Genetic Structure of Invasive Baby’s Breath (Gypsophila paniculata L.) Populations in a Michigan Dune System
Abstract
:1. Introduction
2. Results
2.1. Microsatellite Genotyping and Genetic Diversity
2.2. Genetic Structure
2.3. Landscape Genetics
3. Discussion
4. Materials and Methods
4.1. Study Area and Sampling Collection
4.2. Microsatellite Genotyping
4.3. Quality Control
4.4. nSSR Genetic Diversity
4.5. nSSR Genetic Structure
4.6. cpSSR Genetic Diversity
4.7. nSSR and cpSSR Genetic Structure
4.8. Landscape Genetics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Arbogast, A.F.; Loope, W.L. Maximum-limiting ages of Lake Michigan coastal dunes: Their correlation with Holocene lake level history. J. Great Lakes Res. 1999, 25, 372–382. [Google Scholar] [CrossRef]
- Everard, M.; Jones, L.; Watts, B. Have we neglected the societal importance of sand dunes? An ecosystem services perspective. Aquat. Conserv. 2010, 20, 476–487. [Google Scholar] [CrossRef]
- Blumer, B.E.; Arbogast, A.F.; Forman, S.L. The OSL chronology of eolian sand deposition in a perched dune field along the northwestern shore of Lower Michigan. Quat. Res. 2012, 77, 445–455. [Google Scholar] [CrossRef]
- Cowles, H.C. The ecological relations of the vegetation on the sand dunes of Lake Michigan. Part I.—Geographical relations of the dune floras. Bot. Gaz. 1899, 27, 95–117. [Google Scholar] [CrossRef] [Green Version]
- Olson, J.S. Rates of succession and soil changes on southern Lake Michigan sand dunes. Bot. Gaz. 1958. [Google Scholar] [CrossRef]
- Arbogast, A.F.; Tonya, C.; Davis, C.F., III; DeVries-Zimmerman, S.; Yurk, B.; Garmon, B.; van Dijk, D.; VanHorn, J. Bringing the Latest Science to the Management of Michigan’s Coastal Dunes 2015. Available online: https://d3n8a8pro7vhmx.cloudfront.net/environmentalcouncil/pages/467/attachments/original/1518314397/Latest_Science-Michigan_Coastal_Dunes.pdf?1518314397%22 (accessed on 10 July 2017).
- Albert, D. Borne of the Wind: Michigan Sand Dunes; University of Michigan Press: Ann Arbor, MI, USA, 2006; ISBN 978-0-472-03172-6. [Google Scholar]
- Jørgensen, R.H.; Kollmann, J. Invasion of coastal dunes by the alien shrub Rosa rugosa is associated with roads, tracks and houses. Flora 2009, 204, 289–297. [Google Scholar] [CrossRef]
- Carranza, M.L.; Carboni, M.; Feola, S.; Acosta, A.T.R. Landscape-scale patterns of alien plant species on coastal dunes: The case of iceplant in central Italy. Appl. Veg. Sci. 2010, 13, 135–145. [Google Scholar] [CrossRef]
- Rand, T.A.; Louda, S.M.; Bradley, K.M.; Crider, K.K. Effects of invasive knapweed (Centaurea stoebe subsp. micranthos) on a threatened native thistle (Cirsium pitcheri) vary with environment and life stage. Botany 2015, 93, 543–558. [Google Scholar] [CrossRef]
- Grime, J.P. Plant Strategies and Vegetation Processes; John Wiley & Sons: Chichester, UK, 1979; ISBN 978-0-471-99695-8. [Google Scholar]
- Baker, H.G. Patterns of plant invasion in North America. In Ecology of Biological Invasions of North America and Hawaii; Mooney, H.A., Drake, J.A., Eds.; Ecological Studies; Springer: New York, NY, USA, 1986; pp. 44–57. ISBN 978-1-4612-4988-7. [Google Scholar]
- Sakai, A.; Allendorf, F.; Holt, J.; Lodge, D.; Molofsky, J.; With, K.; Baughman, S.; Cabin, R.; Cohen, J.; Ellstrand, N.; et al. The population biology of invasive species. Annu. Rev. Ecol. Syst. 2001, 32, 305–332. [Google Scholar] [CrossRef] [Green Version]
- Cusseddu, V.; Ceccherelli, G.; Bertness, M. Hierarchical organization of a Sardinian sand dune plant community. PeerJ 2016, 4, e2199. [Google Scholar] [CrossRef] [Green Version]
