Molecular Insights into FaEG1, a Strawberry Endoglucanase Enzyme Expressed during Strawberry Fruit Ripening
Abstract
1. Introduction
2. Methods
2.1. Plant Material
2.2. RNA Isolation, Reverse Transcription and Reverse-Transcription Quantitative Polymerase Chain Reaction (RT-qPCR) Analysis
2.3. Scanning Electron Microscopy (SEM)
2.4. Sequence Comparison and Phylogenic Analysis
2.5. FaEG1 Structural Model Obtained by Comparative Modeling
2.6. Protein–Ligand Interaction Determinations
3. Results and Discussion
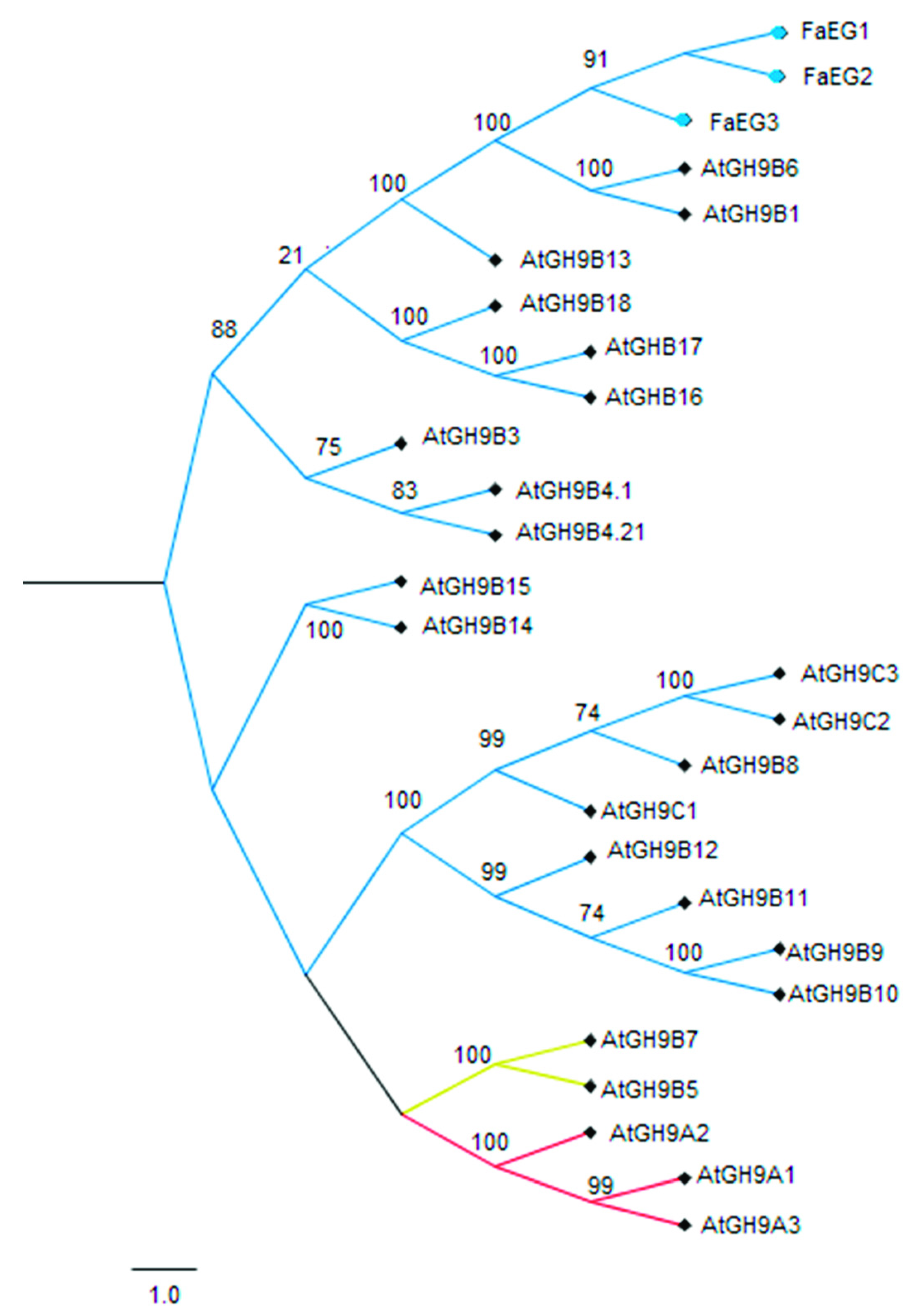
3.1. Sequence and Phylogenetic Analyses
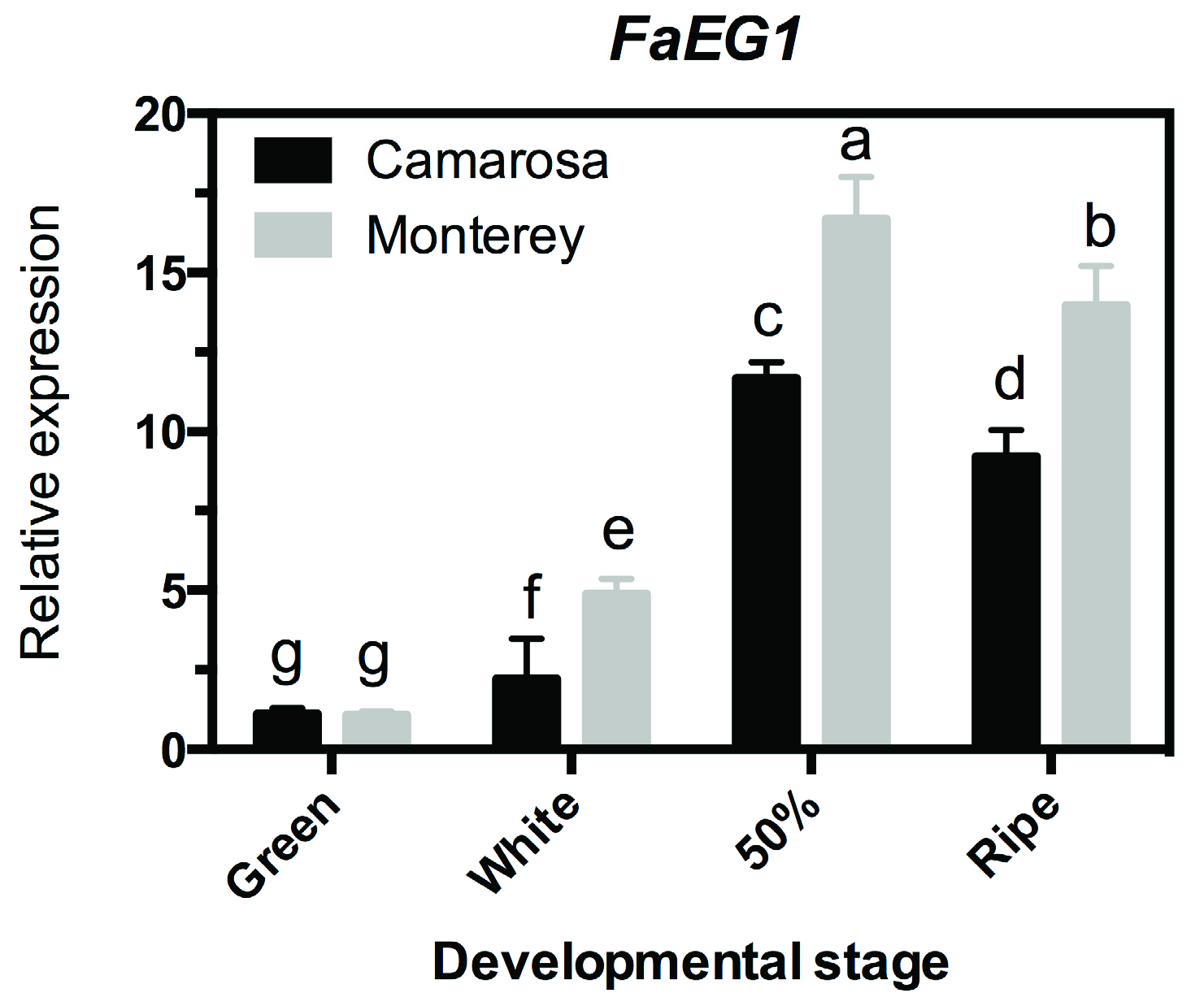
3.2. Transcriptional Profile Analysis of the FaEG1 Gene
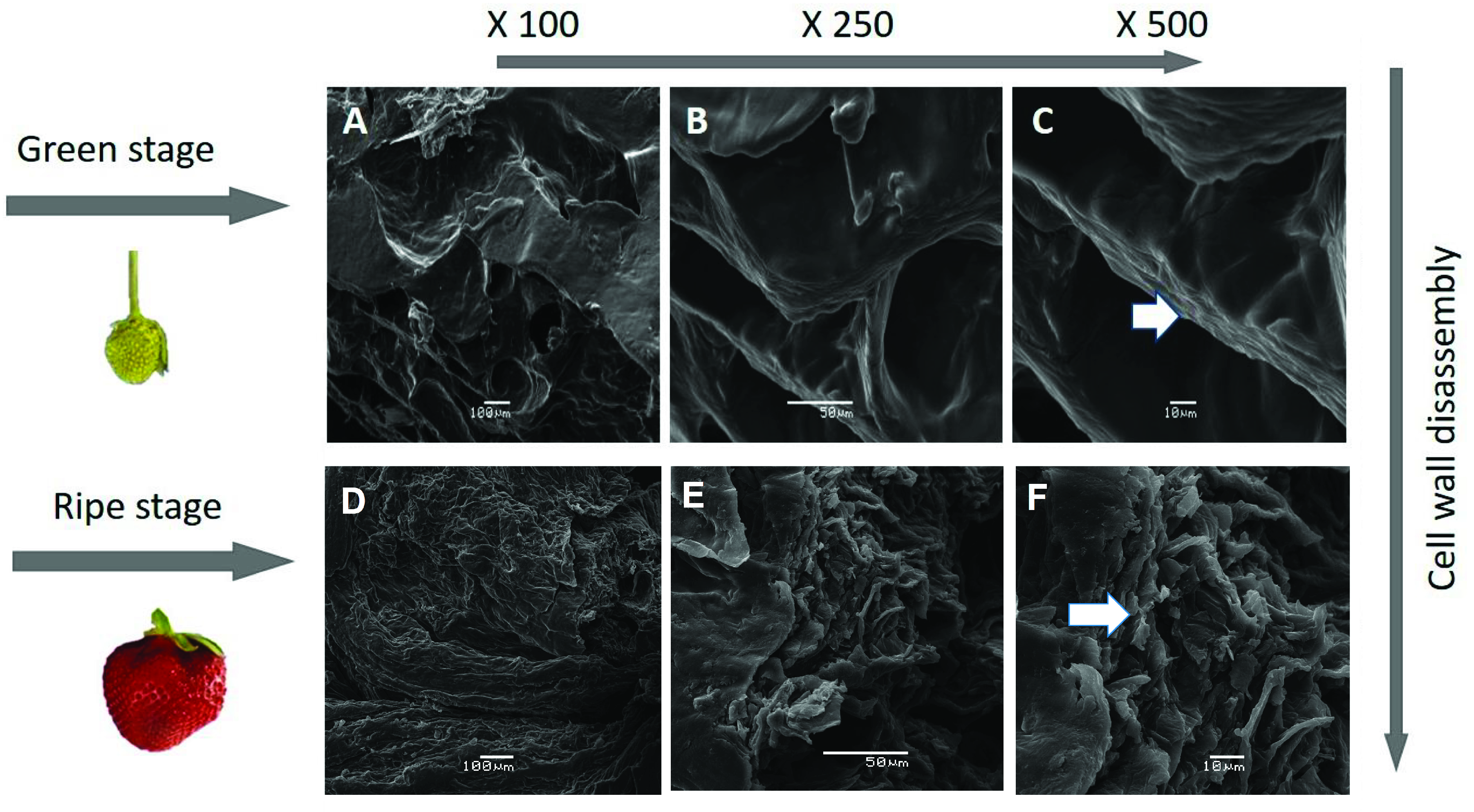
3.3. SEM Studies of the Surface Morphology of Strawberries
3.4. Obtaining the FaEG1 Protein Model
3.5. 3D Structure of the FaEG1 Protein Model
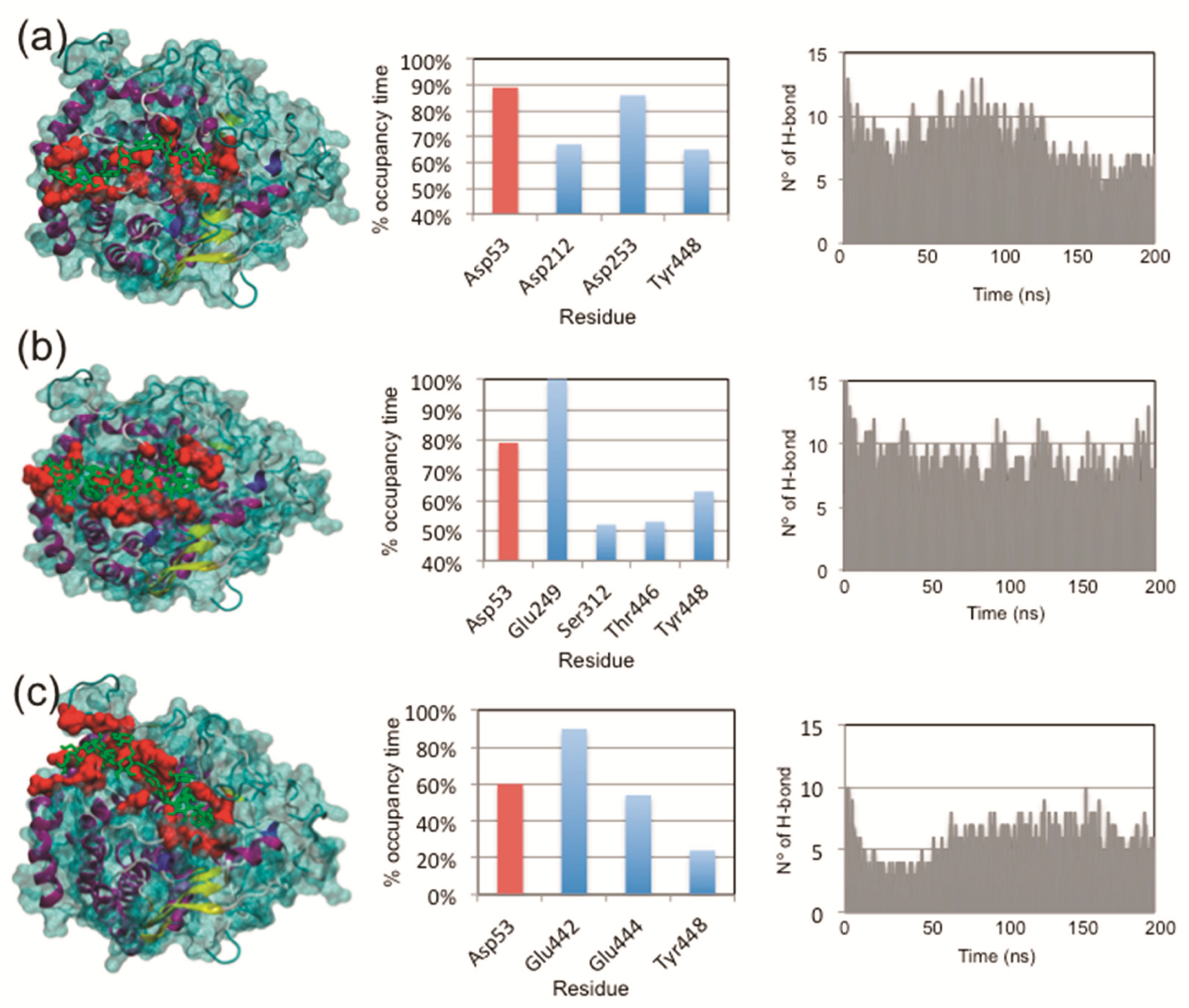
3.6. Protein–Ligand Interactions
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rosli, H.G.; Civello, P.M.; Martínez, G.A. Changes in cell wall composition of three Fragaria x ananassa cultivars with different softening rate during ripening. Plant Physiol. Biochem. 2004, 42, 823–831. [Google Scholar] [CrossRef] [PubMed]
- Ramos, P.; Parra-Palma, C.; Figueroa, C.R.; Zuñiga, P.E.; Valenzuela-Riffo, F.; Gonzalez, J.; Gaete-Eastman, C.; Morales-Quintana, L. Cell wall-related enzymatic activities and transcriptional profiles in four strawberry (Fragaria x ananassa) cultivars during fruit development and ripening. Sci. Hortic. 2018, 238, 325–332. [Google Scholar] [CrossRef]
