Genome-Wide Analysis of the Maize LBD Gene Family Reveals a Role for ZmLBD12 in the Development of Lateral Roots
Abstract
1. Introduction
2. Results
2.1. Identification of the ZmLBD Gene Family
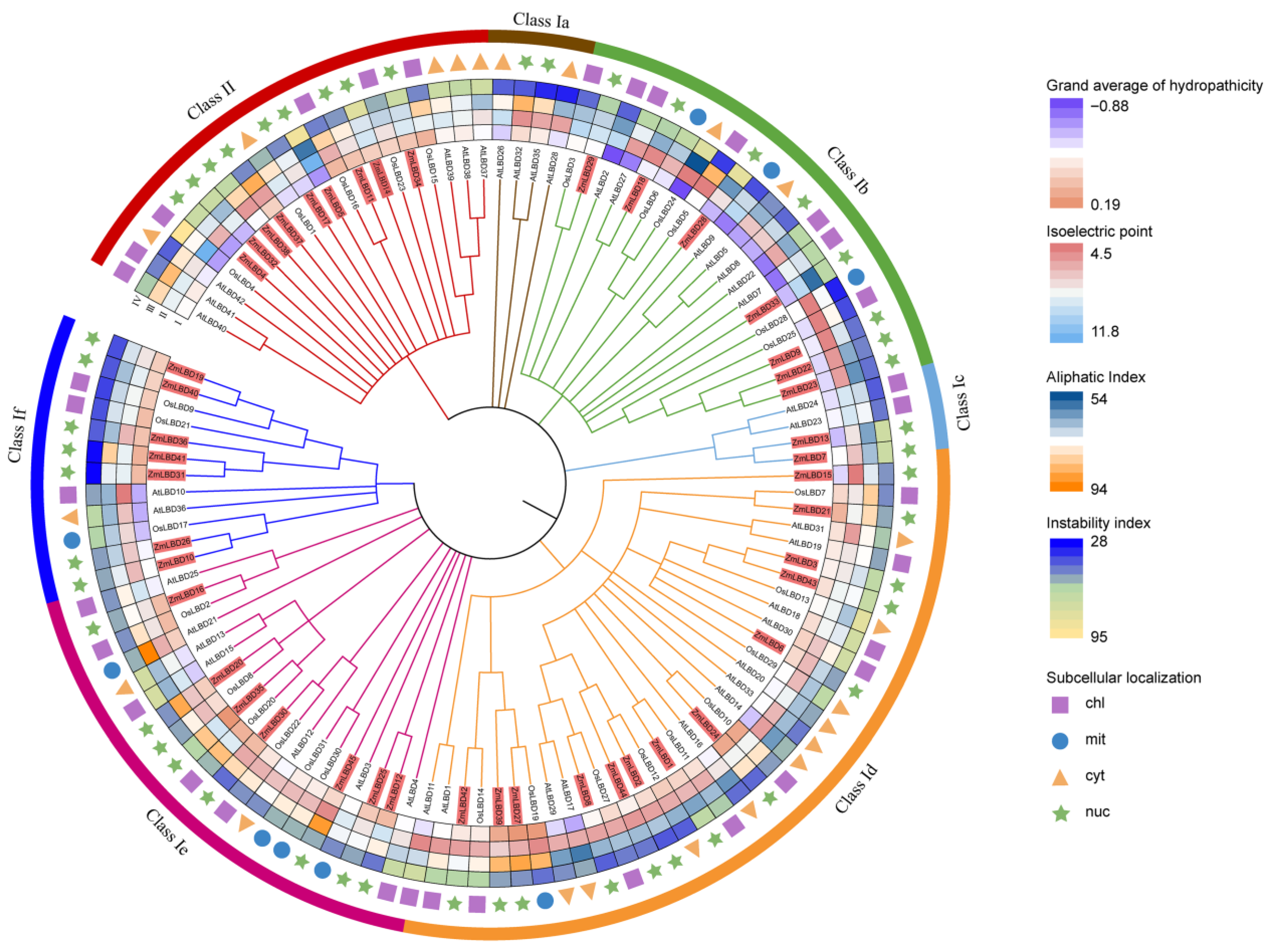
2.2. Phylogenetic Relationship of the LBD Proteins Among Maize, Rice, and Arabidopsis thaliana
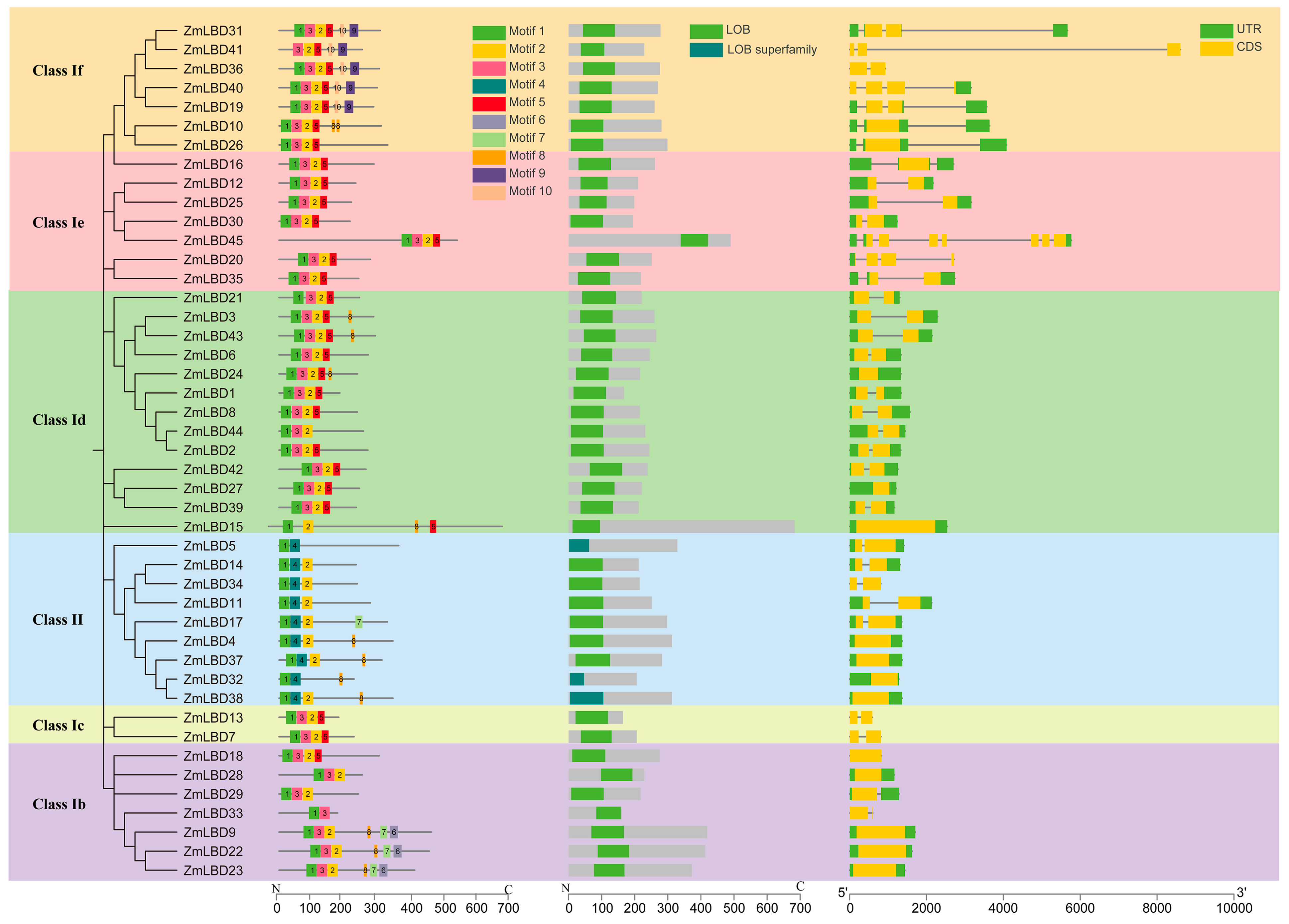
2.3. Gene Structure and Protein Motifs of the ZmLBDs
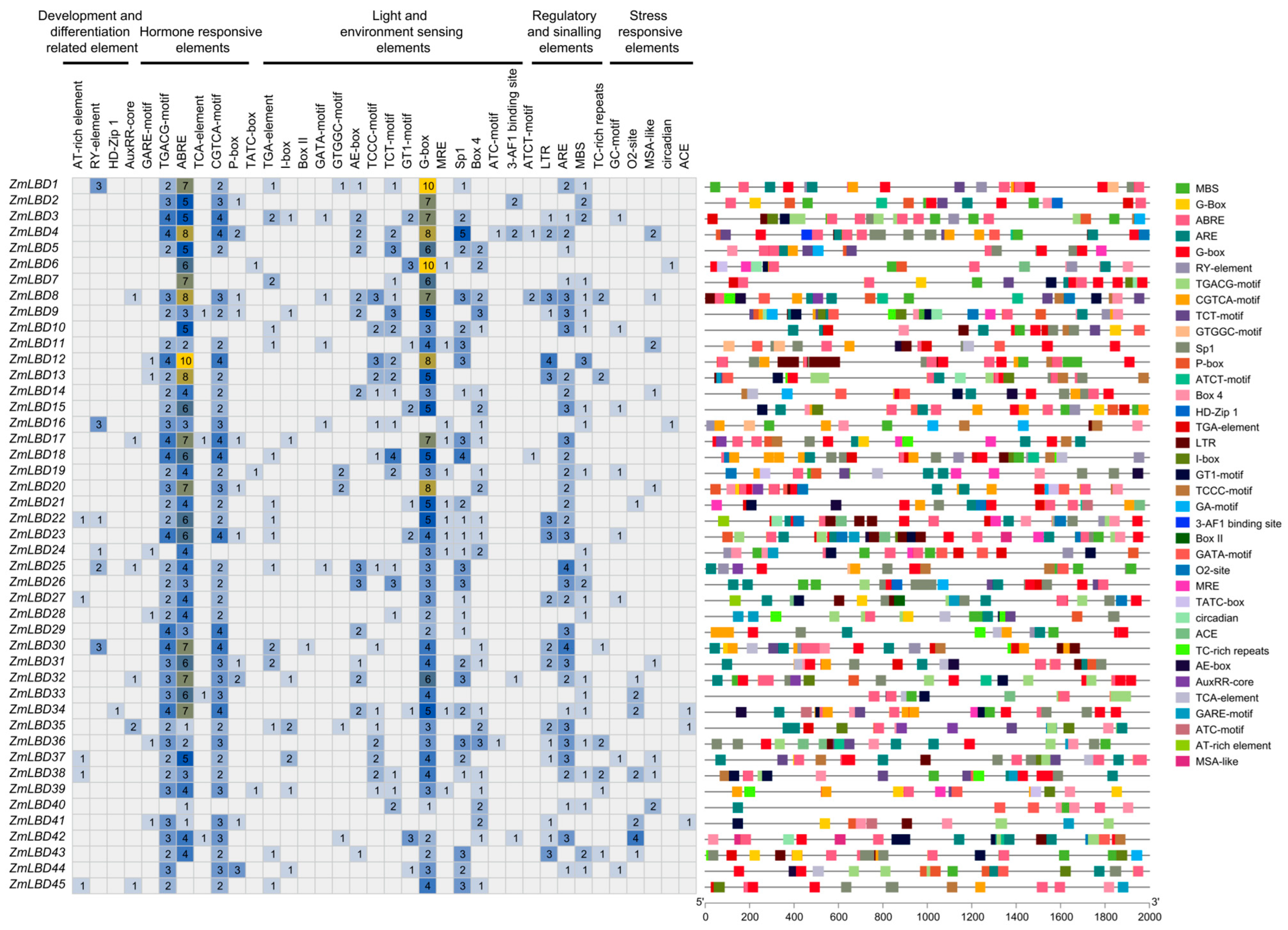
2.4. cis-Regulatory Elements in the Promoters of ZmLBD Genes
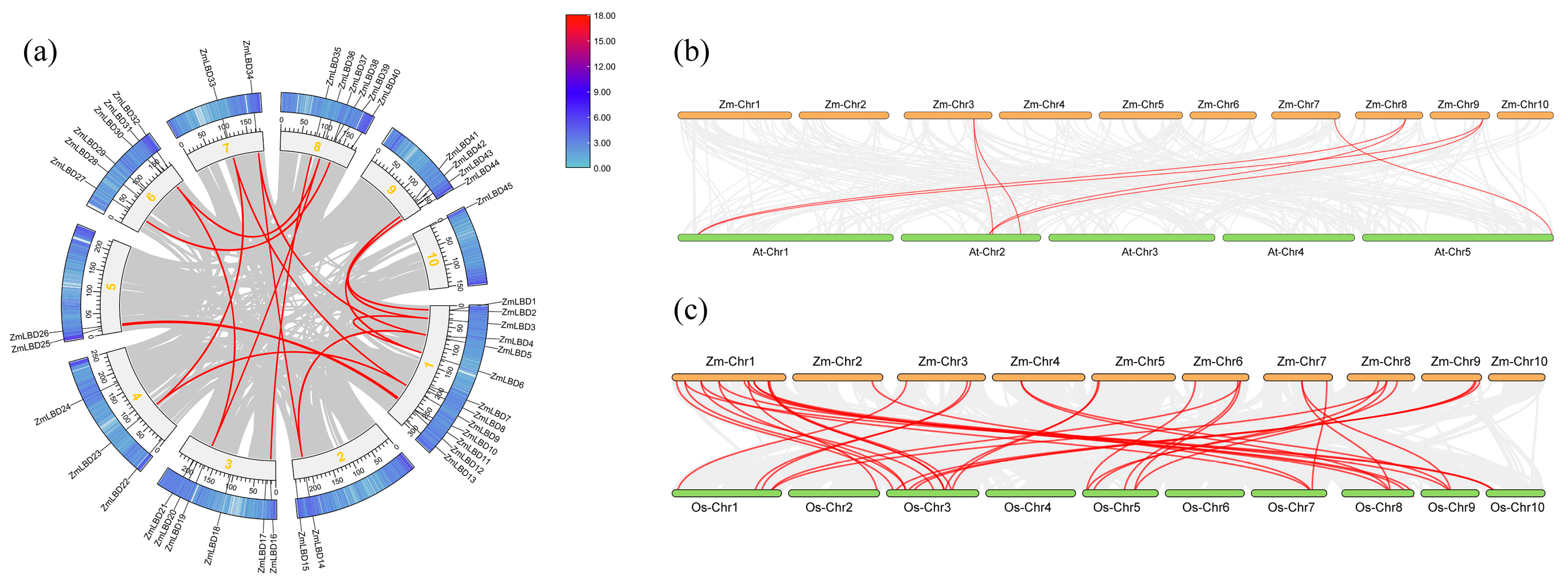
2.5. Synteny and Collinearity of the ZmLBD Proteins
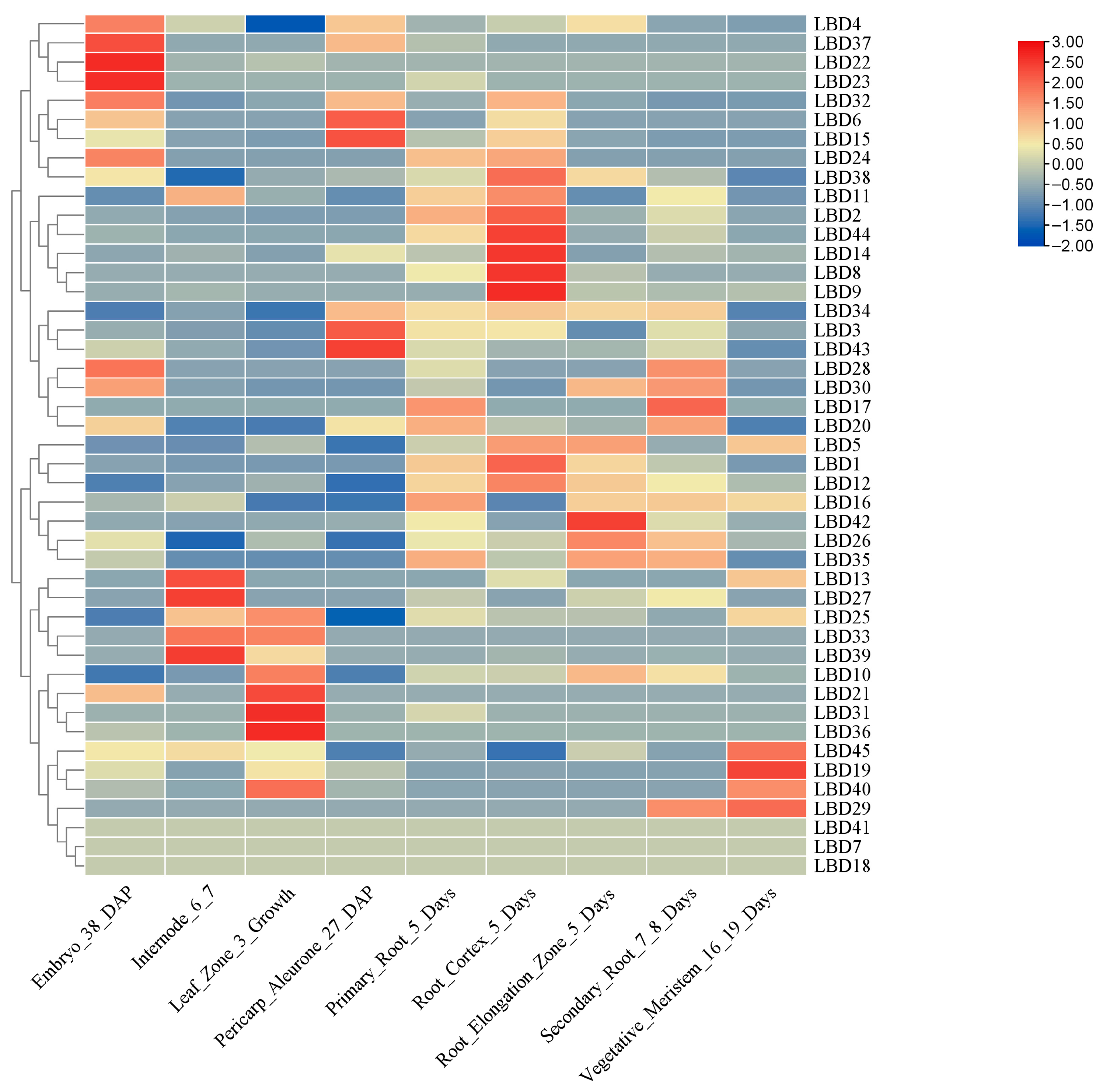
2.6. Analysis of the Pattern of Expression of the ZmLBD Genes
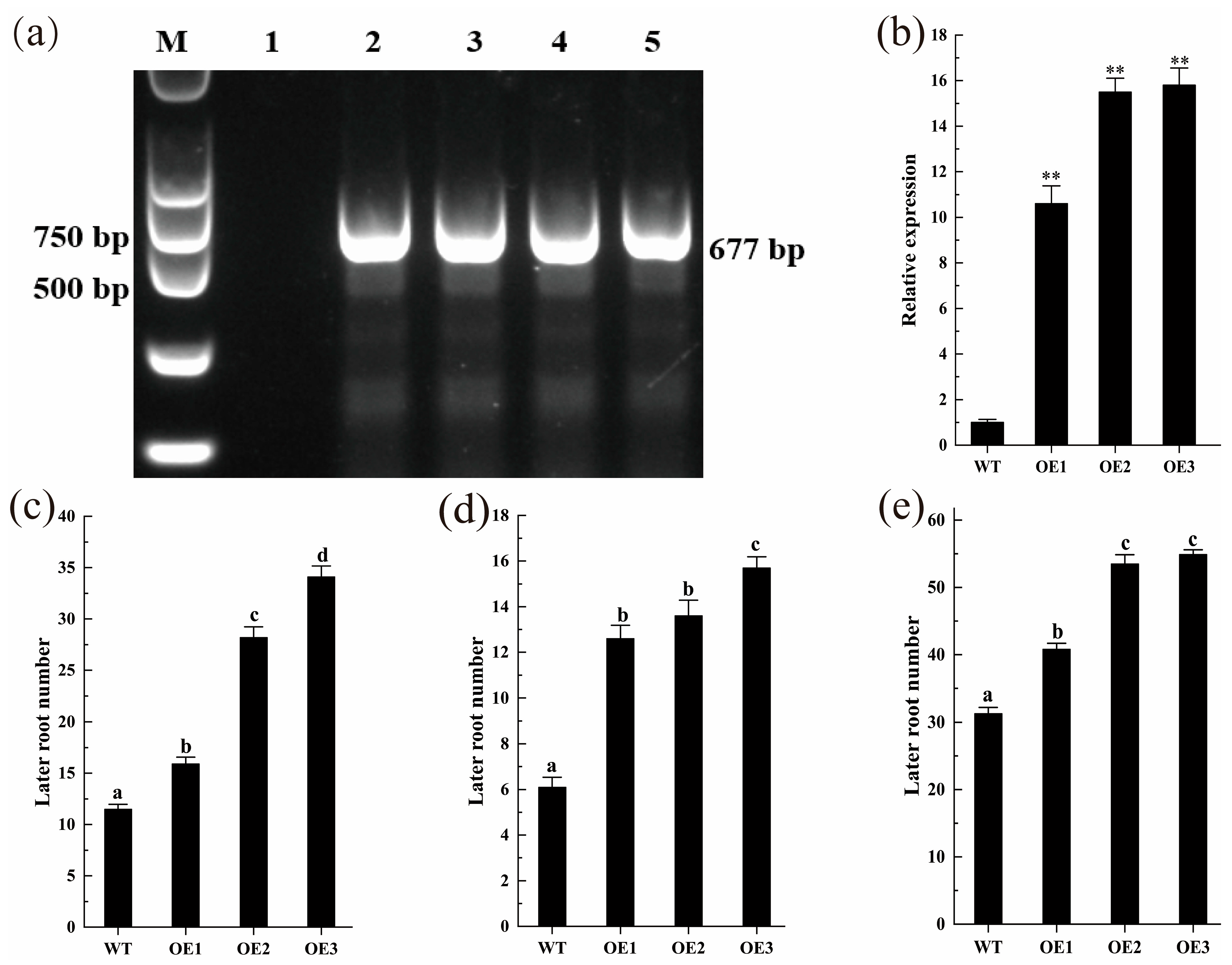
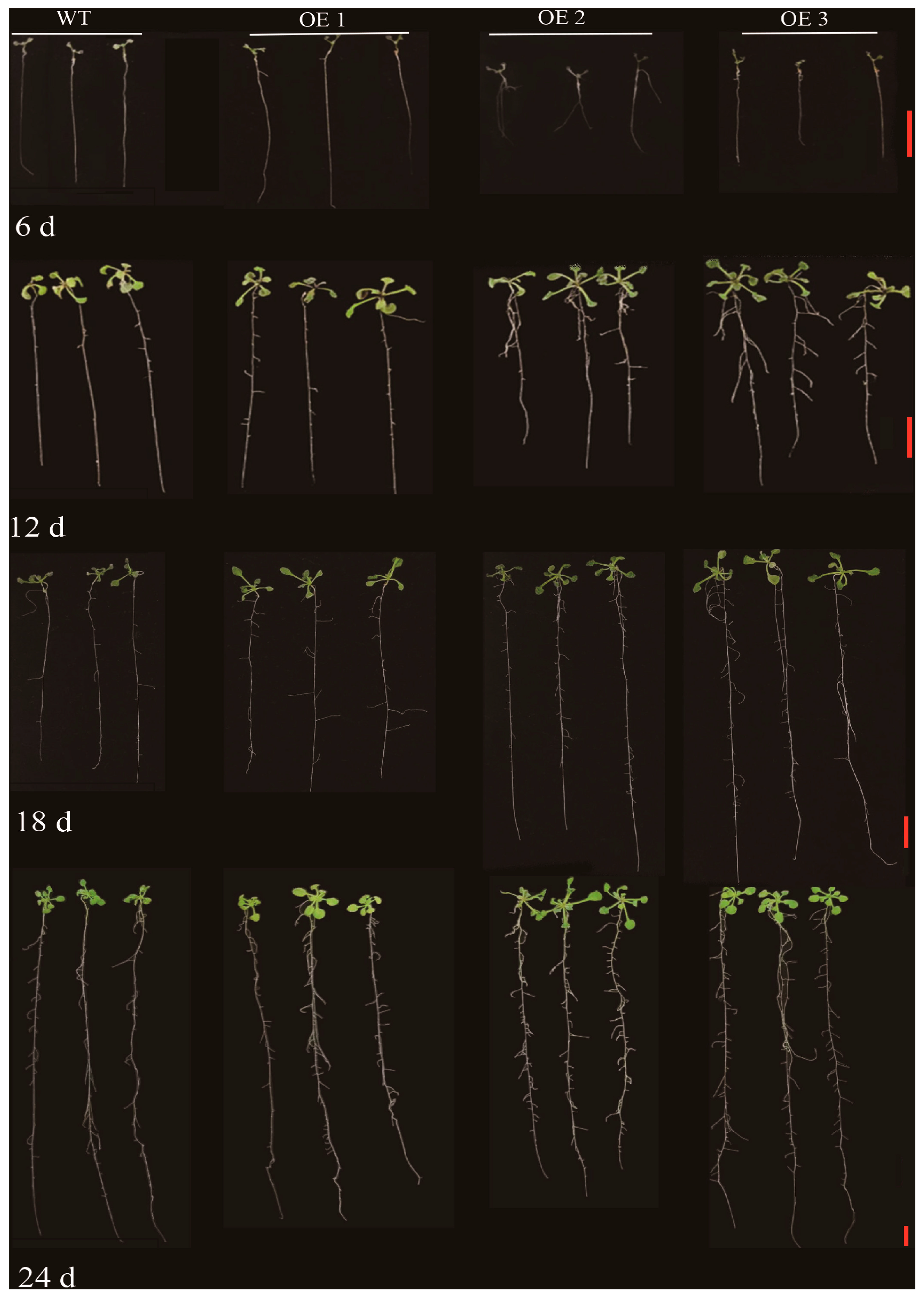
2.7. Transgenic ZmLBD12 Affects the Growth of Lateral Roots in Arabidopsis thaliana
3. Discussion
4. Materials and Methods
4.1. Identification of the LBD Family Genes and Sequence Analysis
4.2. Analysis of Gene Structural and cis-Regulatory Elements in the ZmLBD Gene Family
4.3. Gene Duplication Events of the LBDs Among Different Plants
4.4. Analysis of the Transcriptome Data for the ZmLBDs
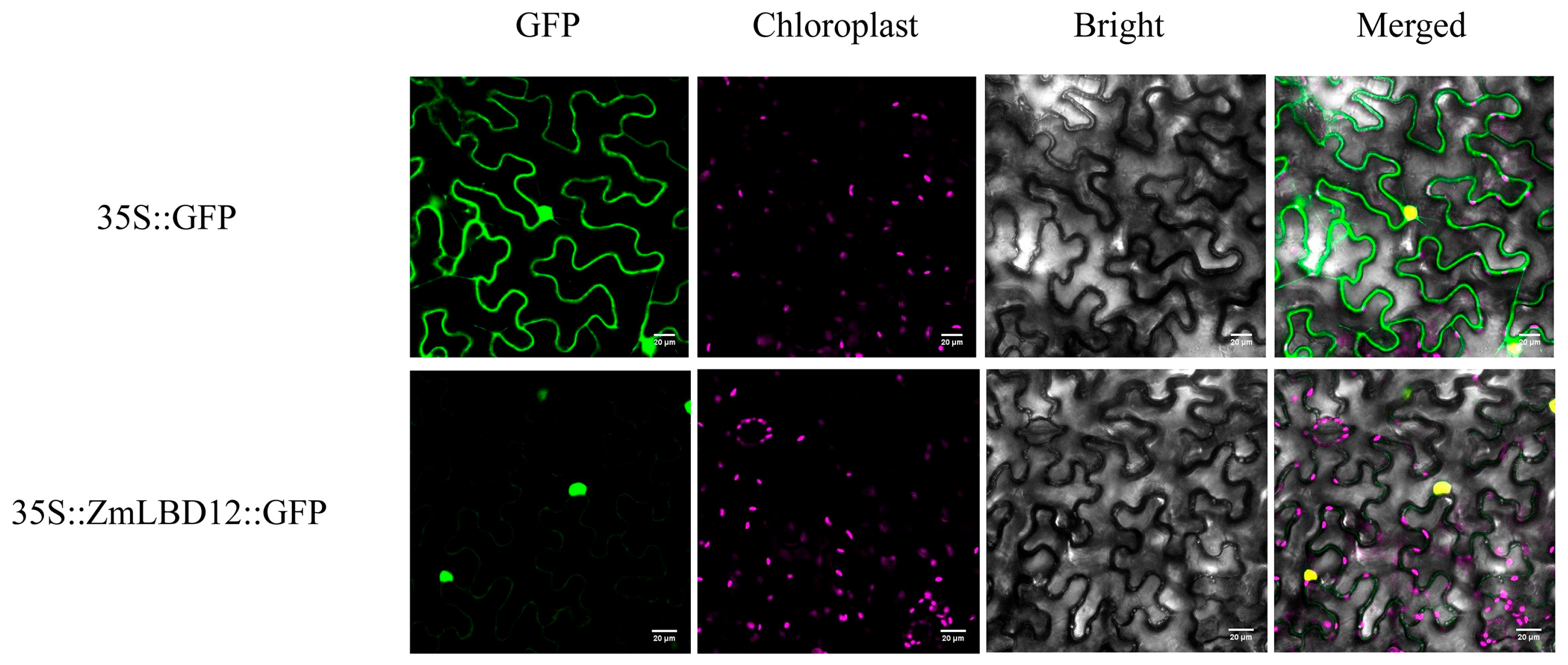
4.5. Determination of the Subcellular Localization of ZmLBD12
4.6. ZmLBD12 Gene Cloning and Real-Time Fluorescent Quantitative PCR
4.7. Genetic Transformation of ZmLBD12 into Arabidopsis thaliana
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wang, L.; Li, H.; Li, J.; Li, G.; Zahid, M.S.; Li, D.; Ma, C.; Xu, W.; Song, S.; Li, X.; et al. Transcriptome analysis revealed the expression levels of genes related to abscisic acid and auxin biosynthesis in grapevine (Vitis vinifera L.) under root restriction. Front. Plant Sci. 2022, 13, 959693. [Google Scholar] [CrossRef]
- Yun, F.; Liu, H.; Deng, Y.; Hou, X.; Liao, W. The Role of Light-Regulated Auxin Signaling in Root Development. Int. J. Mol. Sci. 2023, 24, 5253. [Google Scholar] [CrossRef]
- Zhang, J.; Tang, W.; Huang, Y.; Niu, X.; Zhao, Y.; Han, Y.; Liu, Y. Down-regulation of a LBD-like gene, OsIG1, leads to occurrence of unusual double ovules and developmental abnormalities of various floral organs and megagametophyte in rice. J. Exp. Bot. 2015, 66, 99–112. [Google Scholar] [CrossRef]
