Genome-Wide Analysis of the AT-Hook Gene Family in Malus sieversii and Functional Characterization of MsAHL13
Abstract
1. Introduction
2. Result
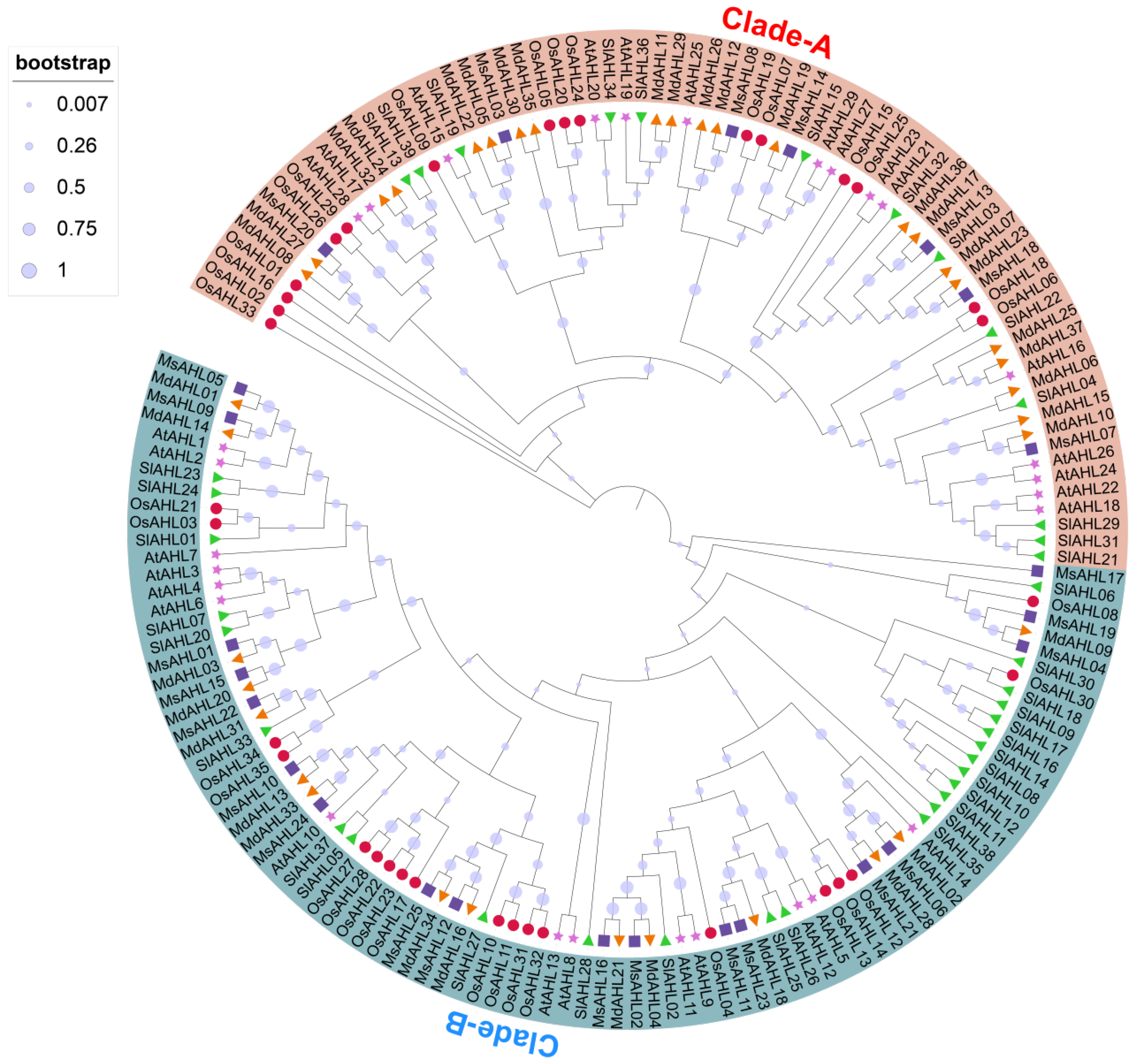
2.1. Identification and Phylogenetic Analysis of Members of the AHL Gene Family in M. sieversii
2.2. Analysis of MsAHL Gene Structure, Motifs, and Domains
2.3. Chromosome Mapping of MsAHLs
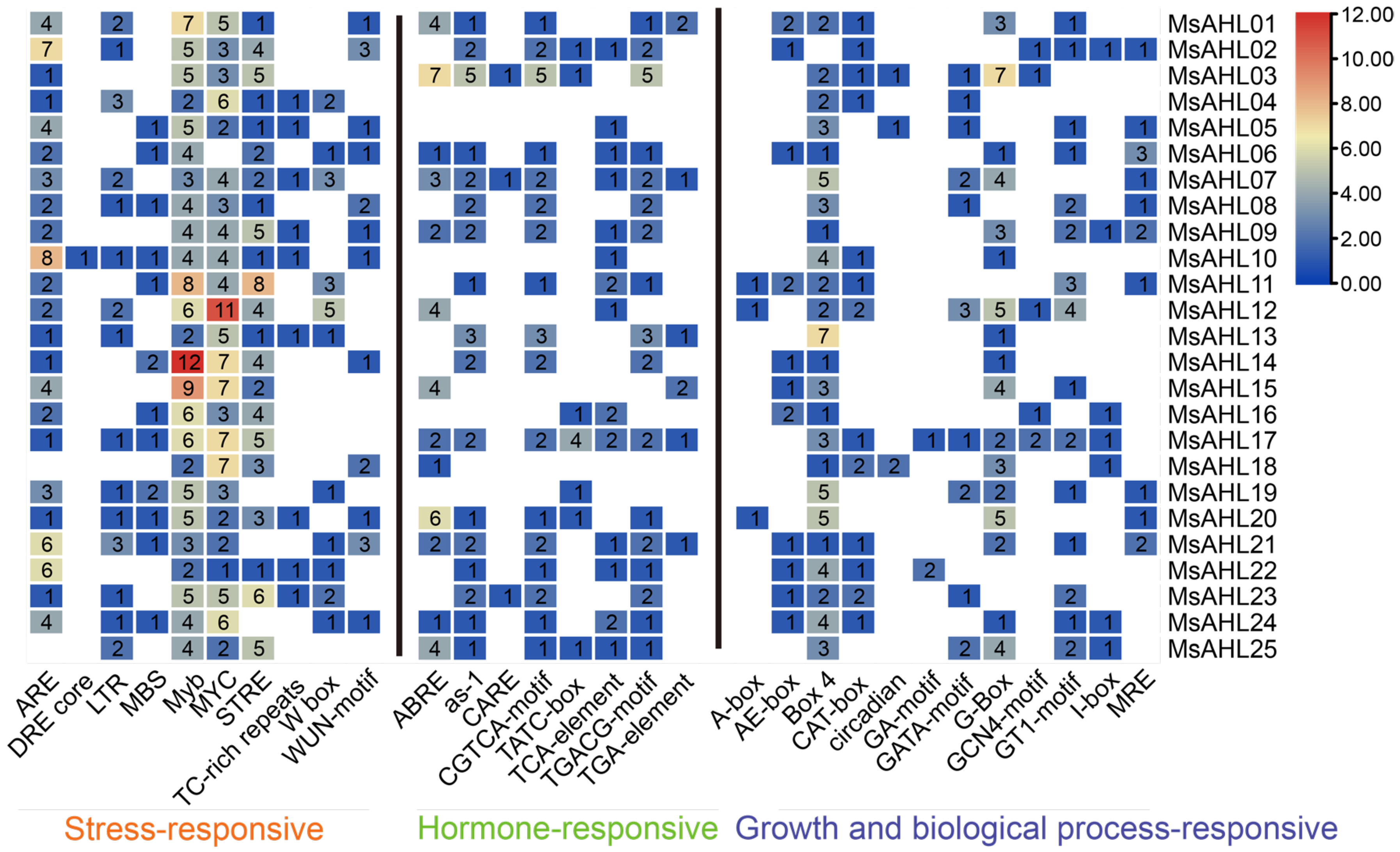
2.4. Analysis of Cis-Acting Elements
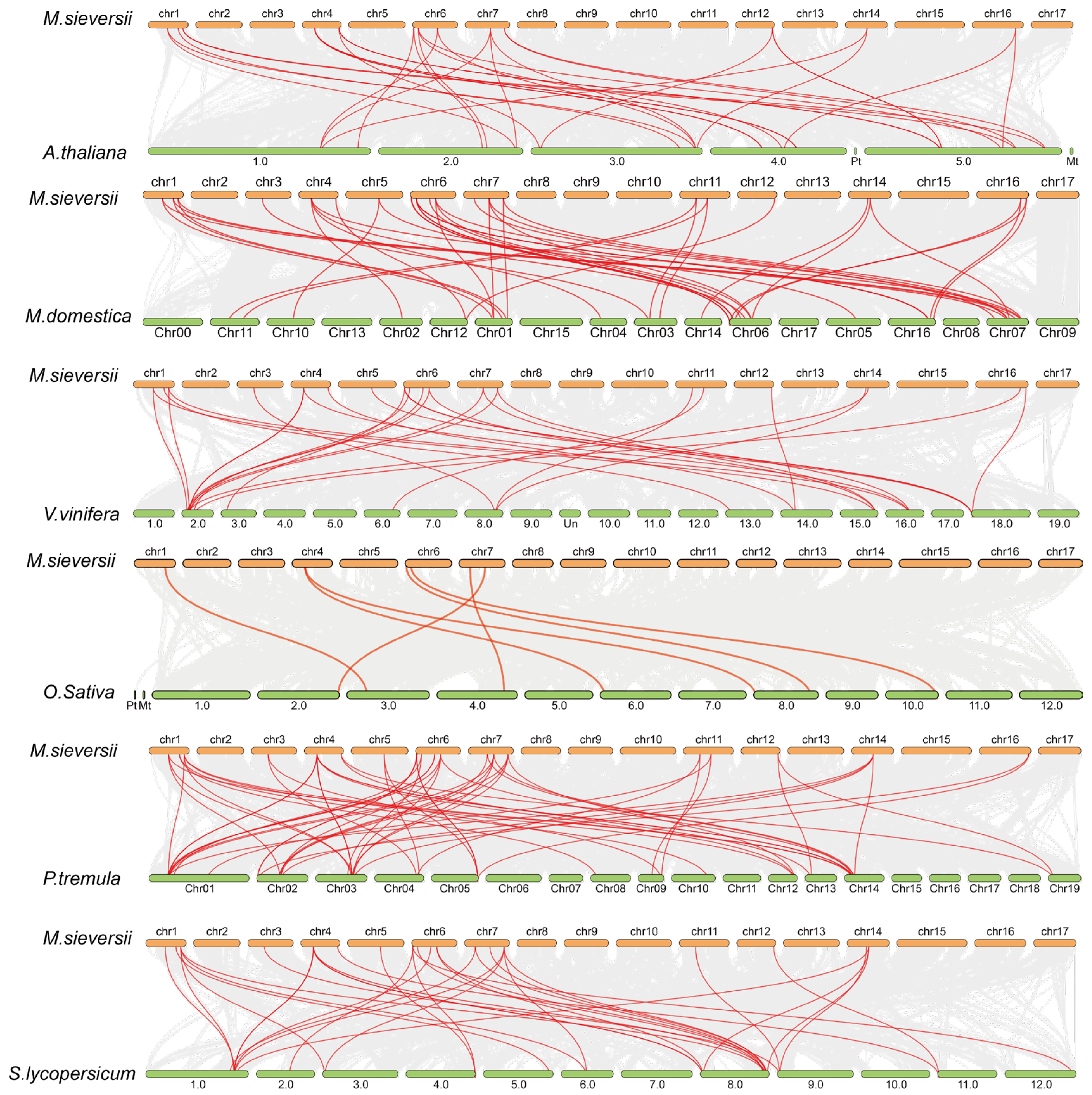
2.5. Collinearity Analysis of MsAHLs
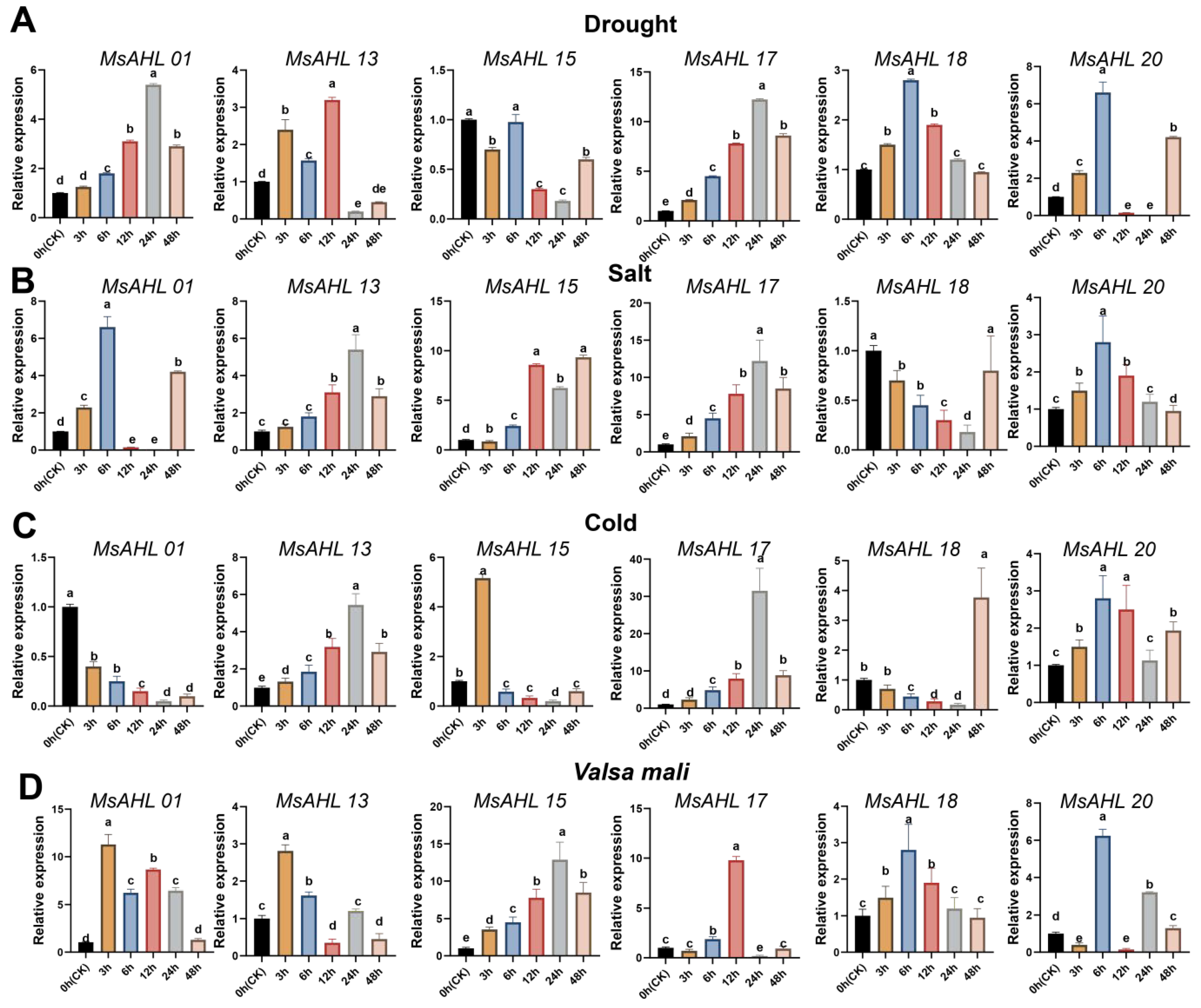
2.6. Expression Analysis of Six MsAHLs
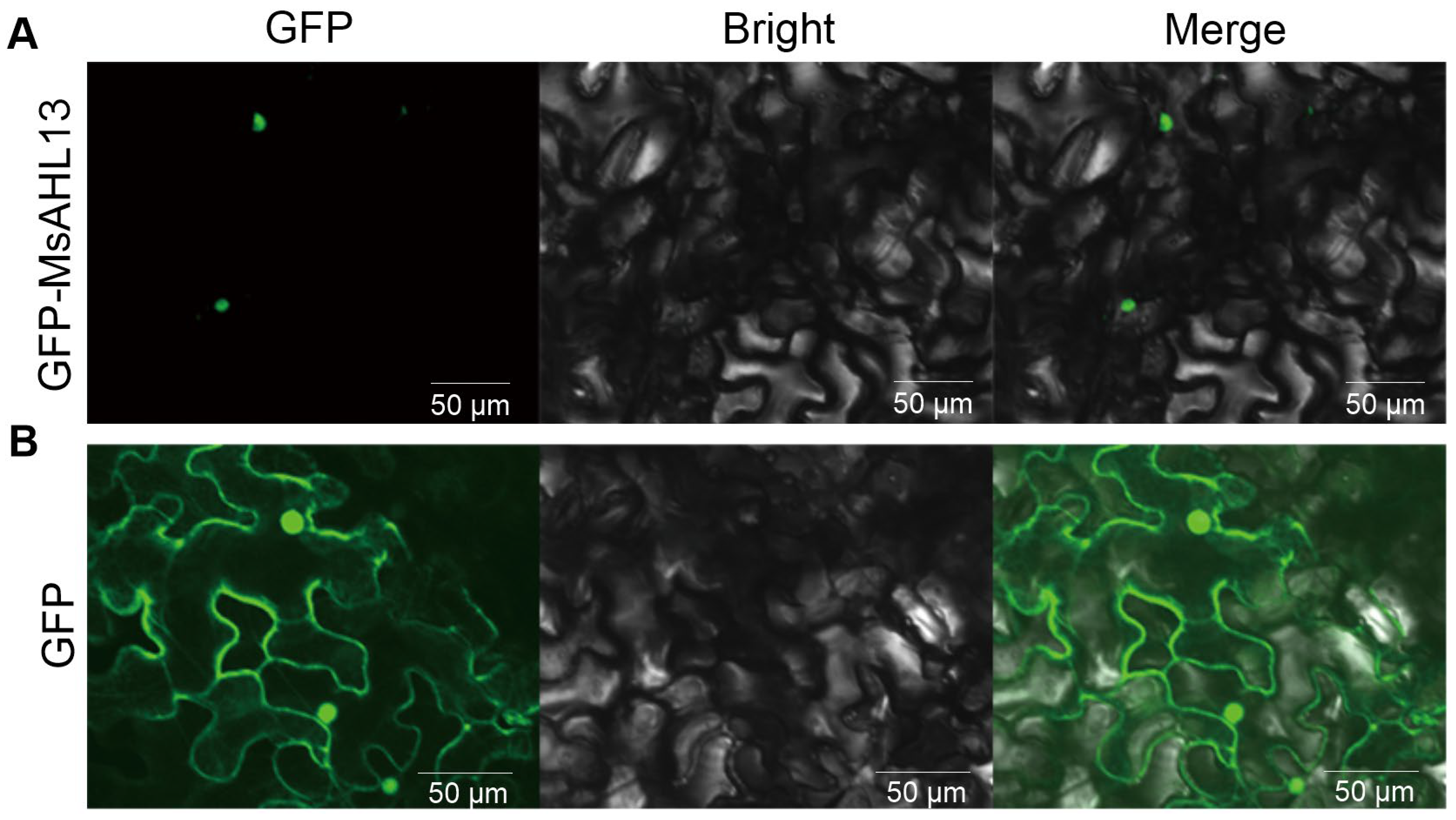
2.7. MsAHL13 Is Localized to the Nucleus
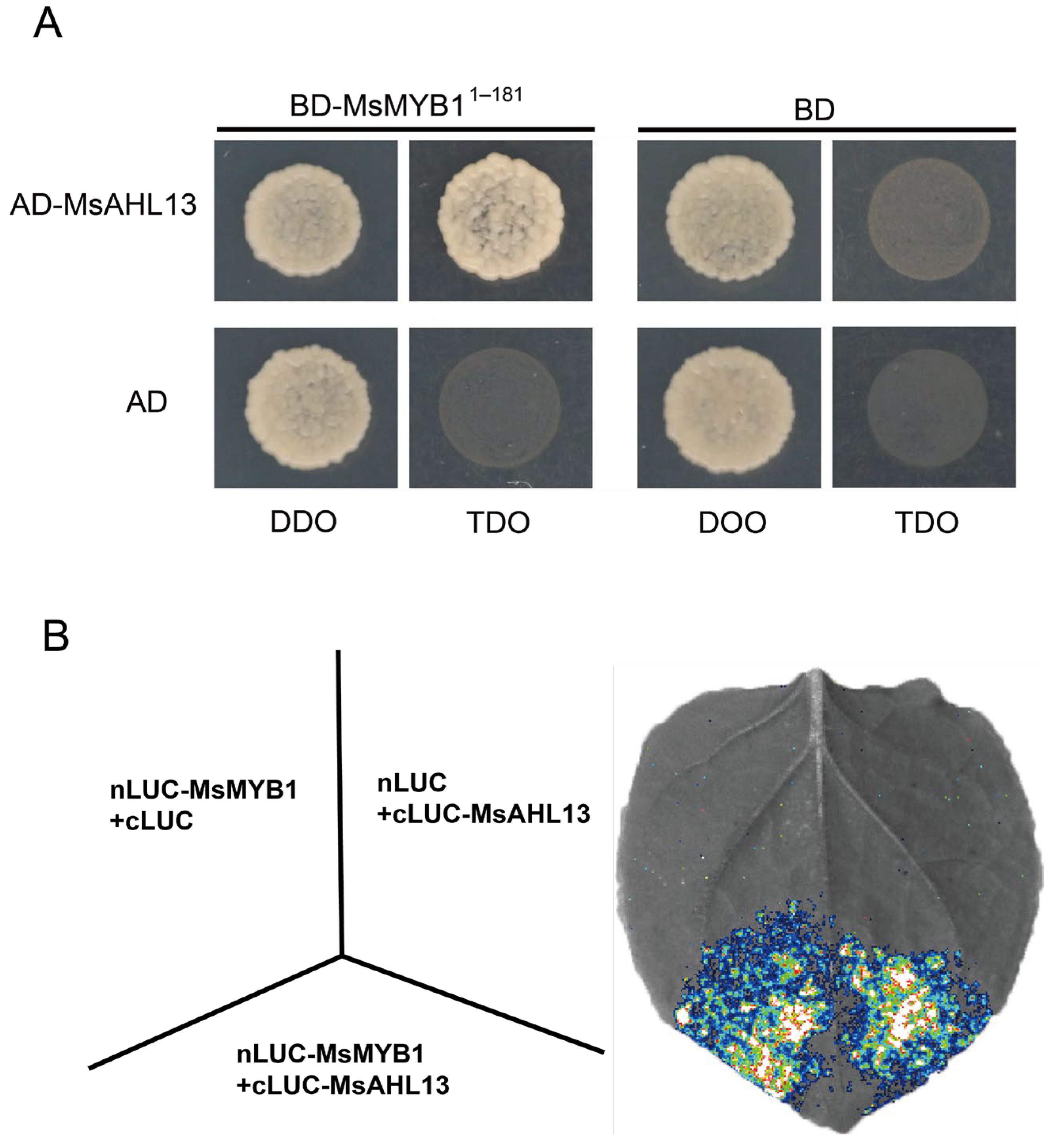
2.8. Physical Interaction Between MsAHL13 and MsMYB1
3. Discussion
4. Material and Methods
4.1. Identification of MsAHL Gene Family Members in M. sieversii
4.2. Phylogenetic Analysis of the AHLs in M. sieversii
4.3. Chromosomal Localization of MsAHLs
4.4. Analysis of Gene Structure, Conserved Motifs, and Phylogenetics of MsAHLs
4.5. Cis-Acting Element Analysis
4.6. Subcellular Localization
4.7. Real-Time Fluorescence Quantitative (qRT-PCR) Analysis
4.8. Yeast Two-Hybrid Assay
4.9. Luciferase Assay
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhang, W.M.; Cheng, X.Z.; Fang, D.; Cao, J. AT-HOOK MOTIF NUCLEAR LOCALIZED (AHL) proteins of ancient origin radiate new functions. Int. J. Biol. Macromol. 2022, 214, 290–300. [Google Scholar] [CrossRef]
- Zhao, J.; Favero, D.S.; Peng, H.; Neff, M.M. Arabidopsis thaliana AHL family modulates hypocotyl growth redundantly by interacting with each other via the PPC/DUF296 domain. Proc. Natl. Acad. Sci. USA 2013, 110, E4688–E4697. [Google Scholar] [CrossRef] [PubMed]
