Litchi LcAP1-1 and LcAP1-2 Exhibit Different Roles in Flowering Time
Abstract
1. Introduction
2. Result
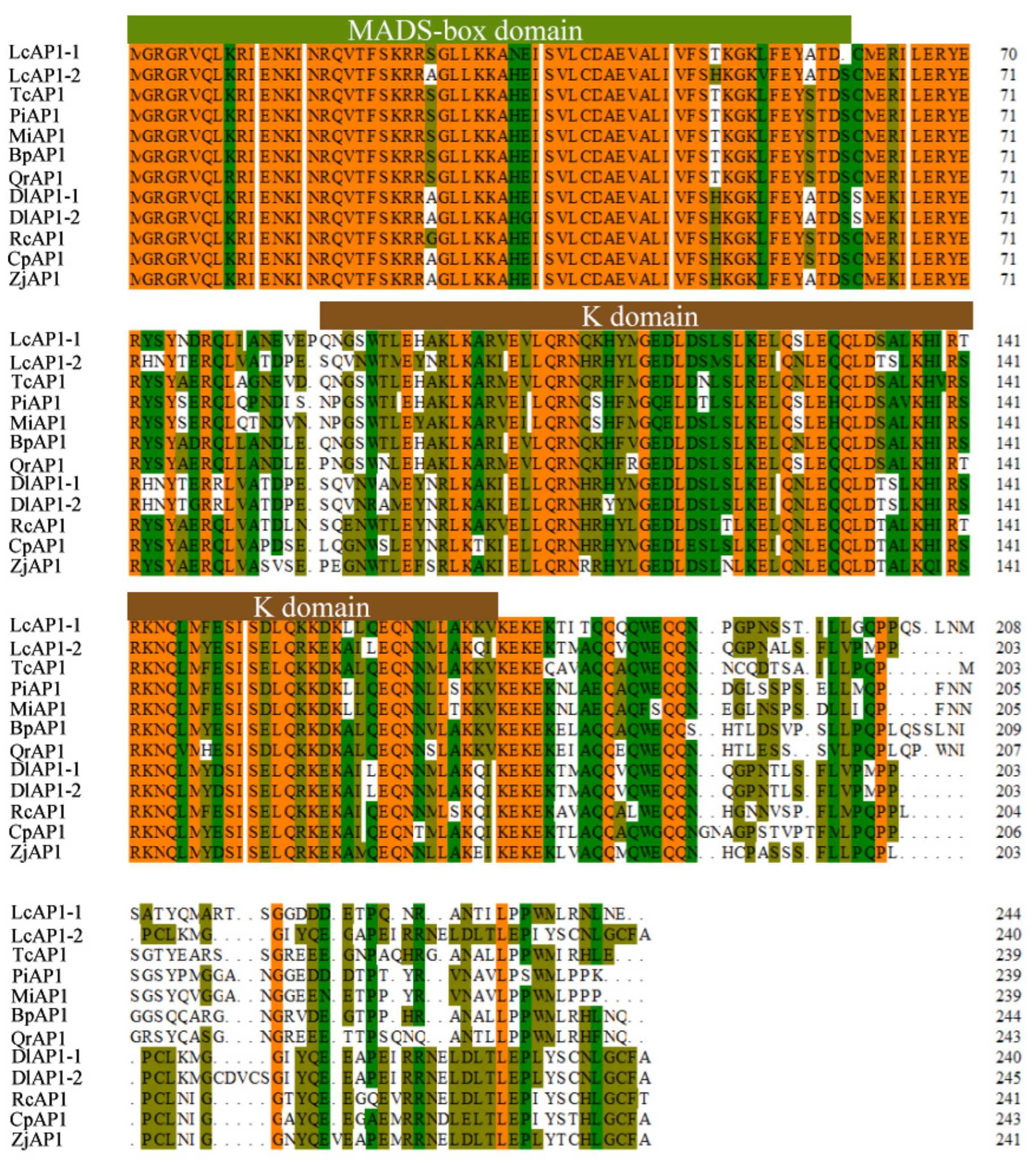
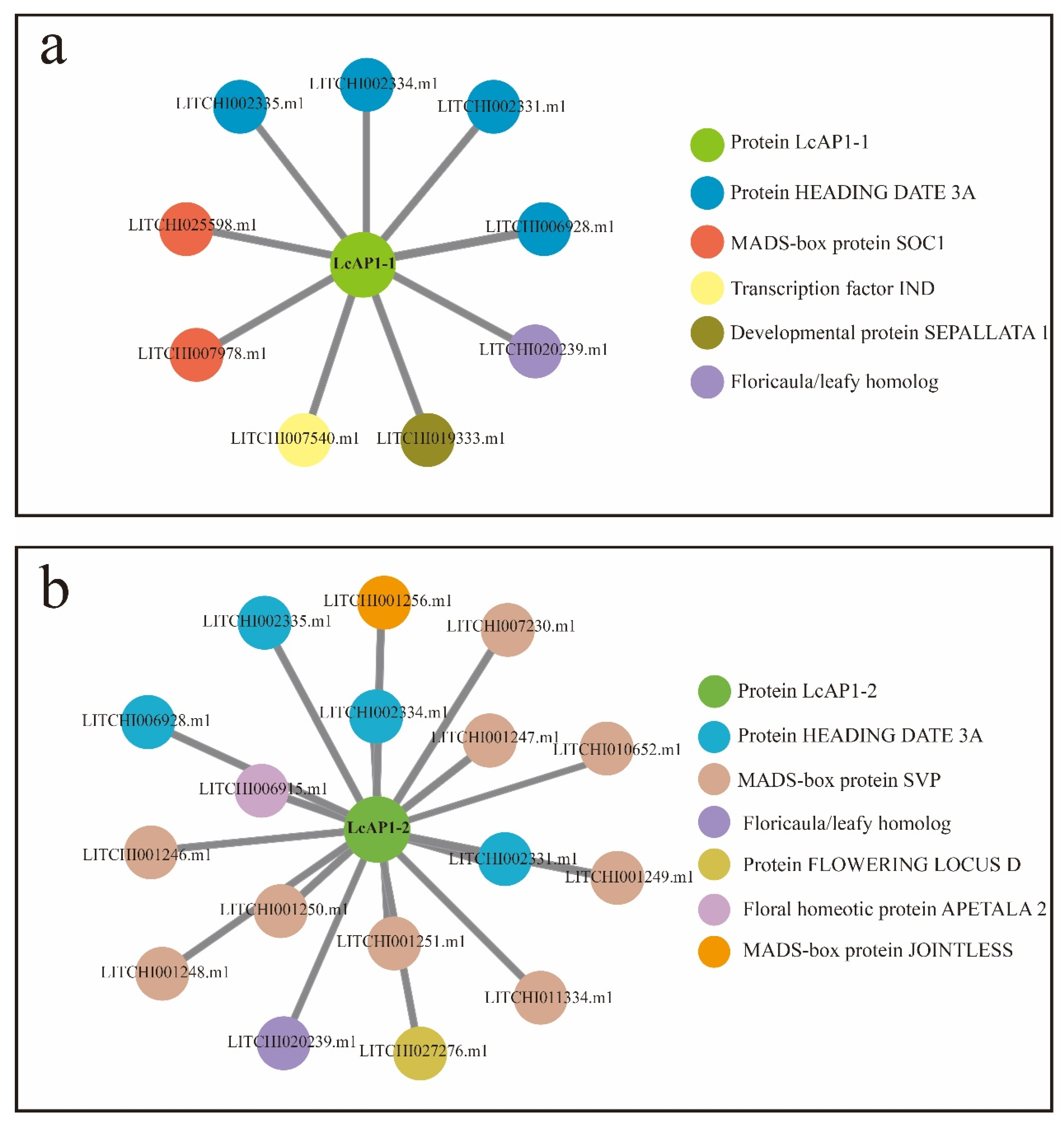
2.1. Isolation and Phylogenetic Analysis of LcAP1-1 and LcAP1-2 Genes in Litchi
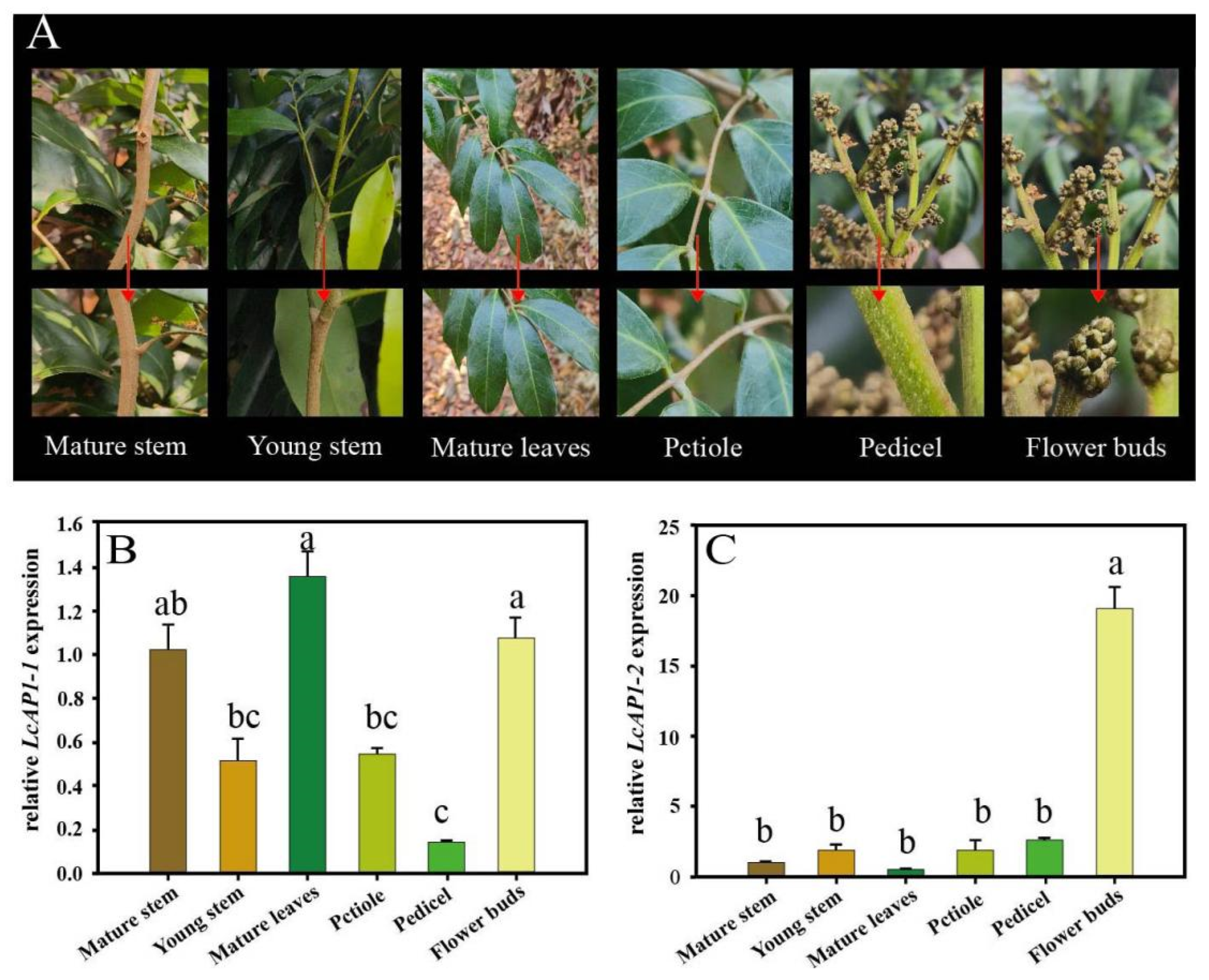
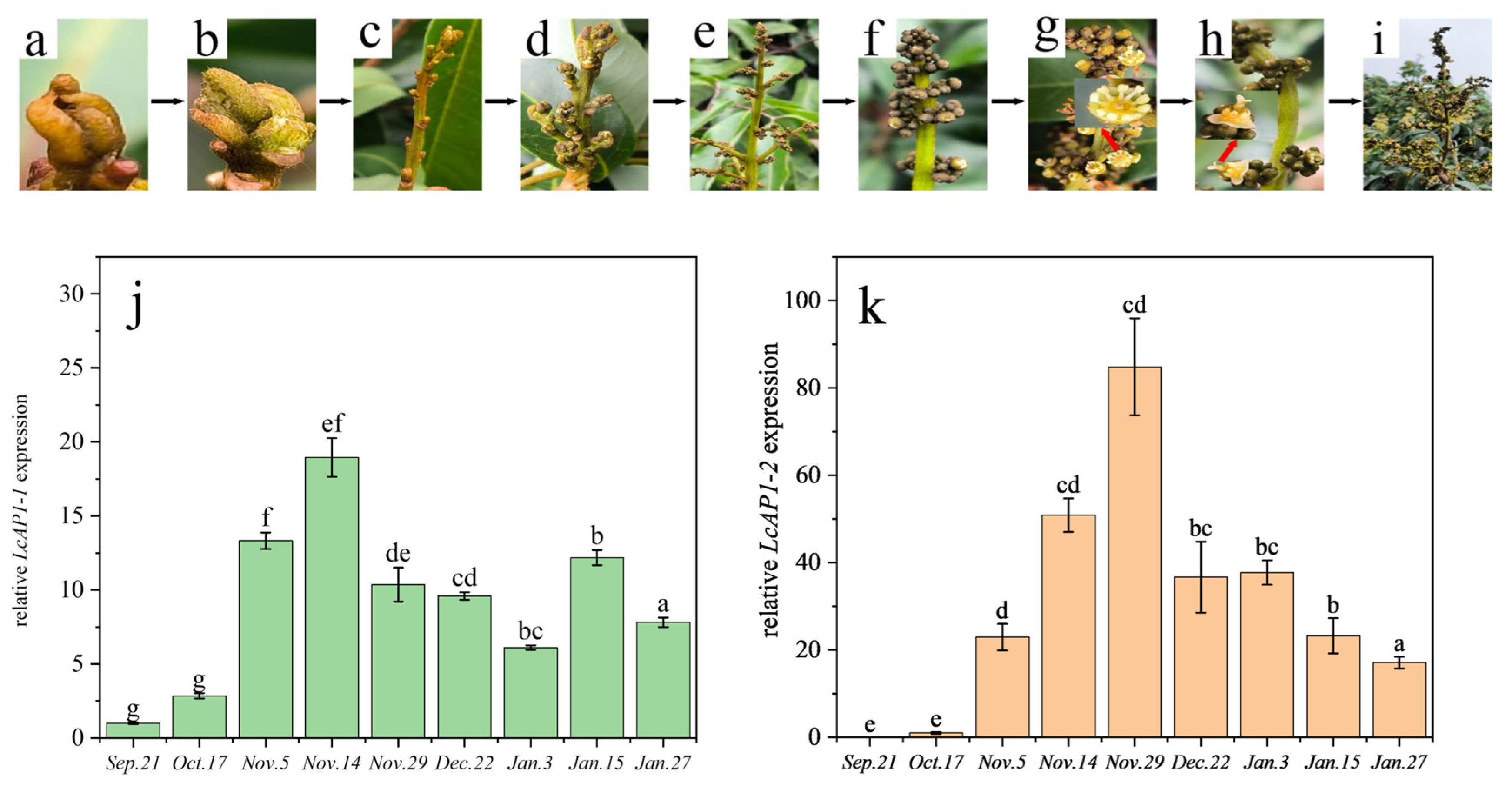
2.2. Expression Pattern Analysis of LcAP1-1 and LcAP1-2 Genes in Litchi
2.3. LcAP1-1 and LcAP1-2 Genes Transformation in Tobacco
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Cloning of LcAP1 Gene
4.3. Quantitative Real-Time PCR (qRT-PCR) Analysis
4.4. Overexpression of LcAP1-1 and LcAP1-2 Genes in Tobacco
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Maple, R.; Zhu, P.; Hepworth, J.; Wang, J.-W.; Dean, C. Flowering Time: From Physiology, through Genetics to Mechanism. Plant Physiol. 2024, 195, 190–212. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Zhang, S.; Gui, Q.; Qing, H.; Li, M.; Yi, C.; Guo, H.; Chen, H.; Xu, J.; Ding, F. The SOC1 Gene Plays an Important Role in Regulating Litchi Flowering Time. Genomics 2024, 116, 110804. [Google Scholar] [CrossRef]
- Rehman, S.; Bahadur, S.; Xia, W. An Overview of Floral Regulatory Genes in Annual and Perennial Plants. Gene 2023, 885, 147699. [Google Scholar] [CrossRef]
- Schwarz-Sommer, Z.; Huijser, P.; Nacken, W.; Saedler, H.; Sommer, H. Genetic Control of Flower Development by Homeotic Genes in Antirrhinum majus. Science 1990, 250, 931–936. [Google Scholar] [CrossRef]
