Allelic Differentiation at the E1/Ghd7 Locus Has Allowed Expansion of Rice Cultivation Area
Abstract
1. Introduction
2. Results
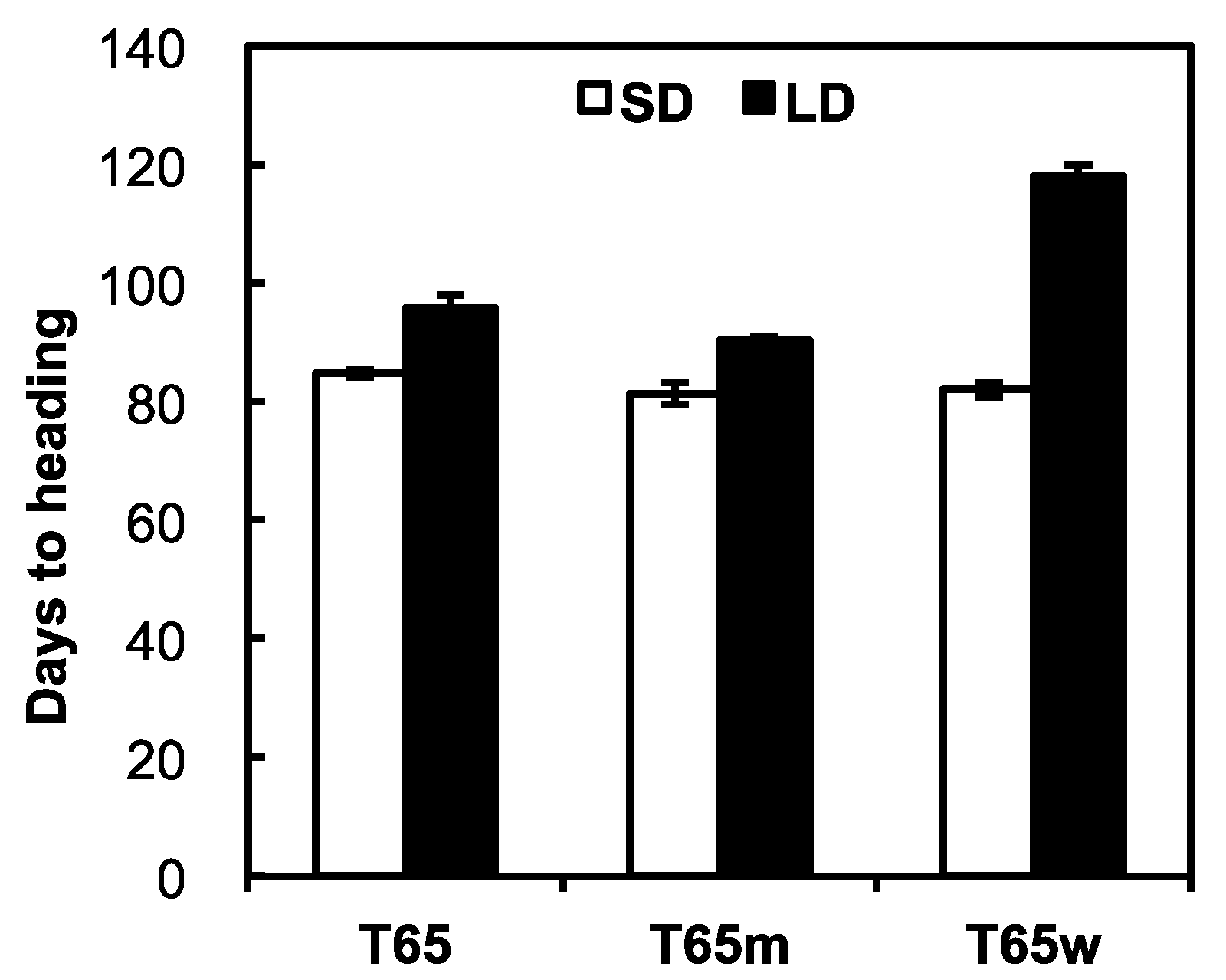
2.1. Photoperiod Sensitivities of T65, T65w, and T65m
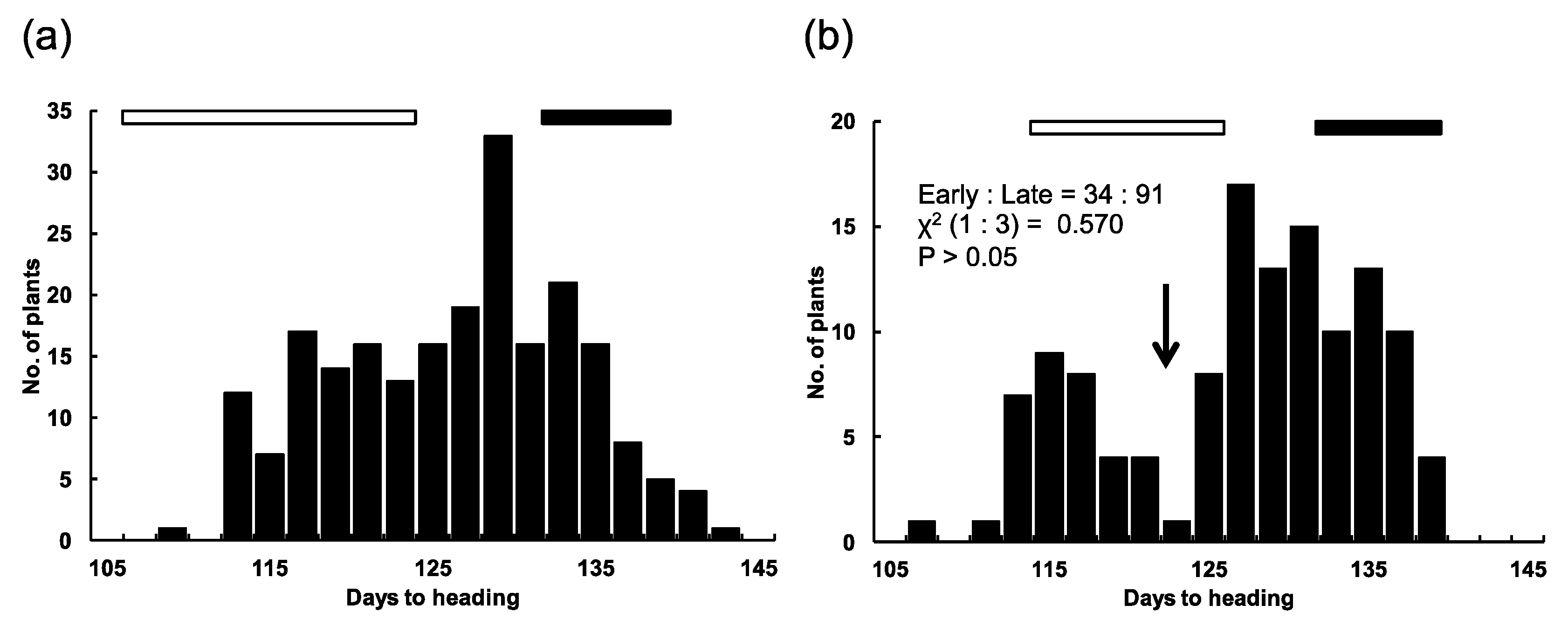
2.2. Chromosomal Location of the E1 Locus
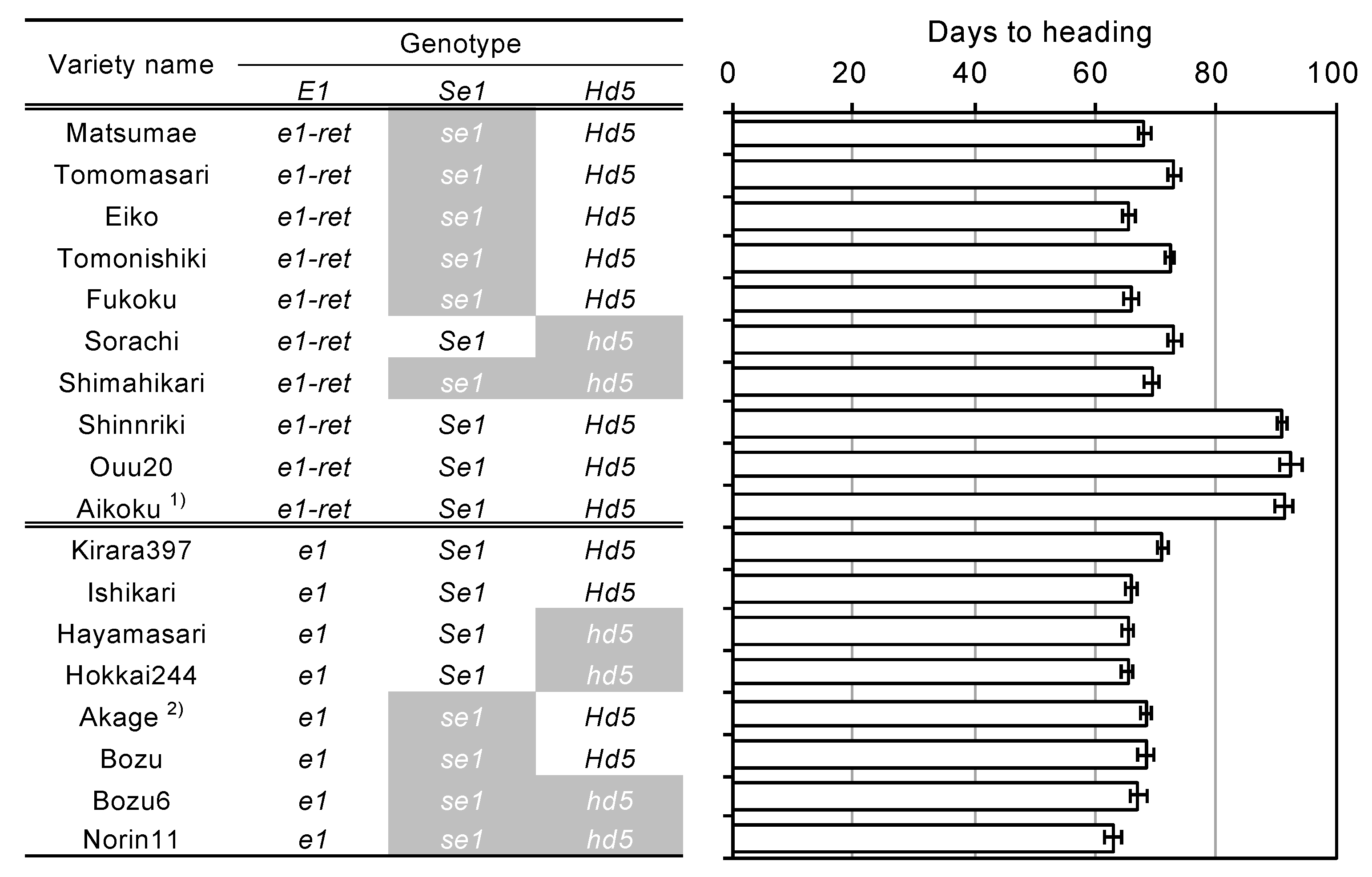
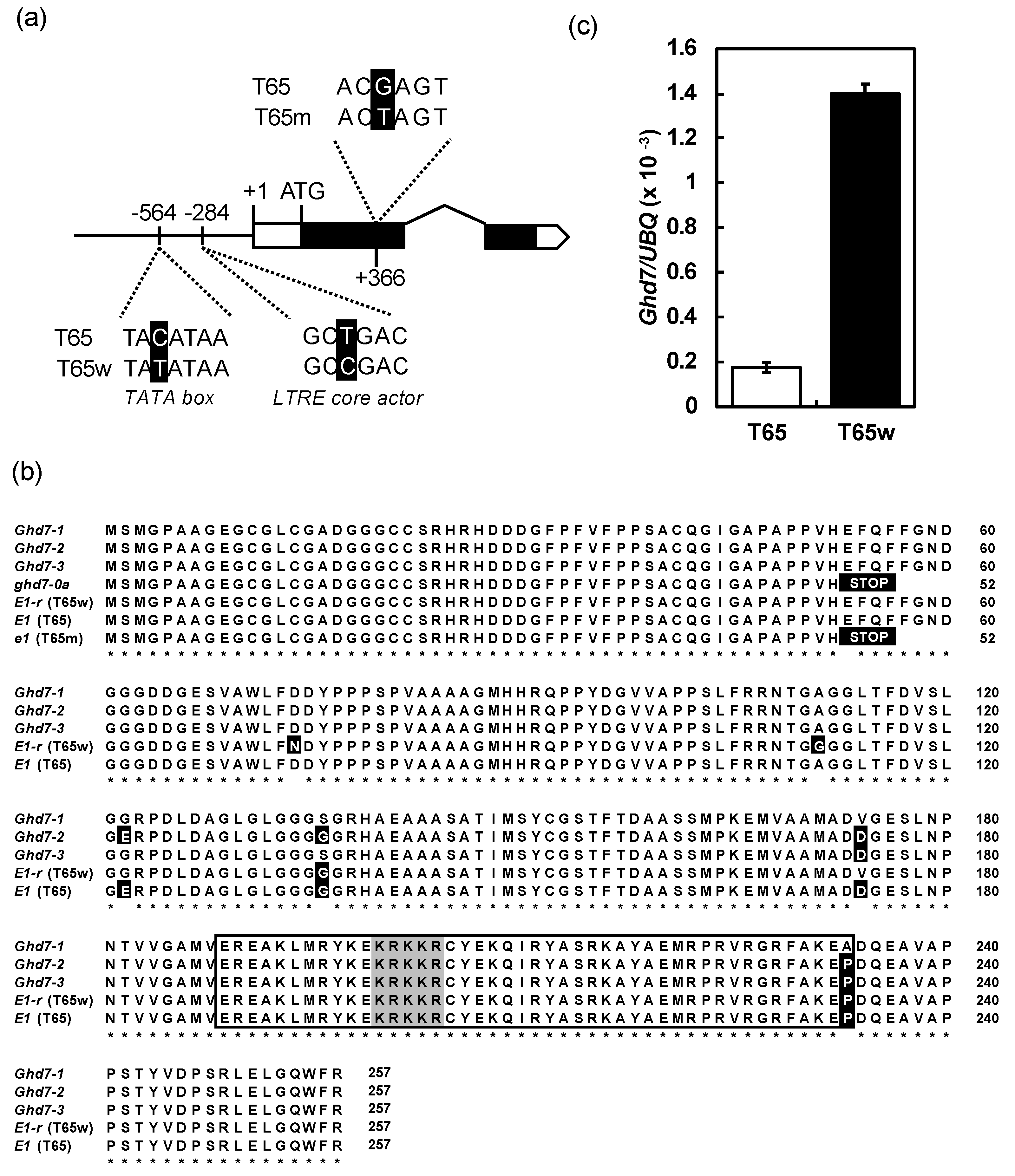
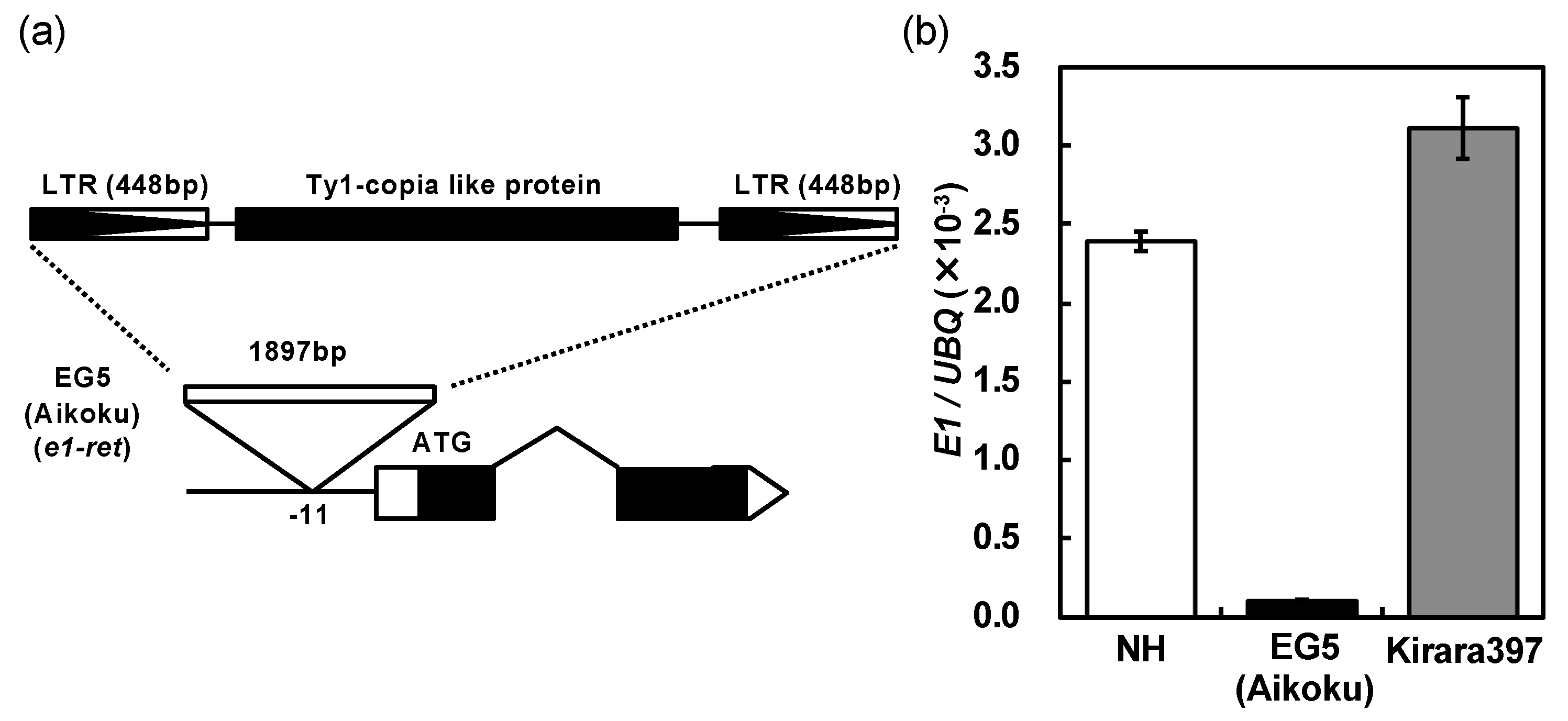
2.3. A Novel Nonfunctional Allele at the E1/Ghd7 Locus
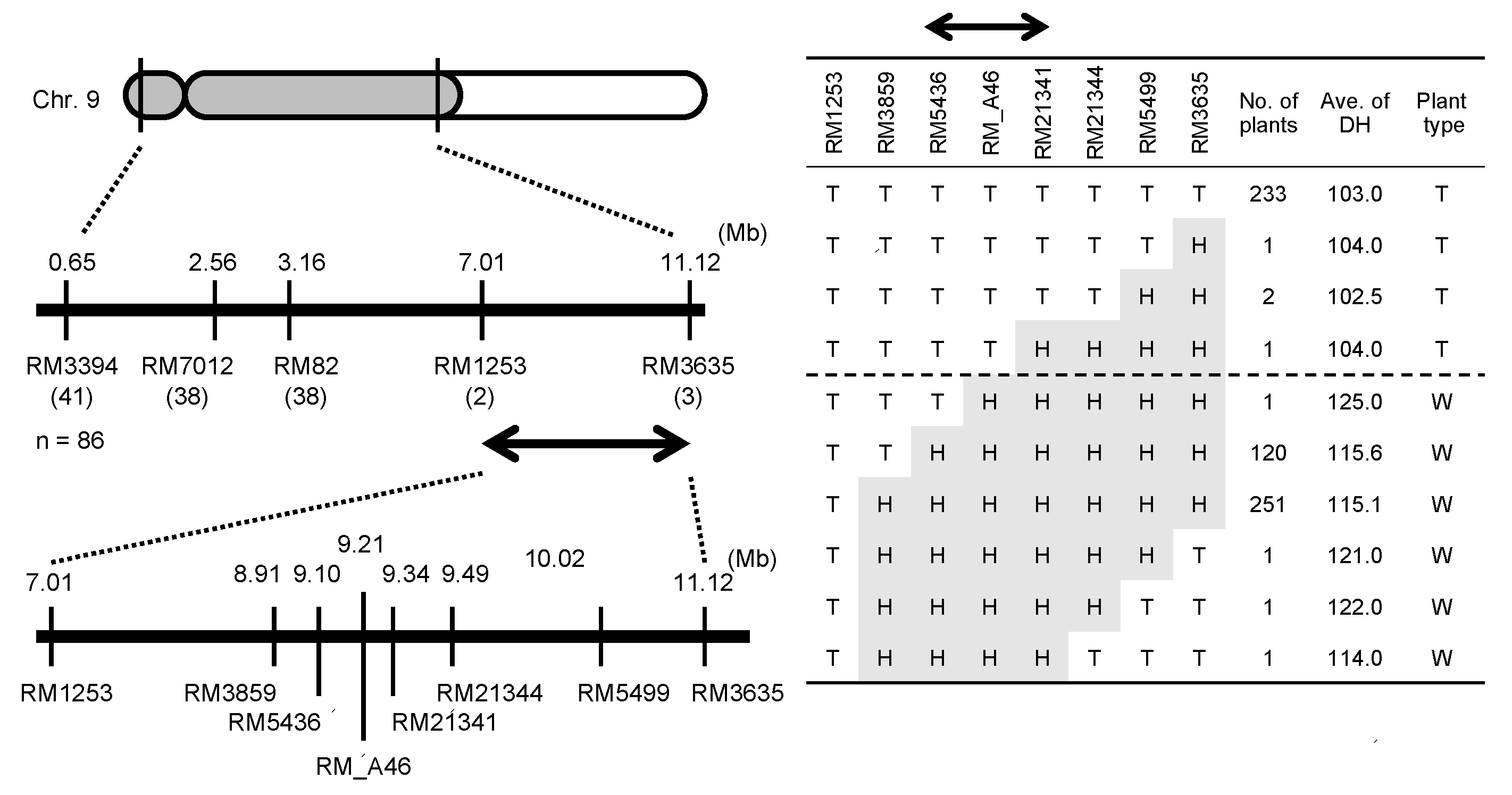
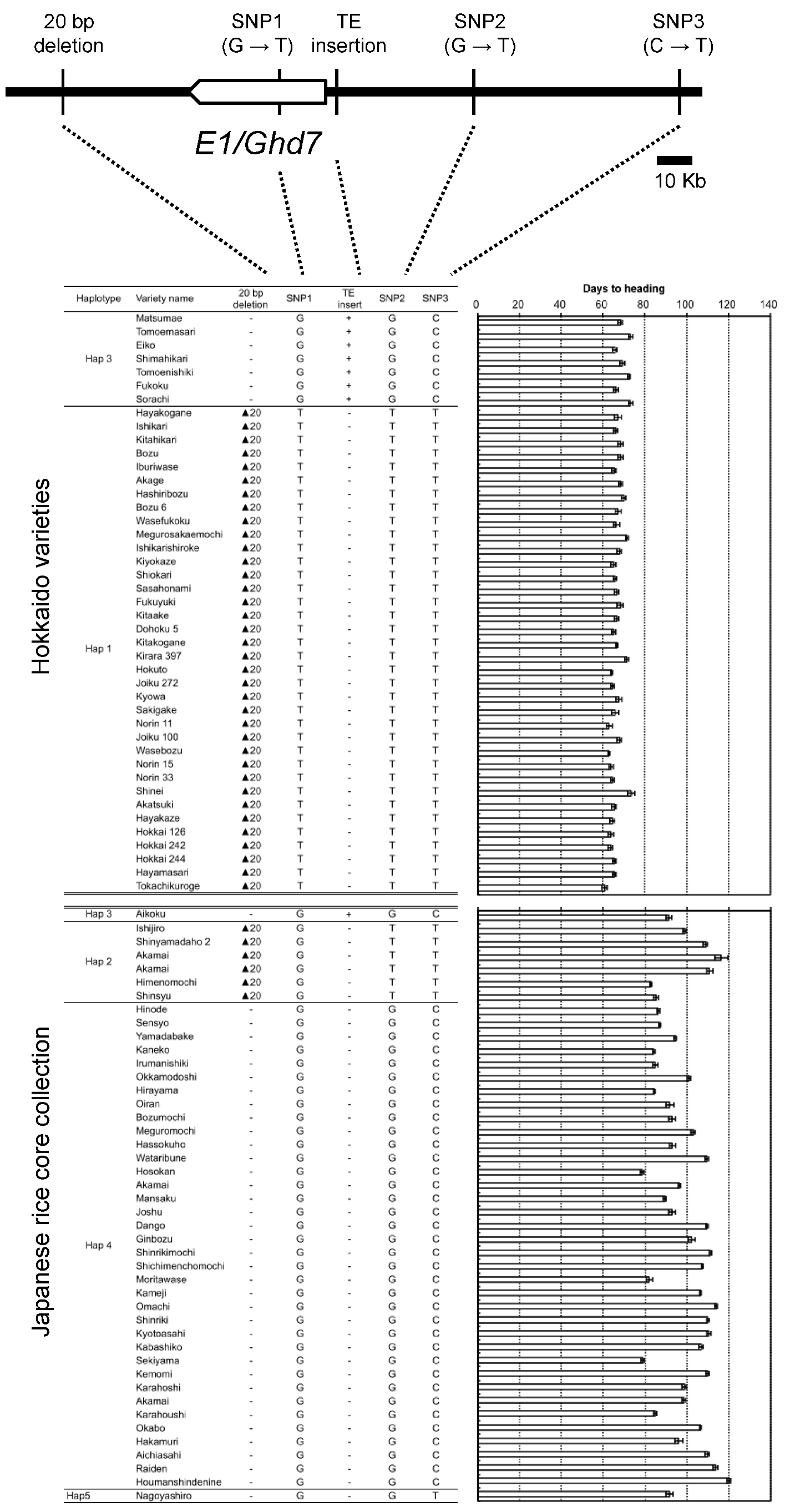
2.4. Haplotype Patterns of the Chromosomal Region Surrounding E1/Ghd7 Locus
3. Discussion
4. Materials and Methods
4.1. Photoperiod Sensitivities of T65, T65m, and T65
4.2. Chromosomal Location of the E1 Locus
4.3. Expression Analysis of E1/Ghd7
4.4. Identification of the Genotype at the E1/Ghd7 Locus
4.5. Haplotype Patterns of the Chromosomal Region around the E1/Ghd7 Locus
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Khush, G.S. Green revolution: The way forward. Nat. Rev. Genet. 2001, 2, 815–820. [Google Scholar] [CrossRef] [PubMed]
- Wada, E. Studies on the response of heading to daylength and temperature in rice plants. : II. Response of varieties and the relation to their geographical distribution in Japan. Jpn. J. Breed. 1952, 2, 55–62. [Google Scholar] [CrossRef]
- Hosoi, T. Studies on meteorological fluctuation in the growth of rice plants. V. Regional references of thermo-sensitivity, photo-sensitivity, basic vegetative growth and factors determining the growth duration of Japanese varieties. Jpn. J. Breed. 1981, 31, 239–250. [Google Scholar] [CrossRef]
- Tanisaka, T.; Inoue, H.; Uozu, S.; Yamagata, H. Basic vegetative growth and photoperiod sensitivity of heading-time mutants induced in rice. Jpn. J. Breed. 1992, 42, 657–668. [Google Scholar] [CrossRef]
