Comparative Transcriptomic Analysis Reveals the Negative Response Mechanism of Peanut Root Morphology and Nitrate Assimilation to Nitrogen Deficiency
Abstract
1. Introduction
2. Results
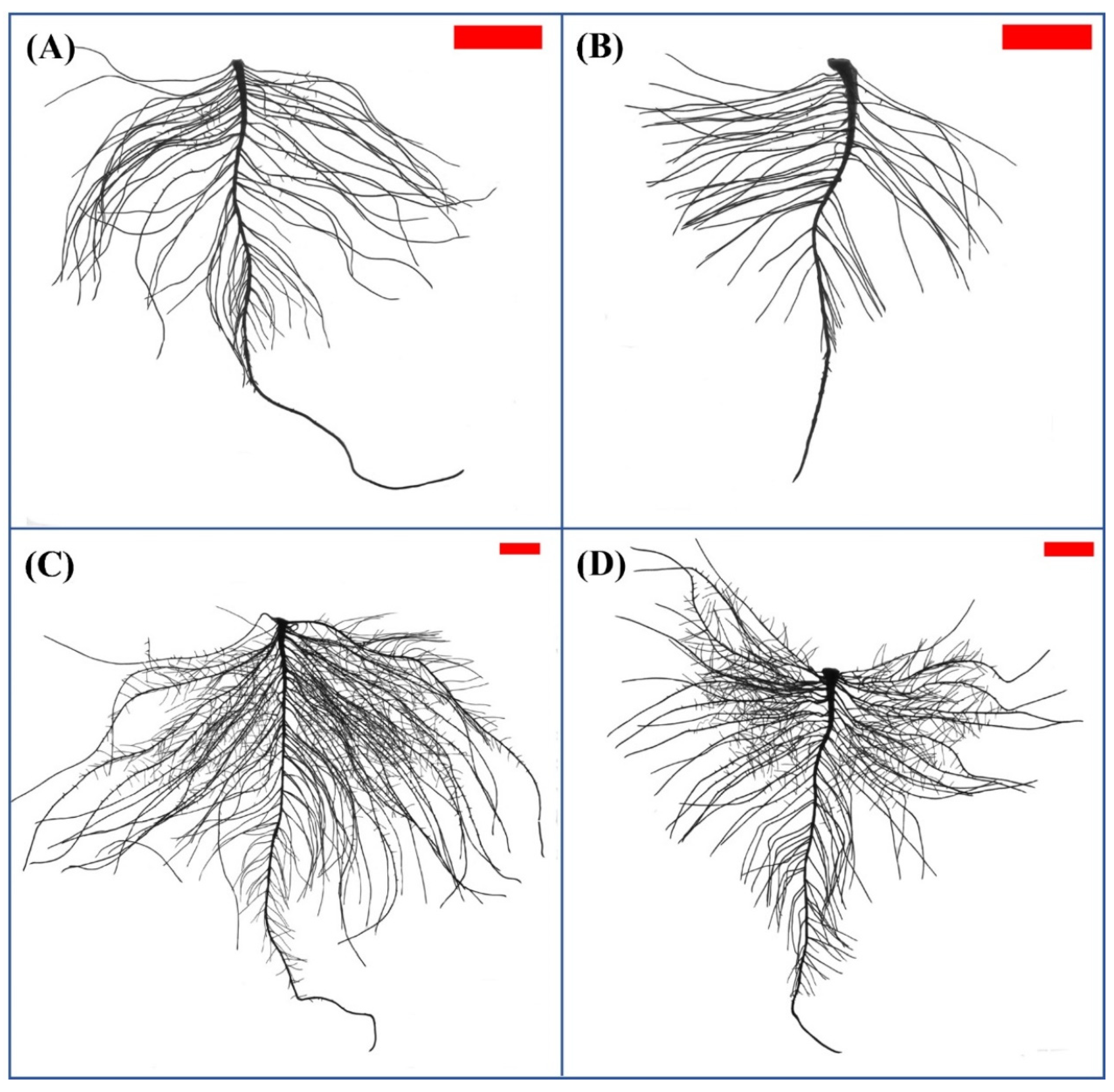
2.1. Nitrogen Deficiency Affected the Morphology of Peanut Roots
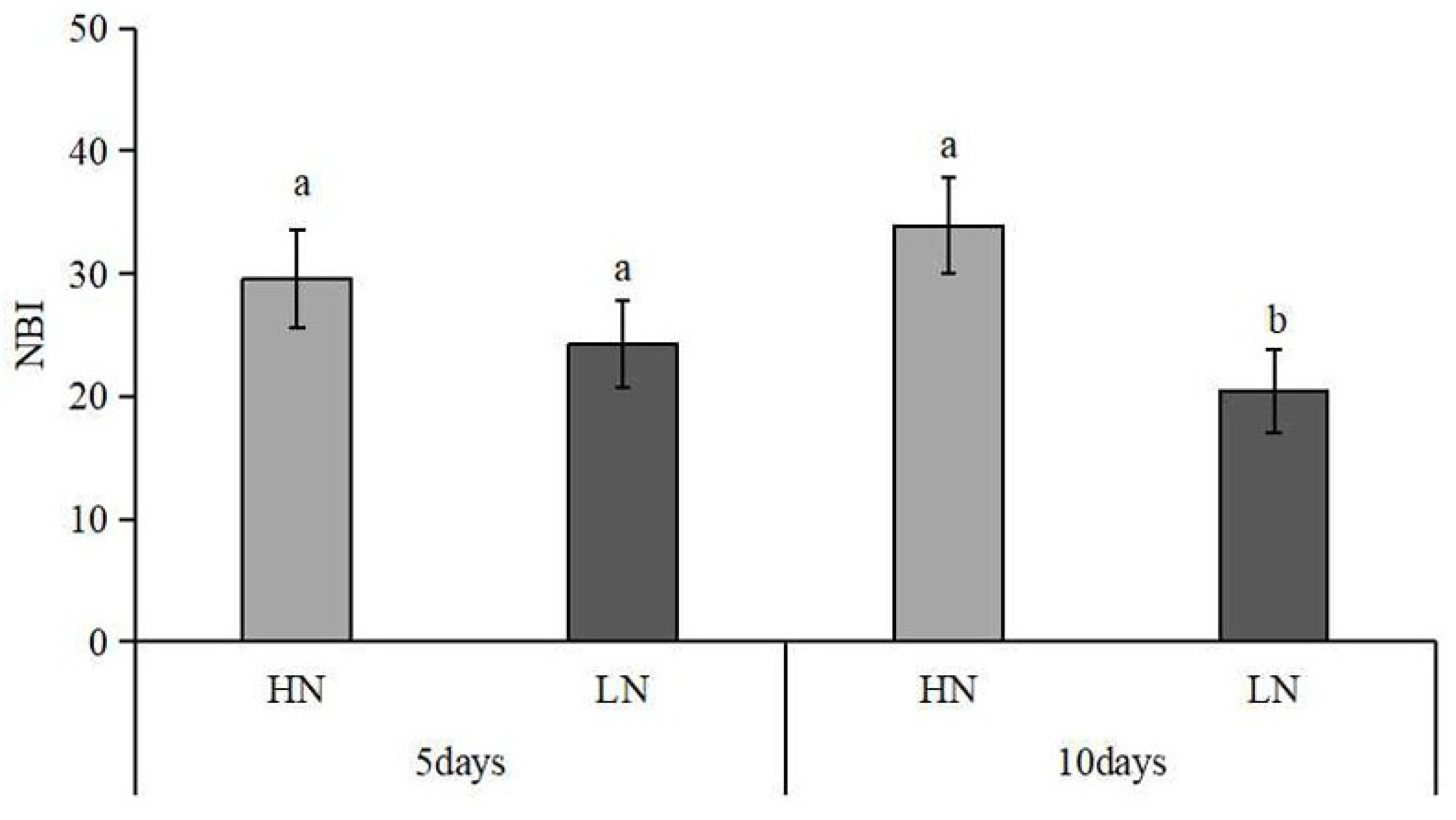
2.2. Nitrogen Deficiency Affected the Nitrogen Content of Peanut Leaves
2.3. Nitrogen Deficiency Affected the Activities of Nitrate Assimilation Enzymes in Peanut Roots
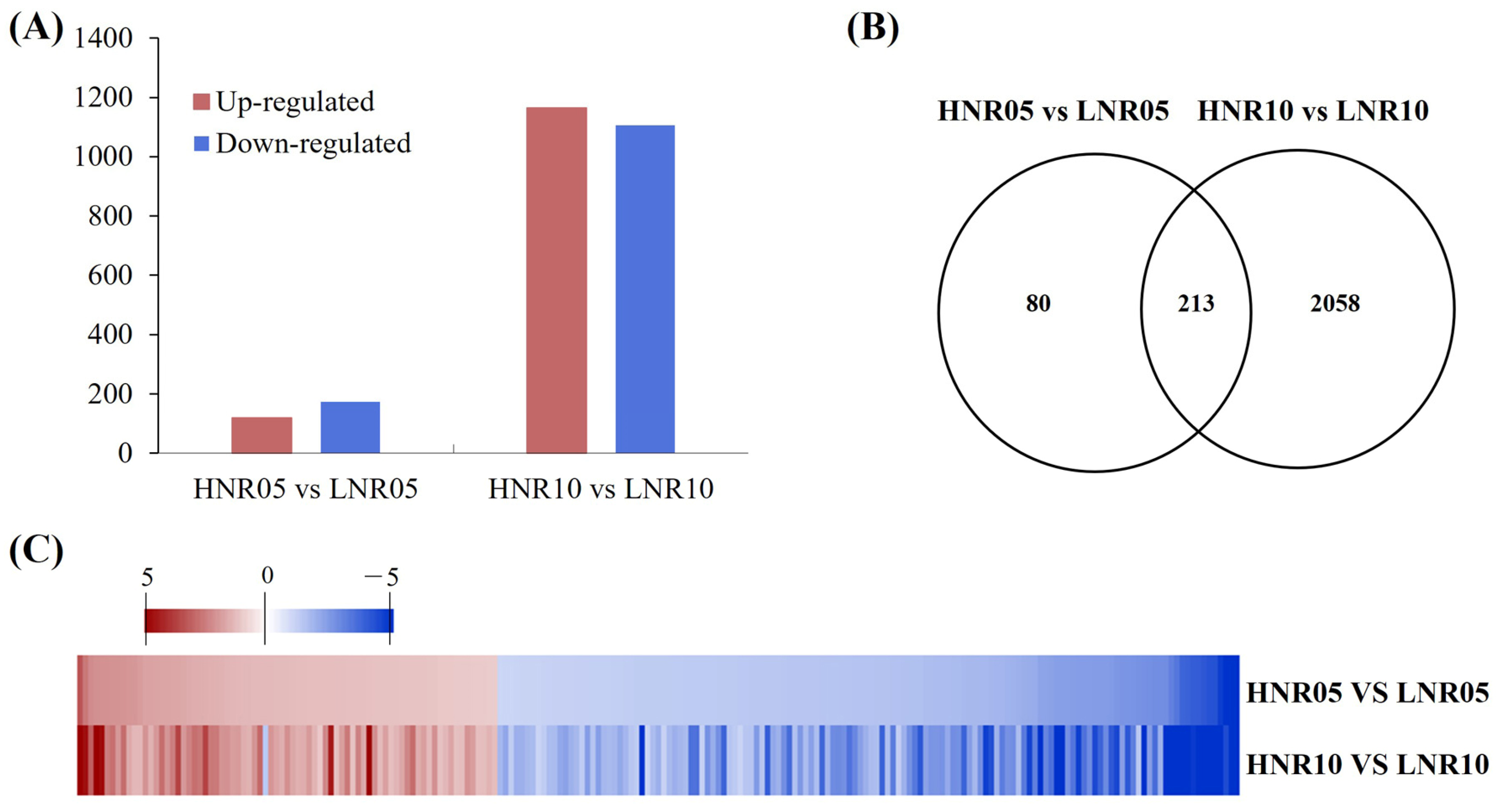
2.4. Nitrogen Deficiency Induced Aberrant Expression of Genes
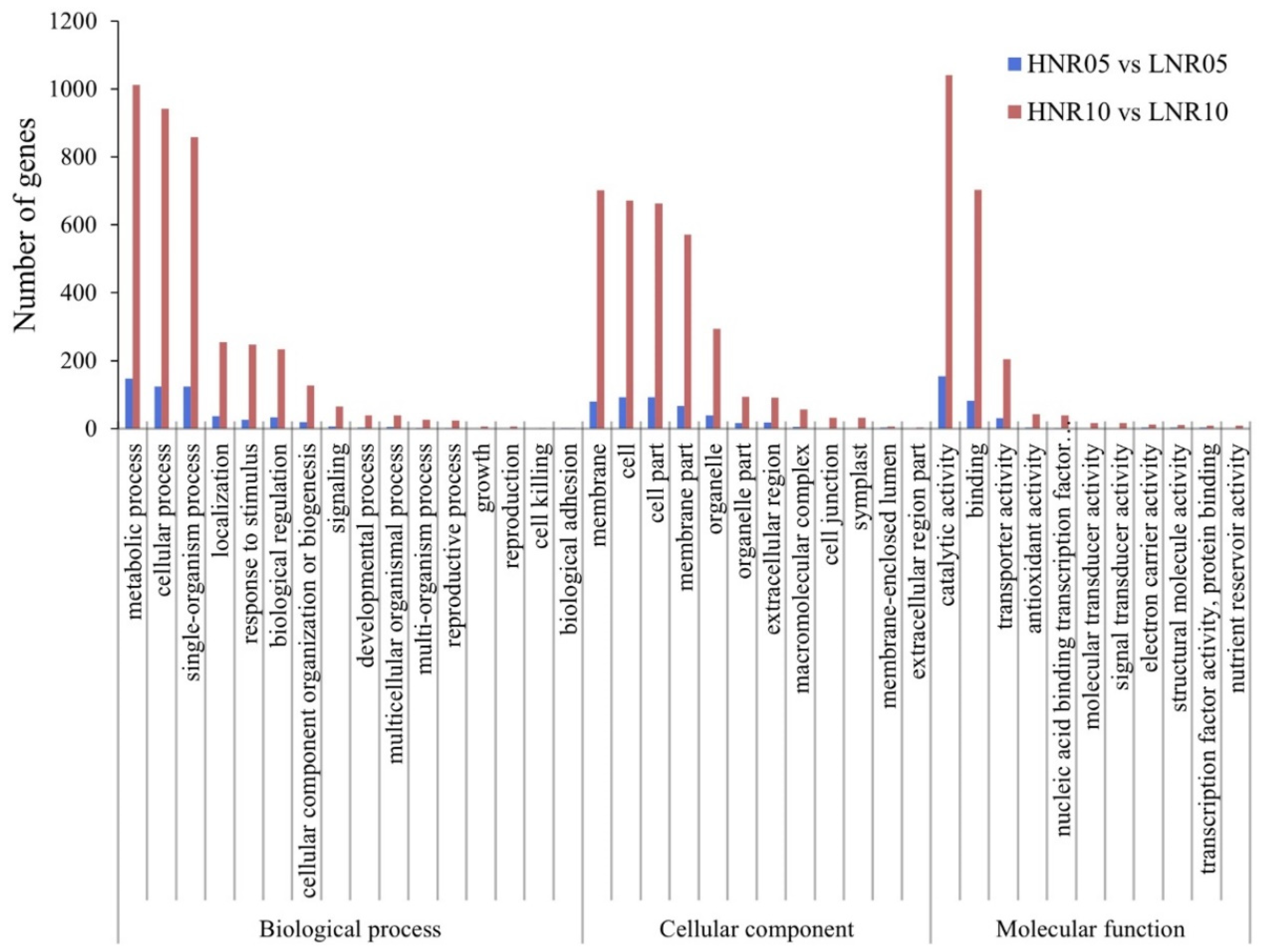
2.5. GO and KEGG Annotation Analysis of DEGs
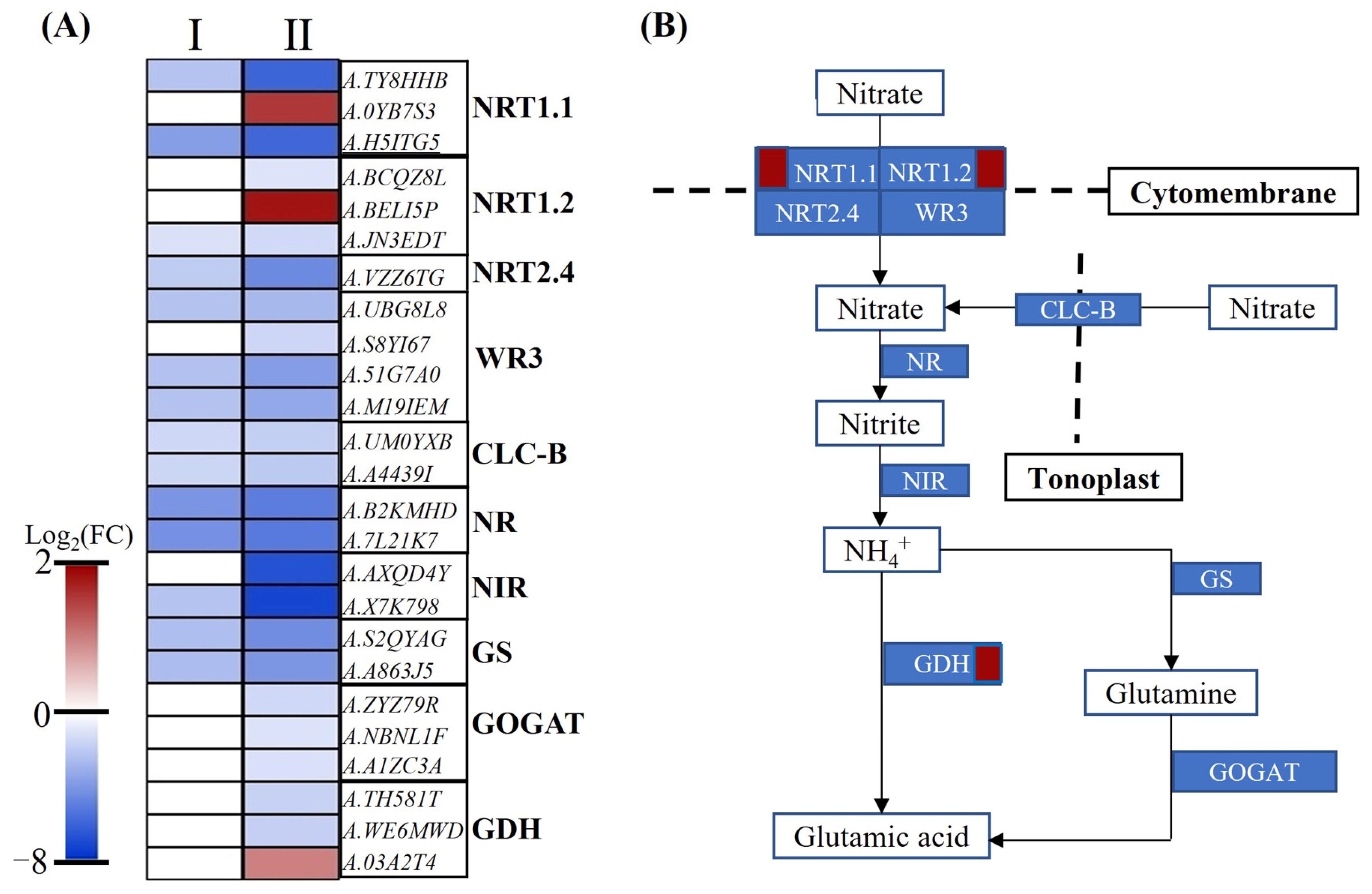
2.6. DEGs Participated in Nitrate Transportation and Assimilation
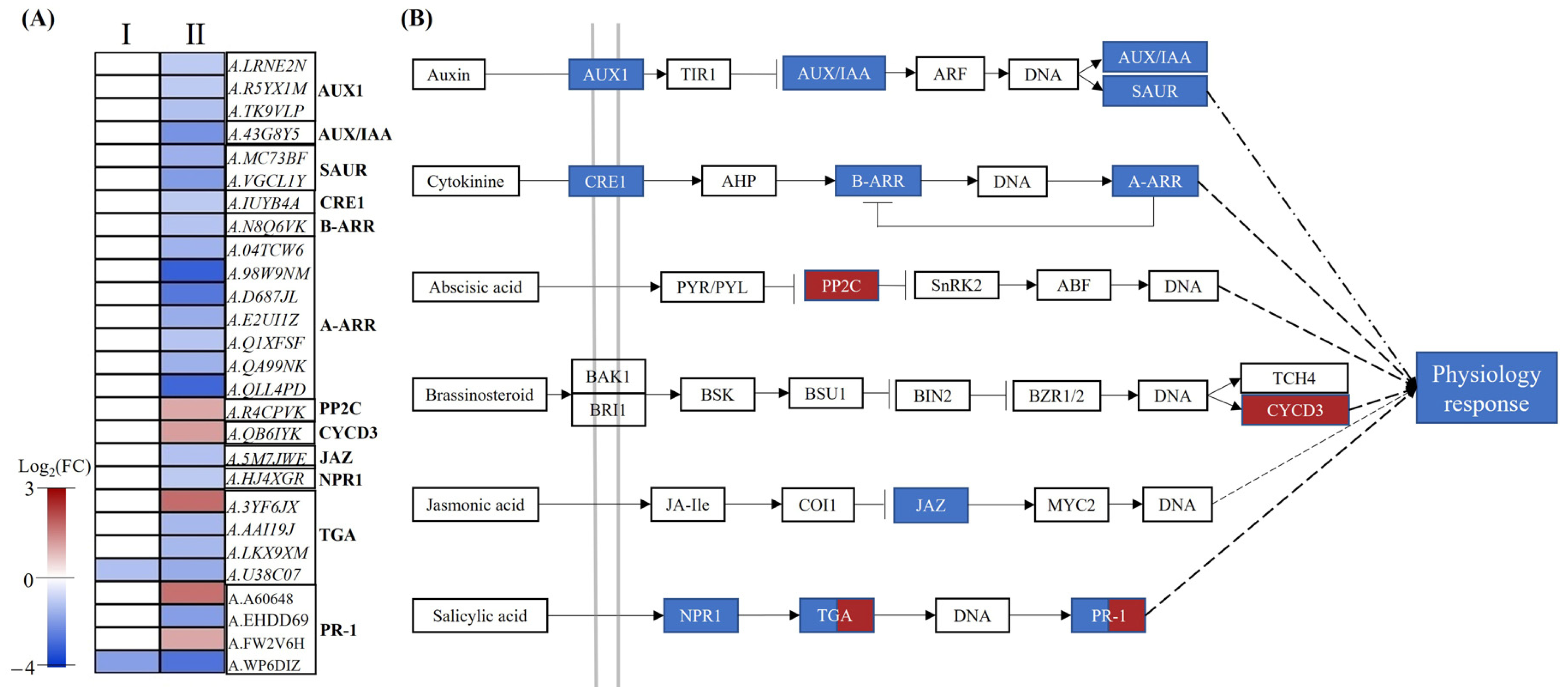
2.7. DEGs Related to Plant Hormone Signal Transduction
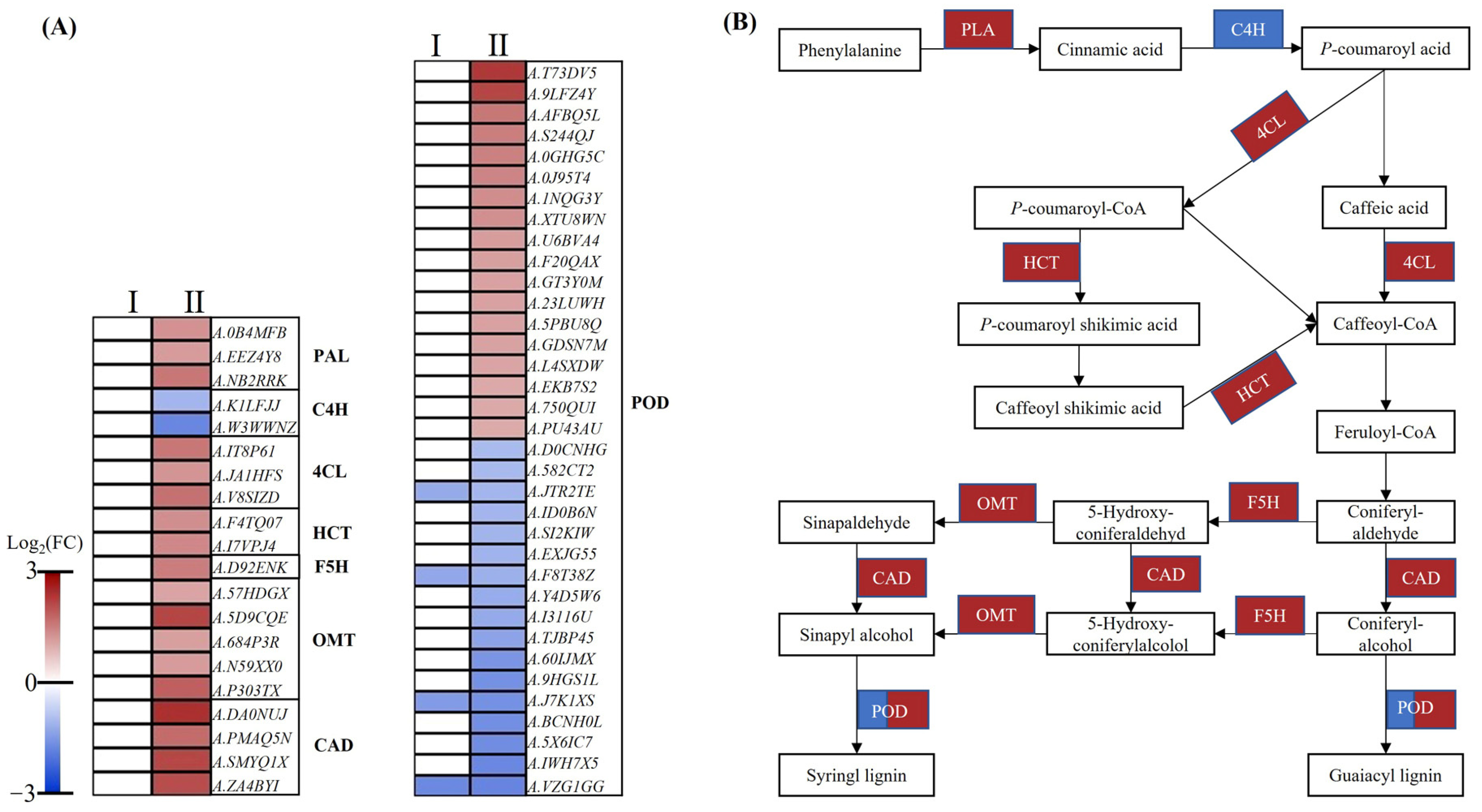
2.8. DEGs Involved in Lignin Biosynthetic Pathway
2.9. DEGs Encoded Transcription Factors (TFs)
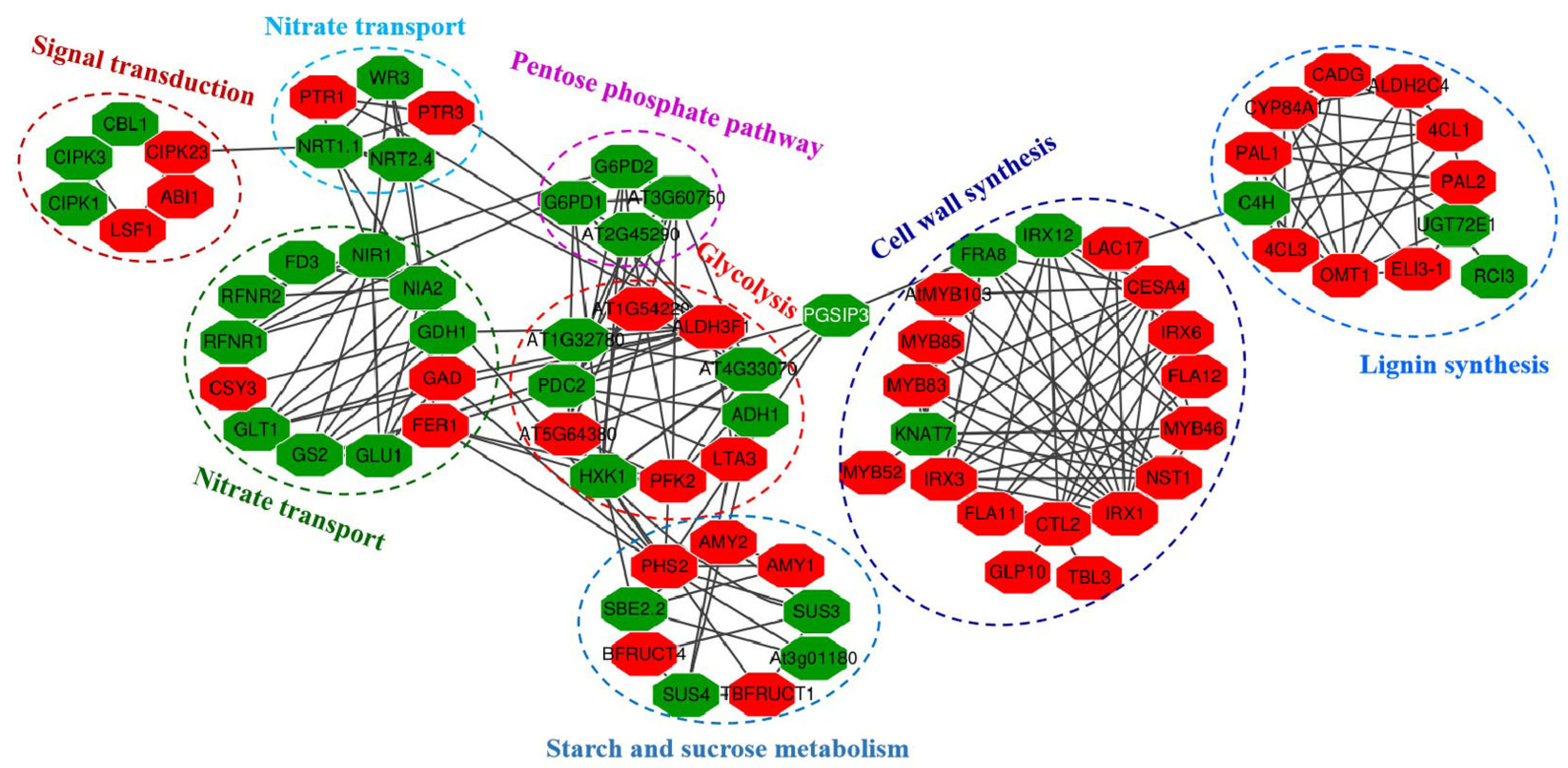
2.10. Putative Interaction Networks of Identified DEGs
2.11. Verification of RNA-Seq by qRT-PCR
3. Discussion
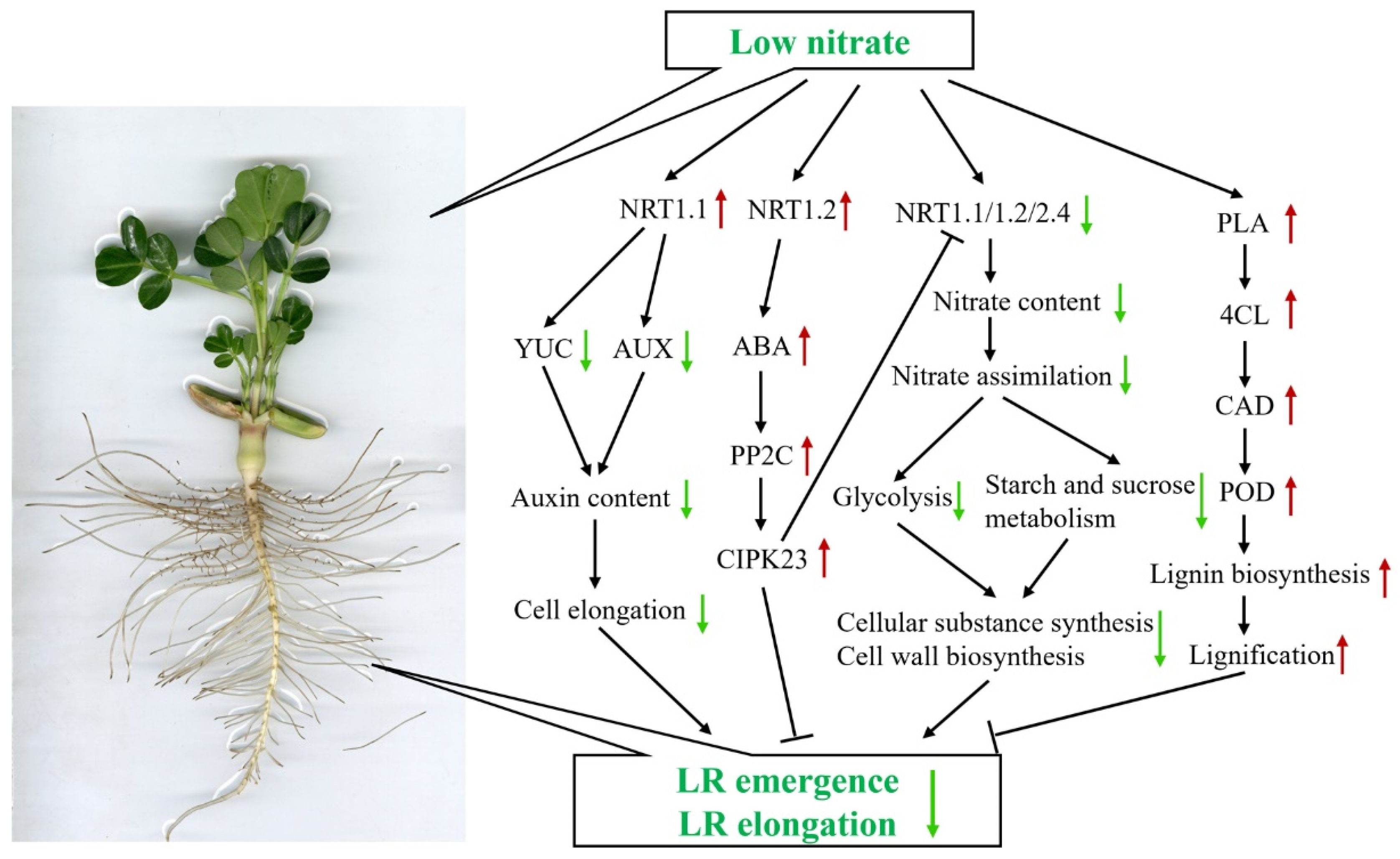
3.1. Nitrate Acting as a Signal Regulating Root Morphology in Peanut Seedlings under Low-Nitrogen Condition
3.2. The Adaptive Mechanisms of Nitrate Transport and Assimilation under Low-Nitrate Condition
3.3. The Mechanism of Plant Hormones Regulating Root Growth under Low-Nitrate Condition
3.4. The Relationship between Lignin Synthesis and Root Growth under Low-Nitrate Condition
3.5. The Response of TFs under Low-Nitrate Condition and Their Relationship with Root Growth
3.6. The Adaptive Mechanisms of Carbon Metabolism under Low-Nitrate Condition
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Determination of Root Morphology and Dry Weight
4.3. Determination of Nitrogen Balance Index (NBI)
4.4. Determination of Nitrate Assimilation Enzyme Activities
4.5. RNA Isolation and cDNA Library Construction
4.6. Genome Mapping and Differential Expression Analysis
4.7. Functional Annotation of DEGs
4.8. Reverse Transcription PCR and Quantitative Reverse-Transcription PCR (qRT-PCR)
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sun, C.-H.; Yu, J.-Q.; Hu, D.-G. Nitrate: A Crucial Signal during Lateral Roots Development. Front. Plant Sci. 2017, 8, 485. [Google Scholar] [CrossRef]
- Li, S.; Wang, Z.; Hu, T.; Gao, Y.; Stewart, B.A. Nitrogen in dryland soils of China and its management. Adv. Agron. 2009, 101, 123–181. [Google Scholar]
