Sequencing and Structural Analysis of the Complete Chloroplast Genome of the Medicinal Plant Lycium chinense Mill
Abstract
:1. Introduction
2. Results
2.1. CP Genome Organization and Gene Content
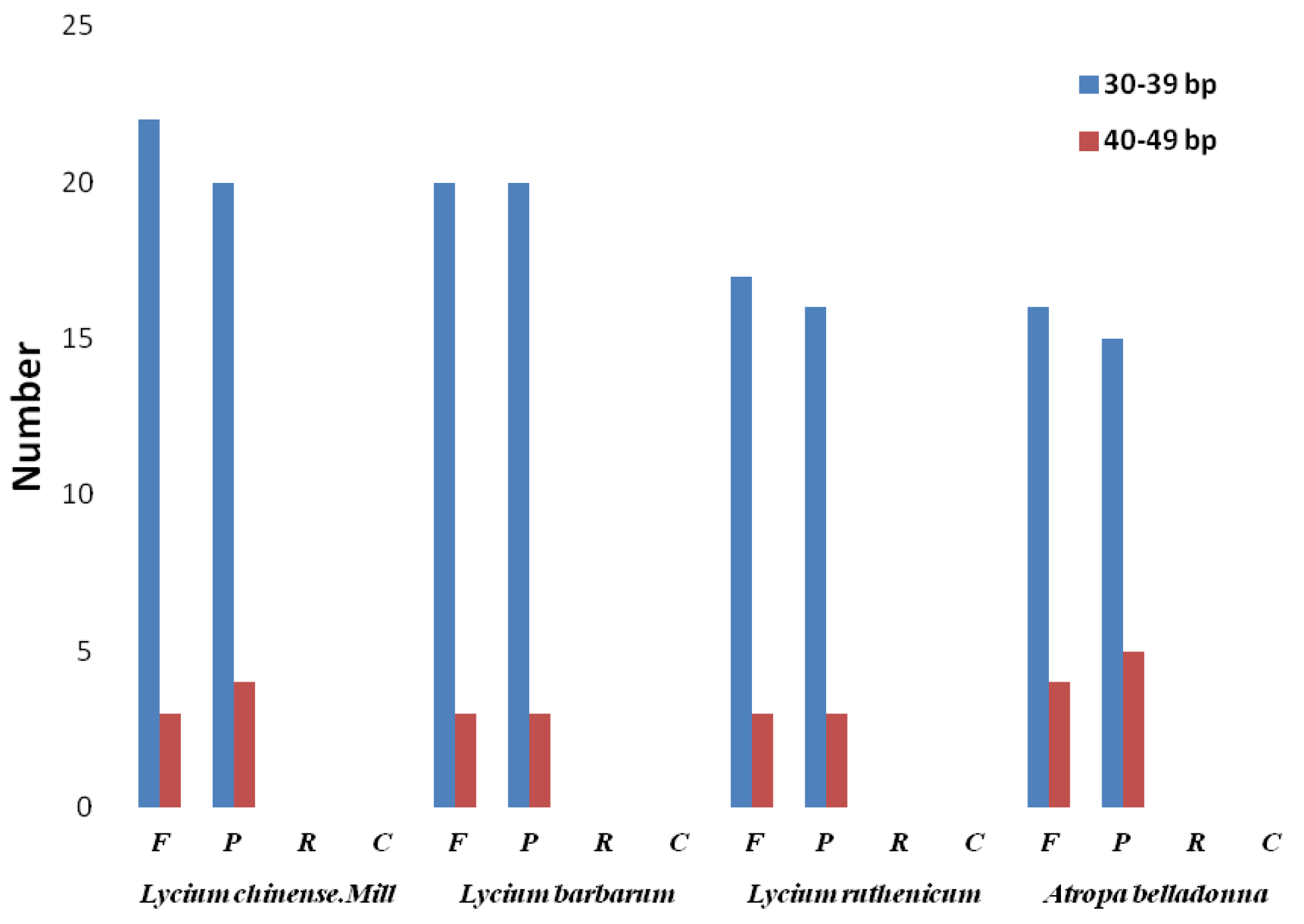
2.2. Repeat Structure and Simple Sequence Repeat (SSR) Analysis
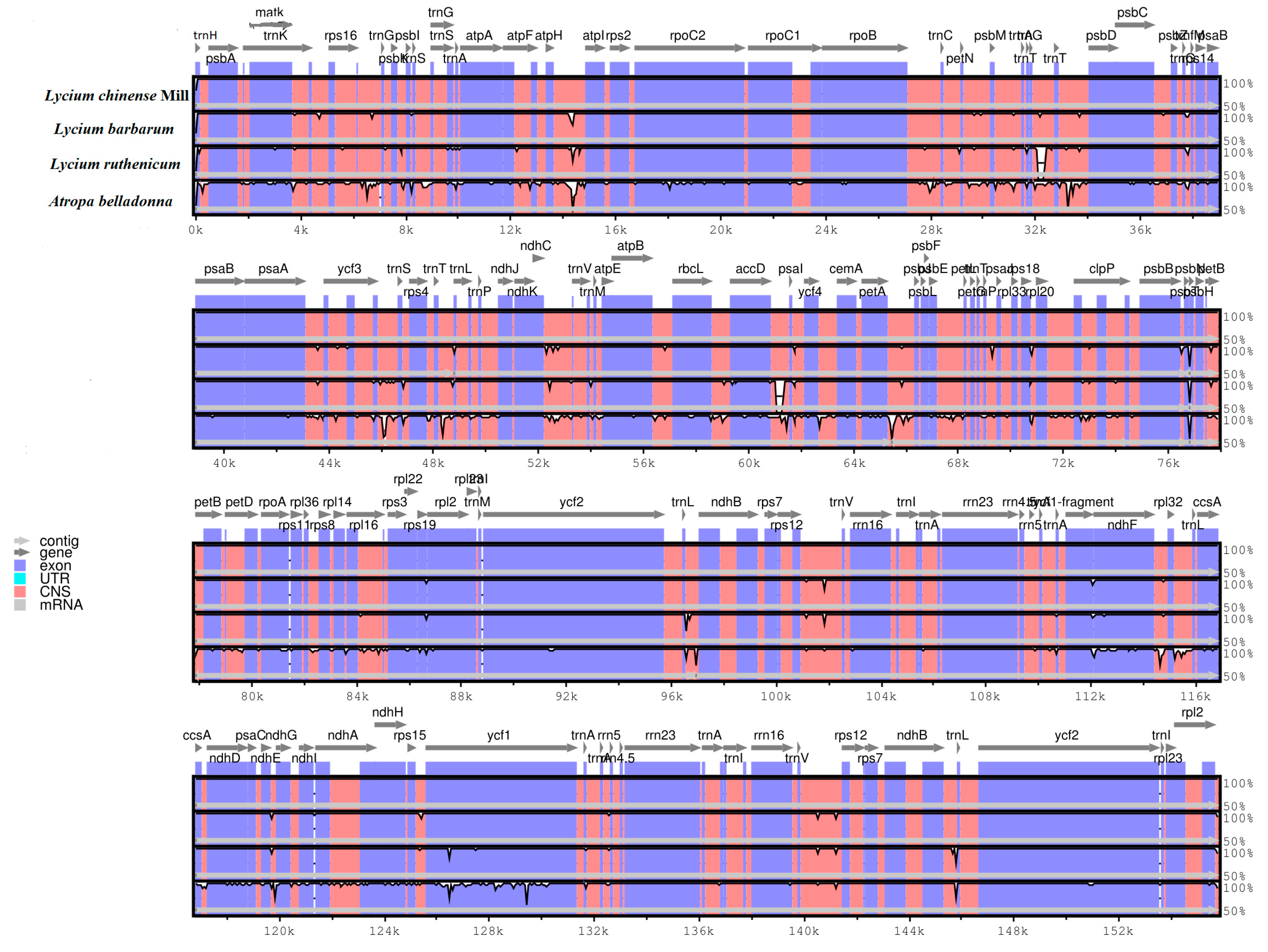
2.3. Comparative Chloroplast Genomic Analysis
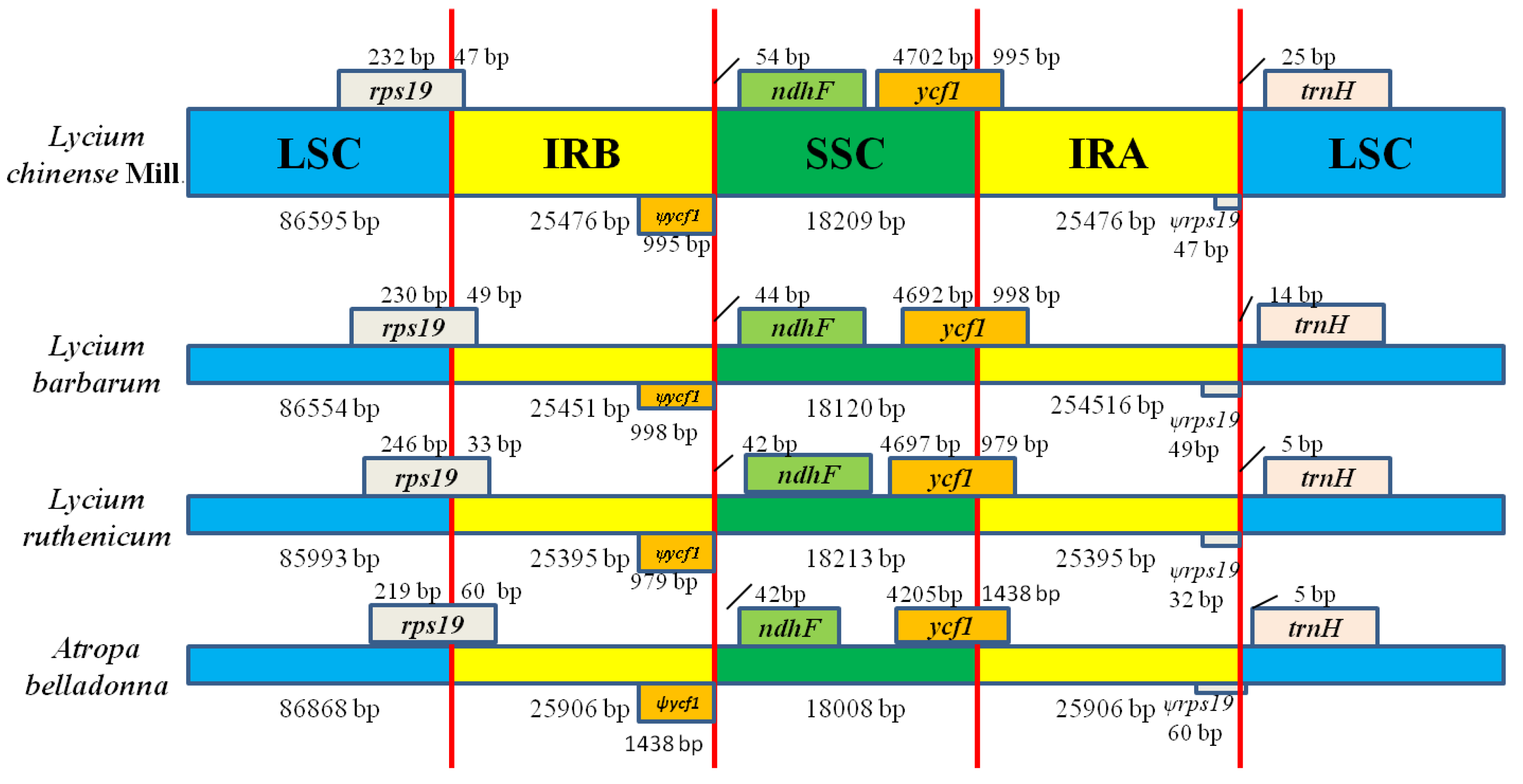
2.4. IR Contraction and Expansion in the L. chinense. CP Genome.
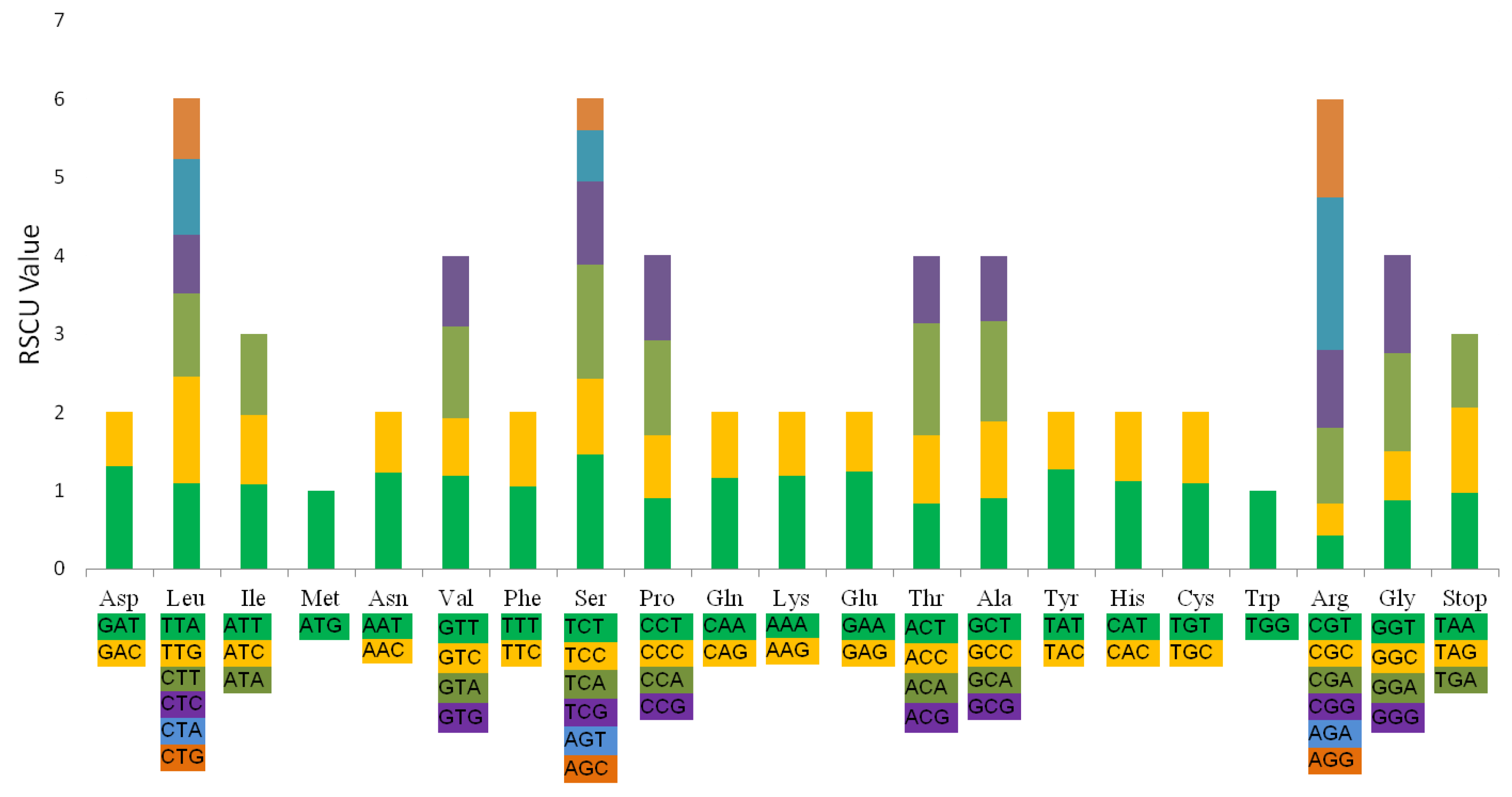
2.5. Codon Usage and RNA Editing Sites
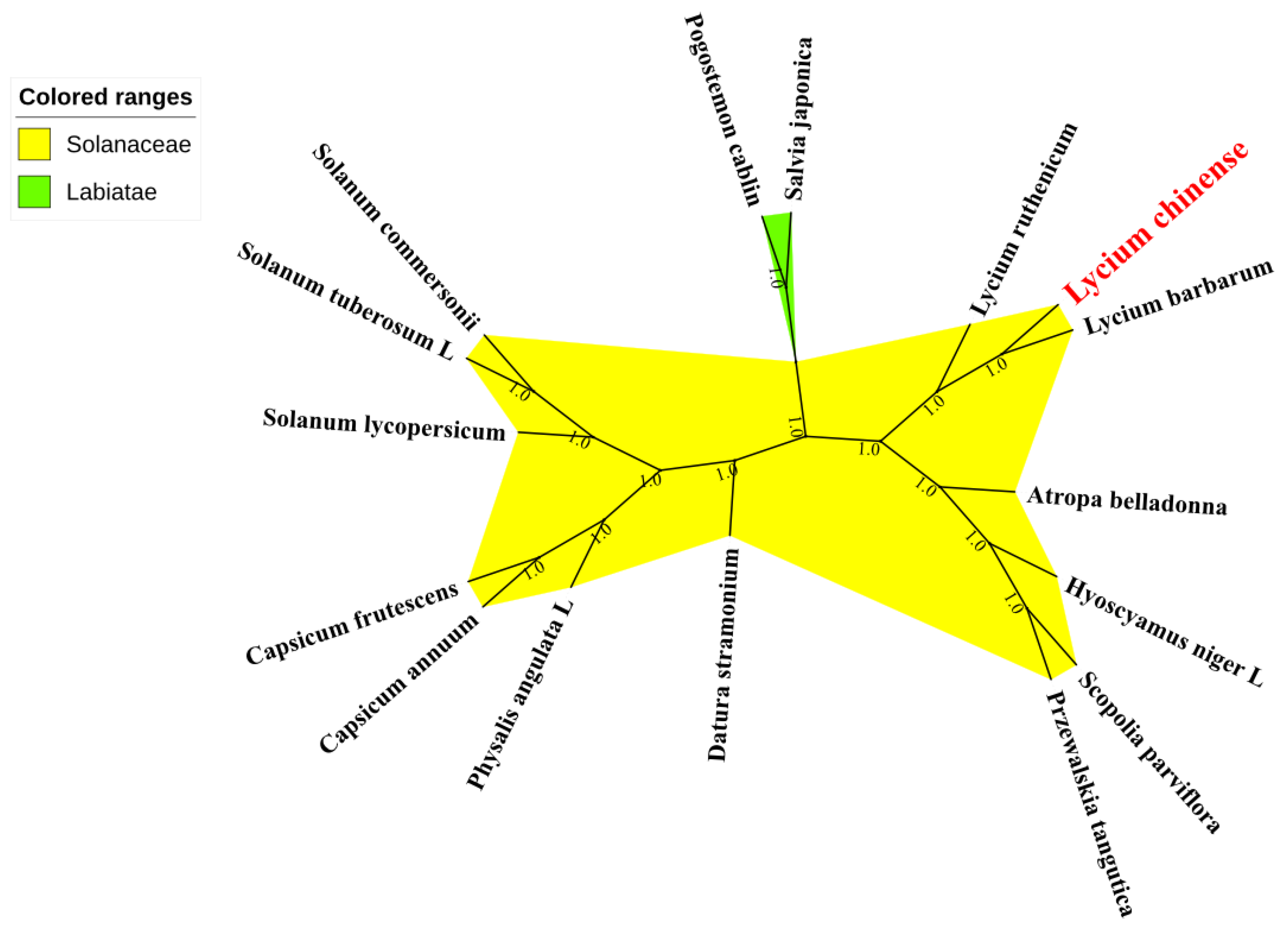
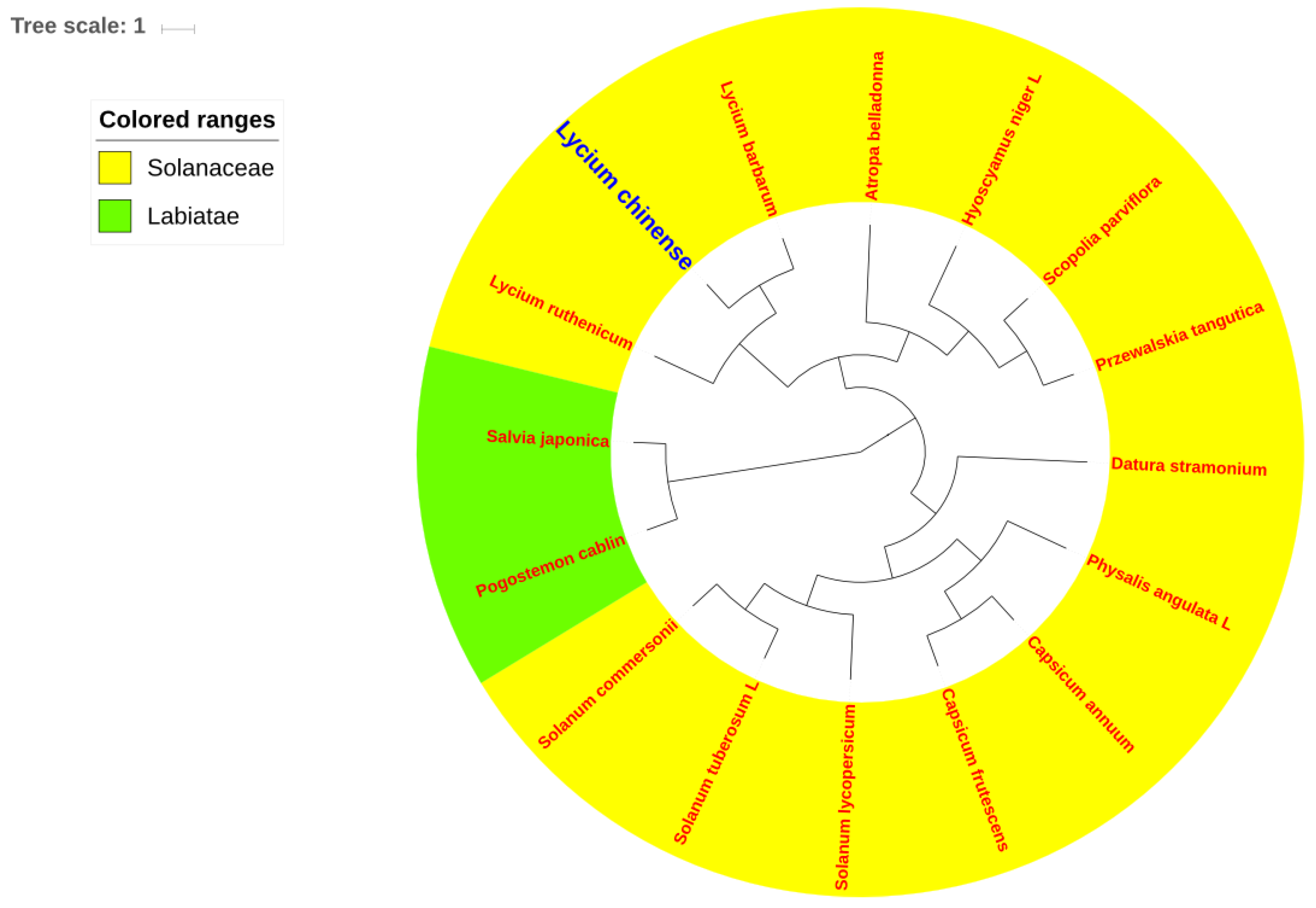
2.6. Phylogenetic Analysis
3. Discussion
4. Materials and Methods
4.1. Plant Material and DNA Extraction
4.2. Chloroplast Genome Sequencing and Assembly
4.3. Gene Annotation and Genome Structure
4.4. Genome Comparison and Phylogenetic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Potterat, O. Goji (Lycium barbarum and L. chinense): Phytochemistry, pharmacology and safety in the perspective of traditional uses and recent popularity. Planta Med. 2010, 76, 7–19. [Google Scholar] [CrossRef]
- Kim, M.H.; Kim, E.J.; Choi, Y.Y.; Hong, J.; Yang, W.M. Lycium chinense Improves Post-Menopausal Obesity via Regulation of PPAR-gamma and Estrogen Receptor-alpha/beta Expressions. Am. J. Chin. Med. 2017, 45, 269–282. [Google Scholar] [CrossRef]
- Qin, X.; Yamauchi, R.; Aizawa, K.; Inakuma, T.; Kato, K. Structural features of arabinogalactan-proteins from the fruit of Lycium chinense Mill. Carbohydr. Res. 2001, 333, 79–85. [Google Scholar] [CrossRef]
- Chinese Pharmacopoeia Commission. Chinese Pharmacopoeia; China Medical Science Press: Beijing, China, 2015. [Google Scholar]
- Yao, R.; Heinrich, M.; Weckerle, C.S. The genus Lycium as food and medicine: A botanical, ethnobotanical and historical review. J. Ethnopharmacol. 2018, 212, 50–66. [Google Scholar] [CrossRef]
- Zhang, J.X.; Guan, S.H.; Feng, R.H.; Wang, Y.; Wu, Z.Y.; Zhang, Y.B.; Chen, X.H.; Bi, K.S.; Guo, D.A. Neolignanamides, lignanamides, and other phenolic compounds from the root bark of Lycium chinense. J. Nat. Prod. 2013, 76, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Turchetto, C.; Fagundes, N.J.; Segatto, A.L.; Kuhlemeier, C.; Solis Neffa, V.G.; Speranza, P.R.; Bonatto, S.L.; Freitas, L.B. Diversification in the South American Pampas: The genetic and morphological variation of the widespread Petunia axillaris complex (Solanaceae). Mol. Ecol. 2014, 23, 374–389. [Google Scholar] [CrossRef]
- Ni, L.L.; Zhao, Z.L.; Lu, J.N. DNA barcoding construction of medicinal plants in genus Lycium L. based on multiple genomic segments. Chin. Tradit. Herb. Drugs 2016, 47, 5. [Google Scholar]
