Transcriptomic Analysis of a Susceptible African Maize Line to Fusarium verticillioides Infection
Abstract
1. Introduction
2. Results and Discussions
2.1. Plant Morphology and Fungal Quantification
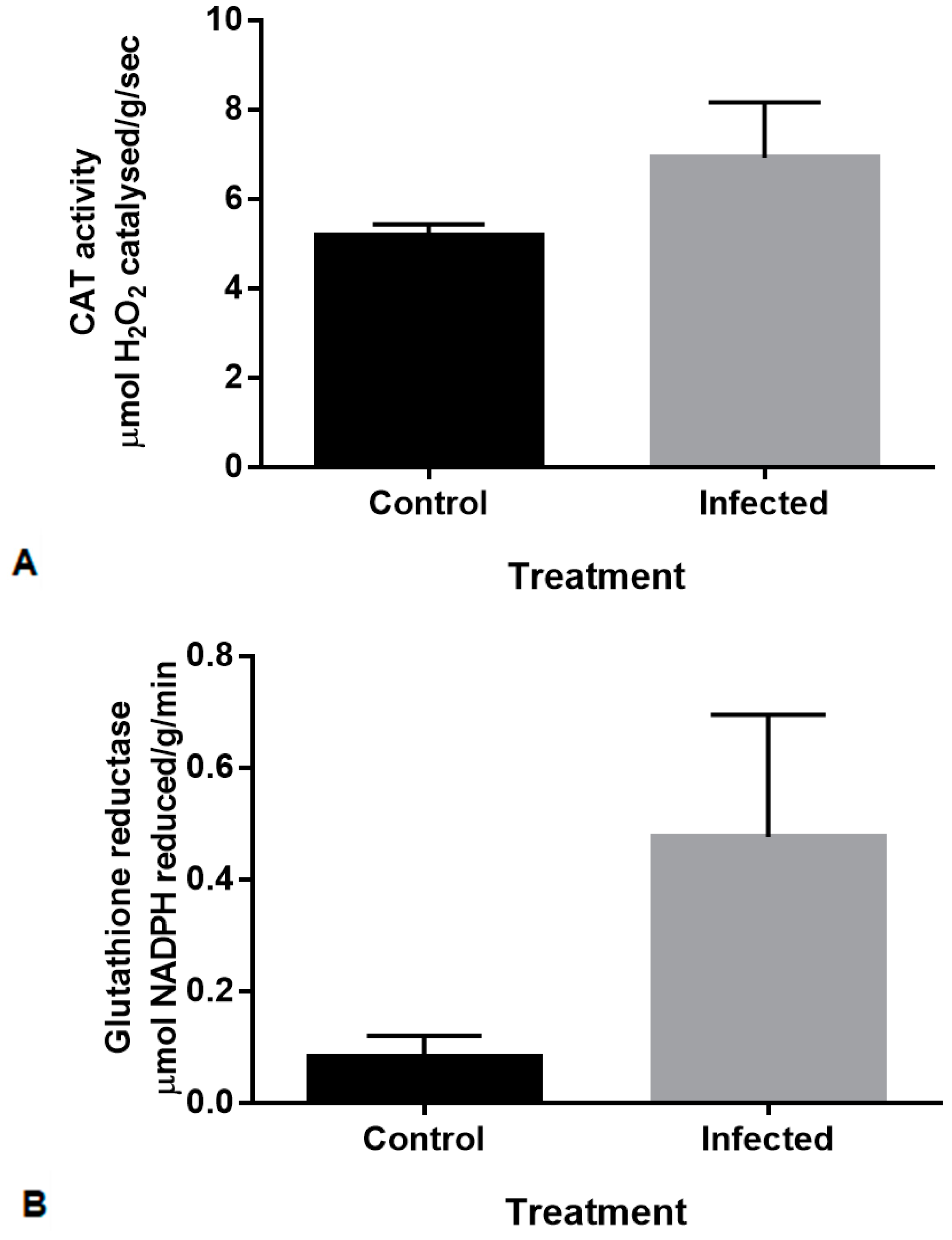
2.2. Production of Antioxidant Enzymes in Maize Shoots in Response to F. verticillioides Inoculation
2.3. Determining Differential Expression between Control and Infected Maize Shoots Using RNA-Sequencing
2.4. Gene Ontology Enrichment Analysis Reveals Possible F. verticillioides Induced Response Mechanisms in Maize
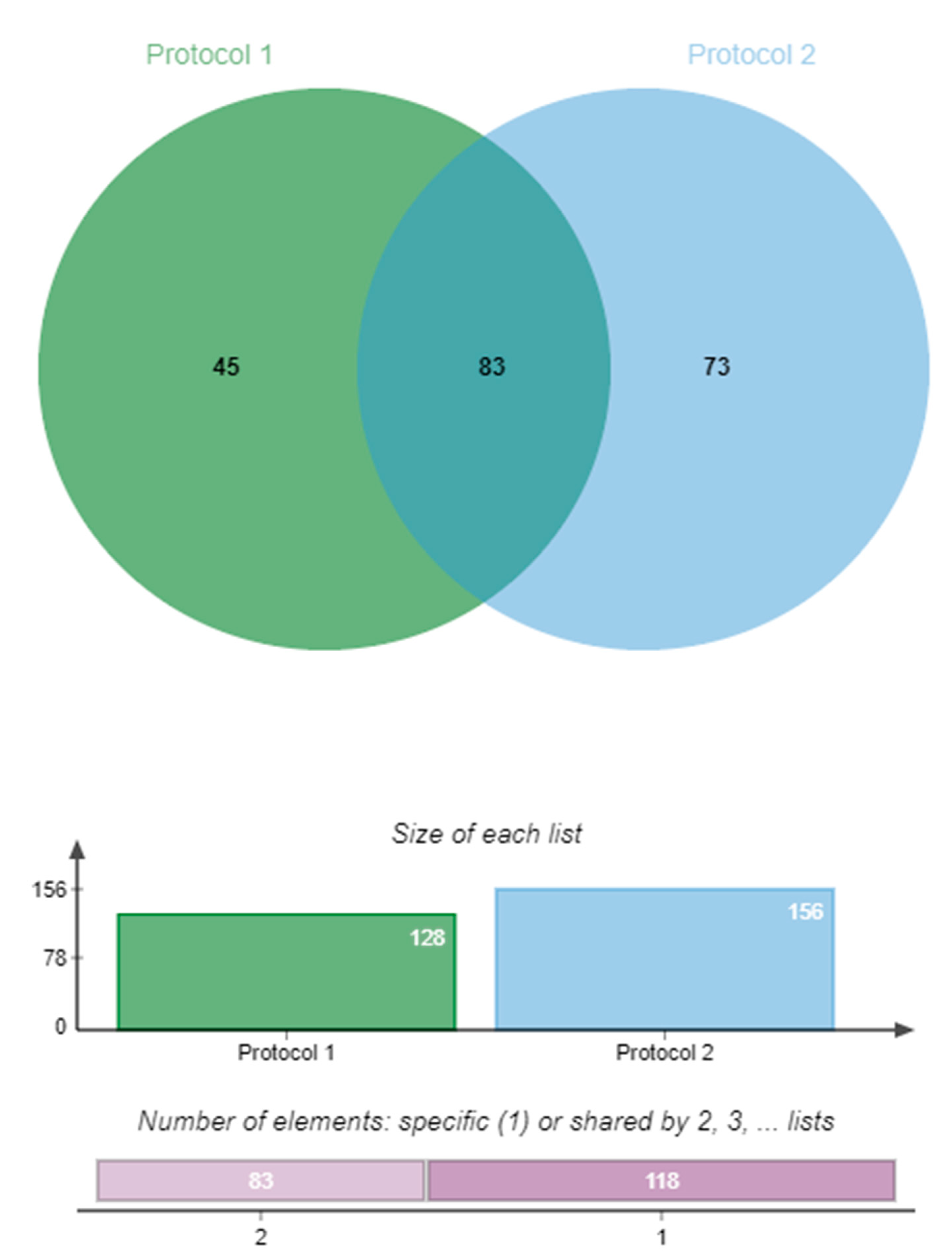
2.5. Comparing DEGs Against a Different Maize Line Infected with F. verticillioides
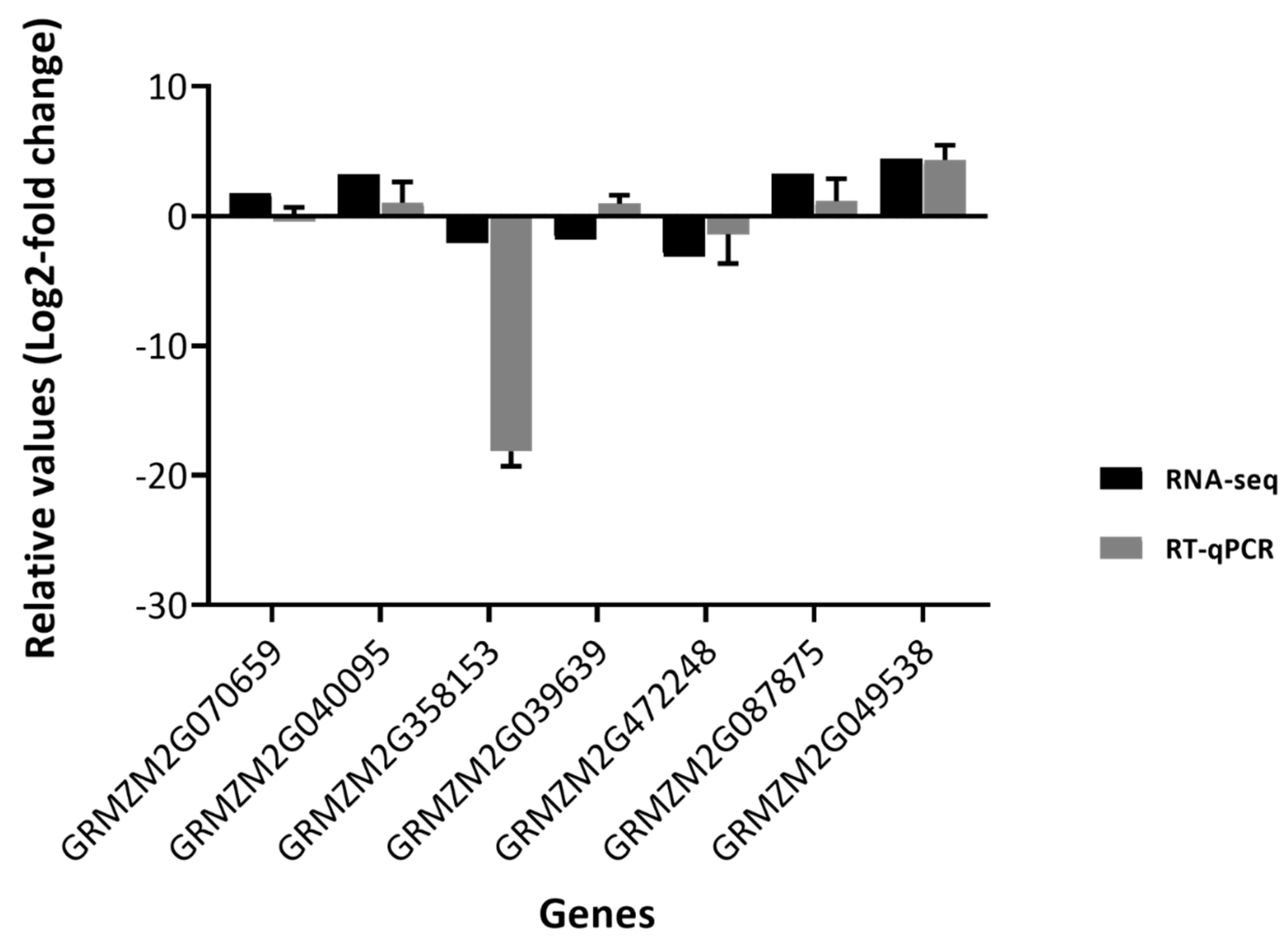
2.6. Validation of RNA-Sequencing by Quantitative Real-Time PCR
2.7. Analysis of Phytoalexin Accumulation
3. Conclusions
4. Materials and Methods
4.1. Fungi
4.2. Plant Growth, Inoculations and Morphology
4.3. RNA Extraction and RNA-Sequencing
4.4. RNA-Sequencing and Data Analysis
4.4.1. Analysis Using Protocol 1
4.4.2. Analysis Using Protocol 2
4.5. Gene Ontology Analysis Using agriGO
4.6. cDNA Synthesis and Quantitative Real-Time PCR
4.7. Phytoalexin Accumulation
4.8. Antioxidant Assays
4.8.1. Catalase (CAT, EC 1.11.1.6)
4.8.2. Glutathione Reductase (GR, EC 1.6.4.2)
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fandohan, P.; Hell, K.; Marasas, W.; Wingfield, M. Infection of maize by Fusarium species and contamination with fumonisin in Africa. Afr. J. Biotechnol. 2003, 2, 570–579. [Google Scholar]
- Rheeder, J.P.; Shephard, G.S.; Vismer, H.F.; Gerderblom, W.C.A. Guidelines on Mycotoxin Control in South Africa Foodstuffs: From the Application of the Hazard Analysis and Critical Control Point (HACCP) System to New National Mycotoxin Regulations. MRC Policy Brief. October 2009. Available online: www.mrc.ac.za/policybriefs/Mycotoxinguidelines.pdf (accessed on 10 March 2014).
- Small, I.; Flett, B.; Marasas, W.; McLeod, A.; Viljoen, A. Use of resistance elicitors to reduce Fusarium ear rot and fumonisin accumulation in maize. Crop Protect. 2012, 41, 10–16. [Google Scholar] [CrossRef]
- Kornher, L. Maize Markets in Eastern and Southern Africa (ESA) in the Context of Climate Change—Background Paper for the State of Agricultural Commodity Markets (SOCO) 2018; FAO: Rome, Italy, 2018. [Google Scholar]
- Du Plessis, J. Maize Production; Department of Agriculture: Pretoria, South Africa, 2003. [Google Scholar]
- Pereira, P.; Nesci, A.; Castillo, C.; Etcheverry, M. Field Studies on the Relationship between Fusarium verticillioides and Maize (Zea mays L.): Effect of Biocontrol Agents on Fungal Infection and Toxin Content of Grains at Harvest. Int. J. Agron. 2011, 1–7. [Google Scholar] [CrossRef]
- Aiyaz, M.; Divakara, S.T.; Nayaka, S.C.; Hariprasad, P.; Niranjana, S.R. Application of beneficial rhizospheric microbes for the mitigation of seed-borne mycotoxigenic fungal infection and mycotoxins in maize. Biocontr. Sci. Technol. 2015, 25, 1105–1119. [Google Scholar] [CrossRef]
- Munkvold, G.P.; Desjardins, A.E. Fumonisins in Maize: Can We Reduce Their Occurrence? Plant Dis. 1997, 81, 556–565. [Google Scholar] [CrossRef]
- Pechanova, O.; Pechan, T. Maize-Pathogen Interactions: An Ongoing Combat from a Proteomics Perspective. Int. J. Mol. Sci. 2015, 16, 28429–28448. [Google Scholar] [CrossRef]