- Gruber, N.; Galloway, J.N. An Earth-system perspective of the global nitrogen cycle. Nature 2008, 451, 293–296. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Li, M.; Liu, Y.; Shi, L. Integration of the metabolome and transcriptome reveals the resistance mechanism to low nitrogen in wild soybean seedling roots. Environ. Exp. Bot. 2020, 175, 104043. [Google Scholar] [CrossRef]
- Konishi, M.; Yanagisawa, S. Emergence of a new step towards understanding the molecular mechanisms under-lying nitrate-regulated gene expression. J. Exp. Bot. 2014, 65, 5589–5600. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Liu, F.; Crawford, N.M.; Wang, Y. Molecular Regulation of Nitrate Responses in Plants. Int. J. Mol. Sci. 2018, 19, 2039. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, Y.; Zhao, Y.; Zhang, Y.; Zhang, J.; Ma, H.; Han, Y. Transcriptome analysis reveals Nitrogen deficiency induced alterations in leaf and root of three cultivars of potato (Solanum tuberosum L.). PLoS ONE 2020, 15, e0240662. [Google Scholar] [CrossRef]
- Ding, H.; Zhang, Z.; Zhang, G.; Xu, Y.; Guo, Q.; Qin, F.; Dai, L. Nitrogen application improved peanut yield and nitrogen use efficiency by optimizing root morphology and distribution under drought stress. Chil. J. Agric. Res. 2022, 82, 256–265. [Google Scholar] [CrossRef]
- Bouguyon, E.; Brun, F.; Meynard, D.; Kubeš, M.; Pervent, M.; Learn, S.; Lacombe, B.; Krouk, G.; Guiderdoni, E.; Zažímalová, E.; et al. Multiple mechanisms of nitrate sensing by Arabidopsis ni-trate transceptor NRT1.1. Nat. Plants 2015, 1, 15015. [Google Scholar] [CrossRef]
- Bouguyon, E.; Perrine-Walker, F.; Pervent, M.; Rochette, J.; Cuesta, C.; Benkova, E.; Martinière, A.; Bach, L.; Krouk, G.; Gojon, A.; et al. Nitrate controls root development through post-transcriptional regulation of the NRT1.1/NPF6.3 transporter/sensor. Plant Physiol. 2016, 172, 1237–1248. [Google Scholar] [CrossRef]
- Lynch, J. Root Architecture and Plant Productivity. Plant Physiol. 1995, 109, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Gifford, M.L.; Banta, J.A.; Katari, M.S.; Hulsmans, J.; Chen, L.; Ristova, D.; Tranchina, D.; Purugganan, M.D.; Coruzzi1, G.M.; Birnbaum, K.D. Plasticity regulators modulate specific root traits in discrete nitrogen environ-ments. PLoS Genet. 2013, 9, e1003760. [Google Scholar] [CrossRef] [PubMed]
- Krouk, G.; Lacombe, B.; Bielach, A.; Perrine-Walker, F.; Malinska, K.; Mounier, E.; Hoyerova, K.; Tillard, P.; Leon, S.; Ljung, K.; et al. Nitrate-regulated auxin transport by NRT1.1 de-fines a mechanism for nutrient sensing in plants. Dev. Cell. 2010, 18, 927–937. [Google Scholar] [CrossRef] [PubMed]