- With, K.A. The landscape ecology of invasive spread. Conserv. Biol. 2002, 16, 1192–1203. [Google Scholar] [CrossRef] [Green Version]
- Theoharides, K.A.; Dukes, J.S. Plant invasion across space and time: Factors affecting nonindigenous species success during four stages of invasion. New Phytol. 2007, 176, 256–273. [Google Scholar] [CrossRef] [PubMed]
- Dlugosch, K.M.; Parker, I.M. Founding events in species invasions: Genetic variation, adaptive evolution, and the role of multiple introductions. Mol. Ecol. 2008, 17, 431–449. [Google Scholar] [CrossRef] [PubMed]
- Crosby, K.; Stokes, T.O.; Latta, R.G. Evolving California genotypes of Avena barbata are derived from multiple introductions but still maintain substantial population structure. PeerJ 2014, 2, e633. [Google Scholar] [CrossRef] [Green Version]
- Hagenblad, J.; Hülskötter, J.; Acharya, K.P.; Brunet, J.; Chabrerie, O.; Cousins, S.A.O.; Dar, P.A.; Diekmann, M.; De Frenne, P.; Hermy, M.; et al. Low genetic diversity despite multiple introductions of the invasive plant species Impatiens glandulifera in Europe. BMC Genet. 2015, 16, 103. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, C.-Y.; Tang, S.; Fatemi, M.; Gross, C.L.; Julien, M.H.; Curtis, C.; van Klinken, R.D. Population structure and genetic diversity of invasive Phyla canescens: Implications for the evolutionary potential. Ecosphere 2015, 6, art162. [Google Scholar] [CrossRef]
- Nagy, A.-M.; Korpelainen, H. Population genetics of Himalayan balsam (Impatiens glandulifera): Comparison of native and introduced populations. Plant Ecol. Divers. 2015, 8, 317–321. [Google Scholar] [CrossRef]
- Moran, E.V.; Reid, A.; Levine, J.M. Population genetics and adaptation to climate along elevation gradients in invasive Solidago canadensis. PLoS ONE 2017, 12. [Google Scholar] [CrossRef] [Green Version]
- Bustamante, R.O.; Durán, A.P.; Peña-Gómez, F.T.; Véliz, D. Genetic and phenotypic variation, dispersal limitation and reproductive success in the invasive herb Eschscholzia californica along an elevation gradient in central Chile. Plant Ecol. Divers. 2017, 10, 419–429. [Google Scholar] [CrossRef]
- DNR. Michigan Invasive Species Grant Program Handbook; Department of Natural Resources (DNR): Lansing, MI, USA, 2014; p. 26.
- Barkoudah, Y.I. A revision of Gypsophila, Bolanthus, Ankyropetalum and Phryna. Wentia 1962, 9, 1–203. [Google Scholar] [CrossRef]
- Darwent, A.L.; Coupland, R.T. Life history of Gypsophila paniculata. Weeds 1966, 14, 313–318. [Google Scholar] [CrossRef]
- Darwent, A.L. The biology of Canadian weeds: 14. Gypsophila paniculata L. Can. J. Plant Sci. 1975, 55, 1049–1058. [Google Scholar] [CrossRef]
- Lamar, S.K.; Partridge, C.G. Old meets new: Combining herbarium databases with genetic methods to evaluate the invasion status of baby’s breath (Gypsophila paniculata) in North America. BioRxiv 2019. [Google Scholar] [CrossRef] [Green Version]
- Baskett, C.A.; Emery, S.M.; Rudgers, J.A. Pollinator visits to threatened species are restored following invasive plant removal. Int. J. Plant Sci. 2011, 172, 411–422. [Google Scholar] [CrossRef]
- Jolls, C.L.; Marik, J.E.; Hamzé, S.I.; Havens, K. Population viability analysis and the effects of light availability and litter on populations of Cirsium pitcheri, a rare, monocarpic perennial of Great Lakes shorelines. Biol. Conserv. 2015, 187, 82–90. [Google Scholar] [CrossRef]
- Emery, S.M.; Doran, P.J. Presence and management of the invasive plant Gypsophila paniculata (baby’s breath) on sand dunes alters arthropod abundance and community structure. Biol. Conserv. 2013, 161, 174–181. [Google Scholar] [CrossRef]
