New Insights on Lilium Phylogeny Based on a Comparative Phylogenomic Study Using Complete Plastome Sequences
Abstract
1. Introduction
2. Results
2.1. Newly Sequenced Plastomes of Three Lilium Species
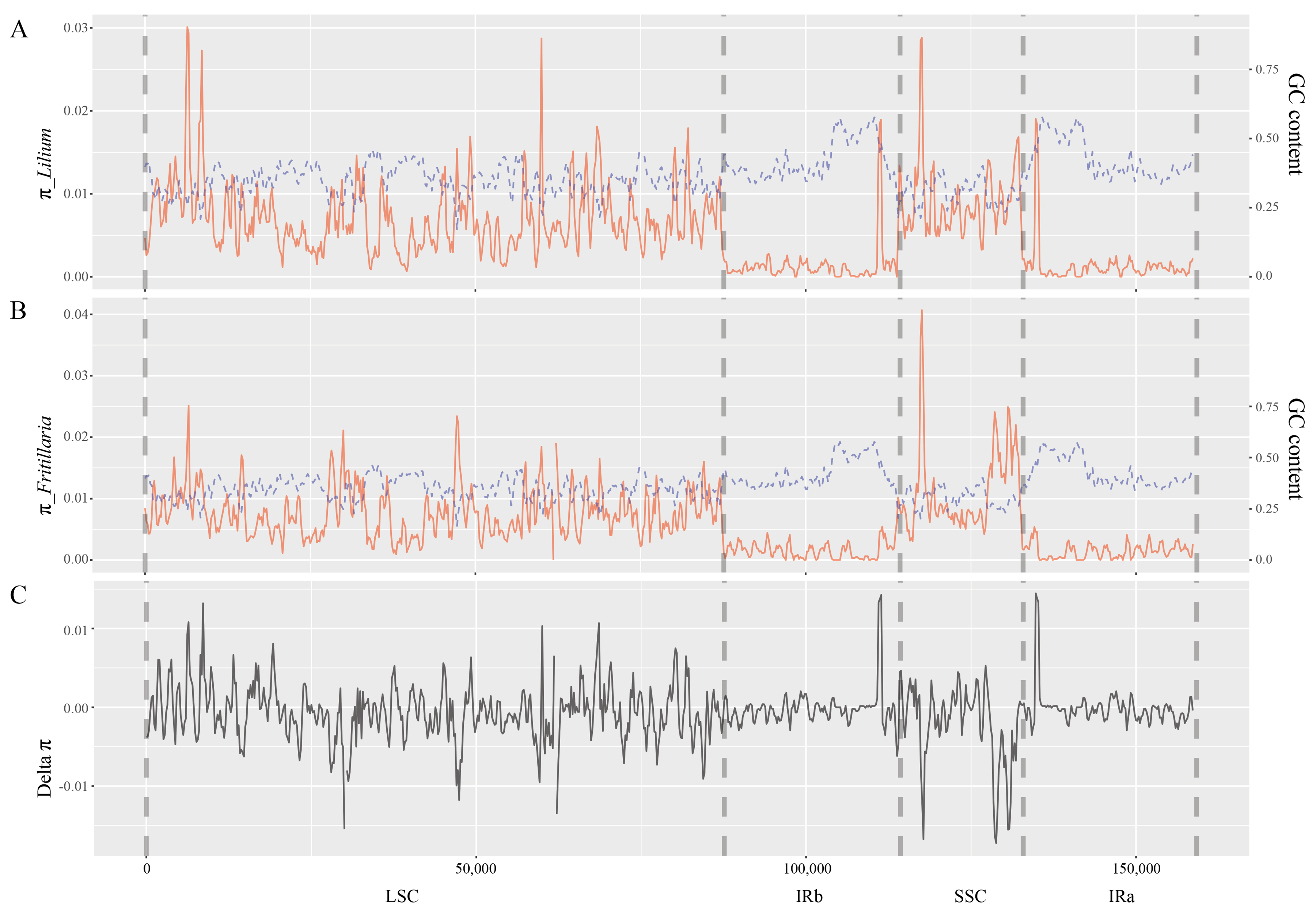
2.2. Nucleotide Diversity within Genera Lilium and Fritillaria
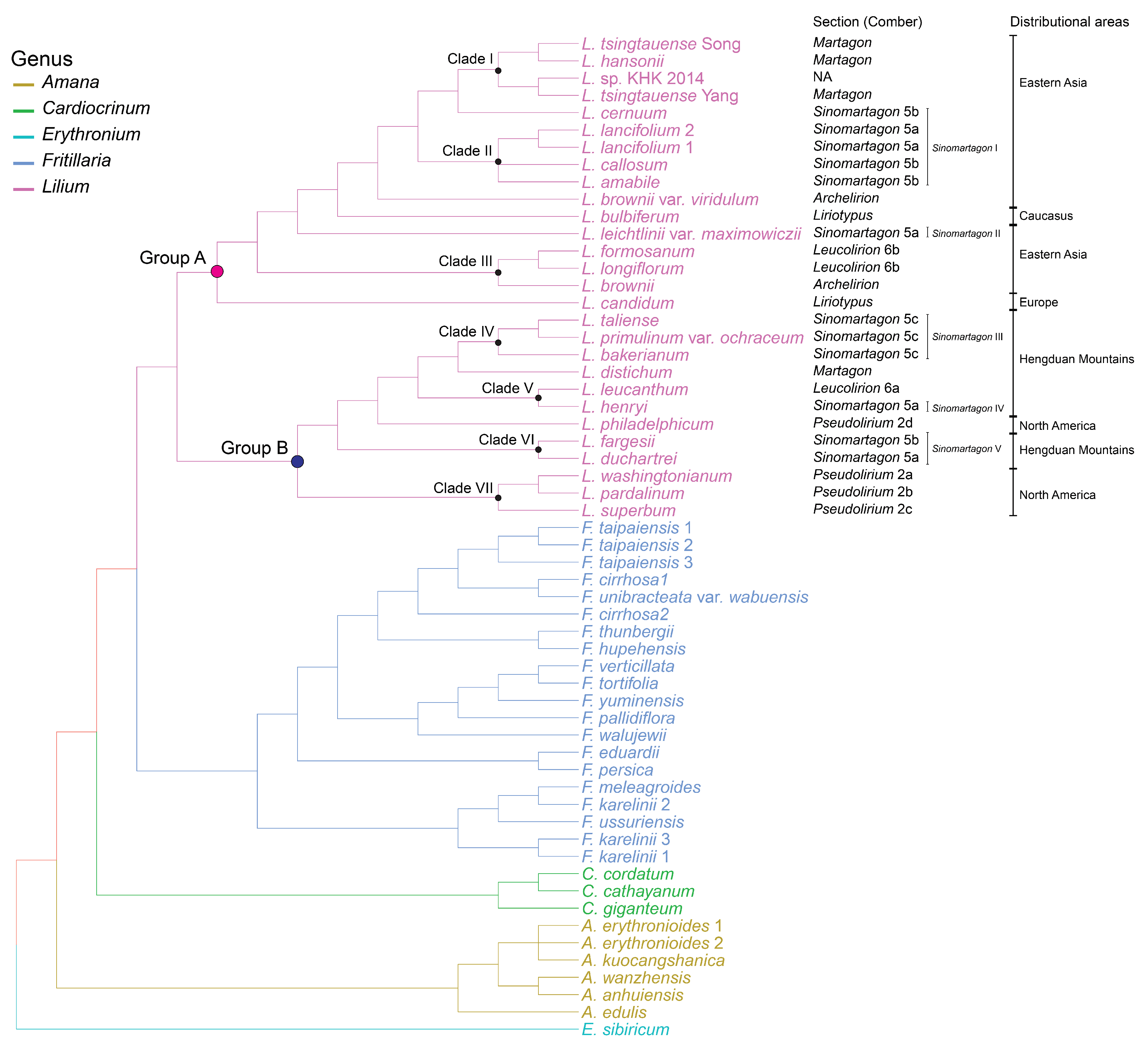
2.3. Phylogeny of Lilium in Liliaceae
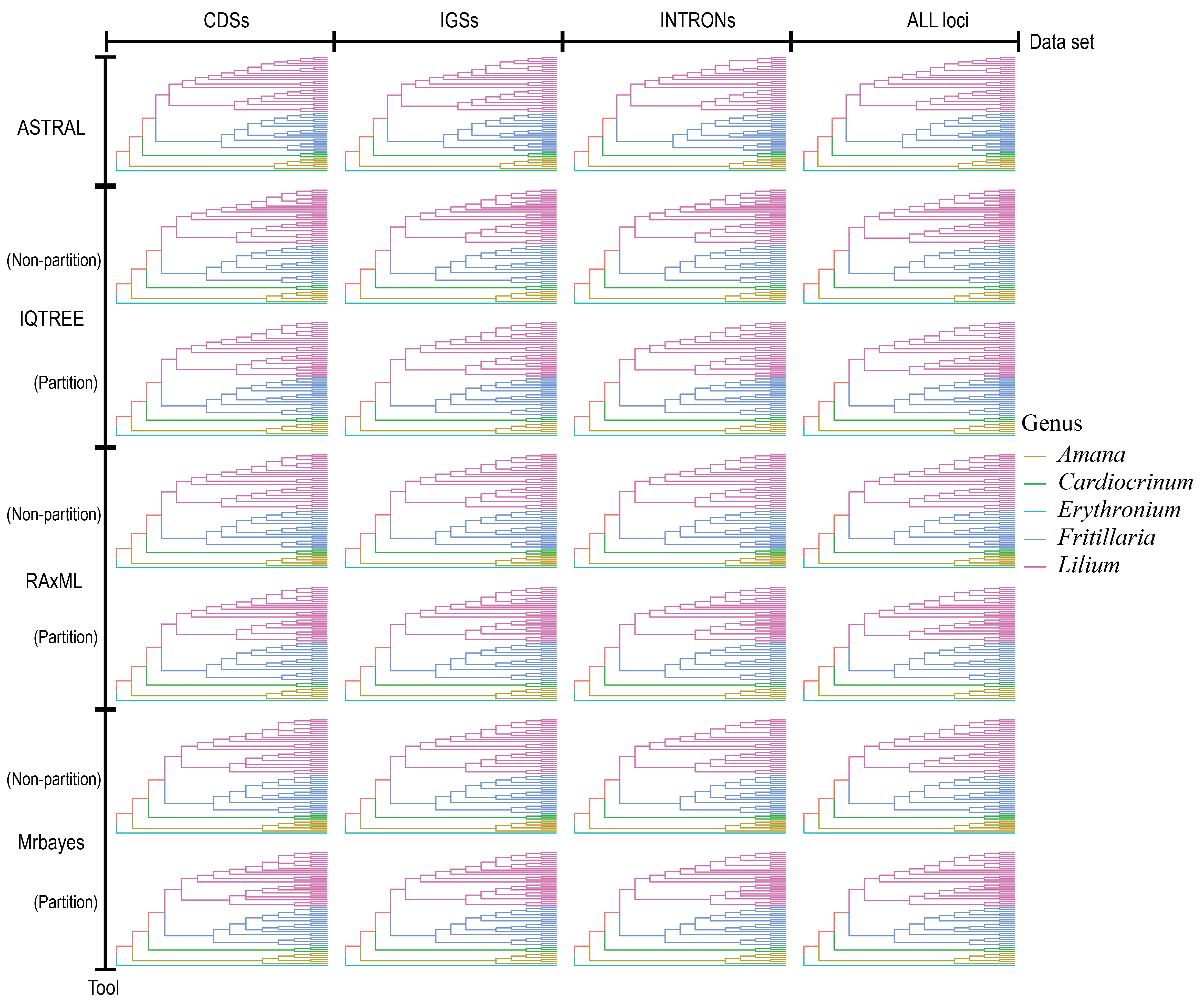
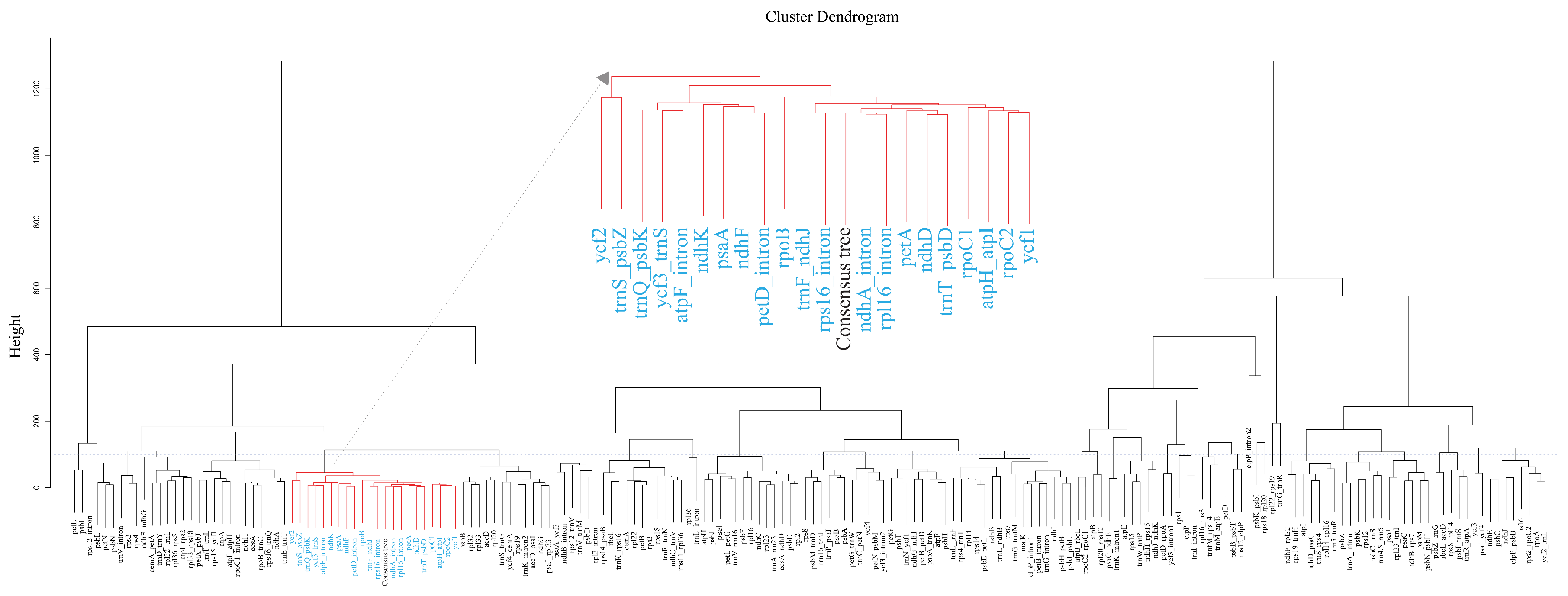
2.4. Comparing Gene Trees to Species Trees
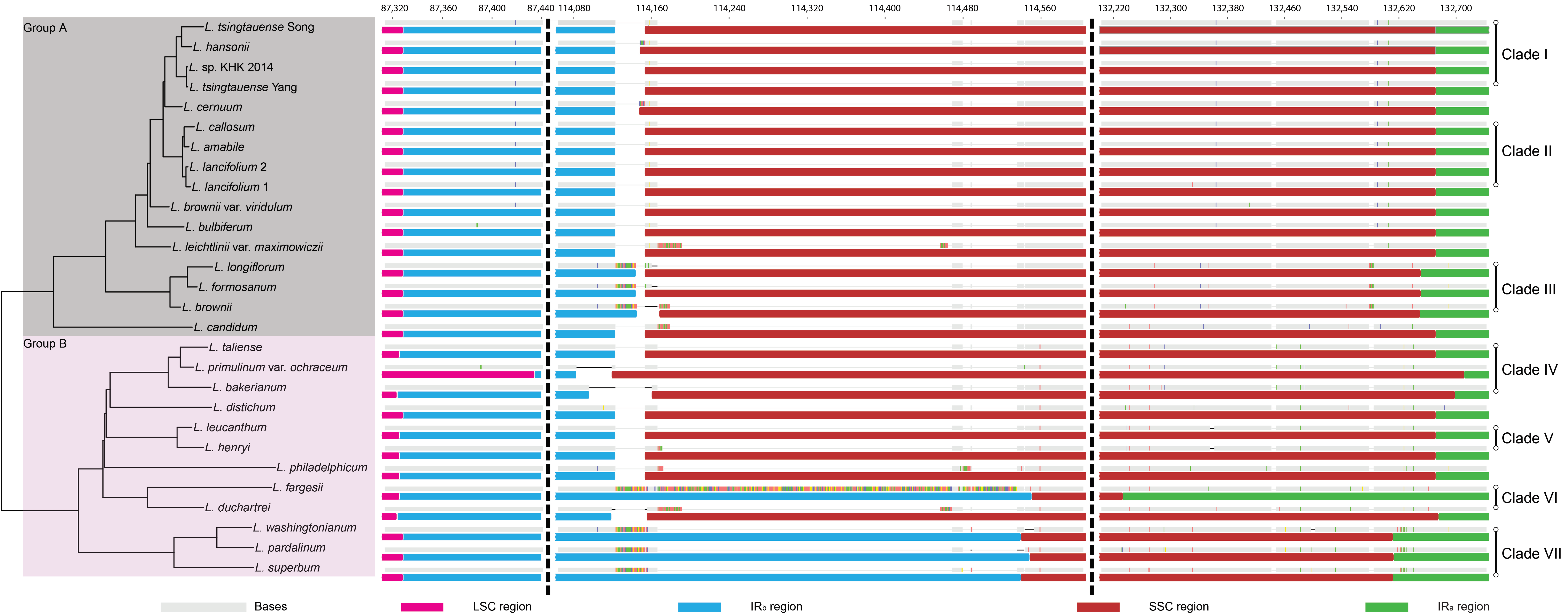
2.5. IR Expansion and Contraction in Lilium
2.6. Insertions/Deletions in Lilium
3. Discussion