- Okumoto, Y.; Ichitani, K.; Inoue, H.; Tanisaka, T. Photoperiod insensitivity gene essential to the varieties grown in the northern limit region of paddy rice (Oryza sativa L.) cultivation. Euphytica. 1996, 92, 63–66. [Google Scholar] [CrossRef]
- Zhang, Y. An Economic Analysis on Growth and Technological Progress of Rice Production in Three Northeastern Provinces of China. J. Rural Problem 2002, 38, 1–12. [Google Scholar] [CrossRef][Green Version]
- Saito, H.; Yuan, Q.; Okumoto, Y.; Doi, K.; Yoshimura, A.; Inoue, H.; Teraishi, M.; Tsukiyama, T.; Tanisaka, T. Multiple alleles at Early flowering 1 locus making variation in the basic vegetative growth period in rice (Oryza sativa L.). Theor. Appl. Genet. 2009, 119, 315–323. [Google Scholar] [CrossRef]
- Saito, H.; Ogiso-Tanaka, E.; Okumoto, Y.; Yoshitake, Y.; Izumi, H.; Yokoo, T.; Matsubara, K.; Hori, K.; Yano, M.; Inoue, H.; et al. Ef7 encodes an ELF3-like protein and promotes rice flowering by negatively regulating the floral repressor gene Ghd7 under both short-and long-day conditions. Plant Cell Physiol. 2012, 53, 717–728. [Google Scholar] [CrossRef]
- Yuan, Q.; Saito, H.; Okumoto, Y.; Inoue, H.; Nishida, H.; Tsukiyama, T.; Teraishi, M.; Tanisaka, T. Identification of a novel gene ef7 conferring an extremely long basic vegetative growth phase in rice. Theor. Appl. Genet. 2009, 119, 675–684. [Google Scholar] [CrossRef]
- Hoshino, Y. On the inheritance of the flowering time in peas and rice. J. Col. Agric. 1915, 6, 229–288. [Google Scholar]
- Syakudo, K.; Kawase, T. Studies on the quantitative inheritance (11): A Rice (Oryza sativa L.) (d) Inheritance of the heading duration and the quantitative function of the causal genes in its determination. : (1) On the quantitative function of the genes E1, E2 and D1. Jpn. J. Breed. 1953, 3, 6–12. [Google Scholar] [CrossRef]