- Hebert, P.D.; Cywinska, A.; Ball, S.L.; de Waard, J.R. Biological identifications through DNA barcodes. Proc. Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef]
- Nielsen, A.Z.; Ziersen, B.; Jensen, K.; Lassen, L.M.; Olsen, C.E.; Moller, B.L.; Jensen, P.E. Redirecting photosynthetic reducing power toward bioactive natural product synthesis. ACS Synth. Biol. 2013, 2, 308–315. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Dang, Y.; Li, Q.; Lu, J.; Li, X.; Wang, Y. Complete chloroplast genome sequence of poisonous and medicinal plant Datura stramonium: Organizations and implications for genetic engineering. PLoS ONE 2014, 9, e110656. [Google Scholar] [CrossRef]
- Sanchez-Puerta, M.V.; Abbona, C.C. The chloroplast genome of Hyoscyamus niger and a phylogenetic study of the tribe Hyoscyameae (Solanaceae). PLoS ONE 2014, 9, e98353. [Google Scholar] [CrossRef] [PubMed]
- Shaw, J.; Lickey, E.B.; Schilling, E.E.; Small, R.L. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: The tortoise and the hare III. Am. J. Bot. 2007, 94, 275–288. [Google Scholar] [CrossRef] [PubMed]
- Shaw, J.; Lickey, E.B.; Beck, J.T.; Farmer, S.B.; Liu, W.; Miller, J.; Siripun, K.C.; Winder, C.T.; Schilling, E.E.; Small, R.L. The tortoise and the hare II: Relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. Am. J. Bot. 2005, 92, 142–166. [Google Scholar] [CrossRef]
- Wang, G.; Du, X.; Ji, J.; Guan, C.; Li, Z.; Josine, T.L. De novo characterization of the Lycium chinense Mill. leaf transcriptome and analysis of candidate genes involved in carotenoid biosynthesis. Gene 2015, 555, 458–463. [Google Scholar] [CrossRef]
- Amiryousefi, A.H.J.; Poczai, P. The chloroplast genome sequence of bittersweet (Solanum dulcamara): Plastid genome structure evolution in Solanaceae. PLoS ONE 2018, 13, 23. [Google Scholar] [CrossRef] [PubMed]
- Cho, K.S.; Cheon, K.S.; Hong, S.Y.; Cho, J.H.; Im, J.S.; Mekapogu, M.; Yu, Y.S.; Park, T.H. Complete chloroplast genome sequences of Solanum commersonii and its application to chloroplast genotype in somatic hybrids with Solanum tuberosum. Plant Cell Rep. 2016, 35, 2113–2123. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Qian, J.; Li, X.; Sun, Z.; Xu, X.; Chen, S. Complete Chloroplast Genome of Medicinal Plant Lonicera japonica: Genome Rearrangement, Intron Gain and Loss, and Implications for Phylogenetic Studies. Molecules 2017, 22. [Google Scholar] [CrossRef] [PubMed]
- Xiang, B.; Li, X.; Qian, J.; Wang, L.; Ma, L.; Tian, X.; Wang, Y. The Complete Chloroplast Genome Sequence of the Medicinal Plant Swertia mussotii Using the PacBio RS II Platform. Molecules 2016, 21. [Google Scholar] [CrossRef]
