THROUGH THE LOOKING GLASS: Real-Time Imaging in Brachypodium Roots and Osmotic Stress Analysis
Abstract
1. Introduction
2. Results
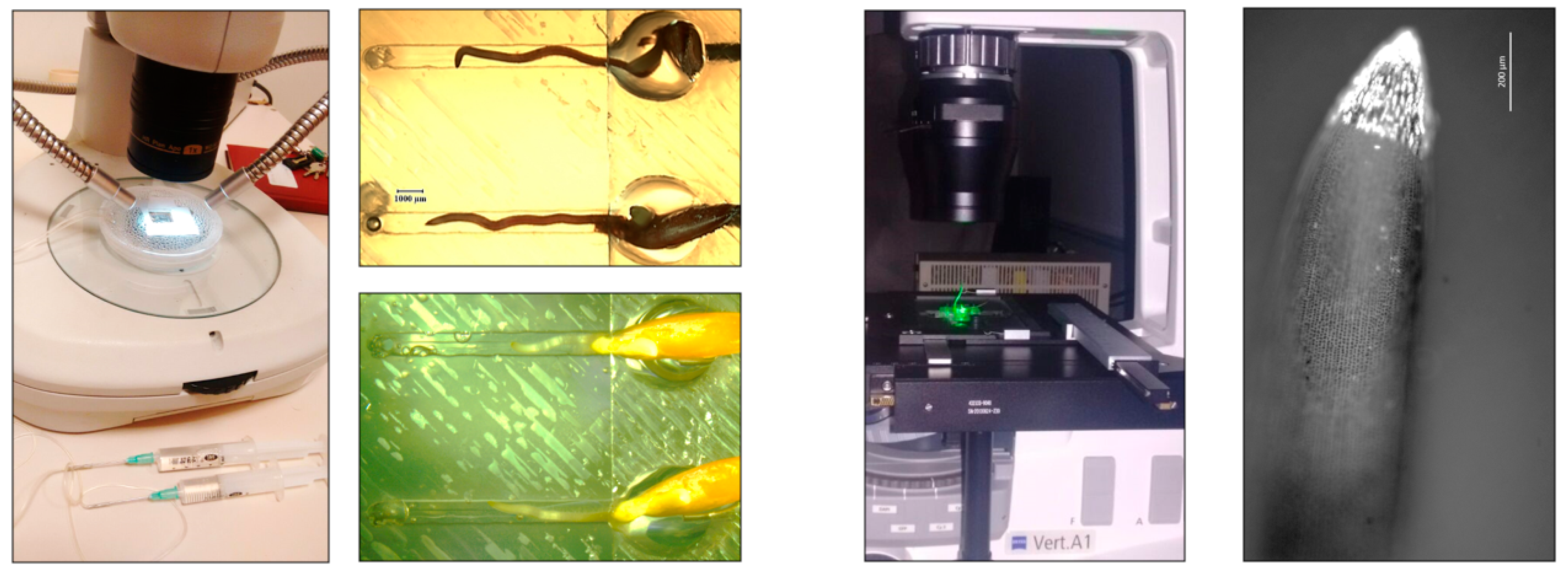
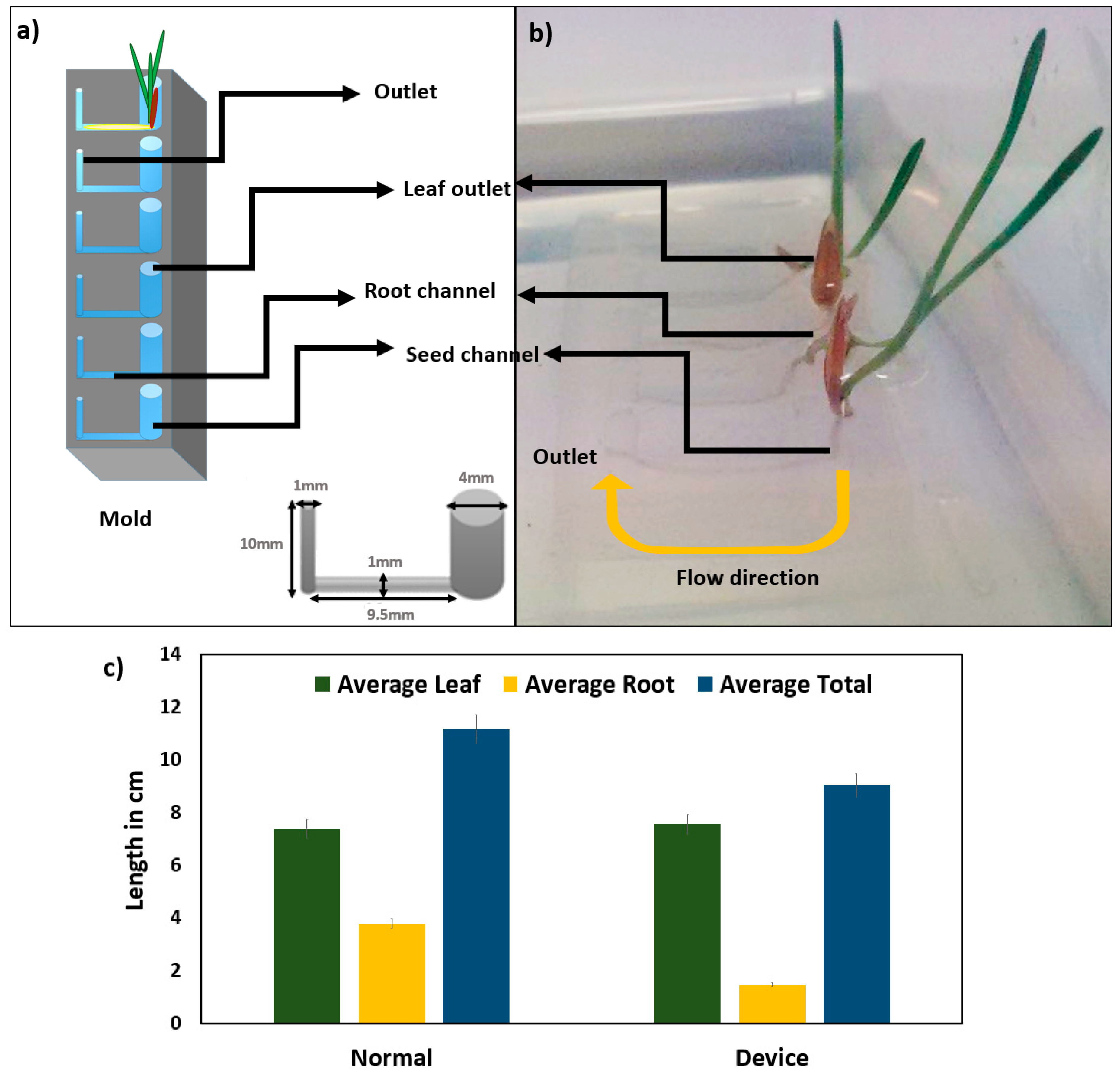
2.1. Plant Growth in PDMS
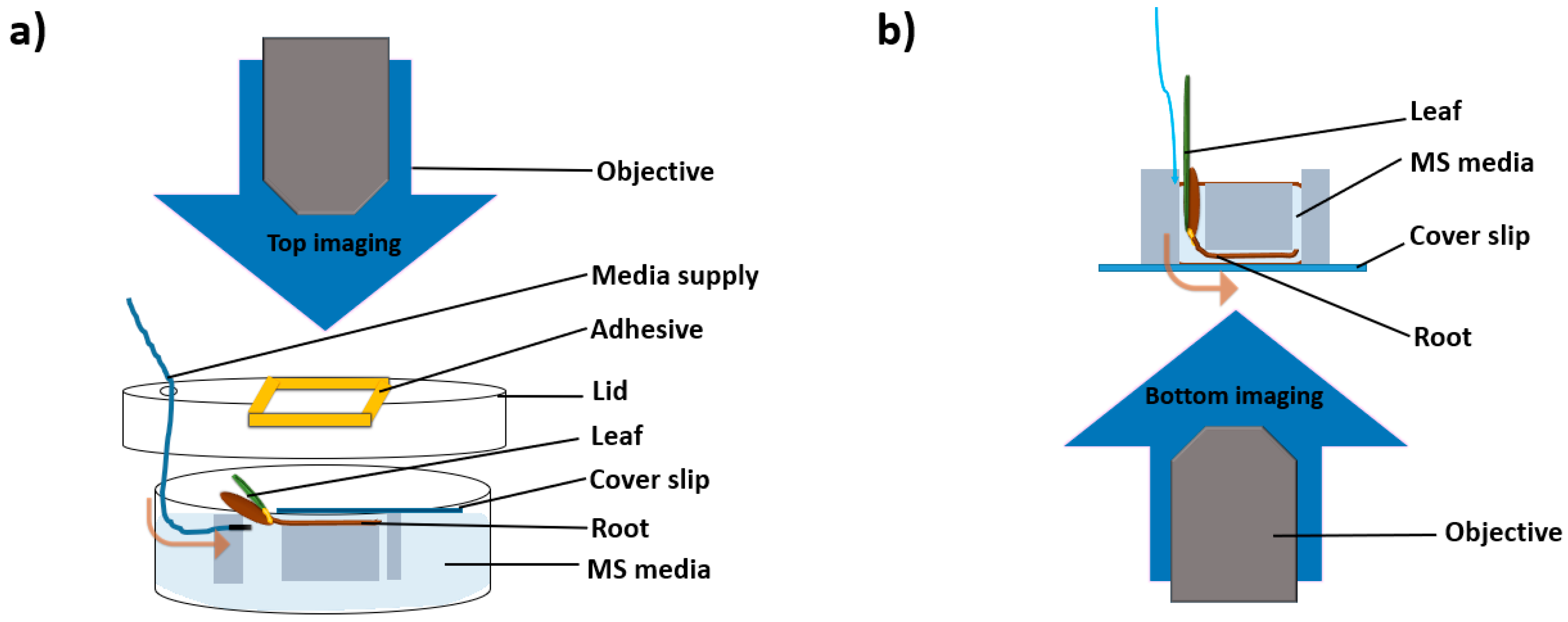
2.2. Microscopy Setups
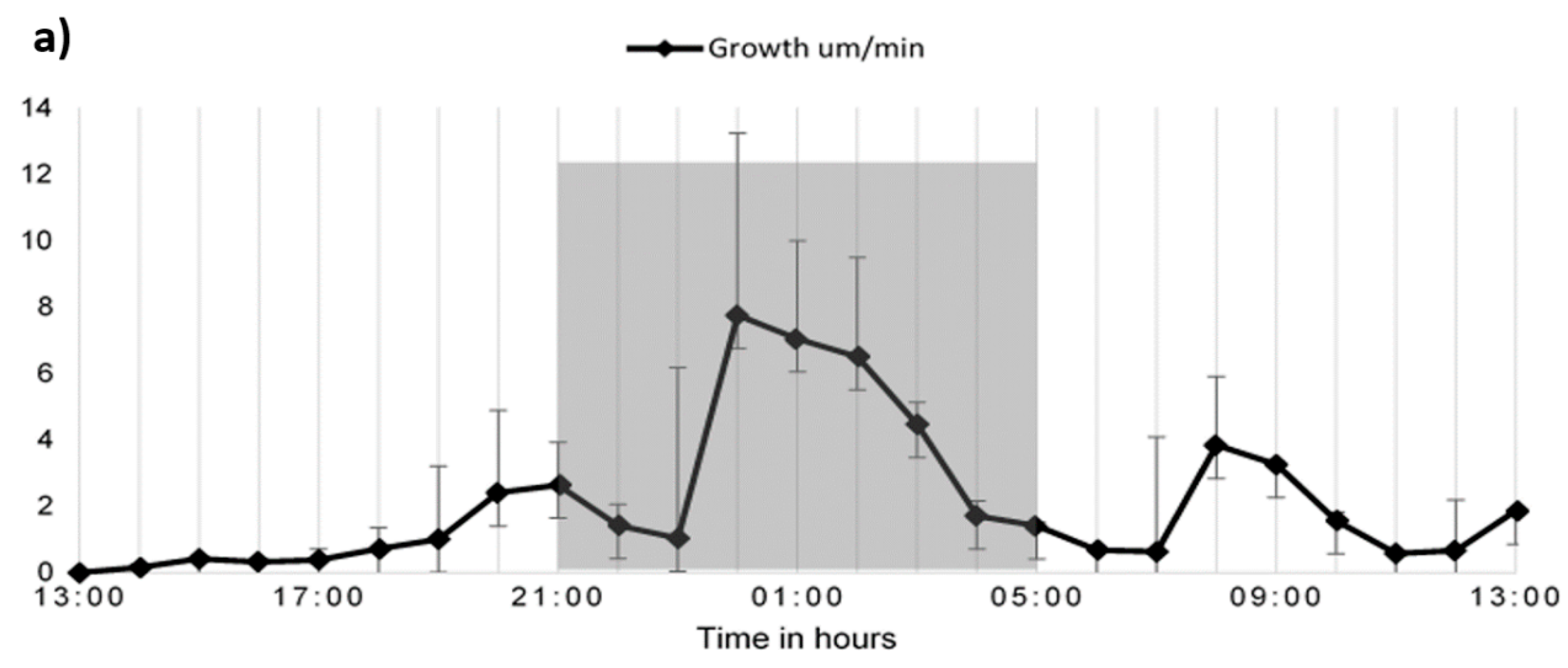
2.3. Time Lapse Growth Curve Analysis
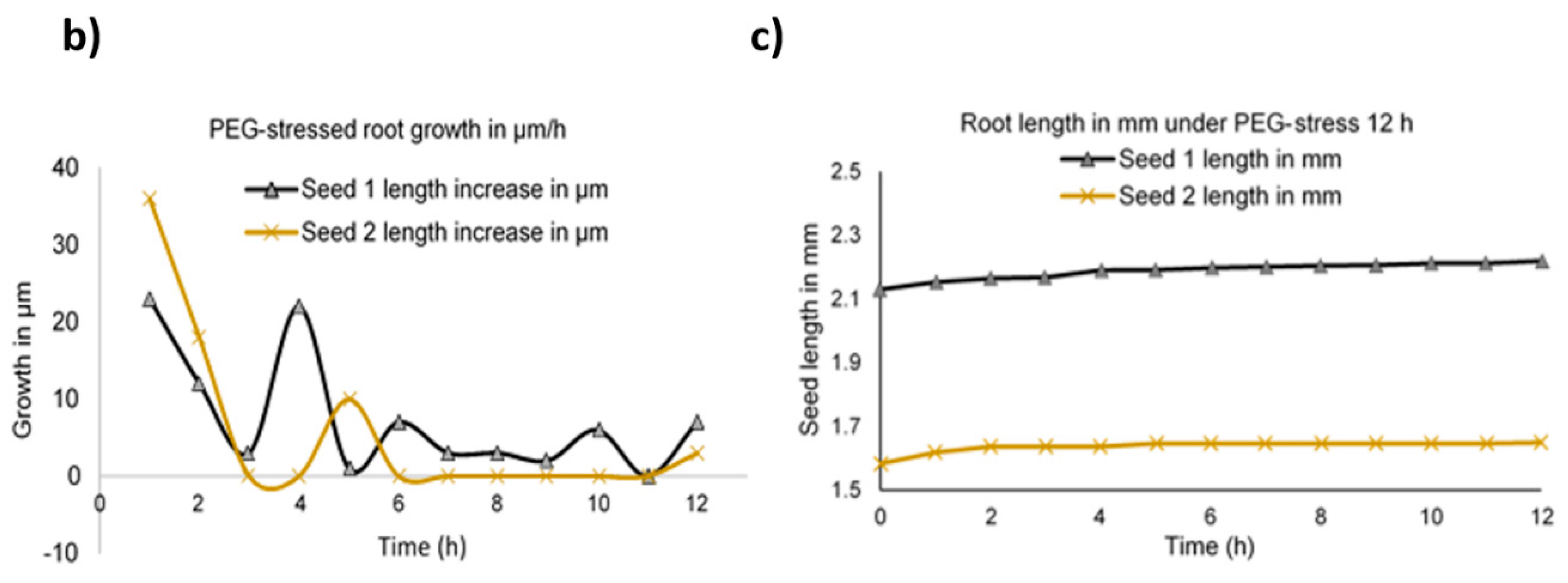
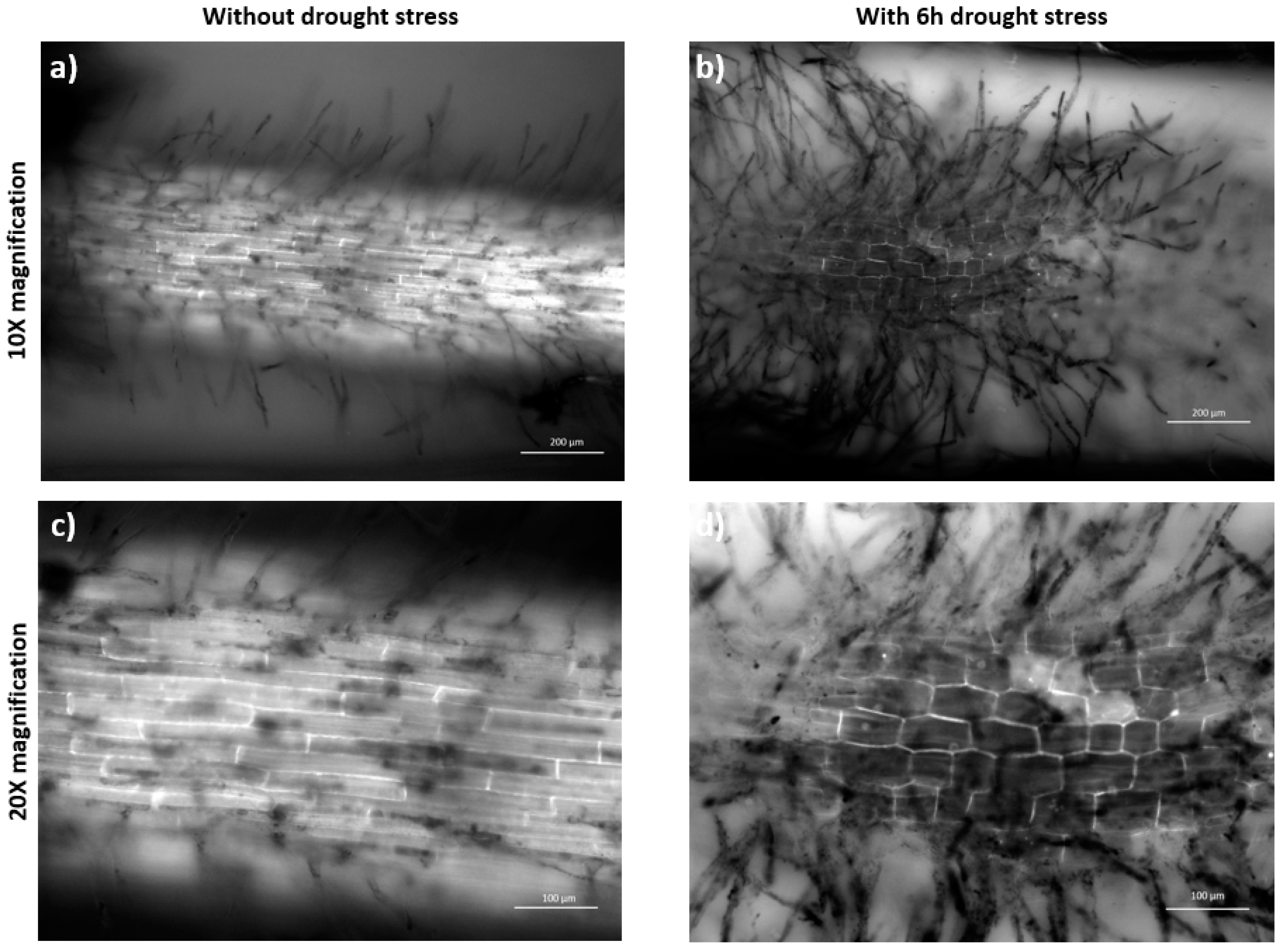
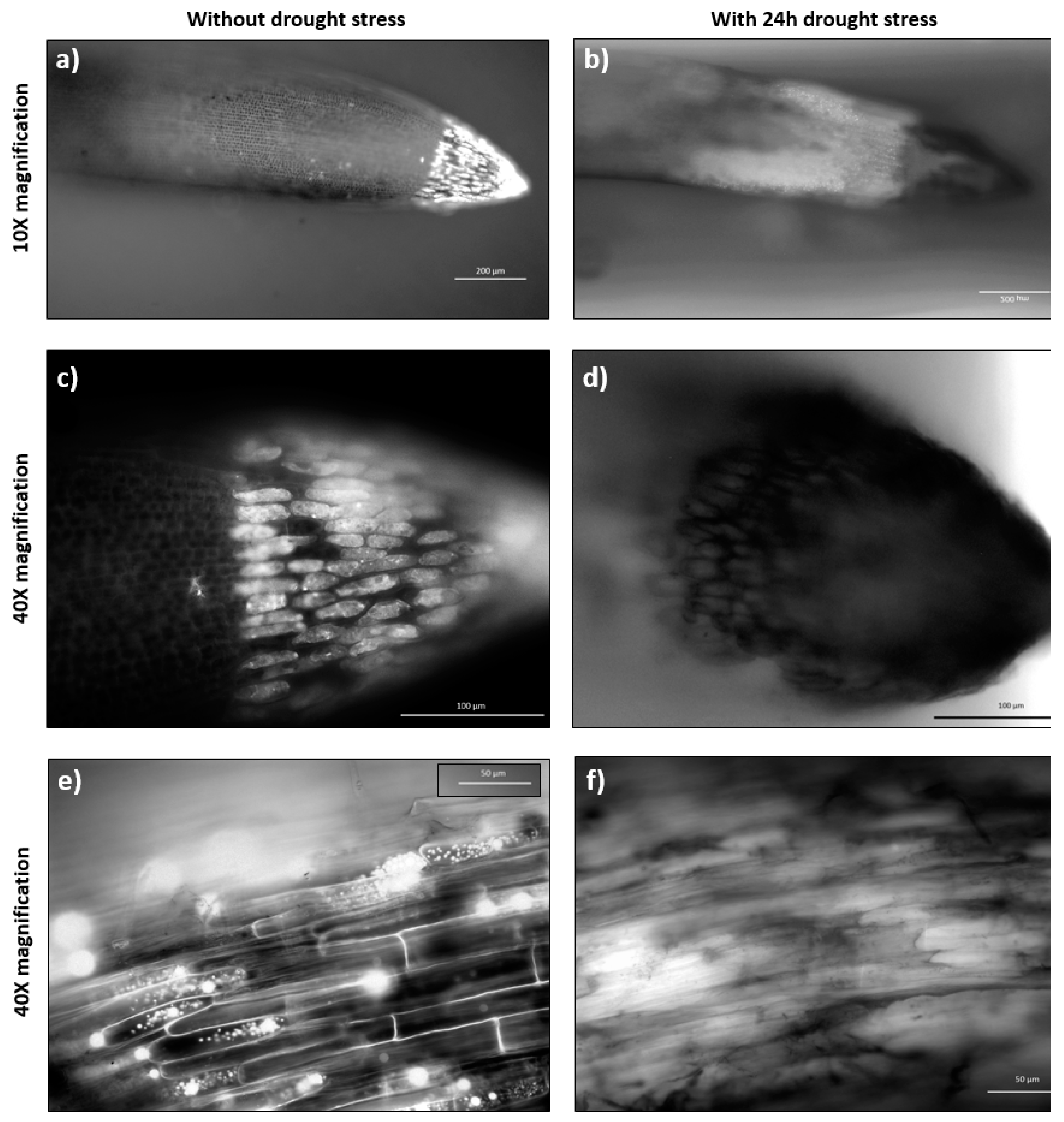
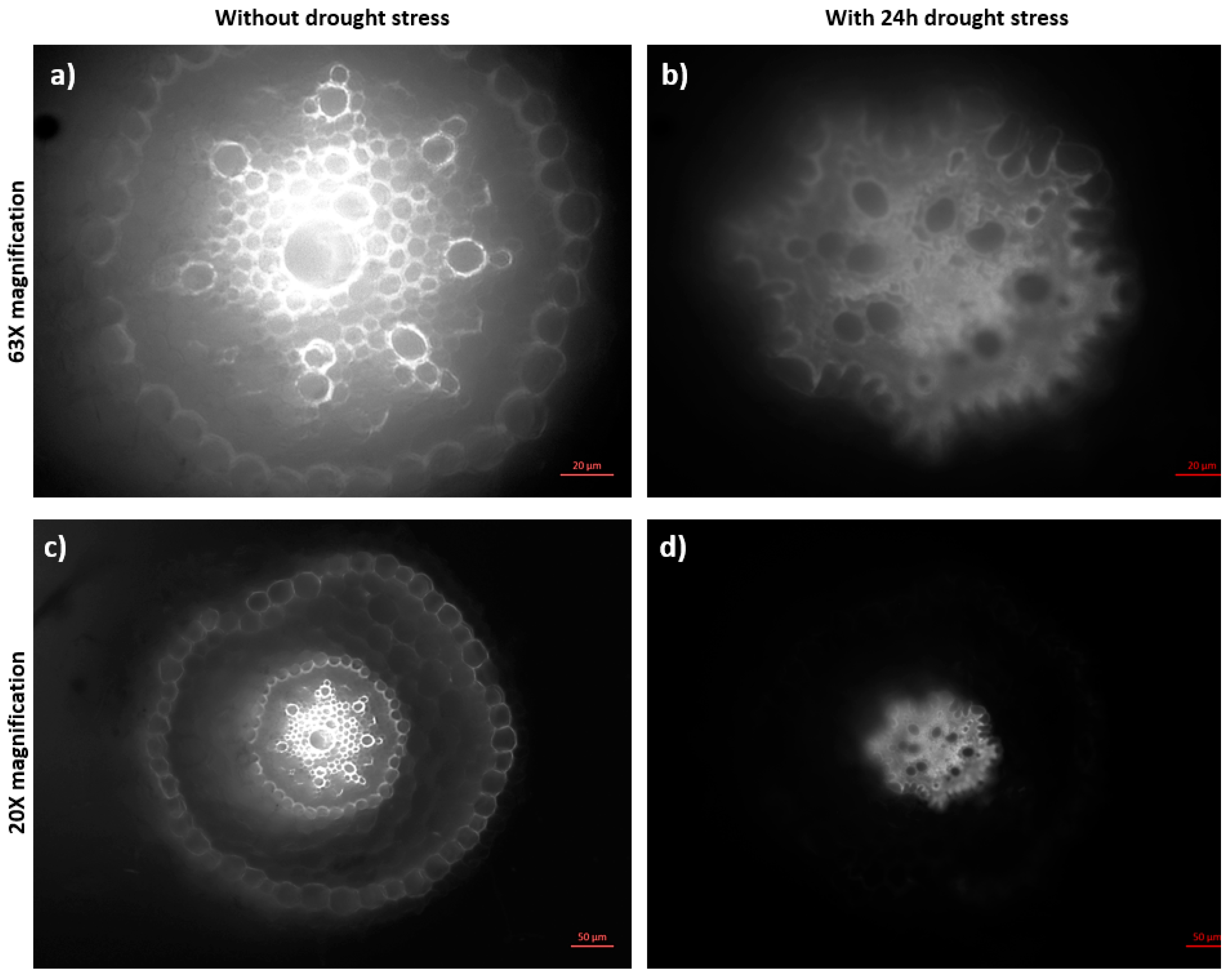
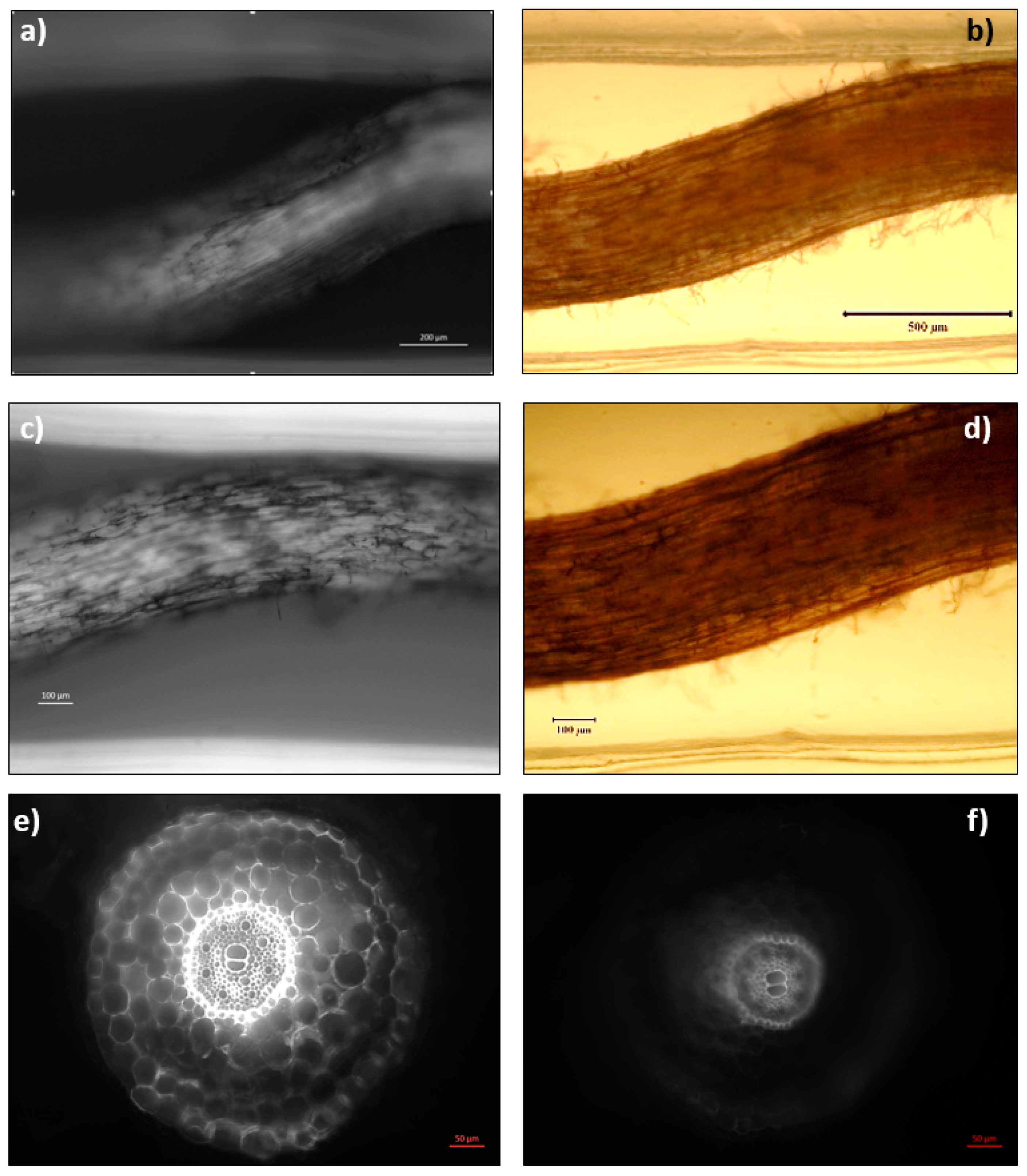
2.4. Abiotic Stress Analysis by Microscopy
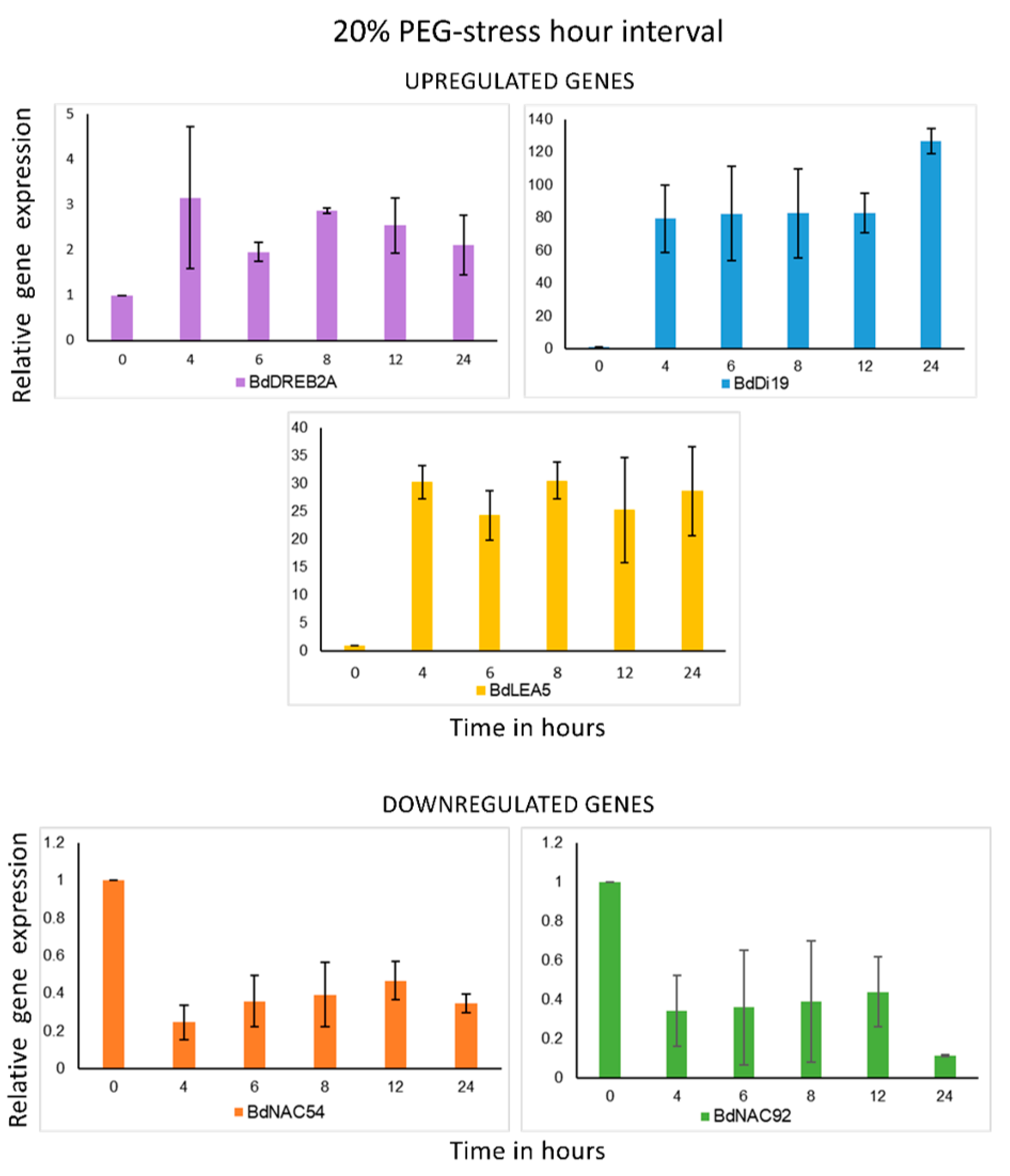
2.5. Gene Expression Analysis
3. Discussion
4. Materials and Methods
4.1. Device Fabrication
4.2. Preparation of Seeds and Measurement of Growth
4.3. Osmotic Stress Application
4.4. Imaging Setups
4.5. RNA Isolation, DNase Treatment and qRT-PCR
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Frazier, T.P.; Sun, G.; Burklew, C.E.; Zhang, B. Salt and drought stresses induce the aberrant expression of microRNA genes in tobacco. Mol. Biotechnol. 2011, 49, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.R.; Pathak, H.; Sharma, S.K.; Kala, Y.K.; Nirjal, M.K.; Singh, G.P.; Goswami, S.; Rai, R.D. Novel and conserved heat-responsive microRNAs in wheat (Triticum aestivum L.). Funct. Integr. Genom. 2015, 15, 323–348. [Google Scholar] [CrossRef] [PubMed]
- Verelst, W.; Bertolini, E.; De Bodt, S.; Vandepoele, K.; Demeulenaere, M.; Enrico Pè, M.; Inzé, D. Molecular and Physiological Analysis of Growth-Limiting Drought Stress in Brachypodium distachyon Leaves. Mol. Plant 2013, 6, 311–322. [Google Scholar] [CrossRef] [PubMed]
- Elitaş, M.; Yüce, M.; Budak, H. Microfabricated tools for quantitative plant biology. Analyst 2017, 142, 835–848. [Google Scholar] [CrossRef] [PubMed]