- Syakudo, K.; Kawase, T.; Yoshino, K. Studies on the quantitative inheritance (13): A Rice (Oryza sativa L.) (d) Inheritance of the heading duration and the quantitative function of the causal genes in its determination. : (2) On the quantitative function of the genes E3, E4 and E5. Jpn. J. Breed. 1954, 4, 83–91. [Google Scholar] [CrossRef]
- Yamagata, H.; Okumoto, Y.; Tanisaka, T. Analysis of genes controlling heading time in Japanese rice. Rice Genet. 1986, 1, 351–359. [Google Scholar]
- Okumoto, Y.; Tanisaka, T.; Yamagata, H. Genotypic difference in response to light interruption in Japanese rice varieties. Rice Genet. 1991, 2, 778–780. [Google Scholar]
- Yokoo, M.; Kikuchi, F.; Nakane, A.; Fujimaki, H. Genetical analysis of heading time by aid of close linkage with blast resistance in rice. Bull. Nat. Inst. Agric. Sci. Ser. 1980, D31, 95–126. [Google Scholar]
- Tsai, K.H. Studies on earliness genes in rice, with special reference to analysis of isoalleles at the E locus. Jpn. J. Genet. 1976, 51, 115–128. [Google Scholar] [CrossRef]
- Okumoto, Y.; Tanisaka, T.; Yamagata, H. Heading-time genes of the rice varieties grown in south-west-warm region in Japan. Jpn. J. Breed. 1991, 41, 135–152. [Google Scholar] [CrossRef]
- Okumoto, T.; Tanisaka, T.; Yamagata, H. Heading-time genes of the rice varieties grown in the Tohoku-Hokuriku region in Japan. Jpn. J. Breed. 1992, 42, 121–135. [Google Scholar] [CrossRef]
- Okumoto, Y.; Yoshimura, A.; Tanisaka, T.; Yamagata, H. Analysis of a rice variety Taichung 65 and its isogenic early-heading lines for late-heading genes E1, E2 and E3. Jpn. J. Breed. 1992, 42, 415–429. [Google Scholar] [CrossRef]
- Ichitani, K.; Okumoto, Y.; Tanisaka, T. Photoperiod sensitivity gene of Se-1 locus found in photoperiod insensitive rice cultivars of the northern limit region of rice cultivation. Breed. Sci. 1997, 47, 145–152. [Google Scholar]
