Genetic Diversity Analysis of Soybean Collection Using Simple Sequence Repeat Markers
Abstract
1. Introduction
2. Results
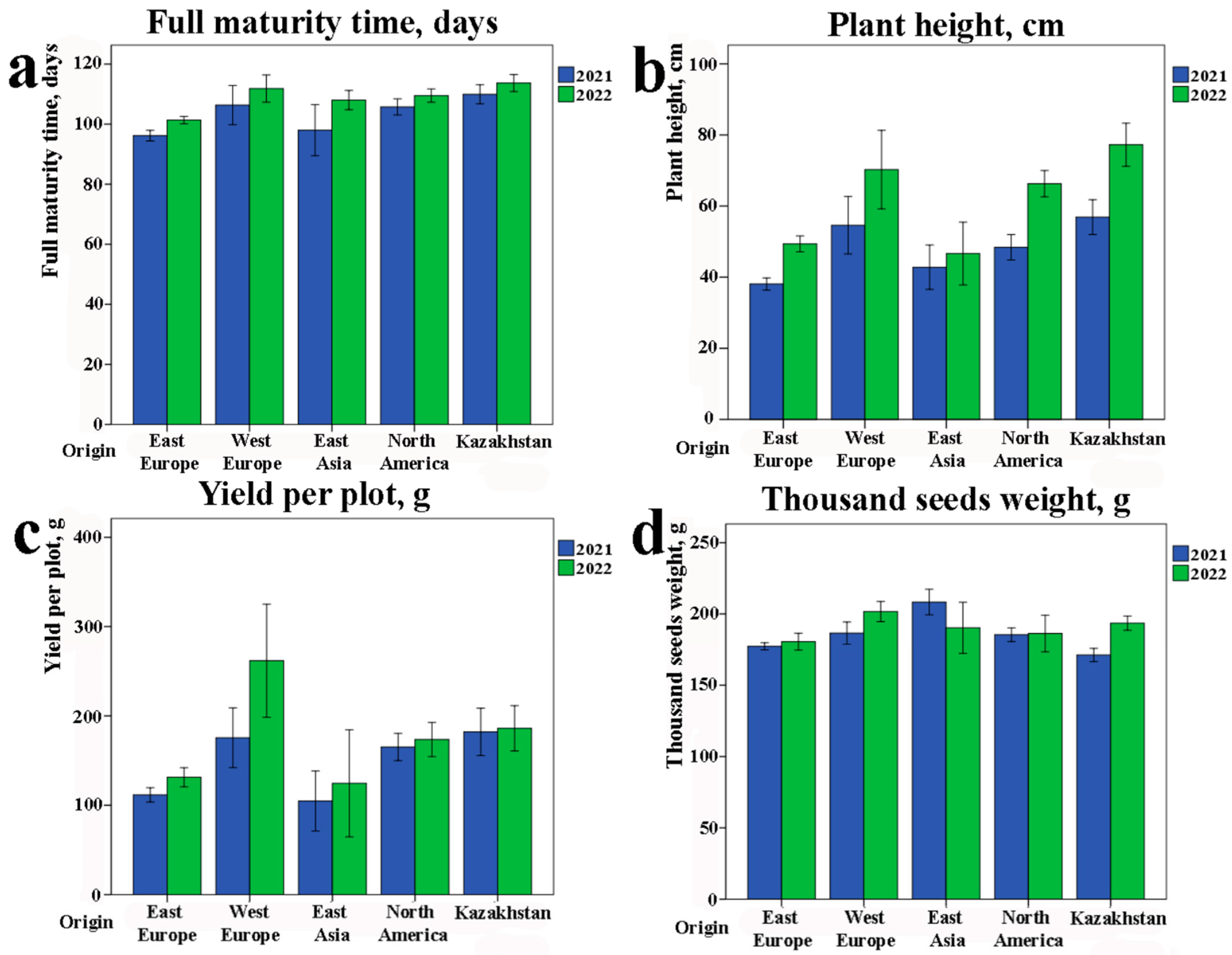
2.1. Field Assessment of the Studied Collection by Using Main Agronomic Traits
2.2. Assessment of the Soybean Collection by Using SSR Markers
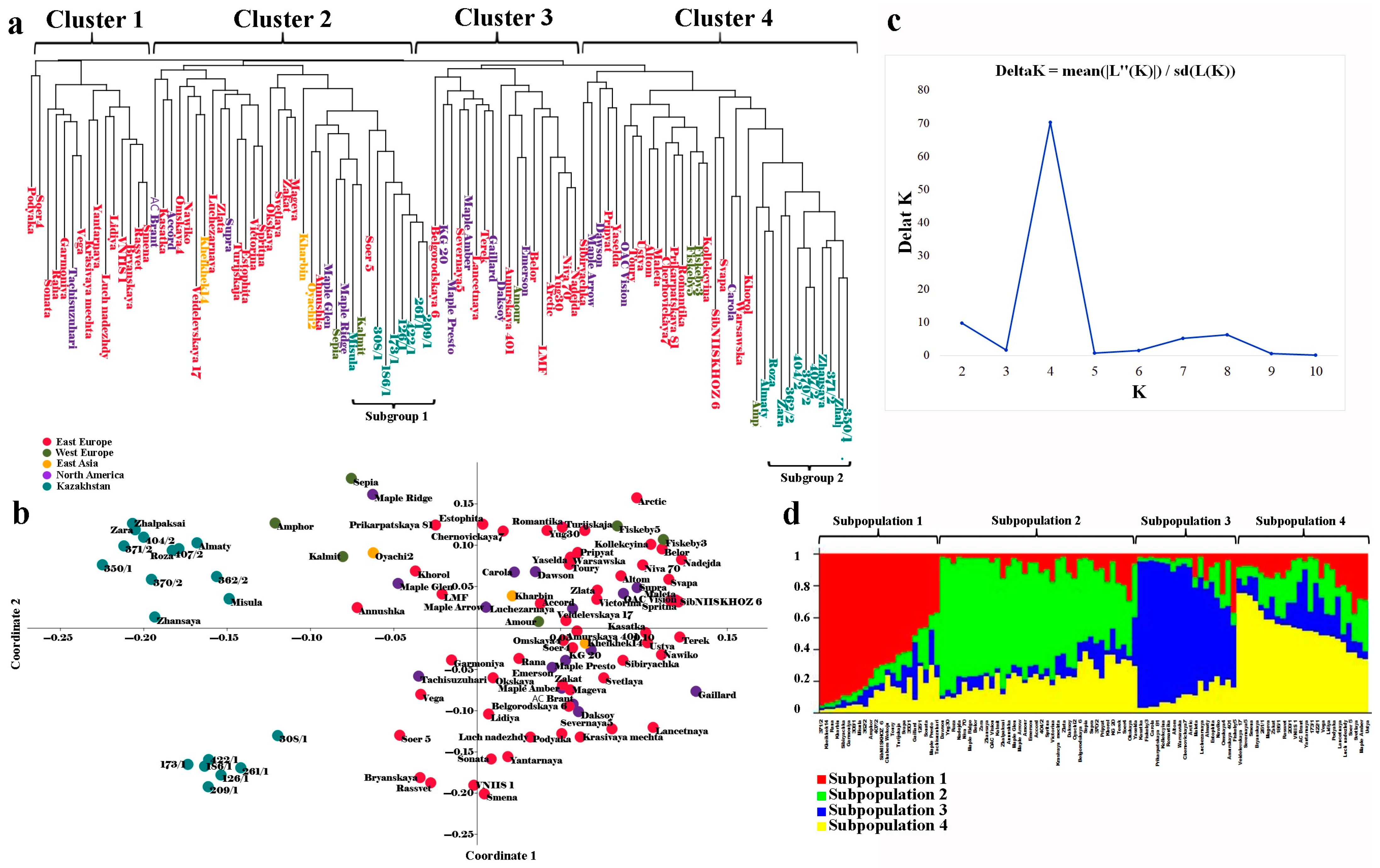
2.3. Clusterization Analysis of the Studied Collection
2.4. The Association of SSR Markers with Main Agronomic Traits
3. Discussion
3.1. Phenotypic Variation in the Studied Soybean Collection
3.2. Analysis of Population Structure and Polymorphism Level in the Studied Soybean Collection
3.3. Association of SSRs with Main Agronomic Traits
4. Materials and Methods
4.1. Studied Collection and Field Experiments
4.2. DNA Extraction and Genotyping by Using SSR Markers
4.3. Statistical and Population Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- FAOSTAS. Available online: https://www.fao.org/faostat/en/#data/QCL (accessed on 17 July 2023).
- USDA. Available online: https://ipad.fas.usda.gov/cropexplorer/cropview/commodityView.aspx?cropid=2222000 (accessed on 17 July 2023).
- Makulbekova, A.; Iskakov, A.; Kulkarni, K.P.; Song, J.T.; Lee, J.D. Current status of future prospects of soybean production in Kazakhstan. Plant Breed. Biotechnol. 2017, 5, 55–66. [Google Scholar] [CrossRef]
- Didorenko, S.V.; Zakiyeva, A.A.; Sidorik, I.V.; Abugalieva, A.I.; Kudaibergenov, M.S.; Iskakov, A.R. Diversification of crop production by means of spreading soybeans to the northern regions of the republic of Kazakhstan. Biosci. Biotechnol. Res. Asia 2016, 13, 23–30. [Google Scholar] [CrossRef][Green Version]
- Li, M.; Liu, Y.; Wang, C.; Yang, X.; Li, D.; Zhang, X.; Xu, C.; Zhang, Y.; Li, W.; Zhao, L. Identification of Traits Contributing to High and Stable Yields in Different Soybean Varieties Across Three Chinese Latitudes. Front. Plant Sci. 2020, 10, 1642. [Google Scholar] [CrossRef] [PubMed]
- Jean, M.; Cober, E.; O’Donoughue, L.; Rajcan, I.; Belzile, F. Improvement of key agronomical traits in soybean through genomic prediction of superior crosses. Crop Sci. 2021, 61, 3908–3918. [Google Scholar] [CrossRef]
- Zatybekov, A.; Abugalieva, S.; Didorenko, S.; Gerasimova, Y.; Sidorik, I.; Anuarbek, S.; Turuspekov, Y. GWAS of agronomic traits in soybean collection included in breeding pool in Kazakhstan. BMC Plant Biol. 2017, 17, 179. [Google Scholar] [CrossRef]
- Moose, S.P.; Mumm, R.H. Molecular plant breeding as the foundation for 21st century crop improvement. Plant Physiol. 2008, 147, 969–977. [Google Scholar] [CrossRef]
- Ahmar, S.; Gill, R.A.; Jung, K.-H.; Faheem, A.; Qasim, M.U.; Mubeen, M.; Zhou, W. Conventional and Molecular Techniques from Simple Breeding to Speed Breeding in Crop Plants: Recent Advances and Future Outlook. Int. J. Mol. Sci. 2020, 21, 2590. [Google Scholar] [CrossRef]
- Abugalieva, S.; Didorenko, S.; Anuarbek, S.; Volkova, L.; Gerasimova, Y.; Sidorik, I.; Turuspekov, Y. Assessment of soybean flowering and seed maturation time in different latitude regions of Kazakhstan. PLoS ONE 2016, 11, e0166894. [Google Scholar] [CrossRef]
- Xue, H.; Tian, X.; Zhang, K.; Li, W.; Qi, Z.; Fang, Y.; Li, X.; Wang, Y.; Song, J.; Li, W.X.; et al. Mapping developmental QTL for plant height in soybean [Glycine max (L.) Merr.] using a four-way recombinant inbred line population. PLoS ONE 2019, 14, e0224897. [Google Scholar] [CrossRef]
