Research Progress on Rice-Blast-Resistance-Related Genes
Abstract
1. Introduction
2. Molecular Mechanisms of Rice Immunity and Rice Blast Invasion
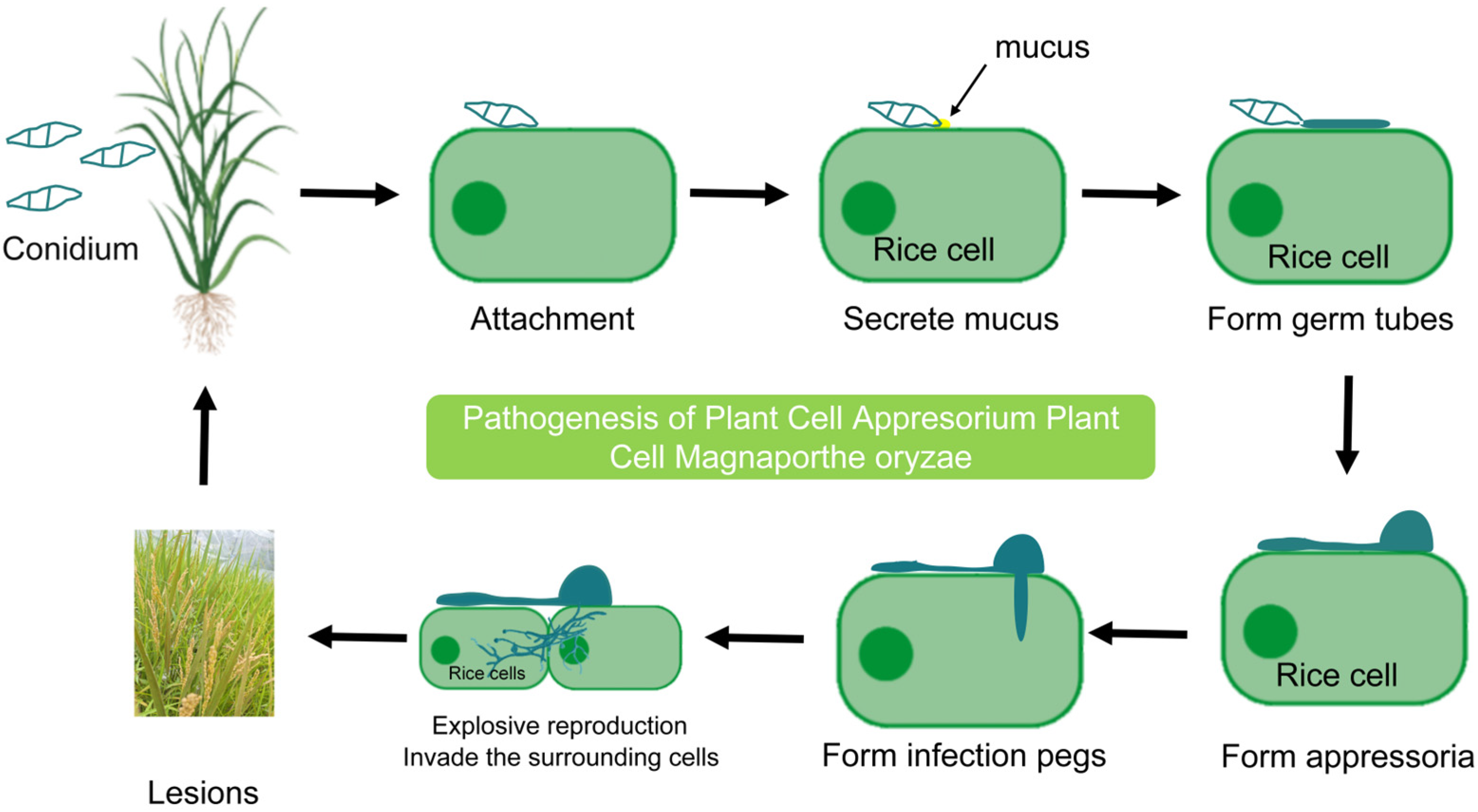
2.1. Infection Mechanism of Rice Blast Fungus
2.2. Mechanisms of Action of PTI and ETI
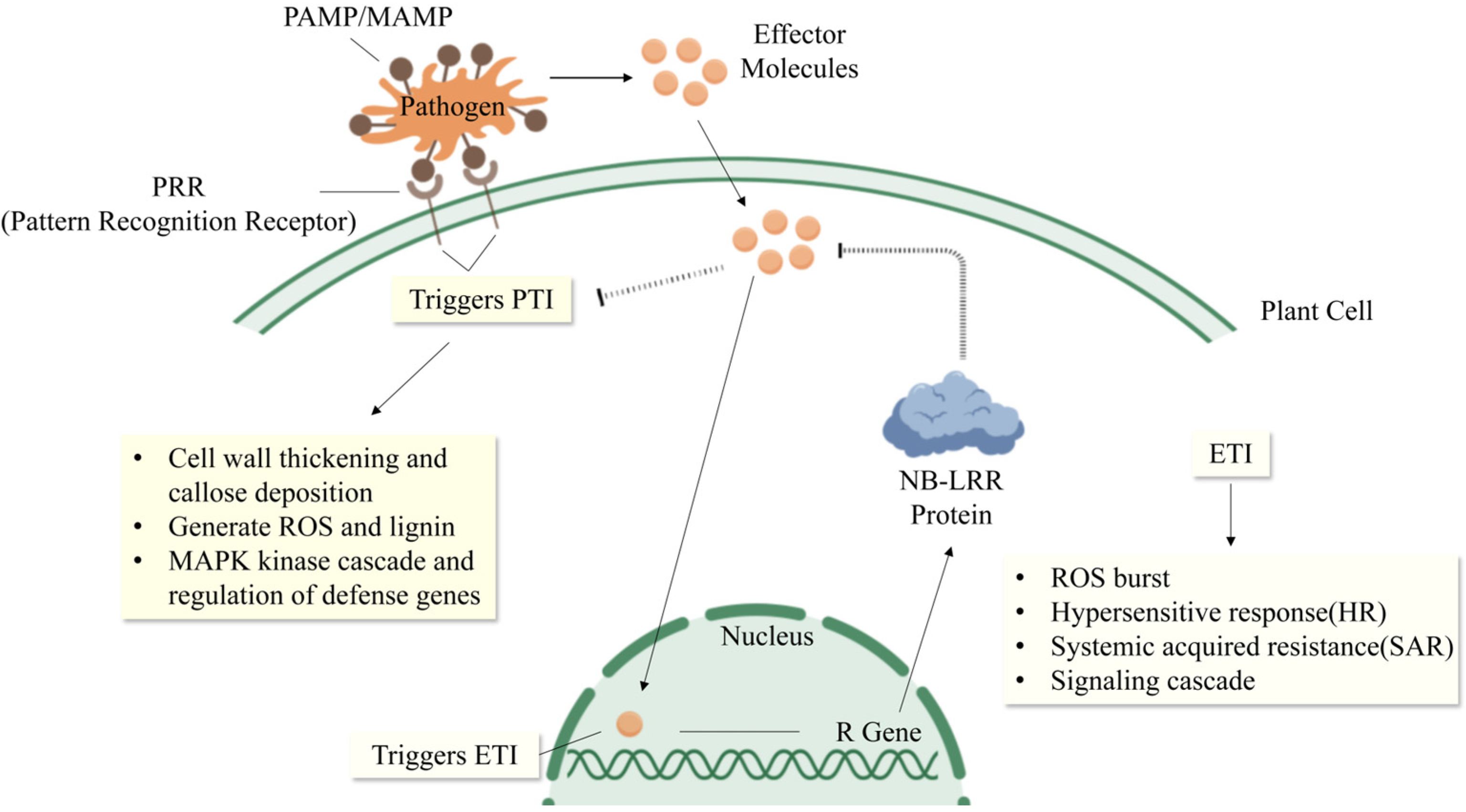
3. Research Progress on Rice Blast Resistance Genes
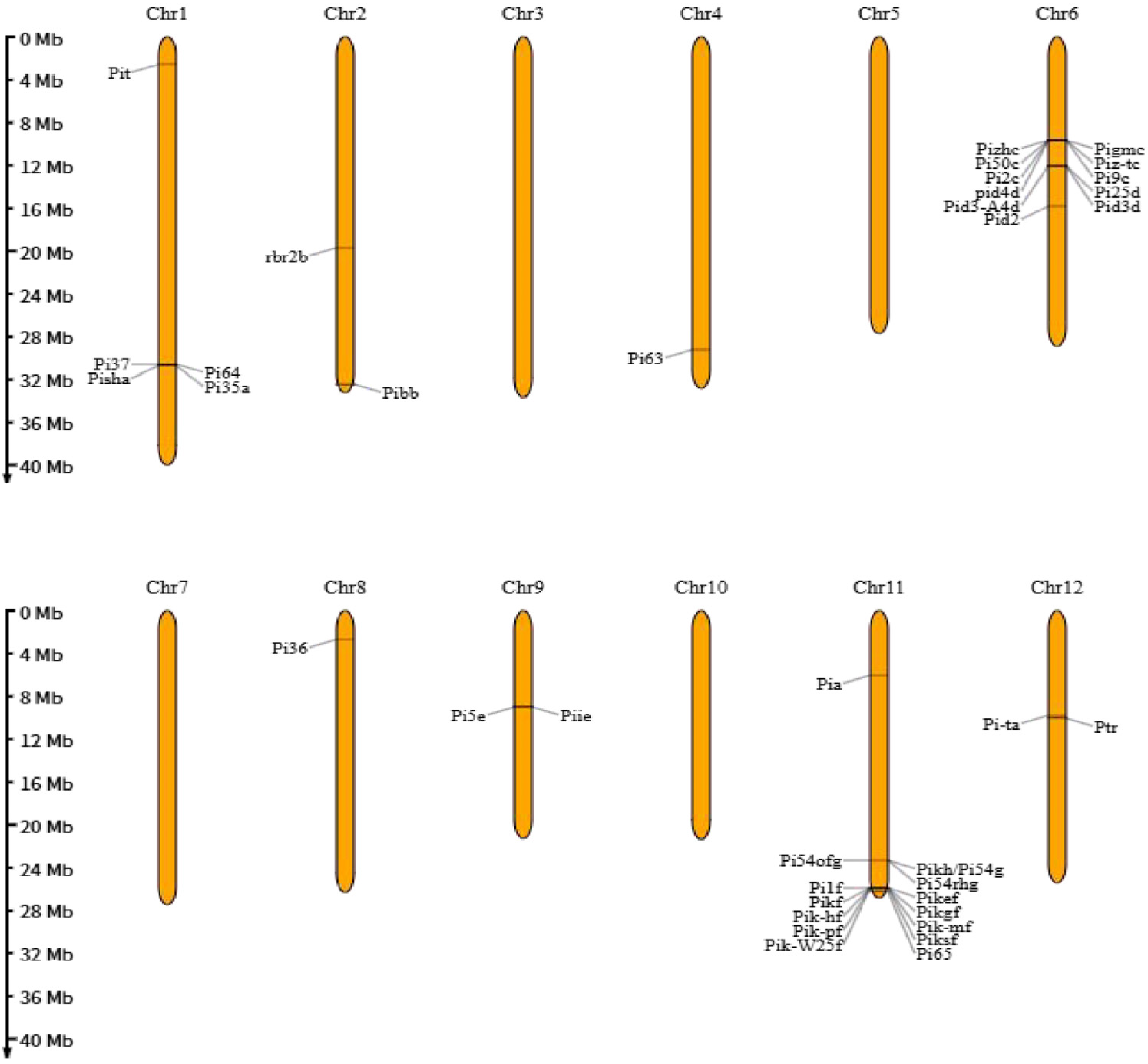
3.1. Mechanism of Rice Blast Resistance Controlled by Major Resistance Genes (R Genes)
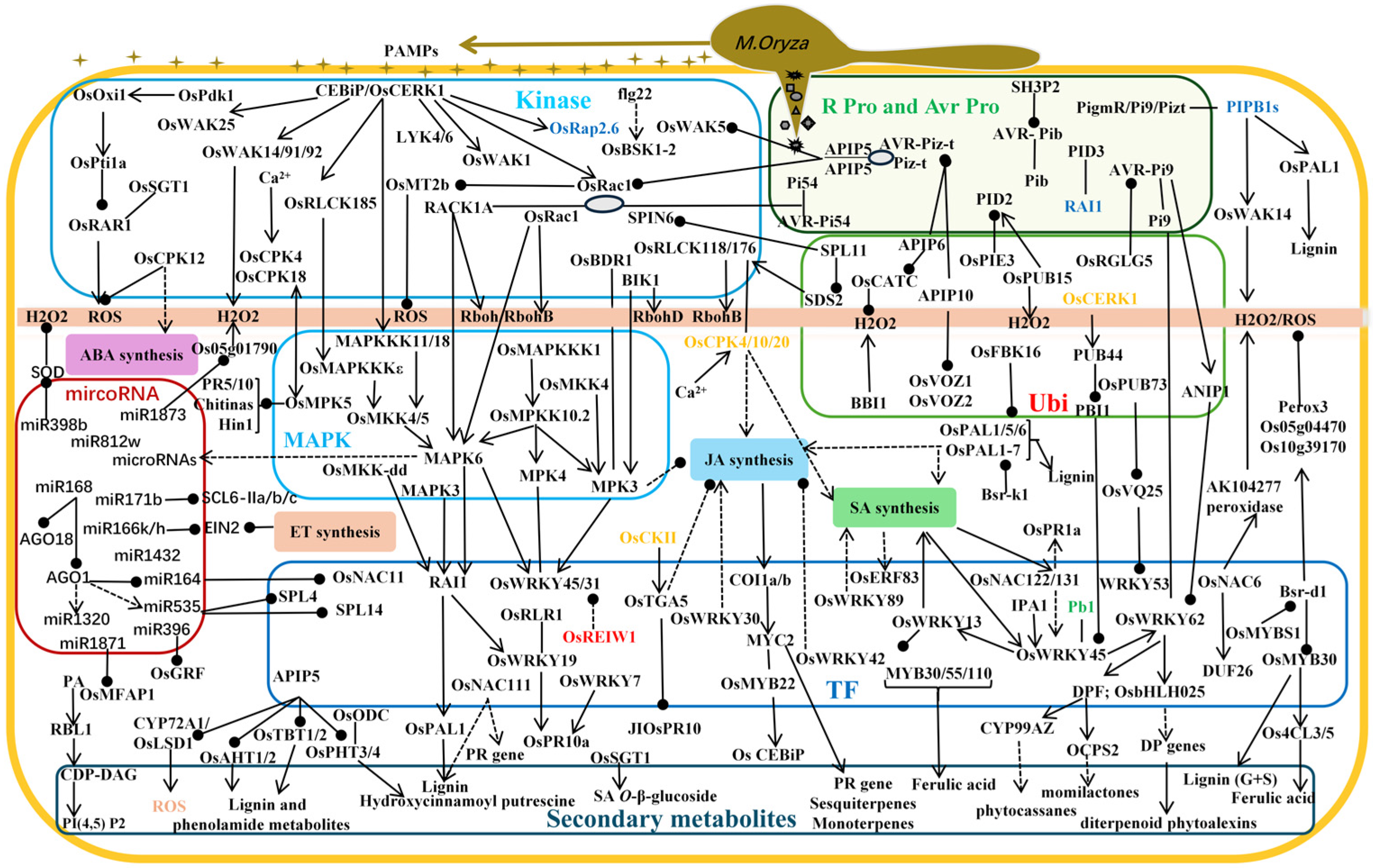
3.2. Resistance Mechanism of Rice Blast Controlled by Protein Kinases
3.3. Partial Resistance Mechanism of Rice Blast Controlled by Ubiquitin Ligases
3.4. Mechanism of Rice Blast Resistance Controlled by Transcription Factors
3.5. Partial Resistance Mechanism of Rice Blast Controlled by microRNA
3.6. Partial Resistance Mechanism of Rice Blast Controlled by Plant Hormones
3.7. Resistance Mechanism of Rice Blast Controlled by Secondary Metabolites
4. Rice Blast Resistance Breeding
4.1. Induce Blast Resistance Through Marker-Assisted Selection Breeding
4.2. Induce Blast Resistance Through Marker-Assisted Selection Breeding
4.3. Inducing Rice Blast Resistance by Genomics-Assisted Breeding
4.4. Induce Blast Resistance Through Gene-Editing Breeding
4.5. Assist Rice Blast Resistance Breeding Through AI Technology

5. Conclusions
6. Future Prospects
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| PTI | PAMP-triggered immunity |
| PAMP | pathogen-associated molecular pattern |
| ETI | elicit effector-triggered immunity |
| PRRs | Pattern recognition receptors |
| NLR | nucleotide-binding domain leucine-rich repeat |
| PAMP | Pathogen-associated molecular patterns |
| aROS | reactive oxygen species |
| MAPK | mitogen-activated protein kinase |
| AVR | avirulence gene |
| NBS | nucleotide binding sites |
| LRR | leucine rich repeat |
| QTL | quantitative trait locus |
| NBS-LRR | nucleotide binding site–leucine-rich repeat protein |
| RLKs | Receptor-like kinase |
| STK | serine/threonine kinase domain |
| ARM | Armadillo |
| RLCK | receptor-like cytoplasmic kinase |
| CDPKs | Calcium-Dependent Protein Kinases |
| WAK-RLKs | WAK-type receptor kinases |
| SA | salicylic acid |
| MeJA | Methyl Jasmonate |
| WAK/WAKL | Wall-associated kinases |
| PAL | Phenylalanine Ammonia-Lyase |
| SPL | Squamosa promoter-binding protein-like |
| TF | transcription factors |
| JA | jasmonic acid |
| ABA | Abscisic acid |
| UPS | ubiquitin–proteasome system |
| miRNAs | MicroRNAs |
| AGO | Argonaute |
| RISC | RNA-induced silencing complex |
| CDP-DAG | cytidine diphosphate glycerol |
| PtdInsP2 | phospholipid inositol diphosphate |
| BIC | biotype interacting complex |
| EIHM | Extra-invasive hyphal membrane |
| TPR | tetratricopeptide repeats |
| IAA | auxin |
| GA | gibberellins |
| SOD | Superoxide Dismutase |
| AOX | aldehyde oxidase |
| CAT | Catalase |
| DAMP | damage-associated molecular patterns |
| GWAS | Genome-wide association analysis |
| MAS | Marker-Assisted Selection |
| GS | Genomic Selection |
| HR | hypersensitive response |
| PCD | programmed cell death |
| CRISPR/Cas9 | Clustered Regularly Interspaced Short Palindromic Repeats |
Appendix A
| Gene Name | Coding Protein | Chr | Cloning Method | Donor | Reference |
|---|---|---|---|---|---|
| Pit | CNL | 1 | Map-based cloning | K59 | [135] |
| Pish a | NLR | 1 | Mutagenesis-based cloning | Nipponbarc | [136] |
| Pi35 a | NLR | 1 | Map-based cloning | Hokkai-188 | [137] |
| Pi37 | NLR | 1 | Map-based cloning | St. No. 1 | [138] |
| Pi64 | CNL | 1 | Map-based cloning | Yangmaogu | [39] |
| Pib b | NLR | 2 | Map-based cloning | BL1 | [28,139] |
| rbr2 b | CNL | 2 | Map-based cloning | Minghui 63 | [140] |
| Pi63 | CNL | 4 | Map-based cloning | Kahei | [141] |
| Pid2 | B-lectin RLK | 6 | Map-based cloning | Digu | [30] |
| Pid3 d | NLR | 6 | Comparative Genomics | MC276 | [142] |
| pid4 d | CNL | 6 | Transcriptome and comparative genomics | Digu | [143] |
| Pid3-A4 d | NLR | 6 | Allelic mining | A4 (Oryza rufipogon) | [144] |
| Pi25 d | NLR | 6 | Map-based cloning | Gumei2 | [145] |
| Pi9 c | CNL | 6 | Map-based cloning | 75-1-127 | [27] |
| Pi2 c | CNL | 6 | Map-based cloning | C101A51 | [146] |
| Piz-t c | CNL | 6 | Map-based cloning | Toride 1 | [146] |
| Pi50 c | CNL | 6 | Map-based cloning | Er-Ba-zhan | [147] |
| Pigm c | CNL | 6 | Map-based cloning | Gumei 4 | [148] |
| Pizh c | CNL | 6 | Map-based cloning | Zhonghua 11 | [149] |
| Pi36 | CNL | 8 | Map-based cloning | Q61 | [150] |
| Pi5 e | CNL | 9 | Map-based cloning | RIL260 | [151] |
| Pii e | CNL | 9 | Allele mining | Fujisaka5 | [36] |
| Pik h/Pi54 g | CNL | 11 | Map-based cloning | Tetep | [152] |
| Pi54rh g | CNL | 11 | Allele mining | Oryza rhizomatis | [153] |
| Pi54of g | CNL | 11 | Allele mining | Oryza officinalis | [154] |
| Pia | NLR | 11 | Multifaceted genomics approach | Nipponbare | [155] |
| Pik f | NLR | 11 | Map-based cloning | Kusabue | [156] |
| Pike f | CNL | 11 | Map-based cloning | Xiangzao 143 | [157] |
| Piks f | CNL | 11 | Allele mining | IRBLKs-F5 | [26] |
| Pi1 f | CNL | 11 | Map-based cloning | C101LAC | [158] |
| Pik-p f | CNL | 11 | Map-based cloning | K60 | [159] |
| Pik-m f | CNL | 11 | Map-based cloning | Tsuyuake | [160] |
| Pik-h f | CNL | 11 | Map-based cloning | K3 | [161] |
| Pikg f | CNL | 11 | Allele mining | accession G9 | [162] |
| Pi65 | LRR-RLK | 11 | Map-based cloning | GangYu65 | [24] |
| Pik-W25 f | CNL | 11 | BSA mapping | WR25 | [108] |
| Pi-ta | NLR | 12 | Map-based cloning | Katy | [163] |
| Ptr | ARM repeats | 12 | Map-based cloning | BHA | [33] |
| S. No | Variety | Breeding Techniques | Researcher or Unit |
|---|---|---|---|
| 1 | Chunjiang 25 | Crossbreeding | Wu Mingguo et al. [164] |
| 2 | shangyou 63 | Crossbreeding | Xiuxia Yang et al. [165] |
| 3 | Tiejing 16 | MAS | Ma et al. [166] |
| 4 | K59 | Crossbreeding | Yang Wangxing et al. [167] |
| 5 | Nipponbarc | Crossbreeding | Aichi Prefectural Agricultural Experiment Station [168] |
| 6 | Hokkai-188 | Crossbreeding | Nguyen, Fukuoka et al. [138] |
| 7 | St. No. 1 | Genome Editing | Hayano-Saito [169] |
| 8 | Yangmaogu | Crossbreeding | Jian Ma et al. [39] |
| 9 | BL1 | Backcross | Sedeek, S et al. [170] |
| 10 | Minghui 63 | Crossbreeding | Wang et al. [171] |
| 11 | Kahei | Mas | Miyamoto M., Yano M., Hirasawa H et al. [141] |
| 12 | Digu | Crossbreeding | Lihuang Zhu, Xuewei Chen et al. [30] |
| 13 | MC276 | Gs | Wan, Moroberekan et al. [172] |
| 14 | Gumei2 | Crossbreeding | Jie Chen [145] |
| 15 | Tetep | Mas | Atul Singh [173] |
| 16 | Zhonghua 11 | Gs | Xue-Feng Yao [174] |
| 17 | Q61 | Crossbreeding | Xie et al. [150] |
| 18 | Fujisaka5 | Backcross | Fujimaki Hiroshi et al. [175] |
| 19 | Oryza rhizomatis | Crossbreeding | Vaughan et al. [154] |
| 20 | Nipponbare | Crossbreeding | Okuyama et al. [155] |
