Genome-Wide Identification and Expression Analysis of the Strawberry FvbZIP Gene Family and the Role of Key Gene FabZIP46 in Fruit Resistance to Gray Mold
Abstract
:1. Introduction
2. Results and Analysis
2.1. Members of the bZIP Transcription Factor Family in Strawberries
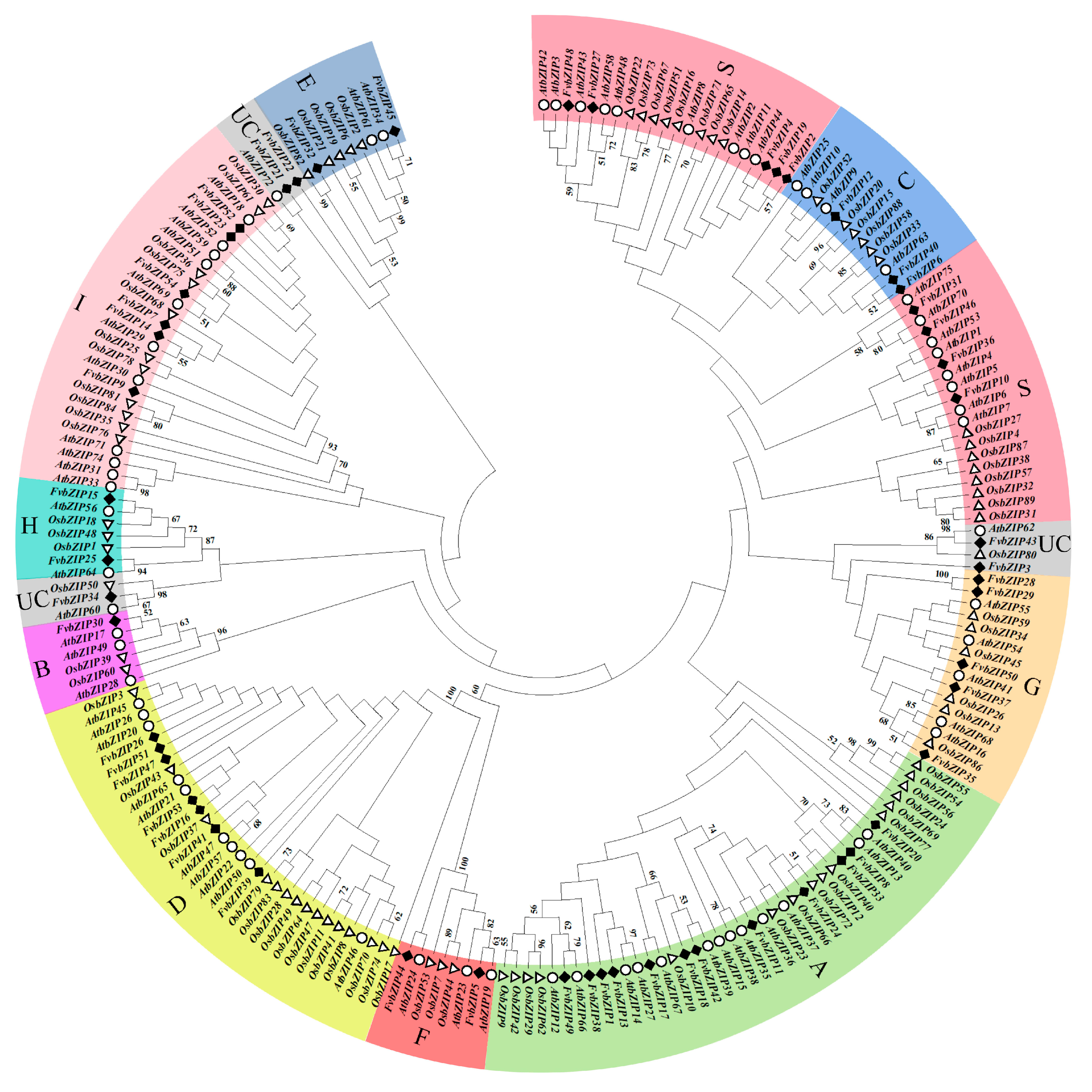
2.2. Phylogenetic Relationships among the Family Members of bZIP Transcription Factors in Strawberries
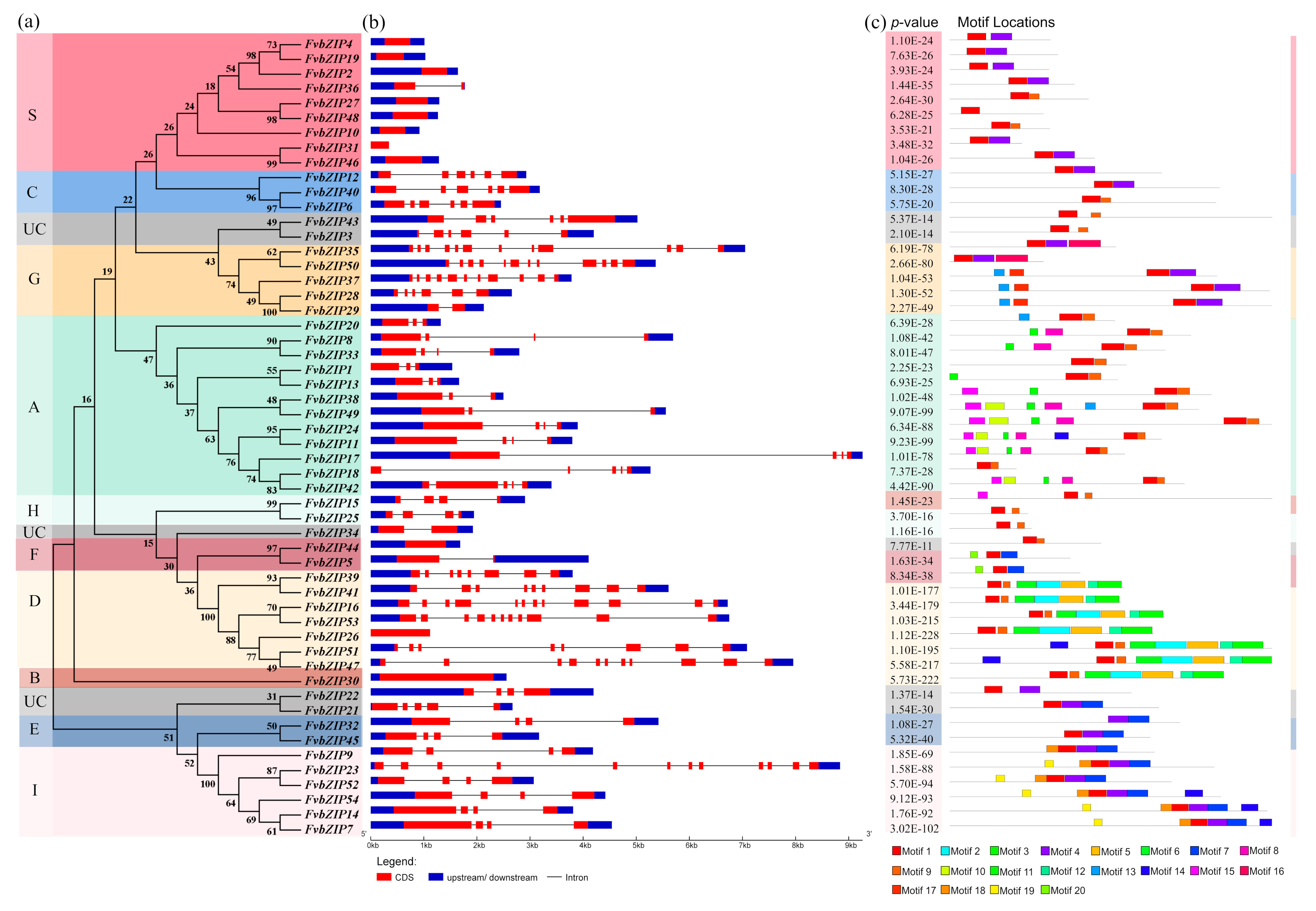
2.3. Chromosomal Localization and Analysis of the Gene Structure of Strawberry bZIP Genes
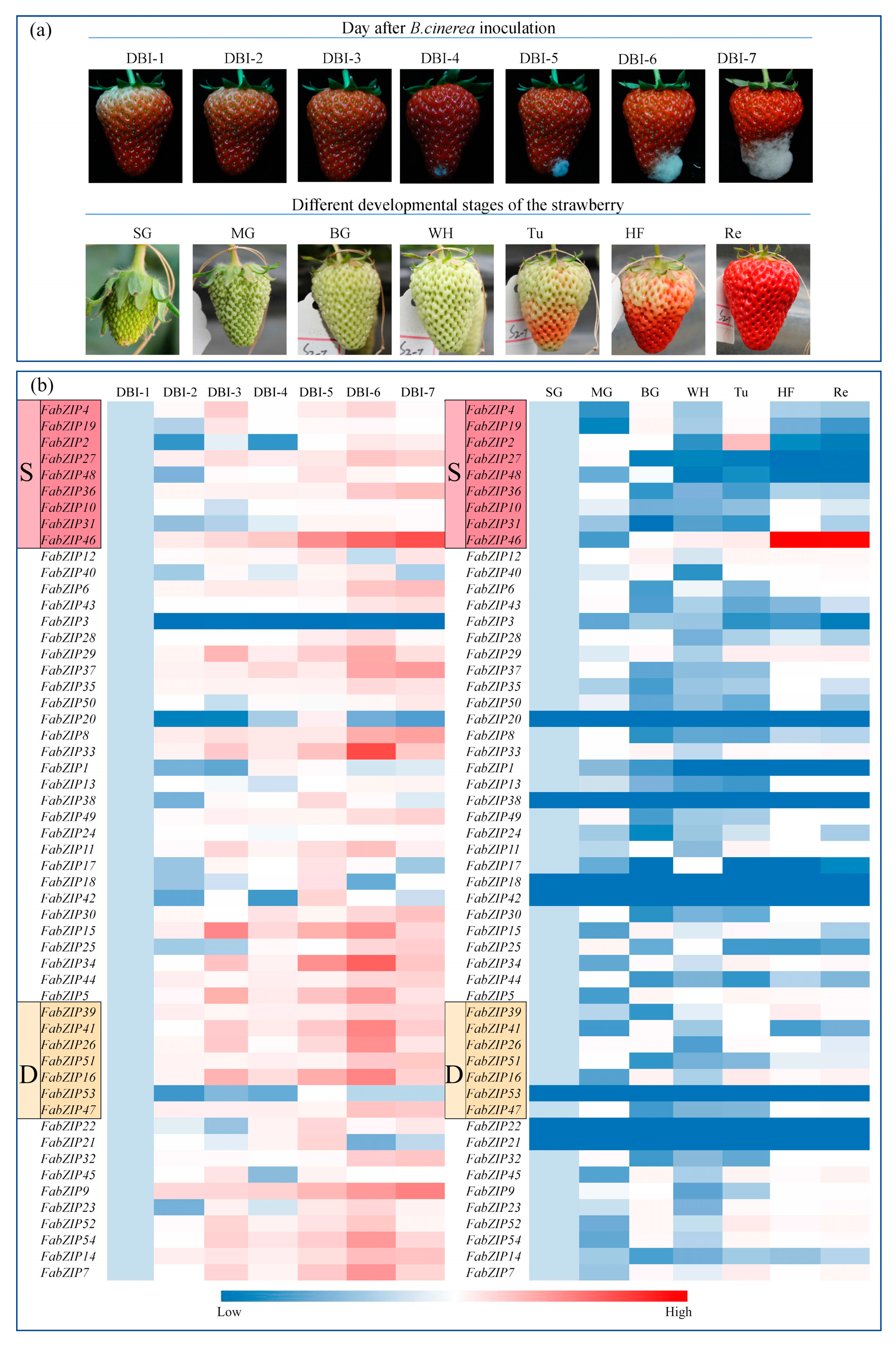
2.4. Response of the FabZIP Genes of Fragaria × Ananassa “Benihoppe” Strawberry to Stress Caused by B. Cinerea
2.5. Analysis of the Transient Expression of FabZIP46 in Strawberries
2.6. Effect of Overexpression and Silencing of FabZIP46 on the Expression of Disease-Resistant Genes
3. Discussion
4. Materials and Methods
4.1. Fungi and Plant Materials
4.2. Extraction of FvbZIP Genes from Strawberries and Prediction of Physicochemical Properties
4.3. Phylogenetic Analysis and Sequence Alignment of Strawberry FvbZIP Genes
4.4. Analysis of the Structure of FvbZIP Genes in the Strawberry and Analysis of the MEME Protein Motif
4.5. Real-Time Fluorescence Quantitative Analysis of Gene Expression
4.6. Gene Cloning and Vector Construction
4.7. Transient Transformation of Strawberries and Experimental Design
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Kumar, R.; Khurana, A.; Sharma, A.K. Role of plant hormones and their interplay in development and ripening of fleshy fruits. J. Exp. Bot. 2014, 65, 4561–4575. [Google Scholar] [CrossRef] [Green Version]
- Symons, G.M.; Chua, Y.J.; Ross, J.J.; Quittenden, L.J.; Davies, N.W.; Reid, J.B. Hormonal changes during non-climacteric ripening in strawberry. J. Exp. Bot. 2012, 63, 4741–4750. [Google Scholar] [CrossRef] [Green Version]
- Amselem, J.; Cuomo, C.A.; Van Kan, J.A.; Viaud, M.; Benito, E.P.; Couloux, A.; Coutinho, P.M.; de Vries, R.P.; Dyer, P.S.; Fillinger, S.; et al. Genomic analysis of the necrotrophic fungal pathogens Sclerotinia sclerotiorum and Botrytis cinerea. PLoS Genet. 2011, 7, e1002230. [Google Scholar] [CrossRef] [Green Version]