- Ichitani, K.; Okumoto, Y.; Tanisaka, T. Genetic analysis of low photoperiod sensitivity of rice cultivars for the northernmost regions of Japan. Plant Breed. 1998, 117, 543–547. [Google Scholar] [CrossRef]
- Ichitani, K.; Inoue, H.; Nishida, H.; Okumoto, Y.; Tanisaka, T. Interactive effects of two heading-time loci, Se1 and Ef1, on pre-flowering developmental phases in rice (Oryza sativa L.). Euphytica. 2002, 12, 227–234. [Google Scholar] [CrossRef]
- Saito, H.; Okumoto, Y.; Teranishi, T.; Yuan, Q.; Nakazaki, T.; Tanisaka, T. Heading time genes responsible for the regional adaptability of ‘Tongil-type short-culmed rice cultivars’ developed in Korea. Breed. Sci. 2007, 57, 135–143. [Google Scholar] [CrossRef][Green Version]
- Kinoshita, T. Report of the committee on gene symbolization, nomenclature and linkage groups. Rice Genet. Let. 1986, 3, 3. [Google Scholar]
- Yano, M.; Sasaki, T. Genetic and molecular dissection of quantitative traits in rice. Plant Mol. Biol. 1997, 35, 145–153. [Google Scholar] [CrossRef]
- Doi, K.; Izawa, T.; Fuse, T.; Yamanouchi, U.; Kubo, T.; Shimatani, Z.; Yano, M.; Yoshimura, A. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Gene Dev. 2004, 18, 926–936. [Google Scholar] [CrossRef]
- Xue, W.; Xing, Y.; Weng, X.; Zhao, Y.; Tang, W.; Wang, L.; Zhou, H.; Yu, S.; Xu, C.; Li, X.; et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 2008, 40, 761–767. [Google Scholar] [CrossRef]
- Yano, K.; Yamamoto, E.; Aya, K.; Takeuchi, H.; Lo, P.C.; Hu, L.; Yamasaki, M.; Yoshida, S.; Kitano, H.; Matsuoka, M. Genome-wide association study using whole genome sequencing rapidly identifies new genes influencing agronomic traits in rice. Nat. Genet. 2016, 48, 927–934. [Google Scholar] [CrossRef]
- Okumoto, Y.; Tanisaka, T. Trisomic analysis of a strong photoperiod-sensitivity gene E1 in rice (Oryza sativa L.). Euphytica 1997, 95, 301–307. [Google Scholar] [CrossRef]
