Root Traits and Phenotyping Strategies for Plant Improvement
Abstract
:1. Introduction
2. Key Elements of Root System Architecture Relevant for Crop Productivity
| Root Traits/Phenes | Description | References |
|---|---|---|
| Rooting depth | ||
| Primary root length | Primary root growth is inhibited during P-limitation. | [27] |
| A moderately high rate of nitrate supplies can be inhibitory under some culture conditions. | [25] | |
| Deeper roots provide plants with better access to stored water in the deeper layers of the soil substratum. | [34] | |
| Root tip diameter | Root tips with large diameters have improved root penetration of hard, drying soils. | [40] |
| Gravitropism | Steeper root angles and more robust seedling gravitropic responses (which translates to deeper root systems) results in plants that are more tolerant to drought. | [8] |
| Root hairs | ||
| Root hairs | Proliferation of root hairs is stimulated in P-limited conditions, root hairs can contribute 70% or more of the total root surface area and can be responsible for up to 90% of P acquired. | [40,41,42] |
| Root hairs protect the water status of young root tissue. | [43] | |
| Root hairs improve root penetration of hard, drying soil. | [40] | |
| Rhizosheaths | These protect the water status of young root tissues. | [44] |
| Rhizosheaths can increase zinc uptake from dry soils. | [45] | |
| Their presence is correlated with the aluminum tolerance of root hairs. | [44] | |
| Root Branching | ||
| Length and number of lateral roots (LRs) | LR initiation and emergence is stimulated during P limitation. | [46,47] |
| External nitrate stimulates LR initiation and elongation, whereas a high plant internal nitrate/N status inhibits LR growth. Early LR development can also be systemically inhibited by N deficiency. Reduced frequency of LR branching and longer LR improve N capture from low-N soils in maize. | [25,48,49] | |
| Lateral roots are considered the most active portion of the root system for water uptake and represent the majority of the length and surface area of root systems in various types of plants. | [50] | |
| Shallow/adventitious roots | A reduced gravitropic trajectory of basal roots, adventitious rooting and altered dispersion of lateral roots enable topsoil foraging in response to low P availability. Recombinant inbred lines of common bean with shallow basal roots have better P acquisition in the field. Maize plants with brace and crown roots growing at a more shallow angle are more efficient in N use. | [41,46,51,52,53,54] |
| Cluster roots | Bottlebrush-like clusters of ephemeral rootlets arising from a persistent mother root. These are characteristic of the Proteaceae and several other plant species adapted to extremely low soil fertility. Cluster roots are better able to access nutrients such as P by producing large amounts of exudates containing phosphatases and carboxylates that help release bound P. | [55,56] |
| Crown root number | Reduced crown root number is associated with greater rooting depth, N capture and yield in low N soil. | [57,58] |
| Anatomical root traits | ||
| Root cortical aerenchyma | Root cortical aerenchyma are induced by drought and N, P or potassium limitation in maize. Their formation reduces respiration, nutrient content of root tissues and the metabolic cost of soil exploration. Root cortical aerenchyma increased rooting depth by 15%–30% that led to a corresponding elevation in N capture and biomass/grain yield under N stress. | [36,54,57,58,59,60] |
| Cortical cell file number and cell size | Reduced root cortical cell file number and large root cortical cell size improve maize drought tolerance. | [61,62] |
| Cell wall modification | Suberization/lignification affect radial water conductance, and may be important in reducing water loss from mature roots into dry soil. | [36] |
| Metabolic traits | ||
| Respiration | Reduced root respiration in P-efficient common bean reduces the metabolic cost of soil exploration during P-limitation. | [63,64] |
2.1. Rooting Depth
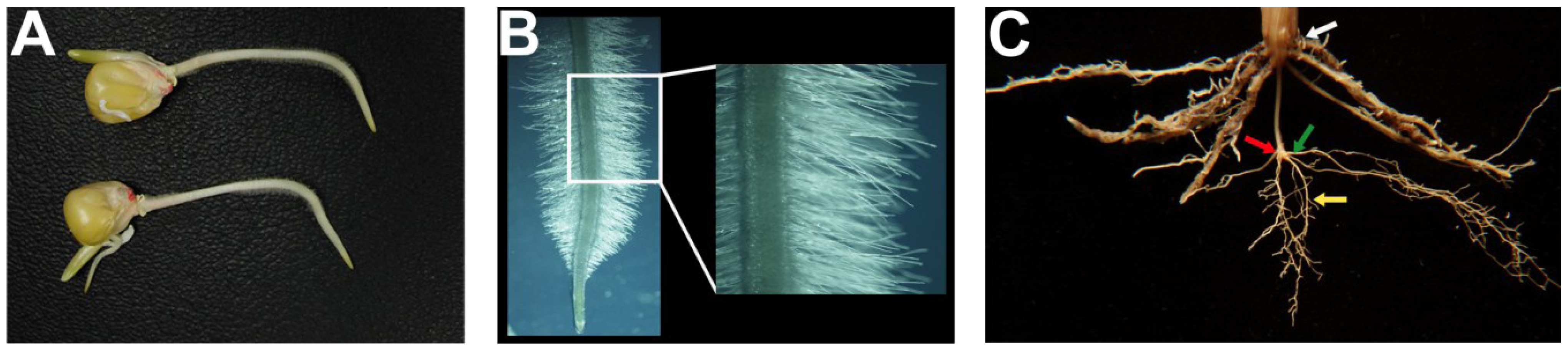

2.2. Root Hairs
2.3. Root Branching
3. Overview of Technologies Available for Phenotyping RSA Traits
