Molecular Cloning and Characterization of SQUAMOSA-Promoter Binding Protein-Like Gene FvSPL10 from Woodland Strawberry (Fragaria vesca)
Abstract
:1. Introduction
2. Results
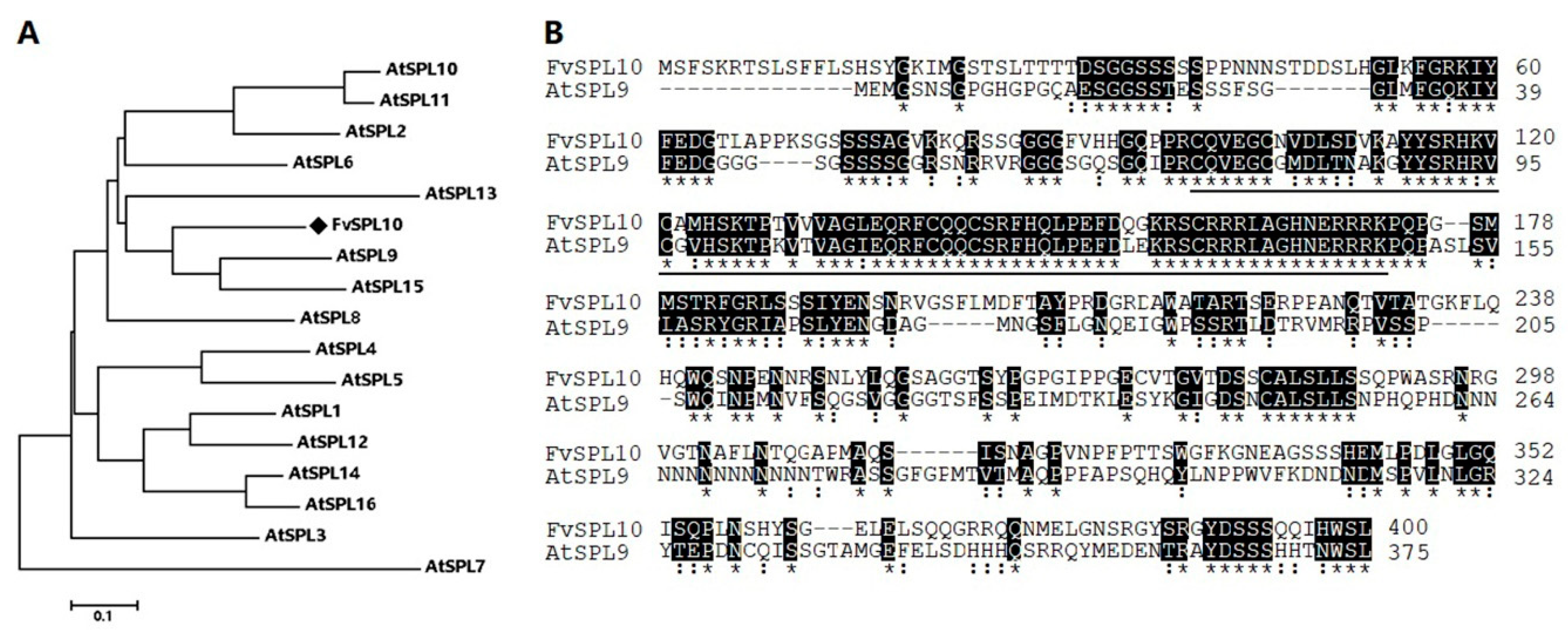
2.1. Cloning and Characterization of FvSPL10 in Woodland Strawberry
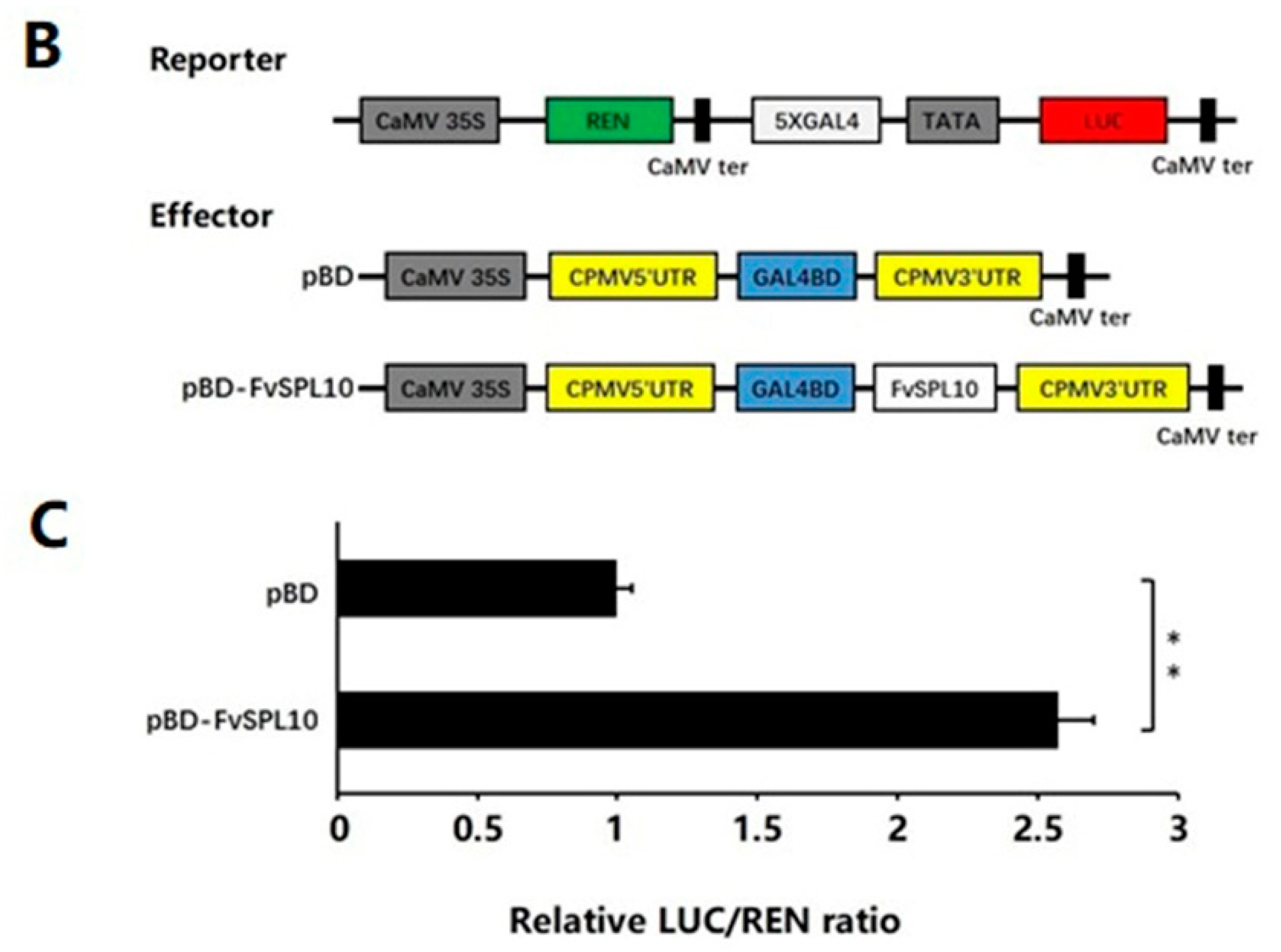
2.2. Subcellular Localization and Transcriptional Activity of FvSPL10
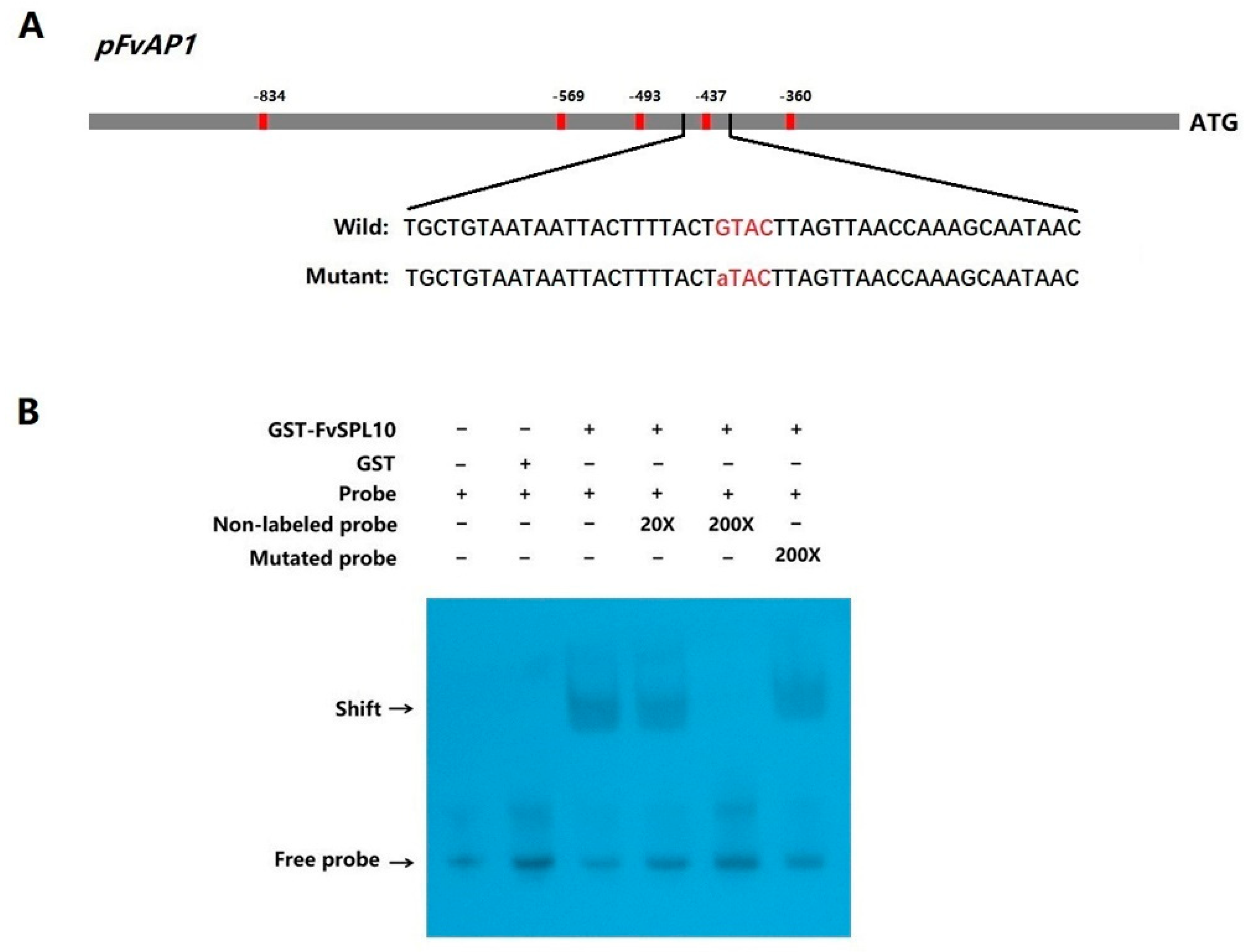
2.3. Interaction of FvSPL10 with the GTAC Motif in FvAP1 Promoter in Vitro
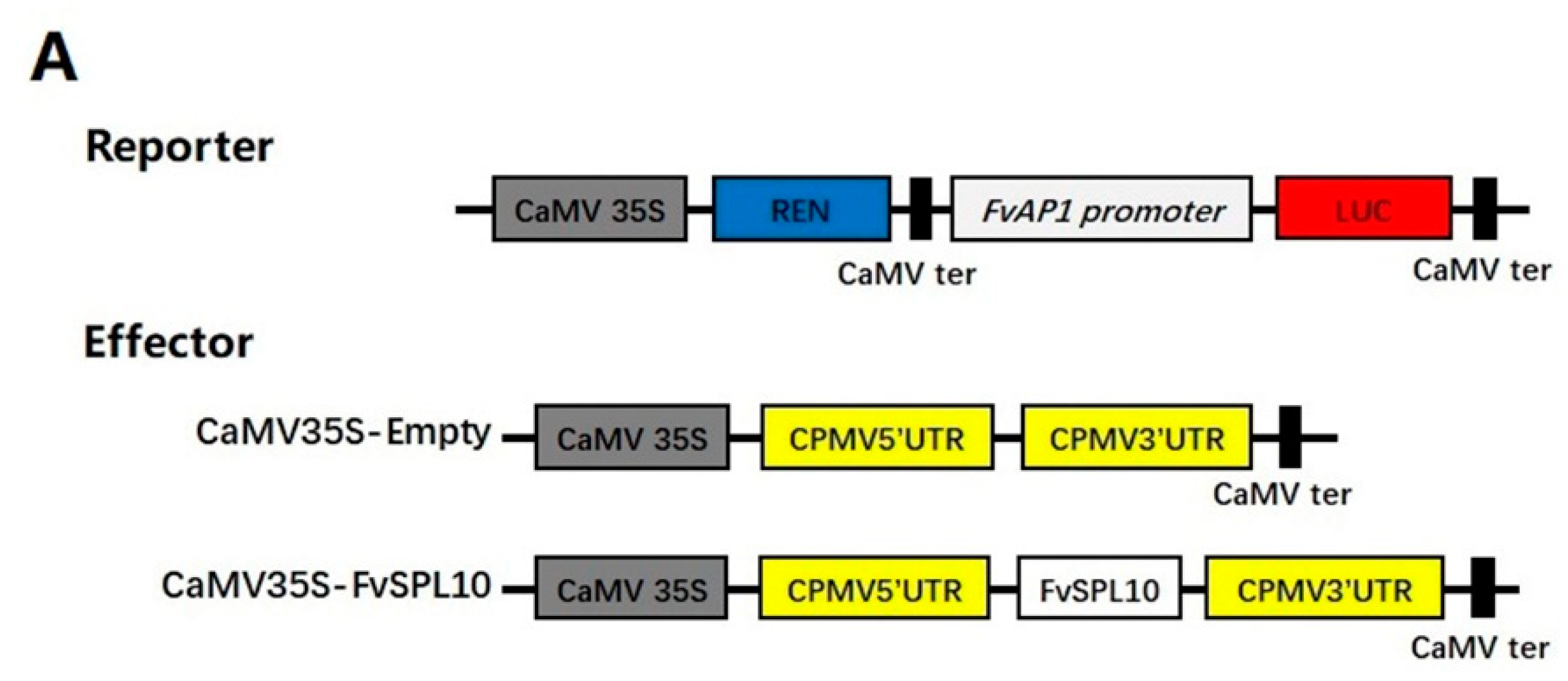
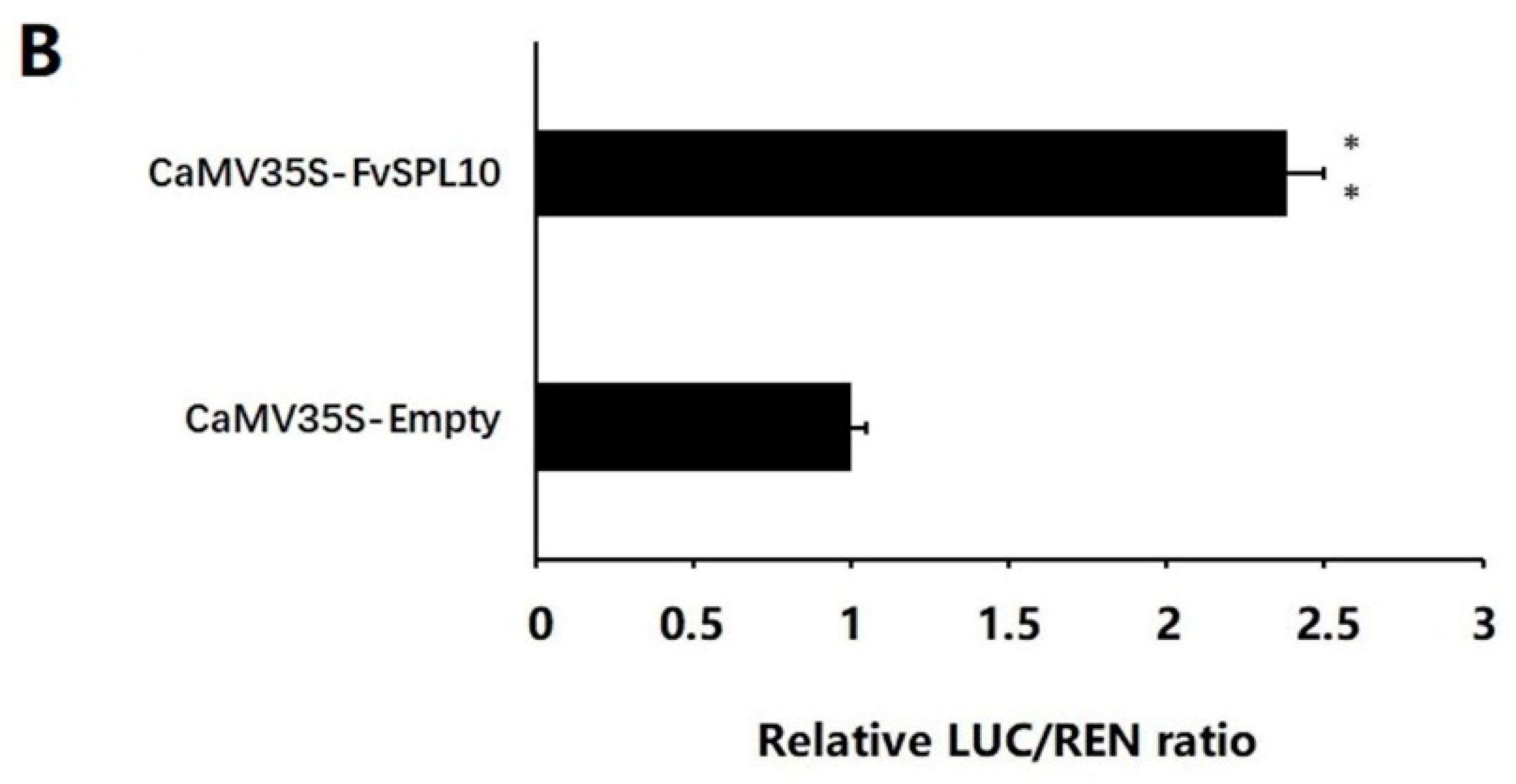
2.4. FvSPL10 Activates the Transcription of FvAP1 in Vivo
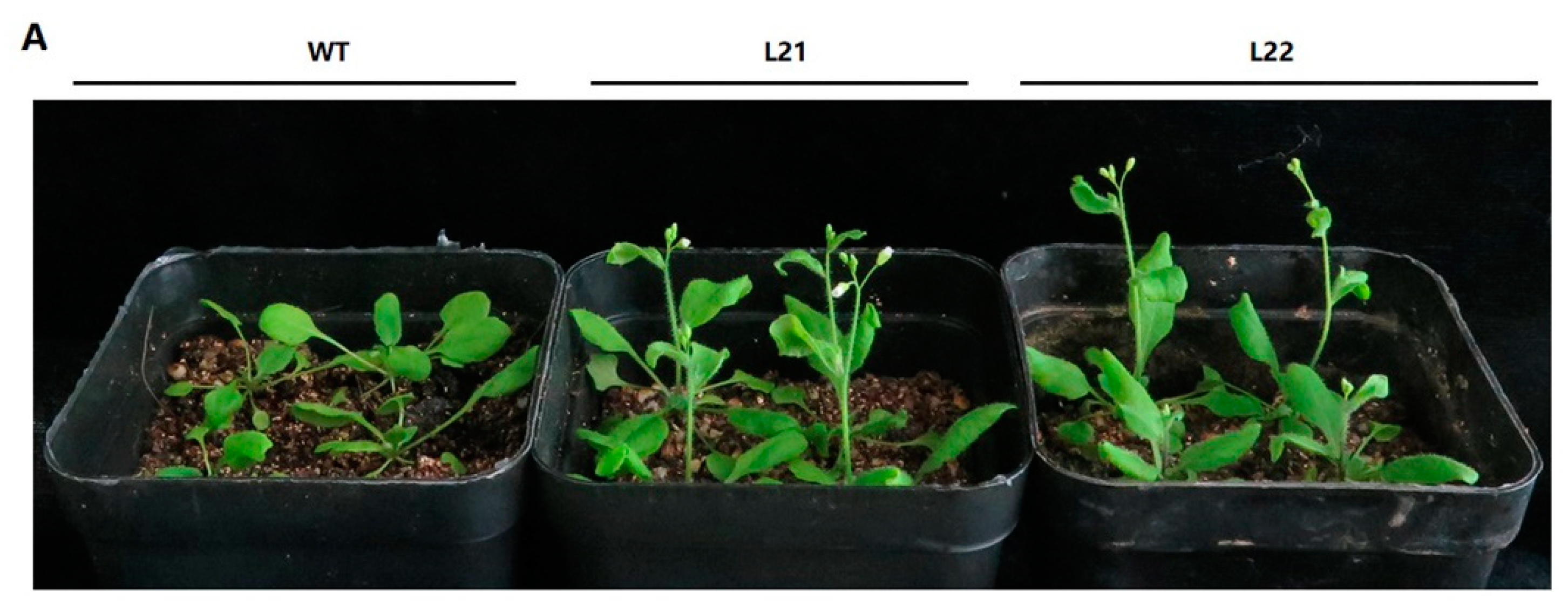
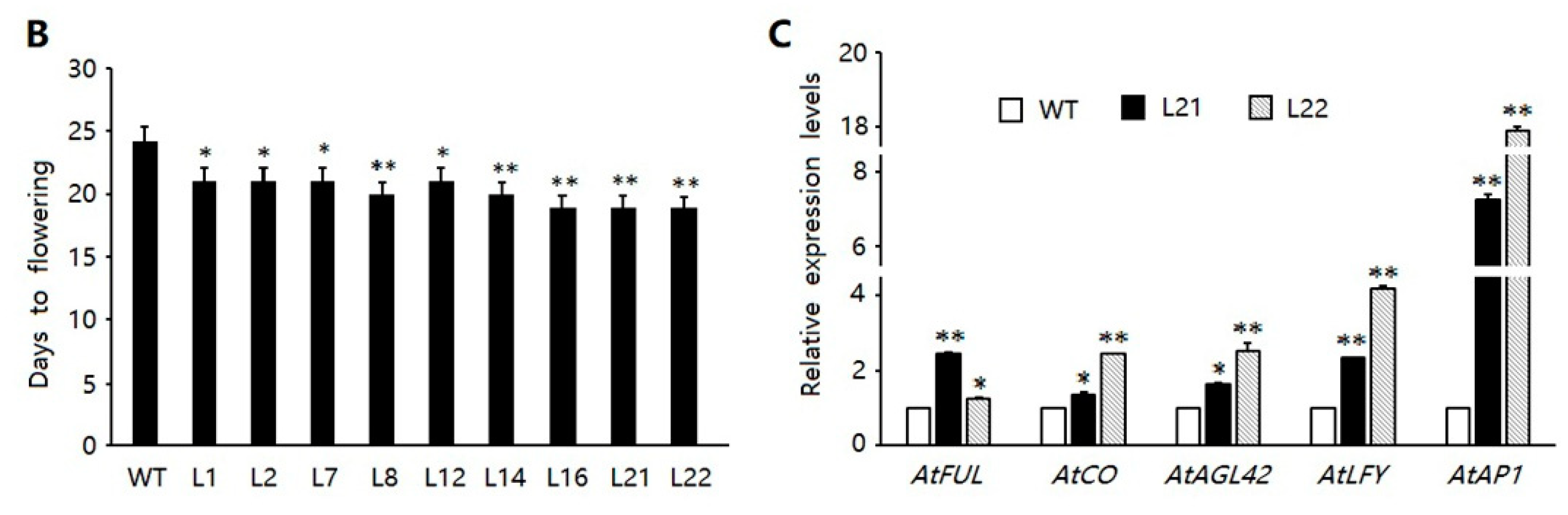
2.5. Ectopic Expression of FvSPL10 in Arabidopsis Promotes Flowering
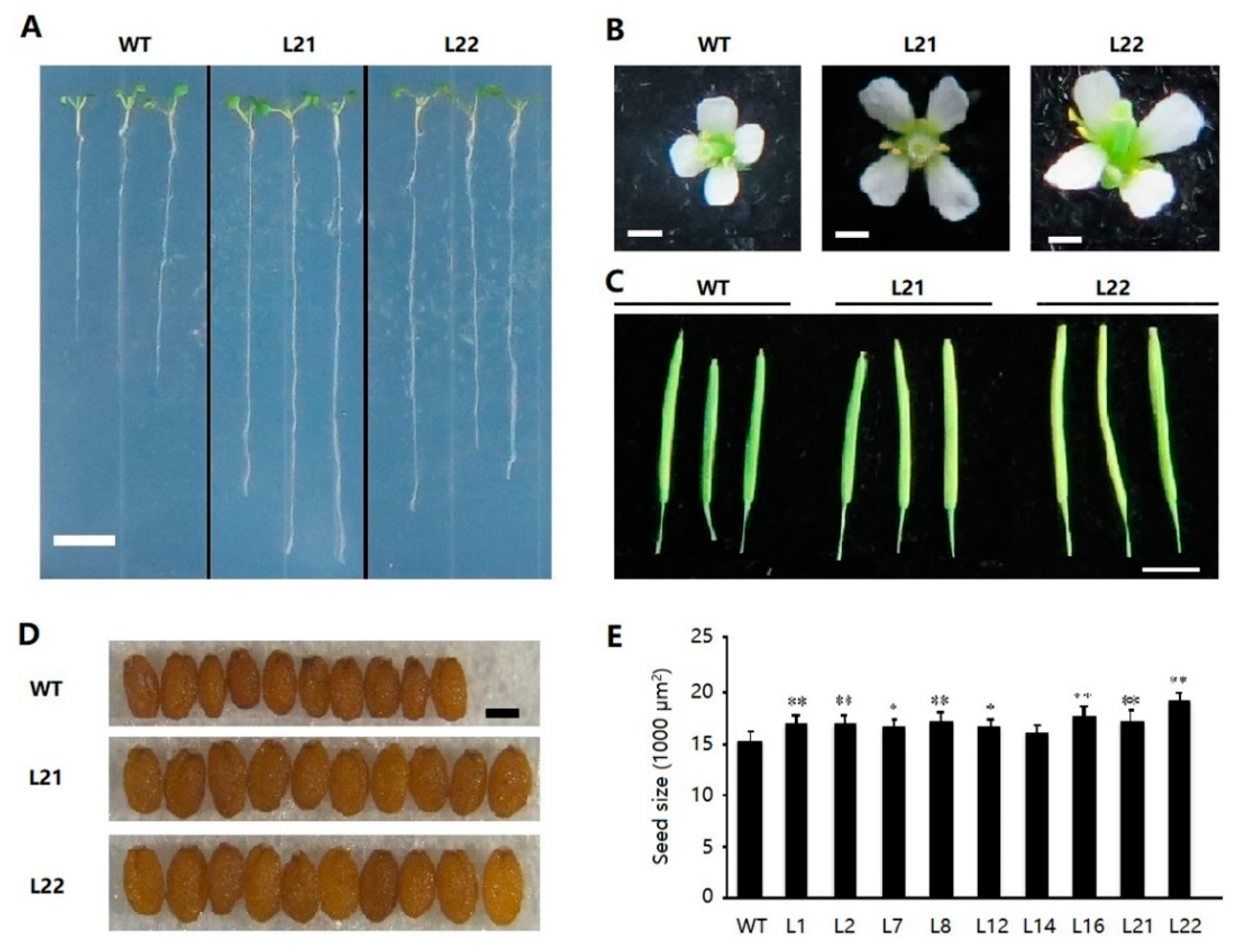
2.6. Ectopic Expression of FvSPL10 in Arabidopsis Enlarges Organs Size
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Nucleic Acid Isolation
4.3. Bioinformatic Analysis
4.4. Subcellular Localization Analysis of FvSPL10
4.5. EMSA
4.6. Transcriptional Activity Analysis of FvSPL10
4.7. Construction of FvSPL10 Overexpression Vector and Arabidopsis Transformation
4.8. Gene Expression Analysis
4.9. Phenotypic Measurement and Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Birkenbihl, R.P.; Jach, G.; Saedler, H.; Huijser, P. Functional dissection of the plant-specific SBP-domain: Overlap of the DNA-binding and nuclear localization domains. J. Mol. Biol. 2005, 352, 585–596. [Google Scholar] [CrossRef] [PubMed]
