Differential Expression Proteins Contribute to Race-Specific Resistant Ability in Rice (Oryza sativa L.)
Abstract
1. Introduction
2. Results
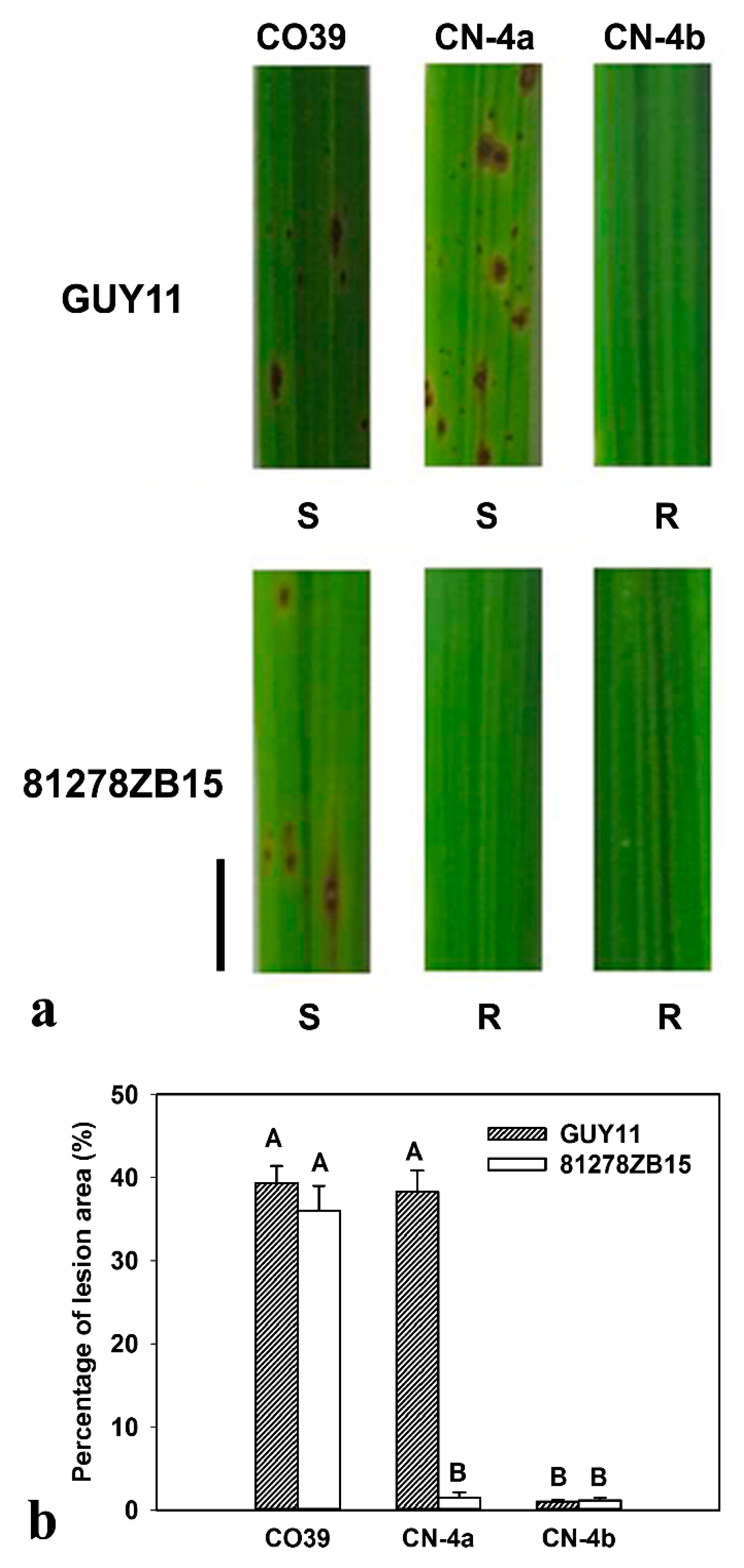
2.1. Phenotype of CO39 and its two NILs in Response to Infection of Different Fungal Isolates
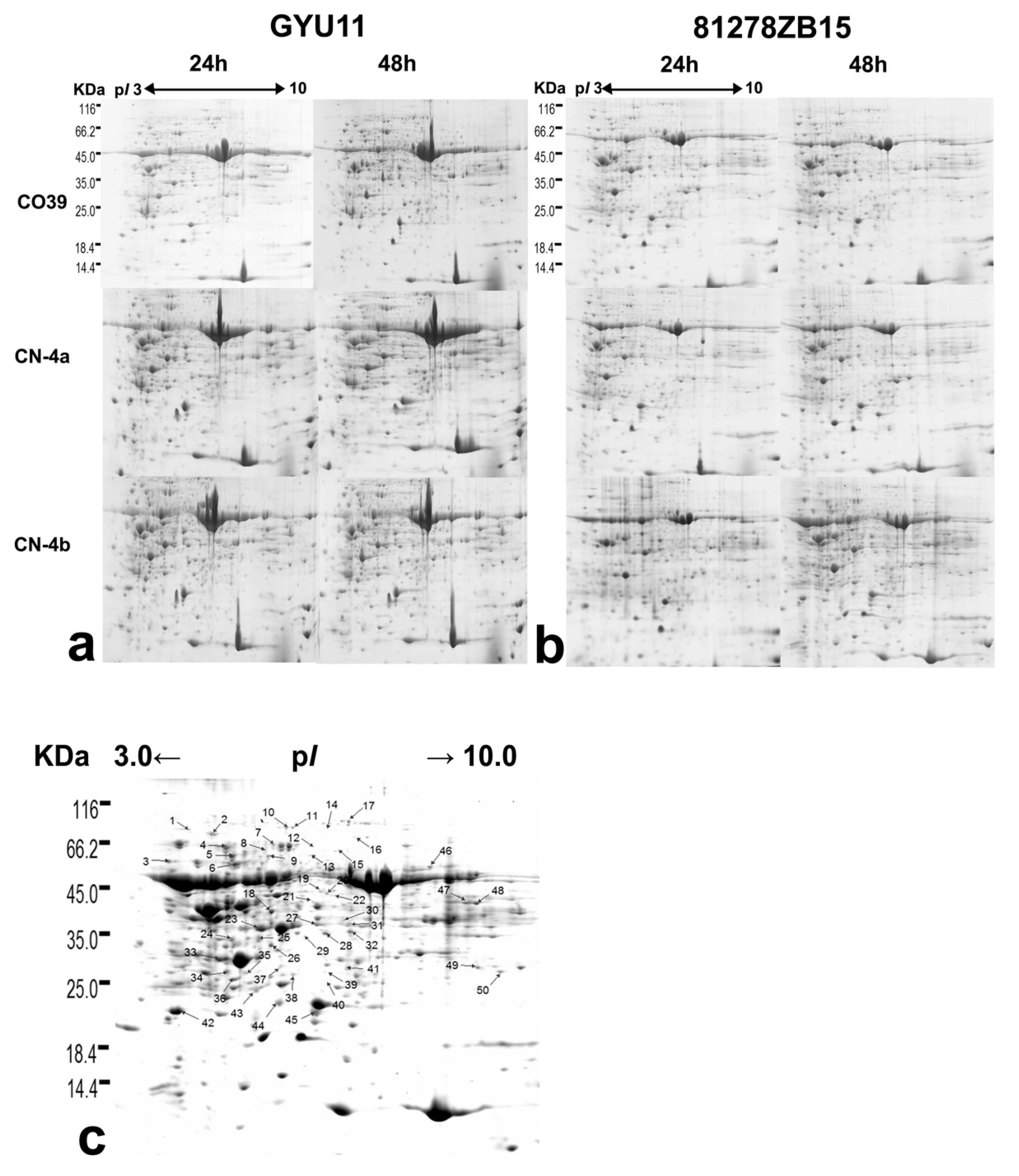
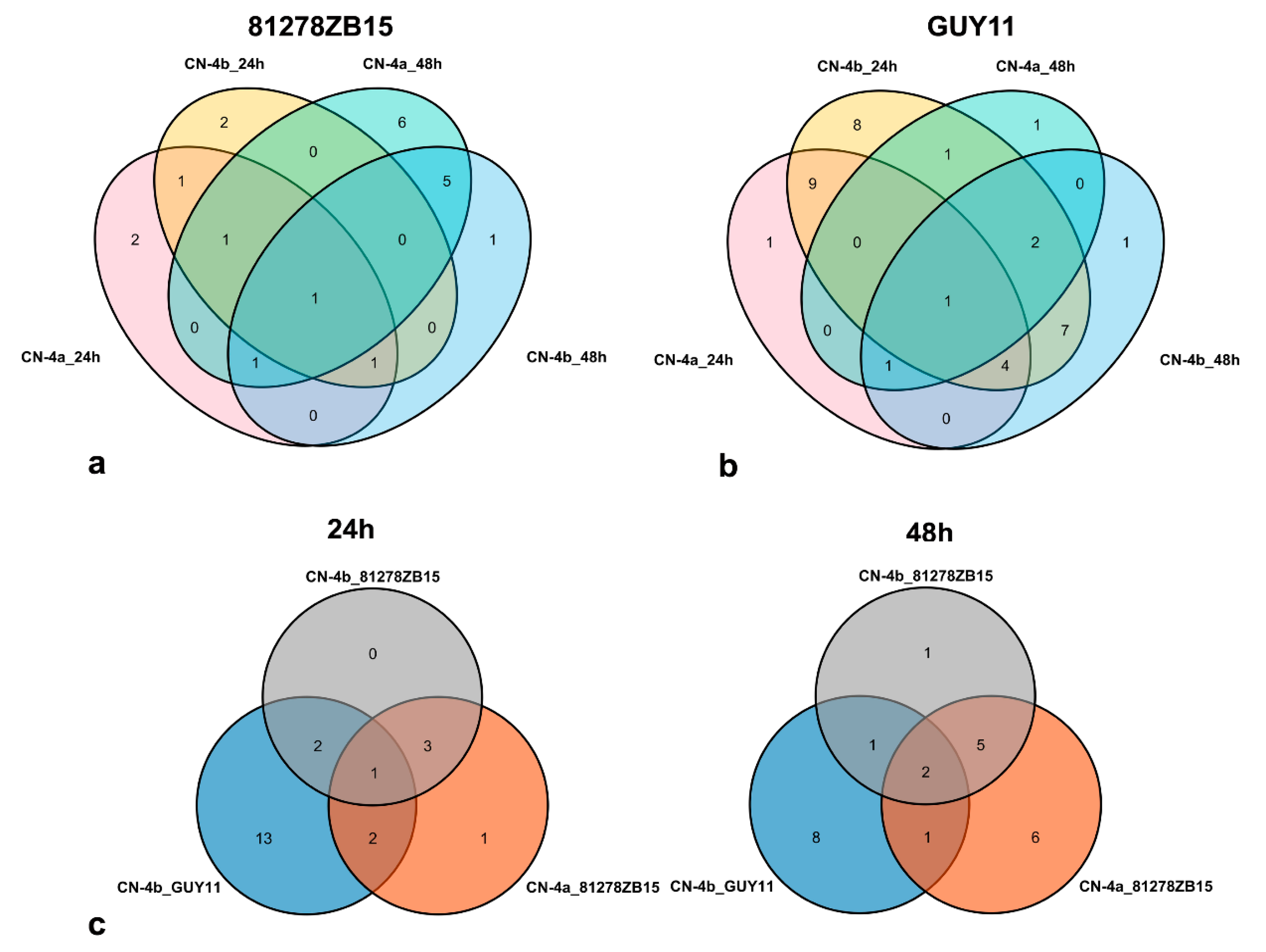
2.2. DEPs in Rice in Response to the Infection of Different Fungal Isolates
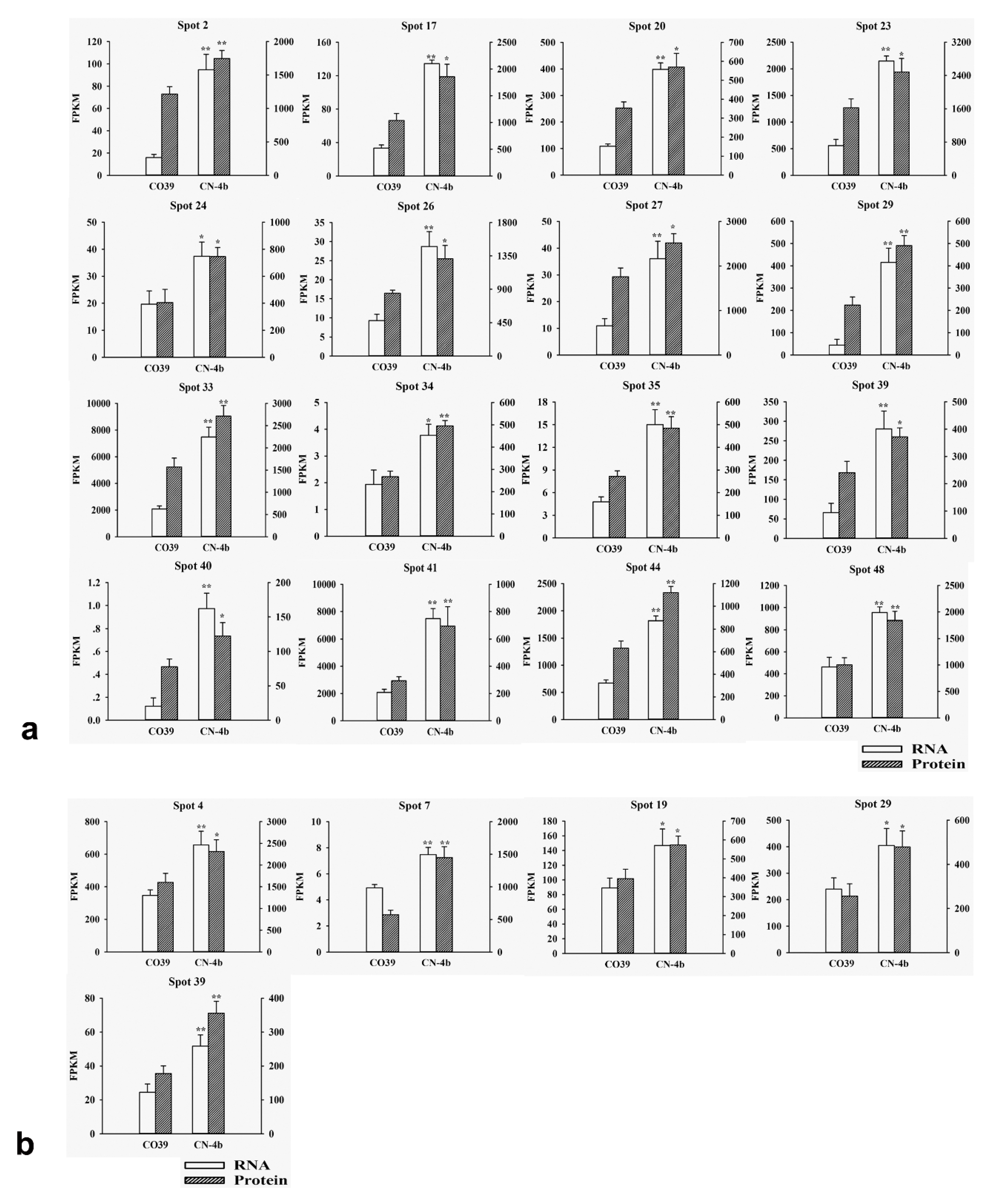
2.3. RNA Expression of DEPs in Rice in Response to Different Fungal Isolate Infection
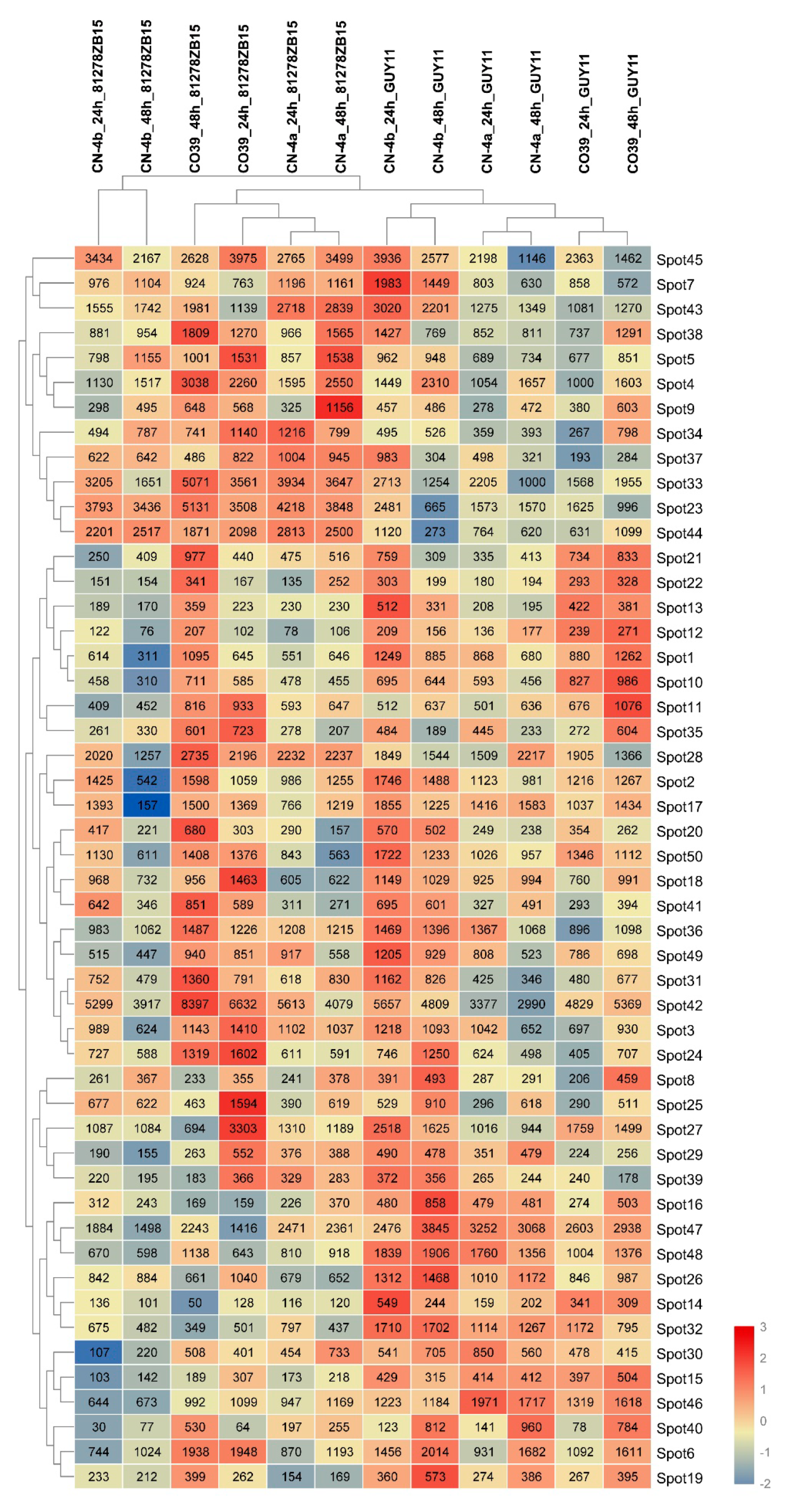
2.4. Clustering Analysis on the Expression Pattern of DEPs in Rice in Response to Different Fungal Isolate Infection
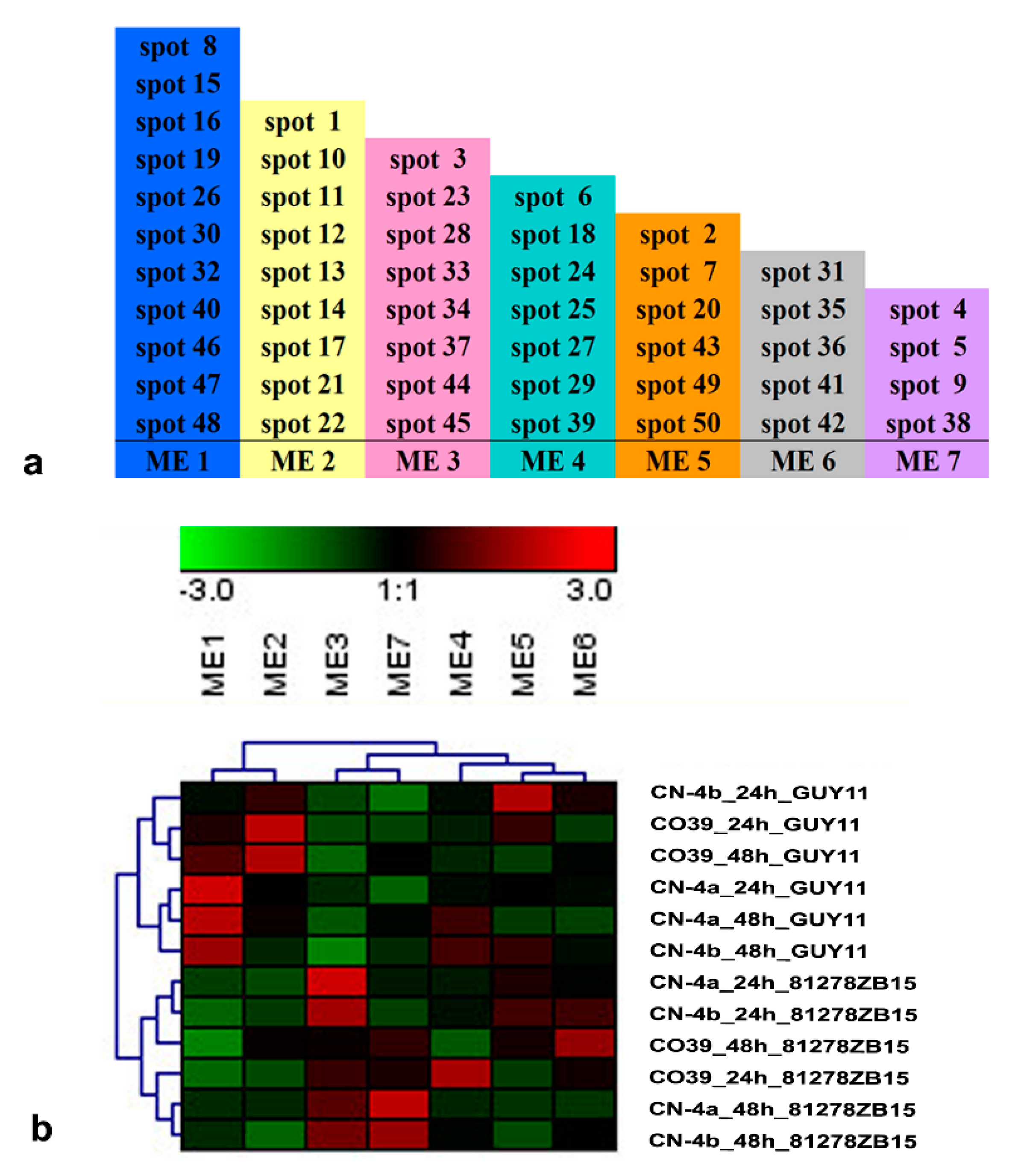
2.5. Functional Module of DEPs in Rice in Response to Different Fungal Isolate Infection
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Fungus Inoculation
4.2. Two-Dimensional Electrophoresis (2-DE) and Spots Selection
4.3. MALDI-TOF-TOF/MS Analysis and Protein Identification
4.4. RNA Extraction, Library Construction
4.5. Transcriptome Sequencing and de novo Assembly
4.6. Identification of Differentially Expressed Genes
4.7. Clustering Analysis of the Expression Pattern of DEPs
4.8. Co-Expression Network and Functional Module Analysis on DEPs
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Azizi, P.; Rafii, M.Y.; Abdullah, S.N.; Nejat, N.; Maziah, M.; Hanafi, M.M.; Latif, M.A.; Sahebi, M. Toward understanding of rice innate immunity against Magnaporthe oryzae. Crit. Rev. Biotechnol. 2016, 36, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Talbot, N.J. On the trail of a cereal killer: Exploring the biology of Magnaporthe grisea. Annu. Rev. Microbiol. 2003, 57, 177–202. [Google Scholar] [CrossRef]
- Dean, R.A.; Talbot, N.J.; Ebbole, D.J.; Farman, M.L.; Mitchell, T.K.; Orbach, M.J.; Thon, M.; Kulkarni, R.; Xu, J.R.; Pan, H.; et al. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 2005, 434, 980–986. [Google Scholar] [CrossRef] [PubMed]
