Characterization of the Isocitrate Dehydrogenase Gene Family and Their Response to Drought Stress in Maize
Abstract
:1. Introduction
2. Results
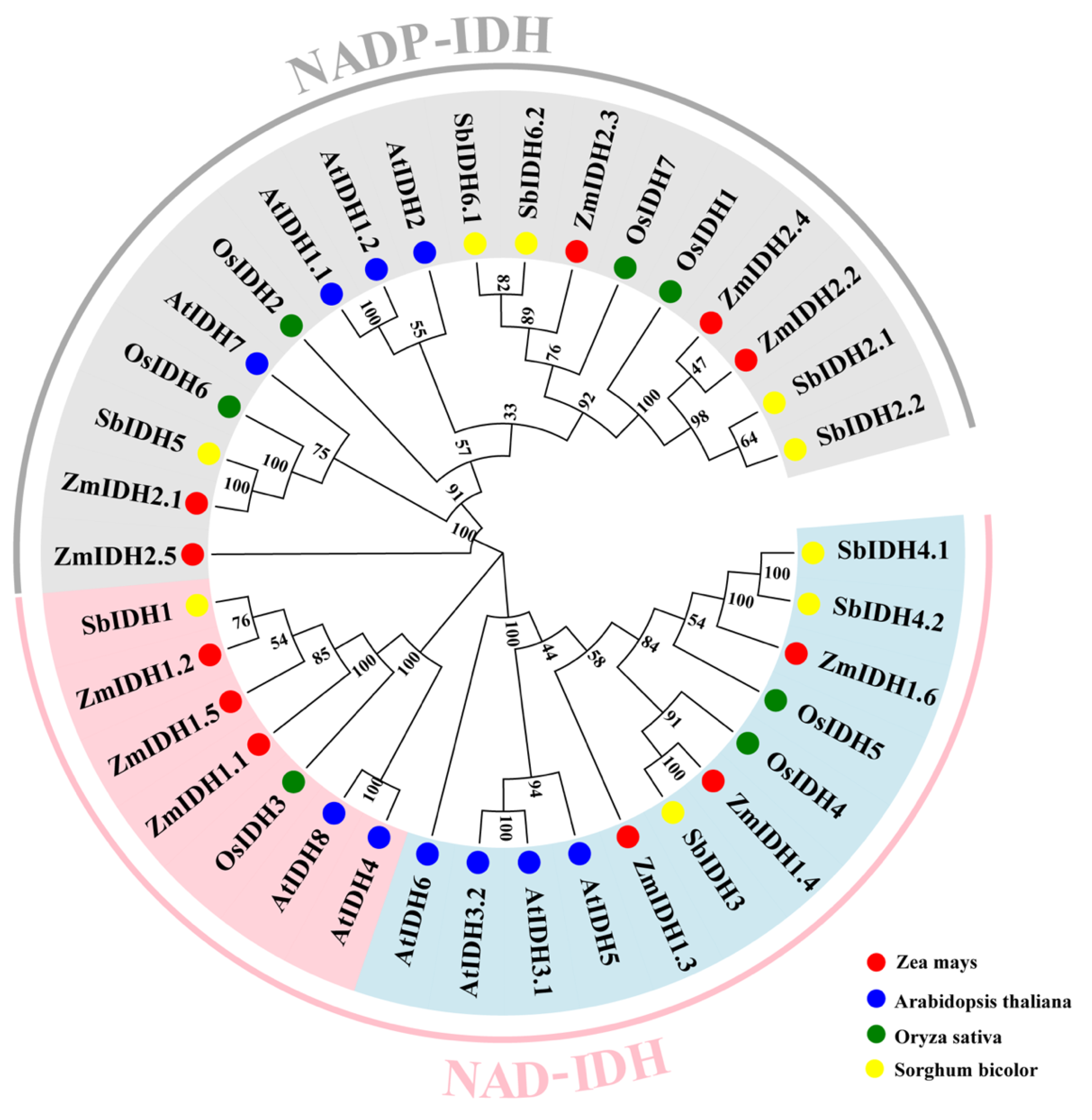
2.1. Identification of ZmIDH in Maize
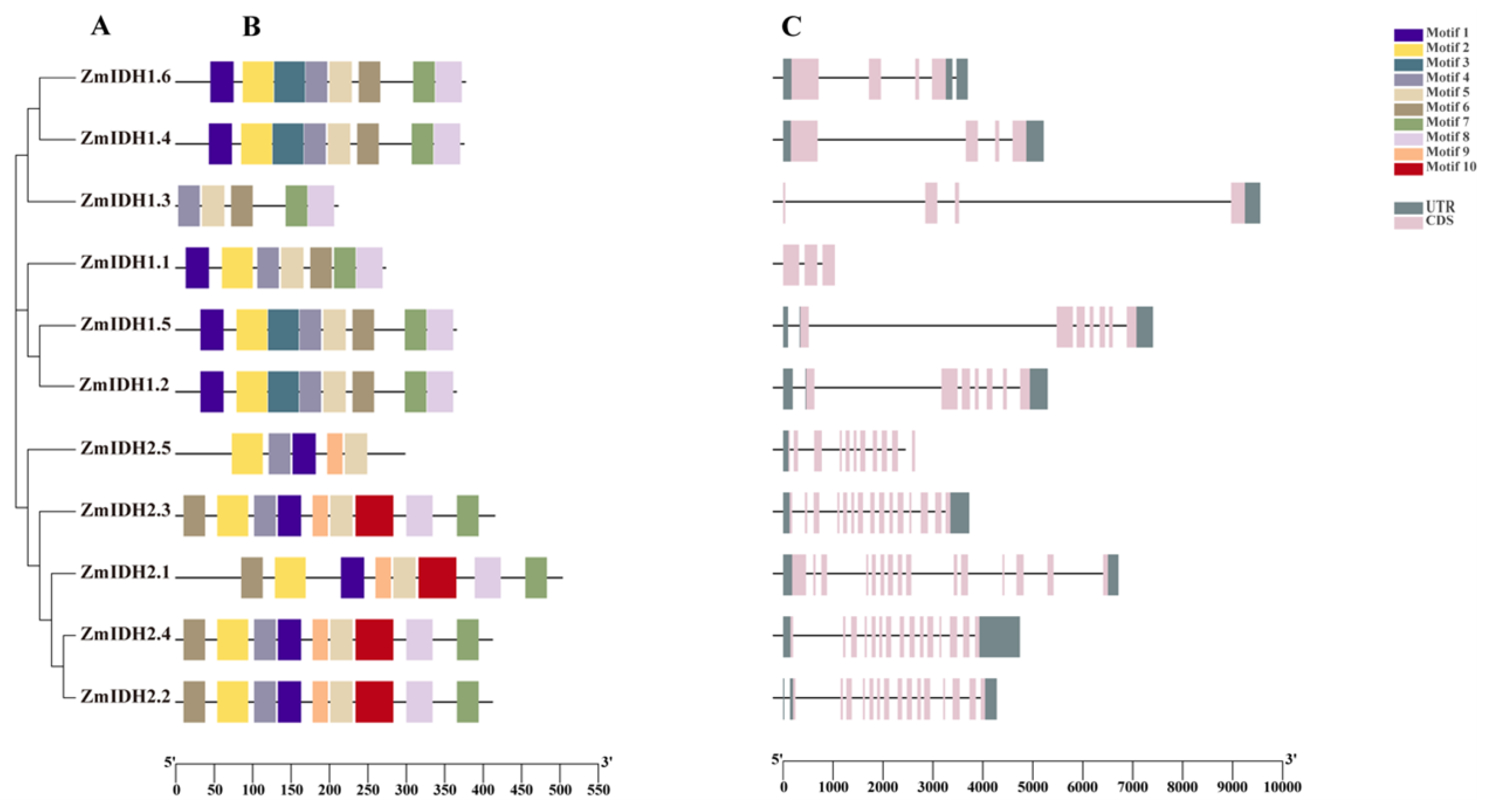
2.2. Gene and Protein Structure Analysis of ZmIDHs
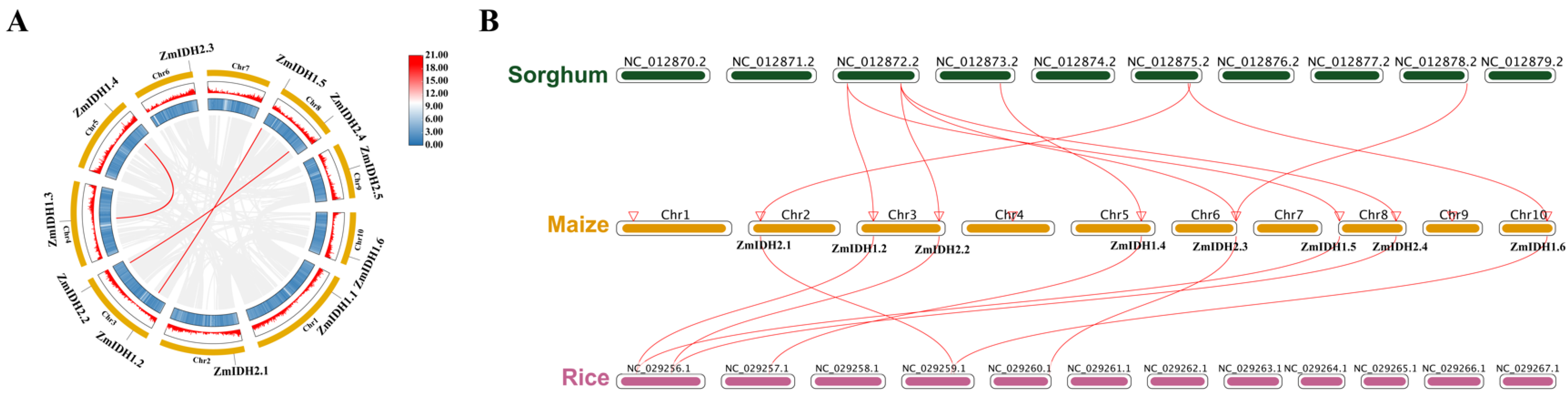
2.3. Colinearity and Duplication of ZmIDH Family Genes
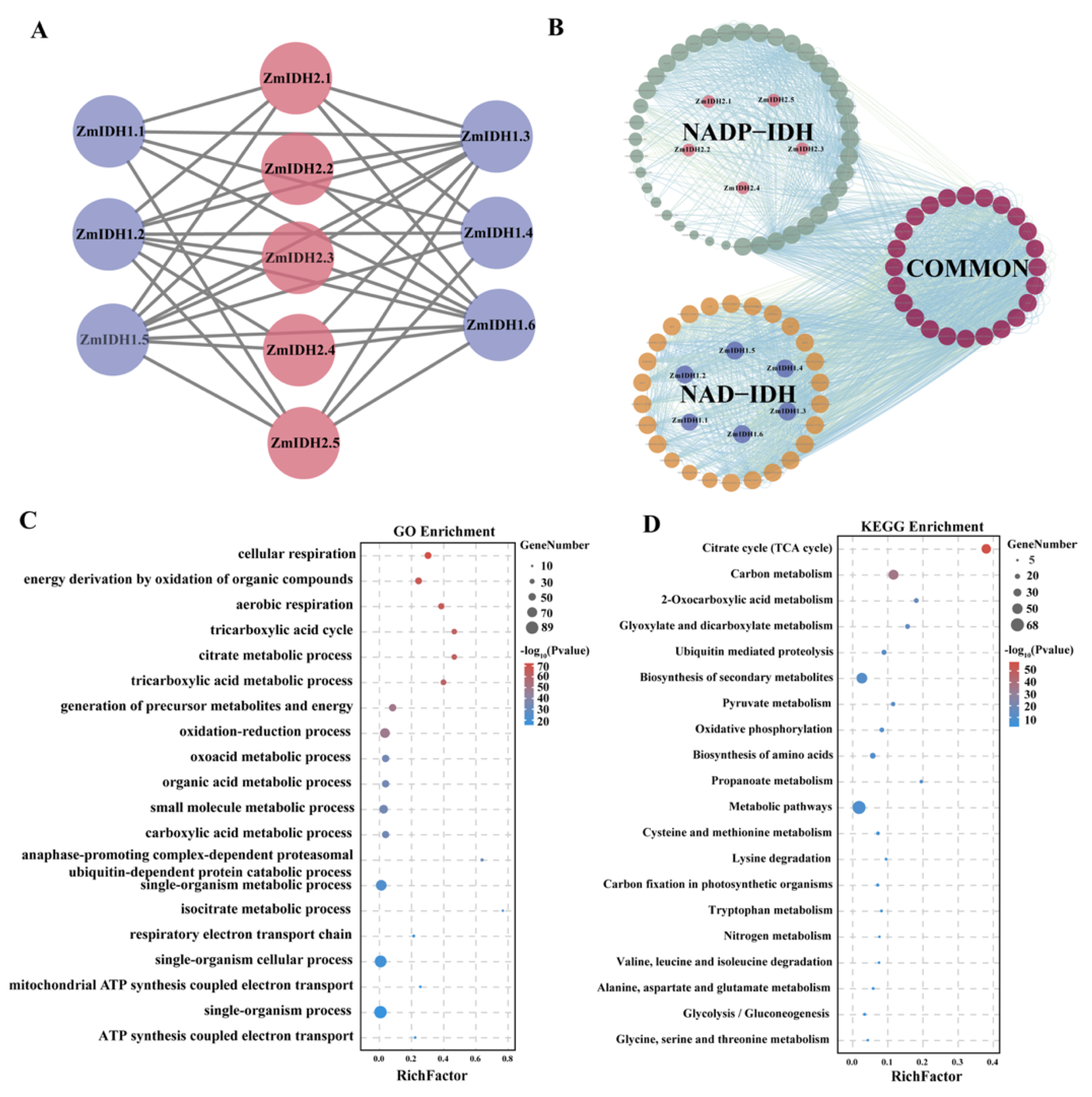
2.4. Analysis of Promoter cis-Elements and the Protein Network of ZmIDH Genes
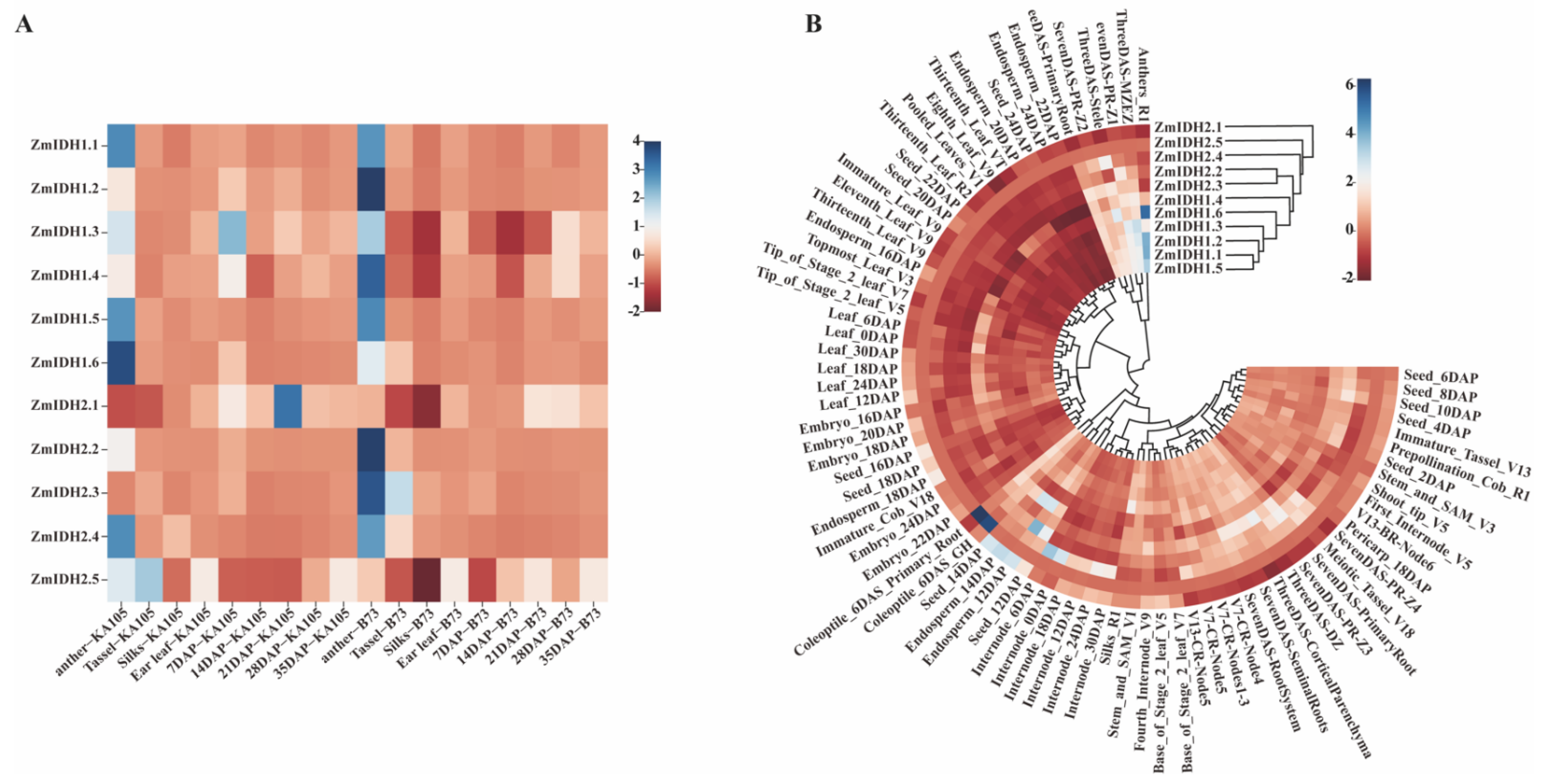
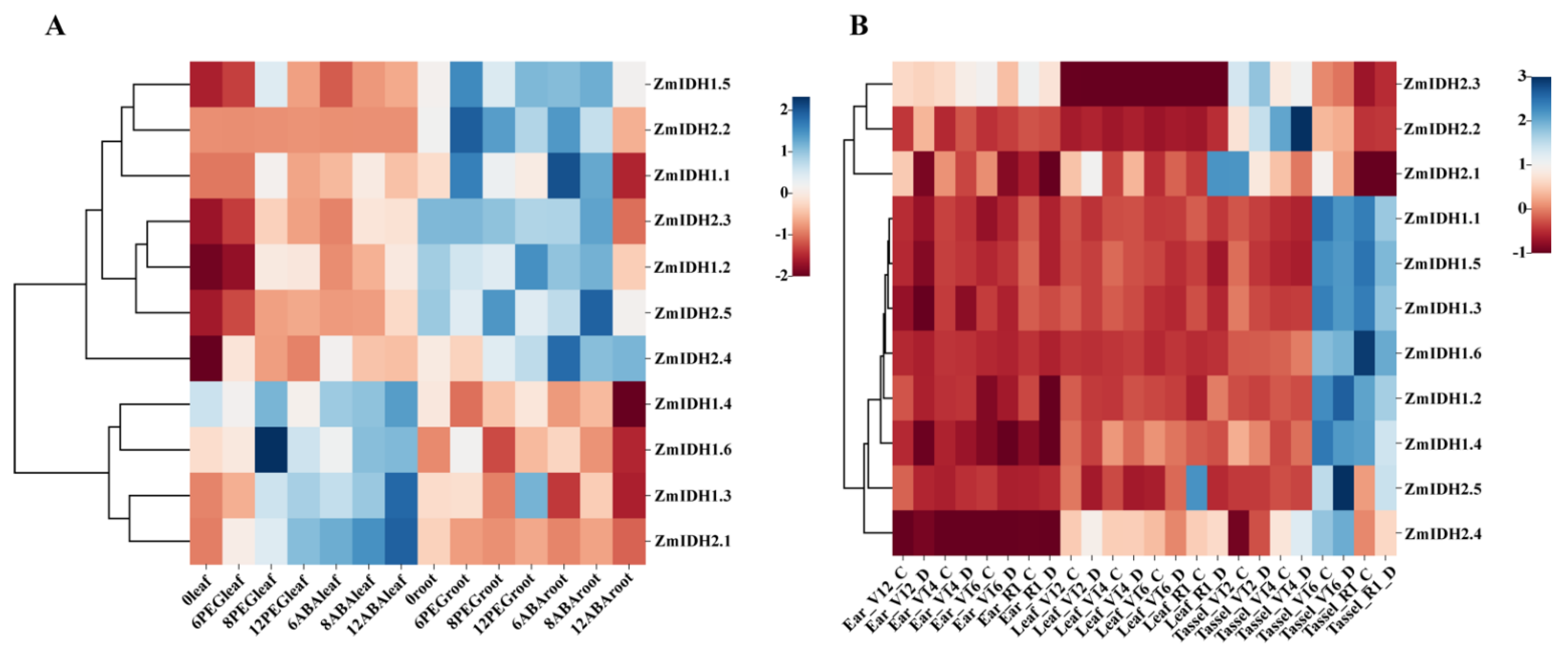
2.5. Expression Patterns of ZmIDHs
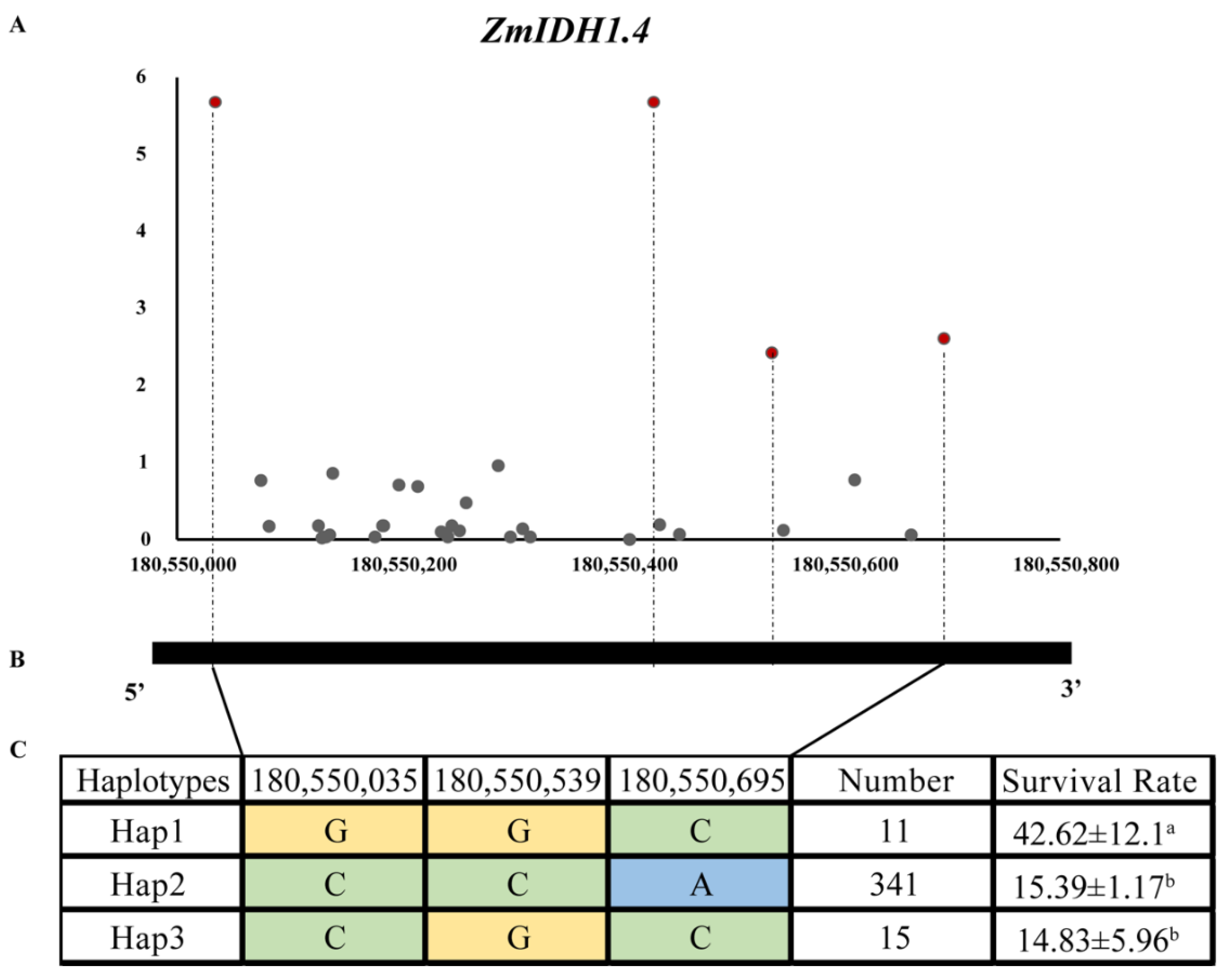
2.6. Candidate Gene Association Analysis of ZmIDHs Gene to Seed Suirvival Rate under Drought
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Identification of IDH Genes
5.2. Characterization of Gene Structure, Protein Tertiary Structure, and Motif Patterns
5.3. Chromosomal Distribution and Gene Duplication of ZmIDH Family Genes
5.4. Prediction and Correlation Analysis of IDH-Interacting Proteins
5.5. Expression Analysis of ZmIDH Genes
5.6. Candidate Gene Association Analysis and Haplotype Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lemaitre, T.; Urbanczyk-Wochniak, E.; Flesch, V.; Bismuth, E.; Fernie, A.R.; Hodges, M. NAD-Dependent Isocitrate Dehydrogenase Mutants of Arabidopsis Suggest the Enzyme Is Not Limiting for Nitrogen Assimilation. Plant Physiol. 2007, 144, 1546–1558. [Google Scholar] [CrossRef] [PubMed]
- Hodges, M.; Flesch, V.; Gálvez, S.; Bismuth, E. Higher plant NADP+-dependent isocitrate dehydrogenases, ammonium assimilation and NADPH production. Plant Physiol. Biochem. 2003, 41, 577–585. [Google Scholar] [CrossRef]
- Akram, M. Citric Acid Cycle and Role of its Intermediates in Metabolism. Cell Biochem. Biophys. 2014, 68, 475–478. [Google Scholar] [CrossRef] [PubMed]