- Ambadas, D.A.; Singh, A.; Jha, R.K.; Chauhan, D.; B, S.; Sharma, V.K. Genome-wide dissection of AT-hook motif nuclear-localized gene family and their expression profiling for drought and salt stress in rice (Oryza sativa). Front. Plant Sci. 2023, 14, 1283555. [Google Scholar] [CrossRef]
- Wang, L.; Li, T.; Liu, N.; Liu, X. Identification of tomato AHL gene families and functional analysis their roles in fruit development and abiotic stress response. Plant Physiol. Biochem. 2023, 202, 107931. [Google Scholar] [CrossRef]
- Bishop, E.H.; Kumar, R.; Luo, F.; Saski, C.; Sekhon, R.S. Genome-wide identification, expression profiling, and network analysis of AT-hook gene family in maize. Genomics 2020, 112, 1233–1244. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Leng, X.; Yang, J.; Zhang, M.; Zeng, M.; Xu, X.; Wang, F.; Li, C. Comprehensive analysis of AHL gene family and their expression under drought stress and ABA treatment in Populus trichocarpa. PeerJ 2021, 9, e10932. [Google Scholar] [CrossRef] [PubMed]
- Jia, P.; Liu, J.; Yan, R.; Yang, K.; Dong, Q.; Luan, H.; Zhang, X.; Li, H.; Guo, S.; Qi, G. Systematical Characterization of the AT-Hook Gene Family in Juglans regia L. and the Functional Analysis of the JrAHL2 in Flower Induction and Hypocotyl Elongation. Int. J. Mol. Sci. 2023, 24, 7244. [Google Scholar] [CrossRef]
- Li, Y.; Jiang, L.; Mo, W.; Wang, L.; Zhang, L.; Cao, Y. AHLs’ life in plants: Especially their potential roles in responding to Fusarium wilt and repressing the seed oil accumulation. Int. J. Biol. Macromol. 2022, 208, 509–519. [Google Scholar] [CrossRef]
- Jin, Y.; Luo, Q.; Tong, H.; Wang, A.; Cheng, Z.; Tang, J.; Li, D.; Zhao, X.; Li, X.; Wan, J.; et al. An AT-hook gene is required for palea formation and floral organ number control in rice. Dev. Biol. 2011, 359, 277–288. [Google Scholar] [CrossRef]
- Kim, H.-B.; Oh, C.-J.; Park, Y.-C.; Lee, Y.; Choe, S.-H.; An, C.-S.; Choi, S.-B. Comprehensive analysis of AHL homologous genes encoding AT-hook motif nuclear localized protein in rice. BMB Rep. 2011, 44, 680–685. [Google Scholar] [CrossRef]