- Weir, B.S.; Cockerham, C.C. Estimating F-Statistics for the analysis of population structure. Evolution 1984, 38, 1358–1370. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [Green Version]
- Estoup, A.; Guillemaud, T. Reconstructing routes of invasion using genetic data: Why, how and so what? Mol. Ecol. 2010, 19, 4113–4130. [Google Scholar] [CrossRef] [PubMed]
- Ellstrand, N.C.; Elam, D.R. Population genetic consequences of small population size: Implications for plant conservation. Annu. Rev. Ecol. Syst. 1993, 24, 217–242. [Google Scholar] [CrossRef]
- Young, A.; Boyle, T.; Brown, T. The population genetic consequences of habitat fragmentation for plants. Trends Ecol. Evol. 1996, 11, 413–418. [Google Scholar] [CrossRef]
- Keller, L.F.; Waller, D.M. Inbreeding effects in wild populations. Trends Ecol. Evol. 2002, 17, 230–241. [Google Scholar] [CrossRef]
- Karamanski, T. A Nationalized Lakeshore: The Creation and Administration of Sleeping Bear Dunes National Lakeshore; US Government Publishing Office: Washington, DC, USA, 2000. Available online: https://www.nps.gov/parkhistory/online_books/slbe/index.htm (accessed on 24 November 2015).
- Emery, S.M.; Doran, P.J.; Legge, J.T.; Kleitch, M.; Howard, S. Aboveground and belowground impacts following removal of the invasive species baby’s breath (Gypsophila paniculata) on Lake Michigan sand dunes: Plant and soil impacts of G. paniculata removal. Restor. Ecol. 2013, 21, 506–514. [Google Scholar] [CrossRef]
- TNC. Lake Michigan Coastal Dune Restoration Project. 2012 Field Season Report; The Nature Conservancy (TNC) and Partners: Lansing, MI, USA, 2012; Internal Report. [Google Scholar]
- Fant, J.B.; Havens, K.; Keller, J.M.; Radosavljevic, A.; Yates, E.D. The influence of contemporary and historic landscape features on the genetic structure of the sand dune endemic, Cirsium pitcheri (Asteraceae). Heredity 2014, 112, 519–530. [Google Scholar] [CrossRef] [Green Version]
- Keddy, C.J.; Keddy, P.A. Reproductive biology and habitat of Cirsium pitcheri. Mich. Bot. 1984, 23, 57–67. [Google Scholar]
- Martínez-Nieto, M.I.; Segarra-Moragues, J.G.; Merlo, E.; Martínez-Hernández, F.; Mota, J.F. Genetic diversity, genetic structure and phylogeography of the Iberian endemic Gypsophila struthium (Caryophyllaceae) as revealed by AFLP and plastid DNA sequences: Connecting habitat fragmentation and diversification: Phylogeography of Gypsophila struthium. Bot. J. Linn. Soc. 2013, 173, 654–675. [Google Scholar] [CrossRef] [Green Version]
- Bang, S.W.; Chung, S.-M. One size does not fit all: The risk of using amplicon size of chloroplast SSR marker for genetic relationship studies. Plant Cell. Rep. 2015, 34, 1681–1683. [Google Scholar] [CrossRef] [Green Version]
- Ansong, M.; Pickering, C. Long-distance dispersal of Black Spear Grass (Heteropogon contortus) seed on socks and trouser legs by walkers in Kakadu National Park. Ecol. Manag. Restor. 2013, 14, 71–74. [Google Scholar] [CrossRef]
- Ansong, M.; Pickering, C. Weed seeds on clothing: A global review. J. Environ. Manag. 2014, 144, 203–211. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keller, S.R.; Taylor, D.R. Genomic admixture increases fitness during a biological invasion. J. Evol. Biol. 2010, 23, 1720–1731. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Stift, M.; van Kleunen, M. Admixture increases performance of an invasive plant beyond first-generation heterosis. J. Ecol. 2018, 106, 1595–1606. [Google Scholar] [CrossRef] [Green Version]
- Qiao, H.; Liu, W.; Zhang, Y.; Zhang, Y.; Li, Q.Q. Genetic admixture accelerates invasion via provisioning rapid adaptive evolution. Mol. Ecol. 2019, 28, 4012–4027. [Google Scholar] [CrossRef] [PubMed]