- Miflin, B.J.; Lea, P.J. 4—Ammonia Assimilation. In Amino Acids and Derivatives; Miflin, B.J., Ed.; Academic Press: Cambridge, MA, USA, 1980; pp. 169–202. [Google Scholar]
- Chen, R.; Greer, A.; Dean, A.M. A highly active decarboxylating dehydrogenase with rationally inverted coenzyme specificity. Proc. Natl. Acad. Sci. USA 1995, 92, 11666–11670. [Google Scholar] [CrossRef] [PubMed]
- Hurley, J.H.; Chen, R.; Dean, A.M. Determinants of Cofactor Specificity in Isocitrate Dehydrogenase: Structure of an Engineered NADP+ → NAD+ Specificity-Reversal Mutant. Biochemistry 1996, 35, 5670–5678. [Google Scholar] [CrossRef] [PubMed]
- Cupp, J.R.; McAlister-Henn, L. Cloning and characterization of the gene encoding the IDH1 subunit of NAD(+)-dependent isocitrate dehydrogenase from Saccharomyces cerevisiae. J. Biol. Chem. 1992, 267, 16417–16423. [Google Scholar] [CrossRef]
- Zhao, W.-N.; McAlister-Henn, L. Affinity Purification and Kinetic Analysis of Mutant Forms of Yeast NAD+-specific Isocitrate Dehydrogenase. J. Biol. Chem. 1997, 272, 21811–21817. [Google Scholar] [CrossRef]
- Lancien, M.; Gadal, P.; Hodges, M. Molecular characterization of higher plant NAD-dependent isocitrate dehydrogenase: Evidence for a heteromeric structure by the complementation of yeast mutants. Plant J. 1998, 16, 325–333. [Google Scholar] [CrossRef]
- Kim, Y.O.; Oh, I.U.; Park, H.S.; Jeng, J.; Song, B.J.; Huh, T.L. Characterization of a cDNA clone for human NAD+-specific isocitrate dehydrogenase α-subunit and structural comparison with its isoenzymes from different species. Biochem. J. 1995, 308, 63–68. [Google Scholar] [CrossRef]
- Nichols, B.J.; Hall, L.; Perry, A.C.F.; Denton, R.M. Molecular cloning and deduced amino acid sequences of the γ-subunits of rat and monkey NAD+-isocitrate dehydrogenases. Biochem. J. 1993, 295, 347–350. [Google Scholar] [CrossRef]