- Cho, S.; Jang, S.; Chae, S.; Chung, K.M.; Moon, Y.-H.; Jang, S.K. Analysis of the C-Terminal Region of Arabidopsis Thaliana APETALA1 as a Transcription Activation Domain. Plant Mol. Biol. 1999, 40, 419–429. [Google Scholar] [CrossRef]
- Mandel, M.A.; Gustafson-Brown, C.; Savidge, B.; Yanofsky, M.F. Molecular Characterization of the Arabidopsis floral Homeotic Gene APETALA1. Nature 1992, 360, 273–277. [Google Scholar] [CrossRef] [PubMed]
- Theißen, G.; Gramzow, L. Chapter 8-Structure and Evolution of Plant MADS Domain Transcription Factors. In Plant Transcription Factors; Elsevier: Amsterdam, The Netherlands, 2016; pp. 127–138. ISBN 978-0-12-800854-6. [Google Scholar]
- Adrian, J.; Torti, S.; Turck, F. From Decision to Commitment: The Molecular Memory of Flowering. Mol. Plant 2009, 2, 628–642. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Thong, Z.; Yu, H. Coming into Bloom: The Specification of Floral Meristems. Development 2009, 136, 3379–3391. [Google Scholar] [CrossRef]
- Yant, L.; Mathieu, J.; Schmid, M. Just Say No: Floral Repressors Help Arabidopsis Bide the Time. Curr. Opin. Plant Biol. 2009, 12, 580–586. [Google Scholar] [CrossRef]
- Irish, V.F. The Flowering of Arabidopsis Flower Development. Plant J. 2010, 61, 1014–1028. [Google Scholar] [CrossRef]
- Kaufmann, K.; Nagasaki, M.; Jáuregui, R. Modelling the Molecular Interactions in the Flower Developmental Network of Arabidopsis thaliana. Silico Biol. 2010, 10, 125–143. [Google Scholar] [CrossRef]