| Approach | Growth Conditions | Advantages | Disadvantages |
|---|---|---|---|
| Laboratory methods | Highly controlled |
|
|
| Greenhouse methods | Moderately controlled |
|
|
| Field methods | Minimally controlled |
|
|
| Plant Cultivation System | Growth Media | Description | References |
|---|---|---|---|
| 1. Growth and luminescence observatory for roots (GLO-Roots) | Soil (lab) | This method combines custom-made growth vessels and new image analysis algorithms to non-destructively monitor RSA development over space (2-D) and time. The technique allows information on soil properties (e.g., moisture) to be integrated with root growth data. The system makes use of luminescence imaging of roots expressing plant codon-optimize luciferase. | [96] |
| 2. X-Ray computed tomography | Soil (lab and greenhouse) | Non-destructively visualizes opaque root structures by measuring the attenuation of ionizing radiation as it passes through the root. A series of projections are acquired and combined to reconstruct a 3D image of the root system. | [97,98] |
| 3. Rhizophonics | Liquid media (lab) | Combines hydroponics and rhizotrons. System is made of a nylon fabric supported by an aluminum frame. The set-up is immersed in a tank filled with liquid media. Allows non-destructive, 2-D imaging of root architecture while simultaneously sampling shoots. | [99] |
| 4. Clear pot method | Soil (greenhouse) | Uses transparent pots filled with soil or other potting media. Seeds are planted close to the pot wall to enable high- throughput imaging of roots along the clear pot wall. To prevent light exposure, the clear pot is placed in black pots while roots are developing. | [100] |
| 5. Rhizoslides | Paper-based (lab, greenhouse) | The set-up consists of a plexiglass sheet covered with moistened germination paper. Seeds are planted on the slit of the plexiglass. The system allows separation of crown roots from embryonic roots. | [101] |
| 6. Shovelomics | Soil (field-based) | Involves manual excavation of plants and separating roots from the shoots. Washed roots are then placed on a phenotyping board for root trait quantification. New algorithms allow extraction of several root traits in a high throughput manner. | [94,95] |
| 7. Soil coring | Soil (field-based) | Uses a tractor-mounted, hydraulic soil corer to drive steel alloy sampling tubes into the soil. When combined with novel planting configurations (e.g., hill plots), this method allows for phenotyping deep rooted crop varieties. | [102] |
| 8. Rhizolysimeters | Soil (field-based) | Elaborate facility consisting of an underground corridor and concrete silos and pipes to house soil-containing soil cores for direct root observation. | [103] |
| 9. Minirhizotrons | Soil (field-based) | A transparent observation tube permanently inserted in the soil. Images of roots growing along the minirhizotron wall at particular locations in the soil profile can be captured over time. | [104,105] |
4. Strategies for Root Phenotyping and Their Utilization in Breeding Programs
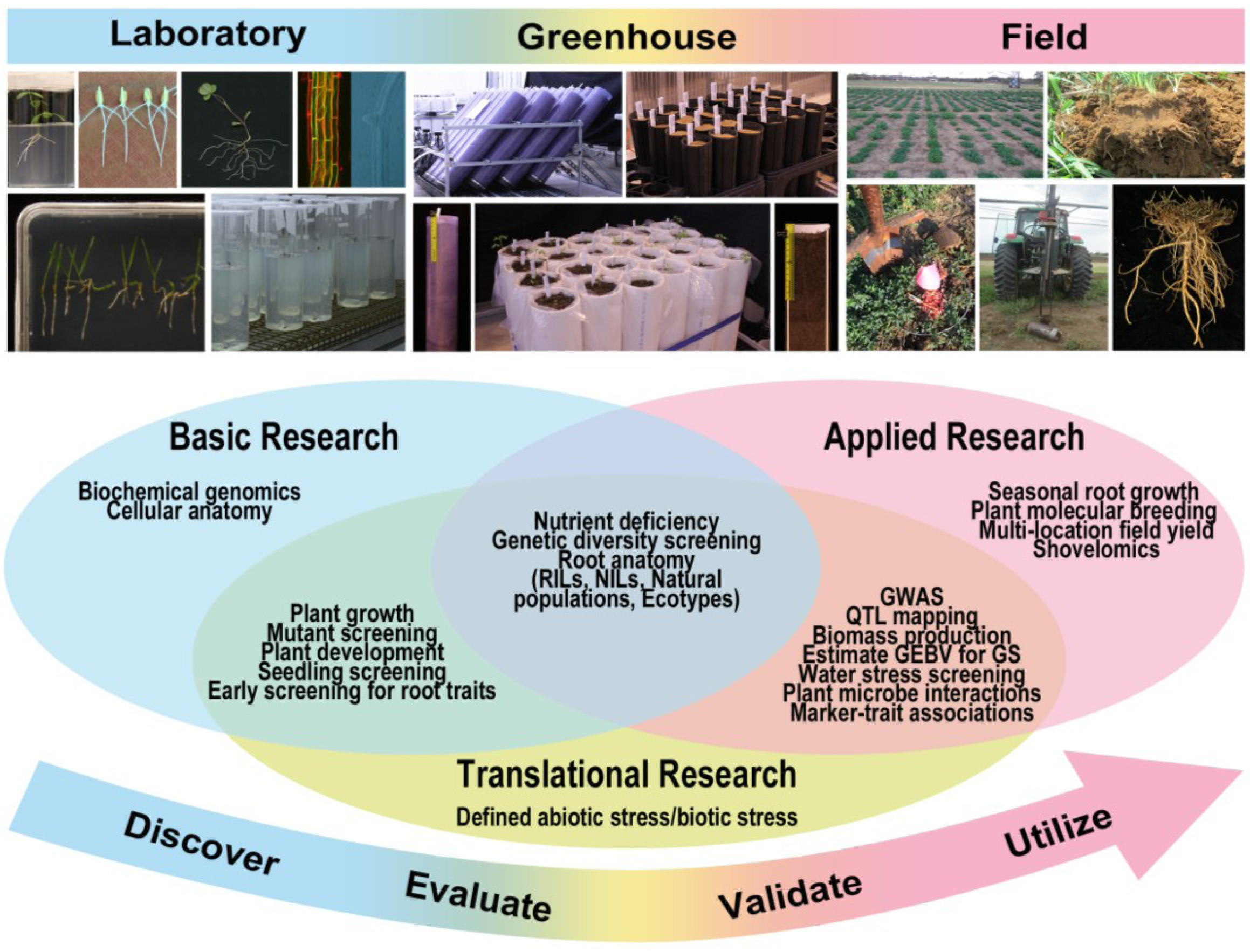
5. Perspectives and Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kano, M.; Inukai, Y.; Kitano, H.; Yamauchi, A. Root plasticity as the key root trait for adaptation to various intensities of drought stress in rice. Plant Soil 2011, 342, 117–128. [Google Scholar] [CrossRef]
- Grossman, J.D.; Rice, K.J. Evolution of root plasticity responses to variation in soil nutrient distribution and concentration. Evolut. Appl. 2012, 5, 850–857. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J. Root architecture and plant productivity. Plant Physiol. 1995, 109, 7–13. [Google Scholar] [PubMed]
- Bao, Y.; Aggarwal, P.; Robbins, N.E.; Sturrock, C.J.; Thompson, M.C.; Tan, H.Q.; Tham, C.; Duan, L.; Rodriguez, P.L.; Vernoux, T. Plant roots use a patterning mechanism to position lateral root branches toward available water. Proc. Natl. Acad. Sci. USA 2014, 111, 9319–9324. [Google Scholar] [CrossRef] [PubMed]
- Robbins, N.E.; Dinneny, J.R. The divining root: Moisture-driven responses of roots at the micro-and macro-scale. J. Exp. Bot. 2015. [Google Scholar] [CrossRef]
