Identification of the Abundant Hydroxyproline-Rich Glycoproteins in the Root Walls of Wild-Type Arabidopsis, an ext3 Mutant Line, and Its Phenotypic Revertant
Abstract
:1. Introduction
2. Results
2.1. Arabidopsis Root Cell Wall Hyp/Protein Content
2.2. HF Deglycosylation of Root Cell Walls
| Amino Acid | WT | WT | WT | WT | ANP4 | ANP4 | ANP4 | ANP4 | ANP10 | ANP10 | ANP10 | ANP10 | MT | MT | MT |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CW | HFI | TI HFI | PI HFI | CW | HFI | TI HFI | PI HFI | CW | HFI | TI HFI | PI HFI | CW | HFI | TI HFI | |
| Asp | 6 | 1 | 2 | 1 | 6 | 6 | 4 | 4 | 6 | 4 | 1 | 1 | 7 | 5 | 5 |
| Glu | 6 | 3 | 2 | 2 | 5 | 4 | 3 | 3 | 6 | 3 | 2 | 2 | 7 | 2 | 4 |
| Hyp | 20 | 25 | 27 | 30 | 17 | 22 | 20 | 22 | 18 | 29 | 22 | 28 | 17 | 15 | 19 |
| Ser | 10 | 14 | 17 | 18 | 11 | 9 | 13 | 18 | 11 | 8 | 15 | 19 | 10 | 2 | 7 |
| Gly | 11 | 11 | 12 | 13 | 14 | 12 | 17 | 23 | 12 | 10 | 12 | 16 | 12 | 18 | 20 |
| His | 2 | 2 | 3 | 2 | 2 | 3 | 1 | 2 | 2 | 4 | 2 | 2 | 2 | 3 | 4 |
| Arg | 2 | 2 | 2 | 2 | 2 | 3 | 2 | 1 | 2 | 3 | 2 | 2 | 3 | 5 | 2 |
| Thr | 3 | 5 | 4 | 3 | 3 | 5 | 3 | 3 | 3 | 4 | 5 | 4 | 3 | 5 | 4 |
| Ala | 6 | 6 | 5 | 5 | 7 | 9 | 9 | 6 | 7 | 9 | 6 | 3 | 7 | 12 | 9 |
| Pro | 7 | 4 | 4 | 4 | 7 | 5 | 4 | 6 | 6 | 5 | 5 | 4 | 6 | 7 | 5 |
| Tyr | 4 | 2 | 2 | 2 | 3 | 2 | 3 | 2 | 3 | 2 | 1 | 2 | 3 | 2 | 1 |
| Val | 7 | 6 | 4 | 4 | 7 | 6 | 3 | 2 | 7 | 6 | 7 | 4 | 7 | 8 | 7 |
| Ile | 3 | 3 | 2 | 1 | 3 | 2 | 2 | 0 | 4 | 2 | 4 | 2 | 3 | 4 | 3 |
| Leu | 5 | 5 | 4 | 3 | 4 | 5 | 5 | 2 | 4 | 4 | 6 | 4 | 5 | 6 | 5 |
| Phe | 4 | 3 | 2 | 2 | 4 | 2 | 3 | 0 | 4 | 2 | 3 | 2 | 4 | 1 | 1 |
| Lys | 4 | 8 | 8 | 8 | 5 | 5 | 8 | 6 | 5 | 5 | 7 | 5 | 4 | 5 | 4 |
| Total | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 |
| % Hyp 1 | 1.5 | 6.9 | 5.2 | 4.5 | 1.1 | 3.9 | 2.1 | 1.1 | 1.3 | 6.3 | 4.3 | 4.1 | 1.1 | 2.5 | 3.8 |
| % Protein 1 | 7.3 | 25.5 | 17.7 | 13.6 | 6.4 | 16.1 | 7 | 4.4 | 6.8 | 20.5 | 17.4 | 13.1 | 6.2 | 13.7 | 14.6 |
| Hyp/ Protein 2 | 0.2 | 0.27 | 0.29 | 0.33 | 0.17 | 0.24 | 0.3 | 0.25 | 0.19 | 0.31 | 0.25 | 0.31 | 0.18 | 0.18 | 0.26 |
| DW (mg) 3 | 50 | 10 | 8 | 7 | 38 | 10 | 7 | 6 | 51 | 10 | 8 | 7 | 4.6 | 1.6 | 0.75 |
| % Hyp Removed 4 | 6.7 | 39.7 | 54.3 | 10.0 | 62.3 | 83.1 | 7.1 | 45.4 | 54.4 | 10.0 | 10.9 | ||||
| % Protein Removed 4 | 30.1 | 44.5 | 62.3 | 34.4 | 69.6 | 83.6 | 41.1 | 32.1 | 55.3 | 20.0 | 60.9 |
2.3. Release of Hyp-Containing Material by Protease Degradation of Root Cell Wall HFI
2.4. Peptide Fractionation from Trypsin Digested HFI of WT and ANP
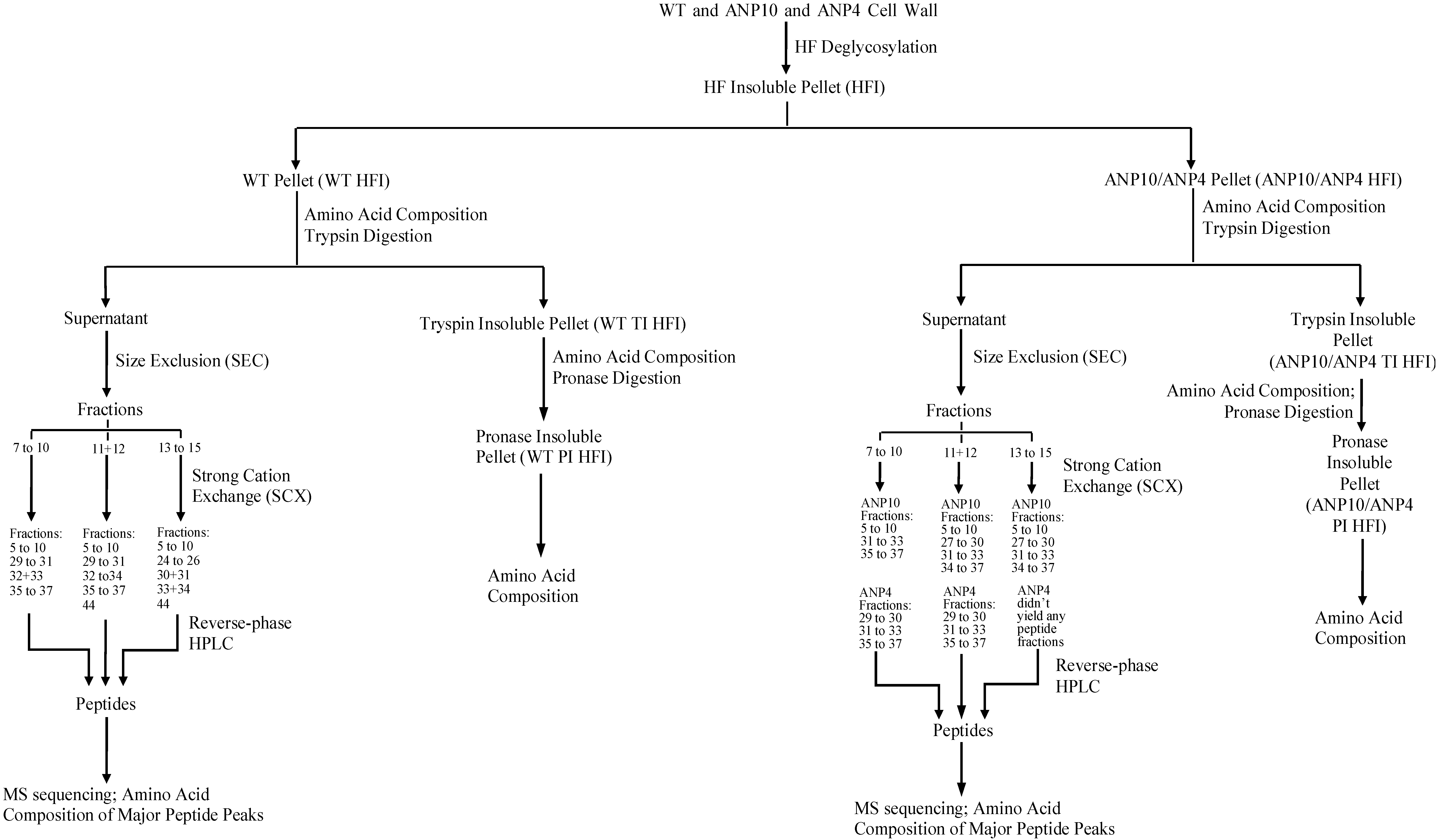
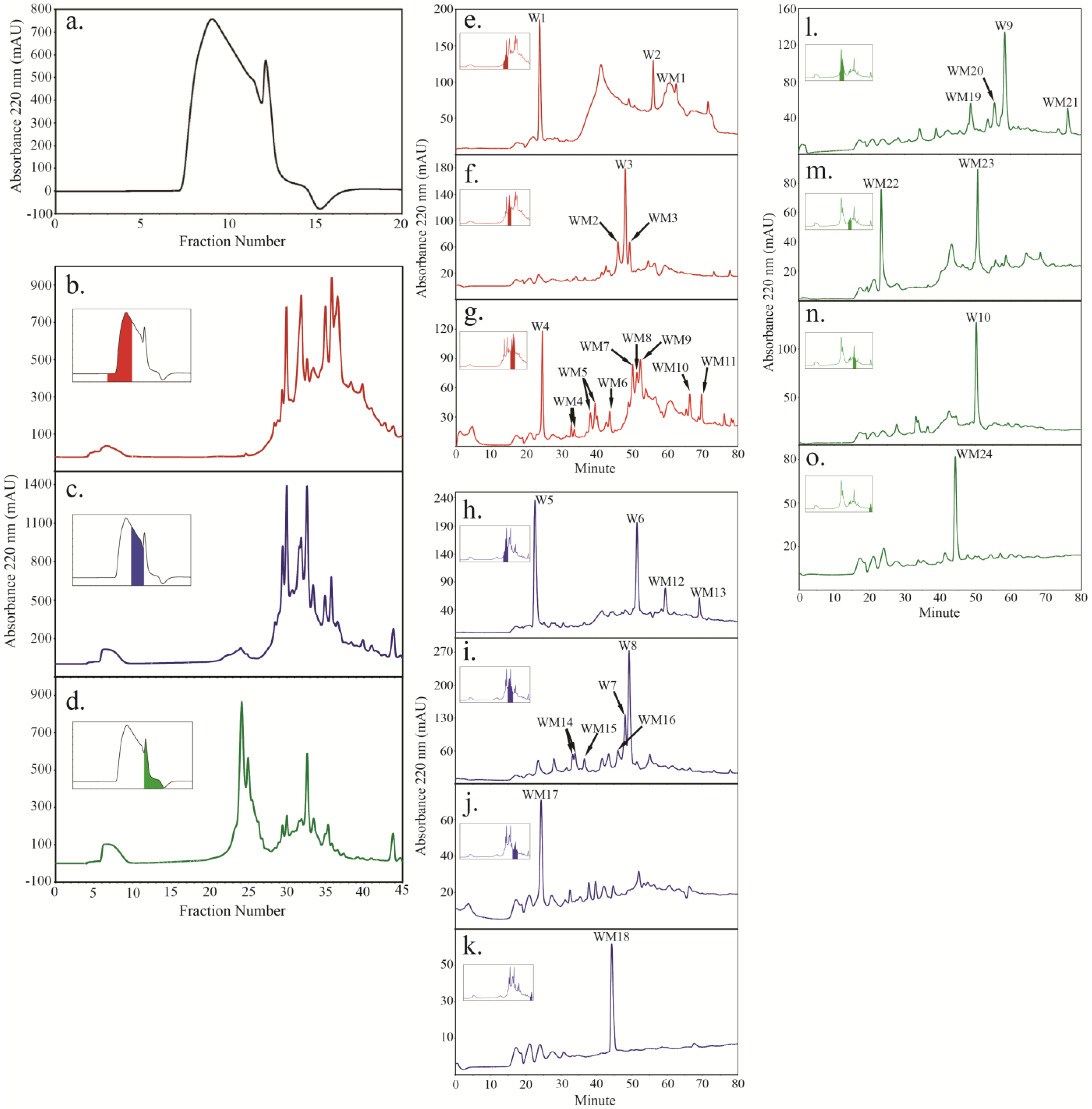
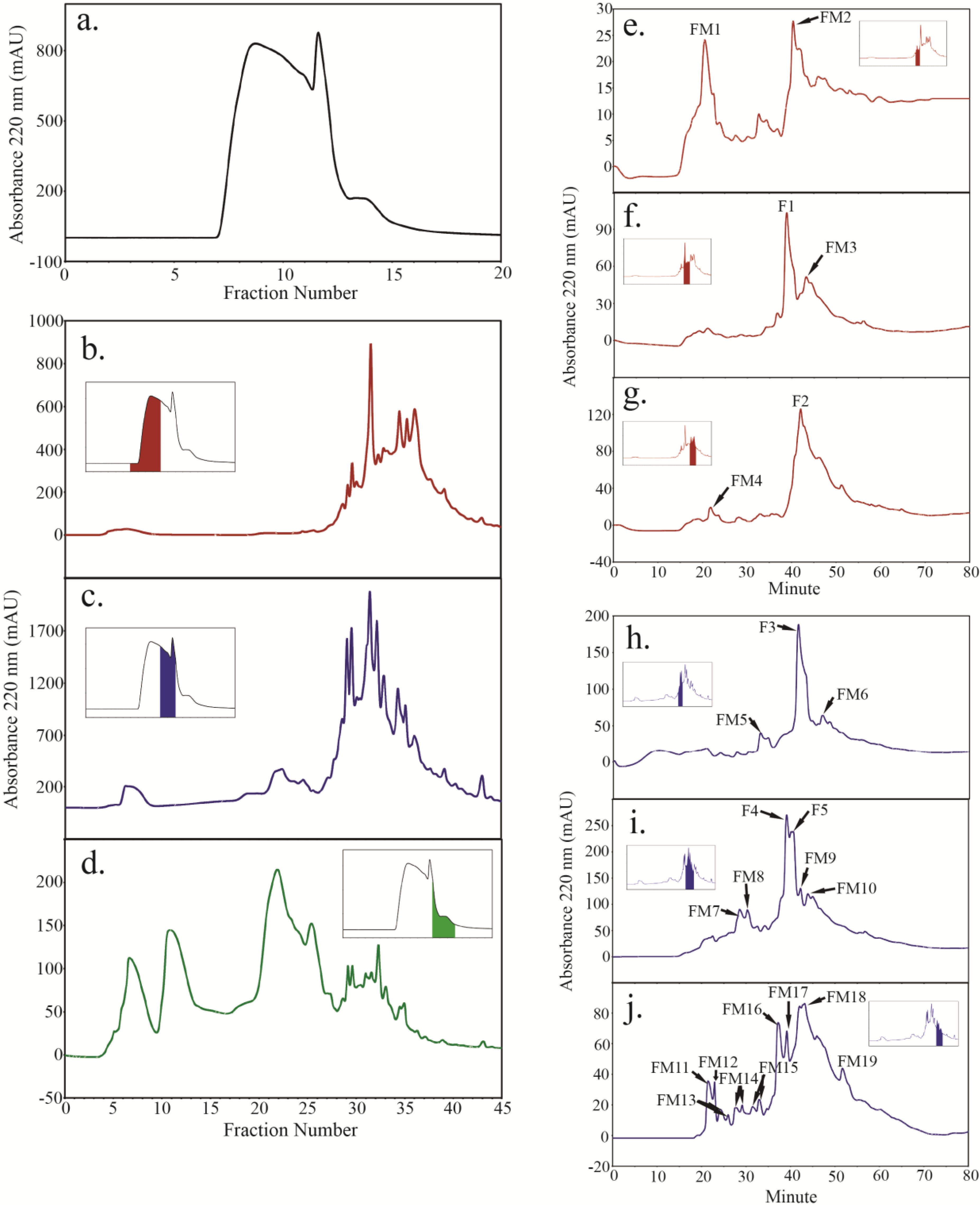
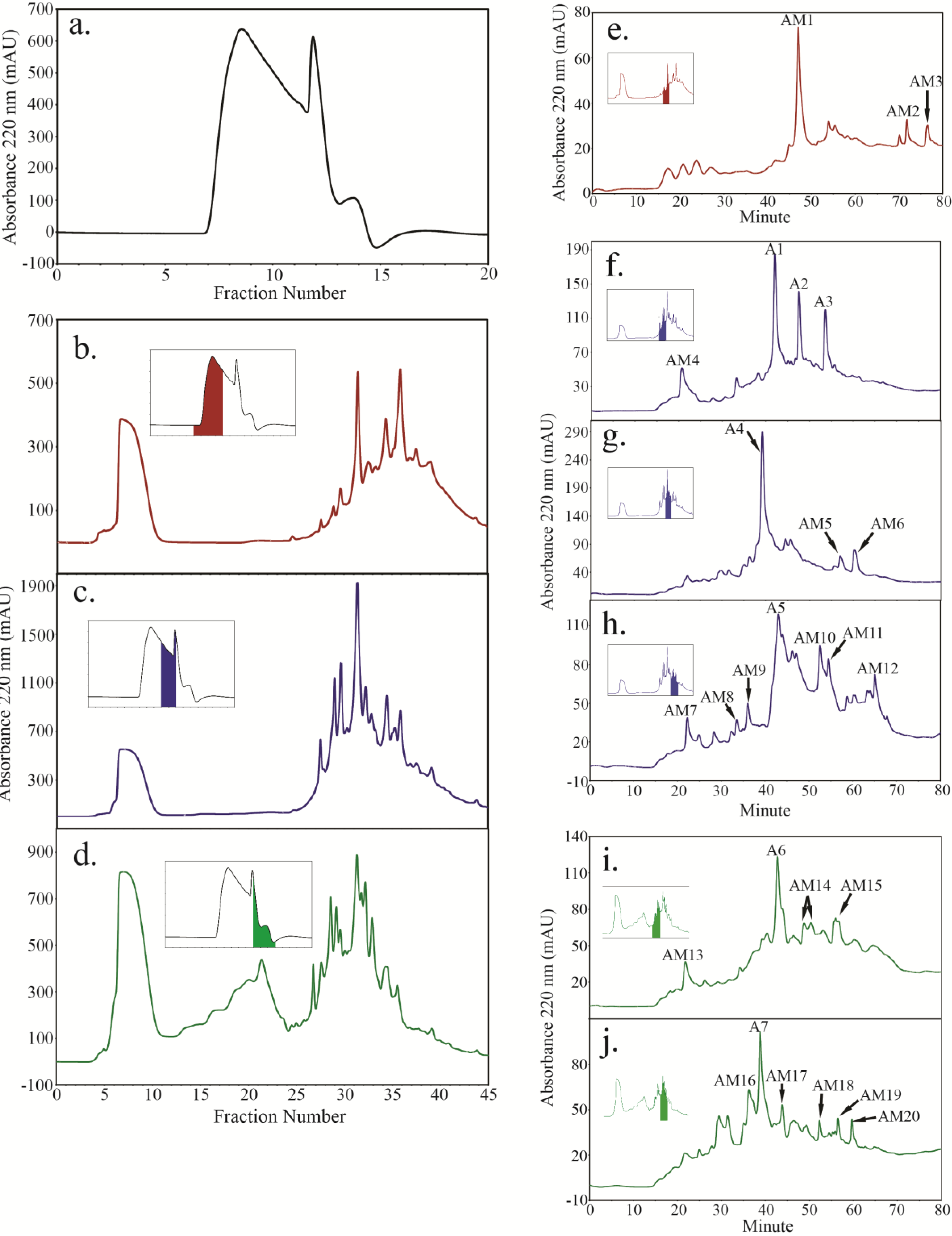
2.5. Peptide Sequencing and HRGP Identification