- Clegg, M.T.; Gaut, B.S.; Learn, G.H., Jr.; Morton, B.R. Rates and patterns of chloroplast DNA evolution. Proc. Natl. Acad. Sci. USA 1994, 91, 6795–6801. [Google Scholar] [CrossRef]
- Morton, B.R. Selection on the codon bias of chloroplast and cyanelle genes in different plant and algal lineages. J. Mol. Evol. 1998, 46, 449–459. [Google Scholar] [CrossRef]
- Shen, X.; Guo, S.; Yin, Y.; Zhang, J.; Yin, X.; Liang, C.; Wang, Z.; Huang, B.; Liu, Y.; Xiao, S.; et al. Complete Chloroplast Genome Sequence and Phylogenetic Analysis of Aster tataricus. Molecules 2018, 23. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Liu, J.; Luo, L.; Wei, X.; Zhang, J.; Qi, Y.; Zhang, B.; Liu, H.; Xiao, P. Complete chloroplast genome sequences of Schisandra chinensis: Genome structure, comparative analysis, and phylogenetic relationship of basal angiosperms. Sci. China Life Sci. 2017, 60, 1286–1290. [Google Scholar] [CrossRef] [PubMed]
- Qian, J.; Song, J.; Gao, H.; Zhu, Y.; Xu, J.; Pang, X.; Yao, H.; Sun, C.; Li, X.; Li, C.; et al. The complete chloroplast genome sequence of the medicinal plant Salvia miltiorrhiza. PLoS ONE 2013, 8, e57607. [Google Scholar] [CrossRef]
- Khakhlova, O.; Bock, R. Elimination of deleterious mutations in plastid genomes by gene conversion. Plant J. Cell Mol. Biol. 2006, 46, 85–94. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Li, J.; Zhang, H.; Cai, B.; Gao, Z.; Qiao, Y.; Mi, L. The complete chloroplast genome sequence of strawberry (Fragaria x ananassa Duch.) and comparison with related species of Rosaceae. PeerJ 2017, 5, e3919. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zheng, Y.; Liu, S.; Zhong, Y.; Wu, Y.; Li, J.; Xu, L.A.; Xu, M. The complete chloroplast genome of Cinnamomum camphora and its comparison with related Lauraceae species. PeerJ 2017, 5, e3820. [Google Scholar] [CrossRef] [PubMed]
- Kong, W.Q.; Yang, J.H. The complete chloroplast genome sequence of Morus cathayana and Morus multicaulis, and comparative analysis within genus Morus L. PeerJ 2017, 5, e3037. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Du, L.; Liu, A.; Chen, J.; Wu, L.; Hu, W.; Zhang, W.; Kim, K.; Lee, S.C.; Yang, T.J.; et al. The Complete Chloroplast Genome Sequences of Five Epimedium Species: Lights into Phylogenetic and Taxonomic Analyses. Front. Plant Sci. 2016, 7, 306. [Google Scholar] [CrossRef]
- Reginato, M.; Neubig, K.M.; Majure, L.C.; Michelangeli, F.A. The first complete plastid genomes of Melastomataceae are highly structurally conserved. PeerJ 2016, 4, e2715. [Google Scholar] [CrossRef]