- Sanati Nezhad, A. Microfluidic platforms for plant cells studies. Lab Chip 2014, 14, 3262–3274. [Google Scholar] [CrossRef]
- Busch, W.; Moore, B.T.; Martsberger, B.; Mace, D.L.; Twigg, R.W.; Jung, J.; Pruteanu-malinici, I.; Kennedy, S.J.; Gregory, K.; Nc, D. A microfluidic device and computational platform for high throughput live imaging of gene expression. Nat. Methods 2012, 9, 1101–1106. [Google Scholar] [CrossRef]
- Yetisen, A.K.; Jiang, L.; Cooper, J.R.; Qin, Y.; Palanivelu, R.; Zohar, Y. A microsystem-based assay for studying pollen tube guidance in plant reproduction. J. Micromech. Microeng. 2011, 21, 054018. [Google Scholar] [CrossRef]
- Grossmann, G.; Guo, W.-J.; Ehrhardt, D.W.; Frommer, W.B.; Sit, R.V.; Quake, S.R.; Meier, M. The RootChip: An integrated microfluidic chip for plant science. Plant Cell 2011, 23, 4234–4240. [Google Scholar] [CrossRef]
- Gooh, K.; Ueda, M.; Aruga, K.; Park, J.; Arata, H.; Higashiyama, T.; Kurihara, D. Live-Cell Imaging and Optical Manipulation of Arabidopsis Early Embryogenesis. Dev. Cell 2015, 34, 242–251. [Google Scholar] [CrossRef]
- Keinath, N.; Waadt, R.; Brugman, R.; Schroeder, J.I.; Grossmann, G.; Schumacher, K.; Krebs, M. Live cell imaging with R-GECO1 sheds light on flg22- and chitin-induced transient [Ca2+]cyt patterns in Arabidopsis. Mol. Plant 2015, 1–13. [Google Scholar] [CrossRef]
- Jiang, H.; Xu, Z.; Aluru, M.R.; Dong, L. Plant chip for high-throughput phenotyping of Arabidopsis. Lab Chip 2014, 14, 1281. [Google Scholar] [CrossRef] [PubMed]
- Meier, M.; Lucchetta, E.M.; Ismagilov, R.F. Chemical stimulation of the Arabidopsis thaliana root using multi-laminar flow on a microfluidic chip. Lab Chip 2010, 10, 2147. [Google Scholar] [CrossRef] [PubMed]
- Massalha, H.; Korenblum, E.; Malitsky, S.; Shapiro, O.H.; Aharoni, A. Live imaging of root–bacteria interactions in a microfluidics setup. Proc. Natl. Acad. Sci. USA 2017, 114, 4549–4554. [Google Scholar] [CrossRef]