| Gene Accession # | Protein Identification (Plant Line of Origin) | Peptides Identified (Number of Occurrence in Protein Sequence; Peak of Peptide Identification) 1 | Peptide Mascot Ion Score 2 | Signal Peptide 3 | MM (kDa) 4 | pI 4 | |
|---|---|---|---|---|---|---|---|
| EXTs | |||||||
| AT1G76930 | EXT1 (WT, ANP4, ANP10) | YYSOOOVYK (3; W2, W6, W8, W10, WM3, WM13; FM16; A2) | 1–24 | 32.9 | 10.1 | ||
| HYSOOOVYK (9; WM6; A5), SOOOOVYK (1; FM8) | |||||||
| SOOOOVK (8; W1, W2, W5, WM22; AM3) | |||||||
| AT1G21310 | EXT3, RSH (WT) | HYSOOOVYHSOOOOK (13; WM6), HYTOOVK (1; WM4), | 1–28 | 49.2 | 10.3 | ||
| SOOOOVK (13; W1, W2, W5, WM22) | |||||||
| AT3G54580 | EXT17 (ANP4) | SOOOOYYSOSOK (15; FM8) | 54.9 | 1–22 | 105.7 | 9.5 | |
| AT2G43150 | EXT215 (WT) | SOOOOYYYHSOOOOVK (6; W8, W10) | 54.3 | 1–27 | 23.4 | 9.6 | |
| SOOOOVK (10; W1, W2, W5, WM22) | |||||||
| AT1G12040 | LRX1 (ANP4, ANP10) | LQGPLPSSVGNMK (1; F3) | 42.3 | 1–26 | 81 | 8.2 | |
| LLYELDLSNNR (1; AM4) | 44.5 | ||||||
| AT1G62440 | LRX2 (ANP4, ANP10) | SLEQLNVANNR (1; FM4, FM12, FM13) | 71.4 | 89.8 | 4.8 | ||
| LSGPLPSSIGNMK (1; A2) | 59.4 | ||||||
| AT4G13340 | LRX3 (ANP10) | FNEFEGTVPK (1; AM5, AM13, AM15, AM20) | 65.9 | 1–20 | 82.2 | 7 | |
| AT3G22800 | LRX6 (ANP4) | SLEQLNIAHNK (1; FM6) | 55.7 | 1–28 | 52.3 | 6.2 | |
| AT3G50580 | HAE2 (ANP10) | SOOOOTOK (1; AM3, AM12) | 41.7 | 1–24 | 27.2 | 9.1 | |
| AGPs | |||||||
| AT2G33790 | AGP30 (WT) | LPAYPOAK (1; W2), TLVAVR (1; WM3), VSSLHDGGK (1; WM4) | 1–25 | 25.8 | 10.6 | ||
| AT1G28290 | AGP31 (WT, ANP4, ANP10) | SLVAVR (1; W8, W10, WM3), SOVKPOVK (1; WM4, WM14), | 1–25 | 38.4 | 10.8 | ||
| NITAETTTDK (1; FM8), | |||||||
| AOVSOOAK(O/P)(O/P)VK(O/P)(O/P)VY(O/P)(O/P)TK7 | |||||||
| (1; W3, W4, W7, W8, W10, WM3, WM18, WM19, WM22; F1, F4, FM4, FM16; A5, AM1), LFGGDVGAELKPEK | |||||||
| (1; FM7, FM8, FM18; AM10, AM11, AM18) | |||||||
| AT2G04780 | FLA7 (WT) | FTDVSGTVR (1; W10, WM3) | 61.4 | 1–22 | 26.8 | 6.5 | |
| AT3G52370 | FLA15 (WT) | HHFNGEAQVK (1; WM24) | 42.3 | 1–20 | 48.1 | 6.7 | |
| AT2G35860 | FLA16 (ANP4) | EETOATEIKPAAOVVK (1; F1, FM4, FM13) | 55.4 | 1–23 | 49.1 | 6.8 | |
| PRPs | |||||||
| AT4G38770 | PRP4 (ANP4) | EVPO(P/O)VOVYK(P/O)(P/O)(P/O)K7 (1; F1), | 1–29 | 49.1 | 10.3 | ||
| IEHPO(P/O)VOVYKOO(P/O)K7 (1; F2, FM18) | |||||||
| AT5G09530 | PRP10 (WT, ANP4, ANP10) | FPENSKPEVPK (1; WM8, WM9; F2, FM18; A5) | 64.8 (in WT) | 1–35 6 | 41.6 | 5.9 | |
| VPEIPKPEETK (1; FM4, FM11, FM12) , MPEIQKPELPK | |||||||
| (1; FM4, FM12, FM13), LPDIPK (1; FM16), | |||||||
| LPEVPK (1; FM16), LPEFPKPELPK (1; AM4, AM18), | |||||||
| MPEIPKPELPK (1; AM20) |
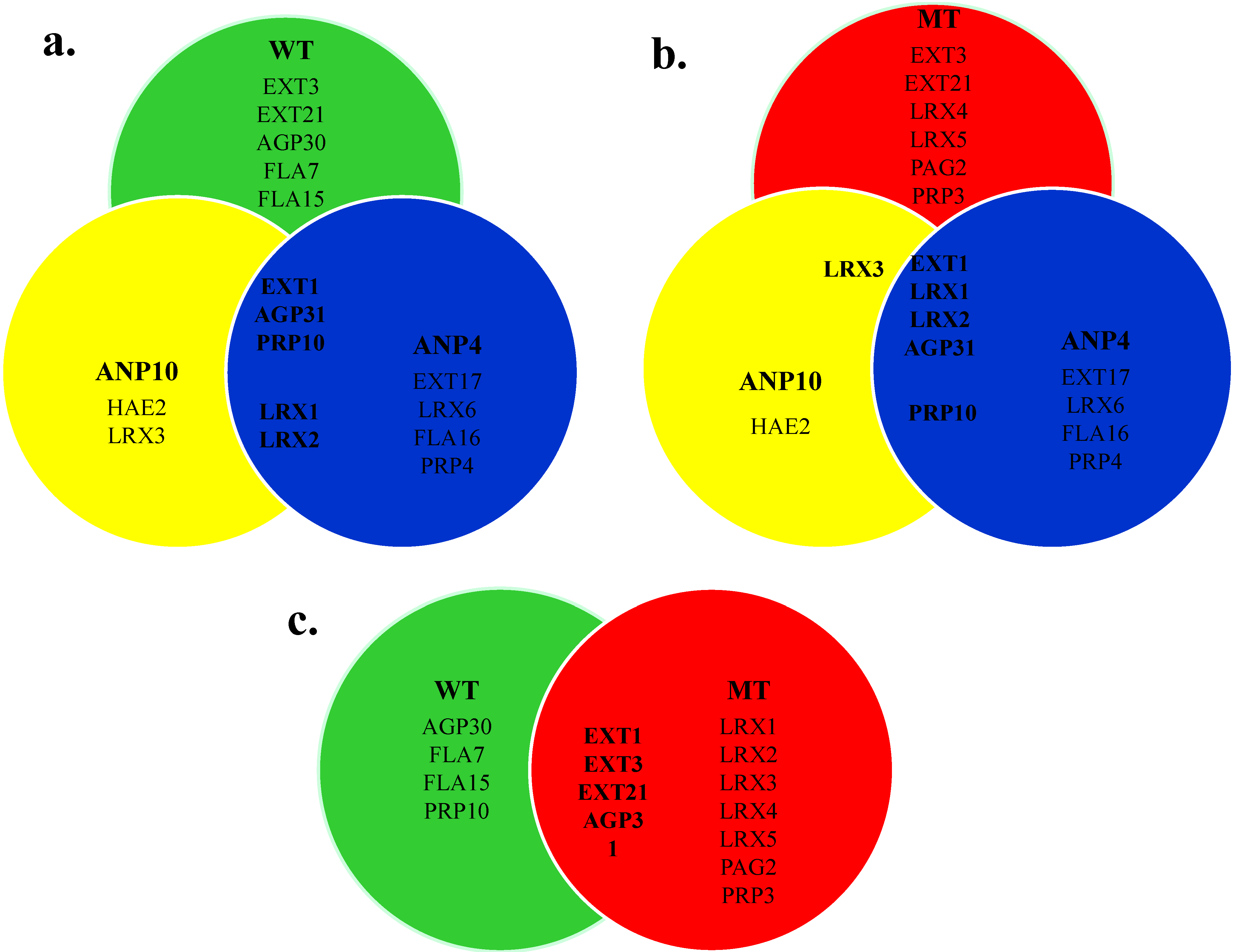
2.6. Comparison of HRGP Networks
2.7. Potential Candidate HRGPs for the Arising of ANP
| Gene Accession # | Protein Identification | Peptides Identified 1 (Number of Occurrence in Protein Sequence) | Peptide Mascot Ion Score 2 | Total Spectra Count 3 | Signal Peptide 4 | MM (kDa) 5 | pI 5 |