- Chang, C.C.; Lin, H.C.; Lin, I.P.; Chow, T.Y.; Chen, H.H.; Chen, W.H.; Cheng, C.H.; Lin, C.Y.; Liu, S.M.; Chang, C.C.; et al. The chloroplast genome of Phalaenopsis aphrodite (Orchidaceae): Comparative analysis of evolutionary rate with that of grasses and its phylogenetic implications. Mol. Biol. Evol. 2006, 23, 279–291. [Google Scholar] [CrossRef]
- Pan, I.C.; Liao, D.C.; Wu, F.H.; Daniell, H.; Singh, N.D.; Chang, C.; Shih, M.C.; Chan, M.T.; Lin, C.S. Complete chloroplast genome sequence of an orchid model plant candidate: Erycina pusilla apply in tropical Oncidium breeding. PLoS ONE 2012, 7, e34738. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.H.; Chan, M.T.; Liao, D.C.; Hsu, C.T.; Lee, Y.W.; Daniell, H.; Duvall, M.R.; Lin, C.S. Complete chloroplast genome of Oncidium Gower Ramsey and evaluation of molecular markers for identification and breeding in Oncidiinae. BMC Plant Biol. 2010, 10, 68. [Google Scholar] [CrossRef]
- Tsudzuki, T.; Wakasugi, T.; Sugiura, M. Comparative analysis of RNA editing sites in higher plant chloroplasts. J. Mol. Evol. 2001, 53, 327–332. [Google Scholar] [CrossRef] [PubMed]
- Yisilam, G.; Mamut, R.; Li, J.; Li, P.; Fu, C. Characterization of the complete chloroplast genome of Lycium ruthenicum (Solanaceae). Mitochondrial DNA Part B 2018, 3, 361–362. [Google Scholar] [CrossRef]
- Jia, G.; Xin, G.; Ren, X.; Du, X.; Liu, H.; Hao, N.; Wang, C.; Ni, X.; Liu, W. Characterization of the complete chloroplast genome of Lycium barbarum (Solanales: Solanaceae), a unique economic plant to China. Mitochondrial DNA Part B 2018, 3, 1062–1063. [Google Scholar] [CrossRef]
- Li, R.; Ma, P.F.; Wen, J.; Yi, T.S. Complete sequencing of five araliaceae chloroplast genomes and the phylogenetic implications. PLoS ONE 2013, 8, e78568. [Google Scholar] [CrossRef]
- Wang, R.J.; Cheng, C.L.; Chang, C.C.; Wu, C.L.; Su, T.M.; Chaw, S.M. Dynamics and evolution of the inverted repeat-large single copy junctions in the chloroplast genomes of monocots. BMC Evol. Biol. 2008, 8, 36. [Google Scholar] [CrossRef]
- Kahlau, S.; Aspinall, S.; Gray, J.C.; Bock, R. Sequence of the tomato chloroplast DNA and evolutionary comparison of solanaceous plastid genomes. J. Mol. Evol. 2006, 63, 194–207. [Google Scholar] [CrossRef]
- An, S.M.; Noh, J.H.; Choi, D.H.; Lee, J.H.; Yang, E.C. Repeat region absent in mitochondrial genome of tube-dwelling diatom Berkeleya fennica (Naviculales, Bacillariophyceae). Mitochondrial DNA Part Adna Mapp. Seq. Anal. 2016, 27, 2137–2138. [Google Scholar] [CrossRef]
- Freyer, R.; Hoch, B.; Neckermann, K.; Maier, R.M.; Kossel, H. RNA editing in maize chloroplasts is a processing step independent of splicing and cleavage to monocistronic mRNAs. Plant J. Cell Mol. Biol. 1993, 4, 621–629. [Google Scholar] [CrossRef]
