New Insights into Evolution of Plant Heat Shock Factors (Hsfs) and Expression Analysis of Tea Genes in Response to Abiotic Stresses
Abstract
1. Introduction
2. Materials and Methods
2.1. Genome-Wide Identification of Plant Genes Encoding Hsf Proteins
2.2. Alignment and Phylogenetic Analysis
2.3. RNA-Seq Data Analysis
2.4. Plant Materials and Treatments
2.5. Expression Analysis of Tea Hsfs Using qRT-PCRs
3. Result
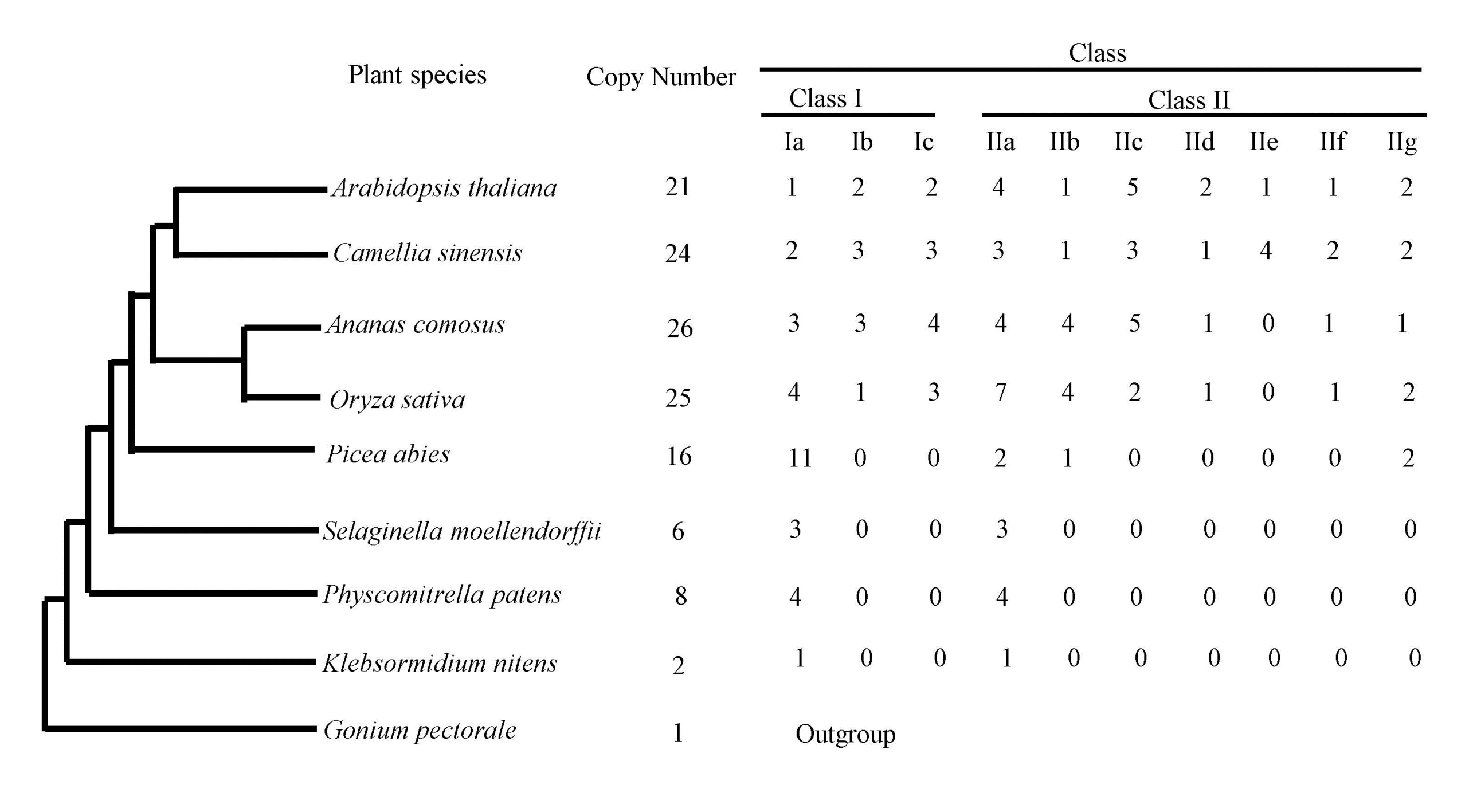
3.1. The Tea and Other Plant Genomes Encode Different Numbers of the Hsf Family Members
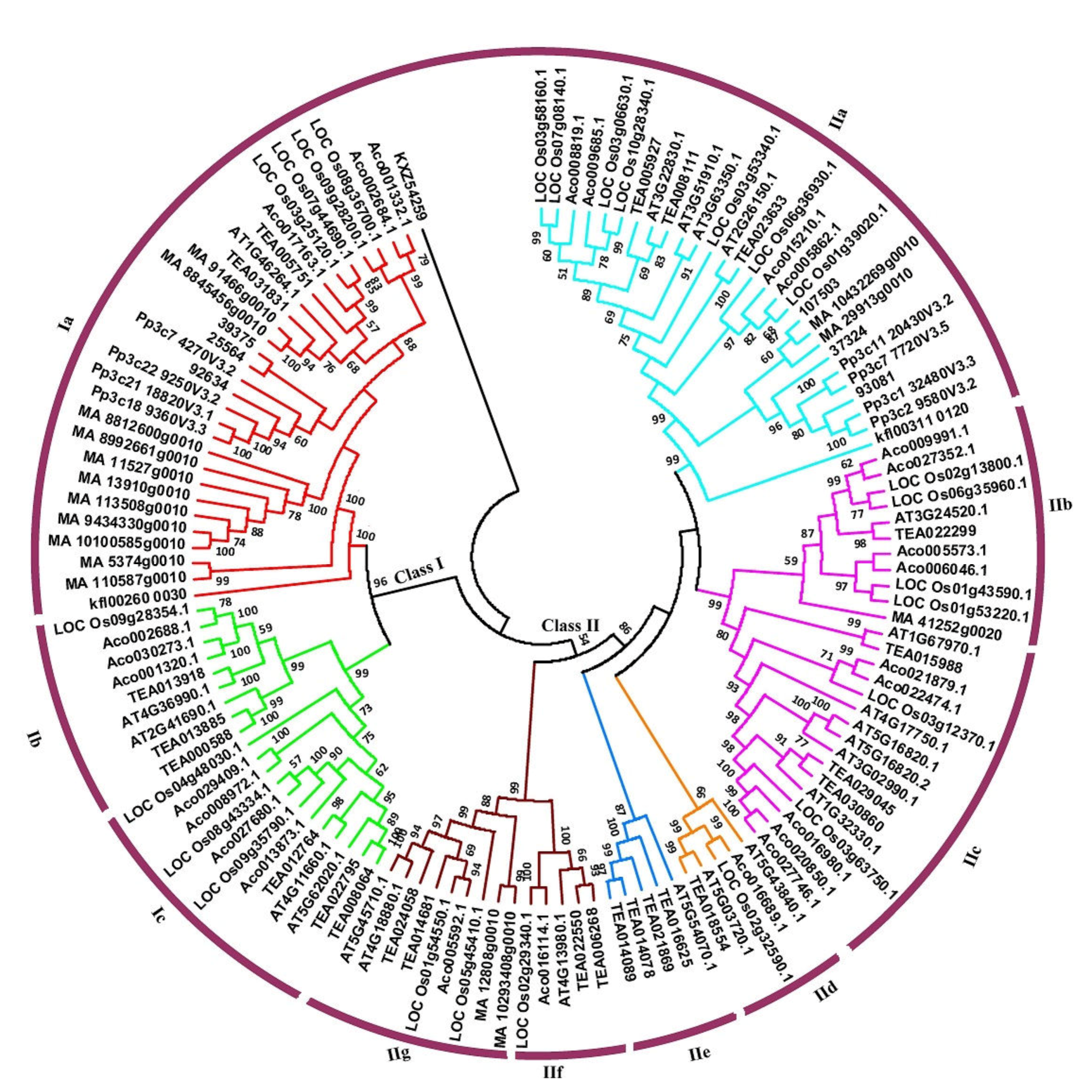
3.2. Phylogenetic Classification of Hsf Genes into Ten Subfamilies
3.3. Multiple Gene Duplication Events of Plant Hsfs
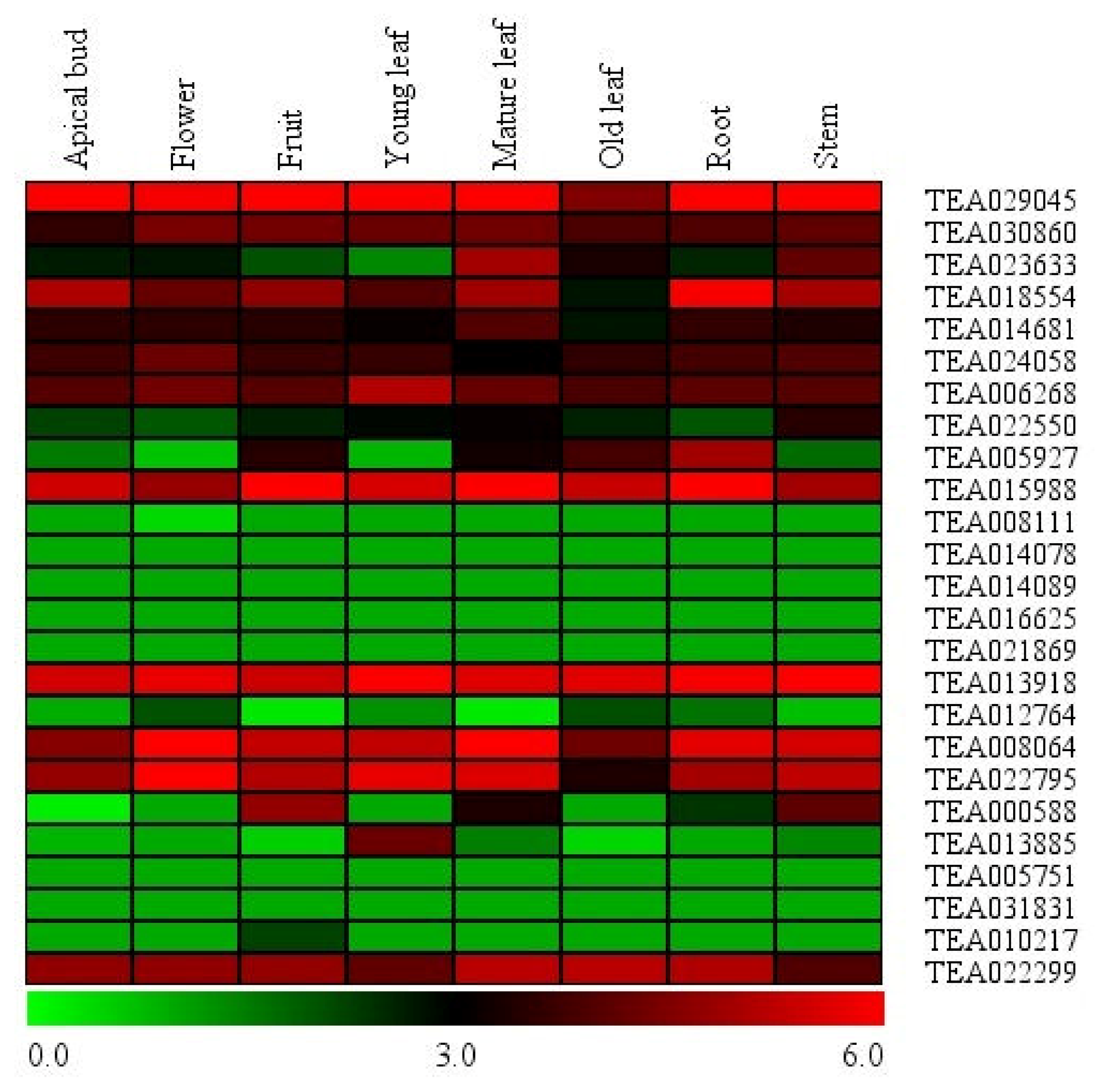
3.4. Expression Analysis of Tea Hsf Transcription Factors in Global Transcriptome
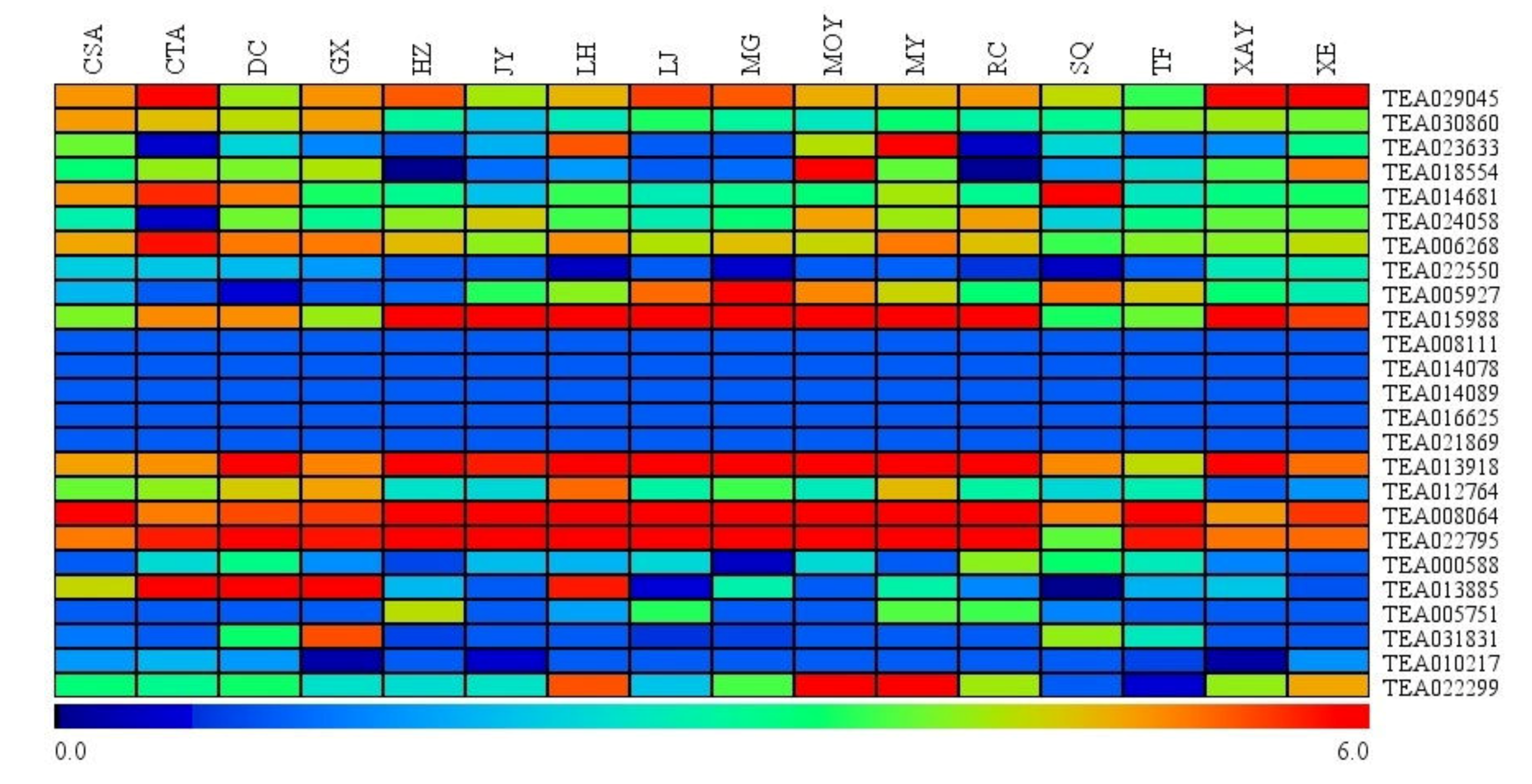
3.5. Expression of Hsfs among Leaves of Closely Representative Camellia Species
3.6. Expression of Hsfs in Response to Hormone Treatment
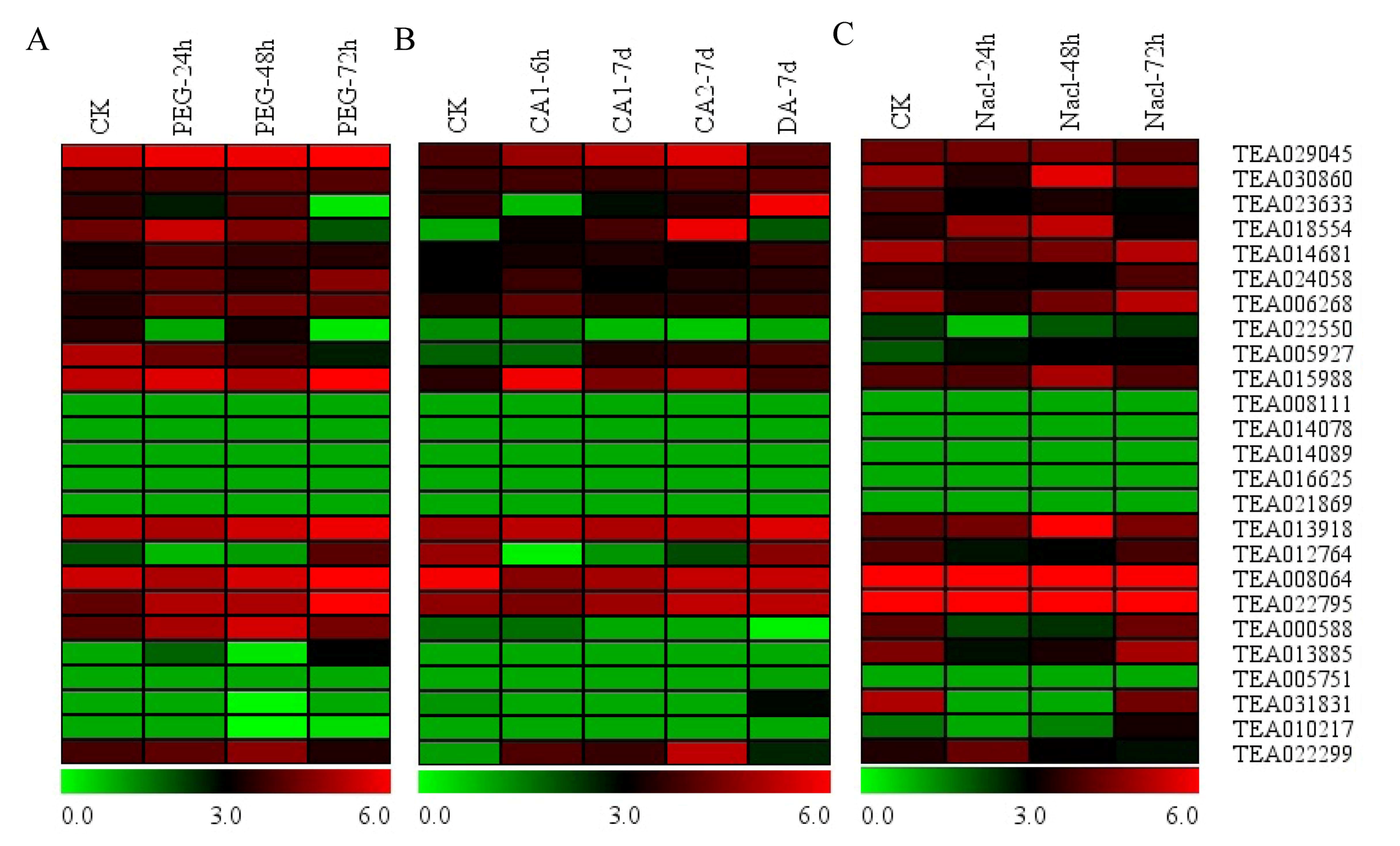
3.7. Expression of Hsfs in Response to PEG, Cold and Nacl Treatments
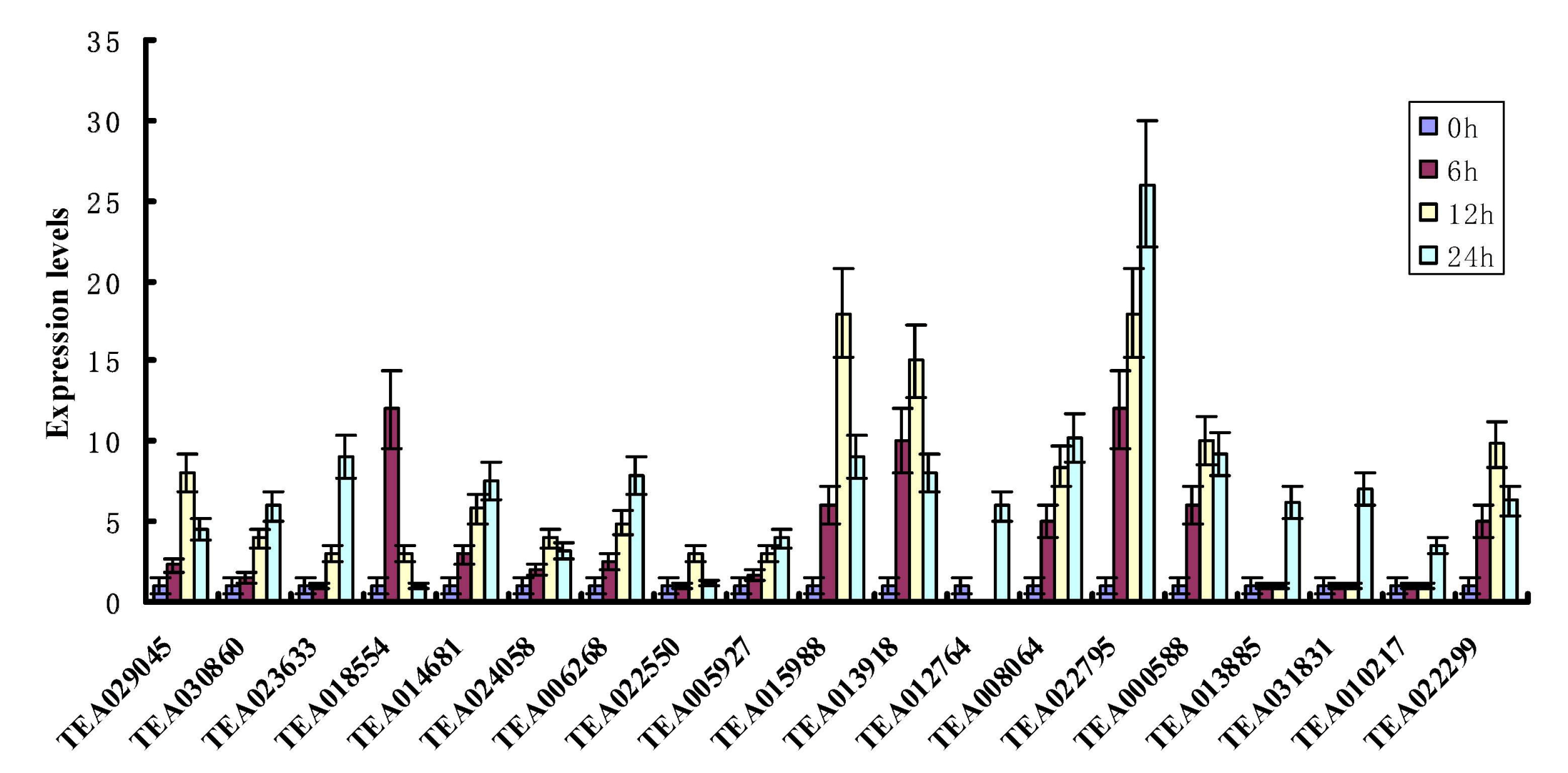
3.8. qRT-PCR Analyses of Tea Hsf Genes under Heat Stress Condition
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Wahid, A.; Gelani, S.; Ashraf, M.; Foolad, M. Heat tolerance in plants: An overview. Environ. Exp. Bot. 2007, 61, 199–223. [Google Scholar] [CrossRef]
- Allakhverdiev, S.I.; Kreslavski, V.D.; Klimov, V.V.; Los, D.A.; Carpentier, R.; Mohanty, P. Heat stress: An overview of molecular responses in photosynthesis. Photosynth. Res. 2008, 98, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, M.; Canales, E.; Borrás-Hidalgo, O. Molecular aspects of abiotic stress in plants. Biotechnol. Appl. 2005, 22, 1–10. [Google Scholar]
- Ashraf, M.; Hafeez, M. Thermotolerance of pearl millet and maize at early growth stages: Growth and nutrient relations. Biol. Plant 2004, 48, 81–86. [Google Scholar] [CrossRef]
- Srivastava, S.; Pathak, A.D.; Gupta, P.S.; Shrivastava, A.K.; Srivastava, A.K. Hydrogen peroxide-scavenging enzymes impart tolerance to high temperature induced oxidative stress in sugarcane. J. Environ. Biol. 2012, 33, 657–661. [Google Scholar]
- Boyer, J.S. Plant productivity and environment. Science 1982, 218, 443–448. [Google Scholar] [CrossRef]
- Century, K.; Reuber, T.L.; Ratcliffe, O.J. Regulating the regulators: The future prospects for transcription-factor-based agricultural biotechnology products. Plant Physiol. 2008, 147, 20–29. [Google Scholar] [CrossRef]
- Chauhan, H.; Khurana, N.; Agarwal, P.; Khurana, P. Heat shock factors in rice (Oryza sativa L.): Genome-wide expression analysis during reproductive development and abiotic stress. Mol. Genet. Genom. 2011, 286, 171–187. [Google Scholar] [CrossRef]
- Chung, E.; Kim, K.M.; Lee, J.H. Genome-wide analysis and molecular characterization of heat shock transcription factor family in Glycine max. J. Genet. Genom. 2013, 40, 127–135. [Google Scholar] [CrossRef]