- Venturini, G.; Toffolatti, S.; Assante, G.; Babazadeh, L.; Campia, P.; Fasoli, E.; Salomoni, D.; Vercesi, A. The influence of flavonoids in maize pericarp on Fusarium ear rot symptoms and fumonisin accumulation under field conditions. Plant Pathol. 2015, 64, 671–679. [Google Scholar] [CrossRef]
- Rose, L.J.; Okoth, S.; Beukes, I.; Ouko, A.; Mouton, M.; Flett, B.C.; Makumbi, D.; Viljoen, A. Determining resistance to Fusarium verticillioides and fumonisin accumulation in African maize inbred lines resistant to Aspergillus flavus and aflatoxins. Euphytica 2017, 213, 93. [Google Scholar] [CrossRef]
- Van Rensburg, B.J.; McLaren, N.; Flett, B.; Schoeman, A. Fumonisin producing Fusarium spp. and fumonisin contamination in commercial South African maize. Eur. J. Plant Pathol. 2015, 141, 491–504. [Google Scholar] [CrossRef]
- Marasas, W. Discovery and occurrence of the fumonisins: A historical perspective. Environ. Health Perspect. 2001, 109, 239–243. [Google Scholar]
- Zorzete, P.; Castro, R.S.; Pozzi, C.R.; Israel, A.L.M.; Fonseca, H.; Yanaguibashi, G.; Corrêa, B. Relative populations and toxin production by Aspergillus flavus and Fusarium verticillioides in artificially inoculated corn at various stages of development under field conditions. J. Sci. Food Agric. 2008, 88, 48–55. [Google Scholar] [CrossRef]
- Lanubile, A.; Pasini, L.; Marocco, A. Differential gene expression in kernels and silks of maize lines with contrasting levels of ear rot resistance after Fusarium verticillioides infection. J. Plant Physiol. 2010, 167, 1398–1406. [Google Scholar] [PubMed]
- Yang, I.S.; Kim, S. Analysis of Whole Transcriptome Sequencing Data: Workflow and Software. Genomote Inform. 2015, 13, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Cox, M.P.; Eaton, C.J.; Scott, D.B. Exploring molecular signaling in plant-fungal symbioses using high throughput RNA sequencing. Plant Signal. Behav. 2010, 5, 1353–1358. [Google Scholar] [CrossRef]
- Marioni, J.C.; Mason, C.E.; Mane, S.M.; Stephens, M.; Gilad, Y. RNA-seq: An assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 2008, 18, 1509–1517. [Google Scholar] [CrossRef] [PubMed]
- Sekhon, R.S.; Briskine, R.; Hirsch, C.N.; Myers, C.L.; Springer, N.M.; Buell, C.R.; de Leon, N.; Kaeppler, S.M. Maize Gene Atlas Developed by RNA Sequencing and Comparative Evaluation of Transcriptomes Based on RNA Sequencing and Microarrays. PLoS ONE 2013, 8, e61005. [Google Scholar] [CrossRef] [PubMed]
- Seyednasrollah, F.; Laiho, A.; Elo, L.L. Comparison of software packages for detecting differential expression in RNA-seq studies. Brief. Bioinf. 2015, 16, 59–70. [Google Scholar] [CrossRef]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef]
- Lanubile, A.; Ferrarini, A.; Maschietto, V.; Delledonne, M.; Marocco, A.; Bellin, D. Functional genomic analysis of constitutive and inducible defense responses to Fusarium verticillioides infection in maize genotypes with contrasting ear rot resistance. BMC Genom. 2014, 15, 710. [Google Scholar] [CrossRef]
- Veenstra, A.; Moola, N.; Wighard, S.; Korsman, J.; Christensen, S.A.; Rafudeen, M.S.; Murray, S.L. Kauralexins and zealexins accumulate in sub-tropical maize lines and play a role in seedling resistance to Fusarium verticillioides. Eur. J. Plant Pathol. 2019, 153, 223–237. [Google Scholar] [CrossRef]