- Ding, F.; Li, H.; Wang, J.; Peng, H.; Chen, H.; Hu, F.; Lai, B.; Wei, Y.; Ma, W.; Li, H.; et al. Development of Molecular Markers Based on the Promoter Difference of LcFT1 to Discriminate Easy- and Difficult-Flowering Litchi Germplasm Resources and Its Application in Crossbreeding. BMC Plant Biol. 2021, 21, 539. [Google Scholar] [CrossRef]
- Wagner, D.; Sablowski, R.W.M.; Meyerowitz, E.M. Transcriptional Activation of APETALA1 by LEAFY. Science 1999, 285, 582–584. [Google Scholar] [CrossRef]
- Goslin, K.; Zheng, B.; Serrano-Mislata, A.; Rae, L.; Ryan, P.T.; Kwaśniewska, K.; Thomson, B.; Ó’Maoiléidigh, D.S.; Madueño, F.; Wellmer, F.; et al. Transcription Factor Interplay between LEAFY and APETALA1/CAULIFLOWER during Floral Initiation. Plant Physiol. 2017, 174, 1097–1109. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, N.; Winter, C.M.; Wu, M.-F.; Kanno, Y.; Yamaguchi, A.; Seo, M.; Wagner, D. Gibberellin Acts Positively Then Negatively to Control Onset of Flower Formation in Arabidopsis. Science 2014, 344, 638–641. [Google Scholar] [CrossRef]
- Zuo, X.; Xiang, W.; Zhang, L.; Gao, C.; An, N.; Xing, L.; Ma, J.; Zhao, C.; Zhang, D. Identification of Apple TFL1-Interacting Proteins Uncovers an Expanded Flowering Network. Plant Cell Rep. 2021, 40, 2325–2340. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Meng, Y.; Cao, Q.; Wada, M.; Liang, A. Molecular Cloning and Expression Analysis of a SQUA/AP1 Homologue in Chinese Jujube (Ziziphus jujube Mill.). Plant Mol. Biol. Report. 2009, 27, 534–541. [Google Scholar] [CrossRef]