- Jin, G.H.; Gho, H.J.; Jung, K.H. A systematic view of rice heat shock transcription factor family using phylogenomic analysis. J. Plant Physiol. 2013, 170, 321–329. [Google Scholar] [CrossRef]
- Von Koskull-Döring, P.; Scharf, K.-D.; Nover, L. The diversity of plant heat stress transcription factors. Trends Plant Sci. 2007, 12, 452–457. [Google Scholar] [CrossRef] [PubMed]
- Scharf, K.-D.; Berberich, T.; Ebersberger, I.; Nover, L. The plant heat stress transcription factor (Hsf) family: Structure, function and evolution. Biochim. Biophys. Acta 2012, 1819, 104–119. [Google Scholar] [CrossRef] [PubMed]
- Kotak, S.; Larkindale, J.; Lee, U.; von Koskull-Döring, P.; Vierling, E.; Scharf, K.-D. Complexity of the heat stress response in plants. Curr. Opin. Plant Biol. 2007, 10, 310–316. [Google Scholar] [CrossRef] [PubMed]
- Kotak, S.; Vierling, E.; Bäumlein, H.; von Koskull-Döring, P. A novel transcriptional cascade regulating expression of heat stress proteins during seed development of Arabidopsis. Plant Cell 2007, 19, 182–195. [Google Scholar] [CrossRef]
- Nishizawa, A.; Yabuta, Y.; Yoshida, E.; Maruta, T.; Yoshimura, K.; Shigeoka, S. Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J. 2006, 48, 535–547. [Google Scholar] [CrossRef]
- Shim, D.; Hwang, J.-U.; Lee, J.; Lee, S.; Choi, Y.; An, G.; Martinoia, E.; Lee, Y. Orthologs of the class A4 heat shock transcription factor HsfA4a confer cadmium tolerance in wheat and rice. Plant Cell 2009, 21, 4031–4043. [Google Scholar] [CrossRef]
- Mishra, S.K.; Tripp, J.; Winkelhaus, S.; Tschiersch, B.; Theres, K.; Nover, L.; Schar, K.D. In the complex family of heat stress transcription factors, HsfA1 has a unique role as master regulator of thermotolerance in tomato. Genes Dev. 2002, 16, 1555–1567. [Google Scholar] [CrossRef]
- Giorno, F.; Wolters-Arts, M.; Grillo, S.; Scharf, K.D.; Vriezen, W.H.; Mariani, C. Developmental and heat stress-regulated expression of HsfA2 and small heat shock proteins in tomato anthers. J. Exp. Bot. 2010, 61, 453–462. [Google Scholar] [CrossRef]
- Chan-Schaminet, K.Y.; Baniwal, S.K.; Bublak, D.; Nover, L.; Scharf, K.D. Specific interaction between tomato HsfA1 and HsfA2 creates hetero-oligomeric superactivator complexes for synergistic activation of heat stress gene expression. J. Biol. Chem. 2009, 284, 20848–20857. [Google Scholar] [CrossRef]
- Ogawa, D.; Yamaguchi, K.; Nishiuchi, T. High-level overexpression of the Arabidopsis HsfA2 gene confers not only increased themotolerance but also salt/osmotic stress tolerance and enhanced callus growth. J. Exp. Bot. 2007, 58, 3373–3383. [Google Scholar] [CrossRef]
- Zhang, L.; Li, Y.; Xing, D.; Gao, C. Characterization of mitochondrial dynami cs and subcellul ar localization of ROS reveal that HsfA2 alleviates oxidative damage caused by heat stress in Arabidopsis. J. Exp. Bot. 2009, 60, 2073–2091. [Google Scholar] [CrossRef]