- Yang, Q.; Lin, G.; Lv, H.; Wang, C.; Yang, Y.; Liao, H. Environmental and genetic regulation of plant height in soybean. BMC Plant Biol. 2021, 21, 63. [Google Scholar] [CrossRef]
- Wang, J.; Hu, B.; Jing, Y.; Hu, X.; Guo, Y.; Chen, J.; Liu, Y.; Hao, J.; Li, W.X.; Ning, H. Detecting QTL and Candidate Genes for Plant Height in Soybean via Linkage Analysis and GWAS. Front. Plant Sci. 2022, 12, 803820. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Nan, H.; Kong, L.; Yue, L.; Yang, H.; Zhao, Q.; Fang, C.; Li, H.; Cheng, Q.; Lu, S.; et al. Soybean AP1 homologs control flowering time and plant height. J. Integr. Plant Biol. 2020, 62, 1868–1879. [Google Scholar] [CrossRef] [PubMed]
- Cregan, P.B.; Jarwik, T.; Bush, A.L.; Shoemaker, R.C.; Lark, K.G.; Kahler, A.L.; Van Toai, T.T.; Lohnes, D.G.; Chung, J.; Specht, J.E. An integrated genetic linkage map of soybean genome. Crop Sci. 1999, 39, 1464–1490. [Google Scholar] [CrossRef]
- Bisen, A.; Khare, D.; Nair, P.; Tripathi, N. SSR analysis of 38 genotypes of soybean (Glycine max (L.) Merr.) genetic diversity in India. Physiol. Mol. Biol. Plants 2014, 21, 109–115. [Google Scholar] [CrossRef]
- Koutu, G.K.; Shrivastava, A.; Singh, Y.; Tiwari, S. Molecular Characterization and Genetic Diversity Assessment of Soybean Varieties using SSR Markers. Int. J. Curr. Microbiol. App. Sci. 2019, 8, 173–182. [Google Scholar] [CrossRef]
- Singh, N.; Choudhury, D.R.; Singh, A.K.; Kumar, S.; Srinivasan, K.; Tyagi, R.K.; Singh, N.K.; Singh, R. Comparison of SSR and SNP markers in estimation of genetic diversity and population structure of Indian rice varieties. PLoS ONE 2013, 8, e84136. [Google Scholar] [CrossRef]
- Priyanatha, C.; Torkamaneh, D.; Rajcan, I. Genome-Wide Association Study of Soybean Germplasm Derived from Canadian × Chinese Crosses to Mine for Novel Alleles to Improve Seed Yield and Seed Quality Traits. Front Plant Sci. 2022, 13, 866300. [Google Scholar] [CrossRef]
- Zatybekov, A.K.; Turuspekov, Y.T.; Doszhanova, B.N.; Abugalieva, S.I. A study of the genetic diversity in the world soybean collection using microsatellite markers associated with fungal disease resistance. Proc. Appl. Bot. Genet. Breed. 2020, 181, 81–90. [Google Scholar] [CrossRef]
- Tanya, P.; Srinives, P.; Toojinda, T.; Vanavichit, A.; Ha, B.-K.; Bae, J.-S.; Moon, J.-K.; Lee, S.-H. Evaluation of genetic diversity among soybean genotypes using SSR and SNP. Korean J. Crop Sci. 2001, 46, 334–340. [Google Scholar]
- Ben-Ari, G.; Lavi, U. Marker-assisted selection in plant breeding. In Plant Biotechnology and Agriculture; Elsevier: Amsterdam, The Netherlands, 2012; pp. 163–184. [Google Scholar] [CrossRef]
- Mishra, A.; Singh, P.K.; Bhandawat, A.; Sharma, V.; Sharma, V.; Singh, P.; Roy, J.; Sharma, H. Analysis of SSR and SNP Markers. In Bioinformatics; Singh, D.B., Patha, R.K., Eds.; Academic Press: Cambridge, MA, USA, 2022; pp. 131–144. [Google Scholar] [CrossRef]
- Malik, M.; Khan, M.N. Analysis of genetic variations in soybean using simple sequence repeat (SSR) markers. Sarhad J. Agric. 2021, 37, 331–339. [Google Scholar] [CrossRef]