- Gruber, B.D.; Giehl, R.F.; Friedel, S.; von Wirén, N. Plasticity of the Arabidopsis root system under nutrient defi-ciencies. Plant Physiol. 2013, 163, 161–179. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.; Walk, T.C.; Han, P.; Chen, L.; Zhang, S.; Li, Y.; Hu, X.; Xie, L.; Yang, Y.; Liu, J.; et al. Adaption of roots to nitrogen deficiency revealed by 3D quantification and proteomic analysis. Plant Physiol. 2019, 179, 329–347. [Google Scholar] [CrossRef]
- Wang, J.; Song, K.; Sun, L.; Qin, Q.; Pan, J.; Xue, Y. Morphological and Transcriptome Analysis of Wheat Seedlings Response to Low Nitrogen Stress. Plants 2019, 8, 98. [Google Scholar] [CrossRef]
- Li, L.; Li, Q.; Davis, K.E.; Patterson, C.; Oo, S.; Liu, W.; Liu, J.; Wang, G.; Fontana, J.E.; Thornburg, T.E.; et al. Response of root growth and development to nitrogen and potassium deficiency as well as microRNA-mediated mechanism in peanut (Arachis hypogaea L.). Front. Plant Sci. 2021, 12, 695234. [Google Scholar] [CrossRef]
- Mounier, E.; Pervent, M.; Ljung, K.; Gojon, A.; Nacry, P. Auxin-mediated nitrate signalling by NRT1.1 partici-pates in the adaptive response of Arabidopsis root architecture to the spatial heterogeneity of nitrate availability. Plant Cell Environ. 2014, 37, 162–174. [Google Scholar] [CrossRef]
- Vidal, E.A.; Araus, V.; Lu, C.; Parry, G.; Green, P.J.; Coruzzi, G.M.; Gutiérreza, R.A. Nitrate-responsive miR393/AFB3 regulatory module controls root system architecture in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2010, 107, 4477–4482. [Google Scholar] [CrossRef]
- von Behrens, I.; Komatsu, M.; Zhang, Y.; Berendzen, K.W.; Niu, X.; Sakai, H.; Taramino, G.; Hochholdinger, F. Rootless with undetectable meristem 1 encodes a monocot-specific AUX/IAA protein that controls embryonic seminal and post-embryonic lateral root initiation in maize. Plant J. 2011, 66, 341–353. [Google Scholar] [CrossRef]
- Gelli, M.; Duo, Y.; Konda, A.R.; Zhang, C.; Holding, D.; Dweikat, I. Identification of differentially expressed genes between sorghum genotypes with contrasting nitrogen stress tolerance by genome-wide transcriptional profiling. BMC Genom. 2014, 15, 179. [Google Scholar] [CrossRef] [PubMed]
- Boddey, R.M.; Urquiaga, S.; Neves, M.C.P. Quantification of the contribution of N2 fixation to field-grown grain legumes—A strategy for the practical application of the 15N isotope dilution technique. Soil Biol. Biochem. 1990, 22, 649–655. [Google Scholar] [CrossRef]
- Lanier, J.E.; Jordan, D.L.; Spears, J.F.; Wells, R.; Johnson, P.D. Peanut response to inoculation and nitrogen fertilizer. Agronomy J. 2005, 97, 79–84. [Google Scholar] [CrossRef]