- Battista, S.; Fedele, M.; Secco, L.; Ingo, A.M.D.; Sgarra, R.; Manfioletti, G. Binding to the Other Side: The AT-Hook DNA-Binding Domain Allows Nuclear Factors to Exploit the DNA Minor Groove. Int. J. Mol. Sci. 2024, 25, 8863. [Google Scholar] [CrossRef] [PubMed]
- Filarsky, M.; Zillner, K.; Araya, I.; Villar-Garea, A.; Merkl, R.; Längst, G.; Németh, A. The extended AT-hook is a novel RNA binding motif. RNA Biol. 2015, 12, 864–876. [Google Scholar] [CrossRef]
- Zhao, J.; Favero, D.S.; Qiu, J.; Roalson, E.H.; Neff, M.M. Insights into the evolution and diversification of the AT-hook Motif Nuclear Localized gene family in land plants. BMC Plant Biol. 2014, 14, 266. [Google Scholar] [CrossRef]
- Favero, D.S.; Kawamura, A.; Shibata, M.; Takebayashi, A.; Jung, J.-H.; Suzuki, T.; Jaeger, K.E.; Ishida, T.; Iwase, A.; Wigge, P.A.; et al. AT-Hook Transcription Factors Restrict Petiole Growth by Antagonizing PIFs. Curr. Biol. 2020, 30, 1454–1466. [Google Scholar] [CrossRef]
- Yin, D.; Liu, X.; Shi, Z.; Li, D.; Zhu, L. An AT-hook protein DEPRESSED PALEA1 physically interacts with the TCP Family transcription factor RETARDED PALEA1 in rice. Biochem. Biophys. Res. Commun. 2018, 495, 487–492. [Google Scholar] [CrossRef]
- Wong, M.M.; Bhaskara, G.B.; Wen, T.-N.; Lin, W.-D.; Nguyen, T.T.; Chong, G.L.; Verslues, P.E. Phosphoproteomics of Arabidopsis Highly ABA-Induced1 identifies AT-Hook-Like10 phosphorylation required for stress growth regulation. Proc. Natl. Acad. Sci. USA 2019, 116, 2354–2363. [Google Scholar] [CrossRef]
- Mazzella, M.A.; Casal, J.J.; Muschietti, J.P.; Fox, A.R. Hormonal networks involved in apical hook development in darkness and their response to light. Front. Plant Sci. 2014, 5, 52. [Google Scholar] [CrossRef]