- Iwakawa, H.; Ueno, Y.; Semiarti, E.; Onouchi, H.; Kojima, S.; Tsukaya, H.; Hasebe, M.; Soma, T.; Ikezaki, M.; Machida, C.; et al. The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana, required for formation of a symmetric flat leaf lamina, encodes a member of a novel family of proteins characterized by cysteine repeats and a leucine zipper. Plant Cell Physiol. 2002, 43, 467–478. [Google Scholar] [CrossRef]
- Shuai, B.; Reynaga-Peña, C.G.; Springer, P.S. The lateral organ boundaries gene defines a novel, plant-specific gene family. Plant Physiol. 2002, 129, 747–761. [Google Scholar] [CrossRef]
- Huang, B.; Huang, Z.; Ma, R.; Ramakrishnan, M.; Chen, J.; Zhang, Z.; Yrjälä, K. Genome-wide identification and expression analysis of LBD transcription factor genes in Moso bamboo (Phyllostachys edulis). BMC Plant Biol. 2021, 21, 296. [Google Scholar] [CrossRef]
- Kim, M.J.; Kim, J. Identification of nuclear localization signal in ASYMMETRIC LEAVES2-LIKE18/LATERAL ORGAN BOUNDARIES DOMAIN16 (ASL18/LBD16) from Arabidopsis. J. Plant Physiol. 2012, 169, 1221–1226. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Yu, X.; Wu, P. Comparison and evolution analysis of two rice subspecies LATERAL ORGAN BOUNDARIES domain gene family and their evolutionary characterization from Arabidopsis. Mol. Phylogenet. Evol. 2006, 39, 248–262. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Luo, F.; Hochholdinger, F. LOB Domain Proteins: Beyond Lateral Organ Boundaries. Trends Plant Sci. 2016, 21, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Iwakawa, H.; Iwasaki, M.; Kojima, S.; Ueno, Y.; Soma, T.; Tanaka, H.; Semiarti, E.; Machida, Y.; Machida, C. Expression of the ASYMMETRIC LEAVES2 gene in the adaxial domain of Arabidopsis leaves represses cell proliferation in this domain and is critical for the development of properly expanded leaves. Plant J. 2007, 51, 173–184. [Google Scholar] [CrossRef]
- Albinsky, D.; Kusano, M.; Higuchi, M.; Hayashi, N.; Kobayashi, M.; Fukushima, A.; Mori, M.; Ichikawa, T.; Matsui, K.; Kuroda, H.; et al. Metabolomic screening applied to rice FOX Arabidopsis lines leads to the identification of a gene-changing nitrogen metabolism. Mol. Plant 2010, 3, 125–142. [Google Scholar] [CrossRef]
- Semiarti, E.; Ueno, Y.; Tsukaya, H.; Iwakawa, H.; Machida, C.; Machida, Y. The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana regulates formation of a symmetric lamina, establishment of venation and repression of meristem-related homeobox genes in leaves. Development 2001, 128, 1771–1783. [Google Scholar] [CrossRef]