- Williamson, B.; Tudzynski, B.; Tudzynski, P.; Van Kan, J.A. Botrytis cinerea: The cause of grey mould disease. Mol. Plant Pathol. 2007, 8, 561–580. [Google Scholar] [CrossRef]
- Dean, R.; Van Kan, J.A.; Pretorius, Z.A.; Hammond, K.E.; Di Pietro, A.; Spanu, P.D.; Rudd, J.J.; Dickman, M.; Kahmann, R.; Ellis, J.; et al. The Top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 2012, 13, 414–430. [Google Scholar] [CrossRef] [Green Version]
- Bestfleisch, M.; Ludererpflimpfl, M.; Hofer, M.; Schulte, E.; Wunsche, J.; Hanke, M.; Flachowsky, H. Evaluation of strawberry (Fragaria L.) genetic resources for resistance to Botrytis cinerea. Plant Physiol. 2015, 64, 396–405. [Google Scholar]
- Windram, O.; Madhou, P.; McHattie, S.; Hill, C.; Hickman, R.; Cooke, E.; Jenkins, D.J.; Penfold, C.A.; Baxter, L.; Breeze, E.; et al. Arabidopsis defense against Botrytis cinerea: Chronology and regulation deciphered by high-resolution temporal transcriptomic analysis. Plant Cell 2012, 24, 3530–3557. [Google Scholar] [CrossRef] [Green Version]
- Pandey, D.; Rajendran, S.R.C.K.; Gaur, M.; Sajeesh, P.K.; Kumar, A. Plant defense signaling and responses against necrotrophic fungal pathogens. J. Plant Growth Regul. 2016, 35, 1159–1174. [Google Scholar] [CrossRef]
- Wei, K.; Chen, J.; Wang, Y.; Chen, Y.; Chen, S.; Lin, Y.; Pan, S.; Zhong, X.; Xie, D. Genome-wide analysis of bZIP-encoding genes in maize. DNA Res. 2012, 19, 463–476. [Google Scholar] [CrossRef] [Green Version]
- Smykowski, A.; Zimmermann, P.; Zentgraf, U. G-Box binding factor1 reduces CATALASE2 expression and regulates the onset of leaf senescence in Arabidopsis. Plant Physiol. 2010, 153, 1321–1331. [Google Scholar] [CrossRef] [Green Version]
- Choi, H.; Hong, J.; Ha, J.; Kang, J.; Kim, S.Y. ABFs, a family of ABA responsive element binding factors. J. Biol. Chem. 2000, 275, 1723–1730. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baena-González, E.; Rolland, F.; Thevelein, J.M.; Sheen, J. A central integrator of transcription networks in plant stress and energy signalling. Nature 2007, 448, 938–942. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.W.; Baek, W.; Lim, S.; Han, S.W.; Lee, S.C. Expression and Functional Roles of the Pepper Pathogen-Induced bZIP Transcription Factor CabZIP2 in Enhanced Disease Resistance to Bacterial Pathogen Infection. Mol. Plant Microbe Interact. 2015, 28, 825–833. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fu, J.; Wu, H.; Ma, S.; Xiang, D.; Liu, R.; Xiong, L. OsJAZ1 Attenuates Drought Resistance by Regulating JA and ABA Signaling in Rice. Front. Plant Sci. 2017, 8, 2108. [Google Scholar] [CrossRef] [Green Version]
- Ma, H.; Liu, C.; Li, Z.; Ran, Q.; Xie, G.; Wang, B.; Fang, S.; Chu, J.; Zhang, J. ZmbZIP4 Contributes to Stress Resistance in Maize by Regulating ABA Synthesis and Root Development. Plant Physiol. 2018, 178, 753–770. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.C.; Choi, H.W.; Hwang, I.S.; Choi, D.S.; Hwang, B.K. Functional roles of the pepper pathogen-induced bZIP transcription factor, CAbZIP1, in enhanced resistance to pathogen infection and environmental stresses. Planta 2006, 224, 1209–1225. [Google Scholar] [CrossRef]
- Liu, C.; Mao, B.; Ou, S.; Wang, W.; Liu, L.; Wu, Y.; Chu, C.; Wang, X. OsbZIP71, a bZIP transcription factor, confers salinity and drought tolerance in rice. Plant Mol. Biol. 2014, 84, 19–36. [Google Scholar] [CrossRef]
- Zhang, N.; Liu, D.; Zheng, W.; He, H.; Ji, B.; Han, Q. A bzip transcription factor, LrbZIP1, is involved in lilium regale wilson defense responses against fusarium oxysporum f. sp. lilii. Genes Genom. 2014, 36, 789–798. [Google Scholar] [CrossRef]
- Wang, X.L.; Chen, X.; Yang, T.B.; Cheng, Q.; Cheng, Z.M. Genome-Wide Identification of bZIP Family Genes Involved in Drought and Heat Stresses in Strawberry (Fragaria vesca). Int. J. Genom. 2017, 2017, 3981031. [Google Scholar]
- Zhang, T.; Lv, W.; Zhang, H.; Ma, L.; Li, P.; Ge, L.; Li, G. Genome-wide analysis of the basic Helix-Loop-Helix (bHLH) transcription factor family in maize. BMC Plant Biol. 2018, 18, 235. [Google Scholar] [CrossRef] [Green Version]
- Jakoby, M.; Weisshaar, B.; Dröge-Laser, W.; Vicente-Carbajosa, J.; Tiedemann, J.; Kroj, T.; Parcy, F. bZIP transcription factors in Arabidopsis. Trends Plant Sci. 2002, 7, 106–111. [Google Scholar] [CrossRef]