- Brummell, D.A.; Harpster, M.H. Cell wall metabolism in fruit softening and quality and its manipulation in transgenic plants. Plant Mol. Biol. 2001, 47, 311–340. [Google Scholar] [CrossRef] [PubMed]
- Castro, R.I.; Morales-Quintana, L. Study of the cell wall components produced during different ripening stages through thermogravimetric analysis. Cellulose 2019, 26, 3009–3020. [Google Scholar] [CrossRef]
- Vicente, A.R.; Saladié, M.; Rose, J.K.C.; Labavitch, J.M. The linkage between cell wall metabolism and fruit softening: looking to the future. J. Sci. Food Agric. 2007, 87, 1435–1448. [Google Scholar] [CrossRef]
- Valenzuela-Riffo, F.; Ramos, P.; Morales-Quintana, L. Computational study of FaEXPA1 a strawberry alpha expansin protein, through molecular modeling and molecular dynamics simulation studies”. Comput. Biol. Chem. 2018, 76, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Mendez-Yañez, A.; Beltrán, D.; Campano-Romero, C.; Molinett, S.; Herrera, R.; Moya-León, M.A.; Morales-Quintana, L. Glycosylation is important for FcXTH1 activity as judged by its structural and biochemical characterization. Plant Physiol. Biochem. 2017, 119, 200–210. [Google Scholar] [CrossRef]
- Goulao, L.F.; Oliveira, C.M. Cell wall modification during fruit ripening: When a fruit is not the fruit. Trends Food Sci. Technol. 2008, 19, 4–25. [Google Scholar] [CrossRef]
- Mercado, J.A.; Pliego-Alfaro, F.; Quesada, M.A. Fruit shelf life and potential for its genetic improvement. In Breeding for Fruit Quality; Jenks, M.A., Bebeli, P.J., Eds.; John Wiley & Sons: Oxford, UK, 2011; pp. 81–104. [Google Scholar]