- Uchida, N.; Townsley, B.; Chung, K.H.; Sinha, N. Regulation of SHOOT MERISTEMLESS genes via an upstream-conserved noncoding sequence coordinates leaf development. Proc. Natl. Acad. Sci. USA 2007, 104, 15953–15958. [Google Scholar] [CrossRef]
- Lin, W.C.; Shuai, B.; Springer, P.S. The Arabidopsis LATERAL ORGAN BOUNDARIES-domain gene ASYMMETRIC LEAVES2 functions in the repression of KNOX gene expression and in adaxial-abaxial patterning. Plant Cell 2003, 15, 2241–2252. [Google Scholar] [CrossRef]
- Ori, N.; Eshed, Y.; Chuck, G.; Bowman, J.L.; Hake, S. Mechanisms that control knox gene expression in the Arabidopsis shoot. Development 2000, 127, 5523–5532. [Google Scholar] [CrossRef]
- Xu, B.; Li, Z.; Zhu, Y.; Wang, H.; Ma, H.; Dong, A.; Huang, H. Arabidopsis genes AS1, AS2, and JAG negatively regulate boundary-specifying genes to promote sepal and petal development. Plant Physiol. 2008, 146, 566–575. [Google Scholar] [CrossRef]
- Scheible, W.R.; Morcuende, R.; Czechowski, T.; Fritz, C.; Osuna, D.; Palacios-Rojas, N.; Schindelasch, D.; Thimm, O.; Udvardi, M.K.; Stitt, M. Genome-wide reprogramming of primary and secondary metabolism, protein synthesis, cellular growth processes, and the regulatory infrastructure of Arabidopsis in response to nitrogen. Plant Physiol. 2004, 136, 2483–2499. [Google Scholar] [CrossRef] [PubMed]
- Jiao, P.; Wei, X.; Jiang, Z.; Liu, S.; Guan, S.; Ma, Y. ZmLBD2 a maize (Zea mays L.) lateral organ boundaries domain (LBD) transcription factor enhances drought tolerance in transgenic Arabidopsis thaliana. Front. Plant Sci. 2022, 13, 1000149. [Google Scholar] [CrossRef] [PubMed]
- Okushima, Y.; Fukaki, H.; Onoda, M.; Theologis, A.; Tasaka, M. ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell 2007, 19, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Taramino, G.; Sauer, M.; Stauffer, J.L., Jr.; Multani, D.; Niu, X.; Sakai, H.; Hochholdinger, F. The maize (Zea mays L.) RTCS gene encodes a LOB domain protein that is a key regulator of embryonic seminal and post-embryonic shoot-borne root initiation. Plant J. 2007, 50, 649–659. [Google Scholar] [CrossRef]
- Liu, J.; Sheng, L.; Xu, Y.; Li, J.; Yang, Z.; Huang, H.; Xu, L. WOX11 and 12 are involved in the first-step cell fate transition during de novo root organogenesis in Arabidopsis. Plant Cell 2014, 26, 1081–1093. [Google Scholar] [CrossRef] [PubMed]
- Sheng, L.; Hu, X.; Du, Y.; Zhang, G.; Huang, H.; Scheres, B.; Xu, L. Non-canonical WOX11-mediated root branching contributes to plasticity in Arabidopsis root system architecture. Development 2017, 144, 3126–3133. [Google Scholar] [CrossRef]