- Wang, L.X.; Guan, R.X.; Li, Y.H.; Lin, F.Y.; Luan, W.J.; Li, W.; Ma, Y.S.; Liu, Z.X.; Chang, R.Z.; Qiu, L.J. Genetic diversity of chinese spring soybean germplasm revealed by SSR markers. Plant Breed. 2008, 127, 56–61. [Google Scholar] [CrossRef]
- Yao, X.; Xu, J.; Liu, Z.; Pachner, M.; Molin, E.M.; Rittler, L.; Hahn, V.; Leiser, W.; Gu, Y.; Lu, Y.; et al. Genetic diversity in early maturity Chinese and European elite soybeans: A comparative analysis. Euphytica 2023, 219, 17. [Google Scholar] [CrossRef]
- Sun, D.; Li, W.; Zhang, Z.; Chen, Q.; Ning, H.; Qiu, L.; Sun, G. Quantitative trait loci analysis for the developmental behavior of Soybean (Glycine max L. Merr.). Theor. Appl. Genet. 2006, 112, 665–673. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.-S.; Diers, B.W.; Hyten, D.L.; Rouf Mian, M.A.; Shannon, J.G.; Nelson, R.L. Identification of positive yield QTL alleles from exotic soybean germplasm in two backcross populations. Theor. Appl. Genet. 2012, 125, 1353–1369. [Google Scholar] [CrossRef] [PubMed]
- Kabelka, E.A.; Diers, B.W.; Fehr, W.R.; LeRoy, A.R.; Baianu, I.C.; You, T.; Neece, D.J.; Nelson, R.L. Putative Alleles for Increased Yield from Soybean Plant Introductions. Crop Sci. 2004, 44, 784. [Google Scholar] [CrossRef]
- Gai, J.; Wang, Y.; Wu, X.; Shouyi, C. A comparative study on segregation analysis and QTL mapping of quantitative traits in plants—With a case in soybean. Front. Agric. China 2007, 1, 1–7. [Google Scholar] [CrossRef]
- Lee, S.; Jun, T.H.; Michel, A.P.; Rouf Mian, M.A. SNP markers linked to QTL conditioning plant height, lodging, and maturity in soybean. Euphytica 2015, 203, 521–532. [Google Scholar] [CrossRef]
- Alcivar, A.; Jacobson, J.; Rainho, J.; Meksem, K.; Lightfoot, D.A.; Kassem, M.A. Genetic Analysis of Soybean Plant Height, Hypocotyl and Internode Lengths. J. Agric. Food Environ. Sci. 2007, 1, 1–20. [Google Scholar]
- Li, W.; Zheng, D.H.; Van, K.; Lee, S.H. QTL Mapping for Major Agronomic Traits across Two Years in Soybean (Glycine max L. Merr.). J. Crop Sci. Biotechnol. 2008, 11, 171–190. [Google Scholar]
- Reinprecht, Y.; Poysa, V.W.; Yu, K.; Rajcan, I.; Ablett, G.R.; Pauls, K.P. Seed and agronomic QTL in low linolenic acid, lipoxygenase-free soybean (Glycine max (L.) Merrill) germplasm. Genome 2006, 49, 1510–1527. [Google Scholar] [CrossRef]
- Orf, J.H.; Chase, K.; Jarvik, T.; Mansur, L.M.; Cregan, P.B.; Adler, F.R.; Lark, K.G. Genetics of Soybean Agronomic Traits. Crop Sci. 1999, 39, 1642–1651. [Google Scholar] [CrossRef]
- Specht, J.E.; Chase, K.; Macrander, M.; Graef, G.L.; Chung, J.; Markwell, J.P.; Germann, M.; Orf, J.H.; Lark, K.G. Soybean Response to Water. Crop Sci. 2001, 41, 493. [Google Scholar] [CrossRef]
- Wang, D.; Procopiuk, A.M.; Diers, B.W.; Graef, G.L. Identification of putative QTL that underlie yield in interspecific soybean backcross populations. TAG Theor. Appl. Genet. 2004, 108, 458–467. [Google Scholar] [CrossRef] [PubMed]
- Yao, D.; Liu, Z.Z.; Zhang, J.; Liu, S.Y.; Qu, J.; Guan, S.Y.; Pan, L.D.; Wang, D.; Liu, J.W.; Wang, P.W. Analysis of quantitative trait loci for main plant traits in soybean. Genet. Mol. Res. 2015, 14, 6101–6109. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.-K.; Wang, Y.-J.; Luo, G.-Z.; Zhang, J.-S.; He, C.-Y.; Wu, X.-L.; Gai, J.-Y.; Chen, S.-Y. QTL mapping of ten agronomic traits on the soybean (Glycine max L. Merr.) genetic map and their association with EST markers. TAG Theor. Appl. Genet. 2004, 108, 1131–1139. [Google Scholar] [CrossRef]