- Rayapuram, N.; Jarad, M.; Alhoraibi, H.M.; Bigeard, J.; Abulfaraj, A.A.; Völz, R.; Mariappan, K.G.; Almeida-Trapp, M.; Schlöffel, M.; Lastrucci, E.; et al. Chromatin phosphoproteomics unravels a function for AT-hook motif nuclear localized protein AHL13 in PAMP-triggered immunity. Proc. Natl. Acad. Sci. USA 2021, 118, e2004670118. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Li, M.; Jia, Z.; Liu, F.; Ma, H.; Huang, Y.; Song, S. AtMYB44 Positively Regulates the Enhanced Elongation of Primary Roots Induced by N-3-Oxo-Hexanoyl-Homoserine Lactone in Arabidopsis thaliana. Mol. Plant Microbe Interact. 2016, 29, 774–785. [Google Scholar] [CrossRef] [PubMed]
- Karami, O.; Rahimi, A.; Mak, P.; Horstman, A.; Boutilier, K.; Compier, M.; van der Zaal, B.; Offringa, R. An Arabidopsis AT-hook motif nuclear protein mediates somatic embryogenesis and coinciding genome duplication. Nat. Commun. 2021, 12, 2508. [Google Scholar] [CrossRef]
- Zhang, X.; Li, J.; Cao, Y.; Huang, J.; Duan, Q. Genome-Wide Identification and Expression Analysis under Abiotic Stress of BrAHL Genes in Brassica rapa. Int. J. Mol. Sci. 2023, 24, 12447. [Google Scholar] [CrossRef]
- Hao, G.; Zhang, H.; Zheng, D.; Burr, T.J. luxR homolog avhR in Agrobacterium vitis affects the development of a grape-specific necrosis and a tobacco hypersensitive response. J. Bacteriol. 2005, 187, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Liu, C.; Liu, J.; Wan, K.; Sun, H.; Liu, B.; Zhang, Y.; Wang, X.; Li, N. The ZmAHL25-ZmPUB19-ZmMPK5 Module Positively Regulates Resistance to Rhizoctonia solani in Maize. Plant Cell Environ. 2025, 48, 4099–4113. [Google Scholar] [CrossRef]
- Kim, S.; Kim, Y.; Seong, E.S.; Lee, Y.; Park, J.M.; Choi, D. The chili pepper CaATL1: An AT-hook motif-containing transcription factor implicated in defence responses against pathogens. Mol. Plant Pathol. 2007, 8, 761–771. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Jiang, S.; Zhang, Z.; Fang, H.; Xu, H.; Wang, Y.; Chen, X. Malus sieversii: The origin, flavonoid synthesis mechanism, and breeding of red-skinned and red-fleshed apples. Hortic. Res. 2018, 5, 70. [Google Scholar] [CrossRef]