- Kell, D.B. Breeding crop plants with deep roots: Their role in sustainable carbon, nutrient and water sequestration. Ann. Bot. 2011. [Google Scholar] [CrossRef] [PubMed]
- Hufnagel, B.; de Sousa, S.M.; Assis, L.; Guimaraes, C.T.; Leiser, W.; Azevedo, G.C.; Negri, B.; Larson, B.G.; Shaff, J.E.; Pastina, M.M.; et al. Duplicate and conquer: Multiple homologs of PHOSPHORUS-STARVATION TOLERANCE1 enhance phosphorus acquisition and sorghum performance on low-phosphorus soils. Plant Physiol. 2014, 166, 659–677. [Google Scholar] [CrossRef] [PubMed]
- Uga, Y.; Sugimoto, K.; Ogawa, S.; Rane, J.; Ishitani, M.; Hara, N.; Kitomi, Y.; Inukai, Y.; Ono, K.; Kanno, N.; et al. Control of root system architecture by DEEPER ROOTING 1 increases rice yield under drought conditions. Nat. Genet. 2013, 45, 1097–1102. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, S.; Mohan, A.; Gill, K.S.; Prasad, P.V. Variability of root traits in spring wheat germplasm. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Meister, R.; Rajani, M.; Ruzicka, D.; Schachtman, D.P. Challenges of modifying root traits in crops for agriculture. Trends Plant Sci. 2014, 19, 779–788. [Google Scholar] [CrossRef] [PubMed]
- Furbank, R.T.; Tester, M. Phenomics-technologies to relieve the phenotyping bottleneck. Trends Plant Sci. 2011, 16, 635–644. [Google Scholar] [CrossRef] [PubMed]
- Rascher, U.; Blossfeld, S.; Fiorani, F.; Jahnke, S.; Jansen, M.; Kuhn, A.J.; Matsubara, S.; Märtin, L.L.; Merchant, A.; Metzner, R. Non-invasive approaches for phenotyping of enhanced performance traits in bean. Funct. Plant Biol. 2011, 38, 968–983. [Google Scholar] [CrossRef]
- York, L.M.; Nord, E.A.; Lynch, J.P. Integration of root phenes for soil resource acquisition. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Fiorani, F.; Schurr, U. Future scenarios for plant phenotyping. Annu. Rev. Plant Biol. 2013, 64, 267–291. [Google Scholar] [CrossRef] [PubMed]
- Canè, M.A.; Maccaferri, M.; Nazemi, G.; Salvi, S.; Francia, R.; Colalongo, C.; Tuberosa, R. Association mapping for root architectural traits in durum wheat seedlings as related to agronomic performance. Mol. Breed. 2014, 34, 1629–1645. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Kaeppler, S.M.; Lynch, J.P. Mapping of QTL controlling root hair length in maize (Zea mays L.) under phosphorus deficiency. Plant Soil 2005, 270, 299–310. [Google Scholar] [CrossRef]
- Kamoshita, A.; Zhang, J.; Siopongco, J.; Sarkarung, S.; Nguyen, H.T.; Wade, L.J. Effects of phenotyping environment on identification of quantitative trait loci for rice root morphology under anaerobic conditions. Crop Sci. 2002, 42, 255–265. [Google Scholar] [CrossRef] [PubMed]
- Doran, J.W.; Zeiss, M.R. Soil health and sustainability: Managing the biotic component of soil quality. Appl. Soil Ecol. 2000, 15, 3–11. [Google Scholar] [CrossRef]
- Saikia, P.; Bhattacharya, S.S.; Baruah, K.K. Organic substitution in fertilizer schedule: Impacts on soil health, photosynthetic efficiency, yield and assimilation in wheat grown in alluvial soil. Agric. Ecosyst. Environ. 2015, 203, 102–109. [Google Scholar] [CrossRef]
- Dias, T.; Dukes, A.; Antunes, P.M. Accounting for soil biotic effects on soil health and crop productivity in the design of crop rotations. J. Sci. Food Agric. 2015, 95, 447–454. [Google Scholar] [CrossRef] [PubMed]
- Teng, Y.; Wang, X.; Li, L.; Li, Z.; Luo, Y. Rhizobia and their bio-partners as novel drivers for functional remediation in contaminated soils. Name Front. Plant Sci. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Subbarao, G.V.; Yoshihashi, T.; Worthington, M.; Nakahara, K.; Ando, Y.; Sahrawat, K.L.; Rao, I.M.; Lata, J.-C.; Kishii, M.; Braun, H.-J. Suppression of soil nitrification by plants. Plant Sci. 2015, 233, 155–164. [Google Scholar] [CrossRef] [PubMed]
- Glick, B.R. Phytoremediation: Synergistic use of plants and bacteria to clean up the environment. Biotechnol. Adv. 2003, 21, 383–393. [Google Scholar] [CrossRef]
- Zhu, J.; Ingram, P.A.; Benfey, P.N.; Elich, T. From lab to field, new approaches to phenotyping root system architecture. Curr. Opin. Plant Biol. 2011, 14, 310–317. [Google Scholar] [CrossRef] [PubMed]