- Banti, V.; Mafessoni, F.; Loreti, E.; Alpi, A.; Perata, P. The heat-inducible transcription factor HsfA2 enhances anoxia tolerance in Arabidopsis. Plant Physiol. 2010, 152, 1471–1483. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Mitsuda, N.; Ohme-Takagi, M. Arabidopsis HsfB1 and HsfB2b act as repressors for the expression of heat-inducible Hsfs but positively regulate the acquired thermotolerance. Plant Physiol. 2011, 157, 1243–1254. [Google Scholar] [CrossRef] [PubMed]
- Almoguera, C.; Rojas, A.; Diaz-Martin, J.; Prieto-Dapena, P.; Carranco, R.; Jordano, J. A seed-specific heat-shock transcription factor involved in developmental regulation during embryogenesis in sunflower. J. Biol. Chem. 2002, 277, 43866–43872. [Google Scholar] [CrossRef]
- Lin, Y.X.; Jiang, H.Y.; Chu, Z.X.; Tang, X.L.; Zhu, S.W.; Cheng, B.J. Genome-wide identification; classification and analysis of heat shock transcription factor family in maize. BMC Genom. 2011, 12, 76. [Google Scholar] [CrossRef]
- Baniwal, S.K.; Bharti, K.; Chan, K.Y.; Fauth, M.; Ganguli, A.; Kotak, S.; Mishra, S.K.; Nover, L.; Port, M.; Scharf, K.D.; et al. Heat stress response in plants: A complex game with chaperones and more than twenty heat stress transcription factors. J. Biosci. 2004, 29, 471–487. [Google Scholar] [CrossRef]
- Harrison, C.J.; Bohm, A.A.; Nelson, H.C. Crystal structure of the DNA binding domain of the heat shock transcription factor. Science 1994, 263, 224–227. [Google Scholar] [CrossRef] [PubMed]
- Vuister, G.W.; Kim, S.J.; Wu, C.; Bax, A. NMR evidence for similaritie s between the DNA-bindi ng regions of Drosophila melanogaster heat shock factor and the helix- turn- helix and HNF-3/forkhead families of transc ription factors. Biochemistry 1994, 33, 10–16. [Google Scholar] [CrossRef] [PubMed]
- Schultheiss, J.; Kunert, O.; Gase, U.; Scharf, K.D.; Nover, L.; Rüterjans, H. Solution structur e of the DNA-binding domain of the tomato heat-stress transcription factor HSF24. Eur. J. Biochem. 1996, 236, 911–921. [Google Scholar] [CrossRef]
- Peteranderl, R.; Rabenstein, M.; Shin, Y.K.; Liu, C.W.; Wemmer, D.E.; King, D.S.; Nelson, H.C. Biochemical and biophysical characterization of the trimerization domain from the heat shock transcription factor. Biochemistry 1999, 38, 3559–3569. [Google Scholar] [CrossRef]
- Nover, L.; Bharti, K.; Döring, P.; Mishra, S.K.; Ganguli, A.; Scharf, K.D. Arabidopsis and the heat stress transcription factor world: How many heat stress transcription factors do we need? Cell Stress Chaperones 2001, 6, 177–189. [Google Scholar] [CrossRef]
- Nover, L.; Scharf, K.D.; Gagliardi, D.; Vergne, P.; Czarnecka-Verner, E.; Gurley, W.B. The Hsf world: Classification and properties of plant heat stress transcription factors. Cell Stress Chaperones 1996, 1, 215–223. [Google Scholar] [CrossRef]
- Lyck, R.; Harmening, U.; Höhfeld, I.; Treuter, E.; Scharf, K.D.; Nover, L. Intracellular distribution and identification of the nuclear localization signals of two plant heat-stress transcription factors. Planta 1997, 202, 117–125. [Google Scholar] [PubMed]