- Yan, R.; Yang, K.; Zhang, T.; Sharif, R.; Yang, S.; Li, S.; Wang, N.; Liu, J.; Zhao, S.; Wang, W.; et al. Comprehensive analysis of AHL genes in Malus domestica reveals the critical role of MdAHL6 in flowering induction. Int. J. Biol. Macromol. 2024, 281, 136387. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Wu, W.; Zheng, X.; Lu, L.; Chen, X.; Hao, Z.; Liu, S.; Chen, Y. AT-Hook Transcription Factors Show Functions in Liriodendron chinense under Drought Stress and Somatic Embryogenesis. Plants 2023, 12, 1353. [Google Scholar] [CrossRef]
- Dahro, B.; Wang, Y.; Khan, M.; Zhang, Y.; Fang, T.; Ming, R.; Li, C.; Liu, J. Two AT-Hook proteins regulate A/NINV7 expression to modulate sucrose catabolism for cold tolerance in Poncirus trifoliata. New Phytol. 2022, 235, 2331–2349. [Google Scholar] [CrossRef]
- Shi, X.; Yang, T.; Ren, M.; Fu, J.; Bai, J.; Cui, H. AT-hook motif nuclear localized transcription factors function redundantly in promoting root growth through modulation of redox homeostasis. Plant J. 2024, 120, 199–217. [Google Scholar] [CrossRef]
- Hao, Y.; Zong, X.; Ren, P.; Qian, Y.; Fu, A. Basic Helix-Loop-Helix (bHLH) Transcription Factors Regulate a Wide Range of Functions in Arabidopsis. Int. J. Mol. Sci. 2021, 22, 7152. [Google Scholar] [CrossRef]
- Castelain, M.; Le Hir, R.; Bellini, C. The non-DNA-binding bHLH transcription factor PRE3/bHLH135/ATBS1/TMO7 is involved in the regulation of light signaling pathway in Arabidopsis. Physiol. Plant. 2012, 145, 450–460. [Google Scholar] [CrossRef]
- Ooka, H.; Satoh, K.; Doi, K.; Nagata, T.; Otomo, Y.; Murakami, K.; Matsubara, K.; Osato, N.; Kawai, J.; Carninci, P.; et al. Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res. 2003, 10, 239–247. [Google Scholar] [CrossRef]