- Liu, W.; Kim, M.Y.; Van, K.; Lee, Y.-H.; Li, H.; Liu, X.; Lee, S.-H. QTL identification of yield-related traits and their association with flowering and maturity in soybean. J. Crop Sci. Biotechnol. 2011, 14, 65–70. [Google Scholar] [CrossRef]
- Chen, Q.; Zhang, Z.; Liu, C.; Xin, D.; Qiu, H.; Shan, D.; Shan, C.; Hu, G. QTL Analysis of Major Agronomic Traits in Soybean. Agric. Sci. China 2007, 6, 399–405. [Google Scholar] [CrossRef]
- Li, D.; Sun, M.; Han, Y.; Teng, W.; Li, W. Identification of QTL underlying soluble pigment content in soybean stems related to resistance to soybean white mold (Sclerotinia sclerotiorum). Euphytica 2010, 172, 49–57. [Google Scholar] [CrossRef]
- Guzman, P.S.; Diers, B.W.; Neece, D.J.; St. Martin, S.K.; LeRoy, A.R.; Grau, C.R.; Hughes, T.J.; Nelson, R.L. QTL Associated with Yield in Three Backcross-Derived Populations of Soybean. Crop Sci. 2007, 47, 111. [Google Scholar] [CrossRef]
- Yuan, J.; Njiti, V.N.; Meksem, K.; Iqbal, M.J.; Triwitayakorn, K.; Kassem, M.A.; Davis, G.T.; Schmidt, M.E.; Lightfoot, D.A. Quantitative trait loci in Two Soybean Recombinant Inbred Line Populations Segregating for Yield and Disease Resistance. Crop Sci. 2002, 42, 271–277. [Google Scholar] [CrossRef]
- Kasu-Bandi, B.T.; Kidinda, L.K.; Kasendue, G.N.; Longanza, L.B.; Emery, K.L.; Lubobo, A.K. Correlations between Growth and Yield Parameters of Soybean (Glycine max (L.) Merr.) under the Influence of Bradyrhizobium japonicum in Kipushi (The Democratic Republic of Congo). Am. J. Agric. Biol. Sci. 2019, 14, 86–94. [Google Scholar] [CrossRef]
- Diers, B.W.; Specht, J.; Rainey, K.M.; Cregan, P.; Song, Q.; Ramasubramanian, V.; Graef, G.; Nelson, R.; Schapaugh, W.; Wang, D.; et al. Genetic Architecture of Soybean Yield and Agronomic Traits. G3 Bethesda 2018, 8, 3367–3375. [Google Scholar] [CrossRef]
- Sulistyo, A.; Purwantoro; Sari, K.P. Correlation, path analysis and heritability estimation for agronomic traits contribute to yield on soybean. IOP Conf. Ser. Earth Environ. Sci. 2018, 102, 012034. [Google Scholar] [CrossRef]
- Ramanatha Rao, V.; Hodgkin, T. Genetic diversity and conservation and utilization of plant genetic resources. Plant Cell Tissue Organ Cult. 2002, 68, 1–19. [Google Scholar] [CrossRef]
- Shaibu, A.S.; Ibrahim, H.; Miko, Z.L.; Mohammed, I.B.; Mohammed, S.G.; Yusuf, H.L.; Kamara, A.Y.; Omoigui, L.O.; Karikari, B. Assessment of the Genetic Structure and Diversity of Soybean (Glycine max L.) Germplasm Using Diversity Array Technology and Single Nucleotide Polymorphism Markers. Plants 2021, 11, 68. [Google Scholar] [CrossRef] [PubMed]
- Samyuktha, S.M.; Kannan Bapu, J.R.; Geethanjali, S. Molecular Genetic Diversity and Population Structure Analysis in Chickpea (Cicer arietinum L.) Germplasm using SSR Markers. Int. J. Curr. Microbiol. Appl. Sci. 2018, 7, 639–651. [Google Scholar] [CrossRef]
- Tribhuvan, K.U.; Amitha Mithra, S.V.; Sharma, P.; Das, A.; Kumar, K.; Tyagi, A.; Solanke, A.U.; Sandhya; Sharma, R.; Jadhav, P.V.; et al. Identification of genomic SSRs in cluster bean (Cyamopsis tetragonoloba) and demonstration of their utility in genetic diversity analysis. Ind. Crops Prod. 2019, 133, 221–231. [Google Scholar] [CrossRef]