- Liu, J.; Chen, N.; Chen, F.; Cai, B.; Dal Santo, S.; Tornielli, G.B.; Pezzotti, M.; Cheng, Z.M. Genome-wide analysis and expression profile of the bZIP transcription factor gene family in grapevine (Vitis vinifera). BMC Genom. 2014, 15, 281. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.Y.; Meng, D.; Li, M.; Cheng, L. Genome-wide identification and expression analysis of the bzip gene family in apple (malus domestica). Tree Genet Genomes 2016, 12, 82. [Google Scholar] [CrossRef]
- Sibéril, Y.; Doireau, P.; Gantet, P. Plant bZIP G-box binding factors. Modular structure and activation mechanisms. Eur. J. Biochem. 2001, 268, 5655–5666. [Google Scholar] [CrossRef]
- Cyrek, M.; Fedak, H.; Ciesielski, A.; Guo, Y.; Sliwa, A.; Brzezniak, L.; Krzyczmonik, K.; Pietras, Z.; Kaczanowski, S.; Liu, F.; et al. Seed Dormancy in Arabidopsis Is Controlled by Alternative Polyadenylation of DOG1. Plant Physiol. 2016, 170, 947–955. [Google Scholar] [CrossRef] [Green Version]
- Nijhawan, A.; Jain, M.; Tyagi, A.K.; Khurana, J.P. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol. 2008, 146, 333–350. [Google Scholar] [CrossRef] [Green Version]
- Rong, S.; Wu, Z.; Cheng, Z.; Zhang, S.; Liu, H.; Huang, Q. Genome-Wide Identification, Evolutionary Patterns, and Expression Analysis of bZIP Gene Family in Olive (Olea europaea L.). Genes 2020, 11, 510. [Google Scholar] [CrossRef]
- Li, X.; Duan, X.; Jiang, H.; Sun, Y.; Tang, Y.; Yuan, Z.; Guo, J.; Liang, W.; Chen, L.; Yin, J.; et al. Genome-wide analysis of basic/helix-loop-helix transcription factor family in rice and Arabidopsis. Plant Physiol. 2006, 141, 1167–1184. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Zhou, J.; Zhang, B.; Vanitha, J.; Ramachandran, S.; Jiang, S.Y. Genome-wide expansion and expression divergence of the basic leucine zipper transcription factors in higher plants with an emphasis on sorghum. J. Integr. Plant Biol. 2011, 53, 212–231. [Google Scholar] [CrossRef]
- Crawford, N.M.; Glass, A.D.M. Molecular and physiological aspects of nitrate uptake in plants. Trends Plant Sci. 1998, 3, 389–395. [Google Scholar] [CrossRef]