- Fujino, K.; Sekiguchi, H. Mapping of QTLs conferring extremely early heading in rice (Oryza sativa L.). Theor. Appl. Genet. 2005, 111, 393–398. [Google Scholar] [CrossRef]
- Nonoue, Y.; Fujino, K.; Hirayama, Y.; Yamanouchi, U.; Lin, S.Y.; Yano, M. Detection of quantitative trait loci controlling extremely early heading in rice. Theor. Appl. Genet. 2008, 116, 715–722. [Google Scholar] [CrossRef] [PubMed]
- Fujino, K.; Yamanouchi, U.; Yano, M. Roles of the Hd5 gene controlling heading date for adaptation to the northern limits of rice cultivation. Theor. Appl. Genet. 2013, 126, 611–618. [Google Scholar] [CrossRef] [PubMed]
- Inoue, H.; Nishida, H.; Okumoto, Y.; Tanisaka, T. Identification of an early heading time gene found in the Taiwanese rice cultivar Taichung 65. Breed. Sci. 1998, 48, 103–108. [Google Scholar] [CrossRef]
- Tsai, K.H.; Oka, H. Genetic studies of yielding capacity and adaptability in crop plants. 4. Effects on an earliness gene, mb in the genetic background of a rice variety, Taichung 65. Bot. Bull. Acad. Sinica. 1970, 11, 16–26. [Google Scholar]
- Itoh, Y.; Sano, Y. Phyllochron dynamics under controlled environments in rice (Oryza sativa L.). Euphytica 2006, 150, 87–95. [Google Scholar] [CrossRef]
- Eiguchi, M.; Sano, Y. Evolutionary significance of chromosome 7 in an annual type of wild rice. Rice Genet. Newsl. 1995, 12, 187. [Google Scholar]
- The Rice Annotation Project Database. Available online: http://rapdb.dna.affrc.go.jp (accessed on 29 August 2019).
- Lu, L.; Yan, W.H.; Shao, D.; Xing, Y.Z. Evolution and association analysis of Ghd7 in rice. PLoS ONE 2012, 7, e34021. [Google Scholar] [CrossRef]
- Vergara, B.S.; Chang, T.T. The Flowering Response of the Rice Plant to Photoperiod: A Review of the Literature, 2nd ed.; IRRI: Manila, Philippine, 1985; pp. 1–65. [Google Scholar]