|---|---|---|---|---|---|---|---|
| EXTs | |||||||
| AT1G76930 | EXT1 | HYSOOOVYK (9), YYSOOOVYK (3) | 6 | 1–24 | 32.9 | 10.1 | |
| AT1G21310 | EXT3, RSH | HYSOOOVYHSOOOOK (13) | 46.6 | 1 | 1–28 | 49.2 | 10.3 |
| AT2G43150 | EXT21 | SOOOOYYYHSOOOOVK (6) | 54.4 | 3 | 1–27 | 23.4 | 9.6 |
| AT1G12040 | LRX1 | KVTVFDITSNR (1), VTVFDITSNR (1), VVLSLPSLK (1) | 5 | 1–26 | 81 | 8.2 | |
| AT1G62440 | LRX2 | ELGLLTDLALFHLNSNR (1), FPNVVLSLPSLK (1), SLEQLNVANNR (1) | 10 | 89.8 | 4.8 | ||
| AT4G13340 | LRX3 | DLDAIFINHNR (1), FNEFEGTVPK (1), FPTVVLQLPSLK (1) | 11 | 1–20 | 82.2 | 7 | |
| FRFELPENFGDSPVSVIVLANNR (1) | |||||||
| AT3G24480 | LRX4 | FPTVVLHLPSLK (1), SAYIALQAWK (1) | 11 | 1–25 | 54.7 | 6.9 | |
| AT4G18670 | LRX5 | FAGIFPTVVLQLPSLK (1), FNEFEGPVPR (1) | 9 | 1–31 6 | 90.8 | 6.7 | |
| AGPs | |||||||
| AT1G28290 | AGP31 | LFGGDVGAELKPEK (1), NGYFLLLAPK (1), TVTNFGFR (1) | 5 | 1–25 | 38.4 | 10.8 | |
| AT2G25060 | PAG2 | LSLVVISPR (1) | 44.7 | 1 | 1–28 | 19.5 | 7.4 |
| PRPs | |||||||
| AT3G62680 | PRP3 | GLTGVPLALYGYR (1), SNTEVVIYSNPTDSK (1) | 2 | 1–22 | 34.4 | 9.7 |
| Protein (Accession #) | Fold Change (Log2) ANP vs. WT 1 | Fold Change (Log2) ANP vs. MT1 | |
|---|---|---|---|
| ANP4 | EXTs: | ||
| AtEXT1 (AT1G76930) | 0.22; (0.72) 2 | −2.50; (0.00) | |
| AtEXT17 (AT3G54580) | −0.94; (0.13) | 1.03; (0.20) | |
| LRX1 (AT1G12040) | −0.45; (0.29) | 5.35; (0.02) | |
| LRX2 (AT1G62440) | −0.85; (0.05) | −0.66; (0.05) | |
| LRX6 (AT3G22800) | −0.50; (0.30) | −0.95; (0.05) | |
| AGPs: | |||
| AGP31 (AT1G28290) | n/d 3 | n/d | |
| FLA16 (AT2G35860) | n/d | n/d | |
| PRPs: | |||
| PRP4 (AT4G38770) | 1.20; (0.00) | 2.25; (0.00) | |
| PRP10 (AT5G09530) | −0.55; (0.05) | 3.30; (0.01) | |
| ANP10 | EXTs: | ||
| AtEXT1 (AT1G76930) | 0.40; (0.04) | −1.69; (0.001) | |
| LRX1 (AT1G12040) | −0.07; (0.86) | 6.90; (0.0001) | |
| LRX2 (AT1G62440) | −0.07; (0.79) | 0.89; (0.01) | |
| LRX3 (AT4G13340) | 0.64; (0.02) | −2.27; (0.0003) | |
| HAE2 (AT3G50580) | 3.33; (0.02) | 0.14; (0.77) | |
| AGPs: | |||
| AGP31 (AT1G28290) | n/d | n/d | |
| PRPs: | |||
| PRP10 (AT5G09530) | −1.33; (0.00) | 3.7; (7.9 × 10−6) |
3. Discussion
3.1. Structural Protein Network in Arabidopsis Root Cell Wall
3.2. Protease Degradation of HFI
3.3. EXT3 in Root Cell Wall
3.4. Cross-linking of HRGPs into the Cell Wall
3.5. Potential Candidate HRGPs for the WT Phenotype Reversion
3.6. Gene Expression vs. Protein Identification by Proteomics
4. Experimental Section
4.1. Arabidopsis Seedling Culture
4.2. Root Cell Wall Isolation
4.3. Wall Deglycosylation with HF
4.4. Trypsin Digestion of HFI
4.5. Pronase Degradation
4.6. Tryptic Peptide Isolation
4.7. Peptide Sequencing by Mass Spectrometry
5. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Cassab, G.I.; Varner, J.E. Cell wall proteins. Ann. Rev. Plant. Physiol. Plant. Mol. Biol. 1998, 39, 321–353. [Google Scholar] [CrossRef]