- Leimbach-Maus, H.B.; Parks, S.R.; Partridge, C.G. Microsatellite primer development for the invasive perennial herb Gypsophila paniculata (Caryophyllaceae). Appl. Plant Sci. 2018, 6, e01203. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chung, S.-M.; Staub, J.E. The development and evaluation of consensus chloroplast primer pairs that possess highly variable sequence regions in a diverse array of plant taxa. Theor. Appl. Genet. 2003, 107, 757–767. [Google Scholar] [CrossRef] [PubMed]
- Calistri, E.; Buiatti, M.; Bogani, P. Characterization of Gypsophila species and commercial hybrids with nuclear whole-genome and cytoplasmic molecular markers. Plant Biosyst. 2016, 150, 11–21. [Google Scholar] [CrossRef]
- Dewoody, J.; Nason, J.D.; Hipkins, V.D. Mitigating scoring errors in microsatellite data from wild populations. Mol. Ecol. Notes 2006, 6, 951–957. [Google Scholar] [CrossRef] [Green Version]
- Guichoux, E.; Lagache, L.; Wagner, S.; Chaumeil, P.; Léger, P.; Lepais, O.; Lepoittevin, C.; Malausa, T.; Revardel, E.; Salin, F.; et al. Current trends in microsatellite genotyping. Mol. Ecol. Resour. 2011, 11, 591–611. [Google Scholar] [CrossRef]
- Van Oosterhout, C.; Hutchinson, W.F.; Wills, D.P.M.; Shipley, P. micro-checker: Software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 2004, 4, 535–538. [Google Scholar] [CrossRef]
- Van Oosterhout, C.; Weetman, D.; Hutchinson, W.F. Estimation and adjustment of microsatellite null alleles in nonequilibrium populations. Mol. Ecol. Notes 2006, 6, 255–256. [Google Scholar] [CrossRef]
- Gomes, I.; Collins, A.; Lonjou, C.; Thomas, N.S.; Wilkinson, J.; Watson, M.; Morton, N. Hardy–Weinberg quality control. Ann. Hum. Genet. 1999, 63, 535–538. [Google Scholar] [CrossRef] [PubMed]
- Archer, F.I.; Adams, P.E.; Schneiders, B.B. STRATAG: An R package for manipulating, summarizing and analysing population genetic data. Mol. Ecol. Resour. 2017, 17, 5–11. [Google Scholar] [CrossRef]
- Raymond, M.; Rousset, F. GENEPOP (version 1.2): Population genetic software for exact tests and ecumenicism. J. Hered. 1995, 86, 248–249. [Google Scholar] [CrossRef]
- Rousset, F. GENEPOP’007: A complete re-implementation of the GENEPOP software for Windows and Linux. Mol. Ecol. Resour. 2008, 8, 103–106. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keenan, K.; McGinnity, P.; Cross, T.F.; Crozier, W.W.; Prodöhl, P.A. diveRsity: An R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol. Evol. 2013, 4, 782–788. [Google Scholar] [CrossRef] [Green Version]
- Kopelman, N.M.; Mayzel, J.; Jakobsson, M.; Rosenberg, N.A.; Mayrose, I. Clumpak: A program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 2015, 15, 1179–1191. [Google Scholar] [CrossRef] [Green Version]
- Jombart, T. adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [Green Version]
- Jombart, T.; Devillard, S.; Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 2010, 11, 94. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Slatkin, M. A measure of population subdivision based on microsatellite allele frequencies. Genetics 1995, 139, 457–462. [Google Scholar] [PubMed]
- Smouse, P.E.; Long, J.C.; Sokal, R.R. Multiple regression and correlation extensions of the mantel test of matrix correspondence. Syst. Zool. 1986, 35, 627–632. [Google Scholar] [CrossRef]
- Smouse, P.E.; Long, J.C. Matrix correlation analysis in anthropology and genetics. Am. J. Phys. Anthropol. 1992, 35, 187–213. [Google Scholar] [CrossRef]
- Eliades, N.-G.; Eliades, D. Haplotype analysis: Software for Analysis of Haplotype Data. Available online: https://www.researchgate.net/publication/221936337_HAPLOTYPE_ANALYSIS_software_for_analysis_of_haplotype_data (accessed on 18 May 2018).