- Wighard, S. Characterisation of a Maize Mutant Deficient in Antifungal Kauralexin Accumulation. Master’s Thesis, University of Cape Town, Cape Town, South Africa, 2017. [Google Scholar]
- Ding, Y.; Murphy, K.M.; Poretsky, E.; Mafu, S.; Yang, B.; Char, S.N.; Christensen, S.A.; Saldivar, E.; Wu, M.; Wang, Q.; et al. Multiple genes recruited from hormone pathways partition maize diterpenoid defences. Nat. Plants 2019, 5, 1043–1056. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Ren, F.; Lu, X.; Mao, H.; Xu, M.; Degenhardt, J.; Peters, R.J.; Wang, Q. A Tandem Array of ent-Kaurene Synthases in Maize with Roles in Gibberellin and More Specialized Metabolism. Plant Physiol. 2016, 170, 742–751. [Google Scholar] [CrossRef] [PubMed]
- Lanubile, A.; Bernardi, J.; Battilani, P.; Logrieco, A.; Marocco, A. Resistant and susceptible maize genotypes activate different transcriptional responses against Fusarium verticillioides. Physiol. Mol. Plant Pathol. 2012, 77, 223–237. [Google Scholar] [CrossRef]
- Maschietto, V.; Lanubile, A.; Leonardis, S.D.; Marocco, A.; Paciolla, C. Constitutive expression of pathogenesis-related proteins and antioxydant enzyme activities triggers maize resistance towards Fusarium verticillioides. J. Plant Physiol. 2016, 200, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhou, Z.; Gao, J.; Wu, Y.; Xia, Z.; Zhang, H.; Wu, J. The Mechanisms of Maize Resistance to Fusarium verticillioides by comprehensive analysis of RNA-seq Data. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef]
- Okello, D.; Manna, R.; Imanyowoha, J.; Pixley, K.; Edema, R. Agronomic performance and breeding potential of selected inbred lines for improvement of protein quality of adapted Ugandan maize germplasm. Afr. Crop Sci. J. 2006, 14, 37–47. [Google Scholar]
- Nicolaisen, M.; Supronienė, S.; Nielsen, L.K.; Lazzaro, I.; Spliid, N.H.; Justesen, A.F. Real-time PCR for quantification of eleven individual Fusarium species in cereals. J. Microbiol. Methods 2009, 76, 234–240. [Google Scholar] [CrossRef]
- Lanubile, A.; Maschietto, V.; De Leonardis, S.; Battilani, P.; Paciolla, C.; Marocco, A. Defense Responses to Mycotoxin-Producing Fungi Fusarium proliferatum, F. subglutinans, and Aspergillus flavus in Kernels of Susceptible and Resistant Maize Genotypes. Mol. Plant Microbe Interact. 2015, 28, 546–557. [Google Scholar] [CrossRef]
- Magbanua, Z.V.; De Moraes, C.M.; Brooks, T.D.; Williams, W.P.; Luthe, D.S. Is Catalase Activity One of the Factors Associated with Maize Resistance to Aspergillus flavus? Mol. Plant Microbe Interact. 2007, 20, 697–706. [Google Scholar] [CrossRef]
- Mhamdi, A.; Queval, G.; Chaouch, S.; Vanderauwera, S.; Van Breusegem, F.; Noctor, G. Catalase function in plants: A focus on Arabidopsis mutants as stress-mimic models. J. Exp. Bot. 2010, 61, 4197–4220. [Google Scholar] [CrossRef]
- Redinbaugh, M.G.; Sabre, M.; Scandalios, J.G. The distribution of catalase activity, isozyme protein, and transcript in the tissues of the developing maize seedling. Plant Physiol. 1990, 92, 375–380. [Google Scholar] [CrossRef] [PubMed]
- Doulis, A.G.; Debian, N.; Kingston-Smith, A.H.; Foyer, C.H. Differential Localization of Antioxidants in Maize Leaves. Plant Physiol. 1997, 114, 1031–1037. [Google Scholar] [CrossRef] [PubMed]
- Lanubile, A.; Bernardi, J.; Marocco, A.; Logrieco, A.; Paciolla, C. Differential activation of defense genes and enzymes in maize genotypes with contrasting levels of resistance to Fusarium verticillioides. Environ. Exp. Bot. 2012, 78, 39–46. [Google Scholar] [CrossRef]