| 21 | Kusabue | Crossbreeding | Zhai et al. [156] |
| 22 | Xiangzao 143 | Gs | Hunan Rice Research Institute [157] |
| 23 | K60 | Crossbreeding | Yuan et al. [159] |
| 24 | YF47 | Backcross | Mao et al. [106] |
| Source Cultivarsa | Year of Isolation | Races | State | Frequency of Occurrence |
|---|---|---|---|---|
| Caffey | 2014 | IC-17 | Arkansas | The most common race |
| CL 151 | 2013 | 1B-25 | Arkansas | Uncommon race |
| HR12 | MG01 | south of indian | The most common race | |
| CL 151 | 2015 | 1B-1 | Arkansas | The most common race |
| CL 261 | 2013 | IA-37 | Arkansas | Uncommon race |
| 70-15 | MG-8 | south of indian | The most common race | |
| CL 261 | 2013 | IA-69 | Arkansas | Uncommon race |
| Jupiter | 2015 | IA-113 | Arkansas | Uncommon race |
| CL 261 | 2012 | 1B-17 | Arkansas | The most common race |
| Jupiter | 2013 | IG-1 | Arkansas | Uncommon race |
| CL 261 | 2012 | 1B-49 | Arkansas | The most common race |
| Jupiter | 2015 | IA-1 | Arkansas | Uncommon race |
| CL 261 | 2012 | IC-9 | Arkansas | Uncommon race |
| CL 261 | 2013 | IE-1 | Arkansas | The most common race |
| FJ81278 | fujian province china | The most common race | ||
| CL 262 | 2013 | 1B-41 | Arkansas | Uncommon race |
| Colorado | 2013 | IC-1 | Texas | Uncommon race |
| CL 261 | 2013 | IA-65 | Arkansas | Uncommon race |
| Jupiter | 2013 | 1B-21 | Louisiana | Uncommon race |
| LA 2025 | 2013 | 1B-37 | Louisiana | Uncommon race |
| HN19311 | hunan province in china | The most common race |
References
- Qi, Z.; Zhou, X.; Tian, L.; Zhang, H.; Cai, L.; Tang, F. Temporal and spatial variation of microbial communities in stored rice grains from two major depots in China. Food Res. Int. 2022, 152, 110876. [Google Scholar] [CrossRef]
- Le Mouël, C.; Forslund, A. How can we feed the world in 2050? A review of the responses from global scenario studies. Eur. Rev. Agric. Econ. 2017, 44, 541–591. [Google Scholar] [CrossRef]
- Li, J.; Lu, L.; Wang, Q.; Shi, Z.; Li, C.; Guo, Z. Genome Re-Sequencing Reveals the Host-Specific Origin of Genetic Variation in Magnaporthe Species. Front. Genet. 2022, 16, 1664–8021. [Google Scholar] [CrossRef]
- Han, X.; Li, S.; Zeng, Q.; Sun, P.; Wu, D.; Wu, J.; Yu, X.; Lai, Z.; Milne, R.J.; Kang, Z.; et al. Genetic engineering, including genome editing, for enhancing broad-spectrum disease resistance in crops. Plant Commun. 2025, 6, 101195. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.; Tsuda, K.; Parker, J.E. Effector-Triggered Immunity: From Pathogen Perception to Robust Defense. Annu. Rev. Plant Biol. 2015, 66, 487–511. [Google Scholar] [CrossRef]
- Odjo, T.; Diagne, D.; Adreit, H.; Milazzo, J.; Raveloson, H.; Andriantsimialona, D.; Kassankogno, A.I.; Ravel, S.; Gumedzoé, Y.M.D.; Ouedraogo, I.; et al. Structure of African Populations of Pyricularia oryzae from Rice. Phytopathology 2021, 111, 1428–1437. [Google Scholar] [CrossRef]
- Wang, L.; Zuo, S.-M.; Zhang, Y.-F.; Chen, Z.-X.; Pan, X.-B.; Huang, S.-W. Genetic Structure of Rice Blast Pathogen Magnaporthe oryzae in Sichuan Province. Chin. J. Rice Sci. 2015, 29, 327–334. [Google Scholar] [CrossRef]
- Gupta, L.; Vermani, M.; Kaur Ahluwalia, S.; Vijayaraghavan, P. Molecular virulence determinants of Magnaporthe oryzae: Disease pathogenesis and recent interventions for disease management in rice plant. Mycology 2021, 12, 174–187. [Google Scholar] [CrossRef]
- Ryder, L.S.; Dagdas, Y.F.; Kershaw, M.J.; Venkataraman, C.; Madzvamuse, A.; Yan, X.; Cruz-Mireles, N.; Soanes, D.M.; Oses-Ruiz, M.; Styles, V.; et al. A sensor kinase controls turgor-driven plant infection by the rice blast fungus. Nature 2019, 574, 423–427. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Tang, B.; Ryder, L.S.; MacLean, D.; Were, V.M.; Eseola, A.B.; Cruz-Mireles, N.; Ma, W.; Foster, A.J.; Osés-Ruiz, M.; et al. The transcriptional landscape of plant infection by the rice blast fungus Magnaporthe oryzae reveals distinct families of temporally co-regulated and structurally conserved effectors. Plant Cell 2023, 35, 1360–1385. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Lubberstedt, T.; Xu, M. The Genetic and Molecular Basis of Plant Resistance to Pathogens. J. Genet. Genom. 2013, 40, 23–35. [Google Scholar] [CrossRef]
- Sun, L.; Zhang, J. Regulatory role of receptor-like cytoplasmic kinases in early immune signaling events in plants. FEMS Microbiol. Rev. 2020, 44, 845–856. [Google Scholar] [CrossRef]
- Shiu, S.-H.; Karlowski, W.M.; Pan, R.; Tzeng, Y.-H.; Mayer, K.F.X.; Li, W.-H. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 2004, 16, 1220–1234. [Google Scholar] [CrossRef]
- Zipfel, C. Plant pattern-recognition receptors. Trends Immunol. 2014, 35, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Roychowdhury, R.; Govta, L.; Jaiwar, S.; Wei, Z.Z.; Shams, I.; Fahima, T. Intracellular Reactive Oxygen Species-Aided Localized Cell Death Contributing to Immune Responses Against Wheat Powdery Mildew Pathogen. Phytopathology 2023, 113, 884–892. [Google Scholar] [CrossRef] [PubMed]
- Agarrwal, R.; Ahire, T.P.; Manjunatha, N.; Pokhare, S.S.; Ojha, R.; Salvi, P. Imperative Role of R-Genes and Associated Molecular Mechanisms in Plant Disease Resistance. In Biotechnological Advances for Disease Tolerance in Plants; Springer: Singapore, 2024; pp. 73–97. [Google Scholar] [CrossRef]
- Dow, M.; Newman, M.A.; von Roepenack, E. The Induction and Modulation of Plant Defense Responses by Bacterial Lipopolysaccharides. Annu. Rev. Phytopathol. 2000, 38, 241–261. [Google Scholar] [CrossRef] [PubMed]
- Tongyu, Z. Advances in plant immunity and disease resistance breeding. SPIE 2023, 12611, 1261107. [Google Scholar] [CrossRef]
- Huang, X.; Yao, W.; Chen, Q.; Lin, J.; Huang, J.; Zou, Y.; Guo, C.; He, B.; Yuan, X.; Xu, C.; et al. A century of advances in molecular genetics and breeding for sustainable resistance to rice blast disease. Theor. Appl. Genet. 2025, 138, 174. [Google Scholar] [CrossRef]
- Zhai, K.; Liang, D.; Li, H.; Jiao, F.; Yan, B.; Liu, J.; Lei, Z.; Huang, L.; Gong, X.; Wang, X.; et al. NLRs guard metabolism to coordinate pattern- and effector-triggered immunity. Nature 2022, 601, 245–251. [Google Scholar] [CrossRef]
- Danso Ofori, A.; Zheng, T.; Titriku, J.; Appiah, C.; Xiang, X.; Kandhro, A.G.; Ahmed, M.I.; Zheng, A. The Role of Genetic Resistance in Rice Disease Management. Int. J. Mol. Sci. 2025, 26, 956. [Google Scholar] [CrossRef]
- Sahu, P.K.; Sao, R.; Choudhary, D.K.; Thada, A.; Kumar, V.; Mondal, S.; Das, B.K.; Jankuloski, L.; Sharma, D. Advancement in the Breeding, Biotechnological and Genomic Tools towards Development of Durable Genetic Resistance against the Rice Blast Disease. Plants 2022, 11, 2386. [Google Scholar] [CrossRef]
- Lv, Q.H.; Huang, Z.; Xu, X.; Tang, L.; Liu, H.; Wang, C.; Zhou, Z.; Xin, Y.; Xing, J.; Peng, Z.; et al. Allelic variation of the rice blast resistance gene Pid3 in cultivated rice worldwide. Sci. Rep. 2017, 7, 10362. [Google Scholar] [CrossRef]