3.1. Evolution of Lilium Plastomes in Liliaceae
3.2. Verification of Comber’s Sectional Classification
3.2.1. The Phylogenetic Position of Martagon
3.2.2. The Polyphyly of Pseudolirium
3.2.3. The Polyphyly of Leucolirion
3.2.4. The Position of Liriotypus in the Genus Lilium
3.2.5. Biogeographic History of the Genus Lilium
3.3. Molecular Markers for the Phylogeny of Lilium and its Relatives
4. Materials and Methods
4.1. Plant Materials and DNA Extraction
4.2. Sequencing, Assembly, and Annotation
4.3. Sequence Diversity and GC Content Analyses
4.4. Phylogenetic Analysis
4.5. Comparison of Gene Trees
4.6. R Packages for Manipulation of Phylogenies
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Haw, S.G.; Liang, S.-Y. The Lilies of China: The Genera Lilium, Cardiocrinum, Nomocharis and Notholirion; Timber Press: Portland, OR, USA, 1986. [Google Scholar]
- McRae, E.A.; Austin-Mcrae, E.; MacRae, E. Lilies: A Guide for Growers and Collectors; Timber Press: Portland, OR, USA, 1998; Volume 105. [Google Scholar]
- Lim, K.-B.; van Tuyl, J.M.L. Flower Breeding and Genetics; Springer: Berlin, Germany, 2007; pp. 517–537. [Google Scholar]
- Du, Y.-P.; Bi, Y.; Yang, F.-P.; Zhang, M.-F.; Chen, X.-Q.; Xue, J.; Zhang, X.-H. Complete chloroplast genome sequences of Lilium: Insights into evolutionary dynamics and phylogenetic analyses. Sci. Rep. 2017, 7, 5751. [Google Scholar] [CrossRef] [PubMed]
- Comber, H.F. A New Classification of the Genus Lilium; Royal Horticultural Society: London, UK, 1949; Volume 13, pp. 85–105. [Google Scholar]
- Endlicher, S. Genera plantarum. Vindobonae. Apud Fr. Beck Universitatis Bibliopolam 1836. Available online: https://bibdigital.rjb.csic.es/records/item/10951-redirection (accessed on 25 November 2019).
- Reichenbach, H. Flora Germanica Excursoria.–Leipzig. 1830. Available online: https://www.biodiversitylibrary.org/item/7359#page/3/mode/1up (accessed on 25 November 2019).