- De Santana Lopes, A.; Pacheco, T.G.; Santos, K.G.D.; Vieira, L.D.N.; Guerra, M.P.; Nodari, R.O.; de Souza, E.M.; de Oliveira Pedrosa, F.; Rogalski, M. The Linum usitatissimum L. plastome reveals atypical structural evolution, new editing sites, and the phylogenetic position of Linaceae within Malpighiales. Plant Cell Rep. 2018, 37, 307–328. [Google Scholar] [CrossRef]
- Hirose, T.; Sugiura, M. Both RNA editing and RNA cleavage are required for translation of tobacco chloroplast ndhD mRNA: A possible regulatory mechanism for the expression of a chloroplast operon consisting of functionally unrelated genes. EMBO J. 1997, 16, 6804–6811. [Google Scholar] [CrossRef]
- Maier, R.M.; Zeltz, P.; Kossel, H.; Bonnard, G.; Gualberto, J.M.; Grienenberger, J.M. RNA editing in plant mitochondria and chloroplasts. Plant Mol. Biol. 1996, 32, 343–365. [Google Scholar] [CrossRef]
- Raman, G.; Park, S. The Complete Chloroplast Genome Sequence of Ampelopsis: Gene Organization, Comparative Analysis, and Phylogenetic Relationships to Other Angiosperms. Front. Plant Sci. 2016, 7, 341. [Google Scholar] [CrossRef]
- Jain, B.P.; Chauhan, P.; Tanti, G.K.; Singarapu, N.; Ghaskadbi, S.; Goswami, S.K. Tissue specific expression of SG2NA is regulated by differential splicing, RNA editing and differential polyadenylation. Gene 2015, 556, 119–126. [Google Scholar] [CrossRef]
- Corneille, S.; Lutz, K.; Maliga, P. Conservation of RNA editing between rice and maize plastids: Are most editing events dispensable? Mol. Gen. Genet. MGG 2000, 264, 419–424. [Google Scholar] [CrossRef]
- Robinson, A.C.; Castaneda, C.A.; Schlessman, J.L.; Garcia-Moreno, E.B. Structural and thermodynamic consequences of burial of an artificial ion pair in the hydrophobic interior of a protein. Proc. Natl. Acad. Sci. USA 2014, 111, 11685–11690. [Google Scholar] [CrossRef]
- Loladze, V.V.; Ermolenko, D.N.; Makhatadze, G.I. Thermodynamic consequences of burial of polar and non-polar amino acid residues in the protein interior. J. Mol. Biol. 2002, 320, 343–357. [Google Scholar] [CrossRef]
- Chen, C.; Zhou, P.; Choi, Y.A.; Huang, S.; Gmitter, F.G., Jr. Mining and characterizing microsatellites from citrus ESTs. Theor. Appl. Genet. 2006, 112, 1248–1257. [Google Scholar] [CrossRef]
- Huang, H.; Shi, C.; Liu, Y.; Mao, S.Y.; Gao, L.Z. Thirteen Camellia chloroplast genome sequences determined by high-throughput sequencing: Genome structure and phylogenetic relationships. BMC Evol. Biol. 2014, 14, 151. [Google Scholar] [CrossRef]