- Fuertes-Aguilar, J.; Matilla, A.J. Transcriptional Control of Seed Life: New Insights into the Role of the NAC Family. Int. J. Mol. Sci. 2024, 25, 5369. [Google Scholar] [CrossRef]
- Panchy, N.; Lehti-Shiu, M.; Shiu, S.H. Evolution of Gene Duplication in Plants. Plant Physiol. 2016, 171, 2294–2316. [Google Scholar] [CrossRef]
- Adams, K.L.; Wendel, J.F. Polyploidy and genome evolution in plants. Curr. Opin. Plant Biol. 2005, 8, 135–141. [Google Scholar] [CrossRef]
- Rahimi, A.; Karami, O.; Balazadeh, S.; Offringa, R. miR156-independent repression of the ageing pathway by longevity-promoting AHL proteins in Arabidopsis. New Phytol. 2022, 235, 2424–2438. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Q.; Song, L.; Xia, M.; Zheng, Z.; Chen, Z.; Che, X.; Liu, D. Overexpression of AHL proteins enhances root hair production by altering the transcription of RHD6-downstream genes. Plant Direct 2023, 7, e517. [Google Scholar] [CrossRef]
- Zhao, L.; Lü, Y.; Chen, W.; Yao, J.; Li, Y.; Li, Q.; Pan, J.; Fang, S.; Sun, J.; Zhang, Y. Genome-wide identification and analyses of the AHL gene family in cotton (Gossypium). BMC Genom. 2020, 21, 69. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Chen, L.; Cui, L.; Liu, Z.; Yuan, W. Genome-wide analysis of radish AHL gene family and functional verification of RsAHL14 in tomato. Front. Plant Sci. 2024, 15, 1401414. [Google Scholar] [CrossRef] [PubMed]
- Alon, S.; Eisenberg, E.; Jacob-Hirsch, J.; Rechavi, G.; Vatine, G.; Toyama, R.; Coon, S.L.; Klein, D.C.; Gothilf, Y. A new cis-acting regulatory element driving gene expression in the zebrafish pineal gland. Bioinformatics 2009, 25, 559–562. [Google Scholar] [CrossRef]