- Forde, B.G. Nitrogen signalling pathways shaping root system architecture: An update. Curr. Opin. Plant Biol. 2014, 21, 30–36. [Google Scholar] [CrossRef] [PubMed]
- Giehl, R.F.; Gruber, B.D.; von Wirén, N. It’s time to make changes: Modulation of root system architecture by nutrient signals. J. Exp. Bot. 2014, 65, 769–778. [Google Scholar] [CrossRef] [PubMed]
- López-Bucio, J.; Cruz-Ramı́rez, A.; Herrera-Estrella, L. The role of nutrient availability in regulating root architecture. Curr. Opin. Plant Biol. 2003, 6, 280–287. [Google Scholar] [CrossRef]
- Gruber, B.D.; Giehl, R.F.; Friedel, S.; von Wirén, N. Plasticity of the Arabidopsis root system under nutrient deficiencies. Plant Physiol. 2013, 163, 161–179. [Google Scholar] [CrossRef] [PubMed]
- Kellermeier, F.; Armengaud, P.; Seditas, T.J.; Danku, J.; Salt, D.E.; Amtmann, A. Analysis of the root system architecture of Arabidopsis provides a quantitative readout of crosstalk between nutritional signals. Plant Cell Online 2014, 26, 1480–1496. [Google Scholar] [CrossRef] [PubMed]
- DoVale, J.; Fritsche-Neto, R. Root Phenomics. In Phenomics; Fritsche-Neto, R., Borém, A., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 49–66. [Google Scholar]
- Tuberosa, R. Phenotyping for drought tolerance of crops in the genomics era. Front. Physiol. 2012, 3. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P. Turner review no. 14. Roots of the second green revolution. Aust. J. Bot. 2007, 55, 493–512. [Google Scholar] [CrossRef]
- Forde, B.G. Is it good noise? The role of developmental instability in the shaping of a root system. J. Exp. Bot. 2009, 60, 3989–4002. [Google Scholar] [CrossRef] [PubMed]
- Wasson, A.; Richards, R.; Chatrath, R.; Misra, S.; Prasad, S.S.; Rebetzke, G.; Kirkegaard, J.; Christopher, J.; Watt, M. Traits and selection strategies to improve root systems and water uptake in water-limited wheat crops. J. Exp. Bot. 2012, 63, 3485–3498. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P. Steep, cheap and deep: An ideotype to optimize water and N acquisition by maize root systems. Ann. Bot. 2013, 112, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P.; Chimungu, J.G.; Brown, K.M. Root anatomical phenes associated with water acquisition from drying soil: Targets for crop improvement. J. Exp. Bot. 2014, 65, 6155–6166. [Google Scholar] [CrossRef] [PubMed]
- Comas, L.H.; Becker, S.R.; von Mark, V.C.; Byrne, P.F.; Dierig, D.A. Root traits contributing to plant productivity under drought. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Brown, L.; George, T.; Dupuy, L.; White, P. A conceptual model of root hair ideotypes for future agricultural environments: What combination of traits should be targeted to cope with limited P availability? Ann. Bot. 2013, 112, 317–330. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P.; Wojciechowski, T. Opportunities and challenges in the subsoil: Pathways to deeper rooted crops. J. Exp. Bot. 2015. [Google Scholar] [CrossRef]
- Haling, R.E.; Brown, L.K.; Bengough, A.G.; Young, I.M.; Hallett, P.D.; White, P.J.; George, T.S. Root hairs improve root penetration, root-soil contact, and phosphorus acquisition in soils of different strength. J. Exp. Bot. 2013, 64, 3711–3721. [Google Scholar] [CrossRef] [PubMed]
- Miguel, M.A.; Postma, J.A.; Lynch, J. Phene synergism between root hair length and basal root growth angle for phosphorus acquisition. Plant Physiol. 2015, 167, 1430–1439. [Google Scholar] [CrossRef] [PubMed]
- Bates, T.R.; Lynch, J.P. Root hairs confer a competitive advantage under low phosphorus availability. Plant Soil 2001, 236, 243–250. [Google Scholar] [CrossRef]
- Tanaka, N.; Kato, M.; Tomioka, R.; Kurata, R.; Fukao, Y.; Aoyama, T.; Maeshima, M. Characteristics of a root hair-less line of Arabidopsis thaliana under physiological stresses. J. Exp. Bot. 2014, 65, 1497–1512. [Google Scholar] [CrossRef] [PubMed]
- Delhaize, E.; Ma, J.F.; Ryan, P.R. Transcriptional regulation of aluminium tolerance genes. Trends Plant Sci. 2012, 17, 341–348. [Google Scholar] [CrossRef] [PubMed]
- Nambiar, E. Uptake of Zn65 from dry soil by plants. Plant Soil 1976, 44, 267–271. [Google Scholar] [CrossRef]