- Nezhad, A.S.; Packirisamy, M.; Bhat, R.; Geitmann, A. In Vitro Study of Oscillatory Growth Dynamics of Camellia Pollen Tubes in Microfluidic Environment. IEEE Trans. Biomed. Eng. 2013, 60, 3185–3193. [Google Scholar] [CrossRef] [PubMed]
- Agudelo, C.G.; Packirisamy, M.; Geitmann, A. Assessing the Influence of Electric Cues and Conductivity on Pollen Tube Growth via Lab-On-A-Chip Technology. Biophys. J. 2014, 106, 574a. [Google Scholar] [CrossRef]
- Agudelo, C.G.; Sanati Nezhad, A.; Ghanbari, M.; Naghavi, M.; Packirisamy, M.; Geitmann, A. TipChip: A modular, MEMS-based platform for experimentation and phenotyping of tip-growing cells. Plant J. 2013, 73, 1057–1068. [Google Scholar] [CrossRef] [PubMed]
- Ghanbari, M.; Nezhad, A.S.; Agudelo, C.G.; Packirisamy, M.; Geitmann, A. Microfluidic positioning of pollen grains in lab-on-a-chip for single cell analysis. J. Biosci. Bioeng. 2014, 117, 504–511. [Google Scholar] [CrossRef]
- Iyer-Pascuzzi, A.S.; Symonova, O.; Mileyko, Y.; Hao, Y.; Belcher, H.; Harer, J.; Weitz, J.S.; Benfey, P.N. Imaging and Analysis Platform for Automatic Phenotyping and Trait Ranking of Plant Root Systems. Plant Physiol. 2010, 152, 1148–1157. [Google Scholar] [CrossRef]
- Ko, J.-M.; Ju, J.; Lee, S.; Cha, H.-C. Tobacco protoplast culture in a polydimethylsiloxane-based microfluidic channel. Protoplasma 2006, 227, 237–240. [Google Scholar] [CrossRef]
- Wu, H.; Liu, W.; Tu, Q.; Song, N.; Li, L.; Wang, J.J.; Wang, J.J. Culture and chemical-induced fusion of tobacco mesophyll protoplasts in a microfluidic device. Microfluidics Nanofluidics 2011, 10, 867–876. [Google Scholar] [CrossRef]
- Hung, M.-S.; Chang, J.-H. Developing microfluidics for rapid protoplasts collection and lysis from plant leaf. Proc. Inst. Mech. Eng. Part N J. Nanoeng. Nanosyst. 2012, 226, 15–22. [Google Scholar] [CrossRef]