- Ehrlich, R.S.; Hayman, S.; Ramachandran, N.; Colman, R.F. Re-evaluation of molecular weight of pig heart NAD-specific isocitrate dehydrogenase. J. Biol. Chem. 1981, 256, 10560–10564. [Google Scholar] [CrossRef]
- Zeng, Y.; Weiss, C.; Yao, T.T.; Huang, J.; Siconolfi-Baez, L.; Hsu, P.; Rushbrook, J.I. Isocitrate dehydrogenase from bovine heart: Primary structure of subunit 3/4. Biochem. J. 1995, 310, 507–516. [Google Scholar] [CrossRef] [PubMed]
- Dalziel, K. Isocitrate dehydrogenase and related oxidative decarboxylases. FEBS Lett. 1980, 117, K45–K55. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Chen, X. Cloning and activity analysis ofin vitro expression of plant NAD-IDH genes. Chin. Sci. Bull. 2004, 49, 328–336. [Google Scholar] [CrossRef]
- Yasutake, Y.; Watanabe, S.; Yao, M.; Takada, Y.; Fukunaga, N.; Tanaka, I. Crystal Structure of the Monomeric Isocitrate Dehydrogenase in the Presence of NADP+: Insight into the Cofactor Recognition, Catalysis, and Evolution. J. Biol. Chem. 2003, 278, 36897–36904. [Google Scholar] [CrossRef]
- Chandel, N. NADPH—The Forgotten Reducing Equivalent. Cold Spring Harb. Perspect. Biol. 2021, 13, a040550. [Google Scholar] [CrossRef]
- Chen, C.; Cui, H.; Wang, H.; Yang, J.; Luo, L.; Gao, Q.; Guo, T.; Xiao, W.; Huang, Y.; Liu, Y.; et al. Transcriptome Analysis of Hormone Signal Transduction and Glutathione Metabolic Pathway in Rice Seeds at Germination Stage. Chin. J. Rice Sci. 2021, 35, 554–564. [Google Scholar] [CrossRef]
- Marino, D.; González, E.M.; Frendo, P.; Puppo, A.; Arrese-Igor, C. NADPH recycling systems in oxidative stressed pea nodules: A key role for the NADP+-dependent isocitrate dehydrogenase. Planta 2007, 225, 413–421. [Google Scholar] [CrossRef]