- Wang, L.; Ma, Z.; Kang, H.; Gu, S.; Mukhina, Z.; Wang, C.; Wang, H.; Bai, Y.; Sui, G.; Zheng, W.; et al. Cloning and functional analysis of the novel rice blast resistance gene Pi65 in japonica rice. Theor. Appl. Genet. 2022, 135, 173–183. [Google Scholar] [CrossRef]
- Yu, Y.; Ma, L.; Wang, X.; Zhao, Z.; Wang, W.; Fan, Y.; Liu, K.; Jiang, T.; Xiong, Z.; Song, Q.; et al. Genome-Wide Association Study Identifies a Rice Panicle Blast Resistance Gene, Pb2, Encoding NLR Protein. Int. J. Mol. Sci. 2022, 23, 5668. [Google Scholar] [CrossRef]
- Xiao, G.; Wang, W.; Liu, M.; Li, Y.; Liu, J.; Franceschetti, M.; Yi, Z.; Zhu, X.; Zhang, Z.; Lu, G.; et al. The Piks allele of the NLR immune receptor Pik breaks the recognition of AvrPik effectors of rice blast fungus. J. Integr. Plant Biol. 2023, 65, 810–824. [Google Scholar] [CrossRef]
- Qu, S.; Liu, G.; Zhou, B.; Bellizzi, M.; Zeng, L.; Dai, L.; Han, B.; Wang, G.L. The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rich repeat protein and is a member of a multigene family in rice. Genetics 2006, 172, 1901–1914. [Google Scholar] [CrossRef]
- Wang, Z.X.; Yano, M.; Yamanouchi, U.; Iwamoto, M.; Monna, L.; Hayasaka, H.; Katayose, Y.; Sasaki, T. The Pib gene for rice blast resistance belongs to the nucleotide binding and leucine-rich repeat class of plant disease resistance genes. Plant J. 1999, 19, 55–64. [Google Scholar] [CrossRef]
- Li, W.; Chern, M.; Yin, J.; Wang, J.; Chen, X. Recent advances in broad-spectrum resistance to the rice blast disease. Curr. Opin. Plant Biol. 2019, 50, 114–120. [Google Scholar] [CrossRef]
- Chen, X.; Shang, J.; Chen, D.; Lei, C.; Zou, Y.; Zhai, W.; Liu, G.; Xu, J.; Ling, Z.; Cao, G.; et al. A B-lectin receptor kinase gene conferring rice blast resistance. Plant J. 2006, 46, 794–804. [Google Scholar] [CrossRef] [PubMed]
- Devi, S.; Singh, K.; Umakanth, B.; Vishalakshi, B.; Rao, K.V.S.; Suneel, B.; Sharma, S.K.; Kadambari, G.K.M.; Prasad, M.S.; Senguttvel, P.; et al. Identification and Characterization of a Large Effect QTL from Oryza glumaepatula Revealed Pi68(t) as Putative Candidate Gene for Rice Blast Resistance. Rice 2020, 13, 17. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Liu, Y.; Jia, M.H.; Jia, Y. An Allelic Variant of the Broad-Spectrum Blast Resistance Gene Ptr in Weedy Rice Is Associated with Resistance to the Most Virulent Blast Race IB-33. Plant Dis. 2022, 106, 1675–1680. [Google Scholar] [CrossRef]
- Zhao, H.; Wang, X.; Jia, Y.; Minkenberg, B.; Wheatley, M.; Fan, J.; Jia, M.H.; Famoso, A.; Edwards, J.D.; Wamishe, Y.; et al. The rice blast resistance gene Ptr encodes an atypical protein required for broad-spectrum disease resistance. Nat. Commun. 2018, 9, 2039. [Google Scholar] [CrossRef]
- Zhou, X.; Liao, H.; Chern, M.; Yin, J.; Chen, Y.; Wang, J.; Zhu, X.; Chen, Z.; Yuan, C.; Zhao, W.; et al. Loss of function of a rice TPR-domain RNA-binding protein confers broad-spectrum disease resistance. Science 2018, 115, 3174–3179. [Google Scholar] [CrossRef]
- Li, W.; Zhu, Z.; Chern, M.; Yin, J.; Yang, C.; Ran, L.; Cheng, M.; He, M.; Wang, K.; Wang, J.; et al. A Natural Allele of a Transcription Factor in Rice Confers Broad-Spectrum Blast Resistance. Cell 2017, 170, 114–126.e15. [Google Scholar] [CrossRef]
- Vo, K.T.X.; Lee, S.K.; Halane, M.K.; Song, M.Y.; Hoang, T.V.; Kim, C.Y.; Park, S.Y.; Jeon, J.; Kim, S.T.; Sohn, K.H.; et al. Pi5 and Pii Paired NLRs Are Functionally Exchangeable and Confer Similar Disease Resistance Specificity. Mol. Cells 2019, 42, 637–645. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, M.; Hirabuchi, A.; Sugihara, Y.; Abe, A.; Takeda, T.; Kobayashi, M.; Hiraka, Y.; Kanzaki, E.; Oikawa, K.; Saitoh, H.; et al. A genetically linked pair of NLR immune receptors shows contrasting patterns of evolution. Proc. Natl. Acad. Sci USA 2022, 119, e2116896119. [Google Scholar] [CrossRef] [PubMed]
- Kovi, B.; Sakai, T.; Abe, A.; Kanzaki, E.; Terauchi, R.; Shimizu, M. Isolation of Pikps, an allele of Pik, from the aus rice cultivar Shoni. Genes Genet. Syst. 2023, 97, 229–235. [Google Scholar] [CrossRef]
- Ma, J.; Lei, C.; Xu, X.; Hao, K.; Wang, J.; Cheng, Z.; Ma, X.; Ma, J.; Zhou, K.; Zhang, X.; et al. Pi64, Encoding a Novel CC-NBS-LRR Protein, Confers Resistance to Leaf and Neck Blast in Rice. Mol. Plant Microbe Interact. 2015, 28, 558–568. [Google Scholar] [CrossRef]
- Wang, Y.; Pruitt, R.N.; Nürnberger, T.; Wang, Y. Evasion of plant immunity by microbial pathogens. Nat. Rev. Microbiol. 2022, 20, 449–464. [Google Scholar] [CrossRef] [PubMed]
- Bundó, M.; Coca, M. Calcium-dependent protein kinase OsCPK10 mediates both drought tolerance and blast disease resistance in rice plants. J. Exp. Bot. 2017, 68, 2963–2975. [Google Scholar] [CrossRef]
- Hamdi, J.; Kmeli, N.; Bettaieb, I.; Bouktila, D. Wall-Associated Kinase (WAK) and WAK-like Kinase Gene Family in Sugar Beet: Genome-Wide Characterization and In Silico Expression Analysis in Response to Beet Cyst Nematode (Heterodera schachtii Schmidt) Infection. J. Plant Growth Regul. 2025, 44, 1713–1727. [Google Scholar] [CrossRef]
- Li, H.; Zhou, S.Y.; Zhao, W.S.; Su, S.C.; Peng, Y.L. A novel wall-associated receptor-like protein kinase gene, OsWAK1, plays important roles in rice blast disease resistance. Plant Mol. Biol. 2009, 69, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Delteil, A.; Gobbato, E.; Cayrol, B.; Estevan, J.; Michel-Romiti, C.; Dievart, A.; Kroj, T.; Morel, J.B. Several wall-associated kinases participate positively and negatively in basal defense against rice blast fungus. BMC Plant Biol. 2016, 16, 17. [Google Scholar] [CrossRef]
- Zhang, F.; Fang, H.; Wang, M.; He, F.; Tao, H.; Wang, R.; Long, J.; Wang, J.; Wang, G.L.; Ning, Y. APIP5 functions as a transcription factor and an RNA-binding protein to modulate cell death and immunity in rice. Nucleic Acids Res. 2022, 50, 5064–5079. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhou, L.; Shi, H.; Chern, M.X.; Yu, H.; Yi, H.; He, M.; Yin, J.; Zhu, X.; Li, Y.; et al. A single transcription factor promotes both yield and immunity in rice. Science 2018, 361, 1026–1028. [Google Scholar] [CrossRef]
- Wang, L.; Xu, G.; Li, L.; Ruan, M.; Bennion, A.; Wang, G.-L.; Li, R.; Qu, S. The OsBDR1-MPK3 module negatively regulates blast resistance by suppressing the jasmonate signaling and terpenoid biosynthesis pathway. Proc. Natl. Acad. Sci. USA 2023, 120, e2211102120. [Google Scholar] [CrossRef]
- Fan, J.; Bai, P.; Ning, Y.; Wang, J.; Shi, X.; Xiong, Y.; Zhang, K.; He, F.; Zhang, C.; Wang, R.; et al. The Monocot-Specific Receptor-like Kinase SDS2 Controls Cell Death and Immunity in Rice. Cell Host Microbe 2018, 23, 498–510.e5. [Google Scholar] [CrossRef] [PubMed]