- Baker, J. The Gardeners’ Chronicle and and agricultural gazette. 1871. Available online: https://www.biodiversitylibrary.org/page/16448237#page/1163/mode/1up (accessed on 27 November 2019).
- De Jong, P. Some notes on the evolution of lilies. Lily Yearb. N. Am. Lily Soc. 1974, 27, 23–28. [Google Scholar]
- Nishikawa, T.; Okazaki, K.; Uchino, T.; Arakawa, K.; Nagamine, T. A molecular phylogeny of Lilium in the internal transcribed spacer region of nuclear ribosomal DNA. J. Mol. Evol. 1999, 49, 238–249. [Google Scholar] [CrossRef] [PubMed]
- Nishikawa, T.; Okazaki, K.; Arakawa, K.; Nagamine, T. Phylogenetic analysis of section Sinomartagon in genus Lilium using sequences of the internal transcribed spacer region in nuclear ribosomal DNA. Breed. Sci. 2001, 51, 39–46. [Google Scholar] [CrossRef]
- Lee, C.S.; Kim, S.-C.; Yeau, S.H.; Lee, N.S. Major lineages of the genus Lilium (Liliaceae) based on nrDNA ITS sequences, with special emphasis on the Korean species. J. Plant Biol. 2011, 54, 159–171. [Google Scholar] [CrossRef]
- Du, Y.P.; He, H.B.; Wang, Z.X.; Li, S.; Wei, C.; Yuan, X.N.; Cui, Q.; Jia, G.X. Molecular phylogeny and genetic variation in the genus Lilium native to China based on the internal transcribed spacer sequences of nuclear ribosomal DNA. J. Plant Res. 2014, 127, 249–263. [Google Scholar] [CrossRef]
- Du Puy, D.; Cribb, P. The Genus Cymbidium; Royal Botanic Gardens: Sydney, Australia, 2007. [Google Scholar]
- Sharma, S.K.; Dkhar, J.; Kumaria, S.; Tandon, P.; Rao, S.R. Assessment of phylogenetic inter-relationships in the genus Cymbidium (Orchidaceae) based on internal transcribed spacer region of rDNA. Gene 2012, 495, 10–15. [Google Scholar] [CrossRef]
- van den Berg, C.; Ryan, A.; Cribb, P.J.; Chase, M.W. Molecular phylogenetics of Cymbidium (Orchidaceae: Maxillarieae): Sequence data from internal transcribed spacers (ITS) of nuclear ribosomal DNA and plastid matK. Lindleyana 2002, 17, 102–111. [Google Scholar]
- Yukawa, T.; Miyoshi, K.; Yokoyama, J. Molecular phylogeny and character evolution of Cymbidium (Orchidaceae). Bull. Natl. Sci. Mus. Tokyo B 2002, 28, 129–139. [Google Scholar]
- Zwickl, D.J.; Hillis, D.M. Increased taxon sampling greatly reduces phylogenetic error. Syst. Biol. 2002, 51, 588–598. [Google Scholar] [CrossRef]
- Graybeal, A. Is it better to add taxa or characters to a difficult phylogenetic problem? Syst. Biol. 1998, 47, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Heath, T.A.; Hedtke, S.M.; Hillis, D.M. Taxon sampling and the accuracy of phylogenetic analyses. J. Syst. Evol. 2008, 46, 239–257. [Google Scholar] [CrossRef]
- Rokas, A.; Carroll, S.B. More genes or more taxa? The relative contribution of gene number and taxon number to phylogenetic accuracy. Mol. Biol. Evol. 2005, 22, 1337–1344. [Google Scholar] [CrossRef]
- Rosenberg, M.S.; Kumar, S. Incomplete taxon sampling is not a problem for phylogenetic inference. Proc. Natl. Acad. Sci. USA 2001, 98, 10751–10756. [Google Scholar] [CrossRef] [PubMed]
- Mower, J.P.; Sloan, D.B.; Alverson, A.J. Plant mitochondrial genome diversity: The genomics revolution. In Plant Genome Diversity Volume 1; Springer: Berlin, Germany, 2012; pp. 123–144. [Google Scholar]
- Gregory, T.R.; Nicol, J.A.; Tamm, H.; Kullman, B.; Kullman, K.; Leitch, I.J.; Murray, B.G.; Kapraun, D.F.; Greilhuber, J.; Bennett, M.D. Eukaryotic genome size databases. Nucleic Acids Res. 2006, 35, D332–D338. [Google Scholar] [CrossRef] [PubMed]
- Ruhlman, T.A.; Jansen, R.K. The plastid genomes of flowering plants. Methods Mol. Biol. 2014, 1132, 3–38. [Google Scholar] [CrossRef]
- Weng, M.L.; Blazier, J.C.; Govindu, M.; Jansen, R.K. Reconstruction of the ancestral plastid genome in Geraniaceae reveals a correlation between genome rearrangements, repeats, and nucleotide substitution rates. Mol. Biol. Evol. 2014, 31, 645–659. [Google Scholar] [CrossRef]