- Quesada, M.A.; Blanco-Portales, R.; Pose, R.S.; Gracía-Gago, J.A.; Jiménez-Bermúdez, S.; Muñóz-Serrano, A.; Caballero, J.L.; Pilego-Alfaro, F.; Mercado, J.A.; Muñóz-Blanco, J. Antisense down-regulation of the FaPG1 gene reveals an unexpected central role for polygalacturonase in strawberry fruit softening. Plant Physiol. 2009, 150, 1022–1032. [Google Scholar] [CrossRef]
- Segato, F.; Dias, B.; Berto, G.L.; de Oliveira, D.M.; de Souza, F.H.M.; Citadini, A.P.; Murakami, M.T.; Damásio, A.R.L.; Squina, F.M.; Polikarpov, I. Cloning, heterologous expression and biochemical characterization of a non-specific endoglucanase family 12 from Aspergillus terreus NIH2624. BBA-Proteins Proteom. 2017, 1865, 395–403. [Google Scholar] [CrossRef]
- Strakowska, J.; Błaszczyk, L.; Chełkowski, J. The significance of cellulolytic enzymes produced by Trichoderma in opportunistic lifestyle of this fungus. J. Basic Microbiol. 2014, 54, S2–S13. [Google Scholar] [CrossRef] [PubMed]
- Ramani, G.; Meera, B.; Vanitha, C.; Rajendhran, J.; Gunasekaran, P. Molecular cloning and expression of thermostable glucose-tolerant β-glucosidase of Penicillium funiculosum NCL1 in Pichia pastoris and its characterization. J. Ind. Microbiol. Biotechnol. 2015, 42, 553–565. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Zhang, R.; Yang, X.; Xu, Y.; Tang, Z.; Tian, W.; Shen, Q. Expression, purification and characterization of two thermostable endoglucanases cloned from a lignocellulosic decomposing fungi Aspergillus fumigatus Z5 isolated from compost. Protein Expr. Purif. 2011, 79, 176–186. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Liu, W.; Li, Y.; Lai, H.L.; Zheng, Y.; Huang, J.W.; Chen, C.C.; Chen, Y.; Jin, J.; Li, H.; et al. Functional and structural analysis of Pichia pastoris-expressed Aspergillus Niger 1,4-β-endoglucanase. Biochem. Biophys. Res. Commun. 2016, 475, 8–12. [Google Scholar] [CrossRef] [PubMed]