- Hwang, T.-Y.; Gwak, B.S.; Sung, J.; Kim, H.-S. Genetic Diversity Patterns and Discrimination of 172 Korean Soybean (Glycine max (L.) Merrill) Varieties Based on SSR Analysis. Agriculture 2020, 10, 77. [Google Scholar] [CrossRef]
- Ullah, A.; Akram, Z.; Malik, S.I.; Khan, K.S.U. Assessment of phenotypic and molecular diversity in soybean [Glycine max (L.) Merr.] germplasm using morpho-biochemical attributes and SSR markers. Genet. Resour. Crop Evol. 2021, 68, 2827–2847. [Google Scholar] [CrossRef]
- Botstein, D.; White, R.L.; Skolnick, M.; Davis, R.W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 1980, 32, 314–331. [Google Scholar]
- Adeboye, K.A.; Oyedeji, O.E.; Alqudah, A.M.; Börner, A.; Oduwaye, O.; Adebambo, O.; Daniel, I.O. Genetic structure and diversity of upland rice germplasm using diversity array technology (DArT)-based single nucleotide polymorphism (SNP) markers. Plant Genet. Resour. 2020, 18, 343–350. [Google Scholar] [CrossRef]
- Tantasawat, P.; Trongchuen, J.; Prajongjai, T.; Jenweerawat, S.; Chaowiset, W. SSR analysis of soybean (Glycine max (L.) Merr.) genetic relationship and variety identification in Thailand. Aust. J. Crop Sci. 2011, 5, 283–290. [Google Scholar]
- Swarnalatha, V.; Swar, B.R.; Reddy, M.R.; Vanisree, S. SSR Based Genetic Diversity in Magic Lines of Soybean (Glycine max (L.) Merrill). Int. J. Plant Soil Sci. 2021, 33, 8–16. [Google Scholar] [CrossRef]
- Rani, R.; Raza, G.; Tung, M.H.; Rizwan, M.; Ashfaq, H.; Shimelis, H.; Razzaq, M.K.; Arif, M. Genetic diversity and population structure analysis in cultivated soybean (Glycine max [L.] Merr.) using SSR and EST-SSR markers. PLoS ONE 2023, 18, e0286099. [Google Scholar] [CrossRef]
- Choi, Y.-M.; Lee, M.-C.; Ro, N.; Lee, S.; Gwag, J.-G.; Yoon, M.-S. Morphological Characteristics and SSR profilings of Soybean Landraces by collecting sites of Korea. Korean J. Breed. Sci. 2014, 46, 353–363. [Google Scholar] [CrossRef]
- Turuspekov, Y.; Nakamura, K.; Yoshikawa, R.; Tuberosa, R. Genetic Diversity of Japanese Barley Cultivars Based on SSR Analysis. Breed. Sci. 2001, 51, 215–218. [Google Scholar] [CrossRef]
- Hipparagi, Y.; Singh, R.; Choudhury, D.R.; Gupta, V. Genetic diversity and population structure analysis of Kala bhat (Glycine max (L.) Merrill) genotypes using SSR markers. Hereditas 2017, 154, 9. [Google Scholar] [CrossRef]
- Kachare, S.; Tiwari, S.; Tripathi, N.; Thakur, V.V. Assessment of genetic diversity of soybean (Glycine max) genotypes using qualitative traits and microsatellite markers. Agric. Res. 2020, 9, 23–34. [Google Scholar] [CrossRef]
- Gupta, S.K.; Manjaya, J.G. Genetic diversity and population structure of Indian soybean [Glycine max (L.) Merr.] revealed by simple sequence repeat markers. J. Crop Sci. Biotechnol. 2017, 20, 221–231. [Google Scholar] [CrossRef]
- Dong, D.; Fu, X.; Yuan, F.; Chen, P.; Zhu, S.; Li, B.; Yang, Q.; Yu, X.; Zhu, D. Genetic diversity and population structure of vegetable soybean (Glycine max (L.) Merr.) in China as revealed by SSR markers. Genet. Resour. Crop Evol. 2014, 61, 173–183. [Google Scholar] [CrossRef]