- Hu, K.M.; Qiu, D.Y.; Shen, X.L.; Li, X.H.; Wang, S.P. Isolation and manipulation of quantitative trait loci for disease resistance in rice using a candidate gene approach. Mol. Plant 2008, 1, 786–793. [Google Scholar] [CrossRef] [PubMed]
- Miah, G.; Rafii, M.Y.; Ismail, M.R.; Puteh, A.B.; Rahim, H.A.; Asfaliza, R.; Latif, M.A. Blast resistance in rice: A review of conventional breeding to molecular approaches. Mol. Biol. Rep. 2013, 40, 2369–2388. [Google Scholar] [CrossRef] [PubMed]
- Hulbert, S.H.; Webb, C.A.; Smith, S.M.; Sun, Q. Resistance gene complexes: Evolution and utilization. Annu. Rev. Phytopathol. 2001, 39, 285–312. [Google Scholar] [CrossRef] [PubMed]
- Kou, Y.; Wang, S. Broad-spectrum and durability: Understanding of quantitative disease resistance. Curr. Opin. Plant Biol. 2010, 13, 181–185. [Google Scholar] [CrossRef]
- Kou, Y.; Wang, S. Toward an understanding of the molecular basis of quantitative disease resistance in rice. J. Biotechnol. 2012, 159, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.H.; Ebbole, D.J.; Wang, Z.H. The arms race between Magnaporthe oryzae and rice: Diversity and interaction of Avr and R genes. J. Integr. Agric. 2017, 16, 2746–2760. [Google Scholar] [CrossRef]
- Deng, Y.; Zhai, K.; Xie, Z.; Yang, D.; Zhu, X.; Liu, J.; Wang, X.; Qin, P.; Yang, Y.; Zhang, G.; et al. Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance. Science 2017, 355, 962–965. [Google Scholar] [CrossRef]
- Li, W.; Zhu, Z.; Chern, M.; Yin, J.; Yang, C.; Ran, L.; Cheng, M.; He, M.; Wang, K.; Wang, J.; et al. A Natural Allele of a Transcription Factor in Rice Confers Broad-Spectrum Blast Resistance. Cell 2017, 170, 114–126.e15. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhou, L.; Shi, H.; Chern, M.; Yu, H.; Yi, H.; He, M.; Yin, J.; Zhu, X.; Li, Y.; et al. A single transcription factor promotes both yield and immunity in rice. Science 2018, 361, 1026–1028. [Google Scholar] [CrossRef] [PubMed]
- Flor, H.H. Current Status of the Gene-For-Gene Concept. Annu. Rev. Phytopathol. 2003, 9, 275–296. [Google Scholar] [CrossRef]
- Orbach, M.J.; Farrall, L.; Sweigard, J.A.; Chumley, F.G.; Valent, B. A telomeric avirulence gene determines efficacy for the rice blast resistance gene Pi-ta. Plant Cell 2000, 12, 2019–2032. [Google Scholar] [CrossRef]