- Hanschmann, E.-M.; Godoy, J.R.; Berndt, C.; Hudemann, C.; Lillig, C.H. Thioredoxins, Glutaredoxins, and Peroxiredoxins—Molecular Mechanisms and Health Significance: From Cofactors to Antioxidants to Redox Signaling. Antioxid. Redox Signal. 2013, 19, 1539–1605. [Google Scholar] [CrossRef]
- Xie, X.; Yang, L.; Wang, S.; Zhang, X.; Li, Y. Cloning and Expression Analysis of Sugarcane NADP+-Dependent lsocitrate Dehydrogenase (SoNADP-IDH) Gene. Sci. Agric. Sin. 2015, 48, 185–196. [Google Scholar] [CrossRef]
- Leterrier, M.; Leterrier, M.; del Río, L.A.; Corpas, F.J. Cytosolic NADP-isocitrate dehydrogenase of pea plants: Genomic clone characterization and functional analysis under abiotic stress conditions. Free. Radic. Res. 2007, 41, 191–199. [Google Scholar] [CrossRef] [PubMed]
- Gayatri; Rani, M.; Mahato, A.K.; Sinha, S.K.; Dalal, M.; Singh, N.K.; Mandal, P.K. Homeologue Specific Gene Expression Analysis of Two Vital Carbon Metabolizing Enzymes—Citrate Synthase and NADP-Isocitrate Dehydrogenase—From Wheat (Triticum aestivum L.) Under Nitrogen Stress. Appl. Biochem. Biotechnol. 2019, 188, 569–584. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Song, J.; Weng, Q.; Ma, H.; Wang, L.; Liu, Y. Cloning and Character Analysis of NADP+-Dependent Isocitrate Dehydrogenase Gene in Maize. Plant Physiol. J. 2015, 51, 481–487. [Google Scholar] [CrossRef]
- Liu, Y.; Shi, Y.; Song, Y.; Wang, T.; Li, Y. Characterization of a Stress-induced NADP-isocitrate Dehydrogenase Gene in Maize Confers Salt Tolerance in Arabidopsis. J. Plant Biol. 2010, 53, 107–112. [Google Scholar] [CrossRef]
- Lin, M.; Behal, R.H.; Oliver, D.J. Characterization of a mutation in the IDH-II subunit of the NAD+-dependent isocitrate dehydrogenase from Arabidopsis thaliana. Plant Sci. 2004, 166, 983–988. [Google Scholar] [CrossRef]