- Gohari, A.A.; Niyaki, S.A.N. Effects of iron and nitrogen fertilizers on yield and yield components of peanut (Arachis hypogaea L.) in Astaneh Ashrafiyeh, Iran. Am. Eurasian J. Agric. Environ. Sci. 2010, 9, 256–262. [Google Scholar]
- Araya, T.; Miyamoto, M.; Wibowo, J.; Suzuki, A.; Kojima, S.; Tsuchiya, Y.N.S.; Fukuda, H.; Wirén, N.V.; Takahashi, H. CLE-CLAVATA1 peptide-receptor signaling module regulates the expansion of plant root systems in a nitrogen-dependent manner. Proc. Natl. Acad. Sci. USA 2014, 111, 2029–2034. [Google Scholar] [CrossRef]
- Forde, B.G. Nitrogen signalling pathways shaping root system architecture: An update. Curr. Opin. Plant Biol. 2014, 21, 30–36. [Google Scholar] [CrossRef]
- Remans, T.; Nacry, P.; Pervent, M.; SFilleur, S.; Diatloff, E.; Mounier, E.; Tillard, P.; Forde, B.G.; Gojon, A. The Arabidopsis NRT1.1 transporter participates in the signaling pathway triggering root colonization of ni-trate-rich patches. Proc. Natl. Acad. Sci. USA 2006, 103, 19206–19211. [Google Scholar] [CrossRef]
- Ho, C.-H.; Lin, S.-H.; Hu, H.-C.; Tsay, Y.-F. CHL1 Functions as a Nitrate Sensor in Plants. Cell 2009, 138, 1184–1194. [Google Scholar] [CrossRef]
- Kanno, Y.; Hanada, A.; Chiba, Y.; Ichikawa, T.; Nakazawa, M.; Matsui, M.; Koshiba, T.; Kamiya, Y.; Seo, M. Iden-tification of an abscisic acid transporter by unctional screening using the receptorcomplex as a sensor. Proc. Natl. Acad. Sci. USA 2012, 109, 9653–9658. [Google Scholar] [CrossRef]
- Guo, D.; Liang, J.; Li, L. Abscisic acid (ABA) inhibition of lateral root formation involves endogenous ABA bio-synthesis in Arachis hypogaea L. Plant Growth Regul. 2009, 58, 173–179. [Google Scholar] [CrossRef]
- Crawford, N.M.; Glass, A.D. Molecular and physiological aspects of nitrate uptake in plants. Trends Plant Sci. 1998, 3, 389–395. [Google Scholar] [CrossRef]
- Fan, X.; Feng, H.; Tan, Y.; Xu, Y.; Miao, Q.; Xu, G. A putative 6-transmembrane nitrate transporter OsNRT1.1b plays a key role in rice under low nitroge. J. Integr. Plant Biol. 2016, 58, 590–599. [Google Scholar] [CrossRef]
- Kiba, T.; Feria-Bourrellier, A.-B.; Lafouge, F.; Lezhneva, L.; Boutet-Mercey, S.; Orsel, M.; Bréhaut, V.; Miller, A.; Daniel-Vedele, F.; Sakakibara, H.; et al. The Arabidopsis Nitrate Transporter NRT2.4 Plays a Double Role in Roots and Shoots of Nitrogen-Starved Plants. Plant Cell 2012, 24, 245–258. [Google Scholar] [CrossRef]
- Geelen, D.; Lurin, C.; Bouchez, D.; Frachisse, J.M.; Lelievre, F.; Courtial, B. Barbier-Brygoo, H.; Maurel, C. Dis-ruption of putative anion channel gene AtCLC-a in Arabidopsis suggests a role in the regulation of nitrate con-tent. Plant J. 2000, 21, 259–267. [Google Scholar] [CrossRef]