- Liu, Y.; Song, H.; Liu, Z.; Hu, G.; Lin, S. Molecular Characterization of Loquat EjAP1 Gene in Relation to Flowering. Plant Growth Regul. 2013, 70, 287–296. [Google Scholar] [CrossRef]
- Jing, D.; Chen, W.; Hu, R.; Zhang, Y.; Xia, Y.; Wang, S.; He, Q.; Guo, Q.; Liang, G. An Integrative Analysis of Transcriptome, Proteome and Hormones Reveals Key Differentially Expressed Genes and Metabolic Pathways Involved in Flower Development in Loquat. Int. J. Mol. Sci. 2020, 21, 5107. [Google Scholar] [CrossRef]
- Zhang, S.; Wang, X.; Xu, Z.; Shi, P.; Gu, M.; Kang, T.; Li, Q.; Zhang, D.; Zhao, C. PrupeFUL4 Regulates Ripening and Softening of Peach Fruits through Ethylene Biosynthesis. Acta Physiol. Plant. 2022, 44, 23. [Google Scholar] [CrossRef]
- Song, G.; Chen, Q. Comparative Transcriptome Analysis of Nonchilled, Chilled, and Late-Pink Bud Reveals Flowering Pathway Genes Involved in Chilling-Mediated Flowering in Blueberry. BMC Plant Biol. 2018, 18, 98. [Google Scholar] [CrossRef]
- Wang, J.; Zhang, X.; Yan, G.; Zhou, Y.; Zhang, K. Over-Expression of the PaAP1 Gene from Sweet Cherry (Prunus avium L.) Causes Early Flowering in Arabidopsis thaliana. J. Plant Physiol. 2013, 170, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Winterhagen, P.; Tiyayon, P.; Samach, A.; Hegele, M.; Wünsche, J.N. Isolation and Characterization of FLOWERING LOCUS T Subforms and APETALA1 of the Subtropical Fruit Tree Dimocarpus longan. Plant Physiol. Biochem. 2013, 71, 184–190. [Google Scholar] [CrossRef]