- Zhao, Y.; Yin, J.; Guo, H.; Zhang, Y.; Xiao, W.; Sun, C.; Wu, J.; Qu, X.; Yu, J.; Wang, X.; et al. The complete chloroplast genome provides insight into the evolution and polymorphism of Panax ginseng. Front. Plant Sci. 2014, 5, 696. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, G.L.; Dorman, H.E.; Buchanan, A.; Challagundla, L.; Wallace, L.E. A review of the prevalence, utility, and caveats of using chloroplast simple sequence repeats for studies of plant biology. Appl. Plant Sci. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Vieira Ldo, N.; Faoro, H.; Rogalski, M.; Fraga, H.P.; Cardoso, R.L.; de Souza, E.M.; de Oliveira Pedrosa, F.; Nodari, R.O.; Guerra, M.P. The complete chloroplast genome sequence of Podocarpus lambertii: Genome structure, evolutionary aspects, gene content and SSR detection. PLoS ONE 2014, 9, e90618. [Google Scholar] [CrossRef] [PubMed]
- Firetti, F.; Zuntini, A.R.; Gaiarsa, J.W.; Oliveira, R.S.; Lohmann, L.G.; Van Sluys, M.A. Complete chloroplast genome sequences contribute to plant species delimitation: A case study of the Anemopaegma species complex. Am. J. Bot. 2017, 104, 1493–1509. [Google Scholar] [CrossRef]
- Wang, W.; Chen, S.; Zhang, X. Whole-Genome Comparison Reveals Divergent IR Borders and Mutation Hotspots in Chloroplast Genomes of Herbaceous Bamboos (Bambusoideae: Olyreae). Molecules 2018, 23. [Google Scholar] [CrossRef]
- Zhou, T.; Wang, J.; Jia, Y.; Li, W.; Xu, F.; Wang, X. Comparative Chloroplast Genome Analyses of Species in Gentiana section Cruciata (Gentianaceae) and the Development of Authentication Markers. Int. J. Mol. Sci. 2018, 19. [Google Scholar] [CrossRef]
- De Las Rivas, J.; Lozano, J.J.; Ortiz, A.R. Comparative analysis of chloroplast genomes: Functional annotation, genome-based phylogeny, and deduced evolutionary patterns. Genome Res. 2002, 12, 567–583. [Google Scholar] [CrossRef]
- Mocan, A.; Vlase, L.; Raita, O.; Hanganu, D.; Paltinean, R.; Dezsi, S.; Gheldiu, A.M.; Oprean, R.; Crisan, G. Comparative studies on antioxidant activity and polyphenolic content of Lycium barbarum L. and Lycium chinense Mill. leaves. Pak. J. Pharm. Sci. 2015, 28, 1511–1515. [Google Scholar]
- Skenderidis, P.; Lampakis, D.; Giavasis, I.; Leontopoulos, S.; Petrotos, K.; Hadjichristodoulou, C.; Tsakalof, A. Chemical Properties, Fatty-Acid Composition, and Antioxidant Activity of Goji Berry (Lycium barbarum L. and Lycium chinense Mill.) Fruits. Antioxidants 2019, 8. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Jackman, S.D.; Vandervalk, B.P.; Mohamadi, H.; Chu, J.; Yeo, S.; Hammond, S.A.; Jahesh, G.; Khan, H.; Coombe, L.; Warren, R.L.; et al. ABySS 2.0: Resource-efficient assembly of large genomes using a Bloom filter. Genome Res. 2017, 27, 768–777. [Google Scholar] [CrossRef]
- Tillich, M.; Lehwark, P.; Pellizzer, T.; Ulbricht-Jones, E.S.; Fischer, A.; Bock, R.; Greiner, S. GeSeq—Versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 2017, 45, W6–W11. [Google Scholar] [CrossRef]