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2014, 2, e281. [Google Scholar] [CrossRef] [Green Version]
- Nei, M. Genetic distance between populations. Am. Nat. 1972, 106, 283–292. [Google Scholar] [CrossRef]
- Excoffier, L.; Smouse, P.E.; Quattro, J.M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: Application to human mitochondrial DNA restriction data. Genetics 1992, 131, 479–491. [Google Scholar]
- Peterman, W.E.; Connette, G.M.; Semlitsch, R.D.; Eggert, L.S. Ecological resistance surfaces predict fine-scale genetic differentiation in a terrestrial woodland salamander. Mol. Ecol. 2014, 23, 2402–2413. [Google Scholar] [CrossRef]
- Peterman, W.E. ResistanceGA: An R package for the optimization of resistance surfaces using genetic algorithms. Methods Ecol. Evol. 2018, 9, 1638–1647. [Google Scholar] [CrossRef] [Green Version]
- Jenness, J.; Brost, B.; Beier, P. Land Facet Corridor Designer: Extension for ArcGIS; Jenness Enterprises: Flagstaff, AZ, USA, 2013. [Google Scholar]
- Evans, J.; Oakleaf, J.; Cushman, S.; Theobald, D. An ArcGIS Toolbox for Surface Gradient and Geomorphometric Modeling, Version 2.0-0. 2014. Available online: https://github.com/jeffreyevans/GradientMetrics (accessed on 12 January 2019).
- McRae, B.H.; Dickson, B.G.; Keitt, T.H.; Shah, V.B. Using circuit theory to model connectivity in ecology, evolution, and conservation. Ecology 2008, 89, 2712–2724. [Google Scholar] [CrossRef]
- Piya, S.; Nepal, M.P.; Butler, J.L.; Larson, G.E.; Neupane, A. Genetic diversity and population structure of sickleweed (Falcaria vulgaris; Apiaceae) in the upper Midwest USA. Biol. Invasions 2014, 16, 2115–2125. [Google Scholar] [CrossRef]