- Kumar, M.; Yadav, V.; Tuteja, N.; Johri, A.K. Antioxidant enzyme activities in maize plants colonized with Piriformospora indica. Microbiology 2009, 155, 780–790. [Google Scholar] [CrossRef] [PubMed]
- Bardou, P.; Mariette, J.; Escudié, F.; Djemiel, C.; Klopp, C. jvenn: An interactive Venn diagram viewer. BMC Bioinform. 2014, 15, 293. [Google Scholar] [CrossRef] [PubMed]
- Du, Z.; Zhou, X.; Ling, Y.; Zhang, Z.; Su, Z. agriGO: A GO analysis toolkit for the agricultural community. Nucleic Acids Res. 2010, 38, W64–W70. [Google Scholar] [CrossRef] [PubMed]
- Carbon, S.; Ireland, A.; Mungall, C.J.; Shu, S.; Marshall, B.; Lewis, S. AmiGO: Online access to ontology and annotation data. Bioinformatics 2008, 25, 288–289. [Google Scholar] [CrossRef]
- Kawaide, H.; Sassa, T.; Kamiya, Y. Functional Analysis of the Two Interacting Cyclase Domains in ent-Kaurene Synthase from the Fungus Phaeosphaeria sp. L487 and a Comparison with Cyclases from Higher Plants. J. Biol. Chem. 2000, 275, 2276–2280. [Google Scholar] [CrossRef]
- Schmelz, E.A.; Kaplan, F.; Huffaker, A.; Dafoe, N.J.; Vaughan, M.M.; Ni, X.; Rocca, J.R.; Alborn, H.T.; Teal, P.E. Identity, regulation, and activity of inducible diterpenoid phytoalexins in maize. Proc. Natl. Acad. Sci. USA 2011, 108, 5455–5460. [Google Scholar] [CrossRef]
- Hedden, P.; Thomas, S. Gibberellin biosynthesis and its regulation. Biochem. J. 2012, 444, 11–25. [Google Scholar] [CrossRef]
- Schmelz, E.A.; Huffaker, A.; Sims, J.W.; Christensen, S.A.; Lu, X.; Okada, K.; Peters, R.J. Biosynthesis, elicitation and roles of monocot terpenoid phytoalexins. Plant J. 2014, 79, 659–678. [Google Scholar] [CrossRef] [PubMed]
- Vaughan, M.M.; Christensen, S.; Schmelz, E.A.; Huffaker, A.; Mcauslane, H.J.; Alborn, H.T.; Romero, M.; Allen, L.H.; Teal, P.E.A. Accumulation of terpenoid phytoalexins in maize roots is associated with drought tolerance. Plant Cell Environ. 2015, 38, 2195–2207. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Liu, W.; Yao, Y.; Wei, Y.; Chan, Z. Alcohol dehydrogenase 1 (ADH1) confers both abiotic and biotic stress resistance in Arabidopsis. Plant Sci. 2017, 262, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Mauch-Mani, B.; Mauch, F. The role of abscisic acid in plant–pathogen interactions. Curr. Opin. Plant Biol. 2005, 8, 409–414. [Google Scholar] [CrossRef]
- Berrocal-Lobo, M.; Molina, A. Ethylene Response Factor 1 Mediates Arabidopsis Resistance to the Soilborne Fungus Fusarium oxysporum. Mol. Plant Microbe Interact. 2004, 17, 763–770. [Google Scholar] [CrossRef]
- Chen, Y.C.; Wong, C.L.; Muzzi, F.; Vlaardingerbroek, I.; Kidd, B.N.; Schenk, P.M. Root defense analysis against Fusarium oxysporum reveals new regulators to confer resistance. Sci. Rep. 2014, 4, 5584. [Google Scholar] [CrossRef]
- Handa, Y.; Nishide, H.; Takeda, N.; Suzuki, Y.; Kawaguchi, M.; Saito, K. RNA-seq Transcriptional Profiling of an Arbuscular Mycorrhiza Provides Insights into Regulated and Coordinated Gene Expression in Lotus japonicus and Rhizophagus irregularis. Plant Cell Physiol. 2015, 56, 1490–1511. [Google Scholar] [CrossRef]
- Campos-Bermudez, V.A.; Fauguel, C.M.; Tronconi, M.A.; Casati, P.; Presello, D.A.; Andreo, C.S. Transcriptional and metabolic changes associated to the infection by Fusarium verticillioides in maize inbreds with contrasting ear rot resistance. PLoS ONE 2013, 8, e61580. [Google Scholar] [CrossRef]