- Li, W.; Wang, B.; Wu, J.; Lu, G.; Hu, Y.; Zhang, X.; Zhang, Z.; Zhao, Q.; Feng, Q.; Zhang, H.; et al. The Magnaporthe oryzae avirulence gene AvrPiz-t encodes a predicted secreted protein that triggers the immunity in rice mediated by the blast resistance gene Piz-t. Mol. Plant Microbe Interact. 2009, 22, 411–420. [Google Scholar] [CrossRef]
- Yoshida, K.; Saitoh, H.; Fujisawa, S.; Kanzaki, H.; Matsumura, H.; Yoshida, K.; Tosa, Y.; Chuma, I.; Takano, Y.; Win, J.; et al. Association genetics reveals three novel avirulence genes from the rice blast fungal pathogen Magnaporthe oryzae. Plant Cell 2009, 21, 1573–1591. [Google Scholar] [CrossRef] [PubMed]
- Miki, S.; Matsui, K.; Kito, H.; Otsuka, K.; Ashizawa, T.; Yasuda, N.; Fukiya, S.; Sato, J.; Hirayae, K.; Fujita, Y.; et al. Molecular cloning and characterization of the AVR-Pia locus from a Japanese field isolate of Magnaporthe oryzae. Mol. Plant Pathol. 2009, 10, 361–374. [Google Scholar] [CrossRef] [PubMed]
- Farman, M.L.; Eto, Y.; Nakao, T.; Tosa, Y.; Nakayashiki, H.; Mayama, S.; Leong, S.A. Analysis of the structure of the AVR1-CO39 avirulence locus in virulent rice-infecting isolates of Magnaporthe grisea. Mol. Plant Microbe Interact. 2002, 15, 6–16. [Google Scholar] [CrossRef] [PubMed]
- Ray, S.; Singh, P.K.; Gupta, D.K.; Mahato, A.K.; Sarkar, C.; Rathour, R.; Singh, N.K.; Sharma, T.R. Analysis of Magnaporthe oryzae Genome Reveals a Fungal Effector, Which Is Able to Induce Resistance Response in Transgenic Rice Line Containing Resistance Gene, Pi54. Front. Plant Sci. 2016, 7, 1140. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhao, J.; Xiang, Y.; Bian, X.; Zuo, Q.; Shen, Q.; Gai, J.; Xing, H. Proteomics study of changes in soybean lines resistant and sensitive to Phytophthora sojae. Proteome Sci. 2011, 9, 52. [Google Scholar] [CrossRef]
- Li, D.X.; Wang, L.J.; Teng, S.L.; Zhang, G.G.; Guo, L.J.; Mao, Q.; Wang, W.; Li, M.; Chen, L. Proteomics analysis of rice proteins up-regulated in response to bacterial leaf streak disease. J. Plant Biol. 2012, 55, 316–324. [Google Scholar] [CrossRef]