- Sakata, Y.; Itami, J.; Isagi, Y.; Ohgushi, T. Multiple and mass introductions from limited origins: Genetic diversity and structure of Solidago altissima in the native and invaded range. J. Plant Res. 2015, 128, 909–921. [Google Scholar] [CrossRef] [PubMed] [Green Version]
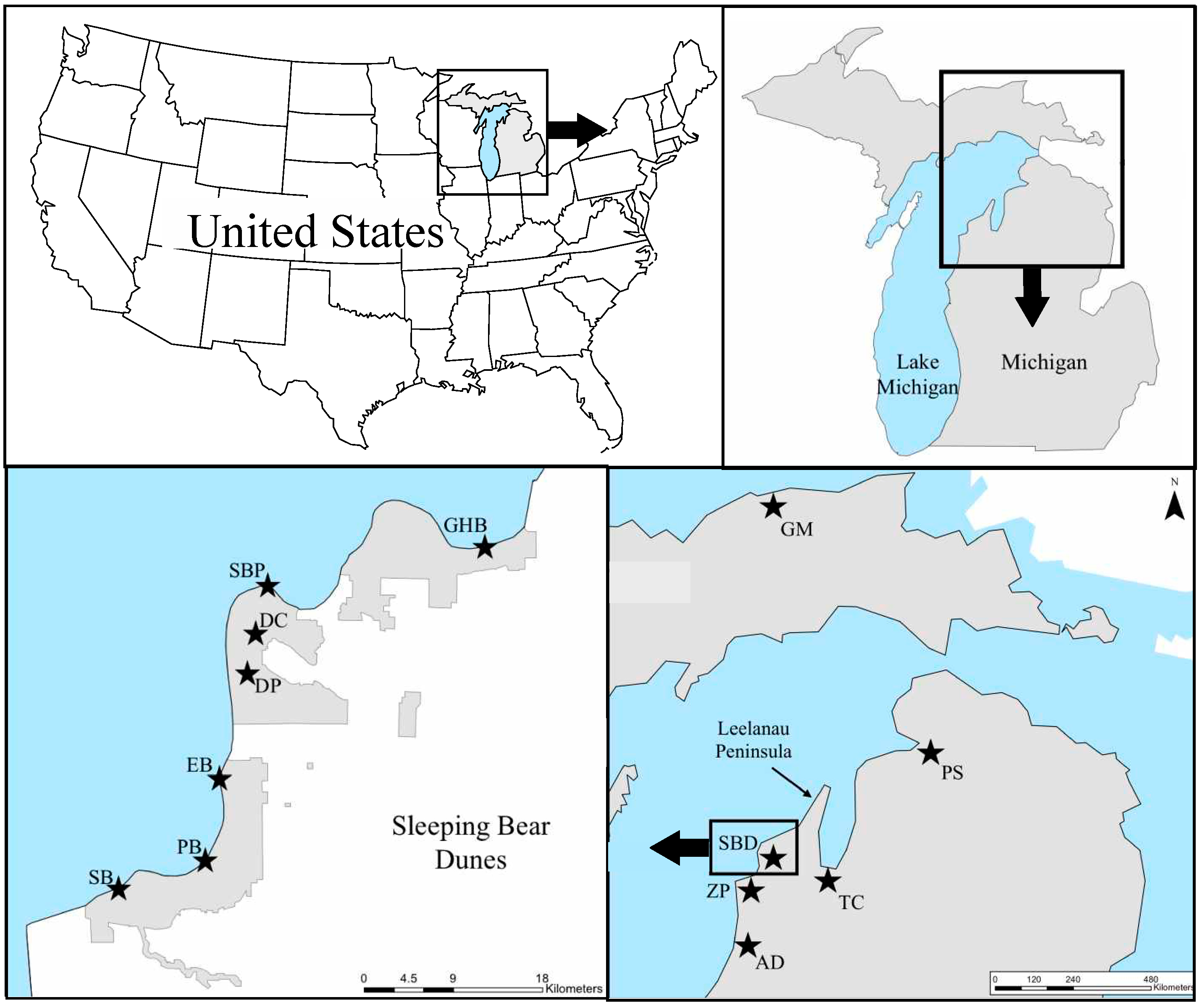

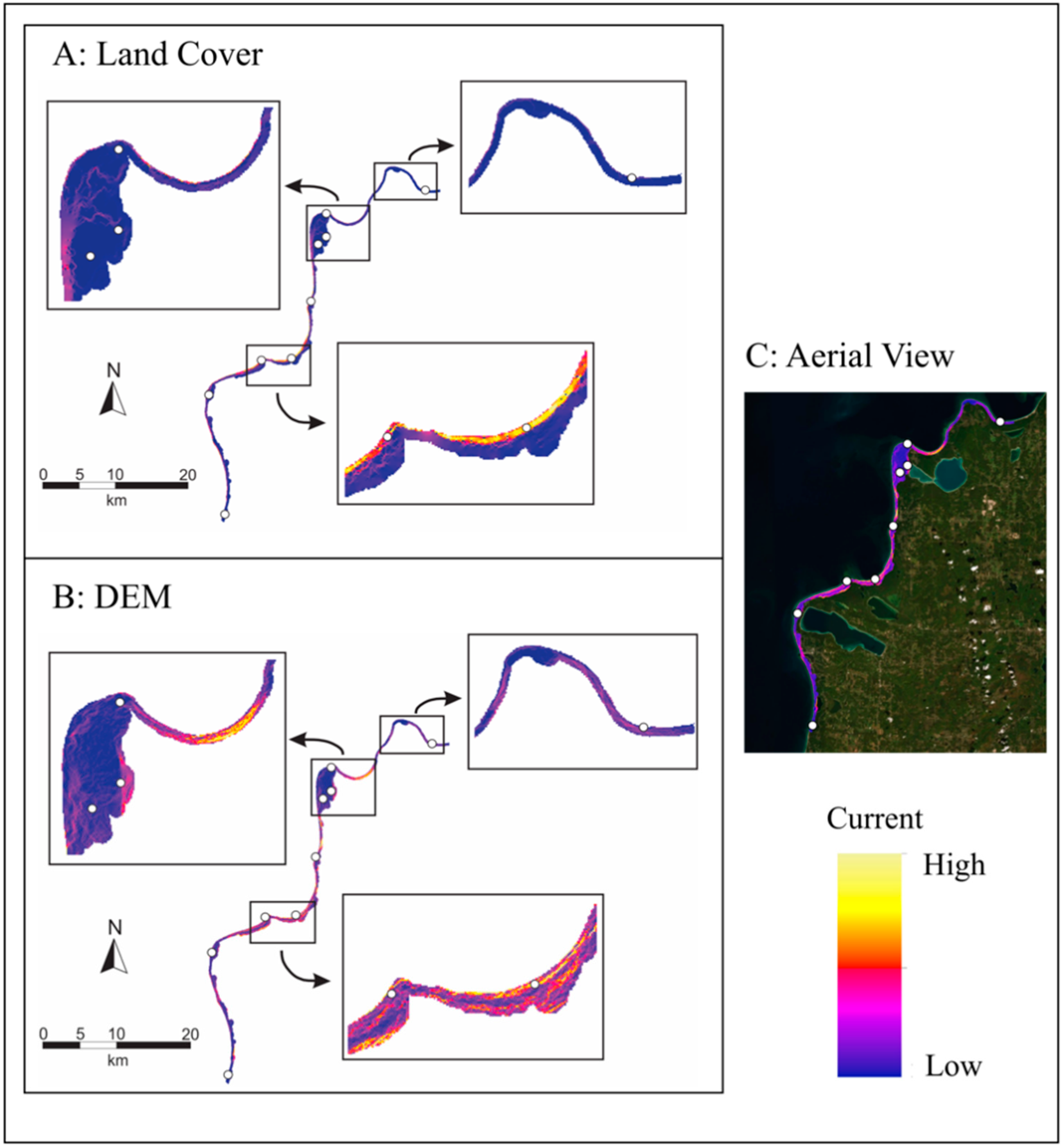
| Sampling Locations | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GM | PS | TC | GHB | SBP | DC | DP | EB | PB | SB | ZP | AD | |
| N | 35 | 30 | 30 | 20 | 25 | 23 | 30 | 20 | 20 | 20 | 30 | 30 |
| nSSR | ||||||||||||
| HO | 0.25 (0.06) | 0.32 (0.07) | 0.31 (0.05) | 0.53 (0.06) | 0.50 (0.05) | 0.49 (0.06) | 0.54 (0.05) | 0.50 (0.07) | 0.56 (0.06) | 0.51 (0.05) | 0.56 (0.05) | 0.50 (0.5) |
| HE | 0.30 (0.07) | 0.33 (0.07) | 0.33 (0.05) | 0.56 (0.05) | 0.55 (0.05) | 0.53 (0.06) | 0.57 (0.05) | 0.05 (0.07) | 0.56 (0.05) | 0.52 (0.04) | 0.55 (0.05) | 0.54 (0.04) |
| FIS | 0.14 (0.05) | 0.02 (0.04) | 0.03 (0.05) | 0.05 (0.06) | 0.09 (0.05) | 0.08 (0.05) | 0.04 (0.03) | −0.02 (0.04) | 0.01 (0.02) | 0.01 (0.04) | −0.05 (0.3) | 0.06 (0.07) |
| I | 0.56 (0.18) | 0.55 (0.11) | 0.58 (0.1) | 1.07 (0.13) | 1.02 (0.11) | 1.01 (0.14) | 1.07 (0.13) | 0.96 (0.14) | 1.06 (0.14) | 1.02 (0.12) | 1.05 (0.12) | 0.92 (0.10) |
| % Poly Loci | 85.71 | 85.7 | 92.9 | 100 | 100 | 100 | 100 | 92.86 | 100 | 100 | 100 | 100 |
| AR | 2.66 | 2.32 | 2.54 | 3.97 | 3.75 | 3.92 | 3.99 | 3.66 | 4.07 | 4.21 | 4.19 | 3.12 |
| cpSSR | ||||||||||||