- Heerklotz, D.; Döring, P.; Bonzelius, F.; Winkelhaus, S.; Nover, L. The balance of nuclear import and export determines the intracellular distribution and function of tomato heat stress transcription factor HsfA2. Mol. Cell Biol. 2001, 21, 1759–1768. [Google Scholar] [CrossRef]
- Kotak, S.; Port, M.; Ganguli, A.; Bicker, F.; von Koskull-Döring, P. Characterization of C-terminal domains of Arabidopsis heat stress transcription factors (Hsfs) and identification of a new signature combination of plant class A Hsfs with AHA and NES motifs essential for activator function and intracellular localization. Plant J. 2004, 39, 98–112. [Google Scholar] [CrossRef]
- Song, X.; Liu, G.; Duan, W.; Liu, T.; Huang, Z.; Ren, J.; Li, Y.; Hou, X.L. Genome-wide identification, classification and expression analysis of the heat shock transcription factor family in Chinese cabbage. Mol. Genet. Genom. 2014, 289, 541–551. [Google Scholar] [CrossRef]
- Xue, G.P.; Sadat, S.; Drenth, J.; McIntyre, C.L. The heat shock factor family from Triticum aestivum in response to heat and other major abiotic stresses and their role in regulation of heat shock protein genes. J. Exp. Bot. 2014, 65, 539–557. [Google Scholar] [CrossRef]
- Lu, T.; Zhang, G.; Sun, L.; Wang, J.; Hao, F. Genome-wide identification of CBL family and expression analysis of CBLs in response to potassium deficiency in cotton. Peer J. 2017, 5, e3653. [Google Scholar] [CrossRef]
- Chen, M.Y.; Li, K.; Li, H.P.; Song, C.P.; Miao, Y.C. The Glutathione peroxidase gene family in Gossypium hirsutum: Genome-wide identification classification gene expression and functional analysis. Sci. Rep. 2017, 7, 44743. [Google Scholar]
- Sun, Q.; Wang, G.H.; Zhang, X.; Zhang, X.R.; Qiao, P.; Long, L.; Yuan, Y.L.; Cai, Y.F. Genome-wide identification of the TIFY gene family in three cultivated Gossypium species and the expression of JAZ genes. Sci. Rep. 2017, 7, 42418. [Google Scholar] [CrossRef]
- Guo, Y.W.; Guo, H.L.; Li, X.; Huang, L.L.; Zhang, B.N.; Pang, X.B.; Liu, B.Y.; Ma, L.Q.; Wang, H. Two type III polyketide synthases from Polygonum cuspidatum: Gene structure evolutionary route and metabolites. Plant Biotech. Rep. 2013, 7, 371–381. [Google Scholar] [CrossRef]
- Wei, C.; Yang, H.; Wang, S.; Zhao, J.; Liu, C.; Gao, L.; Xia, E.; Lu, Y.; Tai, Y.; She, G.; et al. Draft genome sequence of Camellia sinensis var. sinensis provides insights into the evolution of the tea genome and tea quality. Proc. Natl. Acad. Sci. USA 2018, 115, E4151–E4158. [Google Scholar] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef]
- Abascal, F.; Zardoya, R.; Posada, D. ProtTest: Selection of best-fit models of protein evolution. Bioinformatics 2005, 21, 2104–2105. [Google Scholar] [CrossRef]
- Wang, L.; Cao, H.; Qian, W.; Yao, L.; Hao, X.; Li, N.; Yang, Y.; Wang, X. Identification of a novel bZIP transcription factor in Camellia sinensis as a negative regulator of freezing tolerance in transgenic Arabidopsis. Ann. Bot. 2017, 119, 1195–1209. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Li, K.; Yang, F.B.; Miao, Y.C.; Song, C.P. Abscisic acid signaling is involved in regulating the mitogen-activated protein kinase cascade module AIK1-MKK5-MPK6. BMC Plant Biol. 2017, 12, 5. [Google Scholar] [CrossRef]