- Liu, Y.; Qu, J.; Zhang, L.; Xu, X.; Wei, G.; Zhao, Z.; Ren, M.; Cao, M. Identification and characterization of the TCA cycle genes in maize. BMC Plant Biol. 2019, 19, 592. [Google Scholar] [CrossRef] [PubMed]
- Skopelitis, D.S.; Paranychianakis, N.V.; Paschalidis, K.A.; Pliakonis, E.D.; Delis, I.D.; Yakoumakis, D.I.; Kouvarakis, A.; Papadakis, A.K.; Stephanou, E.G.; Roubelakis-Angelakis, K.A. Abiotic Stress Generates ROS That Signal Expression of Anionic Glutamate Dehydrogenases to Form Glutamate for Proline Synthesis in Tobacco and Grapevine. Plant Cell 2006, 18, 2767–2781. [Google Scholar] [CrossRef]
- Wu, X.; Feng, H.; Wu, D.; Yan, S.; Zhang, P.; Wang, W.; Zhang, J.; Ye, J.; Dai, G.; Fan, Y.; et al. Using high-throughput multiple optical phenotyping to decipher the genetic architecture of maize drought tolerance. Genome Biol. 2021, 22, 185. [Google Scholar] [CrossRef]
- Lemaitre, T.; Hodges, M. Expression Analysis of Arabidopsis thaliana NAD-dependent Isocitrate Dehydrogenase Genes Shows the Presence of a Functional Subunit That Is Mainly Expressed in the Pollen and Absent from Vegetative Organs. Plant Cell Physiol. 2006, 47, 634–643. [Google Scholar] [CrossRef]
- Hu, W.; Ren, Q.; Chen, Y.; Xu, G.; Qian, Y. Genome-wide identification and analysis of WRKY gene family in maize provide insights into regulatory network in response to abiotic stresses. BMC Plant Biol. 2021, 21, 427. [Google Scholar] [CrossRef]
- Holub, E.B. The arms race is ancient history in Arabidopsis, the wildflower. Nat. Rev. Genet. 2001, 2, 516–527. [Google Scholar] [CrossRef] [PubMed]
- dos Santos, T.B.; Ribas, A.F.; de Souza, S.G.H.; Budzinski, I.G.F.; Domingues, D.S. Physiological Responses to Drought, Salinity, and Heat Stress in Plants: A Review. Stresses 2022, 2, 113–135. [Google Scholar] [CrossRef]
- Hoopes, G.M.; Hamilton, J.P.; Wood, J.C.; Esteban, E.; Pasha, A.; Vaillancourt, B.; Provart, N.J.; Buell, C.R. An updated gene atlas for maize reveals organ-specific and stress-induced genes. Plant J. 2019, 97, 1154–1167. [Google Scholar] [CrossRef] [PubMed]