- Zhao, Y.; Hu, Y.; Dai, M.; Huang, L.; Zhou, D.X. The WUSCHEL-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice. Plant Cell 2009, 21, 736–748. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Xiong, J.; Zhang, W.; Guan, H.; Zheng, D.; Xiong, H.; Jia, L.; Hu, Y.; Zhou, H.; Wen, Y.; et al. ZmLBD5, a class-II LBD gene, negatively regulates drought tolerance by impairing abscisic acid synthesis. Plant J. 2022, 112, 1364–1376. [Google Scholar] [CrossRef]
- Lee, H.W.; Kim, N.Y.; Lee, D.J.; Kim, J. LBD18/ASL20 regulates lateral root formation in combination with LBD16/ASL18 downstream of ARF7 and ARF19 in Arabidopsis. Plant Physiol. 2009, 151, 1377–1389. [Google Scholar] [CrossRef] [PubMed]
- Chanderbali, A.S.; He, F.; Soltis, P.S.; Soltis, D.E. Out of the Water: Origin and Diversification of the LBD Gene Family. Mol. Biol. Evol. 2015, 32, 1996–2000. [Google Scholar] [CrossRef]
- Kong, Y.; Xu, P.; Jing, X.; Chen, L.; Li, L.; Li, X. Decipher the ancestry of the plant-specific LBD gene family. BMC Genomics 2017, 18, 951. [Google Scholar] [CrossRef]
- Liu, H.; Cao, M.; Chen, X.; Ye, M.; Zhao, P.; Nan, Y.; Li, W.; Zhang, C.; Kong, L.; Kong, N.; et al. Genome-Wide Analysis of the Lateral Organ Boundaries Domain (LBD) Gene Family in Solanum tuberosum. Int. J. Mol. Sci. 2019, 20, 5360. [Google Scholar] [CrossRef]
- Dong, L.; Manghwar, H. Genome-wide expression analysis of LBD genes in tomato (Solanum lycopersicum L.) under different light conditions. Plant Signal Behav. 2023, 18, 2290414. [Google Scholar] [CrossRef]
- Yu, Q.; Hu, S.; Du, J.; Yang, Y.; Sun, X. Genome-wide identification and characterization of the lateral organ boundaries domain gene family in Brassica rapa var. rapa. Plant Divers. 2020, 42, 52–60. [Google Scholar] [CrossRef]
- Li, J.; Meng, X.; Zong, Y.; Chen, K.; Zhang, H.; Liu, J.; Li, J.; Gao, C. Gene replacements and insertions in rice by intron targeting using CRISPR-Cas9. Nat. Plants 2016, 2, 16139. [Google Scholar] [CrossRef]
- Staiger, D.; Brown, J.W. Alternative splicing at the intersection of biological timing, development, and stress responses. Plant Cell 2013, 25, 3640–3656. [Google Scholar] [CrossRef]
- Feng, X.; Zhou, B.; Wu, X.; Wu, H.; Zhang, S.; Jiang, Y.; Wang, Y.; Zhang, Y.; Cao, M.; Guo, B.; et al. Molecular characterization of SPL gene family during flower morphogenesis and regulation in blueberry. BMC Plant Biol. 2023, 23, 40. [Google Scholar] [CrossRef]