- Zhang, G.; Lu, T.; Miao, W.; Sun, L.; Tian, M.; Wang, J.; Hao, F. Genome-wide identification of ABA receptor PYL family and expression analysis of PYLs in response to ABA and osmotic stress in Gossypium. Peer J. 2017, 5, e4126. [Google Scholar] [CrossRef]
- Liu, L.Y.; Li, N.; Yao, C.P.; Meng, S.S.; Song, C.P. Functional analysis of the ABA-responsive protein family in ABA and stress signal transduction in Arabidopsis. Chin. Sci. Bull. 2013, 58, 3721–3730. [Google Scholar] [CrossRef]
- Hao, F.; Zhao, S.; Dong, H.; Zhang, H.; Sun, L.; Miao, C. Nia1 and Nia2 are involved in exogenous salicylic acid-induced nitric oxide generation and stomatal closure in Arabidopsis. J. Integr. Plant Biol. 2010, 52, 298–307. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Xiang, F.; Zhang, G.; Miao, Y.; Miao, C.; Song, C.P. Abscisic acid as an internal integrator of multiple physiological processes modulates leaf senescence onset in Arabidopsis thaliana. Front. Plant Sci. 2016, 7, 181. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Li, Y.Y.; Xiao, H.L.; Xu, C.S.; Zhang, X. Nitric oxide blocks blue light-induced K(+) influx by elevating the cytosolic Ca2+ concentration in Vicia faba L. Guard Cells. J. Integr. Plant Biol. 2013, 55, 527–536. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; He, J.N.; Zhao, Y.; Wu, T.; Zhou, X.; Ding, Y.; Kong, L.; Wang, X.; Wang, Y.; Li, J.; et al. EAR1 negatively regulates ABA signaling by enhancing 2C protein phosphatase activity. Plant Cell 2018, 30, 815–834. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; de Ollas, C.; Dodd, I.C. Long-distance ABA transport can mediate distal tissue responses by affecting local ABA concentrations. J. Integr. Plant Biol. 2018, 60, 16–33. [Google Scholar] [CrossRef]
- Xu, L.H.; Wang, W.Y.; Guo, J.J.; Qin, J.; Shi, D.Q.; Li, Y.L.; Xu, J. Zinc improves salt tolerance by increasing reactive oxygen species scavenging and reducing Na+ accumulation in wheat seedlings. Biol. Plant. 2014, 58, 751–757. [Google Scholar] [CrossRef]
- Gao, W.; Xu, F.C.; Guo, D.D.; Zhao, J.R.; Liu, J.; Guo, Y.W.; Singh, P.K.; Ma, X.N.; Long, L.; Botella, J.R.; et al. Calcium-dependent protein kinases in cotton: Insights into early plant responses to salt stress. BMC Plant Biol. 2018, 18, 15. [Google Scholar] [CrossRef]
- Li, W.; Zhao, F.; Fang, W.; Xie, D.; Hou, J.; Yang, X.; Zhao, Y.; Tang, Z.; Nie, L.; Lv, S. Identification of early salt stress responsive proteins in seedling roots of upland cotton (Gossypium hirsutum L) employing iTRAQ-based proteomic technique. Front. Plant Sci. 2015, 6, 732. [Google Scholar] [CrossRef]
- Zhao, X.; Wang, Y.J.; Wang, Y.L.; Wang, X.L.; Zhang, X. Extracellular Ca2+ alleviates NaCl-induced stomatal opening through a pathway involving H2O2-blocked Na+ influx in Vicia guard cells. J. Plant Physiol. 2011, 168, 903–910. [Google Scholar] [CrossRef]