- Ferrario-Méry, S.; Masclaux, C.; Suzuki, A.; Valadier, M.H.; Hirel, B.; Foyer, C.H. Glutamine and alpha-ketoglutarate are metabolite signals involved in nitrate reductase gene transcription in untransformed and transformed tobacco plants deficient in ferredoxin-glutamine-alpha-ketoglutarate aminotransferase. Planta 2001, 213, 265–271. [Google Scholar] [CrossRef]
- Song, T.; Das, D.; Yang, F.; Chen, M.; Tian, Y.; Cheng, C.; Sun, C.; Xu, W.; Zhang, J. Genome-wide transcriptome analysis of roots in two rice varieties in response to alternate wetting and drying irrigation. Crop. J. 2020, 8, 586–601. [Google Scholar] [CrossRef]
- Fukaki, H.; Tasaka, M. Hormone interactions during lateral root formation. Plant Mol. Biol. 2008, 69, 437–449. [Google Scholar] [CrossRef]
- Marchant, A.; Bhalerao, R.; Casimiro, I.; Eklöf, J.; Casero, P.J.; Bennett, M.; Sandberg, G. AUX1 promotes lateral root formation by facilitating indole-3-acetic acid distribution between sink and source tissues in the Arabidopsis seedling. Plant Cell 2002, 14, 589–597. [Google Scholar] [CrossRef]
- Swarup, K.; Benková, E.; Swarup, R.; Casimiro, I.; Péret, B.; Yang, Y.; Parry, G.; Nielsen, E.; De Smet, I.; Vanneste, S.; et al. The auxin influx carrier LAX3 promotes lateral root emergence. Nature 2008, 10, 946–954. [Google Scholar] [CrossRef]
- Peng, Y.; Li, X.; Yuan, L.; Li, C. A novel morphological response of maize (Zea mays) adult roots to heterogene-ous nitrate supply revealed by a split root experiment. Physiol. Plantarum. 2013, 150, 133–144. [Google Scholar]
- Joshi-Saha, A.; Valon, C.; Leung, J. A Brand New START: Abscisic Acid Perception and Transduction in the Guard Cell. Sci. Signal. 2011, 29, 201. [Google Scholar] [CrossRef]
- Leran, S.; Edel, K.H.; Pervent, M.; Hashimoto, K.; Corratge-Faillie, C.; Offenborn, J.N.; Tillard, P.; Gojon, A.; Kudla, J.; Lacombe, B. Nitrate sensing and uptake in Arabidopsis are enhanced by ABI2, a phosphatase inactivated by the stress hormone abscisic acid. Sci. Signal. 2015, 8, ra43. [Google Scholar] [CrossRef]
- Park, H.L.; Bhoo, S.H.; Kwon, M.; Lee, S.W.; Cho, M.H. Biochemical and expression analyses of the rice cin-namoyl-CoA reductase gene family. Front. Plant Sci. 2017, 8, 2099. [Google Scholar] [CrossRef]
- Huang, W.-N.; Liu, H.-K.; Zhang, H.-H.; Chen, Z.; Guo, Y.-D.; Kang, Y.-F. Ethylene-induced changes in lignification and cell wall-degrading enzymes in the roots of mungbean (Vigna radiata) sprouts. Plant Physiol. Biochem. 2013, 73, 412–419. [Google Scholar] [CrossRef]