- Deokar, A.A.; Tar’an, B. Genome-Wide Analysis of the Aquaporin Gene Family in Chickpea (Cicer arietinum L.). Front. Plant Sci. 2016, 7, 1802. [Google Scholar] [CrossRef]
- Huang, X.; Yan, H.; Liu, Y.; Yi, Y. Genome-wide analysis of LATERAL ORGAN BOUNDARIES DOMAIN-in Physcomitrella patens and stress responses. Genes. Genomics 2020, 42, 651–662. [Google Scholar] [CrossRef]
- Song, B.; Tang, Z.; Li, X.; Li, J.; Zhang, M.; Zhao, K.; Liu, H.; Zhang, S.; Wu, J. Mining and evolution analysis of lateral organ boundaries domain (LBD) genes in Chinese white pear (Pyrus bretschneideri). BMC Genomics 2020, 21, 644. [Google Scholar] [CrossRef]
- Lian, C.; Li, Q.; Yao, K.; Zhang, Y.; Meng, S.; Yin, W.; Xia, X. Populus trichocarpa PtNF-YA9, A Multifunctional Transcription Factor, Regulates Seed Germination, Abiotic Stress, Plant Growth and Development in Arabidopsis. Front. Plant Sci. 2018, 9, 954. [Google Scholar] [CrossRef] [PubMed]
- Fan, M.; Xu, C.; Xu, K.; Hu, Y. LATERAL ORGAN BOUNDARIES DOMAIN transcription factors direct callus formation in Arabidopsis regeneration. Cell Res. 2012, 22, 1169–1180. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Salasini, B.C.; Khan, M.; Devi, B.; Bush, M.; Subramaniam, R.; Hepworth, S.R. Clade I TGACG-Motif Binding Basic Leucine Zipper Transcription Factors Mediate BLADE-ON-PETIOLE-Dependent Regulation of Development. Plant Physiol. 2019, 180, 937–951. [Google Scholar] [CrossRef]
- Li, B.J.; Shi, Y.N.; Xiao, Y.N.; Jia, H.R.; Yang, X.F.; Dai, Z.R.; Sun, Y.F.; Shou, J.H.; Jiang, G.H.; Grierson, D.; et al. AUXIN RESPONSE FACTOR 2 mediates repression of strawberry receptacle ripening via auxin-ABA interplay. Plant Physiol. 2024, 196, 2638–2653. [Google Scholar] [CrossRef] [PubMed]
- Shin, J.; Seo, P.J. Varying Auxin Levels Induce Distinct Pluripotent States in Callus Cells. Front. Plant Sci. 2018, 9, 1653. [Google Scholar] [CrossRef]
- Liu, Q.; Li, Z.; Wang, Z.; Lu, Y.; Jiang, S.; Xia, C.; An, P.; Zhao, L.; Deng, K.; Xia, Z.; et al. Construction of an ultrahigh-density genetic linkage map for Manihot esculenta Crantz and identification of QTL for root quantity traits. BMC Plant Biol. 2025, 25, 534. [Google Scholar] [CrossRef]
- Zhu, Y.; Wu, N.; Song, W.; Yin, G.; Qin, Y.; Yan, Y.; Hu, Y. Soybean (Glycine max) expansin gene superfamily origins: Segmental and tandem duplication events followed by divergent selection among subfamilies. BMC Plant Biol. 2014, 14, 93. [Google Scholar] [CrossRef]