- Klein, J.; Saedler, H.; Huijser, P. A new family of DNA binding proteins includes putative transcriptional regulators of the Antirrhinum majus floral meristem identity gene SQUAMOSA. Mol. Gen. Genet. 1996, 250, 7–16. [Google Scholar] [PubMed]
- Guo, A.Y.; Zhu, Q.H.; Gu, X.C.; Ge, S.; Yang, J.; Luo, J.C. Genome-wide identification and evolutionary analysis of the plant specific SBP-box transcription factor family. Gene 2008, 418, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hultquist, J.F.; Dorweiler, J.E. Feminized tassels of maize mop1 and ts1 mutants exhibit altered levels of miR156 and specific SBP-box genes. Planta 2008, 229, 99–113. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Wang, X.; Gu, S.; Hu, Z.; Xu, H.; Xu, C. Comparative study of SBP-box gene family in Arabidopsis and rice. Gene 2008, 407, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Salinas, M.; Xing, S.; Hohmann, S.; Berndtgen, R.; Huijser, P. Genomic organization, phylogenetic comparison and differential expression of the SBP-box family of transcription factors in tomato. Planta 2012, 235, 1171–1184. [Google Scholar] [CrossRef] [PubMed]
- Hou, H.M.; Li, J.; Gao, M.; Singer, S.D.; Wang, H.; Mao, L.Y.; Fei, Z.J.; Wang, X.P. Genomic Organization, Phylogenetic Comparison and Differential Expression of the SBP-Box Family Genes in Grape. PLoS One 2013, 8, e59358. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.S.; Zheng, D.; Zhu, H.Y.; Chen, J.Q.; Na, R.; Cheng, Z.M. Genome-wide identification and expression analysis of the SPL gene family in woodland strawberry Fragaria vesca. Genome 2018, 61, 675–683. [Google Scholar] [CrossRef]
- Cardon, G.; Hohmann, S.; Klein, J.; Nettesheim, K.; Saedler, H.; Huijser, P. Molecular characterisation of the Arabidopsis SBP-box genes. Gene 1999, 237, 91–104. [Google Scholar] [CrossRef]
- Chao, L.M.; Liu, Y.Q.; Chen, D.Y.; Xue, X.Y.; Mao, Y.B.; Chen, X.Y. Arabidopsis Transcription Factors SPL1 and SPL12 Confer Plant Thermotolerance at Reproductive Stage. Mol. Plant 2017, 10, 735–748. [Google Scholar] [CrossRef]
- Shikata, M.; Koyama, T.; Mitsuda, N.; Ohme-Takagi, M. Arabidopsis SBP-box genes SPL10, SPL11 and SPL2 control morphological change in association with shoot maturation in the reproductive phase. Plant Cell Physiol. 2009, 50, 2133–2145. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.W.; Czech, B.; Weigel, D. miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 2009, 138, 738–749. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, A.; Wu, M.F.; Yang, L.; Wu, G.; Poethig, R.S.; Wagner, D. The MicroRNA-Regulated SBP-Box Transcription Factor SPL3 Is a Direct Upstream Activator of LEAFY, FRUITFULL, and APETALA1. Dev. Cell 2009, 17, 268–278. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.L.; Hu, T.Q.; Zhao, J.F.; Park, M.Y.; Earley, K.W.; Wu, G.; Yang, L.; Poethig, R.S. Developmental Functions of miR156-Regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) Genes in Arabidopsis thaliana. PLoS Genet. 2016, 12, e1006263. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, M.S.; Ma, S.S.; Burch-Smith, T.M.; Czymmek, K.; Huijser, P.; Dinesh-Kumar, S.P. Novel Positive Regulatory Role for the SPL6 Transcription Factor in the N TIR-NB-LRR Receptor-Mediated Plant Innate Immunity. PLoS Pathog. 2013, 9, e1003235. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.Y.; Zhao, X.; Li, J.G.; Cai, H.Q.; Deng, X.W.; Li, L. MicroRNA408 Is Critical for the HY5-SPL7 Gene Network That Mediates the Coordinated Response to Light and Copper. Plant Cell 2014, 26, 4933–4953. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.Y.; Lin, S.; Qiu, Z.B.; Cao, D.C.; Wen, J.L.; Deng, X.; Wang, X.H.; Lin, J.X.; Li, X.J. MicroRNA857 Is Involved in the Regulation of Secondary Growth of Vascular Tissues in Arabidopsis. Plant Physiol. 2015, 169, 2539–2552. [Google Scholar] [CrossRef] [PubMed]
- Unte, U.S.; Sorensen, A.M.; Pesaresi, P.; Gandikota, M.; Leister, D.; Saedler, H.; Huijser, P. SPL8, an SBP-Box gene that affects pollen sac development in Arabidopsis. Plant Cell 2003, 15, 1009–1019. [Google Scholar] [CrossRef]