- Bascom, C.S.; Wu, S.-Z.; Nelson, K.; Oakey, J.; Bezanilla, M. Long-Term Growth of Moss in Microfluidic Devices Enables Subcellular Studies in Development. Plant Physiol. 2016, 172, 28–37. [Google Scholar] [CrossRef] [PubMed]
- Shiono, K.; Yamada, S. Waterlogging tolerance and capacity for oxygen transport in Brachypodium distachyon (Bd21). Plant Root 2014, 8, 5–12. [Google Scholar] [CrossRef]
- Hardtke, C.S.; Pacheco-Villalobos, D. The Brachypodium distachyon Root System: A Tractable Model to Investigate Grass Roots. In Genetics and Genomics of Brachypodium; Vogel, J.P., Ed.; Springer International Publishing: Cham, Switzerland, 2016; pp. 245–258. ISBN 978-3-319-26944-3. [Google Scholar]
- Hong, S.-Y.; Seo, P.; Yang, M.-S.; Xiang, F.; Park, C.-M. Exploring valid reference genes for gene expression studies in Brachypodium distachyon by real-time PCR. BMC Plant Biol. 2008, 8, 112. [Google Scholar] [CrossRef] [PubMed]
- You, J.; Zhang, L.; Song, B.; Qi, X.; Chan, Z. Systematic Analysis and Identification of Stress-Responsive Genes of the NAC Gene Family in Brachypodium distachyon. PLoS ONE 2015, 10, e0122027. [Google Scholar] [CrossRef] [PubMed]
- Gordon, S.P.; Priest, H.; Des Marais, D.L.; Schackwitz, W.; Figueroa, M.; Martin, J.; Bragg, J.N.; Tyler, L.; Lee, C.-R.; Bryant, D.; et al. Genome diversity in Brachypodium distachyon: Deep sequencing of highly diverse inbred lines. Plant J. 2014, 79, 361–374. [Google Scholar] [CrossRef] [PubMed]
- Vogel, J.P.; Garvin, D.F.; Mockler, T.C.; Schmutz, J.; Rokhsar, D.; Bevan, M.W.; Barry, K.; Lucas, S.; Harmon-Smith, M.; Lail, K.; et al. Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature 2010, 463, 763–768. [Google Scholar] [CrossRef] [PubMed]
- Budak, H.; Hernandez, P.; Schulman, A. Analysis and Exploitation of Cereal Genomes with the Aid of Brachypodium. In Genomics of Plant Genetic Resources; Springer: Cham, Switzerland, 2014. [Google Scholar]