- Ventelon-Debout, M.; Delalande, F.; Brizard, J.P.; Diemer, H.; Van Dorsselaer, A.; Brugidou, C. Proteome analysis of cultivar-specific deregulations of Oryza sativa indica and O. sativa japonica cellular suspensions undergoing rice yellow mottle virus infection. Proteomics 2004, 4, 216–225. [Google Scholar] [CrossRef] [PubMed]
- Mackill, D.; Bonman, J. Inheritance of blast resistance in near-isogenic lines of rice. Phytopathology 1992, 82, 746–749. [Google Scholar] [CrossRef]
- Wei, X.Y.; Xu-Ming, X.U.; Zhang, R.; Chen, M.L.; Bin-Lin, M.A.; Zou, W.G.; Yang, W.X.; Zhuo, W.; Wang, Z.H.; Liang, K.J. Inheritance of Blast Resistance in New Germplasm Kangfeng A from Indica-Japonica Crosses. J. Plant Genet. Resour. 2014, 15, 1133–1137. [Google Scholar]
- Kim, S.T.; Kim, S.G.; Hwang, D.H.; Kang, S.Y.; Kim, H.J.; Lee, B.H.; Lee, J.J.; Kang, K.Y. Proteomic analysis of pathogen-responsive proteins from rice leaves induced by rice blast fungus, Magnaporthe grisea. Proteomics 2004, 4, 3569–3578. [Google Scholar] [CrossRef] [PubMed]
- Xue, W.M.; Mao, X.H.; Wei, Y.D.; Ling, L.; Zhu, Y.S.; He, S.L.; Zhang, J.F.; Xie, H.A. Proteomic analysis of blast-resistant near-isogenic lines derived from japonica rice, var. Yunyin, infected with Magnaporthe oryzae. Chin. Sci. Bull. 2014, 59, 4312–4322. [Google Scholar] [CrossRef]
- Tian, D.; Yang, L.; Chen, Z.; Chen, Z.; Wang, F.; Zhou, Y.; Luo, Y.; Yang, L.; Chen, S. Proteomic analysis of the defense response to Magnaporthe oryzae in rice harboring the blast resistance gene Piz-t. Rice (N. Y.) 2018, 11, 47. [Google Scholar] [CrossRef]
- Sasaki, K.; Iwai, T.; Hiraga, S.; Kuroda, K.; Seo, S.; Mitsuhara, I.; Miyasaka, A.; Iwano, M.; Ito, H.; Matsui, H.; et al. Ten rice peroxidases redundantly respond to multiple stresses including infection with rice blast fungus. Plant Cell Physiol. 2004, 45, 1442–1452. [Google Scholar] [CrossRef]
- Nishizawa, Y.; Nishio, Z.; Nakazono, K.; Soma, M.; Nakajima, E.; Ugaki, M.; Hibi, T. Enhanced resistance to blast (Magnaporthe grisea) in transgenic Japonica rice by constitutive expression of rice chitinase. Theor. Appl. Genet. 1999, 99, 383–390. [Google Scholar] [CrossRef]