- Bundo, M.; Coca, M. Enhancing blast disease resistance by overexpression of the calcium-dependent protein kinase OsCPK4 in rice. Plant Biotechnol. J. 2016, 14, 1357–1367. [Google Scholar] [CrossRef]
- Wongsa, T.; Chankaew, S.; Monkham, T.; Sanitchon, J. Broad-Spectrum Resistance and Monogenic Inheritance of Bacterial Blight Resistance in an Indigenous Upland Rice Germplasm ULR207. Agronomy 2024, 14, 898. [Google Scholar] [CrossRef]
- Wang, B.; Xue, P.; Zhang, Y.; Zhan, X.; Wu, W.; Yu, P.; Chen, D.; Fu, J.; Hong, Y.; Shen, X.; et al. OsCPK12 phosphorylates OsCATA and OsCATC to regulate H2O2 homeostasis and improve oxidative stress tolerance in rice. Plant Commun. 2024, 5, 100780. [Google Scholar] [CrossRef]
- Li, H.; Zhang, Y.; Wu, C.; Bi, J.; Chen, Y.; Jiang, C.; Cui, M.; Chen, Y.; Hou, X.; Yuan, M.; et al. Fine-tuning OsCPK18/OsCPK4 activity via genome editing of phosphorylation motif improves rice yield and immunity. Plant Biotechnol. J. 2022, 20, 2258–2271. [Google Scholar] [CrossRef]
- Chen, M.; Ni, L.; Chen, J.; Sun, M.; Qin, C.; Zhang, G.; Zhang, A.; Jiang, M. Rice calcium/calmodulin-dependent protein kinase directly phosphorylates a mitogen-activated protein kinase kinase to regulate abscisic acid responses. Plant Cell 2021, 33, 1790–1812. [Google Scholar] [CrossRef]
- Ahmad, S.; Wei, X.; Sheng, Z.; Hu, P.; Tang, S. CRISPR/Cas9 for development of disease resistance in plants: Recent progress, limitations and future prospects. Brief. Funct. Genom. 2020, 19, 26–39. [Google Scholar] [CrossRef]
- Gao, C.; Tang, D.; Wang, W. The Role of Ubiquitination in Plant Immunity: Fine-Tuning Immune Signaling and Beyond. Plant Cell Physiol. 2022, 63, 1405–1413. [Google Scholar] [CrossRef]
- Wang, C.; Wang, G.; Zhang, C.; Zhu, P.; Dai, H.; Yu, N.; He, Z.; Xu, L.; Wang, E. OsCERK1-Mediated Chitin Perception and Immune Signaling Requires Receptor-like Cytoplasmic Kinase 185 to Activate an MAPK Cascade in Rice. Mol. Plant 2017, 10, 619–633. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Wang, T.; Meng, G.; Liu, J.; Lu, D.; Liu, X.; Zeng, Y. Cytological observation and transcriptome analysis reveal dynamic changes of Rhizoctonia solani colonization on leaf sheath and different genes recruited between the resistant and susceptible genotypes in rice. Front. Plant Sci. 2022, 13, 599–612. [Google Scholar] [CrossRef]
- Niu, Y.; Huang, X.; He, Z.; Zhang, Q.; Meng, H.; Shi, H.; Feng, B.; Zhou, Y.; Zhang, J.; Lu, G.; et al. Phosphorylation of OsTGA5 by casein kinase II compromises its suppression of defense-related gene transcription in rice. Plant Cell 2022, 34, 3425–3442. [Google Scholar] [CrossRef]
- Ban, Z.; Estelle, M. CUL3 E3 ligases in plant development and environmental response. Nat. Plants 2021, 7, 6–16. [Google Scholar] [CrossRef] [PubMed]
- Younas, M.U.; Ahmad, I.; Qasim, M.; Ijaz, Z.; Rajput, N.; Parveen Memon, S.; ULZaman, W.; Jiang, X.; Zhang, Y.; Zuo, S. Progress in the Management of Rice Blast Disease: The Role of Avirulence and Resistance Genes through Gene-for-Gene Interactions. Agronomy 2024, 14, 163. [Google Scholar] [CrossRef]
- Zheng, Y.; Zhang, X.; Liu, Y.; Zhu, T.; Wu, X.; Ning, Y.; Liu, J.; Wang, D. Crystal structure of rice APIP6 reveals a new dimerization mode of RING-type E3 ligases that facilities the construction of its working model. Phytopathol. Res. 2023, 5, 31. [Google Scholar] [CrossRef]
- Wang, J.; Wang, R.; Fang, H.; Zhang, C.; Zhang, F.; Hao, Z.; You, X.; Shi, X.; Park, C.H.; Hua, K.; et al. Two VOZ transcription factors link an E3 ligase and an NLR immune receptor to modulate immunity in rice. Mol. Plant 2021, 14, 253–266. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Qiu, J.; Shen, Z.; Wang, C.; Jiang, N.; Shi, H.; Kou, Y. The E3 ubiquitin ligase OsRGLG5 targeted by the Magnaporthe oryzae effector AvrPi9 confers basal resistance against rice blast. Plant Commun. 2023, 4, 100626. [Google Scholar] [CrossRef]
- Gou, M.; Balint-Kurti, P.; Xu, M.; Yang, Q. Quantitative disease resistance: Multifaceted players in plant defense. Cell Res. 2023, 65, 594–610. [Google Scholar] [CrossRef]
- Wang, R.; You, X.; Zhang, C.; Fang, H.; Wang, M.; Zhang, F.; Kang, H.; Xu, X.; Liu, Z.; Wang, J.; et al. An ORFeome of rice E3 ubiquitin ligases for global analysis of the ubiquitination interactome. Genome Biol. 2022, 23, 154. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Li, S.; Chen, L.; Tian, H.; Chen, C.; Fu, Y.; Du, H.; Hu, Z.; Li, R.; Du, Y.; et al. E3 ubiquitin ligase OsPIE3 destabilises the B-lectin receptor-like kinase PID2 to control blast disease resistance in rice. New Phytol. 2023, 237, 1826–1842. [Google Scholar] [CrossRef]
- Wang, S.; Han, S.; Zhou, X.; Zhao, C.; Guo, L.; Zhang, J.; Liu, F.; Huo, Q.; Zhao, W.; Guo, Z.; et al. Phosphorylation and ubiquitination of OsWRKY31 are integral to OsMKK10-2-mediated defense responses in rice. Plant Cell 2023, 35, 2391–2412. [Google Scholar] [CrossRef]
- Ichimaru, K.; Yamaguchi, K.; Harada, K.; Nishio, Y.; Hori, M.; Ishikawa, K.; Inoue, H.; Shigeta, S.; Inoue, K.; Shimada, K.; et al. Cooperative regulation of PBI1 and MAPKs controls WRKY45 transcription factor in rice immunity. Nat. Commun. 2022, 13, 2397. [Google Scholar] [CrossRef] [PubMed]
- Son, S.; Song, G.; Nam, S.; Lee, J.; Hwang, D.-J.; Suh, E.-J.; Park, S.R. OsWRKY65 enhances immunity against fungal and bacterial pathogens in rice. Crop J. 2024, 12, 470–481. [Google Scholar] [CrossRef]
- Li, W.; Wang, K.; Chern, M.; Liu, Y.; Zhu, Z.; Liu, J.; Zhu, X.; Yin, J.; Ran, L.; Xiong, J.; et al. Sclerenchyma cell thickening through enhanced lignification induced by OsMYB30 prevents fungal penetration of rice leaves. New Phytol. 2020, 226, 1850–1863. [Google Scholar] [CrossRef]
- Qiu, J.; Xie, J.; Chen, Y.; Shen, Z.; Shi, H.; Naqvi, N.I.; Qian, Q.; Liang, Y.; Kou, Y. Warm temperature compromises JA-regulated basal resistance to enhance Magnaporthe oryzae infection in rice. Mol. Plant 2022, 15, 723–739. [Google Scholar] [CrossRef]
- Huang, L.; Ökmen, B.; Stolze, S.C.; Kastl, M.; Khan, M.; Hilbig, D.; Nakagami, H.; Djamei, A.; Doehlemann, G. The fungal pathogen Ustilago maydis targets the maize corepressor RELK2 to modulate host transcription for tumorigenesis. New Phytol. 2024, 241, 1747–1762. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Zhang, H.; Li, D.; Huang, L.; Hong, Y.; Ding, X.S.; Nelson, R.S.; Zhou, X.; Song, F. Functions of rice NAC transcriptional factors, ONAC122 and ONAC131, in defense responses against Magnaporthe grisea. Plant Mol. Biol. 2013, 81, 41–56. [Google Scholar] [CrossRef] [PubMed]
- Bui, H.; Dong, G.; Le, C.; Chu, H.; Duong, D.; Phung, L.; Phung, H.; Nguyen, C.; Nguyen, N.; Nguyen, P.; et al. Designing CRISPR/Cas9 System Targeting OsERF922 Gene of J02 Rice Variety. Asian J. Plant Sci. 2023, 22, 66–74. [Google Scholar] [CrossRef]
- Yang, X.; Zhang, L.; Yang, Y.; Schmid, M.; Wang, Y. miRNA Mediated Regulation and Interaction between Plants and Pathogens. Int. J. Mol. Sci. 2021, 22, 2913. [Google Scholar] [CrossRef]