- Brkljacic, J.; Grotewold, E.; Scholl, R.; Mockler, T.; Garvin, D.F.; Vain, P.; Brutnell, T.; Sibout, R.; Bevan, M.; Budak, H.; et al. Brachypodium as a Model for the Grasses: Today and the Future. Plant Physiol. 2011, 157, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Girin, T.; David, L.C.; Chardin, C.; Sibout, R.; Krapp, A.; Ferrario-Méry, S.; Daniel-Vedele, F. Brachypodium: A promising hub between model species and cereals. J. Exp. Bot. 2014, 65, 5683–5696. [Google Scholar] [CrossRef]
- Filiz, E.; Ozdemir, B.S.; Budak, F.; Vogel, J.P.; Tuna, M.; Budak, H. Molecular, morphological, and cytological analysis of diverse Brachypodium distachyon inbred lines. Genome 2009, 52, 876–890. [Google Scholar] [CrossRef]
- Oliveira, E.J.; Koehler, A.D.; Rocha, D.I.; Vieira, L.M.; Pinheiro, M.V.M.; de Matos, E.M.; da Cruz, A.C.F.; da Silva, T.C.R.; Tanaka, F.A.O.; Nogueira, F.T.S.; et al. Morpho-histological, histochemical, and molecular evidences related to cellular reprogramming during somatic embryogenesis of the model grass Brachypodium distachyon. Protoplasma 2017, 254, 2017–2034. [Google Scholar] [CrossRef] [PubMed]
- Barrero, J.M.; Jacobsen, J.V.; Talbot, M.J.; White, R.G.; Swain, S.M.; Garvin, D.F.; Gubler, F. Grain dormancy and light quality effects on germination in the model grass Brachypodium distachyon. New Phytol. 2012, 193, 376–386. [Google Scholar] [CrossRef] [PubMed]
- Guillon, F.; Larré, C.; Petipas, F.; Berger, A.; Moussawi, J.; Rogniaux, H.; Santoni, A.; Saulnier, L.; Jamme, F.; Miquel, M.; et al. A comprehensive overview of grain development in Brachypodium distachyon variety Bd21. J. Exp. Bot. 2012, 63, 739–755. [Google Scholar] [CrossRef] [PubMed]