- Bluhm, B.H.; Woloshuk, C.P. Amylopectin induces fumonisin B1 production by Fusarium verticillioides during colonization of maize kernels. Mol. Plant Microbe Interact. 2005, 18, 1333–1339. [Google Scholar] [CrossRef]
- Shetty, N.P.; Mehrabi, R.; Lütken, H.; Haldrup, A.; Kema, G.H.; Collinge, D.B.; Jørgensen, H.J.L. Role of hydrogen peroxide during the interaction between the hemibiotrophic fungal pathogen Septoria tritici and wheat. New Phytol. 2007, 174, 637–647. [Google Scholar] [CrossRef]
- Ma, K.-W.; Ma, W. Phytohormone pathways as targets of pathogens to facilitate infection. Plant Mol. Biol. 2016, 91, 713–725. [Google Scholar] [CrossRef] [PubMed]
- Christensen, S.A.; Sims, J.; Vaughan, M.M.; Hunter, C.; Block, A.; Willett, D.; Alborn, H.T.; Huffaker, A.; Schmelz, E.A. Commercial hybrids and mutant genotypes reveal complex protective roles for inducible terpenoid defenses in maize. J. Exp. Bot. 2018, 69, 1693–1705. [Google Scholar] [CrossRef] [PubMed]
- Mafu, S.; Ding, Y.; Murphy, K.M.; Yaacoobi, O.; Addison, J.B.; Wang, Q.; Shen, Z.; Briggs, S.P.; Bohlmann, J.; Castro-Falcon, G.; et al. Discovery, Biosynthesis and Stress-Related Accumulation of Dolabradiene-Derived Defenses in Maize. Plant Physiol. 2018, 176, 2677–2690. [Google Scholar] [CrossRef] [PubMed]
- Block, A.K.; Vaughan, M.M.; Schmelz, E.A.; Christensen, S.A. Biosynthesis and function of terpenoid defense compounds in maize (Zea mays). Planta 2019, 249, 21–30. [Google Scholar] [CrossRef]
- Oren, L.; Ezrati, S.; Cohen, D.; Sharon, A. Early events in the Fusarium verticillioides-maize interaction characterized by using a green fluorescent protein-expressing transgenic isolate. Appl. Environ. Microbiol. 2003, 69, 1695–1701. [Google Scholar] [CrossRef]
- Williams, J. DNA Subway–An Educational Bioinformatics Platform for Gene and Genome Analysis: DNA Barcoding, and RNA-Seq. In Proceedings of the 10th World Congress on Genetics Applied to Livestock Production, Vancouver, BC, Canada, 17–22 August 2014; ASAS: Champaign, IL, USA, 2014. [Google Scholar]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 16 November 2016).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28. [Google Scholar] [CrossRef]
- Bailly, C.; Benamar, A.; Corbineau, F.; Come, D. Changes in malondialdehyde content and in superoxide dismutase, catalase and glutathione reductase activities in sunflower seeds as related to deterioration during accelerated aging. Physiol. Plant. 1996, 97, 104–110. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Claiborne, A. Catalase activity. In CRC Handbook of Methods for Oxygen Radical Research; CRC Press: Boca Raton, FL, USA, 1985; Volume 1, pp. 283–284. [Google Scholar]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Manoli, A.; Sturaro, A.; Trevisan, S.; Quaggiotti, S.; Nonis, A. Evaluation of candidate reference genes for qPCR in maize. J. Plant Physiol. 2012, 169, 807–815. [Google Scholar] [CrossRef]
- Ma, J.; Morrow, D.J.; Fernandes, J.; Walbot, V. Comparative profiling of the sense and antisense transcriptome of maize lines. Genome Biol. 2006, 7, R22. [Google Scholar] [CrossRef] [PubMed]