- Mort, A.J.; Lamport, D.T.A. Anhydrous hydrogen fluoride deglycosylates glycoproteins. Anal. Biochem. 1977, 82, 289–309. [Google Scholar] [CrossRef] [PubMed]
- Komalavilas, P.; Zhu, J.K.; Nothnagel, E.A. Arabinogalactan-proteins from the suspension culture medium and plasma membrane of rose cell. J. Biol. Chem. 1991, 268, 15956–15965. [Google Scholar]
- Showalter, A.M.; Bell, J.N.; Cramer, C.L.; Bailey, J.A.; Varner, J.E.; Lamb, C.J. Accumulation of Hydroxyproline-rich glycoprotein mRNAs in response to fungal elicitor and infection. Proc. Natl. Acad. Sci. USA 1985, 82, 6551–6555. [Google Scholar] [CrossRef] [PubMed]
- Bradley, D.J.; Kjellbom, P.; Lamb, C.J. Elicitor- and wound-induced oxidative cross-linking of a proline-rich plant cell wall protein: A novel, rapid defense response. Cell 1992, 70, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Clarke, A.; Gleeson, P.; Harrison, S.; Knox, R.B. Pollen-stigma interactions: identification and characterization of surface components with recognition potential (cell surface receptors/plant glycoproteins/adhesion/concanavalin A/tridacnin). Proc. Natl. Acad. Sci. USA 1979, 76, 3358–3362. [Google Scholar] [CrossRef] [PubMed]
- Levitin, B.; Richter, D.; Markovich, I.; Zik, M. Arabinogalactan proteins 6 and 11 are required for stamen and pollen function in Arabidopsis. Plant J. 2008, 56, 351–363. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Eberhard, S.; Pattathil, S.; Warder, C.; Glushka, J.; Yuan, C.; Hao, Z.; Zhu, X.; Avci, U.; Miller, J.S.; et al. An Arabidopsis cell wall proteoglycan consists of pectin and Arabinoxylan covalently linked to an Arabinogalactan protein. Plant Cell 2013, 25, 270–287. [Google Scholar] [CrossRef] [PubMed]
- Lamport, D.T.A.; Varnai, P. Periplasmic Arabinogalactan glycoproteins act as a calcium capacitor that regulates plant growth and development. New Phytol. 2013, 197, 58–64. [Google Scholar] [CrossRef] [PubMed]
- Lamport, D.T.A.; Varnai, P.; Seal, C.E. Back to the future with the AGP-Ca2+ flux capacitor. Ann. Bot. 2014. [Google Scholar] [CrossRef]
- Shpak, E.; Leykam, J.F.; Kieliszewsk, M.J. Synthetic genes for glycoprotein design and the elucidation of hydroxyproline-O-glycosylation codes. Proc. Natl. Acad. Sci. USA 1999, 96, 14736–14741. [Google Scholar] [CrossRef] [PubMed]
- Du, H.; Simpson, R.J.; Moritz, R.L.; Clarke, A.E.; Bacic, A. Isolation of the protein backbone of an Arabinogalactan-protein from the styles of Nicotiana alata and characterization of a corresponding cDNA. Plant Cell 1994, 6, 1643–1653. [Google Scholar] [CrossRef] [PubMed]
- Youl, J.J.; Bacic, A.; Oxley, D. Arabinogalactan-proteins from Nicotiana alata and Pyrus communis contain glycosylphosphatidylinositol membrane anchors. Proc. Natl. Acad. Sci. USA 1998, 95, 7921–7926. [Google Scholar] [CrossRef] [PubMed]