- Nakashima, A.; Chen, L.; Thao, N.P.; Fujiwara, M.; Wong, H.L.; Kuwano, M.; Umemura, K.; Shirasu, K.; Kawasaki, T.; Shimamoto, K. RACK1 functions in rice innate immunity by interacting with the Rac1 immune complex. Plant Cell 2008, 20, 2265–2279. [Google Scholar] [CrossRef] [PubMed]
- Bagnaresi, P.; Biselli, C.; Orrù, L.; Urso, S.; Crispino, L.; Abbruscato, P.; Piffanelli, P.; Lupotto, E.; Cattivelli, L.; Valè, G. Comparative Transcriptome Profiling of the Early Response to Magnaporthe oryzae in Durable Resistant vs Susceptible Rice (Oryza sativa L.) Genotypes. PLoS ONE 2012, 7, e51609. [Google Scholar] [CrossRef] [PubMed]
- Ravet, K.; Touraine, B.; Boucherez, J.; Briat, J.F.; Gaymard, F.; Cellier, F. Ferritins control interaction between iron homeostasis and oxidative stress in Arabidopsis. Plant J. 2009, 57, 400–412. [Google Scholar] [CrossRef] [PubMed]
- Salekdeh, G.H.; Siopongco, J.; Wade, L.J.; Ghareyazie, B.; Bennett, J. Proteomic analysis of rice leaves during drought stress and recovery. Proteomics 2002, 2, 1131–1145. [Google Scholar] [CrossRef]
- Lowry, O.H.; Rosebrough, N.J.; Farr, A.L.; Randall, R.J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 1951, 193, 265–275. [Google Scholar] [PubMed]
- Bolwell, G.P.; Bindschedler, L.V.; Blee, K.A.; Butt, V.S.; Davies, D.R.; Gardner, S.L.; Gerrish, C.; Minibayeva, F. The apoplastic oxidative burst in response to biotic stress in plants: A three-component system. J. Exp. Bot. 2002, 53, 1367–1376. [Google Scholar] [PubMed]
- Yoda, H.; Yamaguchi, Y.; Sano, H. Induction of hypersensitive cell death by hydrogen peroxide produced through polyamine degradation in tobacco plants. Plant Physiol. 2003, 132, 1973–1981. [Google Scholar] [CrossRef] [PubMed]
- Valent, B.; Farrall, L.; Chumley, F.G. Magnaporthe grisea genes for pathogenicity and virulence identified through a series of backcrosses. Genetics 1991, 127, 87–101. [Google Scholar]