- Martin, C. Transcription factors and the manipulation of plant traits. Curr. Opin. Biotechnol. 1996, 7, 130–138. [Google Scholar] [CrossRef]
- Lee, H.W.; Kim, N.Y.; Lee, D.J.; Kim, J. LBD18/ASL20 regulates lateral root formation in combination with LBD16/ASL18 downstream of ARF7 and ARF19 in Arabidopsis. Plant Physiol. 2009, 151, 1377–1389. [Google Scholar] [CrossRef]
- Rubin, G.; Tohge, T.; Matsuda, F.; Saito, K.; Scheible, W. Members of the LBD Family of Transcription factors repress anthocyanin synthesis and affect additional nitrogen responses in Arabidopsis. Plant Cell 2009, 21, 3567–3584. [Google Scholar] [CrossRef]
- Liu, X.; Yin, C.; Xiang, L.; Jiang, W.; Xu, S.; Mao, Z. Transcription strategies related to photosynthesis and nitrogen metabolism of wheat in response to nitrogen deficiency. BMC Plant Biol. 2020, 20, 448. [Google Scholar] [CrossRef]
- Li, J.; Baroja-Fernández, E.; Bahaji, A.; Muñoz, F.J.; Ovecka, M.; Montero, M.; Sesma, M.T.; Alonso-Casajús, N.; Almagro, G.; Sánchez-López, A.M.; et al. Enhancing sucrose syn-thase activity results in increased levels of starch and ADP-glucose in maize (Zea mays L.) seed endosperms. Plant Cell Physiol. 2013, 54, 282–294. [Google Scholar] [CrossRef]
- Chandra, A.; Jain, R.; Solomon, S. Complexities of invertases controlling sucrose accumulation and retention in sugarcane. Curr. Sci. 2012, 102, 857–866. [Google Scholar]
- Zhang, H.; Hou, J.; Liu, J.; Zhang, J.; Song, B.; Xie, C. The roles of starch metabolic pathways in the cold-induced sweetening process in potatoes. Starch-Starke 2017, 69, 1600194. [Google Scholar] [CrossRef]
- Akgul, R.; Morgil, H.; Kizilkaya, I.T.; Sarayloo, E.; Cevahir, G.; Akgul, F.; Kavakli, I.H. Transcriptomic and fatty acid analyses of Neochloris aquatica grown under different nitrogen concentration. Funct. Integr. Genom. 2022, 22, 407–421. [Google Scholar] [CrossRef]
- Li, C.; Brant, E.; Budak, H.; Zhang, B. CRISPR/Cas: A Nobel Prize award-winning precise genome editing technology for gene therapy and crop improvement. J. Zhejiang Univ. Sci. B 2021, 22, 253–284. [Google Scholar] [CrossRef]
- Zhang, D.; Zhang, Z.; Unver, T.; Zhang, B. CRISPR/Cas: A powerful tool for gene function study and crop improvement. J. Adv. Res. 2021, 29, 207–221. [Google Scholar] [CrossRef]
- Fontana, J.E.; Wang, G.; Sun, R.; Xue, H.; Li, Q.; Liu, J.; Davis, K.E.; Thornburg, T.E.; Zhang, B.; Zhang, Z.; et al. Impact of potassium deficiency on cotton growth, development and potential microRNA-mediated mechanism. Plant Physiol. Biochem. 2020, 153, 72–80. [Google Scholar] [CrossRef]