- Oxley, D.; Bacic, A. Structure of the glycosylphosphatidylinositol anchor of an arabinogalactan protein from Pyrus communis suspension-cultured cells. Proc. Natl. Acad. Sci. USA 1999, 96, 14246–14251. [Google Scholar] [CrossRef] [PubMed]
- Schultz, C.J.; Johnson, K.L.; Currie, G.; Bacic, A. The classical arabinogalactan protein gene family of arabidopsis. Plant Cell 2000, 12, 1751–1767. [Google Scholar] [CrossRef] [PubMed]
- Fowler, T.J.; Bernhardt, C.; Tierney, M.L. Characterization and expression of four proline-rich cell wall protein genes in arabidopsis encoding two distinct subsets of multiple domain proteins. Plant Physiol. 1999, 121, 1081–1091. [Google Scholar] [PubMed]
- Wyatt, R.E.; Nagao, R.; Key, J.L. Patterns of soybean proline-rich protein gene expression. Plant Cell 1992, 4, 99–110. [Google Scholar] [CrossRef] [PubMed]
- Lamport, D.T.A.; Kieliszewski, M.J.; Chen, Y.; Cannon, M.C. Role of the extensin superfamily in primary cell wall architecture. Plant Physiol. 2011, 156, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Sadava, D.; Chrispeels, M.J. Hydroxyproline-rich cell wall protein (extensin): Role in the cessation of elongation in excised pea epicotyls. Dev. Biol. 1973, 30, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Lamport, D.T.A.; Katona, L.; Roerig, S. Galactosylserine in extensin. Biochem. J. 1973, 133, 125–131. [Google Scholar] [PubMed]
- Brady, J.D.; Sadler, I.H.; Fry, S.C. Di-isodityrosine, a novel tetrameric derivative of tyrosine in plant cell wall proteins: A new potential cross-link. Biochem. J. 1996, 315, 323–327. [Google Scholar] [PubMed]
- Held, M.A.; Tan, L.; Kamyab, A.; Hare, M.; Shpak, E.; Kieliszewski, M.J. Di-isodityrosine is the intermolecular cross-link of Isodityrosine-rich extensin analogs cross-linked in vitro. J. Biol. Chem. 2004, 279, 55474–55482. [Google Scholar] [CrossRef] [PubMed]
- Brady, J.D.; Sadler, I.H.; Fry, S.C. Pulcherosine, an oxidatively coupled trimer of tyrosine in plant cell wall: Its role in cross-link formation. Phytochemistry 1998, 47, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Schnabelrauch, L.S.; Kieliszewski, M.J.; Upham, B.L.; Alizedeh, H.; Lamport, D.T.A. Isolation of pI 4.6 extensin peroxidase from tomato cell suspension cultures and identification of Val-Tyr-Lys as putative intermolecular cross-link site. Plant J. 1996, 9, 477–489. [Google Scholar] [CrossRef] [PubMed]
- Brady, J.D.; Fry, S.C. Formation of di-lsodityrosine and loss of lsodityrosine in the cell walls of tomato cell-suspension cultures treated with fungal elicitors or H2O2. Plant Physiol. 1997, 11, 87–92. [Google Scholar]
- Hall, Q.; Cannon, M.C. The cell wall hydroxyproline-rich glycoprotein RSH is essential for normal embryo development in Arabidopsis. Plant Cell 2002, 14, 1161–1172. [Google Scholar] [CrossRef] [PubMed]