- Schwarz, S.; Grande, A.V.; Bujdoso, N.; Saedler, H.; Huijser, P. The microRNA regulated SBP-box genes SPL9 and SPL15 control shoot maturation in Arabidopsis. Plant Mol. Biol. 2008, 67, 183–195. [Google Scholar] [CrossRef]
- Gou, J.Y.; Felippes, F.F.; Liu, C.J.; Weigel, D.; Wang, J.W. Negative Regulation of Anthocyanin Biosynthesis in Arabidopsis by a miR156-Targeted SPL Transcription Factor. Plant Cell 2011, 23, 1512–1522. [Google Scholar] [CrossRef]
- Zhang, Q.Q.; Wang, J.G.; Wang, L.Y.; Wang, J.F.; Wang, Q.; Yu, P.; Bai, M.Y.; Fan, M. Gibberellin repression of axillary bud formation in Arabidopsis by modulation of DELLA-SPL9 complex activity. J. Integr. Plant Biol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Yu, N.; Cai, W.J.; Wang, S.C.; Shan, C.M.; Wang, L.J.; Chen, X.Y. Temporal Control of Trichome Distribution by MicroRNA156-Targeted SPL Genes in Arabidopsis thaliana. Plant Cell 2010, 22, 2322–2335. [Google Scholar] [CrossRef] [PubMed]
- Stone, J.M.; Liang, X.; Nekl, E.R.; Stiers, J.J. Arabidopsis AtSPL14, a plant-specific SBP-domain transcription factor, participates in plant development and sensitivity to fumonisin B1. Plant J. 2005, 41, 744–754. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.Q.; Wang, Y.H.; Xue, D.W.; Wang, J.; Yan, M.X.; Liu, G.F.; Dong, G.J.; Zeng, D.L.; Lu, Z.F.; Zhu, X.D.; et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 2010, 42, 541–544. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Ikeda, M.; Matsubara, A.; Song, X.J.; Ito, M.; Asano, K.; Matsuoka, M.; Kitano, H.; Ashikari, M. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 2010, 42, 545–549. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.K.; Wu, K.; Yuan, Q.B.; Liu, X.Y.; Liu, Z.B.; Lin, X.Y.; Zeng, R.Z.; Zhu, H.T.; Dong, G.J.; Qian, Q.; et al. Control of grain size, shape and quality by OsSPL16 in rice. Nat. Genet. 2012, 44, 950–954. [Google Scholar] [CrossRef]
- Manning, K.; Tor, M.; Poole, M.; Hong, Y.; Thompson, A.J.; King, G.J.; Giovannoni, J.J.; Seymour, G.B. A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening. Nat. Genet. 2006, 38, 948–952. [Google Scholar] [CrossRef]
- Lu, Z.F.; Yu, H.; Xiong, G.S.; Wang, J.; Jiao, Y.Q.; Liu, G.F.; Jing, Y.H.; Meng, X.B.; Hu, X.M.; Qian, Q.; et al. Genome-Wide Binding Analysis of the Transcription Activator IDEAL PLANT ARCHITECTURE1 Reveals a Complex Network Regulating Rice Plant Architecture. Plant Cell 2013, 25, 3743–3759. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, J.; Cheng, X.L.; Liu, P.; Sun, J.Q. miR156-Targeted SBP-Box Transcription Factors Interact with DWARF53 to Regulate TEOSINTE BRANCHED1 and BARREN STALK1 Expression in Bread Wheat. Plant Physiol. 2017, 174, 1931–1948. [Google Scholar] [CrossRef]