- Guisinger, M.M.; Kuehl, J.V.; Boore, J.L.; Jansen, R.K. Extreme reconfiguration of plastid genomes in the angiosperm family Geraniaceae: Rearrangements, repeats, and codon usage. Mol. Biol. Evol. 2011, 28, 583–600. [Google Scholar] [CrossRef]
- Westwood, J.H.; Yoder, J.I.; Timko, M.P.; dePamphilis, C.W. The evolution of parasitism in plants. Trends Plant Sci. 2010, 15, 227–235. [Google Scholar] [CrossRef]
- Shaw, J.; Lickey, E.B.; Schilling, E.E.; Small, R.L. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: The tortoise and the hare III. Am. J. Bot. 2007, 94, 275–288. [Google Scholar] [CrossRef]
- Shaw, J.; Lickey, E.B.; Beck, J.T.; Farmer, S.B.; Liu, W.; Miller, J.; Siripun, K.C.; Winder, C.T.; Schilling, E.E.; Small, R.L. The tortoise and the hare II: Relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. Am. J. Bot. 2005, 92, 142–166. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.B.; Tang, M.; Li, H.T.; Zhang, Z.R.; Li, D.Z. Complete chloroplast genome of the genus Cymbidium: Lights into the species identification, phylogenetic implications and population genetic analyses. BMC Evol. Biol. 2013, 13, 84. [Google Scholar] [CrossRef]
- Zhang, Y.J.; Ma, P.F.; Li, D.Z. High-throughput sequencing of six bamboo chloroplast genomes: Phylogenetic implications for temperate woody bamboos (Poaceae: Bambusoideae). PLoS ONE 2011, 6, e20596. [Google Scholar] [CrossRef] [PubMed]
- Wysocki, W.P.; Clark, L.G.; Attigala, L.; Ruiz-Sanchez, E.; Duvall, M.R. Evolution of the bamboos (Bambusoideae; Poaceae): A full plastome phylogenomic analysis. BMC Evol. Biol. 2015, 15, 50. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.; Liu, J.; Yu, J.; Wang, L.; Zhou, S. Highly variable chloroplast markers for evaluating plant phylogeny at low taxonomic levels and for DNA barcoding. PLoS ONE 2012, 7, e35071. [Google Scholar] [CrossRef] [PubMed]
- Sarkinen, T.; George, M. Predicting plastid marker variation: Can complete plastid genomes from closely related species help? PLoS ONE 2013, 8, e82266. [Google Scholar] [CrossRef] [PubMed]
- Ku, C.; Hu, J.M.; Kuo, C.H. Complete plastid genome sequence of the basal asterid Ardisia polysticta Miq. and comparative analyses of asterid plastid genomes. PLoS ONE 2013, 8, e62548. [Google Scholar] [CrossRef] [PubMed]
- Ku, C.; Chung, W.C.; Chen, L.L.; Kuo, C.H. The complete plastid genome sequence of madagascar periwinkle Catharanthus roseus (L.) G. Don: Plastid genome evolution, molecular marker identification, and phylogenetic implications in asterids. PLoS ONE 2013, 8, e68518. [Google Scholar] [CrossRef]
- Kim, H.T.; Kim, J.S.; Lee, Y.M.; Mun, J.-H.; Kim, J.-H. Molecular markers for phylogenetic applications derived from comparative plastome analysis of Prunus. J. Syst. Evol. 2019, 57, 15–22. [Google Scholar] [CrossRef]
- Kim, J.-H.; Lee, S.-I.; Kim, B.-R.; Choi, I.-Y.; Ryser, P.; Kim, N.-S. Chloroplast genomes of Lilium lancifolium, L. amabile, L. callosum, and L. philadelphicum: Molecular characterization and their use in phylogenetic analysis in the genus Lilium and other allied genera in the order Liliales. PLoS ONE 2017, 12, e0186788. [Google Scholar] [CrossRef]
- Maddison, W.P. Gene trees in species trees. Syst. Biol. 1997, 46, 523–536. [Google Scholar] [CrossRef]
- Gao, Y.D.; Harris, A.J.; Zhou, S.D.; He, X.J. Evolutionary events in Lilium (including Nomocharis, Liliaceae) are temporally correlated with orogenies of the Q-T plateau and the Hengduan Mountains. Mol. Phylogenet. Evol. 2013, 68, 443–460. [Google Scholar] [CrossRef] [PubMed]
- Yu, G. A tidy tool for phylogenetic tree data manipulation [R package tidytree version 0.1. 9]. Compr. R Arch. Netw. 2019. Available online: https://rdrr.io/cran/tidytree/ (accessed on 25 November 2019).