- Lohse, M.; Drechsel, O.; Kahlau, S.; Bock, R. OrganellarGenomeDRAW—A suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res. 2013, 41, W575–W581. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Kumar, S.; Nei, M.; Dudley, J.; Tamura, K. MEGA: A biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 2008, 9, 299–306. [Google Scholar] [CrossRef]
- Mower, J.P. The PREP suite: Predictive RNA editors for plant mitochondrial genes, chloroplast genes and user-defined alignments. Nucleic Acids Res. 2009, 37, W253–W259. [Google Scholar] [CrossRef]
- Yang, X.M.; Sun, J.T.; Xue, X.F.; Zhu, W.C.; Hong, X.Y. Development and characterization of 18 novel EST-SSRs from the western flower Thrips, Frankliniella occidentalis (Pergande). Int. J. Mol. Sci. 2012, 13, 2863–2876. [Google Scholar] [CrossRef]
- Kurtz, S.; Choudhuri, J.V.; Ohlebusch, E.; Schleiermacher, C.; Stoye, J.; Giegerich, R. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef]
- Mayor, C.; Brudno, M.; Schwartz, J.R.; Poliakov, A.; Rubin, E.M.; Frazer, K.A.; Pachter, L.S.; Dubchak, I. VISTA: Visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 2000, 16, 1046–1047. [Google Scholar] [CrossRef]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2017. [Google Scholar] [CrossRef] [PubMed]

| Region | Positions | T/U (%) | C (%) | A (%) | G (%) | Total (bp) |
|---|---|---|---|---|---|---|
| LSC | 32.7 | 18.3 | 31.4 | 17.5 | 86,595 | |
| IRB | 28.4 | 20.7 | 28.5 | 22.4 | 25,476 | |
| SSC | 33.9 | 16.8 | 33.7 | 15.5 | 18,209 | |
| IRA | 28.5 | 22.4 | 28.4 | 20.7 | 25,476 | |
| Total | 31.5 | 19.2 | 30.7 | 18.6 | 155,756 | |
| CDS | 31.3 | 17.9 | 30.5 | 20.3 | 79,700 | |
| 1st position | 25.0 | 18.6 | 30.9 | 25.3 | 26,567 | |
| 2nd position | 34.0 | 19.6 | 28.5 | 18.3 | 26,567 | |
| 3rd position | 35.0 | 15.5 | 32.7 | 17.6 | 26,566 |
| Classification of Genes | Gene Names | Number |
|---|---|---|
| Photosystem I | psaA, psaB, psaC, psaI, psaJ | 5 |
| Photosystem II | psbA, psbB, psbC, psbD, psbE, psbF, psbH, psbI, psbJ, psbK, psbL, psbM, psbN, psbT, psbZ | 15 |
| Cytochrome b/f complex | petA, petB *, petD *, petG, petL, petN | 6 |
| ATP synthase | atpA, atpB, atpE, atpF, atpH, atpI | 6 |
| NADH dehydrogenase | ndhA *, ndhB * (×2), ndhC, ndhD, ndhE, ndhF, ndhG, ndhH, ndhI, ndhJ, ndhK | 12 (1) |
| RubisCO large subunit | rbcL | 1 |
| RNA polymerase | rpoA, rpoB, rpoC1, rpoC2 | 4 |
| Ribosomal proteins (SSC) | rps2, rps3, rps4, rps7 (×2), rps8, rps11, rps12 ** (×2), rps14, rps15, rps16 *, rps18, rps19 | 14 (2) |
| Ribosomal proteins (LSC) | rpl2 (×2), rpl14, rpl16, rpl20, rpl22, rpl23 (×2), rpl32, rpl33, rpl36 | 11 |