- Jin, C.; Du, S.T.W.; Shamsi, I.H.; Luo, B.; Lin, X. NO synthase-generated NO acts downstream of auxin in regulating Fe-deficiency-induced root branching that enhances Fe-deficiency tolerance in tomato plants. J. Exp. Bot. 2011, 62, 3875–3884. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Florea, L.; Song, L.; Salzberg, S.L. Thousands of exon skipping events differentiate among splicing patterns in sixteen human tissues. F1000 Res. 2013, 2, 188. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Oshlack, A. Gene ontology analysis for RNA-seq: Accounting for selection bias. Genome Biol. 2010, 11, R14. [Google Scholar] [CrossRef]
- Mao, X.; Cai, T.; Olyarchuk, J.G.; Wei, L. Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 2005, 21, 3787–3793. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.; Schmittgen, T. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2000, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Morgante, C.V.; Guimarães, P.M.; Martins, A.C.; Araújo, A.C.; Leal-Bertioli, S.C.; Bertioli, D.J.; Brasileiro, A.C. Reference genes for quantitative reverse transcription-polymerase chain reaction expression studies in wild and cultivated peanut. BMC Res. Notes 2011, 4, 339. [Google Scholar] [CrossRef] [PubMed]

| Treatment | Processing Time (Day) | RDW (g/Plant) | TRL (cm/Plant) | TRSA (cm2/Plant) | TRV (cm3/Plant) | NLR (No./Plant) | LRD (cm/Plant) | PRL (cm/Plant) |
|---|---|---|---|---|---|---|---|---|
| HN | 5 | 0.12 ± 0.03 a | 478.32 ± 41.36 a | 99.47 ± 13.36 a | 1.65 ± 0.30 a | 86 ± 9 a | 13.2 ± 0.51 a | 17.8 ± 1.06 a |
| LN | 0.11 ± 0.02 a | 456.44 ± 25.48 a | 92.88 ± 9.47 a | 1.51 ± 0.22 a | 83 ± 3 a | 13.0 ± 1.24 a | 16.6 ± 1.40 a | |
| HN | 10 | 0.23 ± 0.02 a | 1315.93 ± 82.63 a | 236.63 ±12.08 a | 3.39 ± 0.15 a | 133 ± 9 a | 22.77 ± 0.38 a | 29.1 ± 2.57 a |
| LN | 0.17 ± 0.03 b | 915.42 ± 47.37 b | 161.11 ± 12.29 b | 2.26 ± 0.24 b | 110 ± 9 b | 16.0 ± 0.81 b | 19.1 ± 0.55 b |
| Comparisons | KEGG Pathway | KO ID | DEGs | Corrected p-Value |
|---|---|---|---|---|
| HNR05 vs. LNR05 | Starch and sucrose metabolism | ko00500 | 14 | 8.55 × 10−3 |
| Pentose and glucuronate interconversions | ko00040 | 13 | 2.98 × 10−5 | |
| Phenylpropanoid biosynthesis | ko00940 | 11 | 4.67 × 10−3 | |
| Glutathione metabolism | ko00480 | 9 | 1.65 × 10−3 | |
| Nitrogen metabolism | ko00910 | 6 | 7.95 × 10−4 | |
| HNR10 vs. LNR10 | Phenylpropanoid biosynthesis | ko00940 | 71 | 0 |
| Starch and sucrose metabolism | ko00500 | 58 | 1.27 × 10−4 | |
| Glutathione metabolism | ko00480 | 36 | 1.24 × 10−8 | |
| Cysteine and methionine metabolism | ko00270 | 22 | 4.31 × 10−2 | |