- Den Herder, G.; van Isterdael, G.; Beeckman, T.; de Smet, I. The roots of a new green revolution. Trends Plant Sci. 2010, 15, 600–607. [Google Scholar] [CrossRef] [PubMed]
- Postma, J.A.; Dathe, A.; Lynch, J.P. The optimal lateral root branching density for maize depends on nitrogen and phosphorus availability. Plant Physiol. 2014, 166, 590–602. [Google Scholar] [CrossRef] [PubMed]
- Zhan, A.; Lynch, J.P. Reduced frequency of lateral root branching improves N capture from low-N soils in maize. J. Exp. Bot. 2015. [Google Scholar] [CrossRef] [PubMed]
- Walch-Liu, P.; Ivanov, I.I.; Filleur, S.; Gan, Y.; Remans, T.; Forde, B.G. Nitrogen regulation of root branching. Ann. Bot. 2006, 97, 875–881. [Google Scholar] [CrossRef] [PubMed]
- Rewald, B.; Ephrath, J.E.; Rachmilevitch, S. A root is a root is a root? Water uptake rates of Citrus root orders. Plant Cell Environ. 2011, 34, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P.; Brown, K.M. Topsoil foraging—An architectural adaptation of plants to low phosphorus availability. Plant Soil 2001, 237, 225–237. [Google Scholar] [CrossRef]
- Zhu, J.; Kaeppler, S.M.; Lynch, J.P. Topsoil foraging and phosphorus acquisition efficiency in maize (Zea mays). Funct. Plant Biol. 2005, 32, 749–762. [Google Scholar] [CrossRef]
- Niu, Y.; Chai, R.; Dong, H.; Wang, H.; Tang, C.; Zhang, Y. Effect of elevated CO2 on phosphorus nutrition of phosphate-deficient Arabidopsis thaliana (L.) Heynh under different nitrogen forms. J. Exp. Bot. 2013, 64, 355–367. [Google Scholar] [CrossRef] [PubMed]
- York, L.M.; Galindo-Castaneda, T.; Schussler, J.R.; Lynch, J.P. Evolution of US maize (Zea mays L.) root architectural and anatomical phenes over the past 100 years corresponds to increased tolerance of nitrogen stress. J. Exp. Bot. 2015. [Google Scholar] [CrossRef]
- Neumann, G.; Martinoia, E. Cluster roots—An underground adaptation for survival in extreme environments. Trends Plant Sci. 2002, 7, 162–167. [Google Scholar] [CrossRef]
- Lambers, H.; Finnegan, P.M.; Laliberté, E.; Pearse, S.J.; Ryan, M.H.; Shane, M.W.; Veneklaas, E.J. Phosphorus nutrition of Proteaceae in severely phosphorus-impoverished soils: Are there lessons to be learned for future crops? Plant Physiol. 2011, 156, 1058–1066. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P. Root phenes that reduce the metabolic costs of soil exploration: Opportunities for 21st century agriculture. Plant Cell Environ. 2014. [Google Scholar] [CrossRef] [PubMed]
- Saengwilai, P.; Tian, X.; Lynch, J.P. Low crown root number enhances nitrogen acquisition from low-nitrogen soils in maize. Plant Physiol. 2014, 166, 581–589. [Google Scholar] [CrossRef] [PubMed]
- Drew, M.C.; He, C.-J.; Morgan, P.W. Decreased ethylene biosynthesis, and induction of aerenchyma, by nitrogen-or phosphate-starvation in adventitious roots of Zea mays L. Plant Physiol. 1989, 91, 266–271. [Google Scholar] [CrossRef] [PubMed]
- Postma, J.A.; Lynch, J.P. Root cortical aerenchyma enhances the growth of maize on soils with suboptimal availability of nitrogen, phosphorus, and potassium. Plant Physiol. 2011, 156, 1190–1201. [Google Scholar] [CrossRef] [PubMed]
- Chimungu, J.G.; Brown, K.M.; Lynch, J.P. Reduced root cortical cell file number improves drought tolerance in maize. Plant Physiol. 2014, 166, 1943–1955. [Google Scholar] [CrossRef] [PubMed]
- Chimungu, J.G.; Brown, K.M.; Lynch, J.P. Large root cortical cell size improves drought tolerance in maize. Plant Physiol. 2014, 166, 2166–2178. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.L.; Bouma, T.J.; Lynch, J.P.; Eissenstat, D.M. Effects of phosphorus availability and vesicular-arbuscular mycorrhizas on the carbon budget of common bean (Phaseolus vulgaris). New Phytol. 1998, 139, 647–656. [Google Scholar] [CrossRef]
- Nielsen, K.L.; Eshel, A.; Lynch, J.P. The effect of phosphorus availability on the carbon economy of contrasting common bean (Phaseolus vulgaris L.) genotypes. J. Exp. Bot. 2001, 52, 329–339. [Google Scholar] [CrossRef] [PubMed]
- Wishart, J.; George, T.S.; Brown, L.K.; White, P.J.; Ramsay, G.; Jones, H.; Gregory, P.J. Field phenotyping of potato to assess root and shoot characteristics associated with drought tolerance. Plant Soil 2014, 378, 351–363. [Google Scholar] [CrossRef]