- Veenstra, A. Evaluation of Southern African Maize Germplasm for Phytoalexin Accumulation Following Inoculation by Fusarium verticillioides. Master’s Thesis, University of Cape Town, Cape Town, South Africa, 2017. [Google Scholar]
- Schnee, C.; Köllner, T.G.; Gershenzon, J.; Degenhardt, J. The Maize Gene terpene synthase 1 Encodes a Sesquiterpene Synthase Catalyzing the Formation of (E)-beta -Farnesene, (E)-Nerolidol, and (E,E)-Farnesol after Herbivore Damage. Am. Soc. Plant Biol. 2002, 130, 2049–2060. [Google Scholar]

| Gene Stable ID | Average Log2 Fold-Change | Ensembl/NCBI Gene Description | ¥ Enriched GO-Term |
|---|---|---|---|
| GRMZM2G049538 | 4.56 | Acyclic sesquiterpene synthase/ent-Kaurene synthase B | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G119975 | 3.98 | Uncharacterised LOC103646336 | |
| GRMZM2G029219 | 3.45 | Carbohydrate transporter/sugar porter/transporter | |
| GRMZM5G874955 | 3.33 | Uncharacterised protein | |
| GRMZM2G062724 | 3.27 | Uncharacterised protein | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G026922 | 3.27 | Hypothetical protein | |
| GRMZM2G117971 | 3.25 | Uncharacterised protein | |
| GRMZM2G087875 | 3.20 | Putative cytochrome P450 superfamily protein; Uncharacterised protein | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G143139 | 3.17 | N/A | |
| GRMZM2G149422 | 3.07 | Hypothetical protein | |
| GRMZM2G154523 | 3.06 | Patatin T5; Uncharacterised protein | |
| GRMZM2G098346 | 3.04 | Alcohol dehydrogenase 2 | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM5G892675 | 3.01 | Uncharacterised protein | |
| GRMZM2G137861 | 2.99 | Wall-associated receptor kinase 2-like | |
| GRMZM2G443728 | 2.95 | Potassium transporter 10 | |
| GRMZM2G070011 | 2.85 | Uncharacterised protein; Vignain | |
| GRMZM2G006973 | 2.73 | Uncharacterised protein | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G036464 | 2.72 | Glutamine synthetase root isozyme 4 | |
| GRMZM2G115451 | 2.63 | Uncharacterised protein | |
| GRMZM2G178546 | 2.61 | Trehalose-phosphate phosphatase | |
| GRMZM2G099049 | 2.56 | N/A | |
| GRMZM2G063431 | 2.53 | N/A | |
| GRMZM2G427815 | 2.51 | Uncharacterised protein | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G062531 | 2.50 | Uncharacterised protein | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G093826 | 2.49 | Potassium high-affinity transporter | |
| GRMZM2G110504 | 2.46 | Uncharacterised LOC100278648 | |
| GRMZM2G007151 | 2.42 | Uncharacterised protein | |
| GRMZM2G130173 | 2.41 | Metallothionein-like protein type 2; Uncharacterised protein | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G477503 | 2.36 | Uncharacterised protein | |
| AC217947.4_FG002.2 | 2.24 | N/A | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G026470 | 2.22 | Soluble inorganic pyrophosphatase; Uncharacterised protein | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G091456 | 2.21 | Putative Uncharacterised protein | |
| GRMZM2G366681 | 2.11 | Hypothetical protein | |
| GRMZM2G034152 | 2.07 | Polyamine oxidase | |
| GRMZM2G130149 | 2.07 | Uncharacterised protein | |
| GRMZM2G144097 | 2.04 | Uncharacterised protein | |
| GRMZM2G125669 | 2.03 | Alternative oxidase | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G144083 | 2.02 | Putative ATP dependent copper transporter | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G034302 | 2.01 | Uncharacterised protein | |
| GRMZM2G036217 | 1.91 | Uncharacterised protein | |