- Thatcher, S.R.; Danilevskaya, O.N.; Meng, X.; Beatty, M.; Zastrow-Hayes, G.; Harris, C.; Van Allen, B.; Habben, J.; Li, B. Genome-Wide Analysis of Alternative Splicing during Development and Drought Stress in Maize. Plant Physiol. 2015, 170, 586–599. [Google Scholar] [CrossRef]
- Devaiah, K.M.; Bali, G.; Athmaram, T.N.; Basha, M.S. Identification of two new genes from drought tolerant peanut up-regulated in response to drought. Plant Growth Regul. 2007, 52, 249–258. [Google Scholar] [CrossRef]
- Foyer, C.H.; Noctor, G.; Hodges, M. Respiration and nitrogen assimilation: Targeting mitochondria-associated metabolism as a means to enhance nitrogen use efficiency. J. Exp. Bot. 2011, 62, 1467–1482. [Google Scholar] [CrossRef]
- Lynch, M.; Conery, J.S. The Evolutionary Fate and Consequences of Duplicate Genes. Science 2000, 290, 1151–1155. [Google Scholar] [CrossRef]
- Yang, W.; Dong, X.; Yuan, Z.; Zhang, Y.; Li, X.; Wang, Y. Genome-Wide Identification and Expression Analysis of the Ammonium Transporter Family Genes in Soybean. Int. J. Mol. Sci. 2023, 24, 3991. [Google Scholar] [CrossRef]
- Lu, B.; Yuan, Y.; Zhang, C.; Ou, J.; Zhou, W.; Lin, Q. Modulation of key enzymes involved in ammonium assimilation and carbon metabolism by low temperature in rice (Oryza sativa L.) roots. Plant Sci. 2005, 169, 295–302. [Google Scholar] [CrossRef]
- Gamrasni, D.; Erov, M.; Saar, L.; Raz, A.; Glikman, M.; Sonawane, P.D.; Aharoni, A.; Goldway, M. The isocitrate dehydrogenase 1 gene is associated with the climacteric response in tomato fruit ripening. Postharvest Biol. Technol. 2020, 166, 111219. [Google Scholar] [CrossRef]