- Yu, S.; Galvao, V.C.; Zhang, Y.C.; Horrer, D.; Zhang, T.Q.; Hao, Y.H.; Feng, Y.Q.; Wang, S.; Schmid, M.; Wang, J.W. Gibberellin Regulates the Arabidopsis Floral Transition through miR156-Targeted SQUAMOSA PROMOTER BINDING-LIKE Transcription Factors. Plant Cell 2012, 24, 3320–3332. [Google Scholar] [CrossRef]
- Cao, R.F.; Guo, L.J.; Ma, M.; Zhang, W.J.; Liu, X.L.; Zhao, H.X. Identification and Functional Characterization of Squamosa Promoter Binding Protein-Like Gene TaSPL16 in Wheat (Triticum aestivum L.). Front. Plant Sci. 2019, 10, 212. [Google Scholar] [CrossRef] [PubMed]
- Nitsch, J.P. Growth and morphogenesis of the strawberry as related to auxin. Am. J. Bot. 1950, 37, 211–215. [Google Scholar] [CrossRef]
- Chang, S.J.; Puryear, J.; Cairney, J. A simple and efficient method for isolating RNA from pine trees. Plant Mol. Biol. Report. 1993, 11, 113–116. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, M.N.; Huang, Z.J.; Hua, Q.Z.; Shan, W.; Kuang, J.F.; Lu, W.J.; Qin, Y.H.; Chen, J.Y. The WRKY transcription factor HpWRKY44 regulates CytP450-like1 expression in red pitaya fruit (Hylocereus polyrhizus). Hortic. Res. 2017, 4, 17039. [Google Scholar] [CrossRef] [PubMed]
- Hellens, R.P.; Allan, A.C.; Friel, E.N.; Bolitho, K.; Grafton, K.; Templeton, M.D.; Karunairetnam, S.; Gleave, A.P.; Laing, W.A. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 2005, 1, 13. [Google Scholar] [CrossRef]