- Ma, L.; Zhang, H.; Sun, L.; Jiao, Y.; Zhang, G.; Miao, C.; Hao, F. NADPH oxidase AtrbohD and AtrbohF function in ROS-dependent regulation of Na+/K+ homeostasis in Arabidopsis under salt stress. J. Exp. Bot. 2012, 63, 305–317. [Google Scholar] [CrossRef]
- Xu, F.C.; Liu, H.L.; Xu, Y.Y.; Zhao, J.R.; Guo, Y.W.; Long, L.; Gao, W.; Song, C.P. Heterogeneous expression of the cotton R2R3-MYB transcription factor GbMYB60 increases salt sensitivity in transgenic Arabidopsis. Plant Cell Tiss. Organ Cul. 2018, 133, 15–25. [Google Scholar] [CrossRef]
- Lv, S.; Yu, D.; Sun, Q.; Jiang, J. Activation of gibberellin 20-oxidase 2 undermines auxin-dependent root and root hair growth in NaCl-stressed Arabidopsis seedlings. Plant Growth Regul. 2018, 84, 225–236. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, F.; Zhang, C.; Zhang, J.; Chen, Y.; Liu, G.; Zhao, Y.; Hao, F. A novel VIGS method by agroinoculation of cotton seeds and application for elucidating functions of GhBI-1 in salt-stress response. Plant. Cell Rep. 2018, 37, 1091–1100. [Google Scholar] [CrossRef] [PubMed]
- Qi, J.; Song, C.P.; Wang, B.; Zhou, J.; Kangasjärvi, J.; Zhu, J.K.; Gong, Z. Reactive oxygen species signaling and stomatal movement in plant responses to drought stress and pathogen attack. J. Integr. Plant Biol. 2018, 60, 805–826. [Google Scholar] [CrossRef]
- Zhao, Q.; Chen, W.; Bian, J.; Xie, H.; Li, Y.; Xu, C.; Ma, J.; Guo, S.; Chen, J.; Cai, X.; et al. Proteomics and phosphoproteomics of heat stress-responsive mechanisms in spinach. Front. Plant Sci. 2018, 9, 800. [Google Scholar] [CrossRef]
- Li, L.; Hou, M.; Cao, L.; Shen, Z.G.; Hu, Z.B. Glutathione S-transferases modulate Cu tolerance in Oryza sativa. Environ. Exp. Bot. 2018, 155, 313–320. [Google Scholar] [CrossRef]
- Pang, Y.Q.; Li, J.T.; Qi, B.S.; Tian, M.; Sun, L.R.; Wang, X.C.; Hao, F.S. Aquaporin AtTIP5;1 as an essential target of gibberellins promotes hypocotyl cell elongation in Arabidopsis thaliana under excess boron stress. Func. Plant Biol. 2017, 45, 305–314. [Google Scholar] [CrossRef]
- Sun, L.; Ma, L.; He, S.; Hao, F. AtrbohD functions downstream of ROP2 and positively regulates waterlogging response in Arabidopsis. Plant Signal. Behav. 2018, 13, 1–5. [Google Scholar] [CrossRef]
- Wang, T.P.; Liu, H.; Hua, H.J.; Wang, L.; Song, C.P. A vacuole localized beta-glucosidase contributes to drought tolerance in Arabidopsis. Chin. Sci. Bull. 2011, 56, 3538–3546. [Google Scholar] [CrossRef]
- Lü, D.; Wang, W.; Miao, C. ATHK1 acts downstream of hydrogen peroxide to mediate ABA signaling through regulation of calcium channel activity in Arabidopsis guard cells. Chin. Sci. Bull. 2013, 58, 336–343. [Google Scholar] [CrossRef]
- Ma, X.N.; Zhang, X.R.; Yang, L.; Tang, M.M.; Wang, K.; Wang, L.; Bai, L.; Song, C.P. Hydrogen peroxide plays an important role in PERK4-mediated abscisic acid-regulated root growth in Arabidopsis. Func. Plant Biol. 2018, 46, 165–174. [Google Scholar] [CrossRef]