- Shani, Z.; Dekel, M.; Roiz, L.; Horowitz, M.; Kolosovski, N.; Lapidot, S.; Alkan, S.; Koltai, H.; Tsabary, G.; Goren, R.; et al. Expression of endo-1,4-beta-glucanase (cel1) in Arabidopsis thaliana isassociated with plant growth, xylem development and cell Wall thickening. Plant Cell Rep. 2006, 25, 1067–1074. [Google Scholar] [CrossRef] [PubMed]
- Rose, J.K.C.; Bennett, A.B. Cooperative disassembly of the cellulose-xyloglucan network of plant cell walls: Parallels between cell expansion and fruit ripening. Trends Plant Sci. 1999, 4, 176–183. [Google Scholar] [CrossRef]
- Bhandari, S.; Fujino, T.; Thammanagowda, S.; Zhang, D.; Xu, F.; Joshi, C.P. Xylem-specific and tension stress-responsive coexpression of KORRIGAN endoglucanase and three secondary wallassociated cellulose synthase genes in aspen trees. Planta 2006, 224, 828–837. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Enzymes and other agents that enhance cell Wall extensibility. Annu. Rev. Plant Biol. 1999, 50, 391–417. [Google Scholar] [CrossRef]
- Libertini, E.; Li, Y.; McQueen–Mason, S.J. Phylogenetic Analysis of the Plant Endo-b-1,4-Glucanase Gene Family. J. Mol. Evol. 2004, 58, 506–515. [Google Scholar] [CrossRef]
- Urbanowicz, B.R.; Bennett, A.B.; del Campillo, E.; Catala, C.; Hayashi, T.; Henrissat, B.; Hofte, H.; McQueen-Mason, S.J.; Patterson, S.E.; Shoseyov, O.; et al. Structural organization and a standardized nomenclature for plant endo-1,4-bglucanases (cellulases) of glycosyl hydrolase family 9. Plant Physiol. 2007, 144, 1693–1696. [Google Scholar] [CrossRef]