- Büttner, M.; Singh, K.B. Arabidopsis thaliana ethylene-responsive element binding protein (AtEBP), an ethylene-inducible, GCC box DNA-binding protein interacts with an ocs element binding protein. Proc. Natl. Acad. Sci. USA 1997, 94, 5961–5966. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.J.; Lee, M.Y.; Yi, S.Y.; Oh, S.K.; Choi, S.H.; Her, N.H.; Choi, D.; Min, B.W.; Yang, S.G.; Harn, C.H. PPI1: A novel pathogen-induced basic region-leucine zipper (bzip) transcription factor from pepper. Mol. Plant Microbe Interact. 2002, 15, 540–548. [Google Scholar] [CrossRef] [Green Version]
- Meng, X.B.; Zhao, W.S.; Lin, R.M.; Wang, M.; Peng, Y.L. Identification of a novel rice bzip-type transcription factor gene, osbzip1, involved in response to infection ofmagnaporthe grisea. Plant Mol. Biol. Rep. 2005, 23, 301–302. [Google Scholar] [CrossRef]
- Tateda, C.; Ozaki, R.; Onodera, Y.; Takahashi, Y.; Yamaguchi, K.; Berberich, T.; Koizumi, N.; Kusano, T. NtbZIP60, an endoplasmic reticulum-localized transcription factor, plays a role in the defense response against bacterial pathogens in Nicotiana tabacum. J. Plant Res. 2008, 121, 603–611. [Google Scholar] [CrossRef]
- Thurow, C.; Schiermeyer, A.; Krawczyk, S.; Butterbrodt, T.; Nickolov, K.; Gatz, C. Tobacco bZIP transcription factor TGA2.2 and related factor TGA2.1 have distinct roles in plant defense responses and plant development. Plant J. 2005, 44, 100–113. [Google Scholar] [CrossRef]
- Tak, H.; Mhatre, M. Cloning and molecular characterization of a putative bZIP transcription factor VvbZIP23 from Vitis vinifera. Protoplasma 2013, 250, 333–345. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, G.; Xia, N.; Wang, X.J.; Huang, L.L.; Kang, Z.S. Cloning and characterization of a bzip transcription factor gene in wheat and its expression in response to stripe rust pathogen infection and abiotic stresses. Physiol. Mol. Plant Pathol. 2008, 73, 88–94. [Google Scholar] [CrossRef]
- Orellana, S.; Yañez, M.; Espinoza, A.; Verdugo, I.; González, E.; Ruiz-Lara, S.; Casaretto, J.A. The transcription factor SlAREB1 confers drought, salt stress tolerance and regulates biotic and abiotic stress-related genes in tomato. Plant Cell Environ. 2010, 33, 2191–2208. [Google Scholar] [CrossRef]