- Li, S.; Zhang, T.; Zang, L.; Qu, G. Transformation of Tobacco with Two AP1 Genes Isolated from Poplar (Populus simonii × Populus nigra). Agric. Biotechnol. 2014, 3, 23–27. [Google Scholar]
- Sun, L.-M.; Zhang, J.-Z.; Mei, L.; Hu, C.-G. Molecular Cloning, Promoter Analysis and Functional Characterization of APETALA 1-like Gene from Precocious Trifoliate Orange (Poncirus trifoliata L. Raf.). Sci. Hortic. 2014, 178, 95–105. [Google Scholar] [CrossRef]
- Yu, H.; Luo, C.; Fan, Y.; Zhang, X.; Huang, F.; Li, M.; He, X. Isolation and Characterization of Two APETALA1-like Genes from Mango (Mangifera indica L.). Sci. Hortic. 2020, 259, 108814. [Google Scholar] [CrossRef]
- Orbović, V.; Ravanfar, S.A.; Acanda, Y.; Narvaez, J.; Merritt, B.A.; Levy, A.; Lovatt, C.J. Stress-Inducible Arabidopsis thaliana RD29A Promoter Constitutively Drives Citrus sinensis APETALA1 and LEAFY Expression and Precocious Flowering in Transgenic Citrus spp. Transgenic Res. 2021, 30, 687–699. [Google Scholar] [CrossRef]
- Li, S.; Zhang, X.; Zhang, L.; Hong, J.; Chen, Y. Effects of H2O2 and NO on Flower Development and AP1 Gene Expression of Off-Season Longan. Agric. Biotechnol. 2016, 5, 2164–4993. [Google Scholar]
- Li, S.; Chen, H.; Hong, J.; Ye, X.; Wang, J.; Chen, Y.; Zhang, L.; Su, Z.; Yang, Z. Chlorate-Induced Molecular Floral Transition Revealed by Transcriptomes. Open Life Sci. 2023, 18, 20220612. [Google Scholar] [CrossRef]