- Blancaflor, E.B. Regulation of plant gravity sensing and signaling by the actin cytoskeleton. Am. J. Bot. 2013, 100, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Sato, E.M.; Hijazi, H.; Bennett, M.J.; Vissenberg, K.; Swarup, R. New insights into root gravitropic signalling. J. Exp. Bot. 2014. [Google Scholar] [CrossRef] [PubMed]
- Shin, H.; Shin, H.S.; Guo, Z.; Blancaflor, E.B.; Masson, P.H.; Chen, R. Complex regulation of Arabidopsis AGR1/PIN2-mediated root gravitropic response and basipetal auxin transport by cantharidin-sensitive protein phosphatases. Plant J. 2005, 42, 188–200. [Google Scholar] [CrossRef] [PubMed]
- Passioura, J. Phenotyping for drought tolerance in grain crops: When is it useful to breeders? Funct. Plant Biol. 2012, 39, 851–859. [Google Scholar] [CrossRef]
- Bolaños, J.; Edmeades, G.; Martinez, L. Eight cycles of selection for drought tolerance in lowland tropical maize. III. Responses in drought-adaptive physiological and morphological traits. Field Crops Res. 1993, 31, 269–286. [Google Scholar] [CrossRef]
- Mendrinna, A.; Persson, S. Root hair growth: It’s a one way street. F1000prime Rep. 2015, 7, 23. [Google Scholar] [CrossRef] [PubMed]
- Velasquez, S.M.; Ricardi, M.M.; Dorosz, J.G.; Fernandez, P.V.; Nadra, A.D.; Pol-Fachin, L.; Egelund, J.; Gille, S.; Harholt, J.; Ciancia, M. O-glycosylated cell wall proteins are essential in root hair growth. Science 2011, 332, 1401–1403. [Google Scholar] [CrossRef] [PubMed]
- Yoo, C.M.; Quan, L.; Cannon, A.E.; Wen, J.; Blancaflor, E.B. AGD1, a class 1 ARF-GAP, acts in common signaling pathways with phosphoinositide metabolism and the actin cytoskeleton in controlling Arabidopsis root hair polarity. Plant J. 2012, 69, 1064–1076. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Li, J.; Song, W.; Tao, J.; Huang, S.; Chen, S.; Hou, M.; Xu, G.; Zhang, Y. Nitric oxide generated by nitrate reductase increases nitrogen uptake capacity by inducing lateral root formation and inorganic nitrogen uptake under partial nitrate nutrition in rice. J. Exp. Bot. 2015, in press. [Google Scholar] [CrossRef]
- Tian, H.; de Smet, I.; Ding, Z. Shaping a root system: Regulating lateral vs. primary root growth. Trends Plant Sci. 2014, 19, 426–431. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Zhao, J.; Li, X.; Qin, L.; Yan, X.; Liao, H. A soybean β-expansin gene GmEXPB2 intrinsically involved in root system architecture responses to abiotic stresses. Plant J. 2011, 66, 541–552. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Xie, J.; Liao, H.; Wang, X. Overexpression of β-expansin gene GmEXPB2 improves phosphorus efficiency in soybean. Physiol. Plant. 2014, 150, 194–204. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.-Y.; Zhou, S.; Chen, Y.-H.; Kong, X.; Xu, Y.; Wang, W. The involvement of expansins in responses to phosphorus availability in wheat, and its potentials in improving phosphorus efficiency of plants. Plant Physiol. Biochem. 2014, 78, 53–62. [Google Scholar] [CrossRef] [PubMed]
- Gamuyao, R.; Chin, J.H.; Pariasca-Tanaka, J.; Pesaresi, P.; Catausan, S.; Dalid, C.; Slamet-Loedin, I.; Tecson-Mendoza, E.M.; Wissuwa, M.; Heuer, S. The protein kinase Pstol1 from traditional rice confers tolerance of phosphorus deficiency. Nature 2012, 488, 535–539. [Google Scholar] [CrossRef] [PubMed]
- Watt, M.; Schneebeli, K.; Dong, P.; Wilson, I.W. The shoot and root growth of Brachypodium and its potential as a model for wheat and other cereal crops. Funct. Plant Biol. 2009, 36, 960–969. [Google Scholar] [CrossRef]
- Richards, R.A. Genetic opportunities to improve cereal root systems for dryland agriculture. Plant Prod. Sci.-Tokyo 2008, 11, 12. [Google Scholar] [CrossRef]
- Burton, A.L.; Williams, M.; Lynch, J.P.; Brown, K.M. RootScan: Software for high-throughput analysis of root anatomical traits. Plant Soil 2012, 357, 189–203. [Google Scholar] [CrossRef]
- Pound, M.P.; French, A.P.; Atkinson, J.A.; Wells, D.M.; Bennett, M.J.; Pridmore, T. RootNav: Navigating images of complex root architectures. Plant Physiol. 2013, 162, 1802–1814. [Google Scholar] [CrossRef] [PubMed]