- Izawa, T. Adaptation of flowering-time by natural and artificial selection in Arabidopsis and rice. J. Exp. Bot. 2011, 58, 3091–3097. [Google Scholar] [CrossRef]
- Tsuji, H.; Taoka, K.; Shimamoto, K. Regulation of flowering in rice: Two florigen genes, a complex gene network, and natural variation. Curr. Opin. Plant Biol. 2011, 14, 45–52. [Google Scholar] [CrossRef]
- Baker, S.S.; Wilhelm, K.S.; Thomashow, M.F. The 5’-region of Arabidopsis thaliana cor15a has cis-acting elements that confer cold-, drought- and ABA-regulated gene expression. Plant Mol. Biol. 1994, 24, 701–713. [Google Scholar] [CrossRef]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 1994, 6, 251–264. [Google Scholar]
- Kim, H.J.; Kim, Y.K.; Park, J.Y.; Kim, J. Light signaling mediated by phytochrome plays an important role in cold-induced gene expression through the C-repeat/dehydration responsive element (C/DRE) in Arabidopsis thaliana. Plant J. 2002, 29, 693–704. [Google Scholar] [CrossRef] [PubMed]
- Itoh, H.; Izawa, T. The coincidence of critical day length recognition for florigen gene expression and floral transition under long-day conditions in rice. Mol. Plant 2013, 6, 635–649. [Google Scholar] [CrossRef] [PubMed]
- Osugi, A.; Itoh, H.; Ikeda-Kawakatsu, K.; Takano, M.; Izawa, T. Molecular dissection of the roles of phytochrome in photoperiodic flowering in rice. Plant Physiol. 2011, 157, 1128–1137. [Google Scholar] [CrossRef] [PubMed]
- Shinada, H.; Yamamoto, T.; Yamamoto, E.; Hori, K.; Yonemaru, J.; Matsuba, S.; Fujino, K. Historical changes in population structure during rice breeding programs in the northern limits of rice cultivation. Theor. Appl. Genet. 2014, 127, 995–1004. [Google Scholar] [CrossRef]
- Yano, M.; Katayose, Y.; Ashikari, M.; Yamanouchi, U.; Monna, L.; Fuse, T.; Baba, T.; Yamamoto, T.; Umehara, Y.; Nagamura, Y.; et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 2001, 12, 2473–2484. [Google Scholar] [CrossRef]