- Bretani, G.; Rossini, L.; Ferrandi, C.; Russell, J.; Waugh, R.; Kilian, B.; Bagnaresi, P.; Cattivelli, L.; Fricano, A. Segmental duplications are hot spots of copy number variants affecting barley gene content. Plant J. 2020, 103, 1073–1088. [Google Scholar] [CrossRef]
- Mascagni, F.; Usai, G.; Cavallini, A.; Porceddu, A. Structural characterization and duplication modes of pseudogenes in plants. Sci. Rep. 2021, 11, 5292. [Google Scholar] [CrossRef]
- Yu, J.; Xie, Q.; Li, C.; Dong, Y.; Zhu, S.; Chen, J. Comprehensive characterization and gene expression patterns of LBD gene family in Gossypium. Planta 2020, 251, 81. [Google Scholar] [CrossRef] [PubMed]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef]
- Chen, Q.; Liu, Y.; Maere, S.; Lee, E.; Van Isterdael, G.; Xie, Z.; Xuan, W.; Lucas, J.; Vassileva, V.; Kitakura, S.; et al. A coherent transcriptional feed-forward motif model for mediating auxin-sensitive PIN3 expression during lateral root development. Nat. Commun. 2015, 6, 8821. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhao, P.; Chen, S.; Sun, L.; Mao, J.; Tan, S.; Xiang, C. The ABI3-ERF1 module mediates ABA-auxin crosstalk to regulate lateral root emergence. Cell Rep. 2023, 42, 112809. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, T.J.; Eddy, S.R. nhmmer: DNA homology search with profile HMMs. Bioinformatics 2013, 29, 2487–2489. [Google Scholar] [CrossRef]
- Letunic, I.; Khedkar, S.; Bork, P. SMART: Recent updates, new developments and status in 2020. Nucleic Acids Res. 2021, 49, D458–D460. [Google Scholar] [CrossRef]
- Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Gonzales, N.R.; Gwadz, M.; Lu, S.; Marchler, G.H.; Song, J.S.; Thanki, N.; Yamashita, R.A.; et al. The conserved domain database in 2023. Nucleic Acids Res. 2023, 51, D384–D388. [Google Scholar] [CrossRef]
- Duvaud, S.; Gabella, C.; Lisacek, F.; Stockinger, H.; Ioannidis, V.; Durinx, C. Expasy, the Swiss Bioinformatics Resource Portal, as designed by its users. Nucleic Acids Res. 2021, 49, W216–W227. [Google Scholar] [CrossRef] [PubMed]
- Pierleoni, A.; Martelli, P.L.; Fariselli, P.; Casadio, R. BaCelLo: A balanced subcellular localization predictor. Bioinformatics 2006, 22, e408–e416. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Walley, J.W.; Sartor, R.C.; Shen, Z.; Schmitz, R.J.; Wu, K.J.; Urich, M.A.; Nery, J.R.; Smith, L.G.; Schnable, J.C.; Ecker, J.R.; et al. Integration of omic networks in a developmental atlas of maize. Science 2016, 353, 814–818. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]