- Nezhad, A.S.; Ghanbari, M.; Agudelo, C.G.; Naghavi, M.; Packirisamy, M.; Bhat, R.B.; Geitmann, A. Optimization of flow assisted entrapment of pollen grains in a microfluidic platform for tip growth analysis. Biomed. Microdevices 2014, 16, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Nezhad, A.S.; Naghavi, M.; Packirisamy, M.; Bhat, R.; Geitmann, A.; Sanati Nezhad, A.; Naghavi, M.; Packirisamy, M.; Bhat, R.; Geitmann, A. Quantification of cellular penetrative forces using lab-on-a-chip technology and finite element modeling. Proc. Natl. Acad. Sci. USA 2013, 110, 8093–8098. [Google Scholar] [CrossRef] [PubMed]
- Nezhad, A.S.; Packirisamy, M.; Geitmann, A. Dynamic, high precision targeting of growth modulating agents is able to trigger pollen tube growth reorientation. Plant J. 2014, 80, 185–195. [Google Scholar] [CrossRef] [PubMed]
- Nezhad, A.S.; Naghavi, M.; Packirisamy, M.; Bhat, R.; Geitmann, A. Quantification of the Young’s modulus of the primary plant cell wall using Bending-Lab-On-Chip (BLOC). Lab Chip 2013, 13, 2599. [Google Scholar] [CrossRef] [PubMed]
- Arata, H.; Higashiyama, T. Poly(dimethylsiloxane)-based microdevices for studying plant reproduction. Biochem. Soc. Trans. 2014, 42, 320–324. [Google Scholar] [CrossRef]
- Horade, M.; Kanaoka, M.M.; Kuzuya, M.; Higashiyama, T.; Kaji, N. A microfluidic device for quantitative analysis of chemoattraction in plants. RSC Adv. 2013, 3, 22301. [Google Scholar] [CrossRef]
- Sato, Y.; Sugimoto, N.; Higashiyama, T.; Arata, H. Quantification of pollen tube attraction in response to guidance by female gametophyte tissue using artificial microscale pathway. J. Biosci. Bioeng. 2015, 120, 697–700. [Google Scholar] [CrossRef]
- Dubrovsky, J.G.; Guttenberger, M.; Saralegui, A.; Napsucialy-Mendivil, S.; Voigt, B.; Baluska, F.; Menzel, D. Neutral red as a probe for confocal laser scanning microscopy studies of plant roots. Ann. Bot. 2006, 97, 1127–1138. [Google Scholar] [CrossRef] [PubMed]