- Higuera, J.J.; Garrido, J.; Lekhbou, A.; Arjona, I.; Amil, F.; Mercado, J.A.; Pliego, F.; Muñoz, J.; López, C.J.; Caballero, J.L. The Strawberry FaWRKY1 Transcription Factor Negatively Regulates Resistance to Colletotrichum acutatum in Fruit Upon Infection. Front Plant Sci. 2019, 10, 480. [Google Scholar] [CrossRef]
- Zheng, Z.; Qamar, S.A.; Chen, Z.; Mengiste, T. Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens. Plant J. 2006, 48, 592–605. [Google Scholar] [CrossRef]
- Hu, Y.; Dong, Q.; Yu, D. Arabidopsis WRKY46 coordinates with WRKY70 and WRKY53 in basal resistance against pathogen Pseudomonas syringae. Plant Sci. 2012, 185, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Mehli, L.; Schaart, J.G.; Kjellsen, T.D.; Tran, D.H.; Salentijn, E.M.J.; Iversen, S.T.H. A gene encoding a polygalacturonase-inhibiting protein (pgip) shows developmental regulation and pathogen-induced expression in strawberry. New Phytol. 2004, 163, 99–110. [Google Scholar] [CrossRef]
- Khan, A.A.; Shih, D.S. Molecular cloning, characterization and expression analysis of two class II chitinase genes from the strawberry plant. Plant Sci. 2004, 166, 753–762. [Google Scholar] [CrossRef]
- Wang, K.; Liao, Y.; Xiong, Q.; Kan, J.; Cao, S.; Zheng, Y. Induction of Direct or Priming Resistance against Botrytis cinerea in Strawberries by β-Aminobutyric Acid and Their Effects on Sucrose Metabolism. J. Agric. Food Chem. 2016, 64, 5855–5865. [Google Scholar] [CrossRef]
- Sornaraj, P.; Luang, S.; Lopato, S.; Hrmova, M. Basic leucine zipper (bZIP) transcription factors involved in abiotic stresses: A molecular model of a wheat bZIP factor and implications of its structure in function. Biochim. Biophys. Acta 2016, 1860, 46–56. [Google Scholar] [CrossRef]
- Yin, W.; Cui, P.; Wei, W.; Lin, Y.; Luo, C. Genome-wide identification and analysis of the basic leucine zipper (bZIP) transcription factor gene family in Ustilaginoidea virens. Genome 2017, 60, 1051–1059. [Google Scholar] [CrossRef] [Green Version]
- Yang, S.; Xu, K.; Chen, S.; Li, T.; Xia, H.; Chen, L.; Liu, H.; Luo, L. A stress-responsive bZIP transcription factor OsbZIP62 improves drought and oxidative tolerance in rice. BMC Plant Biol. 2019, 19, 260. [Google Scholar] [CrossRef] [Green Version]