- He, H.; Li, J. Proteomic analysis of phosphoproteins regulated by abscisic acid in rice leaves. Biochem. Biophys. Res. Commun. 2008, 371, 883–888. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Chen, X.; Lin, S.; Liu, Q.; Huang, J.; Zhang, W.; Lin, J.; Wang, Y.; Ke, Y.; He, H. Expression and interaction of small heat shock proteins (sHsps) in rice in response to heat stress. Biochim. Biophys. Acta 2014, 1844, 818–828. [Google Scholar] [CrossRef]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Xia, R.; Chen, H.; He, Y. TBtools, a Toolkit for Biologists integrating various HTS-data handling tools with a user-friendly interface. bioRxiv 2018. [Google Scholar] [CrossRef]
- Wiech, H.; Buchner, J.; Zimmermann, R.; Jakob, U. Hsp90 chaperones protein folding in vitro. Nature 1992, 358, 169–170. [Google Scholar] [CrossRef] [PubMed]
| No a | Gi b | Protein Description | MW c (KDa) | pI d | Peptides matched | Cov (%) e |
|---|---|---|---|---|---|---|
| 1 | gi|115479725 | Heat shock 90 kDa protein | 75.0 | 8.94 | 10 | 18 |
| 2 | gi|115459660 | Elongation factor G (EF-G) family | 42.1 | 5.47 | 12 | 41 |
| 3 | gi|573913556 | Histone-lysine N-methyltransferase ATX4-like | 105.0 | 7.65 | 3 | 5 |
| 4 | gi|115486793 | Heat shock 70 kDa protein | 71.5 | 5.10 | 7 | 16 |
| 5 | gi|115469362 | V-type proton ATPase catalytic subunit A | 68.7 | 5.20 | 16 | 39 |
| 6 | gi|115466004 | Similar to 60 kDa chaperonin (Protein Cpn60) | 64.3 | 5.60 | 10 | 25 |
| 7 | gi|115450022 | Zn-dependent oligopeptidases | 86.5 | 5.76 | 9 | 14 |
| 8 | gi|115448989 | Molecular chaperone DnaK | 73.1 | 5.49 | 13 | 26 |
| 9 | gi|115440691 | 2,3-bisphosphoglycerate-independent phosphoglycerate mutase | 61.0 | 5.42 | 6 | 14 |
| 10 | gi|347602486 | Chaperone protein ClpC1, chloroplastic | 101.9 | 6.14 | 33 | 41 |
| 11 | gi|115487910 | Chaperone protein ClpC2, chloroplastic | 102.1 | 6.62 | 9 | 12 |
| 12 | gi|115454943 | NADH dehydrogenase subunit G | 82.1 | 5.86 | 15 | 33 |
| 13 | gi|115458140 | Similar to Glycyl-tRNA synthetase | 32.8 | 5.47 | 5 | 22 |
| 14 | gi|75225211 | Putative aconitate hydratase | 98.6 | 5.67 | 18 | 22 |
| 15 | gi|115439655 | NADP-dependent malic enzyme | 65.6 | 8.59 | 9 | 23 |