- Li, Y.; Jeyakumar, J.M.J.; Feng, Q.; Zhao, Z.-X.; Fan, J.; Khaskheli, M.I.; Wang, W.-M. The roles of rice microRNAs in rice-Magnaporthe oryzae interaction. Phytopathol. Res. 2019, 1, 33. [Google Scholar] [CrossRef]
- Zhang, L.L.; Huang, Y.Y.; Zheng, Y.P.; Liu, X.X.; Zhou, S.X.; Yang, X.M.; Liu, S.L.; Li, Y.; Li, J.L.; Zhao, S.L.; et al. Osa-miR535 targets SQUAMOSA promoter binding protein-like 4 to regulate blast disease resistance in rice. Plant J. 2022, 110, 166–178. [Google Scholar] [CrossRef]
- Salvador-Guirao, R.; Hsing, Y.I.; San Segundo, B. The Polycistronic miR166k-166h Positively Regulates Rice Immunity via Post-transcriptional Control of EIN2. Front. Plant Sci. 2018, 9, 337. [Google Scholar] [CrossRef]
- Li, Y.; Tong, Y.; He, X.; Zhu, Y.; Li, T.; Lin, X.; Mao, W.; Gishkori, Z.G.N.; Zhao, Z.; Zhang, J.; et al. The rice miR171b–SCL6-IIs module controls blast resistance, grain yield, and flowering. Crop J. 2022, 10, 117–127. [Google Scholar] [CrossRef]
- Wang, H.; Li, Y.; Chern, M.; Zhu, Y.; Zhang, L.L.; Lu, J.H.; Li, X.P.; Dang, W.Q.; Ma, X.C.; Yang, Z.R.; et al. Suppression of rice miR168 improves yield, flowering time and immunity. Nat. Plants 2021, 7, 129–136. [Google Scholar] [CrossRef]
- Zhou, J.; Zhang, R.; Jia, X.; Tang, X.; Guo, Y.; Yang, H.; Zheng, X.; Qian, Q.; Qi, Y.; Zhang, Y. CRISPR-Cas9 mediated OsMIR168a knockout reveals its pleiotropy in rice. Plant Biotechnol. J. 2022, 20, 310–322. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, T.T.; He, X.R.; Zhu, Y.; Feng, Q.; Yang, X.M.; Zhou, X.H.; Li, G.B.; Ji, Y.P.; Zhao, J.H.; et al. Blocking Osa-miR1871 enhances rice resistance against Magnaporthe oryzae and yield. Plant Biotechnol. J. 2022, 20, 646–659. [Google Scholar] [CrossRef]
- Chandran, V.; Wang, H.; Gao, F.; Cao, X.L.; Chen, Y.P.; Li, G.B.; Zhu, Y.; Yang, X.M.; Zhang, L.L.; Zhao, Z.X.; et al. miR396-OsGRFs Module Balances Growth and Rice Blast Disease-Resistance. Front. Plant Sci. 2018, 9, 1999. [Google Scholar] [CrossRef] [PubMed]
- Barna, B.; Fodor, J.; Harrach, B.D.; Pogany, M.; Kiraly, Z. The Janus face of reactive oxygen species in resistance and susceptibility of plants to necrotrophic and biotrophic pathogens. Plant Physiol. Biochem. 2012, 59, 37–43. [Google Scholar] [CrossRef]
- Sha, G.; Sun, P.; Kong, X.; Han, X.; Sun, Q.; Fouillen, L.; Zhao, J.; Li, Y.; Yang, L.; Wang, Y.; et al. Genome editing of a rice CDP-DAG synthase confers multipathogen resistance. Nature 2023, 618, 1017–1023. [Google Scholar] [CrossRef]
- Meng, F.; Yang, C.; Cao, J.; Chen, H.; Pang, J.; Zhao, Q.; Wang, Z.; Qing Fu, Z.; Liu, J. A bHLH transcription activator regulates defense signaling by nucleo-cytosolic trafficking in rice. J. Integr. Plant Biol. 2020, 62, 1552–1573. [Google Scholar] [CrossRef]
- Hu, B.; Zhou, Y.; Zhou, Z.; Sun, B.; Zhou, F.; Yin, C.; Ma, W.; Chen, H.; Lin, Y. Repressed OsMESL expression triggers reactive oxygen species-mediated broad-spectrum disease resistance in rice. Plant Biotechnol. J. 2021, 19, 1511–1522. [Google Scholar] [CrossRef] [PubMed]
- Peng, X.; Hu, Y.; Tang, X.; Zhou, P.; Deng, X.; Wang, H.; Guo, Z. Constitutive expression of rice WRKY30 gene increases the endogenous jasmonic acid accumulation, PR gene expression and resistance to fungal pathogens in rice. Planta 2012, 236, 1485–1498. [Google Scholar] [CrossRef]
- Ying, W.; Liao, L.; Wei, H.; Gao, Y.; Liu, X.; Sun, L. Structural basis for abscisic acid efflux mediated by ABCG25 in Arabidopsis thaliana. Nat. Plants 2023, 9, 1697–1708. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.J.; Shimono, M.; Sugano, S.; Kojima, M.; Yazawa, K.; Yoshida, R.; Inoue, H.; Hayashi, N.; Sakakibara, H.; Takatsuji, H. Abscisic Acid Interacts Antagonistically with Salicylic Acid Signaling Pathway in Rice–Magnaporthe grisea Interaction. Mol. Plant Microbe Interact. 2010, 23, 791–798. [Google Scholar] [CrossRef]
- Tian, H.; Xu, L.; Li, X.; Zhang, Y. Salicylic acid: The roles in plant immunity and crosstalk with other hormones. J. Integr. Plant Biol. 2025, 67, 773–785. [Google Scholar] [CrossRef]
- Yang, Q.; He, Y.; Kabahuma, M.; Chaya, T.; Kelly, A.; Borrego, E.; Bian, Y.; El Kasmi, F.; Yang, L.; Teixeira, P.; et al. A gene encoding maize caffeoyl-CoA O-methyltransferase confers quantitative resistance to multiple pathogens. Nat. Genet. 2017, 49, 1364–1372. [Google Scholar] [CrossRef]
- Fang, H.; Zhang, F.; Zhang, C.; Wang, D.; Shen, S.; He, F.; Tao, H.; Wang, R.; Wang, M.; Wang, D.; et al. Function of hydroxycinnamoyl transferases for the biosynthesis of phenolamides in rice resistance to Magnaporthe oryzae. J. Genet. Genom. 2022, 49, 776–786. [Google Scholar] [CrossRef]
- Yang, C.; Liu, R.; Pang, J.; Ren, B.; Zhou, H.; Wang, G.; Wang, E.; Liu, J. Poaceae-specific cell wall-derived oligosaccharides activate plant immunity via OsCERK1 during Magnaporthe oryzae infection in rice. Nat. Commun. 2021, 12, 2178. [Google Scholar] [CrossRef] [PubMed]
- Shen, S.; Peng, M.; Fang, H.; Wang, Z.; Zhou, S.; Jing, X.; Zhang, M.; Yang, C.; Guo, H.; Li, Y.; et al. An Oryza-specific hydroxycinnamoyl tyramine gene cluster contributes to enhanced disease resistance. Sci. Bull. 2021, 66, 2369–2380. [Google Scholar] [CrossRef]
- Xu, X.-M.; Liang, K.-J.; Zhang, S.-G.; Shang, W.; Zhang, Y.-Y.; Wei, X.-Y.; Ke, B. Analysis of Indica-Japonica Differentiation in Rice Parents and Derived Lines Using ILP Markers. Agric. Sci. China 2009, 8, 1409–1418. [Google Scholar] [CrossRef]
- He, X.; Liu, X.; Wang, L.; Wang, L.; Lin, F.; Cheng, Y.; Chen, Z.; Liao, Y.; Pan, Q. Identification of the novel recessive gene pi55(t) conferring resistance to Magnaporthe oryzae. Sci. China Life Sci. 2012, 55, 141–149. [Google Scholar] [CrossRef][Green Version]
- Romero, F.M.; Gatica-Arias, A. CRISPR/Cas9: Development and Application in Rice Breeding. Rice Sci. 2019, 26, 265–281. [Google Scholar] [CrossRef]
- Xie, B.; Wang, L.; Li, H.; Huo, H.; Cui, C.; Sun, B.; Ma, Y.; Wang, J.; Yin, G.; Zuo, P. An interface-reinforced rhombohedral Prussian blue analogue in semi-solid state electrolyte for sodium-ion battery. Energy Storage Mater. 2021, 36, 99–107. [Google Scholar] [CrossRef]
- Zhao, C.; Wang, Y.; Li, Z.; Chen, W.; Xu, Q.; He, D.; Xi, D.; Zhang, Q.; Yuan, T.; Qu, Y.; et al. Solid-Diffusion Synthesis of Single-Atom Catalysts Directly from Bulk Metal for Efficient CO2 Reduction. Joule 2019, 3, 584–594. [Google Scholar] [CrossRef]
- Pedrozo, R.; Osakina, A.; Huang, Y.; Nicolli, C.P.; Wang, L.; Jia, Y. Status on Genetic Resistance to Rice Blast Disease in the Post-Genomic Era. Plants 2025, 14, 807. [Google Scholar] [CrossRef]
- Deng, Z.; Qin, P.; Liu, K.; Jiang, N.; Yan, T.; Zhang, X.; Fu, C.; He, G.; Wang, K.; Yang, Y. The Development of Multi-Resistant Rice Restorer Lines and Hybrid Varieties by Pyramiding Resistance Genes against Blast and Brown Planthopper. Agronomy 2024, 14, 878. [Google Scholar] [CrossRef]