- Tasma, I.M.; Yani, N.M.; Purwaningdyah, R.; Satyawan, D.; Nugroho, K.; Lestari, P.; Trijatmiko, K.R.; Mastur, M. Genetic diversity analysis and F2 population development for breeding of long juvenile trait in soybean. J. AgroBiogen 2018, 14, 11–22. [Google Scholar] [CrossRef]
- Zigene, Z.D.; Asfaw, B.T.; Bitima, T.D. Analysis of genetic diversity in rosemary (Salvia rosemarinus Schleid.) using SSR molecular marker for its management and sustainable use in Ethiopian genebank. Genet. Resour. Crop Evol. 2020, 68, 279–293. [Google Scholar] [CrossRef]
- Minerdi, D.; Savoi, S.; Sabbatini, P. Role of Cytochrome P450 Enzyme in Plant Microorganisms’ Communication: A Focus on Grapevine. Int. J. Mol. Sci. 2023, 24, 4695. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, G.; Zheng, K.; Zhu, X.; Ma, J.; Wang, D.; Tang, K.; Feng, X.; Leng, J.; Yu, H.; et al. The Soybean Laccase Gene Family: Evolution and Possible Roles in Plant Defense and Stem Strength Selection. Genes 2019, 10, 701. [Google Scholar] [CrossRef]
- Han, Y.; Li, D.; Zhu, D.; Li, H.; Li, X.; Teng, W.; Li, W. QTL analysis of soybean seed weight across multi-genetic backgrounds and environments. Theor. Appl. Genet. 2012, 125, 671–683. [Google Scholar] [CrossRef]
- Sun, Y.; Pan, J.; Shi, X.; Du, X.; Wu, Q.; Qi, Z.; Jiang, H.; Xin, D.; Liu, C.; Hu, G.; et al. Multi-environment mapping and meta-analysis of 100-seed weight in soybean. Mol. Biol. Rep. 2012, 39, 9435–9443. [Google Scholar] [CrossRef]
- Du, W.; Wang, M.; Fu, S.; Yu, D. Mapping QTLs for seed yield and drought susceptibility index in soybean (Glycine max L.) across different environments. J. Genet. Genom. 2009, 36, 721–731. [Google Scholar] [CrossRef]
- Korsakov, N.I.; Makasheva, R.H.; Adamova, O.P. Methodology for Studying the Collection of Legumes (Metodika Izucheniya Kollektsii Zernobobovykh Kultur); VIR: Saint Petersburg, Rusia, 1968. (In Russian) [Google Scholar]
- Dellaporta, S.L.; Wood, J.; Hicks, J.B. A plant DNA minipreparation: Version II. Plant Mol. Biol. Report. 1983, 1, 19–21. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. Past: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontol. Electron. 2001, 4, 4–9. [Google Scholar]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; von Holdt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef] [PubMed]

| PH Groups | n | PH Range, cm | Average PH, cm | Average YpP, g |
|---|---|---|---|---|
| A | 58 | 20–49.9 | 39.1 ± 7.2 | 110.0 ± 6.4 |
| B | 31 | 50–79.9 | 60.5 ± 6.3 | 179.3 ± 11.0 |
| C | 11 | 80–111.1 | 91.5 ± 7.7 | 248.2 ± 19.4 |
| Factors | Traits | SS | d.f. | MS | F | P |
|---|---|---|---|---|---|---|
| Year | VER2 | 272.997 | 1 | 272.997 | 28.666 | 0.000 |
| R2R8 | 72.147 | 1 | 72.147 | 0.623 | 0.431 | |
| VER8 | 625.829 | 1 | 625.829 | 4.577 | 0.034 | |
| PH | 3760.536 | 1 | 3760.536 | 12.360 | 0.001 | |
| HLP | 27.930 | 1 | 27.930 | 4.857 | 0.029 | |
| NLN | 23.365 | 1 | 23.365 | 47.827 | 0.000 | |
| NFN | 2171.070 | 1 | 2171.070 | 35.081 | 0.000 | |
| NSP | 19,794.491 | 1 | 19,794.491 | 48.245 | 0.000 | |
| YpP | 15,047.475 | 1 | 15,047.475 | 2.137 | 0.145 | |
| TSW | 437.906 | 1 | 437.906 | 0.434 | 0.511 | |
| Origin | VER2 | 43.218 | 4 | 10.805 | 1.135 | 0.342 |
| R2R8 | 5435.516 | 4 | 1358.879 | 11.727 | 0.000 | |
| VER8 | 6063.577 | 4 | 1515.894 | 11.086 | 0.000 | |
| PH | 18,794.750 | 4 | 4698.687 | 15.444 | 0.000 | |