- Jombart, T.; Kendall, M.; Almagro-Garcia, J.; Colijn, C. Treespace: Statistical exploration of landscapes of phylogenetic trees. Mol. Ecol. Resour. 2017, 17, 1385–1392. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Wang, S.; Ding, Y.; Xu, J.; Li, M.F.; Zhu, S.; Chen, N. Chloroplast genomic resource of Paris for species discrimination. Sci. Rep. 2017, 7, 3427. [Google Scholar] [CrossRef] [PubMed]
- Lighty, R. The lilies of Korea. Lily Yearb. RHS 1969, 31, 31–39. [Google Scholar]
- Hayashi, K.; Kawano, S. Molecular systematics of Lilium and allied genera (Liliaceae): Phylogenetic relationships among Lilium and related genera based on the rbcL and matK gene sequence data. Plant Species Biol. 2000, 15, 73–93. [Google Scholar] [CrossRef]
- Kim, H.T.; Zale, P.J.; Lim, K.-B. Complete plastome sequence of Lilium pardalinum Kellogg (Liliaceae). Mitochondrial DNA Part B 2018, 3, 478–479. [Google Scholar] [CrossRef]
- İkinci, N.; Oberprieler, C.; Güner, A. On the origin of European lilies: Phylogenetic analysis of Lilium section Liriotypus (Liliaceae) using sequences of the nuclear ribosomal transcribed spacers. Willdenowia 2006, 36, 647–656. [Google Scholar] [CrossRef][Green Version]
- Muratović, E.; Hidalgo, O.; Garnatje, T.; Siljak-Yakovlev, S. Molecular phylogeny and genome size in European lilies (Genus Lilium, Liliaceae). Adv. Sci. Lett. 2010, 3, 180–189. [Google Scholar] [CrossRef]
- Patterson, T.B.; Givnish, T.J. Phylogeny, concerted convergence, and phylogenetic niche conservatism in the core Liliales: Insights from rbcL and ndhF sequence data. Evolution 2002, 56, 233–252. [Google Scholar] [CrossRef]
- Rønsted, N.; Law, S.; Thornton, H.; Fay, M.F.; Chase, M.W. Molecular phylogenetic evidence for the monophyly of Fritillaria and Lilium (Liliaceae; Liliales) and the infrageneric classification of Fritillaria. Mol. Phylogenet. Evol. 2005, 35, 509–527. [Google Scholar] [CrossRef] [PubMed]
- Fay, M.F.; Chase, M.W.; Rønsted, N.; Devey, D.S.; Pillon, Y.; Pires, J.C.; Peterson, G.; Seberg, O.; Davis, J.I. Phylogenetics of Lilliales. Aliso J. Syst. Evol. Bot. 2006, 22, 559–565. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 2004, 5, 113. [Google Scholar] [CrossRef]
- Team, R.C. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- Lanfear, R.; Frandsen, P.B.; Wright, A.M.; Senfeld, T.; Calcott, B. PartitionFinder 2: New methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 2016, 34, 772–773. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the 2010 Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; pp. 1–8. [Google Scholar]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Hohna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2014, 32, 268–274. [Google Scholar] [CrossRef]
- Zhang, C.; Rabiee, M.; Sayyari, E.; Mirarab, S. ASTRAL-III: Polynomial time species tree reconstruction from partially resolved gene trees. BMC Bioinform. 2018, 19, 153. [Google Scholar] [CrossRef]
- Mirarab, S.; Warnow, T. ASTRAL-II: Coalescent-based species tree estimation with many hundreds of taxa and thousands of genes. Bioinformatics 2015, 31, i44–i52. [Google Scholar] [CrossRef]
- Mirarab, S.; Reaz, R.; Bayzid, M.S.; Zimmermann, T.; Swenson, M.S.; Warnow, T. ASTRAL: Genome-scale coalescent-based species tree estimation. Bioinformatics 2014, 30, i541–i548. [Google Scholar] [CrossRef]
- Kendall, M.; Colijn, C. A tree metric using structure and length to capture distinct phylogenetic signals. arXiv 2015, arXiv:preprint/1507.05211. [Google Scholar]
- Paradis, E.; Claude, J.; Strimmer, K. APE: Analyses of phylogenetics and evolution in R language. Bioinformatics 2004, 20, 289–290. [Google Scholar] [CrossRef]
- Pagès, H.; Aboyoun, P.; Gentleman, R.; DebRoy, S. Biostrings: Efficient manipulation of biological strings. R Package Version 2018, 2. Available online: https://rdrr.io/bioc/Biostrings/ (accessed on 25 November 2019).
- Wickham, H.; Francois, R.; Henry, L.; Müller, K. Dplyr: A grammar of data manipulation. R Package Version 2018, 3. Available online: https://rdrr.io/cran/dplyr/ (accessed on 25 November 2019).
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer: Berlin, Germany, 2016. [Google Scholar]
- Yu, G.; Smith, D.K.; Zhu, H.; Guan, Y.; Lam, T.T.Y. Ggtree: An R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 2017, 8, 28–36. [Google Scholar] [CrossRef]
- Yu, G.; Lam, T.T.-Y.; Zhu, H.; Guan, Y. Two methods for mapping and visualizing associated data on phylogeny using ggtree. Mol. Biol. Evol. 2018, 35, 3041–3043. [Google Scholar] [CrossRef]
- Auguie, B. GridExtra: Functions in Grid graphics. R Package Version 0.9 2012, 1. Available online: http://CRAN.R-project.org/package=gridExtra (accessed on 25 November 2019).