- Gao, H.; Cui, J.; Liu, S.; Wang, S.; Lian, Y.; Bai, Y.; Zhu, T.; Wu, H.; Wang, Y.; Yang, S.; et al. Natural variations of ZmSRO1d modulate the trade-off between drought resistance and yield by affecting ZmRBOHC-mediated stomatal ROS production in maize. Mol. Plant 2022, 15, 1558–1574. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Wang, X.; Wang, H.; Xin, H.; Yang, X.; Yan, J.; Li, J.; Tran, L.-S.P.; Shinozaki, K.; Yamaguchi-Shinozaki, K.; et al. Genome-Wide Analysis of ZmDREB Genes and Their Association with Natural Variation in Drought Tolerance at Seedling Stage of Zea mays L. PLoS Genet. 2013, 9, e1003790. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]

| Gene Name | Gene ID | Position | Protein Length (aa) | pI | MW | GRAVY | Subcellular Location |
|---|---|---|---|---|---|---|---|
| ZmIDH1.1 | Zm00001d028735 | Chr1:44718501–44719536 | 273 | 6.72 | 29,983.39 | −0.158 | Mitochondrion |
| ZmIDH1.2 | Zm00001d040438 | Chr3:43449762–43455057 | 365 | 6.52 | 39,724.58 | −0.091 | Mitochondrion |
| ZmIDH1.3 | Zm00001d050965 | Chr4:134060297–134069849 | 211 | 9.21 | 22,949.69 | −0.068 | Mitochondrion |
| ZmIDH1.4 | Zm00001d017091 | Chr5:185084105–185089320 | 375 | 8.91 | 40,361.7 | −0.049 | Mitochondrion |
| ZmIDH1.5 | Zm00001d008244 | Chr8:2452376–2459944 | 365 | 6.33 | 39,907.84 | −0.109 | Mitochondrion |
| ZmIDH1.6 | Zm00001d025690 | Chr10:126319920–126323617 | 377 | 6.72 | 40,565.61 | −0.046 | Mitochondrion |
| ZmIDH2.1 | Zm00001d003083 | Chr2:31519654–31526365 | 503 | 8.24 | 56,084.57 | −0.179 | Cytoplasmic |
| ZmIDH2.2 | Zm00001d044021 | Chr3:216967820–216972096 | 412 | 6.57 | 46,097.73 | −0.231 | Cytoplasmic |
| ZmIDH2.3 | Zm00001d039079 | Chr6:169685197–169688921 | 415 | 6.24 | 46,223.72 | −0.238 | Cytoplasmic |
| ZmIDH2.4 | Zm00001d011487 | Chr8:152072966–152077705 | 412 | 6.11 | 46,196.77 | −0.245 | Cytoplasmic |
| ZmIDH2.5 | Zm00001d046262 | Chr9:77157097–77159734 | 298 | 8.19 | 33,719.93 | −0.116 | Cytoplasmic; Chloroplast |