| GRMZM2G116079 | 1.86 | Uncharacterised protein | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G076537 | 1.82 | Polynucleotidyl transferase, ribonuclease H-like superfamily protein | |
| GRMZM2G176433 | 1.80 | Putative Uncharacterised protein | |
| GRMZM2G008247 | 1.78 | Beta-glucosidase2 | |
| GRMZM2G070659 | 1.77 | Hypersensitive-induced response protein | |
| GRMZM2G147243 | 1.72 | IAA17-auxin-responsive Aux/IAA family member; Uncharacterised protein | |
| GRMZM2G099767 | 1.63 | ATMAP70-2 | |
| GRMZM2G057823 | 1.55 | Fructose-bisphosphate aldolase, cytoplasmic isozyme | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G040369 | 1.54 | Elongation factor 2 | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G168552 | 1.53 | Bundle sheath cell specific protein 1 | |
| GRMZM2G020146 | 1.51 | Uncharacterised protein | |
| GRMZM2G473001 | 1.48 | Phosphoenolpyruvate carboxylase 2 | |
| GRMZM2G113332 | 1.42 | Uncharacterised protein | GO: 0043167 |
| GO: 0046872 | |||
| GO: 0043169 | |||
| GRMZM2G141353 | 1.41 | Uncharacterised LOC100194210 | |
| GRMZM2G146004 | −FC | Uncharacterised protein | GO: 0050896 |
| AC214438.3_FG002.1 | −FC | N/A | |
| GRMZM2G177077 | −1.40 | Glucose-6-phosphate 1-dehydrogenase | GO: 0005975 |
| GRMZM2G047474 | −1.42 | TLD-domain containing nucleolar protein | |
| GRMZM2G366659 | −1.49 | Putative trehalose phosphatase/synthase family protein | GO: 0005975 |
| GRMZM5G870170 | −1.57 | MATE1 | GO: 0050896 |
| GO: 0042221 | |||
| GRMZM2G024733 | −1.57 | Uncharacterised LOC100304285 | |
| GRMZM2G478568 | −1.63 | Nicotianamine synthase 3 | |
| GRMZM2G154278 | −1.68 | Pre-mRNA-splicing factor cwc15 | |
| GRMZM2G103812 | −1.70 | Uncharacterised protein | GO: 0050896 |
| GO: 0042221 | |||
| GRMZM2G121264 | −1.74 | Uncharacterised protein | |
| GRMZM2G173085 | −1.85 | Lipase/lipooxygenase, PLAT/LH2 family protein | |
| GRMZM2G079381 | −1.86 | Ferredoxin-nitrite reductase, chloroplastic | GO: 0050896 |
| GO: 0042221 | |||
| GRMZM2G053669 | −1.99 | Asparagine synthetase | |
| GRMZM2G147687 | −2.02 | Uncharacterised protein | GO: 0005975 |
| GRMZM2G358153 | −2.09 | Chitinase 1; Uncharacterised protein | GO: 0005975 |
| GRMZM2G181081 | −2.21 | CIPK-like protein 1 | GO: 0050896 |
| GO: 0042221 | |||
| GRMZM2G422955 | −2.26 | N/A | |
| GRMZM2G097641 | −2.27 | Sucrose-phosphatase 2 | GO: 0005975 |
| GO: 0050896 | |||
| GO: 0042221 | |||
| GRMZM2G078472 | −2.32 | Asparagine synthetase | |
| GRMZM2G124495 | −2.38 | Putative MYB DNA-binding domain superfamily protein; Transfactor; Uncharacterised protein | |
| GRMZM2G058612 | −2.39 | F-box/LRR-repeat protein 3-like | |
| GRMZM2G125775 | −2.40 | AN17 | GO: 0050896 |
| GO: 0042221 | |||
| GRMZM2G133675 | −2.65 | Putative HLH DNA-binding domain superfamily protein; Uncharacterised protein | |
| GRMZM2G004161 | −2.92 | Uncharacterised protein | GO: 0050896 |
| GO: 0042221 | |||
| GRMZM2G472248 | −3.17 | Protein induced upon tuberization | GO: 0050896 |
| GRMZM2G176430 | −3.19 | Uncharacterised protein | |
| GRMZM2G468111 | −3.58 | Uncharacterised LOC100277849 | |
| GRMZM2G070172 | −3.79 | Uncharacterised protein | GO: 0005975 |
| GO: 0050896 | |||
| GO: 0042221 |
| Up-Regulated Genes | ||||||
|---|---|---|---|---|---|---|
| GO Term | Ontology 1 | Description | Number in Input List | Number in BG/Ref | p-Value 2 | FDR |
| GO:0043169 | F | cation binding | 16 /45 | 4135/25864 | 0.0011 * | 0.028 |