| NH | 2 P | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 3 | 3 | 1 |
| HR | 0.991 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 1.897 | 0 |
| GM | PS | TC | GHB | SBP | DC | DP | EB | PB | SB | ZP | AD | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GM | − | − | − | − | − | − | − | − | − | − | − | − |
| PS | 0.22 | − | − | − | − | − | − | − | − | − | − | − |
| TC | 0.15 | 0.12 | − | − | − | − | − | − | − | − | − | − |
| GHB | 0.26 | 0.25 | 0.22 | − | − | − | − | − | − | − | − | − |
| SBP | 0.26 | 0.25 | 0.23 | 0.05 | − | − | − | − | − | − | − | − |
| DC | 0.32 | 0.32 | 0.28 | 0.06 | 0.04 | − | − | − | − | − | − | − |
| DP | 0.27 | 0.27 | 0.25 | 0.03 | 0.03 | 0.03 | − | − | − | − | − | − |
| EB | 0.22 | 0.24 | 0.18 | 0.08 | 0.07 | 0.09 | 0.08 | − | − | − | − | − |
| PB | 0.29 | 0.28 | 0.26 | 0.06 | 0.04 | 0.05 | 0.05 | 0.11 | − | − | − | − |
| SB | 0.25 | 0.27 | 0.21 | 0.09 | 0.06 | 0.08 | 0.07 | 0.07 | 0.07 | − | − | − |
| ZP | 0.24 | 0.24 | 0.21 | 0.07 | 0.04 | 0.09 | 0.06 | 0.07 | 0.08 | 0.02 | − | − |
| AD | 0.30 | 0.25 | 0.23 | 0.12 | 0.12 | 0.14 | 0.12 | 0.17 | 0.13 | 0.13 | 0.13 | − |
| Surface | k | AIC | AICc | R2m | R2c | LL | Delta.AICc | Weight |
|---|---|---|---|---|---|---|---|---|
| SAR, SE, TPI | 10 | −166.71 | −264.71 | 0.28 | 0.68 | 87.36 | 0.00 | 1.00 |
| LC, SAR, SE | 11 | −167.23 | −241.23 | 0.30 | 0.68 | 87.62 | 23.48 | 0.00 |
| LC, SAR, TPI | 11 | −166.75 | −240.75 | 0.26 | 0.67 | 87.38 | 23.96 | 0.00 |
| LC, SE, TPI | 11 | −166.04 | −240.04 | 0.21 | 0.69 | 87.02 | 24.67 | 0.00 |
| Distance | 2 | −164.88 | −166.88 | 0.19 | 0.71 | 86.44 | 97.83 | 0.00 |