| Gene Name | Gene ID (Zm-B73-REFERENCE-NAM-5.0) | Amino Acid Length | Molecular Weight | Isoelectric Point | Instability Index | Aliphatic Index | Grand Average of Hydropathy (GRAVY) |
|---|---|---|---|---|---|---|---|
| ZmLBD1 | Zm00001eb003910 | 167 | 17,442.63 | 6.27 | 66.36 | 63.95 | −0.131 |
| ZmLBD2 | Zm00001eb003920 | 244 | 25,470.18 | 5.66 | 42.7 | 67.91 | −0.165 |
| ZmLBD3 | Zm00001eb010570 | 260 | 26,838.36 | 7.76 | 67.18 | 71 | −0.269 |
| ZmLBD4 | Zm00001eb019930 | 313 | 33,961.79 | 6.14 | 71.23 | 71.5 | −0.637 |
| ZmLBD5 | Zm00001eb020740 | 329 | 36,067.36 | 11.74 | 89.52 | 58.81 | −0.661 |
| ZmLBD6 | Zm00001eb026870 | 245 | 25,337.54 | 6.36 | 54.46 | 69.63 | −0.149 |
| ZmLBD7 | Zm00001eb035850 | 206 | 21,561.19 | 6.64 | 72.17 | 66.94 | −0.338 |
| ZmLBD8 | Zm00001eb038820 | 215 | 23,764.66 | 5.7 | 43.93 | 65.91 | −0.305 |
| ZmLBD9 | Zm00001eb042180 | 419 | 45,005.73 | 4.84 | 47.25 | 71.36 | −0.399 |
| ZmLBD10 | Zm00001eb051480 | 281 | 30,137.82 | 6.41 | 55.28 | 72.28 | −0.342 |
| ZmLBD11 | Zm00001eb051620 | 251 | 25,553.79 | 7.59 | 52.44 | 76.85 | −0.037 |
| ZmLBD12 | Zm00001eb052620 | 211 | 21,657.7 | 9.13 | 47.7 | 76.49 | 0.004 |
| ZmLBD13 | Zm00001eb060210 | 164 | 17,625.74 | 7.61 | 73.43 | 63.9 | −0.538 |
| ZmLBD14 | Zm00001eb108240 | 212 | 21,648.61 | 9.03 | 81.23 | 71.46 | −0.096 |
| ZmLBD15 | Zm00001eb117380 | 683 | 74,595.77 | 4.52 | 52.19 | 71.92 | −0.466 |
| ZmLBD16 | Zm00001eb123060 | 261 | 26,781.33 | 7.63 | 55.93 | 74.87 | −0.011 |
| ZmLBD17 | Zm00001eb125420 | 298 | 32,356.28 | 7.66 | 50.71 | 72.75 | −0.471 |
| ZmLBD18 | Zm00001eb134030 | 275 | 29,785.4 | 4.94 | 61.88 | 65.05 | −0.388 |
| ZmLBD19 | Zm00001eb145150 | 260 | 26,721 | 7.64 | 41.33 | 71.08 | −0.085 |
| ZmLBD20 | Zm00001eb149360 | 251 | 27,297.14 | 6.71 | 52.26 | 87.93 | −0.091 |
| ZmLBD21 | Zm00001eb151910 | 221 | 22,980.32 | 8.1 | 55.62 | 80.05 | 0.038 |
| ZmLBD22 | Zm00001eb167220 | 413 | 43,899.25 | 5.38 | 49.32 | 64.84 | −0.46 |
| ZmLBD23 | Zm00001eb178740 | 373 | 40,024.3 | 5.58 | 46.77 | 61.8 | −0.461 |
| ZmLBD24 | Zm00001eb191160 | 216 | 22,426.46 | 8.1 | 45.27 | 78.52 | 0.095 |
| ZmLBD25 | Zm00001eb218120 | 199 | 20,689.5 | 7.65 | 54.94 | 73.67 | −0.07 |
| ZmLBD26 | Zm00001eb219100 | 299 | 31,795.7 | 6.82 | 53.72 | 68.93 | −0.368 |
| ZmLBD27 | Zm00001eb265490 | 221 | 23,002.21 | 6.03 | 52.15 | 88.64 | 0.18 |
| ZmLBD28 | Zm00001eb269620 | 229 | 24,347.37 | 7.6 | 84.18 | 62.36 | −0.556 |