| 16 | gi|115489652 | 5-methyltetrahydropteroyltriglutamate--homocysteine methyltransferase | 84.9 | 5.93 | 12 | 24 |
| 17 | gi|115468926 | glycine dehydrogenase | 97.9 | 5.98 | 9 | 10 |
| 18 | gi|115448531 | Glutamine synthetase | 39.4 | 5.51 | 5 | 17 |
| 19 | gi|3024122 | S-adenosylmethionine synthase 2 | 43.3 | 5.68 | 10 | 31 |
| 20 | gi|115450493 | Glyceraldehyde-3-phosphate dehydrogenase B | 47.5 | 6.22 | 8 | 23 |
| 21 | gi|115482032 | GDP-mannose-3,5-epimerase (GME)-like | 43.1 | 5.75 | 7 | 25 |
| 22 | gi|115461951 | S-denosylmethionine synthetase 1 | 43.6 | 5.74 | 14 | 37 |
| 23 | gi|115484401 | Fructose-bisphosphate aldolase | 42.2 | 6.38 | 8 | 35 |
| 24 | gi|115445243 | Similar to Class III peroxidase | 34.8 | 5.32 | 4 | 23 |
| 25 | gi|115482534 | Cytoplasmic malate dehydrogenase | 35.9 | 5.75 | 10 | 36 |
| 26 | gi|115477837 | Copper/zinc superoxide dismutase (SOD) | 21.4 | 5.79 | 5 | 41 |
| 27 | gi|937924719 | Similar to Photosystem II stability/assembly factor HCF136, chloroplastic | 42.9 | 9.05 | 8 | 27 |
| 28 | gi|125537696 | ferredoxin-NADP(+) reductase | 41.1 | 8.26 | 4 | 10 |
| 29 | gi|115435022 | Similar to Acid phosphatase | 33.2 | 6.33 | 10 | 30 |
| 30 | gi|115450565 | Similar to Glutathione S-transferase GSTF14 | 35.7 | 7.04 | 6 | 27 |
| 31 | gi|115439261 | Similar to Guanine nucleotide-binding protein subunit beta-like protein | 36.7 | 5.97 | 8 | 34 |
| 32 | gi|115443911 | Similar to NADPH-dependent mannose 6-phosphate reductase | 42.1 | 8.16 | 6 | 16 |
| 33 | gi|115436780 | Similar to Photosystem II oxygen-evolving complex protein 1 | 34.8 | 4.96 | 3 | 21 |
| 34 | gi|115488968 | Nucleoside diphosphate kinase | 23.6 | 9.51 | 3 | 11 |
| 35 | gi|115453797 | Haloacid dehalogenase-like hydrolases | 34.1 | 8.36 | 6 | 25 |
| 36 | gi|115474285 | L-ascorbate peroxidase | 27.2 | 5.21 | 7 | 45 |
| 37 | gi|218189522 | Phage shock protein A (IM30) | 35.1 | 9.08 | 5 | 16 |
| 38 | gi|115447465 | Similar to ATP-dependent Caseinolytic protease (ClpP) proteolytic subunit | 32.1 | 6.71 | 7 | 22 |
| 39 | gi|297604125 | Similar to Chitinase | 32.8 | 6.08 | 10 | 26 |