| GO:0043167 | F | ion binding | 16 /45 | 4136/25864 | 0.0011 * | 0.028 |
| GO:0046872 | F | metal ion binding | 16 /45 | 4124/25864 | 0.0011 * | 0.028 |
| Down-regulated genes | ||||||
| GO:0050896 | P | response to stimulus | 10/22 | 3551/25864 | 0.00032 * | 0.0059 |
| GO:0042221 | P | response to chemical stimulus | 8/22 | 2052/25864 | 0.00018 * | 0.0059 |
| GO:0005975 | P | carbohydrate metabolic process | 6/22 | 1246/25864 | 0.00048 * | 0.0060 |
| Gene Stable ID | Ensembl/NCBI Gene Description | Susceptible/Resistant/Common 1 | Functional Category 2 |
|---|---|---|---|
| GRMZM2G125669 | Alternative oxidase | Common | Response to stress |
| GRMZM2G093826 | Potassium high-affinity transporter | Common | Transport |
| GRMZM5G874955 | Uncharacterised protein | Common | Transport |
| GRMZM2G029219 | Carbohydrate transporter/sugar porter/transporter | Common | Transport |
| GRMZM2G036464 | Glutamine synthetase root isozyme 4 | Common | Metabolic process |
| GRMZM2G008247 | Beta-glucosidase 2 | Common | Cell wall |
| GRMZM2G034152 | Polyamine oxidase | Common | Metabolic process |
| GRMZM2G427815 | Uncharacterised protein | Common | Resistance |
| GRMZM5G892675 | Uncharacterised protein | Common | Transport |
| GRMZM2G087875 | Putative cytochrome P450 superfamily protein | Common | Metabolic process |
| GRMZM2G020146 | Uncharacterised protein | Resistant | Metabolic process |
| GRMZM2G130173 | Metallothionein-like protein type 2; Uncharacterised protein | Resistant | Unknown function |
| GRMZM2G026470 | Soluble inorganic pyrophosphatase | Resistant | Metabolic process |
| GRMZM2G062724 | Uncharacterised protein | Resistant | Signal transduction |
| GRMZM2G178546 | Trehalose-phosphate phosphatase | Susceptible | Metabolic process |
| GRMZM2G007151 | Uncharacterised protein | Susceptible | Cell component |
| GRMZM2G119975 | Uncharacterised LOC103646336 | Susceptible | Metabolic process |
| Metabolite | ¥ Logged Average Metabolite Accumulation | Standard Deviation | p-Value (Control/Infected) | |
|---|---|---|---|---|
| Control | Infected | |||
| KA1 | 1.42 | 2.07 | 0.59 | 0.09 |
| KA2 | 1.68 | 1.60 | 1.11 | 0.90 |
| KA3 | 1.90 | 2.89 | 0.67 | 0.03 * |
| KB1 | 1.68 | 2.60 | 0.54 | 0.02 * |
| KB2 | 1.60 | 2.51 | 0.90 | 0.11 |
| KB3 | 1.78 | 3.06 | 0.93 | 0.04 * |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lambarey, H.; Moola, N.; Veenstra, A.; Murray, S.; Suhail Rafudeen, M. Transcriptomic Analysis of a Susceptible African Maize Line to Fusarium verticillioides Infection. Plants 2020, 9, 1112. https://doi.org/10.3390/plants9091112
Lambarey H, Moola N, Veenstra A, Murray S, Suhail Rafudeen M. Transcriptomic Analysis of a Susceptible African Maize Line to Fusarium verticillioides Infection. Plants. 2020; 9(9):1112. https://doi.org/10.3390/plants9091112
Chicago/Turabian StyleLambarey, Humaira, Naadirah Moola, Amy Veenstra, Shane Murray, and Mohamed Suhail Rafudeen. 2020. "Transcriptomic Analysis of a Susceptible African Maize Line to Fusarium verticillioides Infection" Plants 9, no. 9: 1112. https://doi.org/10.3390/plants9091112
APA StyleLambarey, H., Moola, N., Veenstra, A., Murray, S., & Suhail Rafudeen, M. (2020). Transcriptomic Analysis of a Susceptible African Maize Line to Fusarium verticillioides Infection. Plants, 9(9), 1112. https://doi.org/10.3390/plants9091112