| ZmLBD29 | Zm00001eb272570 | 218 | 23,124.11 | 9.15 | 61.78 | 69.27 | −0.348 |
| ZmLBD30 | Zm00001eb286410 | 195 | 20,845.92 | 5.9 | 46.23 | 80.15 | 0.005 |
| ZmLBD31 | Zm00001eb288160 | 278 | 28,814.73 | 8.5 | 28.84 | 72.66 | 0.003 |
| ZmLBD32 | Zm00001eb292990 | 206 | 21,875.46 | 8.53 | 71.27 | 66.55 | −0.554 |
| ZmLBD33 | Zm00001eb311030 | 161 | 17,275.49 | 9.72 | 65.09 | 57.83 | −0.546 |
| ZmLBD34 | Zm00001eb326200 | 215 | 22,175.3 | 8.85 | 72.25 | 70.42 | −0.054 |
| ZmLBD35 | Zm00001eb345960 | 219 | 22,316.3 | 9.1 | 63.05 | 83.2 | 0.189 |
| ZmLBD36 | Zm00001eb349310 | 276 | 28,184.69 | 6.29 | 43.66 | 66.99 | −0.035 |
| ZmLBD37 | Zm00001eb353210 | 283 | 29,602.54 | 5.57 | 50.58 | 87.39 | −0.106 |
| ZmLBD38 | Zm00001eb353230 | 313 | 33,373.53 | 5.37 | 67.38 | 80.54 | −0.354 |
| ZmLBD39 | Zm00001eb355150 | 212 | 22,149.29 | 7.06 | 54.3 | 85.42 | 0.086 |
| ZmLBD40 | Zm00001eb365880 | 270 | 27,773.26 | 8.52 | 42.82 | 67.04 | −0.109 |
| ZmLBD41 | Zm00001eb395110 | 229 | 23,611.66 | 8.48 | 29.86 | 81.79 | 0.049 |
| ZmLBD42 | Zm00001eb397130 | 239 | 25,340.56 | 5.6 | 62.97 | 73.56 | −0.227 |
| ZmLBD43 | Zm00001eb398670 | 265 | 27,061.63 | 8.26 | 69.94 | 73.7 | −0.18 |
| ZmLBD44 | Zm00001eb403030 | 232 | 24,697.23 | 5.37 | 47.76 | 66.68 | −0.271 |
| ZmLBD45 | Zm00001eb405850 | 490 | 54,339.82 | 4.99 | 56.15 | 90.31 | −0.138 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, S.; Wang, Y.; Zhong, J.; Xu, W.; Gong, Q.; Zhai, L.; Li, G.; Huang, J. Genome-Wide Analysis of the Maize LBD Gene Family Reveals a Role for ZmLBD12 in the Development of Lateral Roots. Plants 2025, 14, 2600. https://doi.org/10.3390/plants14162600
Wang S, Wang Y, Zhong J, Xu W, Gong Q, Zhai L, Li G, Huang J. Genome-Wide Analysis of the Maize LBD Gene Family Reveals a Role for ZmLBD12 in the Development of Lateral Roots. Plants. 2025; 14(16):2600. https://doi.org/10.3390/plants14162600
Chicago/Turabian StyleWang, Shifeng, Yang Wang, Jianbing Zhong, Wenlin Xu, Qingyou Gong, Lihong Zhai, Gaoke Li, and Jun Huang. 2025. "Genome-Wide Analysis of the Maize LBD Gene Family Reveals a Role for ZmLBD12 in the Development of Lateral Roots" Plants 14, no. 16: 2600. https://doi.org/10.3390/plants14162600
APA StyleWang, S., Wang, Y., Zhong, J., Xu, W., Gong, Q., Zhai, L., Li, G., & Huang, J. (2025). Genome-Wide Analysis of the Maize LBD Gene Family Reveals a Role for ZmLBD12 in the Development of Lateral Roots. Plants, 14(16), 2600. https://doi.org/10.3390/plants14162600