- Chen, H.; He, J.; Li, W.; Chen, Y.; Gai, J. Expression of AP1 Gene during Offseason Flower Bud Differentiation of Mango. Agric. Biotechnol. 2016, 5, 2164–4993. [Google Scholar]
- Hu, G.; Feng, J.; Xiang, X.; Wang, J.; Salojärvi, J.; Liu, C.; Wu, Z.; Zhang, J.; Liang, X.; Jiang, Z.; et al. Two Divergent Haplotypes from a Highly Heterozygous Lychee Genome Suggest Independent Domestication Events for Early and Late-Maturing Cultivars. Nat. Genet. 2022, 54, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Cronje, R.B.; Mostert, P.G. A Holistic Approach to Improve Litchi Orchard Management in South Africa. Acta Hortic. 2010, 2010, 433–436. [Google Scholar] [CrossRef]
- Wigge, P.A.; Kim, M.C.; Jaeger, K.E.; Busch, W.; Schmid, M.; Lohmann, J.U.; Weigel, D. Integration of Spatial and Temporal Information during Floral Induction in Arabidopsis. Science 2005, 309, 1056–1059. [Google Scholar] [CrossRef]
- Corbesier, L.; Vincent, C.; Jang, S.; Fornara, F.; Fan, Q.; Searle, I.; Giakountis, A.; Farrona, S.; Gissot, L.; Turnbull, C.; et al. FT Protein Movement Contributes to Long-Distance Signaling in Floral Induction of Arabidopsis. Science 2007, 316, 1030–1033. [Google Scholar] [CrossRef]
- Ding, F.; Zhang, S.; Chen, H.; Su, Z.; Zhang, R.; Xiao, Q.; Li, H. Promoter Difference of LcFT1 Is a Leading Cause of Natural Variation of Flowering Timing in Different Litchi Cultivars (Litchi chinensis Sonn.). Plant Sci. 2015, 241, 128–137. [Google Scholar] [CrossRef]