- Paradis, E. Pegas: An R package for population genetics with an integrated–modular approach. Bioinformatics 2010, 26, 419–420. [Google Scholar] [CrossRef]
- Revell, L.J. Phytools: An R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 2012, 3, 217–223. [Google Scholar] [CrossRef]
- Xinqi, C.; Songyun, L.; Jiemei, X.; Tamura, M. Liliaceae. Flora China 2000, 24, 73–263. [Google Scholar]
| Taxon | No. of Raw Reads (≥50 bp) | No. of Mapped Reads | Average Coverage | SRA a Accession |
|---|---|---|---|---|
| Lilium candidum | 89,224,524 | 1,187,028 | 1134.7 | SRR7617960, SRR7617961 |
| Lilium leichtlinii var. maximowiczii | 35,028,408 | 568,878 | 520.1 | SRR7617965 |
| Lilium formosanum | 80,985,606 | 1,945,534 | 1840.6 | SRR7617963, SRR7617964 |
| Cluster | Clustered Trees | No. of Trees |
|---|---|---|
| 1 | accD; ndhG; psbB; rpl20; rpl32; rpl33; rps19; accD_psaI; psaJ_rpl33; trnS_trnG; ycf4_cemA; trnK_intron2 | 12 |
| 2 | atpA; ccsA; ndhA; ndhH; atpF_atpH; petA_psbJ; rpoB_trnC; rps15_ycf1; rps16_trnQ; trnE_trnT; trnT_trnL; rpoC1_intron | 12 |
| 3 | atpE; rps15; ndhH_rps15; ndhJ_ndhK; trnW_trnP | 5 |
| 4 | atpF; petG; psbH; psbT; ndhG_ndhI; petB_petD; psbA_trnK; trnN_ycf1 | 8 |
| 5 | atpH; ndhC; psaI; psbE; psbF; psbJ; rpl2; rpl16; rpl23; ccsA_ndhD; petL_petG; trnA_rrn23; trnV_rrn16 | 13 |
| 6 | atpI; ndhD_psaC; ndhF_rpl32; rpl14_rpl16; rps19_trnH; rrn5_trnR; trnS_rps4 | 7 |
| 7 | cemA; petB; rbcL; rpl22; rps3; rps18; ndhC_trnV; rps11_rpl36; rps14_psaB; trnK_rps16; trnR_trnN | 11 |
| 8 | clpP; rps11; trnI_intron | 3 |
| 9 | matK; ndhB; ndhI; rpl14; rps7; rps14; psbE_petL; psbH_petB; psbJ_psbL; rps4_trnT; trnG_trnfM; trnL_ndhB; trnL_trnF; clpP_intron1; petB_intron; trnG_intron | 16 |
| 10 | ndhD; ndhF; ndhK; petA; psaA; rpoB; rpoC1; rpoC2; ycf1; ycf2; atpH_atpI; trnF_ndhJ; trnQ_psbK; trnS_psbZ; trnT_psbD; ycf3_trnS; atpF_intron; ndhA_intron; petD_intron; rpl16_intron; rps16_intron; Consensus tree of species trees | 22 |
| 11 | ndhE; ndhJ; psbC; rpoA; rps16; clpP_psbB; rps2_rpoC2; ycf2_trnL | 8 |
| 12 | petN; psbL; psbN; rps12_intron | 4 |
| 13 | psaB; psbA; rps8; psbM_trnD; rrn16_trnI; trnP_psaJ | 6 |
| 14 | psaC; psaJ; psbM; ndhB_rps7; psbN_psbH; rpl23_trnI | 6 |
| 15 | psbD; psaA_ycf3; rps12_trnV; trnV_trnM; ndhB_intron; rpl2_intron | 6 |
| 16 | psbK; psbZ; rps12; psbC_trnS; rrn4.5_rrn5; trnA_intron | 6 |
| 17 | rps2; rps4; trnV_intron | 3 |
| 18 | ycf4; petG_trnW; petN_psbM; trnC_petN; ycf3_intron2 | 5 |
| 19 | atpI_rps2; cemA_petA; ndhE_ndhG; rpl32_trnL; rpl33_rps18; rpl36_rps8; trnD_trnY | 7 |
| 20 | psbI_trnS; rps8_rpl14; trnR_atpA | 3 |
| 21 | rpl16_rps3; trnfM_rps14; trnM_atpE | 3 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, H.T.; Lim, K.-B.; Kim, J.S. New Insights on Lilium Phylogeny Based on a Comparative Phylogenomic Study Using Complete Plastome Sequences. Plants 2019, 8, 547. https://doi.org/10.3390/plants8120547
Kim HT, Lim K-B, Kim JS. New Insights on Lilium Phylogeny Based on a Comparative Phylogenomic Study Using Complete Plastome Sequences. Plants. 2019; 8(12):547. https://doi.org/10.3390/plants8120547
Chicago/Turabian StyleKim, Hyoung Tae, Ki-Byung Lim, and Jung Sung Kim. 2019. "New Insights on Lilium Phylogeny Based on a Comparative Phylogenomic Study Using Complete Plastome Sequences" Plants 8, no. 12: 547. https://doi.org/10.3390/plants8120547
APA StyleKim, H. T., Lim, K.-B., & Kim, J. S. (2019). New Insights on Lilium Phylogeny Based on a Comparative Phylogenomic Study Using Complete Plastome Sequences. Plants, 8(12), 547. https://doi.org/10.3390/plants8120547