- Cannon, M.C.; Terneus, K.; Hall, Q.; Tan, L.; Wang, Y.; Wegenhart, B.L.; Chen, L.; Lamport, D.T.A.; Chen, Y.; Kieliszewski, M.J. Self-assembly of the plant cell wall requires an extensin scaffold. Proc. Natl. Acad. Sci. USA 2008, 105, 2226–2231. [Google Scholar] [CrossRef] [PubMed]
- Saha, P.; Ray, T.; Tang, Y.; Duttal, I.; Evangelous, N.R.; Kieliszewski, M.J.; Chen, Y.; Cannon, M.C. Self-rescue of an EXTENSIN mutant reveals alternative gene expression programs and candidate proteins for new cell wall assembly in Arabidopsis. Plant J. 2013, 75, 104–116. [Google Scholar] [CrossRef] [PubMed]
- Hijazi, M.; Durand, J.; Pichereaux, C.; Pont, F.; Jamet, E.; Albenne, C. Characterization of the arabinogalactan protein 31 (AGP31) of Arabidopsis thaliana: New advances on the Hyp-O-glycosylation of the pro-rich domain. J. Biol. Chem. 2012, 287, 9623–9632. [Google Scholar] [CrossRef] [PubMed]
- Sadava, D.; Walker, F.; Chrispeels, M.J. Hydroxyproline-rich cell wall protein (extensin): Biosynthesis and accumulation in growing pea epicotyls. Dev. Biol. 1973, 30, 42–48. [Google Scholar] [CrossRef]
- O’Neill, M.A.; Selvendran, R.R. Glycoproteins from the cell wall of Phaseolus coccineus. Biochem. J. 1980, 187, 53–63. [Google Scholar] [PubMed]
- Lamport, D.T.A. The isolation and partial characterization of hydroxyproline-rich glycopeptides obtained by enzymatic degradation of primary cell wall. Biochemistry 1969, 8, 1115–1163. [Google Scholar] [CrossRef]
- Heath, M.F.; Northcote, D.H. Glycoprotein of the wall of sycamore tissue-culture cells. Biochem. J. 1971, 125, 953–961. [Google Scholar] [PubMed]
- Keegstra, K.; Talmadge, K.W.; Bauer, W.D.; Albersheim, P. The structure of plant cell walls III. A model of the walls of suspension-cultured sycamore cells based on the interconnections of the macromolecular components. Plant Physiol. 1973, 51, 188–196. [Google Scholar] [CrossRef] [PubMed]
- Frueauf, J.B.; Dolata, M.; Leykam, J.F.; Lloyd, E.A.; Gonzales, M.; VandenBosch, K.; Kieliszewski, M.J. Peptides isolated from cell walls of Medicago. truncatula nodules and uninfected root. Phytochemistry 2000, 55, 429–438. [Google Scholar]
- Mort, A.J.; Grover, P.B.J. Characterization of root hair cell walls as potential barriers to the infection of plants by rhizobia: the carbohydrate component. Plant Physiol. 1988, 86, 638–641. [Google Scholar] [CrossRef] [PubMed]
- Dolan, L.; Janmaat, K.; Willemsen, V.; Linstead, P.; Poethig, S.; Roberts, K.; Scheres, B. Cellular organization of the Arabidopsis thaliana root. Development 1993, 119, 71–84. [Google Scholar] [PubMed]