- Xie, G.; Yang, B.; Xu, Z.; Li, F.; Guo, K.; Zhang, M.; Wang, L.; Zou, W.; Wang, Y.; Peng, L. Global identification of multiple OsGH9 family members and their involvement in cellulose crystallinity modification in rice. PLoS ONE 2013, 8, e50171. [Google Scholar] [CrossRef] [PubMed]
- Harpster, M.H.; Brummell, D.A.; Dunsmuir, P. Expression analysis of a ripening-specific, auxin-repressed endo-b-1,4 glucanase gene in strawberry. Plant Physiol. 1998, 118, 1307–1316. [Google Scholar] [CrossRef] [PubMed]
- Manning, K. Isolation of a set of ripening-related genes from strawberry: Their identification and possible relationship to fruit quality traits. Planta 1998, 205, 622–631. [Google Scholar] [CrossRef] [PubMed]
- Llop-Tous, I.; Domínguez-Puigjaner, E.; Palomer, X.; Vendrell, M. Characterization of two divergent endo-b-1,4-glucanase cDNA clones highly expressed in the nonclimateric strawberry fruit. Plant Physiol. 1999, 119, 1415–1421. [Google Scholar] [CrossRef] [PubMed]
- Trainotti, L.; Spolaore, S.; Pavanello, A.; Baldan, B.; Casadoro, G. A novel E-type endo-b-1,4-glucanase with a putative cellulose-binding domain is highly expressed in ripening strawberry fruits. Plant Mol. Biol. 1999, 40, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Fisher, R.L.; Bennett, A.B. Role of cell wall hydrolases in frui ripening. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1991, 42, 675–703. [Google Scholar] [CrossRef]
- Ferrarese, L.; Trainotti, L.; Moretto, P.; Polverino de Laureto, P.; Rascio, N.; Casadoro, G. Differential ethylene-inducible expression of cellulase in pepper plants. Plant Mol. Biol. 1995, 29, 735–747. [Google Scholar] [CrossRef]