- Uno, Y.; Furihata, T.; Abe, H.; Yoshida, R.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc. Natl. Acad. Sci. USA 2000, 97, 11632–11637. [Google Scholar] [CrossRef] [Green Version]
- Kesarwani, M.; Yoo, J.; Dong, X. Genetic interactions of TGA transcription factors in the regulation of pathogenesis-related genes and disease resistance in Arabidopsis. Plant Physiol. 2007, 144, 336–346. [Google Scholar] [CrossRef] [Green Version]
- Liao, Y.; Zou, H.F.; Wei, W.; Hao, Y.J.; Tian, A.G.; Huang, J.; Liu, Y.F.; Zhang, J.S.; Chen, S.Y. Soybean GmbZIP44, GmbZIP62 and GmbZIP78 genes function as negative regulator of ABA signaling and confer salt and freezing tolerance in transgenic Arabidopsis. Planta 2008, 228, 225–240. [Google Scholar] [CrossRef]
- Rügner, A.; Frohnmeyer, H.; Näke, C.; Wellmer, F.; Kircher, S.; Schäfer, E.; Harter, K. Isolation and characterization of four novel parsley proteins that interact with the transcriptional regulators CPRF1 and CPRF2. Mol. Genet. Genom. 2001, 265, 964–976. [Google Scholar] [CrossRef] [PubMed]
- Chuang, C.F.; Running, M.P.; Williams, R.W.; Meyerowitz, E.M. The PERIANTHIA gene encodes a bZIP protein involved in the determination of floral organ number in Arabidopsis thaliana. Genes Dev. 1999, 13, 334–344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wellmer, F.; Schäfer, E.; Harter, K. The DNA binding properties of the parsley bZIP transcription factor CPRF4a are regulated by light. J. Biol. Chem. 2001, 276, 6274–6279. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chern, M.S.; Bobb, A.J.; Bustos, M.M. The regulator of MAT2 (ROM2) protein binds to early maturation promoters and represses PvALF-activated transcription. Plant Cell 1996, 8, 305–321. [Google Scholar]
- Hardtke, C.S.; Gohda, K.; Osterlund, M.T.; Oyama, T.; Okada, K.; Deng, X.W. HY5 stability and activity in arabidopsis is regulated by phosphorylation in its COP1 binding domain. EMBO J. 2000, 19, 4997–5006. [Google Scholar] [CrossRef]
- Yin, Y.; Zhu, Q.; Dai, S.; Lamb, C.; Beachy, R.N. RF2a, a bZIP transcriptional activator of the phloem-specific rice tungro bacilliform virus promoter, functions in vascular development. EMBO J. 1997, 16, 5247–5259. [Google Scholar] [CrossRef] [Green Version]