- Baxter, I.; Hosmani, P.S.; Rus, A.; Lahner, B.; Borevitz, J.O.; Muthukumar, B.; Mickelbart, M.V.; Schreiber, L.; Franke, R.B.; Salt, D.E. Root suberin forms an extracellular barrier that affects water relations and mineral nutrition in Arabidopsis. PLOS Genet. 2009, 5, e1000492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ranathunge, K.; Schreiber, L. Water and solute permeabilities of Arabidopsis roots in relation to the amount and composition of aliphatic suberin. J. Exp. Bot. 2011, 62, 1961–1974. [Google Scholar] [CrossRef] [PubMed]
- Hardt, H.; Lamport, D.T.A. Hydrogen fluoride saccharification of wood: Lignin fluoride content, isolation of alpha-D-glucopyranosyl fluoride and posthydrolysis of reversion products. Biotechnol. Bioeng. 1982, 24, 903–918. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.; Qiu, F.; Yan, B.; Wang, H.; Mort, A.J.; Stark, R.E. NMR studies of molecular structure in fruit cuticle polyesters. Phytochemistry 2001, 57, 1035–1042. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; Fang, X.; Wang, W.; Yu, B.; Cheng, X.; Qiu, F.; Mort, A.J.; Stark, R.E. Isolation and identification of oligomers from partial degradation of lime fruit cutin. J. Agric. Food Chem. 2008, 56, 10318–10325. [Google Scholar] [CrossRef] [PubMed]
- Mort, A.J. Partial Characterization of Extensin by Selective Destruction of Cell Walls. Ph.D. Thesis, Michigan State University, East Lancing, MI, USA, 1978. [Google Scholar]
- Whitmore, F.W. Lignin-carbohydrate complex formed in isolated cell walls of callus. Phytochemistry 1978, 17, 421–425. [Google Scholar] [CrossRef]
- McDougall, G.J.; Stewart, D.; Morrison, I.M. Tyrosine residues enhance cross-linking of synthetic proteins into lignin-like dehydrogenation products. Phytochemistry 1996, 41, 43–47. [Google Scholar] [CrossRef]
- Terneus, K.A. The Isolation and Characterization of a Tobacco Extensin Precursor and Two Arabidopsis Hydroxyproline-rich Glycoproteins. Ph.D. Thesis, Ohio University, Athens, OH, USA, 2006. [Google Scholar]
- Fry, S.C. Isodityrosine, A new cross-linking amino acid from plant cell-wall glycoprotein. Biochem. J. 1982, 204, 449–455. [Google Scholar] [PubMed]
- Epstein, L.; Lamport, D.T.A. An intramolecular linkage involving isodityrosine in extensin. Phytochemistry 1984, 23, 1241–1246. [Google Scholar] [CrossRef]
- Smith, J.J.; Muldoon, E.P.; Lamport, D.T.A. Isolation of extensin precursors by direct elution of intact tomato cell suspension cultures. Phytochemistry 1984, 23, 1233–1239. [Google Scholar] [CrossRef]
- Smith, J.J.; Muldoon, E.P.; Willard, J.J.; Lamport, D.T.A. Tomato extensin precursors P1 and P2 are highly periodic structures. Phytochemistry 1986, 25, 1021–1030. [Google Scholar] [CrossRef]
- Lloyd, E.A.; Diezman, J.M.; Leykam, L.F.; Kieliszewski, M.J. Synthetic Genes for Elucidating Cross-linking Amino Acids in P1 Type Extensins. In Proceedings of the Annual Meeting of the American Society of Plant Biologists, Honolulu, HA, USA, 25–30 July 2003.