- Qin, P.; Jiang, N.; Hu, X.; Zhu, Y.; Deng, Z.; Bai, Z.; Fu, J.; Zhang, X.; Wang, K.; Chen, S.; et al. Breeding a series of new rice restorer lines with blast resistance and low-to-intermediate apparent amylose contents by integrating marker-assisted and phenotypic selection. J. Phytopathol. 2021, 169, 667–677. [Google Scholar] [CrossRef]
- Kumar, M.; Singh, R.P.; Jena, D.; Singh, V.; Rout, D.; Arsode, P.B.; Choudhary, M.; Singh, P.; Chahar, S.; Samantaray, S.; et al. Marker-Assisted Improvement for Durable Bacterial Blight Resistance in Aromatic Rice Cultivar HUR 917 Popular in Eastern Parts of India. Plants 2023, 12, 1363. [Google Scholar] [CrossRef]
- Ye, J.; Wang, K.; Wang, Y.; Zhao, Z.; Yan, Y.; Yang, H.; Zhang, L.; Hu, Z.; Shi, Z.; Sun, D.; et al. Improving panicle blast resistance and fragrance in a high-quality japonica rice variety through breeding. Front. Plant Sci. 2024, 15, 1507827. [Google Scholar] [CrossRef]
- Mao, T.; Zhu, M.; Ahmad, S.; Ye, G.; Sheng, Z.; Hu, S.; Jiao, G.; Xie, L.; Tang, S.; Wei, X.; et al. Superior japonica rice variety YJ144 with improved rice blast resistance, yield, and quality achieved using molecular design and multiple breeding strategies. Mol. Breed. 2021, 41, 65. [Google Scholar] [CrossRef]
- Wang, K.; Fu, C.; Fu, X.; Qin, P.; Hu, X.; Zhang, X.; Deng, Z.; Yan, T.; Jiang, N.; Li, Y.; et al. Enhancing the blast resistance of an elite thermo-sensitive genic male sterile line (TGMS) Longke638S and its derived hybrid varieties by incorporating Pigm gene. Mol. Breed. 2025, 45, 35. [Google Scholar] [CrossRef]
- Qi, Z.; Meng, X.; Xu, M.; Du, Y.; Yu, J.; Song, T.; Pan, X.; Zhang, R.; Cao, H.; Yu, M.; et al. A novel Pik allele confers extended resistance to rice blast. Plant Cell Environ. 2024, 12, 4800–4814. [Google Scholar] [CrossRef] [PubMed]
- Meuwissen, T.H.E.; Hayes, B.J.; Goddard, M.E. Prediction of total genetic value using genome-wide dense marker maps. Genetics 2001, 157, 1819–1829. [Google Scholar] [CrossRef]
- Sasaki, T. The map-based sequence of the rice genome. Nature 2005, 436, 793–800. [Google Scholar] [CrossRef]
- Ma, X.; Wang, H.; Yan, S.; Zhou, C.; Zhou, K.; Zhang, Q.; Li, M.; Yang, Y.; Li, D.; Song, P.; et al. Large-scale genomic and phenomic analyses of modern cultivars empower future rice breeding design. Mol. Plant 2025, 18, 651–668. [Google Scholar] [CrossRef] [PubMed]
- Xiao, N.; Pan, C.; Li, Y.; Wu, Y.; Cai, Y.; Lu, Y.; Wang, R.; Yu, L.; Shi, W.; Kang, H.; et al. Genomic insight into balancing high yield, good quality, and blast resistance of japonica rice. Genome Biol. 2021, 22, 283. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Bu, Q.; Liu, G.; Wang, M.; Wang, H.; Liu, H.; Li, X.; Li, H.; Fang, J.; Liang, Y.; et al. Genomic decoding of breeding history to guide breeding-by-design in rice. Natl. Sci. Rev. 2023, 10, nwad029. [Google Scholar] [CrossRef]
- Yu, S.; Ali, J.; Zhou, S.; Ren, G.; Xie, H.; Xu, J.; Yu, X.; Zhou, F.; Peng, S.; Ma, L.; et al. From Green Super Rice to green agriculture: Reaping the promise of functional genomics research. Mol. Plant 2022, 15, 9–26. [Google Scholar] [CrossRef]
- He, Z.; Xin, Y.; Wang, C.; Yang, H.; Xu, Z.; Cheng, J.; Li, Z.; Ye, C.; Yin, H.; Xie, Z.; et al. Genomics-Assisted improvement of super high-yield hybrid rice variety “super 1000” for resistance to bacterial blight and blast diseases. Front. Plant Sci. 2022, 13, 881244. [Google Scholar] [CrossRef]
- Feng, Z.; Gao, P.; Wang, G.; Kang, H.; Zhao, J.; Xie, W.; Chen, R.; Ju, R.; Wang, X.; Wei, Z.; et al. Natural variation in SBRR1 shows high potential for sheath blight resistance breeding in rice. Nat. Genet. 2025, 57, 2004–2016. [Google Scholar] [CrossRef]
- Greenwood, J.R.; Lacorte-Apostol, V.; Kroj, T.; Padilla, J.; Telebanco-Yanoria, M.J.; Glaus, A.N.; Roulin, A.; Padilla, A.; Zhou, B.; Keller, B.; et al. Genome-wide association analysis uncovers rice blast resistance alleles of Ptr and Pia. Commun. Biol. 2024, 7, 607. [Google Scholar] [CrossRef]
- Zhang, Y.; Lin, X.-F.; Li, L.; Piao, R.-H.; Wu, S.; Song, A.; Gao, M.; Jin, Y.-M. CRISPR/Cas9-mediated knockout of Bsr-d1 enhances the blast resistance of rice in Northeast China. Plant Cell Rep. 2024, 43, 100. [Google Scholar] [CrossRef]
- Li, G.; Gao, Q.; Li, W.; Zhang, W.; Wang, J.; Chen, B.; Wang, D.; Gao, H.; Xu, W.; Chen, H.; et al. Directed Knockout of SD1 Gene Improves Lodging Resistance and Blast Resistance of Rice. Chin. J. Rice Sci. 2023, 37, 359–367. [Google Scholar] [CrossRef]
- Tao, H.; Shi, X.; He, F.; Wang, D.; Xiao, N.; Fang, H.; Wang, R.; Zhang, F.; Wang, M.; Li, A.; et al. Engineering broad-spectrum disease-resistant rice by editing multiple susceptibility genes. J. Integr. Plant Biol. 2021, 63, 1639–1648. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Xu, S.; Jiang, N.; Zhao, X.; Bai, Z.; Liu, J.; Yao, W.; Tang, Q.; Xiao, G.; Lv, C.; et al. Engineering of rice varieties with enhanced resistances to both blast and bacterial blight diseases via CRISPR/Cas9. Plant Biotechnol. J. 2022, 20, 876–885. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhou, Z.; Xiong, X.; Li, C.; Li, C.; Shen, E.; Wang, J.; Zha, W.; Wu, B.; Chen, H.; et al. Development of a multi-resistance and high-yield rice variety using multigene transformation and gene editing. Plant Biotechnol. J. 2024, 22, 3118–3120. [Google Scholar] [CrossRef]
- Xu, P.; Wang, H.; Tu, R.; Liu, Q.; Wu, W.; Fu, X.; Cao, L.; Shen, X. Orientation Improvement of Blast Resistance in Rice via CRISPR/Cas9 System. Chin. J. Rice Sci. 2019, 33, 313–322. [Google Scholar] [CrossRef]
- Xu, L.; Tang, Y.; Wang, M.; Peng, X.; Shi, H.; Zhang, Z.; Zhang, Y.; Yin, J.; Xiong, Q.; Lu, X.; et al. U-box E3 ubiquitin ligase OsPUB41 represses rice blast resistance by promoting degradation of OsPALs. New Phytol. 2025, 247, 2312–2327. [Google Scholar] [CrossRef]
- Jung, Y.J.; Kim, J.Y.; Cho, Y.G.; Kang, K.K. CRISPR/Cas9-Mediated Knockout of OsbZIP76 Reveals Its Role in ABA-Associated Immune Signaling in Rice. Int. J. Mol. Sci. 2025, 26, 6374. [Google Scholar] [CrossRef]
- Bezerra, C. Science Uses Gene Editing to Develop Blast-Resistant Rice. Available online: https://www.seedtoday.com/article/1030135/science-uses-gene-editing-to-develop-blast-resistant-rice (accessed on 20 August 2025).
- Yu, T.; Sun, M.K.; Liang, L.K. Receptors in the Induction of the Plant Innate Immunity. Mol. Plant Microbe Interact. 2021, 34, 587–601. [Google Scholar] [CrossRef]
- Golicz, A.A.; Bayer, P.E.; Bhalla, P.L.; Batley, J.; Edwards, D. Pangenomics Comes of Age: From Bacteria to Plant and Animal Applications. Trends Genet. 2020, 36, 132–145. [Google Scholar] [CrossRef] [PubMed]