- Harpster, M.H.; Lee, K.Y.; Dunsmuir, P. Isolation and characterization of a gene encoding endo-b-1,4-glucanase from pepper (Capsicum annuum L.). Plant Mol. Biol. 1997, 33, 47–59. [Google Scholar] [CrossRef] [PubMed]
- Mercado, J.A.; Trainotti, L.; Jiménez-Bermúdez, L.; Santiago-Doménech, N.; Posé, S.; Donolli, R.; Barceló, M.; Casadoro, G.; Pliego-Alfaro, F.; Quesada, M.A. Evaluation of the role of the endo-β-(1,4)-glucanase gene FaEG3 in strawberry fruit softening. Postharvest Biol. Technol. 2010, 55, 8–14. [Google Scholar] [CrossRef]
- Parra-Palma, C.; Ubeda, C.; Gil, M.; Ramos, P.; Castro, R.I.; Morales-Quintana, L. Comparative study of the volatile organic compounds of four strawberry cultivars and it relation to alcohol acyltransferase enzymatic activity. Sci. Hortic. 2019, 251, 65–72. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef]
- Kumar, S.; Nei, M.; Dudley, J.; Tamura, K. MEGA: A biologistcentric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 2008, 9, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Morales-Quintana, L.; Fuentes, L.; Gaete-Eastman, C.; Herrera, R.; Moya-León, M.A. Structural characterization and substrate specificity of VpAAT1 protein related to ester biosynthesis in mountain papaya fruit. J. Mol. Graph. Model. 2011, 29, 635–642. [Google Scholar] [CrossRef] [PubMed]
- Šali, A.; Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 1993, 234, 779–815. [Google Scholar] [CrossRef] [PubMed]
- Jorgensen, W.L.; Maxwell, D.S.; Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc. 1996, 118, 11225–11236. [Google Scholar] [CrossRef]