- Yazdanbakhsh, N.; Sulpice, R.; Graf, A.; Stitt, M.; Fisahn, J. Circadian control of root elongation and C partitioning in Arabidopsis thaliana. Plant. Cell Environ. 2011, 34, 877–894. [Google Scholar] [CrossRef] [PubMed]
- Paez-Garcia, A.; Motes, C.; Scheible, W.-R.; Chen, R.; Blancaflor, E.; Monteros, M. Root Traits and Phenotyping Strategies for Plant Improvement. Plants 2015, 4, 334–355. [Google Scholar] [CrossRef] [PubMed]
- Akmal, M.; Hirasawa, T. Growth responses of seminal roots of wheat seedlings to a reduction in the water potential of vermiculite. Plant Soil 2004, 267, 319–328. [Google Scholar] [CrossRef]
- Ji, H.; Liu, L.; Li, K.; Xie, Q.; Wang, Z.; Zhao, X.; Li, X. PEG-mediated osmotic stress induces premature differentiation of the root apical meristem and outgrowth of lateral roots in wheat. J. Exp. Bot. 2014, 65, 4863–4872. [Google Scholar] [CrossRef] [PubMed]
- Hackenberg, M.; Gustafson, P.; Langridge, P.; Shi, B.J. Differential expression of microRNAs and other small RNAs in barley between water and drought conditions. Plant Biotechnol. J. 2015, 13, 2–13. [Google Scholar] [CrossRef]
- Jiang, Y.; Wang, X.; Yu, X.; Zhao, X.; Luo, N.; Pei, Z.; Liu, H.; Garvin, D.F. Quantitative Trait Loci Associated with Drought Tolerance in Brachypodium distachyon. Front. Plant Sci. 2017, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lei, R.; Qiao, W.; Hu, F.; Jiang, H.; Zhu, S. A simple and effective method to encapsulate tobacco mesophyll protoplasts to maintain cell viability. MethodsX 2015, 2, 24–32. [Google Scholar] [CrossRef]
- Li, S.; Xu, C.; Yang, Y.; Xia, G. Functional analysis of TaDi19A, a salt-responsive gene in wheat. Plant. Cell Environ. 2010, 33, 117–129. [Google Scholar] [CrossRef]
- Rice Expression Data Spatio-Temporal Gene Expression of Various Tissues/Organs throughout Entire growth in the Field. Available online: http://ricexpro.dna.affrc.go.jp/GGEP/graph-view.php?featurenum=12093#tabs-2 (accessed on 10 December 2018).