| Null | 1 | −159.82 | −163.25 | 0.00 | 0.67 | 82.91 | 101.46 | 0.00 |
| SAR | 4 | −166.90 | −156.90 | 0.29 | 0.68 | 87.45 | 107.82 | 0.00 |
| SE | 4 | −166.66 | −156.66 | 0.26 | 0.67 | 87.33 | 108.06 | 0.00 |
| TPI | 4 | −166.03 | −156.03 | 0.24 | 0.70 | 87.01 | 108.69 | 0.00 |
| LC | 5 | −168.28 | −146.28 | 0.52 | 0.77 | 88.14 | 118.44 | 0.00 |
| SAR, SE | 7 | −166.82 | −48.82 | 0.28 | 0.68 | 87.41 | 215.90 | 0.00 |
| SAR, TPI | 7 | −166.82 | −48.82 | 0.28 | 0.68 | 87.41 | 215.90 | 0.00 |
| SE, TPI | 7 | −166.30 | −48.30 | 0.24 | 0.68 | 87.15 | 216.41 | 0.00 |
| LC, SAR | 8 | −167.49 | Inf | 0.31 | 0.68 | 87.74 | Inf | 0.00 |
| LC, SE | 8 | −166.01 | Inf | 0.21 | 0.68 | 87.00 | Inf | 0.00 |
| LC, TPI | 8 | −165.88 | Inf | 0.18 | 0.69 | 86.94 | Inf | 0.00 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Leimbach-Maus, H.B.; McCluskey, E.M.; Locher, A.; Parks, S.R.; Partridge, C.G. Genetic Structure of Invasive Baby’s Breath (Gypsophila paniculata L.) Populations in a Michigan Dune System. Plants 2020, 9, 1123. https://doi.org/10.3390/plants9091123
Leimbach-Maus HB, McCluskey EM, Locher A, Parks SR, Partridge CG. Genetic Structure of Invasive Baby’s Breath (Gypsophila paniculata L.) Populations in a Michigan Dune System. Plants. 2020; 9(9):1123. https://doi.org/10.3390/plants9091123
Chicago/Turabian StyleLeimbach-Maus, Hailee B., Eric M. McCluskey, Alexandra Locher, Syndell R. Parks, and Charlyn G. Partridge. 2020. "Genetic Structure of Invasive Baby’s Breath (Gypsophila paniculata L.) Populations in a Michigan Dune System" Plants 9, no. 9: 1123. https://doi.org/10.3390/plants9091123