- Pick, T.; Jaskiewicz, M.; Peterhänsel, C.; Conrath, U. Heat shock factor HsfB1 primes gene transcription and systemic acquired resistance in Arabidopsis. Plant Physiol. 2012, 159, 52–55. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.Z.; Cheng, Y.W.; Ya, H.Y.; Gao, K.; Han, J.M. Identification and analysis of the heat shock transcriptor PsHsfB1 in tree peony. Appl. Mech. Mater. 2014, 675–677, 1133–1136. [Google Scholar] [CrossRef]
- Liang, J.Y.; Xia, J.Y.; Liu, L.L.; Wan, S.Q. Global patterns of the responses of leaf-level photosynthesis and respiration in terrestrial plants to experimental warming. J. Plant Ecol. 2013, 6, 437–447. [Google Scholar] [CrossRef]
- Zhao, X.; Wang, Y.L.; Qiao, X.R.; Wang, J.; Wang, L.D.; Xu, C.S.; Zhang, X. Phototropins function in high-intensity blue light-induced hypocotyl phototropism in Arabidopsis by altering cytosolic calcium. Plant Physiol. 2013, 162, 1539–1551. [Google Scholar] [CrossRef]
- Han, Y.J.; Wang, X.H.; Chen, W.C.; Dong, M.F.; Yuan, W.J.; Liu, X.; Shang, F.D. Differential expression of carotenoid-related genes determines diversified carotenoid coloration in flower petal of Osmanthus fragrans. Tree Genet. Genomes 2014, 10, 329–338. [Google Scholar] [CrossRef]
- Han, Y.; Wu, M.; Cao, L.; Yuan, W.; Dong, M.; Wang, X.; Chen, W.; Shang, F. Characterization of OfWRKY3 a transcription factor that positively regulates the carotenoid cleavage dioxygenase gene OfCCD4 in Osmanthus fragrans. Plant Mol. Biol. 2016, 91, 485–496. [Google Scholar] [CrossRef]
- Guo, S.; Dai, S.; Singh, P.K.; Wang, H.; Wang, Y.; Tan, J.L.H.; Wee, W.; Ito, T. A membrane-bound NAC-like transcription factor OsNTL5 represses the flowering in Oryza sativa. Front. Plant Sci. 2018, 9, 555. [Google Scholar] [CrossRef]
- Shang, B.S.; Zang, Y.H.; Zhao, X.; Zhu, J.D.; Fan, C.; Guo, X.N.; Zhang, X. Functional characterization of GhPHOT2 in chloroplast avoidance of Gossypium hirsutum. Plant Physiol. Biochem. 2019, 135, 51–60. [Google Scholar] [CrossRef]
- Wan, H.J.; Yuan, W.; Ruan, M.Y.; Ye, Q.J.; Wang, R.Q.; Li, Z.M.; Zhou, G.; Yao, Z.; Zhao, J.; Liu, S.; et al. Identification of reference genes for reverse transcription quantitative real-time PCR normalization in pepper (Capsicum annuum L.). Biochem. Biophys. Res. Commun. 2011, 416, 24–30. [Google Scholar] [CrossRef]
- Moreno, A.A.; Orellana, A. The physiological role of the unfolded protein response in plants. Biol. Res. 2011, 44, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Hasanuzzaman, M.; Hossain, M.A.; Fujita, M. Physiological and biochemical mechanisms of nitric oxide induced abiotic stress tolerance in plants. Am. J. Plant Physiol. 2010, 5, 295–324. [Google Scholar]
- Xue, G.P.; Drenth, J.; Lynne, M.I.C. TaHsfA6f is a transcriptional activator that regulates a suite of heat stress protection genes in wheat (Triticum aestivum L.) including previously unknown Hsf targets. J. Exp. Bot. 2015, 66, 1025–1039. [Google Scholar] [CrossRef] [PubMed]

| Gene Name | Locus | Scaffold | ORF Length a (bp) | Deduced Polypeptide | ||
|---|---|---|---|---|---|---|
| Length b (aa) | Mol wt c (kDa) | pId | ||||
| CsHSF01 | TEA000588 | Scaffold4738:242479-245432 | 723 | 240 | 27.83 | 5.59 |