- Kawahara, Y.; Oono, Y.; Wakimoto, H.; Ogata, J.; Kanamori, H.; Sasaki, H.; Mori, S.; Matsumoto, T.; Itoh, T. TENOR: Database for Comprehensive mRNA-Seq Experiments in Rice. Plant Cell Physiol. 2016, 57, e7. [Google Scholar] [CrossRef]
- Schiefelbein Lab. Rapid Preparation of Transverse Sections of Plant Roots|Schiefelbein Lab. Available online: http://sites.lsa.umich.edu/schiefelbein-lab/rapid-preparation-of-transverse-sections-of-plant-roots/ (accessed on 1 April 2017).

| Species Name | Species | Organ Studied | Device Type | Physical Parameters | Ref. |
|---|---|---|---|---|---|
| Camellia japonica | Dicot | Pollen tube | Lab-on-a-chip (LOC) technology | Influence of electric fields and conductivity | [15] |
| Dicot, fungus | Pollen grains, root hairs or fungal spores | Tip Chip (serially arranged microchannels) | Experimentation and phenotyping of chemical gradients, microstructural features, integrated biosensors or directional triggers within the modular microchannels | [16] | |
| Dicot | Pollen grains | Microchannels and inlets/outlets | Protuberance growth of single plant cells in a micro- vitro environment | [36] | |
| Dicot | Pollen grains | Tip Chip | penetrative forces generated in pollen tubes | [37] | |
| Dicot | Pollen tube | Laminar flow based microfluidic device | Ca2+, pectin methylesterase (PME) application for quantitative assessment of chemoattraction | [38] | |
| Dicot | Pollen tube | Device with a knot-shaped microchannel microfluidic | Trapping probability and uniformity of fluid flow conditions | [17] | |
| Dicot | Pollen tube | Trapping microfluidic device | Primary and secondary peak frequencies in oscillatory growth dynamics | [14] | |
| Dicot | Pollen tube | Bending-Lab-On-a-Chip (BLOC) | Flexural rigidity of the pollen tube and Young’s modulus of the cell wall | [39] | |
| Arabidopsis thaliana | Dicot | Plant body/Root | Microfluidic chip platform Root Chip | Monitoring time-resolved growth and cytosolic sugar levels at a subcellular resolution | [8] |
| Dicot | Embryo | PDMS micropillar array | Live-Cell Imaging and Optical Manipulation | [9] | |
| Dicot | Root/Plants | Root Array | Imaged by confocal microscopy | [6] | |
| Dicot | Root | RootChip16 | Identification of defined [Ca2+] cyt oscillations, Forster resonance energy transfer (FRET) | [10] | |
| Dicot | Plant body- pathogen interaction | Plant Chip: vertical and transparent microfluidic for high-throughput phenotyping | Quantitative monitoring of plant phenotypes | [11] | |
| Dicot | Live Root | Plant on-chip platform | Stimuli and Phytohormones 2,4-dichlorophenoxyacetic acid (2,4-D), and its inhibitorN-1-naphthylphthalamic acid (NPA) | [12] | |
| Dicot | Pollen-ovule | Mimicry of in vivo microenvironment of ovule fertilization | Chemoattraction | [7] | |
| Torenia fournieri | Dicot | Pollen tube, ovules | T-shaped microchannel device, micro cage array | Pollen tube chemoattraction, long-term live imaging of ovules | [40] |
| Dicot | Pollen tubes | T-shaped channel | Quantitate the effect of chemoattractants on directional pollen tube growth, UV-irradiation | [41] | |
| Dicot | Pollen Tube | Crossroad device | Net guidance response ratio (GRR) | [42] | |
| Tobacco Nicotiana tabacum | Dicot | Mesophyll Protoplast | Microcolumn array | Microscopic real-time optimization and dynamics of protoplast growth including size change, organelle motion, and cell mass formation | [20] |
| Phalaenopsis | Dicot | Protoplasts | Convex-concave sieving array | Real-time collection and lysis of Phalaenopsis protoplasts | [21] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khan, Z.; Karamahmutoğlu, H.; Elitaş, M.; Yüce, M.; Budak, H. THROUGH THE LOOKING GLASS: Real-Time Imaging in Brachypodium Roots and Osmotic Stress Analysis. Plants 2019, 8, 14. https://doi.org/10.3390/plants8010014
Khan Z, Karamahmutoğlu H, Elitaş M, Yüce M, Budak H. THROUGH THE LOOKING GLASS: Real-Time Imaging in Brachypodium Roots and Osmotic Stress Analysis. Plants. 2019; 8(1):14. https://doi.org/10.3390/plants8010014
Chicago/Turabian StyleKhan, Zaeema, Hande Karamahmutoğlu, Meltem Elitaş, Meral Yüce, and Hikmet Budak. 2019. "THROUGH THE LOOKING GLASS: Real-Time Imaging in Brachypodium Roots and Osmotic Stress Analysis" Plants 8, no. 1: 14. https://doi.org/10.3390/plants8010014
APA StyleKhan, Z., Karamahmutoğlu, H., Elitaş, M., Yüce, M., & Budak, H. (2019). THROUGH THE LOOKING GLASS: Real-Time Imaging in Brachypodium Roots and Osmotic Stress Analysis. Plants, 8(1), 14. https://doi.org/10.3390/plants8010014