| HLP | 25.838 | 4 | 6.460 | 1.123 | 0.347 | |
| NLN | 2.907 | 4 | 0.727 | 1.488 | 0.207 | |
| NFN | 1218.281 | 4 | 304.570 | 4.921 | 0.001 | |
| NSP | 11,136.326 | 4 | 2784.081 | 6.786 | 0.000 | |
| YpP | 209,326.569 | 4 | 52,331.642 | 7.432 | 0.000 | |
| TSW | 5080.853 | 4 | 1270.213 | 1.258 | 0.288 | |
| Year xOrigin | VER2 | 37.244 | 4 | 9.311 | 0.978 | 0.421 |
| R2R8 | 9.090 | 4 | 2.272 | 0.020 | 0.999 | |
| VER8 | 65.186 | 4 | 16.297 | 0.119 | 0.976 | |
| PH | 875.544 | 4 | 218.886 | 0.719 | 0.580 | |
| HLP | 18.167 | 4 | 4.542 | 0.790 | 0.533 | |
| NLN | 0.246 | 4 | 0.062 | 0.126 | 0.973 | |
| NFN | 135.675 | 4 | 33.919 | 0.548 | 0.701 | |
| NSP | 932.594 | 4 | 233.149 | 0.568 | 0.686 | |
| YpP | 16,622.417 | 4 | 4155.604 | 0.590 | 0.670 | |
| TSW | 4055.344 | 4 | 1013.836 | 1.004 | 0.407 |
| SSR Loci | Chr | na | ne | Ho | uHe | uh | PIC |
|---|---|---|---|---|---|---|---|
| Satt428 | 2 | 4 | 2.45 | 0.128 | 0.531 | 0.550 | 0.539 |
| Satt600 | 2 | 5 | 1.57 | 0.061 | 0.346 | 0.357 | 0.340 |
| Satt387 | 3 | 2 | 1.50 | 0.000 | 0.318 | 0.333 | 0.322 |
| Satt307 | 6 | 6 | 2.49 | 0.065 | 0.538 | 0.582 | 0.731 |
| Satt371 | 6 | 6 | 3.13 | 0.134 | 0.663 | 0.684 | 0.703 |
| Satt460 | 6 | 3 | 1.43 | 0.011 | 0.281 | 0.295 | 0.290 |
| Satt489 | 6 | 5 | 3.19 | 0.000 | 0.600 | 0.626 | 0.684 |
| Satt557 | 6 | 4 | 2.25 | 0.000 | 0.497 | 0.526 | 0.563 |
| Satt150 | 7 | 3 | 1.66 | 0.000 | 0.414 | 0.453 | 0.319 |
| Satt308 | 7 | 5 | 2.35 | 0.064 | 0.590 | 0.634 | 0.660 |
| Satt567 | 7 | 5 | 2.85 | 0.000 | 0.683 | 0.747 | 0.661 |
| Satt153 | 10 | 5 | 2.02 | 0.046 | 0.448 | 0.465 | 0.455 |
| Satt243 | 10 | 5 | 2.79 | 0.061 | 0.650 | 0.699 | 0.675 |
| Satt197 | 11 | 4 | 2.08 | 0.032 | 0.527 | 0.572 | 0.493 |
| Satt509 | 11 | 3 | 1.70 | 0.075 | 0.378 | 0.391 | 0.423 |
| Satt335 | 13 | 3 | 1.75 | 0.025 | 0.437 | 0.479 | 0.357 |
| Satt263 | 15 | 4 | 2.32 | 0.046 | 0.585 | 0.630 | 0.598 |
| Satt244 | 16 | 7 | 2.77 | 0.048 | 0.642 | 0.689 | 0.732 |
| Satt547 | 16 | 4 | 2.47 | 0.063 | 0.532 | 0.551 | 0.642 |
| Sat_308 | 18 | 3 | 1.19 | 0.007 | 0.148 | 0.152 | 0.140 |
| Satt288 | 18 | 8 | 2.35 | 0.198 | 0.557 | 0.567 | 0.624 |
| Satt309 | 18 | 3 | 1.57 | 0.033 | 0.336 | 0.347 | 0.338 |
| Satt324 | 18 | 5 | 2.51 | 0.101 | 0.591 | 0.619 | 0.667 |
| Satt440 | 20 | 4 | 2.73 | 0.000 | 0.560 | 0.584 | 0.644 |
| Sct189 | 20 | 3 | 2.19 | 0.060 | 0.565 | 0.608 | 0.499 |
| Mean value | 4.36 | 2.21 | 0.050 | 0.497 | 0.526 | 0.524 | |
| Standard error | 0.33 | 0.33 | 0.007 | 0.019 | 0.021 | 0.028 | |
| Region | ne | Ho | He | uHe | Fst | uh |
|---|---|---|---|---|---|---|
| Eastern Europe | 2.60 | 0.078 | 0.567 | 0.573 | 0.854 | 0.580 |
| Western Europe | 2.23 | 0.048 | 0.504 | 0.551 | 0.894 | 0.625 |
| East Asia | 1.44 | 0.040 | 0.229 | 0.275 | 0.764 | 0.240 |
| North America | 2.51 | 0.045 | 0.546 | 0.564 | 0.929 | 0.583 |
| Kazakhstan | 2.29 | 0.040 | 0.507 | 0.521 | 0.921 | 0.605 |
| Mean value | 2.21 | 0.050 | 0.471 | 0.497 | 0.884 | 0.527 |
| Standard error | 0.08 | 0.007 | 0.020 | 0.020 | 0.019 | 0.024 |
| Subgroups | N | Year | VER2 | R2R8 | VER8 | PH | HLP | NLB | NFN | NSP | TSW | YpP |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Subgroup 1 | 8 | 2021 | 31.75 | 68.75 | 100.50 | 44.45 | 7.75 | 0.38 | 11.63 | 24.99 | 170.38 | 105.41 |