- Zhang, X.R.; Henriques, R.; Lin, S.S.; Niu, Q.W.; Chua, N.H. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 2006, 1, 641–646. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Schindelin, J.; Rueden, C.T.; Hiner, M.C.; Eliceiri, K.W. The ImageJ ecosystem: An open platform for biomedical image analysis. Mol. Reprod. Dev. 2015, 82, 518–529. [Google Scholar] [CrossRef] [Green Version]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xiong, J.; Bai, Y.; Ma, C.; Zhu, H.; Zheng, D.; Cheng, Z. Molecular Cloning and Characterization of SQUAMOSA-Promoter Binding Protein-Like Gene FvSPL10 from Woodland Strawberry (Fragaria vesca). Plants 2019, 8, 342. https://doi.org/10.3390/plants8090342
Xiong J, Bai Y, Ma C, Zhu H, Zheng D, Cheng Z. Molecular Cloning and Characterization of SQUAMOSA-Promoter Binding Protein-Like Gene FvSPL10 from Woodland Strawberry (Fragaria vesca). Plants. 2019; 8(9):342. https://doi.org/10.3390/plants8090342
Chicago/Turabian StyleXiong, Jinsong, Yibo Bai, Chuangju Ma, Hongyu Zhu, Dan Zheng, and Zongming Cheng. 2019. "Molecular Cloning and Characterization of SQUAMOSA-Promoter Binding Protein-Like Gene FvSPL10 from Woodland Strawberry (Fragaria vesca)" Plants 8, no. 9: 342. https://doi.org/10.3390/plants8090342
APA StyleXiong, J., Bai, Y., Ma, C., Zhu, H., Zheng, D., & Cheng, Z. (2019). Molecular Cloning and Characterization of SQUAMOSA-Promoter Binding Protein-Like Gene FvSPL10 from Woodland Strawberry (Fragaria vesca). Plants, 8(9), 342. https://doi.org/10.3390/plants8090342