- Fujino, K.; Wu, J.; Sekiguchi, H.; Ito, T.; Izawa, T.; Matsumoto, T. Multiple introgression events surrounding the Hd1 flowering-time gene in cultivated rice, Oryza sativa L. Mol. Gen. Genom. 2010, 284, 137–146. [Google Scholar] [CrossRef]
- Wu, W.; Zheng, X.M.; Lu, G.; Zhong, Z.; Gao, H.; Chen, L.; Wu, C.; Wang, H.J.; Wang, Q.; Zhou, K.; et al. Association of functional nucleotide polymorphisms at DTH2 with the northward expansion of rice cultivation in Asia. Proc. Natl. Acad. Sci. USA 2013, 110, 2775–2780. [Google Scholar] [CrossRef]
- Gao, H.; Jin, M.; Zheng, X.M.; Chen, J.; Yuan, D.; Xin, Y.; Wang, M.; Huang, D.; Zhang, Z.; Zhou, K.; et al. Days to heading 7, a major quantitative locus determining photoperiod sensitivity and regional adaptation in rice. Proc. Natl. Acad. Sci. USA 2014, 111, 16337–16342. [Google Scholar] [CrossRef]
- Zhang, J.; Zhou, X.; Yan, W.; Zhang, Z.; Lu, L.; Han, Z.; Zhao, H.; Liu, H.; Song, P.; Hu, Y.; et al. Combinations of the Ghd7, Ghd8 and Hd1 genes largely define the ecogeographical adaptation and yield potential of cultivated rice. New Phytol. 2015, 208, 1056–1066. [Google Scholar] [CrossRef]
- Zheng, X.M.; Feng, L.; Wang, J.; Qiao, W.; Zhang, L.; Cheng, Y.; Yang, Q. Nonfunctional alleles of long-day suppressor genes independently regulate flowering time. J. Int. Plant Biol. 2016, 58, 540–548. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, Y.; Teshima, K.M.; Yokoi, S.; Innan, H.; Shimamoto, K. Variations in Hd1 proteins, Hd3a promoters, and Ehd1 expression levels contribute to diversity of flowering time in cultivated rice. Proc. Natl. Acad. Sci. USA 2009, 106, 4555–4560. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Xu, J.; Guo, H.; Jiang, L.; Chen, S.; Yu, C.; Zhou, Z.; Hu, P.; Zhai, H.; Wan, J. DTH8 suppresses flowering in rice, influencing plant height and yield potential simultaneously. Plant Physiol. 2010, 153, 1747–1758. [Google Scholar] [CrossRef] [PubMed]