- Wang, C.X.; Wang, L.; McQueen-Mason, S.J.; Pritchard, J.; Thomas, C.R. pH and expansin action on single suspension-cultured tomato (Lycopersicon esculentum) cells. J. Plant Res. 2008, 121, 527–534. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R.A.; MacArthur, M.W.; Moss, D.S.; Thornton, J.M. PROCHECK: A program to check the stereochemical quality of protein structures. J. Appl. Cryst. 1993, 26, 283–291. [Google Scholar] [CrossRef]
- Zhang, Y.; Skolnick, J. Scoring function for automated assessment of protein structure template quality. Proteins 2004, 57, 702–710. [Google Scholar] [CrossRef] [PubMed]
- Sippl, M.J. Recognition of errors in three-dimensional structures of proteins. Proteins 1993, 17, 355–362. [Google Scholar] [CrossRef] [PubMed]
- Wiederstein, M.; Sippl, M.J. ProSA-web: Interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 2007, 35, W407–W410. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [PubMed]
- Georgelis, N.; Yennawar, N.H.; Cosgrove, D.J. Structural basis for entropy-driven cellulose binding by a type—A cellulose-binding module (CBM) and bacterial expansin. Proc. Natl. Acad. Sci. USA 2012, 109, 14830–14835. [Google Scholar] [CrossRef] [PubMed]
- Valenzuela-Riffo, F.; Gaete-Eastman, C.; Stappung, Y.; Lizana, R.; Herrera, R.; Moya-León, M.A.; Morales-Quintana, L. A Comparative in silico study of the differences in the structure and ligand interaction properties of three alpha-expansin proteins from Fragaria chiloensis fruit”. J. Biomol. Struct. Dyn. 2018. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]

| Complex | Binding Energy (Autodock Vina Score) | |
|---|---|---|