| CsHSF02 | TEA005751 | Scaffold3668:1248898-1251806 | 849 | 282 | 32.80 | 6.42 |
| CsHSF03 | TEA005927 | Scaffold372:5052384-5054777 | 1059 | 352 | 40.55 | 5.37 |
| CsHSF04 | TEA006268 | Scaffold1096:351535-356817 | 1440 | 479 | 53.66 | 5.76 |
| CsHSF05 | TEA008064 | Scaffold385:66932-69032 | 900 | 299 | 34.02 | 6.27 |
| CsHSF06 | TEA008111 | Scaffold736:2759744-2761297 | 1023 | 340 | 38.81 | 5.07 |
| CsHSF07 | TEA010217 | Scaffold4444:1140231-1146979 | 519 | 172 | 19.44 | 10.01 |
| CsHSF08 | TEA012764 | Scaffold474:257888-260307 | 945 | 314 | 35.08 | 5.37 |
| CsHSF09 | TEA013885 | Scaffold598:1002357-1004559 | 723 | 240 | 27.87 | 6.67 |
| CsHSF10 | TEA013918 | Scaffold598:1726479-1739146 | 1569 | 522 | 57.83 | 5.16 |
| CsHSF11 | TEA014078 | Scaffold2655:1213850-1216317 | 996 | 331 | 37.20 | 5.14 |
| CsHSF12 | TEA014089 | Scaffold2655:1252903-1263984 | 1401 | 466 | 51.34 | 4.49 |
| CsHSF13 | TEA014681 | Scaffold2339:167132-178145 | 1998 | 665 | 74.98 | 5.91 |
| CsHSF14 | TEA015988 | Scaffold2960:495401-502597 | 1167 | 388 | 44.26 | 4.72 |
| CsHSF15 | TEA016625 | Scaffold2245:1443845-1445616 | 1116 | 371 | 41.43 | 4.71 |
| CsHSF16 | TEA018554 | Scaffold367:1317907-1320377 | 1722 | 573 | 63.81 | 5.02 |
| CsHSF17 | TEA021869 | Scaffold151:757381-759282 | 1266 | 421 | 47.11 | 5.16 |
| CsHSF18 | TEA022299 | Scaffold881:562868-564244 | 966 | 321 | 36.25 | 5.35 |
| CsHSF19 | TEA022550 | Scaffold502:2406177-2409890 | 684 | 207 | 23.56 | 5.68 |
| CsHSF20 | TEA022795 | Scaffold3997:691331-693719 | 1104 | 367 | 40.70 | 5.35 |
| CsHSF21 | TEA023633 | Scaffold1243:1037225-1040235 | 1203 | 400 | 45.39 | 4.85 |
| CsHSF22 | TEA024058 | Scaffold1471:1329221-1337467 | 1470 | 489 | 55.56 | 5.00 |
| CsHSF23 | TEA029045 | Scaffold1047:271898-276789 | 1587 | 508 | 56.46 | 5.13 |
| CsHSF24 | TEA030860 | Scaffold2384:70221-80010 | 1542 | 513 | 56.78 | 4.78 |
| CsHSF25 | TEA031831 | Scaffold2436:227948-229420 | 906 | 301 | 33.38 | 6.49 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, P.; Guo, Q.; Pang, X.; Zhang, P.; Kong, D.; Liu, J. New Insights into Evolution of Plant Heat Shock Factors (Hsfs) and Expression Analysis of Tea Genes in Response to Abiotic Stresses. Plants 2020, 9, 311. https://doi.org/10.3390/plants9030311
Xu P, Guo Q, Pang X, Zhang P, Kong D, Liu J. New Insights into Evolution of Plant Heat Shock Factors (Hsfs) and Expression Analysis of Tea Genes in Response to Abiotic Stresses. Plants. 2020; 9(3):311. https://doi.org/10.3390/plants9030311
Chicago/Turabian StyleXu, Ping, Qinwei Guo, Xin Pang, Peng Zhang, Dejuan Kong, and Jia Liu. 2020. "New Insights into Evolution of Plant Heat Shock Factors (Hsfs) and Expression Analysis of Tea Genes in Response to Abiotic Stresses" Plants 9, no. 3: 311. https://doi.org/10.3390/plants9030311
APA StyleXu, P., Guo, Q., Pang, X., Zhang, P., Kong, D., & Liu, J. (2020). New Insights into Evolution of Plant Heat Shock Factors (Hsfs) and Expression Analysis of Tea Genes in Response to Abiotic Stresses. Plants, 9(3), 311. https://doi.org/10.3390/plants9030311