- Ding, F.; Zhang, S.; Chen, H.; Peng, H.; Lu, J.; He, X.; Pan, J. Functional Analysis of a Homologue of the FLORICAULA/LEAFY Gene in Litchi (Litchi chinensis Sonn.) Revealing Its Significance in Early Flowering Process. Genes Genom. 2018, 40, 1259–1267. [Google Scholar] [CrossRef]
- Han, Y.; Tang, A.; Yu, J.; Cheng, T.; Wang, J.; Yang, W.; Pan, H.; Zhang, Q. RcAP1, a Homolog of APETALA1, Is Associated with Flower Bud Differentiation and Floral Organ Morphogenesis in Rosa chinensis. Int. J. Mol. Sci. 2019, 20, 3557. [Google Scholar] [CrossRef]
- Ng, M.; Yanofsky, M.F. Activation of the Arabidopsis B Class Homeotic Genes by APETALA. Plant Cell. 2001, 13, 739–753. [Google Scholar]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the Sensitivity of Progressive Multiple Sequence Alignment through Sequence Weighting, Position-Specific Gap Penalties and Weight Matrix Choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Holsters, M.; De Waele, D.; Depicker, A.; Messens, E.; Van Montagu, M.; Schell, J. Transfection and Transformation of Agrobacterium tumefaciens. Molec. Gen. Genet. 1978, 163, 181–187. [Google Scholar] [CrossRef] [PubMed]





| Plant Traits at Flowering | WT | OE-LcAP1-1 | OE-LcAP1-2 |
|---|---|---|---|
| The time of the first flower (day) | 135 ± 2.6 | 132 ± 1.5 | 80 ± 1.3 ** |
| Plant height at flowering (cm) | 82 ± 3.1 | 78 ± 2.4 | 42 ± 2.2 ** |
| Number of leaves at flowering | 24 ± 0.8 | 22 ± 0.6 | 14 ± 1.2 ** |
| Primer ID | Sequences 5′-3′ |
|---|---|
| KLLcAP1-1_F1 | GGTGAAGGAAAAGGAGAAGACA |