- Le Bot, J.; Serra, V.; Fabre, J.; Draye, X.; Adamowicz, S.; Pagès, L. DART: A software to analyse root system architecture and development from captured images. Plant Soil 2010, 326, 261–273. [Google Scholar] [CrossRef]
- Galkovskyi, T.; Mileyko, Y.; Bucksch, A.; Moore, B.; Symonova, O.; Price, C.A.; Topp, C.N.; Iyer-Pascuzzi, A.S.; Zurek, P.R.; Fang, S. GiA Roots: Software for the high throughput analysis of plant root system architecture. BMC Plant Biol. 2012, 12, 116. [Google Scholar] [CrossRef] [PubMed]
- Pierret, A.; Gonkhamdee, S.; Jourdan, C.; Maeght, J.-L. IJ_Rhizo: An open-source software to measure scanned images of root samples. Plant Soil 2013, 373, 531–539. [Google Scholar] [CrossRef]
- Leitner, D.; Felderer, B.; Vontobel, P.; Schnepf, A. Recovering root system traits using image analysis exemplified by two-dimensional neutron radiography images of lupine. Plant Physiol. 2014, 164, 24–35. [Google Scholar] [CrossRef]
- Clark, R.T.; Famoso, A.N.; Zhao, K.; Shaff, J.E.; Craft, E.J.; Bustamante, C.D.; Mccouch, S.R.; Aneshansley, D.J.; Kochian, L.V. High-throughput two-dimensional root system phenotyping platform facilitates genetic analysis of root growth and development. Plant Cell Environ. 2013, 36, 454–466. [Google Scholar] [CrossRef] [PubMed]
- Clark, R.T.; MacCurdy, R.B.; Jung, J.K.; Shaff, J.E.; McCouch, S.R.; Aneshansley, D.J.; Kochian, L.V. Three-dimensional root phenotyping with a novel imaging and software platform. Plant Physiol. 2011, 156, 455–465. [Google Scholar] [CrossRef] [PubMed]
- Mairhofer, S.; Zappala, S.; Tracy, S.R.; Sturrock, C.; Bennett, M.; Mooney, S.J.; Pridmore, T. RooTrak: Automated recovery of three-dimensional plant root architecture in soil from x-ray microcomputed tomography images using visual tracking. Plant Physiol. 2012, 158, 561–569. [Google Scholar] [CrossRef] [PubMed]
- Lobet, G.; Pound, M.P.; Diener, J.; Pradal, C.; Draye, X.; Godin, C.; Javaux, M.; Leitner, D.; Meunier, F.; Nacry, P. Root System Markup Language: Toward a unified root architecture description language. Plant Physiol. 2015, 167, 617–627. [Google Scholar] [CrossRef] [PubMed]
- Lobet, G.; Draye, X. Novel scanning procedure enabling the vectorization of entire rhizotron-grown root systems. Plant Methods 2013, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, R.; Al-Shugeairy, Z.; Al-Ogaidi, F.; Munasinghe, M.; Radermacher, M.; Vandenhirtz, J.; Price, A. Comparing simple root phenotyping methods on a core set of rice genotypes. Plant Biol. 2014, 16, 632–642. [Google Scholar] [CrossRef] [PubMed]
- Bucksch, A.; Burridge, J.; York, L.M.; Das, A.; Nord, E.; Weitz, J.S.; Lynch, J.P. Image-based high-throughput field phenotyping of crop roots. Plant Physiol. 2014, 166, 470–486. [Google Scholar] [CrossRef] [PubMed]
- Trachsel, S.; Kaeppler, S.M.; Brown, K.M.; Lynch, J.P. Shovelomics: High throughput phenotyping of maize (Zea mays L.) root architecture in the field. Plant Soil 2011, 341, 75–87. [Google Scholar] [CrossRef]
- Rellán-Álvarez, R.; Lobet, G.; Lindner, H.; Pradier, P.-L.M.; Yee, M.-C.; Sebastian, J.; Geng, Y.; Trontin, C.; LaRue, T.; Lavelle, A.S. Multidimensional mapping of root responses to soil environmental cues using a luminescence-based imaging system. bioRxiv 2015, 016931. [Google Scholar] [CrossRef]
- Mairhofer, S.; Zappala, S.; Tracy, S.; Sturrock, C.; Bennett, M.J.; Mooney, S.J.; Pridmore, T.P. Recovering complete plant root system architectures from soil via X-ray μ-Computed Tomography. Plant Methods 2013, 9, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Mooney, S.; Pridmore, T.; Helliwell, J.; Bennett, M. Developing X-ray computed tomography to non-invasively image 3-D root systems architecture in soil. Plant Soil 2012, 352, 1–22. [Google Scholar] [CrossRef]
- Mathieu, L.; Lobet, G.; Tocquin, P.; Perilleux, C. “Rhizoponics”: A novel hydroponic rhizotron for root system analyses on mature Arabidopsis thaliana plants. Plant Methods 2015, 11, 3. [Google Scholar] [CrossRef] [PubMed]
- Richard, C.A.; Hickey, L.T.; Fletcher, S.; Jennings, R.; Chenu, K.; Christopher, J.T. High-throughput phenotyping of seminal root traits in wheat. Plant Methods 2015, 11, 13. [Google Scholar] [CrossRef] [PubMed]