- Ito, M.; Iwase, M.; Kodama, H.; Lavisse, P.; Komamine, A.; Nishihama, R.; Machida, Y.; Watanabe, A. A novel cis-acting element in promoters of plant B-type cyclin genes activates M phase-specific transcription. Plant Cell 1998, 10, 331–341. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ma, Z.; Hu, L.; Jiang, W. Understanding AP2/ERF Transcription Factor Responses and Tolerance to Various Abiotic Stresses in Plants: A Comprehensive Review. Int. J. Mol. Sci. 2024, 25, 893. [Google Scholar] [CrossRef] [PubMed]
- Chung, S.; Kwon, C.; Lee, J.H. Epigenetic control of abiotic stress signaling in plants. Genes Genom. 2022, 44, 267–278. [Google Scholar] [CrossRef]
- Ahmed, S.; Hulbert, A.K.; Xin, X.; Neff, M.M. AHL26, an AT-hook gene, negatively regulates hypocotyl growth and flowering time in Arabidopsis thaliana. BMC Plant Biol. 2025, 25, 730. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Luo, M.; Li, P.; Zhao, Q.; Chao, L.; Jiao, Y.; Tian, S.; Tian, X.; Ye, S.; Yuan, Z.; et al. CsAHL20 negatively regulates epi-catechins biosynthesis under drought stress in Camellia sinensis. Plant Sci. 2025, 359, 112613. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y.; et al. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Mol. Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef]
- Liu, W.; Zhao, C.; Liu, L.; Huang, D.; Ma, C.; Li, R.; Huang, L. Genome-wide identification of the TGA gene family in kiwifruit (Actinidia chinensis spp.) and revealing its roles in response to Pseudomonas syringae pv. actinidiae (Psa) infection. Int. J. Biol. Macromol. 2022, 222, 101–113. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Meng, J.F.; Xu, T.F.; Wang, Z.Z.; Fang, Y.L.; Xi, Z.M.; Zhang, Z.W. The ameliorative effects of exogenous melatonin on grape cuttings under water-deficient stress: Antioxidant metabolites, leaf anatomy, and chloroplast morphology. J. Pineal Res. 2014, 57, 200–212. [Google Scholar] [CrossRef]