- Ebana, K.; Shibaya, T.; Wu, J.; Matsubara, K.; Kanamori, H.; Yamane, H.; Yamanouchi, U.; Mizubayashi, T.; Kono, I.; Shomura, A.; et al. Uncovering of major genetic factors generating naturally occurring variation in heading date among Asian rice cultivars. Theor. Appl. Genet. 2011, 122, 1199–1210. [Google Scholar] [CrossRef]
- Koo, B.H.; Yoo, S.C.; Park, J.W.; Kwon, C.T.; Lee, B.D.; An, G.; Zhang, Z.; Li, J.; Li, Z.; Paek, N.C. Natural variation in OsPRR37 regulates heading date and contributes to rice cultivation at a wide range of latitudes. Mol. Plant 2013, 6, 1877–1888. [Google Scholar] [CrossRef]
- Ebana, K.; Kojima, K.; Fukuoka, S.; Nagamine, T.; Kawase, M. Development of mini core collection of Japanese rice landrace. Breed Sci. 2008, 58, 281–291. [Google Scholar] [CrossRef]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Saito, H.; Okumoto, Y.; Tsukiyama, T.; Xu, C.; Teraishi, M.; Tanisaka, T. Allelic Differentiation at the E1/Ghd7 Locus Has Allowed Expansion of Rice Cultivation Area. Plants 2019, 8, 550. https://doi.org/10.3390/plants8120550
Saito H, Okumoto Y, Tsukiyama T, Xu C, Teraishi M, Tanisaka T. Allelic Differentiation at the E1/Ghd7 Locus Has Allowed Expansion of Rice Cultivation Area. Plants. 2019; 8(12):550. https://doi.org/10.3390/plants8120550
Chicago/Turabian StyleSaito, Hiroki, Yutaka Okumoto, Takuji Tsukiyama, Chong Xu, Masayoshi Teraishi, and Takatoshi Tanisaka. 2019. "Allelic Differentiation at the E1/Ghd7 Locus Has Allowed Expansion of Rice Cultivation Area" Plants 8, no. 12: 550. https://doi.org/10.3390/plants8120550
APA StyleSaito, H., Okumoto, Y., Tsukiyama, T., Xu, C., Teraishi, M., & Tanisaka, T. (2019). Allelic Differentiation at the E1/Ghd7 Locus Has Allowed Expansion of Rice Cultivation Area. Plants, 8(12), 550. https://doi.org/10.3390/plants8120550