| KLLcAP1-1_R1 | CCCTAAAAGAATAGTGGACGAGTT |
| KLLcAP1-2_F1 | GCAGCAACAACAACAACTACCT |
| KLLcAP1-2_R1 | CCACAAATGGAAAACCTGAAGA |
| Lcactin_F1 | ACCGTATGAGCAAGGAAATCACTG |
| Lcactin_R1 | TCGTCGTACTCACCCTTTGAAATC |
| DLLcAP1-1_F2 | CACGGGGGACTCTAGAATGGGGAGAGGTCGGGTGCAGCT |
| DLLcAP1-1_R2 | AGGGACTGACCACCCGGGTTATTCGTTAAGGTTGCGAAGCA |
| DLLcAP1-2_F2 | CACGGGGGACTCTAGAATGGGGAGAGGTAGGGTTCAGTT |
| DLLcAP1-2_R2 | AGGGACTGACCACCCGGGTCATGTAGCGAAACATCCAAGAT |
| 35S_F | GCACAATCCCACTATCCTTCG |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gui, Q.; Wei, J.; Wu, Z.; Mo, X.; Qing, H.; Shi, Y.; Guo, H.; Sheng, J.; Ding, F.; Zhang, S. Litchi LcAP1-1 and LcAP1-2 Exhibit Different Roles in Flowering Time. Plants 2025, 14, 2697. https://doi.org/10.3390/plants14172697
Gui Q, Wei J, Wu Z, Mo X, Qing H, Shi Y, Guo H, Sheng J, Ding F, Zhang S. Litchi LcAP1-1 and LcAP1-2 Exhibit Different Roles in Flowering Time. Plants. 2025; 14(17):2697. https://doi.org/10.3390/plants14172697
Chicago/Turabian StyleGui, Qiulin, Jinju Wei, Ziang Wu, Xiao Mo, Haowei Qing, Yuyu Shi, Huiqin Guo, Jingwen Sheng, Feng Ding, and Shuwei Zhang. 2025. "Litchi LcAP1-1 and LcAP1-2 Exhibit Different Roles in Flowering Time" Plants 14, no. 17: 2697. https://doi.org/10.3390/plants14172697
APA StyleGui, Q., Wei, J., Wu, Z., Mo, X., Qing, H., Shi, Y., Guo, H., Sheng, J., Ding, F., & Zhang, S. (2025). Litchi LcAP1-1 and LcAP1-2 Exhibit Different Roles in Flowering Time. Plants, 14(17), 2697. https://doi.org/10.3390/plants14172697