- Xiao, H.; Siddiqua, M.; Braybrook, S.; Nassuth, A. Three grape CBF/DREB1 genes respond to low temperature, drought and abscisic acid. Plant Cell Environ. 2006, 29, 1410–1421. [Google Scholar] [CrossRef]
- Lu, X.; Ma, L.; Zhang, C.; Yan, H.; Bao, J.; Gong, M.; Wang, W.; Li, S.; Ma, S.; Chen, B. Grapevine (Vitis vinifera) responses to salt stress and alkali stress: Transcriptional and metabolic profiling. BMC Plant Biol. 2022, 22, 528. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Liu, W.; Yao, C.; Zhang, Y.; Du, X.; Ma, C.; Li, R.; Wang, H.; Huang, L. AcNAC10, regulated by AcTGA07, enhances kiwifruit resistance to Pseudomonas syringae pv. actinidiae via inhibiting jasmonic acid pathway. Mol. Hortic. 2025, 5, 21. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, D.; Zhao, C.; Liu, X.; Wang, H.; Zhu, B.; Zhao, G.; Chen, D.; Zhao, T.; Xu, H.; Wang, Y.; et al. Genome-Wide Analysis of the AT-Hook Gene Family in Malus sieversii and Functional Characterization of MsAHL13. Plants 2025, 14, 2625. https://doi.org/10.3390/plants14172625
Zhang D, Zhao C, Liu X, Wang H, Zhu B, Zhao G, Chen D, Zhao T, Xu H, Wang Y, et al. Genome-Wide Analysis of the AT-Hook Gene Family in Malus sieversii and Functional Characterization of MsAHL13. Plants. 2025; 14(17):2625. https://doi.org/10.3390/plants14172625
Chicago/Turabian StyleZhang, Da, Chao Zhao, Xin Liu, Han Wang, Bowei Zhu, Guodong Zhao, Dongmei Chen, Tongsheng Zhao, Haijiao Xu, Yingjie Wang, and et al. 2025. "Genome-Wide Analysis of the AT-Hook Gene Family in Malus sieversii and Functional Characterization of MsAHL13" Plants 14, no. 17: 2625. https://doi.org/10.3390/plants14172625
APA StyleZhang, D., Zhao, C., Liu, X., Wang, H., Zhu, B., Zhao, G., Chen, D., Zhao, T., Xu, H., Wang, Y., Zhang, C., & Zhang, X. (2025). Genome-Wide Analysis of the AT-Hook Gene Family in Malus sieversii and Functional Characterization of MsAHL13. Plants, 14(17), 2625. https://doi.org/10.3390/plants14172625