- Baumberger, N.; Ringli, C.; Keller, B. The chimeric leucine-rich repeat/extensin cell wall protein LRX1 is required for root hair morphogenesis in Arabidopsis thaliana. Gene Dev. 2001, 15, 1128–1139. [Google Scholar] [CrossRef] [PubMed]
- Baumberger, N.; Doesseger, B.; Guyot, R.; Diet, A.; Parsons, R.L.; Clark, M.A.; Simmons, M.P.; Bedinger, P.; Goff, S.A.; Ringli, C.; et al. Whole-genome comparison of leucine-rich repeat extensins in Arabidopsis and rice. A conserved family of cell wall proteins form a vegetative and a reproductive clade. Plant Physiol. 2003, 131, 1313–1326. [Google Scholar] [CrossRef] [PubMed]
- Baumberger, N.; Steiner, M.; Ryser, U.; Keller, B.; Ringli, C. Synergistic interaction of the two paralogous Arabidopsis genes LRX1 and LRX2 in cell wall formation during root hair development. Plant J. 2003, 35, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Ringli, C. The hydroxyproline-rich glycoprotein domain of the Arabidopsis LRX1 requires Tyr for function but not for insolubilization in the cell wall. Plant J. 2010, 63, 662–669. [Google Scholar] [CrossRef] [PubMed]
- Bayer, E.; Bottrill, A.; Walshaw, J.; Vigouroux, M.; Naldrett, M.; Thomas, C.; Maule, A. Arabidopsis cell wall proteome defined using multidimensional protein identification technology. Proteomics 2006, 6, 301–311. [Google Scholar] [CrossRef] [PubMed]
- Irshad, M.; Canut, H.; Borderies, G.; Pont-Lezica, R.; Jamet, E. A new picture of cell wall protein dynamics in elongating cells of Arabidopsis thaliana: Confirmed actors and newcomers. BMC Plant Biol. 2008, 8. [Google Scholar] [CrossRef] [PubMed]
- Baldwin, T.C.; McCann, M.C.; Roberts, K. A Novel Hydroxyproline-deficient arabinogalactan protein secreted by suspension-cultured cells of Daucus carota. Plant Physiol. 1993, 103, 115–123. [Google Scholar] [PubMed]
- Immerzeela, P.; Eppink, M.M.; de Vries, S.C.; Schols, H.A.; Voragen, A.G.J. Carrot arabinogalactan proteins are interlinked with pectins. Physiol. Plantarum. 2006, 128, 18–28. [Google Scholar] [CrossRef]
- Hijazi, M.; Roujol, D.; Nguyen-Kim, H.; Del Rocio Cisneros Castillo, L.; Saland, E.; Jamet, E.; Albenne, C. Arabinogalactan protein 31 (AGP31), a putative network-forming protein in Arabidopsis thaliana cell walls? Ann. Bot. 2014. [Google Scholar] [CrossRef]
- Johnson, K.L.; Jones, B.J.; Bacic, A.; Schultz, C.J. The fasciclin-like arabinogalactan proteins of Arabidopsis. A multigene family of putative cell adhesion molecules. Plant Physiol. 2003, 133, 1911–1925. [Google Scholar] [CrossRef] [PubMed]
- Johnson, K.L.; Kibble, N.A.J.; Bacic, A.; Schultz, C.J. A fasciclin-like arabinogalactan-protein (FLA) mutant of Arabidopsis thaliana, FLA1, shows defects in shoot regeneration. PLOS ONE 2011, 6, e25154. [Google Scholar] [CrossRef] [PubMed]
- Roubal, W.T.; Tappel, A.L. Polymerization of proteins induced by free-radical lipid peroxidation. Arch. Biochem. Biophys. 1966, 113, 150–155. [Google Scholar] [CrossRef] [PubMed]
- Kjellbom, P.; Snogerup, L.; Stöhr, C.; Reuzeau, C.; McCabe, P.F.; Pennell, R.I. Oxidative cross-linking of plasma membrane arabinogalactan proteins. Plant J. 1997, 12, 1189–1196. [Google Scholar] [CrossRef] [PubMed]
- Stafstrom, J.P.; Staehelin, L.A. The role of carbohydrate in maintaining extensin in an extended conformation. Plant Physiol. 1986, 81, 242–246. [Google Scholar] [CrossRef] [PubMed]
- Cooper, J.B.; Varner, J.E. Cross-linking of soluble extensin in isolated cell walls. Plant Physiol. 1984, 76, 414–417. [Google Scholar] [CrossRef] [PubMed]
- Valentin, R.; Cerclier, C.; Geneix, N.; Aguié-Béghin, V.; Gaillard, C.; Ralet, M.C.; Cathala, B. Elaboration of extensin-pectin thin film model of primary plant cell wall. Langmuir 2010, 26, 9891–9898. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Shalongo, W.; Stallwagen, E. The role of PII conformations in the calculation of peptide fractional helix content. Protein Sci. 1997, 6, 1694–1700. [Google Scholar] [CrossRef] [PubMed]
- Chuang, C.F.; Meyerowitz, E.M. Specific and heritable genetic interference by double-stranded RNA in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2000, 97, 4985–4990. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Reichel, M.; Li, Y.; Millar, A.A. The functional scope of plant microRNA-mediated silencing. Trends Plant Sci. 2014, 19, 785–756. [Google Scholar]
- York, W.S.; Darvill, A.G.; McNeil, M.; Stevenson, T.T.; Albersheim, P. Isolation and characterization of plant cell walls and cell wall components. Methods Enzymol. 1986, 118, 3–40. [Google Scholar]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, Y.; Ye, D.; Held, M.A.; Cannon, M.C.; Ray, T.; Saha, P.; Frye, A.N.; Mort, A.J.; Kieliszewski, M.J. Identification of the Abundant Hydroxyproline-Rich Glycoproteins in the Root Walls of Wild-Type Arabidopsis, an ext3 Mutant Line, and Its Phenotypic Revertant. Plants 2015, 4, 85-111. https://doi.org/10.3390/plants4010085
Chen Y, Ye D, Held MA, Cannon MC, Ray T, Saha P, Frye AN, Mort AJ, Kieliszewski MJ. Identification of the Abundant Hydroxyproline-Rich Glycoproteins in the Root Walls of Wild-Type Arabidopsis, an ext3 Mutant Line, and Its Phenotypic Revertant. Plants. 2015; 4(1):85-111. https://doi.org/10.3390/plants4010085
Chicago/Turabian StyleChen, Yuning, Dening Ye, Michael A. Held, Maura C. Cannon, Tui Ray, Prasenjit Saha, Alexandra N. Frye, Andrew J. Mort, and Marcia J. Kieliszewski. 2015. "Identification of the Abundant Hydroxyproline-Rich Glycoproteins in the Root Walls of Wild-Type Arabidopsis, an ext3 Mutant Line, and Its Phenotypic Revertant" Plants 4, no. 1: 85-111. https://doi.org/10.3390/plants4010085
APA StyleChen, Y., Ye, D., Held, M. A., Cannon, M. C., Ray, T., Saha, P., Frye, A. N., Mort, A. J., & Kieliszewski, M. J. (2015). Identification of the Abundant Hydroxyproline-Rich Glycoproteins in the Root Walls of Wild-Type Arabidopsis, an ext3 Mutant Line, and Its Phenotypic Revertant. Plants, 4(1), 85-111. https://doi.org/10.3390/plants4010085