| Gene ID | Gene Name | Polymorphic Number | MLM (p ≤ 0.05) | MLM (p ≤ 0.01) | MLM (p ≤ 0.001) | Max R2 (%) |
|---|---|---|---|---|---|---|
| Zm00001d028735 | ZmIDH1.1 | 14 | 3 | 1 | 0 | 1.956 |
| Zm00001d040438 | ZmIDH1.2 | 30 | 1 | 0 | 0 | 1.611 |
| Zm00001d050965 | ZmIDH1.3 | 23 | 0 | 0 | 0 | 1.029 |
| Zm00001d017091 | ZmIDH1.4 | 32 | 4 | 4 | 2 | 6.706 |
| Zm00001d008244 | ZmIDH1.5 | 27 | 4 | 1 | 0 | 2.094 |
| Zm00001d025690 | ZmIDH1.6 | 55 | 3 | 1 | 0 | 2.329 |
| Zm00001d003083 | ZmIDH2.1 | 20 | 0 | 0 | 0 | 1.024 |
| Zm00001d044021 | ZmIDH2.2 | 42 | 1 | 1 | 0 | 2.372 |
| Zm00001d039079 | ZmIDH2.3 | 66 | 10 | 1 | 0 | 2.032 |
| Zm00001d011487 | ZmIDH2.4 | 28 | 1 | 0 | 0 | 1.833 |
| Zm00001d046262 | ZmIDH2.5 | 52 | 1 | 0 | 0 | 1.213 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wei, N.; Zhang, Z.; Yang, H.; Hu, D.; Wu, Y.; Xue, J.; Guo, D.; Xu, S. Characterization of the Isocitrate Dehydrogenase Gene Family and Their Response to Drought Stress in Maize. Plants 2023, 12, 3466. https://doi.org/10.3390/plants12193466
Wei N, Zhang Z, Yang H, Hu D, Wu Y, Xue J, Guo D, Xu S. Characterization of the Isocitrate Dehydrogenase Gene Family and Their Response to Drought Stress in Maize. Plants. 2023; 12(19):3466. https://doi.org/10.3390/plants12193466
Chicago/Turabian StyleWei, Ningning, Ziran Zhang, Haoxiang Yang, Die Hu, Ying Wu, Jiquan Xue, Dongwei Guo, and Shutu Xu. 2023. "Characterization of the Isocitrate Dehydrogenase Gene Family and Their Response to Drought Stress in Maize" Plants 12, no. 19: 3466. https://doi.org/10.3390/plants12193466
APA StyleWei, N., Zhang, Z., Yang, H., Hu, D., Wu, Y., Xue, J., Guo, D., & Xu, S. (2023). Characterization of the Isocitrate Dehydrogenase Gene Family and Their Response to Drought Stress in Maize. Plants, 12(19), 3466. https://doi.org/10.3390/plants12193466