- Weltmeier, F.; Rahmani, F.; Ehlert, A.; Dietrich, K.; Schütze, K.; Wang, X.; Chaban, C.; Hanson, J.; Teige, M.; Harter, K.; et al. Expression patterns within the Arabidopsis C/S1 bZIP transcription factor network: Availability of heterodimerization partners controls gene expression during stress response and development. Plant Mol. Biol. 2009, 69, 107–119. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.L.; Zhong, Y.; Cheng, Z.M.; Xiong, J.S. Divergence of the bZIP Gene Family in Strawberry, Peach, and Apple Suggests Multiple Modes of Gene Evolution after Duplication. Int. J. Genom. 2015, 2015, 536943. [Google Scholar] [CrossRef] [Green Version]
- Van Verk, M.C.; Neeleman, L.; Bol, J.F.; Linthorst, H.J. Tobacco Transcription Factor NtWRKY12 Interacts with TGA2.2 in vitro and in vivo. Front Plant Sci. 2011, 2, 32. [Google Scholar] [CrossRef] [Green Version]
- Jia, H.; Wang, Y.; Sun, M.; Li, B.; Han, Y.; Zhao, Y.; Li, X.; Ding, N.; Li, C.; Ji, W.; et al. Sucrose functions as a signal involved in the regulation of strawberry fruit development and ripening. New Phytol. 2013, 198, 453–465. [Google Scholar] [CrossRef]




© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, B.; Wang, Y.; Zhang, G.; Feng, Y.; Yan, Z.; Wu, J.; Chen, X. Genome-Wide Identification and Expression Analysis of the Strawberry FvbZIP Gene Family and the Role of Key Gene FabZIP46 in Fruit Resistance to Gray Mold. Plants 2020, 9, 1199. https://doi.org/10.3390/plants9091199
Lu B, Wang Y, Zhang G, Feng Y, Yan Z, Wu J, Chen X. Genome-Wide Identification and Expression Analysis of the Strawberry FvbZIP Gene Family and the Role of Key Gene FabZIP46 in Fruit Resistance to Gray Mold. Plants. 2020; 9(9):1199. https://doi.org/10.3390/plants9091199
Chicago/Turabian StyleLu, Bei, Yuanhua Wang, Geng Zhang, Yingna Feng, Zhiming Yan, Jianhua Wu, and Xuehao Chen. 2020. "Genome-Wide Identification and Expression Analysis of the Strawberry FvbZIP Gene Family and the Role of Key Gene FabZIP46 in Fruit Resistance to Gray Mold" Plants 9, no. 9: 1199. https://doi.org/10.3390/plants9091199