| 40 | gi|115461679 | NAD-dependent epimerase/dehydratase family protein | 31.4 | 9.13 | 6 | 31 |
| 41 | gi|115463555 | Beta-glucanase | 34.7 | 5.92 | 5 | 23 |
| 42 | gi|115446541 | Similar to 2-cys peroxiredoxin BAS1 | 28.3 | 5.67 | 3 | 42 |
| 43 | gi|115486898 | Eukaryotic ferritins | 28.3 | 5.47 | 6 | 29 |
| 44 | gi|115470529 | photosystem II oxygen-evolving enhancer protein 2 | 27.1 | 8.66 | 5 | 28 |
| 45 | gi|115476760 | Cupin_1; Similar to Germin-like protein 1 | 22.0 | 6.01 | 1 | 10 |
| 46 | gi|115436320 | Dihydrolipoamide dehydrogenase family protein | 53.0 | 7.21 | 5 | 18 |
| 47 | gi|115460656 | Similar to Aminomethyltransferase | 44.3 | 8.53 | 7 | 19 |
| 48 | gi|115477148 | Serine-glyoxylate aminotransaminase | 44.4 | 8.19 | 4 | 18 |
| 49 | gi|115478779 | Outer mitochondrial membrane protein porin (Voltage-dependent anion-selective channel protein) (VDAC) | 29.2 | 7.07 | 6 | 28 |
| 50 | gi|115460338 | Heme-dependent peroxidases | 38.4 | 8.67 | 14 | 42 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, S.; Lin, S.; Wang, M.; Zou, Y.; Tao, H.; Liu, W.; Zhang, L.; Liang, K.; Ai, Y.; He, H. Differential Expression Proteins Contribute to Race-Specific Resistant Ability in Rice (Oryza sativa L.). Plants 2019, 8, 29. https://doi.org/10.3390/plants8020029
Ma S, Lin S, Wang M, Zou Y, Tao H, Liu W, Zhang L, Liang K, Ai Y, He H. Differential Expression Proteins Contribute to Race-Specific Resistant Ability in Rice (Oryza sativa L.). Plants. 2019; 8(2):29. https://doi.org/10.3390/plants8020029
Chicago/Turabian StyleMa, Shiwei, Shoukai Lin, Menglin Wang, Yang Zou, Huan Tao, Wei Liu, Lina Zhang, Kangjing Liang, Yufang Ai, and Huaqin He. 2019. "Differential Expression Proteins Contribute to Race-Specific Resistant Ability in Rice (Oryza sativa L.)" Plants 8, no. 2: 29. https://doi.org/10.3390/plants8020029
APA StyleMa, S., Lin, S., Wang, M., Zou, Y., Tao, H., Liu, W., Zhang, L., Liang, K., Ai, Y., & He, H. (2019). Differential Expression Proteins Contribute to Race-Specific Resistant Ability in Rice (Oryza sativa L.). Plants, 8(2), 29. https://doi.org/10.3390/plants8020029