- Tettelin, H.; Masignani, V.; Cieslewicz, M.J.; Donati, C.; Medini, D.; Ward, N.L.; Angiuoli, S.V.; Crabtree, J.; Jones, A.L.; Durkin, A.S.; et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: Implications for the microbial “pan-genome”. Proc. Natl. Acad. Sci. USA 2005, 102, 13950–13955. [Google Scholar] [CrossRef] [PubMed]
- Shang, L.; Li, X.; He, H.; Yuan, Q.; Song, Y.; Wei, Z.; Lin, H.; Hu, M.; Zhao, F.X.; Zhang, C.; et al. A super pan-genomic landscape of rice. Cell Res. 2022, 32, 878–896. [Google Scholar] [CrossRef]
- Watson, A.; Ghosh, S.; Williams, M.J.; Cuddy, W.S.; Simmonds, J.; Rey, M.D.; Asyraf Md Hatta, M.; Hinchliffe, A.; Steed, A.; Reynolds, D.; et al. Speed breeding is a powerful tool to accelerate crop research and breeding. Nat. Plants 2018, 4, 23–29. [Google Scholar] [CrossRef]
- Jiang, S.; Li, H.; Zhang, L.; Mu, W.; Zhang, Y.; Chen, T.; Wu, J.; Tang, H.; Zheng, S.; Liu, Y.; et al. Generic Diagramming Platform (GDP). A comprehensive database of high-quality biomedical graphics. Nucleic Acids Res. 2025, 53, 1670–1676. [Google Scholar] [CrossRef]
- Feng, Q.; Wang, H.; Yang, X.; Hu, Z.; Zhou, X.; Xiang, L.; Xiong, X.; He, X.; Zhu, Y.; Li, G.; et al. Osa-miR160a confers broad-spectrum resistance to fungal and bacterial pathogens in rice. New Phytol. 2022, 236, 2216–2232. [Google Scholar] [CrossRef]
- Maidment, J.; Shimizu, M.; Bentham, A.; Vera, S.; Franceschetti, M.; Longya, A.; Stevenson, C.; De la Concepcion, J.; Białas, A.; Kamoun, S.; et al. Effector target-guided engineering of an integrated domain expands the disease resistance profile of a rice NLR immune receptor. Elife 2023, 12, e81123. [Google Scholar] [CrossRef]
- Hayashi, K.; Yasuda, N.; Fujita, Y.; Koizumi, S.; Yoshida, H. Identification of the blast resistance gene Pit in rice cultivars using functional markers. Theor. Appl. Genet. 2010, 121, 1357–1367. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, A.; Hayashi, N.; Miyao, A.; Hirochika, H. Unique features of the rice blast resistance Pish locus revealed by large scale retrotransposon-tagging. BMC Plant Biol. 2010, 10, 175. [Google Scholar] [CrossRef] [PubMed]
- Fukuoka, S.; Yamamoto, S.-I.; Mizobuchi, R.; Yamanouchi, U.; Ono, K.; Kitazawa, N.; Yasuda, N.; Fujita, Y.; Thi Thanh Nguyen, T.; Koizumi, S.; et al. Multiple functional polymorphisms in a single disease resistance gene in rice enhance durable resistance to blast. Sci. Rep. 2014, 4, 4550. [Google Scholar] [CrossRef]
- Lin, F.; Chen, S.; Que, Z.; Wang, L.; Liu, X.; Pan, Q. The blast resistance gene Pi37 encodes a nucleotide binding site leucine-rich repeat protein and is a member of a resistance gene cluster on rice chromosome 1. Genetics 2007, 177, 1871–1880. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Wang, L.; Wu, W.; He, L.; Yang, X.; Pan, Q. Function and evolution of Magnaporthe oryzae avirulence gene AvrPib responding to the rice blast resistance gene Pib. Sci Rep 2015, 5, 11642. [Google Scholar] [CrossRef]
- Chen, H.; Wang, S.; Xing, Y.; Xu, C.; Hayes, P.M.; Zhang, Q. Comparative analyses of genomic locations and race specificities of loci for quantitative resistance to Pyricularia grisea in rice and barley. Proc. Natl. Acad. Sci. USA 2003, 100, 2544–2549. [Google Scholar] [CrossRef]
- Xu, X.; Hayashi, N.; Wang, C.-T.; Fukuoka, S.; Kawasaki, S.; Takatsuji, H.; Jiang, C.-J. Rice blast resistance gene Pikahei-1(t), a member of a resistance gene cluster on chromosome 4, encodes a nucleotide-binding site and leucine-rich repeat protein. Mol. Breed. 2014, 34, 691–700. [Google Scholar] [CrossRef]
- Inukai, T.; Nagashima, S.; Kato, M. Pid3-I1 is a race-specific partial-resistance allele at the Pid3 blast resistance locus in rice. Theor. Appl. Genet. 2019, 132, 395–404. [Google Scholar] [CrossRef]
- Chen, Z.; Zhao, W.; Zhu, X.; Zou, C.; Yin, J.; Chern, M.; Zhou, X.; Ying, H.; Jiang, X.; Li, Y.; et al. Identification and characterization of rice blast resistance gene Pid4 by a combination of transcriptomic profiling and genome analysis. J. Genet. Genom. 2018, 45, 663–672. [Google Scholar] [CrossRef]
- Lv, Q.; Xu, X.; Shang, J.; Jiang, G.; Pang, Z.; Zhou, Z.; Wang, J.; Liu, Y.; Li, T.; Li, X.; et al. Functional analysis of Pid3-A4, an ortholog of rice blast resistance gene Pid3 revealed by allele mining in common wild rice. Phytopathology 2013, 103, 594–599. [Google Scholar] [CrossRef]
- Chen, J.; Shi, Y.; Liu, W.; Chai, R.; Fu, Y.; Zhuang, J.; Wu, J. A Pid3 allele from rice cultivar Gumei2 confers resistance to Magnaporthe oryzae. J. Genet. Genom. 2011, 38, 209–216. [Google Scholar] [CrossRef]
- Zhou, B.; Qu, S.; Liu, G.; Dolan, M.; Sakai, H.; Lu, G.; Bellizzi, M.; Wang, G.-L. The eight amino-acid differences within three leucine-rich repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to Magnaporthe grisea. Mol. Plant Microbe Interact. 2006, 19, 1216–1228. [Google Scholar] [CrossRef]
- Su, J.; Wang, W.; Han, J.; Chen, S.; Wang, C.; Zeng, L.; Feng, A.; Yang, J.; Zhou, B.; Zhu, X. Functional divergence of duplicated genes results in a novel blast resistance gene Pi50 at the Pi2/9 locus. Theor. Appl. Genet. 2015, 128, 2213–2225. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Zhai, K.; Xie, Z.; Yang, D.; Zhu, X.; Liu, J.; Wang, X.; Qin, P.; Yang, Y.; Zhang, G.; et al. Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance. Science 2017, 355, 962–965. [Google Scholar] [CrossRef]
- Xie, Z.; Yan, B.; Shou, J.; Tang, J.; Wang, X.; Zhai, K.; Liu, J.; Li, Q.; Luo, M.; Deng, Y.; et al. A nucleotide-binding site-leucine-rich repeat receptor pair confers broad-spectrum disease resistance through physical association in rice. Philos. Trans. R Soc. Lond. B Biol. Sci. 2019, 374, 20180308. [Google Scholar] [CrossRef]
- Liu, X.; Lin, F.; Wang, L.; Pan, Q. The in silico map-based cloning of Pi36, a rice coiled-coil nucleotide-binding site leucine-rich repeat gene that confers race-specific resistance to the blast fungus. Genetics 2007, 176, 2541–2549. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.K.; Song, M.Y.; Seo, Y.S.; Kim, H.K.; Ko, S.; Cao, P.J.; Suh, J.P.; Yi, G.; Roh, J.H.; Lee, S.; et al. Rice Pi5-mediated resistance to Magnaporthe oryzae requires the presence of two coiled-coil-nucleotide-binding-leucine-rich repeat genes. Genetics 2009, 181, 1627–1638. [Google Scholar] [CrossRef] [PubMed]
- Zhai, C.; Zhang, Y.; Yao, N.; Lin, F.; Liu, Z.; Dong, Z.; Wang, L.; Pan, Q. Function and interaction of the coupled genes responsible for Pik-h encoded rice blast resistance. PLoS ONE 2014, 9, e98067. [Google Scholar] [CrossRef]
- Das, A.; Soubam, D.; Singh, P.K.; Thakur, S.; Singh, N.K.; Sharma, T.R. A novel blast resistance gene, Pi54rh cloned from wild species of rice, Oryza rhizomatis confers broad spectrum resistance to Magnaporthe oryzae. Funct. Integr. Genom. 2012, 12, 215–228. [Google Scholar] [CrossRef]
- Devanna, N.B.; Vijayan, J.; Sharma, T.R. The blast resistance gene Pi54of cloned from Oryza officinalis interacts with Avr-Pi54 through its novel non-LRR domains. PLoS ONE 2014, 9, e104840. [Google Scholar] [CrossRef]