| Ribosomal RNAs | rrn 4.5 (×2), rrn 5 (×2), rrn 16 (×2), rrn 23 (×2) | 8 (4) |
| Protein of unkown function | ycf1 (×2), ycf2 (×2), ycf3 **, ycf4 | 6 (2) |
| Transfer RNAs | 37 tRNAs (8 contain an intron, 7 in the inverted repeats region) | 37 (7) |
| Other genes | accD, ccsA, cemA, clpP, matK | 5 |
| Total | 130 |
| Gene | Location | Exon I (bp) | Intron I (bp) | Exon II (bp) | Intron II (bp) | Exon III (bp) |
|---|---|---|---|---|---|---|
| atpF | LSC | 410 | 704 | 145 | ||
| clpP | LSC | 228 | 640 | 292 | 808 | 71 |
| ndhA | SSC | 539 | 1154 | 553 | ||
| ndhB | IR | 777 | 679 | 756 | ||
| petB | LSC | 6 | 750 | 642 | ||
| petD | LSC | 8 | 742 | 475 | ||
| rpl16 | LSC | 396 | 1016 | 9 | ||
| rpl2 | IR | 434 | 666 | 391 | ||
| rpoC1 | LSC | 1616 | 737 | 430 | ||
| rps12 | IR | 26 | 536 | 232 | ||
| rps16 | LSC | 227 | 822 | 40 | ||
| trnA-UGC | IR | 38 | 681 | 35 | ||
| trnI-GAU | IR | 34 | 717 | 37 | ||
| trnK-UUU | LSC | 36 | 2513 | 37 | ||
| trnL-UAA | LSC | 35 | 497 | 50 | ||
| trnV-UAC | LSC | 35 | 565 | 38 | ||
| ycf3 | LSC | 154 | 756 | 229 | 744 | 124 |
| SSR Type | Repeat Unit | Amount | Ratio (%) |
|---|---|---|---|
| Mono | A/T | 106 | 99.1 |
| C/G | 1 | 0.9 | |
| Di | AC/GT | 1 | 2.2 |
| AG/CT | 16 | 34.8 | |
| AT/TA | 29 | 63.0 | |
| Tri | AAC/GTT | 9 | 13.4 |
| AAG/CTT | 20 | 30.0 | |
| AAT/ATT | 21 | 31.3 | |
| ACC/GGT | 1 | 1.4 | |
| ACG/CGT | 1 | 1.4 | |
| ACT/AGT | 2 | 3.0 | |
| AGC/CTG | 5 | 7.5 | |
| AGG/CCT | 4 | 6.0 | |
| ATC/ATG | 4 | 6.0 | |
| Tetra | AAAC/GTTT | 3 | 27.3 |
| AAAG/CTTT | 1 | 9.1 | |
| AAAT/ATTT | 5 | 45.4 | |
| AATC/ATTG | 1 | 9.1 | |
| AGAT/ATCT | 1 | 9.1 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Z.; Huang, Y.; An, W.; Zheng, X.; Huang, S.; Liang, L. Sequencing and Structural Analysis of the Complete Chloroplast Genome of the Medicinal Plant Lycium chinense Mill. Plants 2019, 8, 87. https://doi.org/10.3390/plants8040087
Yang Z, Huang Y, An W, Zheng X, Huang S, Liang L. Sequencing and Structural Analysis of the Complete Chloroplast Genome of the Medicinal Plant Lycium chinense Mill. Plants. 2019; 8(4):87. https://doi.org/10.3390/plants8040087
Chicago/Turabian StyleYang, Zerui, Yuying Huang, Wenli An, Xiasheng Zheng, Song Huang, and Lingling Liang. 2019. "Sequencing and Structural Analysis of the Complete Chloroplast Genome of the Medicinal Plant Lycium chinense Mill" Plants 8, no. 4: 87. https://doi.org/10.3390/plants8040087
APA StyleYang, Z., Huang, Y., An, W., Zheng, X., Huang, S., & Liang, L. (2019). Sequencing and Structural Analysis of the Complete Chloroplast Genome of the Medicinal Plant Lycium chinense Mill. Plants, 8(4), 87. https://doi.org/10.3390/plants8040087





