [Google Scholar] [CrossRef] [PubMed]
- Hunsucker, S.W.; Duncan, M.W. Is protein overlap in two-dimensional gels a serious practical problem? Proteomics 2006, 6, 1374–1375. [Google Scholar] [CrossRef] [PubMed]
- Colignon, B.; Raes, M.; Dieu, M.; Delaive, E.; Mauro, S. Evaluation of three-dimensional gel electrophoresis to improve quantitative profiling of complex proteomes. Proteomics 2013, 13, 2077–2082. [Google Scholar] [CrossRef] [PubMed]
- Rabilloud, T. When 2D is not enough, go for an extra dimension. Proteomics 2013, 13, 2065–2068. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cao, B.Y.; Li, J.; Zhu, J.Y.; Shen, M.Y.; Han, K.K.; Zhang, Z.B.; Yu, Y.; Wang, Y.L.; Wu, D.P.; Chen, S.N.; et al. The Antiparasitic Clioquinol Induces Apoptosis in Leukemia and Myeloma Cells by Inhibiting Histone Deacetylase Activity. J. Biol. Chem. 2013, 288, 34181–34189. [Google Scholar] [CrossRef]
- Cao, B.Y.; Li, J.; Zhou, X.M.; Juan, J.X.; Han, K.K.; Zhang, Z.B.; Kong, Y.; Wang, J.R.; Mao, X.L. Clioquinol induces pro-death autophagy in leukemia and myeloma cells by disrupting the mTOR signaling pathway. Sci. Rep. 2014, 4. [Google Scholar] [CrossRef]
- He, M.L.; Luo, M.; Liu, Q.Y.; Chen, J.K.; Li, K.S.; Zheng, M.G.; Weng, Y.L.; Ouyang, L.P.; Liu, A.M. Combination treatment with fasudil and clioquinol produces synergistic anti-tumor effects in U87 glioblastoma cells by activating apoptosis and autophagy. J. Neuro-Oncol. 2016, 127, 261–270. [Google Scholar] [CrossRef]
- Du, T.; Filiz, G.; Caragounis, A.; Crouch, P.J.; White, A.R. Clioquinol promotes cancer cell toxicity through tumor necrosis factor alpha release from macrophages. J. Pharmacol. Exp. Ther. 2008, 324, 360–367. [Google Scholar] [CrossRef]
- Triboulet, S.; Aude-Garcia, C.; Armand, L.; Collin-Faure, V.; Chevallet, M.; Diemer, H.; Gerdil, A.; Proamer, F.; Strub, J.M.; Habert, A.; et al. Comparative proteomic analysis of the molecular responses of mouse macrophages to titanium dioxide and copper oxide nanoparticles unravels some toxic mechanisms for copper oxide nanoparticles in macrophages. PLoS ONE 2015, 10, e0124496. [Google Scholar] [CrossRef] [PubMed]
- Laws, K.; Suntharalingam, K. The Next Generation of Anticancer Metallopharmaceuticals: Cancer Stem Cell-Active Inorganics. Chembiochem 2018, 19, 2246–2253. [Google Scholar] [CrossRef] [PubMed]

| Condition | Control | OHQ | Qcu |
|---|---|---|---|
| Zn | 263 ± 81 | 257 ± 23 | 280 ± 79 |
| Cu | 16 ± 17 | 13 ± 10 | 1076 ± 258 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dalzon, B.; Bons, J.; Diemer, H.; Collin-Faure, V.; Marie-Desvergne, C.; Dubosson, M.; Cianferani, S.; Carapito, C.; Rabilloud, T. A Proteomic View of Cellular Responses to Anticancer Quinoline-Copper Complexes. Proteomes 2019, 7, 26. https://doi.org/10.3390/proteomes7020026
Dalzon B, Bons J, Diemer H, Collin-Faure V, Marie-Desvergne C, Dubosson M, Cianferani S, Carapito C, Rabilloud T. A Proteomic View of Cellular Responses to Anticancer Quinoline-Copper Complexes. Proteomes. 2019; 7(2):26. https://doi.org/10.3390/proteomes7020026
Chicago/Turabian StyleDalzon, Bastien, Joanna Bons, Hélène Diemer, Véronique Collin-Faure, Caroline Marie-Desvergne, Muriel Dubosson, Sarah Cianferani, Christine Carapito, and Thierry Rabilloud. 2019. "A Proteomic View of Cellular Responses to Anticancer Quinoline-Copper Complexes" Proteomes 7, no. 2: 26. https://doi.org/10.3390/proteomes7020026
APA StyleDalzon, B., Bons, J., Diemer, H., Collin-Faure, V., Marie-Desvergne, C., Dubosson, M., Cianferani, S., Carapito, C., & Rabilloud, T. (2019). A Proteomic View of Cellular Responses to Anticancer Quinoline-Copper Complexes. Proteomes, 7(2), 26. https://doi.org/10.3390/proteomes7020026