- Okuyama, Y.; Kanzaki, H.; Abe, A.; Yoshida, K.; Tamiru, M.; Saitoh, H.; Fujibe, T.; Matsumura, H.; Shenton, M.; Galam, D.C.; et al. A multifaceted genomics approach allows the isolation of the rice Pia-blast resistance gene consisting of two adjacent NBS-LRR protein genes. Plant J. 2011, 66, 467–479. [Google Scholar] [CrossRef]
- Zhai, C.; Lin, F.; Dong, Z.; He, X.; Yuan, B.; Zeng, X.; Wang, L.; Pan, Q. The isolation and characterization of Pik, a rice blast resistance gene which emerged after rice domestication. New Phytol. 2011, 189, 321–334. [Google Scholar] [CrossRef]
- Chen, J.; Peng, P.; Tian, J.; He, Y.; Zhang, L.; Liu, Z.; Yin, D.; Zhang, Z. Pike, a rice blast resistance allele consisting of two adjacent NBS–LRR genes, was identified as a novel allele at the Pik locus. Mol. Breed. 2015, 35, 117. [Google Scholar] [CrossRef]
- Hua, L.; Wu, J.; Chen, C.; Wu, W.; He, X.; Lin, F.; Wang, L.; Ashikawa, I.; Matsumoto, T.; Wang, L.; et al. The isolation of Pi1, an allele at the Pik locus which confers broad spectrum resistance to rice blast. Theor. Appl. Genet. 2012, 125, 1047–1055. [Google Scholar] [CrossRef]
- Yuan, B.; Zhai, C.; Wang, W.; Zeng, X.; Xu, X.; Hu, H.; Lin, F.; Wang, L.; Pan, Q. The Pik-p resistance to Magnaporthe oryzae in rice is mediated by a pair of closely linked CC-NBS-LRR genes. Theor. Appl. Genet. 2011, 122, 1017–1028. [Google Scholar] [CrossRef] [PubMed]
- Ashikawa, I.; Hayashi, N.; Yamane, H.; Kanamori, H.; Wu, J.; Matsumoto, T.; Ono, K.; Yano, M. Two adjacent nucleotide-binding site-leucine-rich repeat class genes are required to confer Pikm-specific rice blast resistance. Genetics 2008, 180, 2267–2276. [Google Scholar] [CrossRef] [PubMed]
- Sharma, T.R.; Madhav, M.S.; Singh, B.K.; Shanker, P.; Jana, T.K.; Dalal, V.; Pandit, A.; Singh, A.; Gaikwad, K.; Upreti, H.C.; et al. High-resolution mapping, cloning and molecular characterization of the Pi-kh gene of rice, which confers resistance to Magnaporthe grisea. Mol. Genet. Genom. 2005, 274, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; He, Y.; Chen, J.; Long, X.; Wang, H.; Zhu, M.; Liu, S.; Cai, Q.; Zhang, Z. Analysis of natural variation of the rice blast resistance gene Pike and identification of a novel allele Pikg. Mol. Genet. Genom. 2021, 296, 939–952. [Google Scholar] [CrossRef]
- Bryan, G.T.; Wu, K.-S.; Farrall, L.; Jia, Y.; Hershey, H.P.; McAdams, S.A.; Faulk, K.N.; Donaldson, G.K.; Tarchini, R.; Valent, B. A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta. Plant Cell 2000, 11, 2033–2046. [Google Scholar] [CrossRef]
- Lin, J.; Song, X.; Wu, M.; Ruan, G. Breeding and Utilization of Japonicalinous CMS Line Chunjiang 19A with Wide Compatibility and High Outcrossing Rate. Agron. For. 2012, 13, 537–540. [Google Scholar] [CrossRef]
- Yang, X.; Li, Y.; Ren, B.; Ding, L.; Gao, C.; Shen, Q.; Guo, S. Drought-induced root aerenchyma formation restricts water uptake in rice seedlings supplied with nitrate. Plant Cell Physiol. 2012, 53, 495–504. [Google Scholar] [CrossRef]
- Liu, G.; Zhu, P.; Liu, Y.; Kong, D.; Wang, J.; Luo, L.; Yu, X. Molecular Marker-Assisted Selection of a New Water-Saving and Drought-Resistant Rice (WDR) Restoration Line, Hanhui 8200, for Enhanced Resistance to Rice Blast. Agronomy 2024, 14, 1504. [Google Scholar] [CrossRef]
- Cui, Y.; Li, C.; Zhou, F.; Chen, H.; Zhang, W.; Ma, W.; Lin, Y. Developing of transgenic glyphosate-tolerant Indica restorer line with commercial application potential. Mol. Breed. 2020, 40, 43. [Google Scholar] [CrossRef]
- Hori, K.; Yamamoto, T.; Yano, M. Genetic dissection of agronomically important traits in closely related temperate japonica rice cultivars. Breed. Sci. 2017, 67, 427–434. [Google Scholar] [CrossRef] [PubMed]
- Hayano-Saito, Y.; Hayashi, K. Stvb-i, a Rice Gene Conferring Durable Resistance to Rice stripe virus, Protects Plant Growth From Heat Stress. Front. Plant Sci. 2020, 11, 519. [Google Scholar] [CrossRef]
- Sedeek, S.; Aboyousef, M.; El-Rafaee, I.; Abdallah, A.; Hammoud, S.; El-Malky, M.; El-Namkay, R.; El-Abd, A.; Ammar, M.; Abdelkhalik, A.; et al. Giza 183 Egyptian rice variety: A step to confront climate change challenges. Egypt. J. Agric. Res. 2023, 101, 519–537. [Google Scholar] [CrossRef]
- Wang, Y.; Fu, X.; Gao, G.; He, Y. Improving the Grain Quality of Minghui63, a Restorer Line of Rice with Good Quality through Marker-Assisted Selection. Mol. Plant Breed. 2009, 7, 661–665. [Google Scholar] [CrossRef]
- Wang, G.L.; Mackill, D.J.; Bonman, J.M.; McCouch, S.R.; Champoux, M.C.; Nelson, R.J. RFLP mapping of genes conferring complete and partial resistance to blast in a durably resistant rice cultivar. Genetics 1994, 136, 1421–1434. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Singh, V.K.; Singh, S.P.; Pandian, R.T.P.; Ellur, R.K.; Singh, D.; Bhowmick, P.K.; Krishnan, S.G.; Nagarajan, M.; Vinod, K.K.; et al. Molecular breeding for the development of multiple disease resistance in Basmati rice. AoB Plants 2012, 2012, pls029. [Google Scholar] [CrossRef]
- Yao, X.F.; Sun, Z.; Xu, X.; Li, H.; Liu, C.M. Complete Genome Assembly of Japonica Rice Variety Zhonghua 11. Plant Commun. 2025, 25, 101463. [Google Scholar] [CrossRef] [PubMed]
- Hasan, M.M.; Rafii, M.Y.; Ismail, M.R.; Mahmood, M.; Rahim, H.A.; Alam, M.A.; Ashkani, S.; Malek, M.A.; Latif, M.A. Marker-assisted backcrossing: A useful method for rice improvement. Biotechnology 2015, 29, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Jia, Y.; Wamishe, Y.; Jia, M.H.; Valent, B. Dynamic Changes in the Rice Blast Population in the United States Over Six Decades. Mol. Plant Microbe Interact. 2017, 30, 803–812. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Lian, B.; Hu, J.; Zhai, H.; Wang, X.; Venu, R.; Liu, E.; Wang, Z.; Chen, M.; Wang, B.; et al. Genome comparison of two Magnaporthe oryzae field isolates reveals genome variations and potential virulence effectors. BMC Genom. 2013, 14, 887. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cheng, B.; Lv, B.; Xuan, Q.; Li, Y.; Li, J.; Liang, W.; Wang, J. Research Progress on Rice-Blast-Resistance-Related Genes. Plants 2025, 14, 2698. https://doi.org/10.3390/plants14172698
Cheng B, Lv B, Xuan Q, Li Y, Li J, Liang W, Wang J. Research Progress on Rice-Blast-Resistance-Related Genes. Plants. 2025; 14(17):2698. https://doi.org/10.3390/plants14172698
Chicago/Turabian StyleCheng, Biaobiao, Beibei Lv, Qiangbing Xuan, Yunfang Li, Jing Li, Weihong Liang, and Junjie Wang. 2025. "Research Progress on Rice-Blast-Resistance-Related Genes" Plants 14, no. 17: 2698. https://doi.org/10.3390/plants14172698
APA StyleCheng, B., Lv, B., Xuan, Q., Li, Y., Li, J., Liang, W., & Wang, J. (2025). Research Progress on Rice-Blast-Resistance-Related Genes. Plants, 14(17), 2698. https://doi.org/10.3390/plants14172698