| Nitrogen metabolism | ko00910 | 20 | 2.18 × 10−8 | |
| Cyanoamino acid metabolism | ko00460 | 19 | 1.41 × 10−2 | |
| Tyrosine metabolism | ko00350 | 18 | 1.46 × 10−3 | |
| Phenylalanine metabolism | ko00360 | 16 | 1.07 × 10−2 | |
| Isoquinoline alkaloid biosynthesis | ko00950 | 10 | 4.76 × 10−2 | |
| Taurine and hypotaurine metabolism | ko00430 | 8 | 4.54 × 10−2 |
| Gene ID | Protein Identify | HNR05 vs. LNR05 | HNR10 vs. LNR10 | ||
|---|---|---|---|---|---|
| RNA-Seq | qRT-PCR | RNA-Seq | qRT-PCR | ||
| A.P1SHIC | BTB/POZ and TAZ domain-containing protein | −5.41 | −1.73 | −7.26 | −3.49 |
| A.X7K798 | ferredoxin--nitrite reductase | −2.35 | −8.29 | −7.27 | −8.46 |
| A.VZZ6TG | high-affinity nitrate transporter 2.4 | −1.99 | −5.92 | −4.58 | −6.10 |
| A.TY8HHB | protein NRT1/ PTR FAMILY 6.3 | −2.34 | −5.16 | −6.04 | −5.75 |
| A.3895TY | LOB domain-containing protein 38 | −5.29 | −5.52 | −8.71 | −6.75 |
| A.T73DV5 | cationic peroxidase 1-like | - | - | 2.35 | 0.63 |
| A.5D9CQE | anthranilate N-methyltransferase | - | - | 2.18 | 1.26 |
| A.6255L4 | cellulose synthase A catalytic subunit 8 | - | - | 2.02 | 1.83 |
| A.R4CPVK | probable protein phosphatase 2C | - | - | 1.03 | 0.41 |
| A.QB6IYK | cyclin-D3-1 | - | - | 1.15 | 1.17 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, L.; Cheng, X.; Kong, X.; Jia, P.; Wang, X.; Zhang, L.; Zhang, X.; Zhang, Y.; Zhang, Z.; Zhang, B. Comparative Transcriptomic Analysis Reveals the Negative Response Mechanism of Peanut Root Morphology and Nitrate Assimilation to Nitrogen Deficiency. Plants 2023, 12, 732. https://doi.org/10.3390/plants12040732
Li L, Cheng X, Kong X, Jia P, Wang X, Zhang L, Zhang X, Zhang Y, Zhang Z, Zhang B. Comparative Transcriptomic Analysis Reveals the Negative Response Mechanism of Peanut Root Morphology and Nitrate Assimilation to Nitrogen Deficiency. Plants. 2023; 12(4):732. https://doi.org/10.3390/plants12040732
Chicago/Turabian StyleLi, Lijie, Xiangguo Cheng, Xiangjun Kong, Peipei Jia, Xiaohui Wang, Lei Zhang, Xiaotian Zhang, Yi Zhang, Zhiyong Zhang, and Baohong Zhang. 2023. "Comparative Transcriptomic Analysis Reveals the Negative Response Mechanism of Peanut Root Morphology and Nitrate Assimilation to Nitrogen Deficiency" Plants 12, no. 4: 732. https://doi.org/10.3390/plants12040732
APA StyleLi, L., Cheng, X., Kong, X., Jia, P., Wang, X., Zhang, L., Zhang, X., Zhang, Y., Zhang, Z., & Zhang, B. (2023). Comparative Transcriptomic Analysis Reveals the Negative Response Mechanism of Peanut Root Morphology and Nitrate Assimilation to Nitrogen Deficiency. Plants, 12(4), 732. https://doi.org/10.3390/plants12040732