| FcEG1 | cellodextrin 8-mer | −7.5 a ± 0.4 |
| XXXGXXXG | −6.9 ab ± 0.5 | |
| XXFGXXFG | −5.9 b ± 0.5 | |
| Complex | ΔHvdWMM (kcal mol−1) | ΔHelecMM (kcal mol−1) | ΔGsol–pol (kcal mol−1) | ΔGsol–npol (kcal mol−1) | ΔGbind (kcal mol−1) | |
|---|---|---|---|---|---|---|
| FcEG1 | Cellulose | −67.21 | −71.11 | 67.30 | −52.30 | −123.32 ± 9 a |
| XXXGXXXG | −55.21 | −58.14 | 63.10 | −25.06 | −75.31 ± 5 ab | |
| XXFGXXFG | −52.85 | −26.51 | 46.68 | −30.97 | −63.65 ± 8 b | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jara, K.; Castro, R.I.; Ramos, P.; Parra-Palma, C.; Valenzuela-Riffo, F.; Morales-Quintana, L. Molecular Insights into FaEG1, a Strawberry Endoglucanase Enzyme Expressed during Strawberry Fruit Ripening. Plants 2019, 8, 140. https://doi.org/10.3390/plants8060140
Jara K, Castro RI, Ramos P, Parra-Palma C, Valenzuela-Riffo F, Morales-Quintana L. Molecular Insights into FaEG1, a Strawberry Endoglucanase Enzyme Expressed during Strawberry Fruit Ripening. Plants. 2019; 8(6):140. https://doi.org/10.3390/plants8060140
Chicago/Turabian StyleJara, Karla, Ricardo I. Castro, Patricio Ramos, Carolina Parra-Palma, Felipe Valenzuela-Riffo, and Luis Morales-Quintana. 2019. "Molecular Insights into FaEG1, a Strawberry Endoglucanase Enzyme Expressed during Strawberry Fruit Ripening" Plants 8, no. 6: 140. https://doi.org/10.3390/plants8060140
APA StyleJara, K., Castro, R. I., Ramos, P., Parra-Palma, C., Valenzuela-Riffo, F., & Morales-Quintana, L. (2019). Molecular Insights into FaEG1, a Strawberry Endoglucanase Enzyme Expressed during Strawberry Fruit Ripening. Plants, 8(6), 140. https://doi.org/10.3390/plants8060140