| 2022 | 34.63 | 69.75 | 104.38 | 58.69 | 7.54 | 1.80 | 23.02 | 60.00 | 194.63 | 130.46 | ||
| Mean | 33.19 | 69.25 | 102.44 | 51.57 | 7.64 | 1.09 | 17.32 | 42.49 | 182.50 | 117.94 | ||
| SE | 1.72 | 0.22 | 1.93 | 6.33 | 0.24 | 0.76 | 5.57 | 15.80 | 11.57 | 1.18 | ||
| Subgroup 2 | 11 | 2021 | 34.55 | 82.27 | 116.82 | 65.94 | 7.15 | 0.74 | 15.08 | 42.29 | 171.82 | 237.95 |
| 2022 | 36.36 | 84.09 | 120.45 | 90.75 | 11.27 | 1.80 | 27.68 | 78.69 | 192.73 | 226.55 | ||
| Mean | 35.45 | 83.18 | 118.64 | 78.34 | 9.21 | 1.27 | 21.38 | 60.49 | 182.27 | 232.25 | ||
| SE | 0.80 | 1.04 | 1.84 | 12.43 | 1.96 | 0.52 | 6.33 | 19.14 | 10.92 | 3.82 | ||
| p-value for 2 subgroups | 0.158 | 0.002 | 0.001 | 0.007 | 0.152 | 0.568 | 0.187 | 0.063 | 0.977 | 0.001 | ||
| SSR Markers | Chr | VER2 | R2R8 | VER8 | PH | HLP | NLB | NFN | NSP | TSW | YpP |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Satt387 | 2 | ns | 0.045 2 | ns | 0.018 1 | ns | 0.018 1 0.004 2 0.008 3 | 0.001 1 | 0.002 1 0.007 3 | ns | 0.013 2 0.002 3 |
| Satt489 | 3 | ns | ns | ns | 0.007 2 0.034 3 | ns | ns | ns | ns | ns | ns |
| Satt557 | 4 | ns | ns | ns | ns | ns | ns | ns | ns | 0.031 1 | ns |
| Satt150 | 4 | ns | 0.044 2 | 0.031 2 | ns | ns | ns | ns | ns | ns | ns |
| Satt567 | 6 | 0.040 1 | ns | ns | ns | ns | ns | ns | ns | ns | ns |
| Satt324 | 6 | Ns | 0.038 1 0.016 2 0.021 3 | 0.022 1 0.017 2 0.016 3 | 0.020 1 0.027 2 0.016 3 | ns | 0.036 2 | ns | 0.030 1 0.028 2 0.016 3 | 0.036 2 0.023 3 | 0.048 1 0.043 2 0.014 3 |
| Satt440 | 6 | 0.000 1 0.000 3 | 0.002 1 0.002 2 0.001 3 | 0.000 1 0.000 2 0.000 3 | 0.002 1 0.006 3 | ns | 0.000 1 0.004 3 | 0.000 1 0.036 3 | 0.000 1 0.019 3 | ns | 0.001 1 0.042 2 0.002 3 |
| Satt600 | 6 | ns | ns | ns | ns | 0.030 2 | ns | ns | ns | ns | ns |
| Satt460 | 7 | ns | 0.038 1 0.042 3 | ns | 0.039 1 0.0413 | ns | ns | ns | ns | ns | 0.024 2 0.046 3 |
| Satt244 | 10 | ns | 0.006 1 0.006 2 0.004 3 | 0.019 1 0.019 2 0.016 3 | ns | ns | 0.026 1 | ns | ns | ns | 0.013 1 |
| Satt288 | 11 | ns | 0.031 1 0.004 2 0.010 3 | 0.003 1 0.011 2 0.016 3 | 0.002 1 0.000 2 0.000 3 | ns | 0.067 1 | 0.005 1 0.043 2 0.014 3 | 0.001 1 0.021 2 0.004 3 | ns | 0.000 2 0.000 3 |
| Satt308 | 17 | ns | ns | ns | ns | ns | ns | ns | ns | ns | 0.029 2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zatybekov, A.; Yermagambetova, M.; Genievskaya, Y.; Didorenko, S.; Abugalieva, S. Genetic Diversity Analysis of Soybean Collection Using Simple Sequence Repeat Markers. Plants 2023, 12, 3445. https://doi.org/10.3390/plants12193445
Zatybekov A, Yermagambetova M, Genievskaya Y, Didorenko S, Abugalieva S. Genetic Diversity Analysis of Soybean Collection Using Simple Sequence Repeat Markers. Plants. 2023; 12(19):3445. https://doi.org/10.3390/plants12193445
Chicago/Turabian StyleZatybekov, Alibek, Moldir Yermagambetova, Yuliya Genievskaya, Svetlana Didorenko, and Saule Abugalieva. 2023. "Genetic Diversity Analysis of Soybean Collection Using Simple Sequence Repeat Markers" Plants 12, no. 19: 3445. https://doi.org/10.3390/plants12193445
APA StyleZatybekov, A., Yermagambetova, M., Genievskaya, Y., Didorenko, S., & Abugalieva, S. (2023). Genetic Diversity Analysis of Soybean Collection Using Simple Sequence Repeat Markers. Plants, 12(19), 3445. https://doi.org/10.3390/plants12193445