- Le Marie, C.; Kirchgessner, N.; Marschall, D.; Walter, A.; Hund, A. Rhizoslides: Paper-based growth system for non-destructive, high throughput phenotyping of root development by means of image analysis. Plant Methods 2014, 10, 13. [Google Scholar] [CrossRef] [PubMed]
- Wasson, A.; Rebetzke, G.; Kirkegaard, J.; Christopher, J.; Richards, R.; Watt, M. Soil coring at multiple field environments can directly quantify variation in deep root traits to select wheat genotypes for breeding. J. Exp. Bot. 2014, 65, 6231–6249. [Google Scholar] [CrossRef] [PubMed]
- Eberbach, P.L.; Hoffmann, J.; Moroni, S.J.; Wade, L.J.; Weston, L.A. Rhizo-lysimetry: Facilities for the simultaneous study of root behaviour and resource use by agricultural crop and pasture systems. Plant Methods 2013, 9, 3. [Google Scholar] [CrossRef] [PubMed]
- Maeght, J.L.; Rewald, B.; Pierret, A. How to study deep roots—And why it matters. Front. Plant Sci. 2013, 4, 299. [Google Scholar] [CrossRef]
- Iversen, C.M.; Murphy, M.T.; Allen, M.F.; Childs, J.; Eissenstat, D.M.; Lilleskov, E.A.; Sarjala, T.M.; Sloan, V.L.; Sullivan, P.F. Advancing the use of minirhizotrons in wetlands. Plant Soil 2011, 352, 23–39. [Google Scholar] [CrossRef]
- Lynch, J.P.; Brown, K.M. New roots for agriculture: Exploiting the root phenome. Philos. Trans. R. Soc. B Biol. Sci. 2012, 367, 1598–1604. [Google Scholar] [CrossRef] [PubMed]
- Steele, K.; Virk, D.; Kumar, R.; Prasad, S.; Witcombe, J. Field evaluation of upland rice lines selected for QTLs controlling root traits. Field Crops Res. 2007, 101, 180–186. [Google Scholar] [CrossRef]
- Bernier, J.; Serraj, R.; Kumar, A.; Venuprasad, R.; Impa, S.; Oane, R.; Spaner, D.; Atlin, G. The large-effect drought-resistance QTL qtl12. 1 increases water uptake in upland rice. Field Crops Res. 2009, 110, 139–146. [Google Scholar] [CrossRef]
- Lamb, J.; Samac, D.; Barnes, D.; Henjum, K. Increased herbage yield in alfalfa associated with selection for fibrous and lateral roots. Crop Sci. 2000, 40, 693–699. [Google Scholar] [CrossRef]
- Lamb, J.; Barnes, D.; Henjum, K. Gain from two cycles of divergent selection for root morphology in alfalfa. Crop Sci. 1999, 39, 1026–1035. [Google Scholar] [CrossRef]
- Johnson, L.; Marquez-Ortiz, J.; Lamb, J.; Barnes, D. Root morphology of alfalfa plant introductions and cultivars. Crop Sci. 1998, 38, 497–502. [Google Scholar] [CrossRef]
- Liu, L.; Zhang, H.; Ju, C.; Xiong, Y.; Bian, J.; Zhao, B.; Yang, J. Changes in Grain Yield and Root Morphology and Physiology of Mid-Season Rice in the Yangtze River Basin of China During the Last 60 Years. J. Agric. Sci. 2014, 6. [Google Scholar] [CrossRef]
- Lamb, J.; Johnson, L.; Barnes, D.; Marquez-Ortiz, J. A method to characterize root morphology traits in alfalfa. Can. J. Plant Sci. 2000, 80, 97–104. [Google Scholar] [CrossRef]
- White, P.J.; George, T.S.; Dupuy, L.X.; Karley, A.J.; Valentine, T.A.; Wiesel, L.; Wishart, J. Root traits for infertile soils. Front. Plant Sci. 2013, 4, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Wishart, J.; George, T.S.; Brown, L.K.; Ramsay, G.; Bradshaw, J.E.; White, P.J.; Gregory, P.J. Measuring variation in potato roots in both field and glasshouse: The search for useful yield predictors and a simple screen for root traits. Plant Soil 2013, 368, 231–249. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Paez-Garcia, A.; Motes, C.M.; Scheible, W.-R.; Chen, R.; Blancaflor, E.B.; Monteros, M.J. Root Traits and Phenotyping Strategies for Plant Improvement. Plants 2015, 4, 334-355. https://doi.org/10.3390/plants4020334
Paez-Garcia A, Motes CM, Scheible W-R, Chen R, Blancaflor EB, Monteros MJ. Root Traits and Phenotyping Strategies for Plant Improvement. Plants. 2015; 4(2):334-355. https://doi.org/10.3390/plants4020334
Chicago/Turabian StylePaez-Garcia, Ana, Christy M. Motes, Wolf-Rüdiger Scheible, Rujin Chen, Elison B. Blancaflor, and Maria J. Monteros. 2015. "Root Traits and Phenotyping Strategies for Plant Improvement" Plants 4, no. 2: 334-355. https://doi.org/10.3390/plants4020334
APA StylePaez-Garcia, A., Motes, C. M., Scheible, W.-R., Chen, R., Blancaflor, E. B., & Monteros, M. J. (2015). Root Traits and Phenotyping Strategies for Plant Improvement. Plants, 4(2), 334-355. https://doi.org/10.3390/plants4020334






