NADP-Dependent Malic Enzyme Genes in Sweet Pepper Fruits: Involvement in Ripening and Modulation by Nitric Oxide (NO)
Abstract
1. Introduction
2. Results
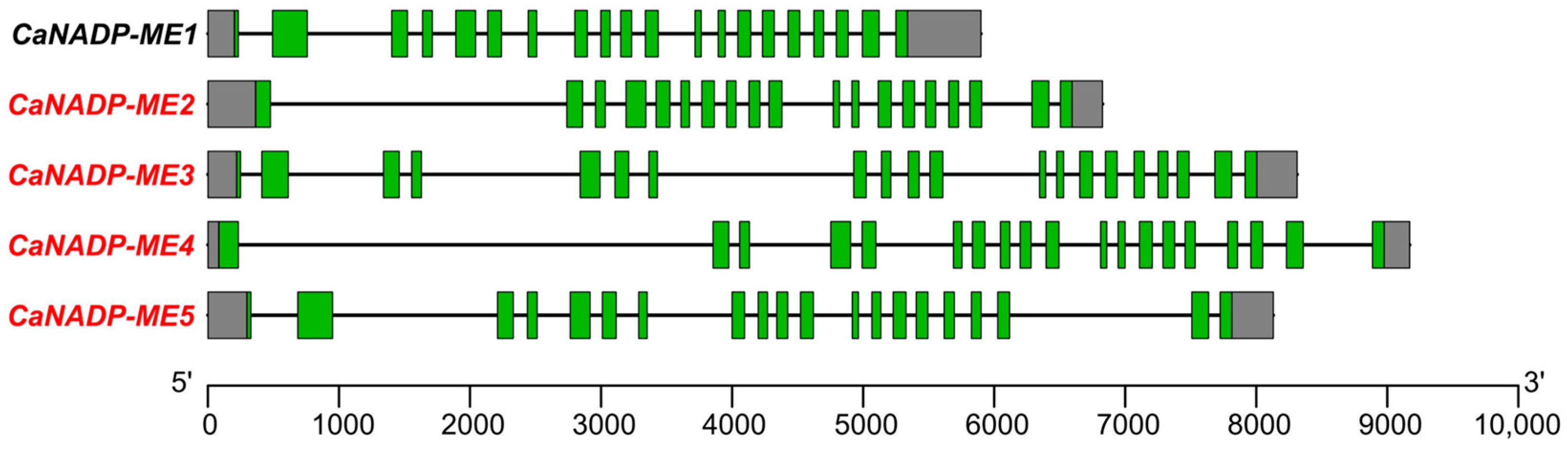
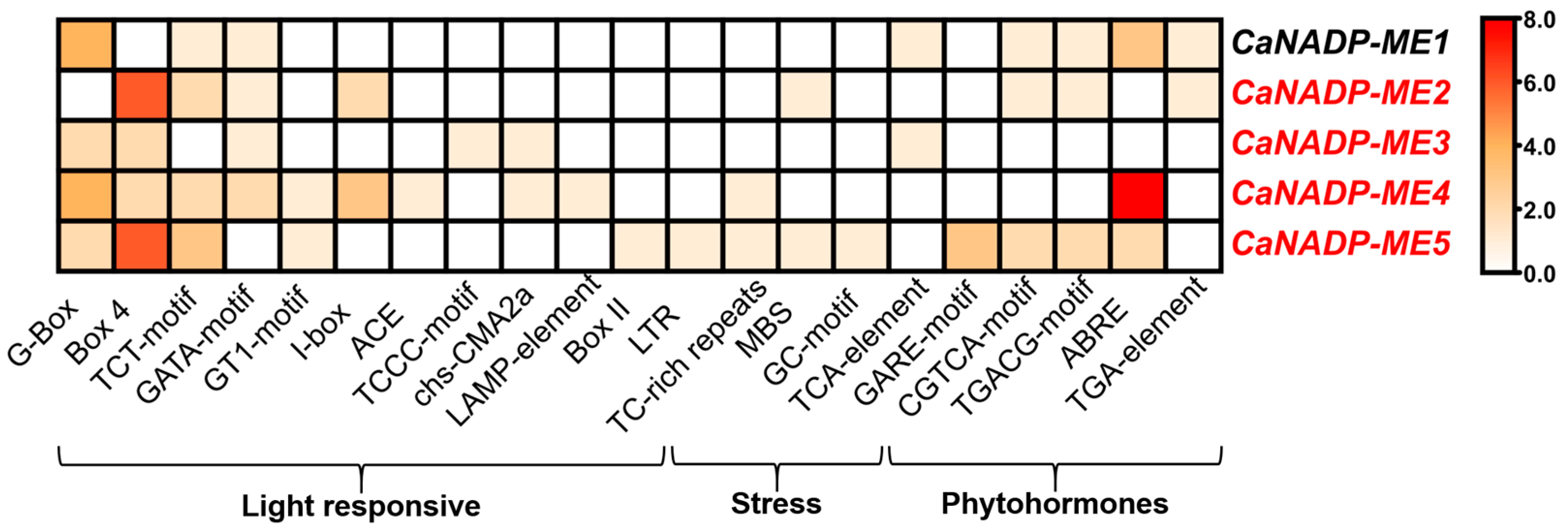
2.1. Identification of the NADP-ME Genes in Pepper (Capsicum annuum L.): Sequence and Cis-Regulatory Elements
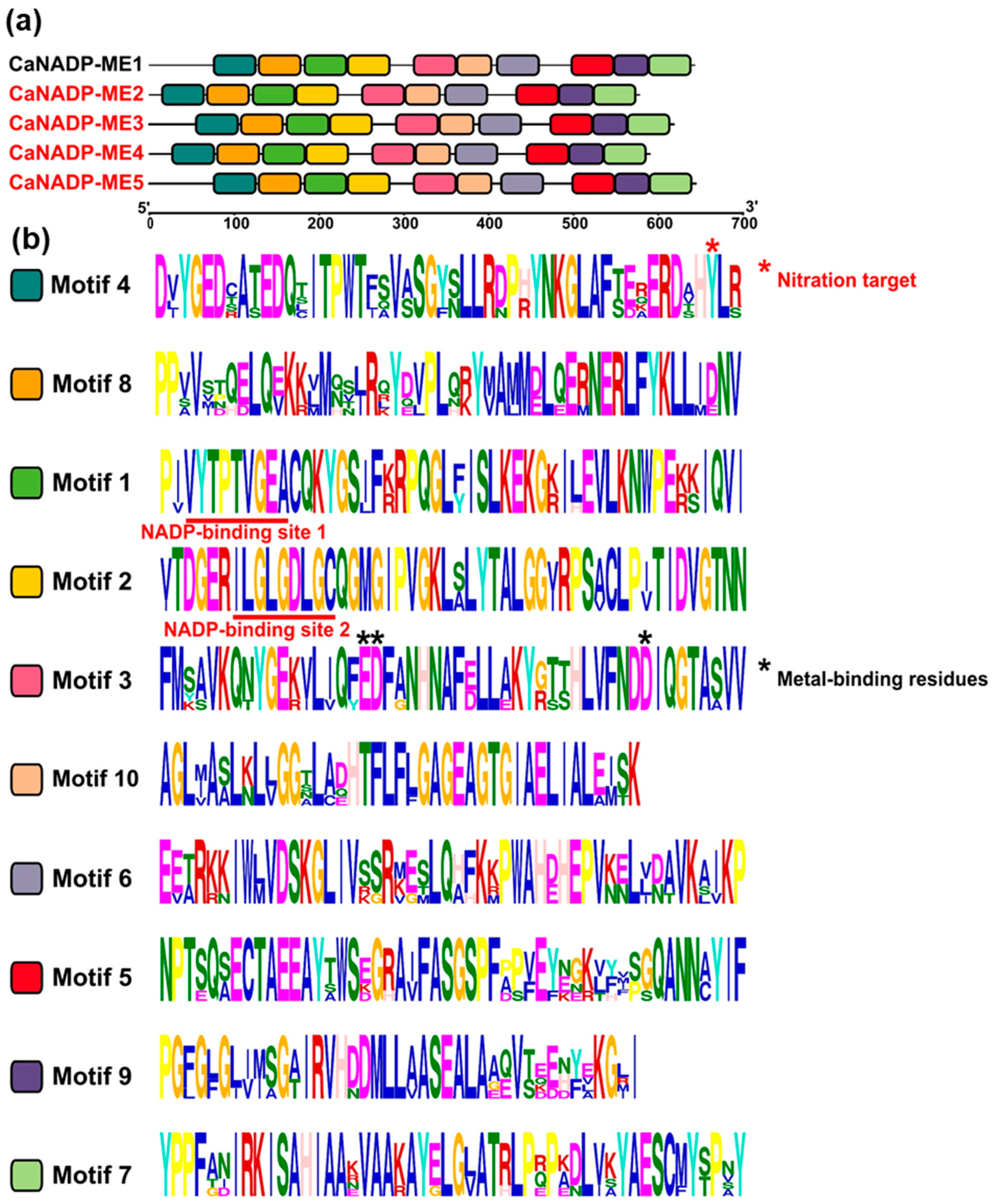
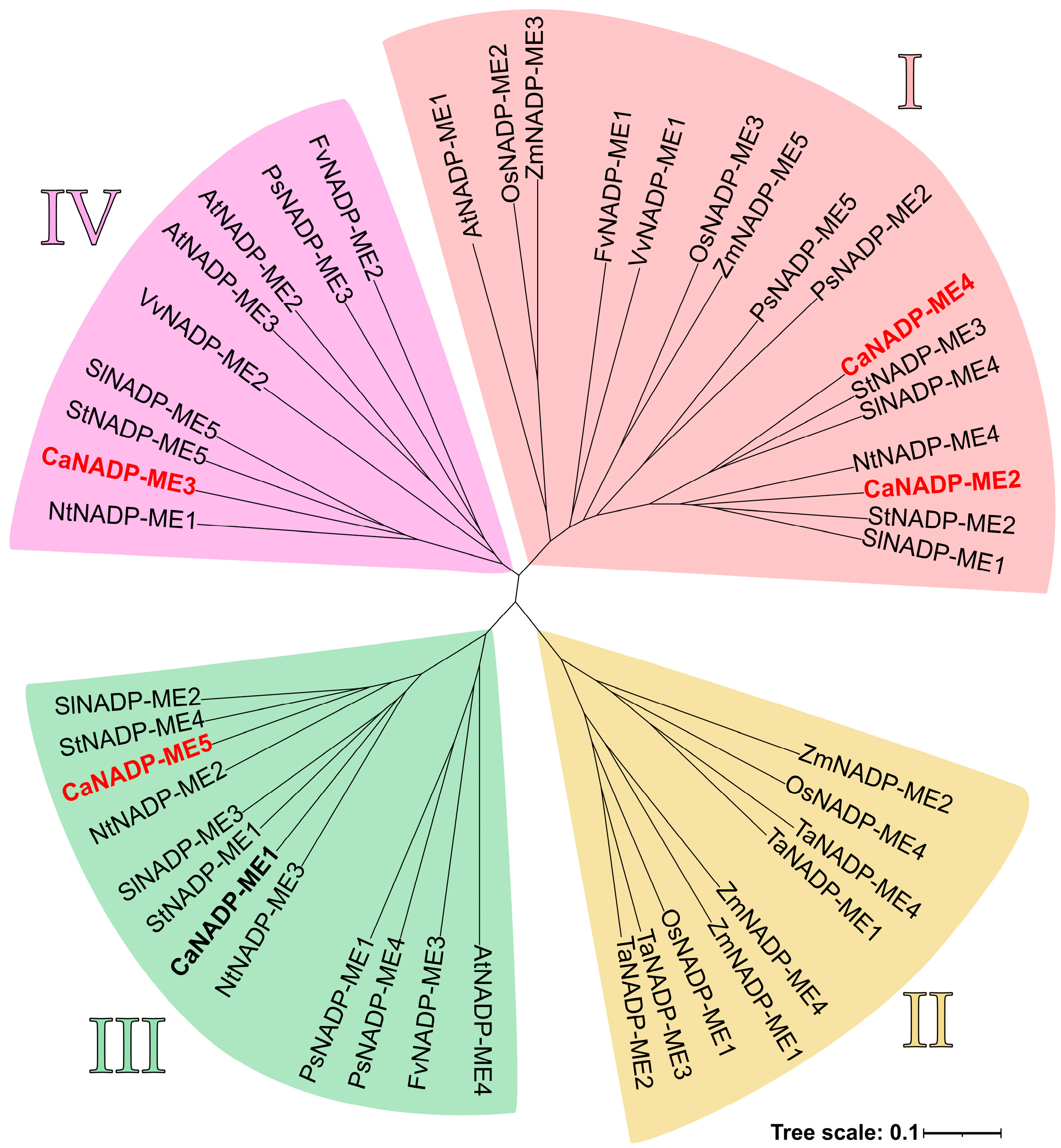
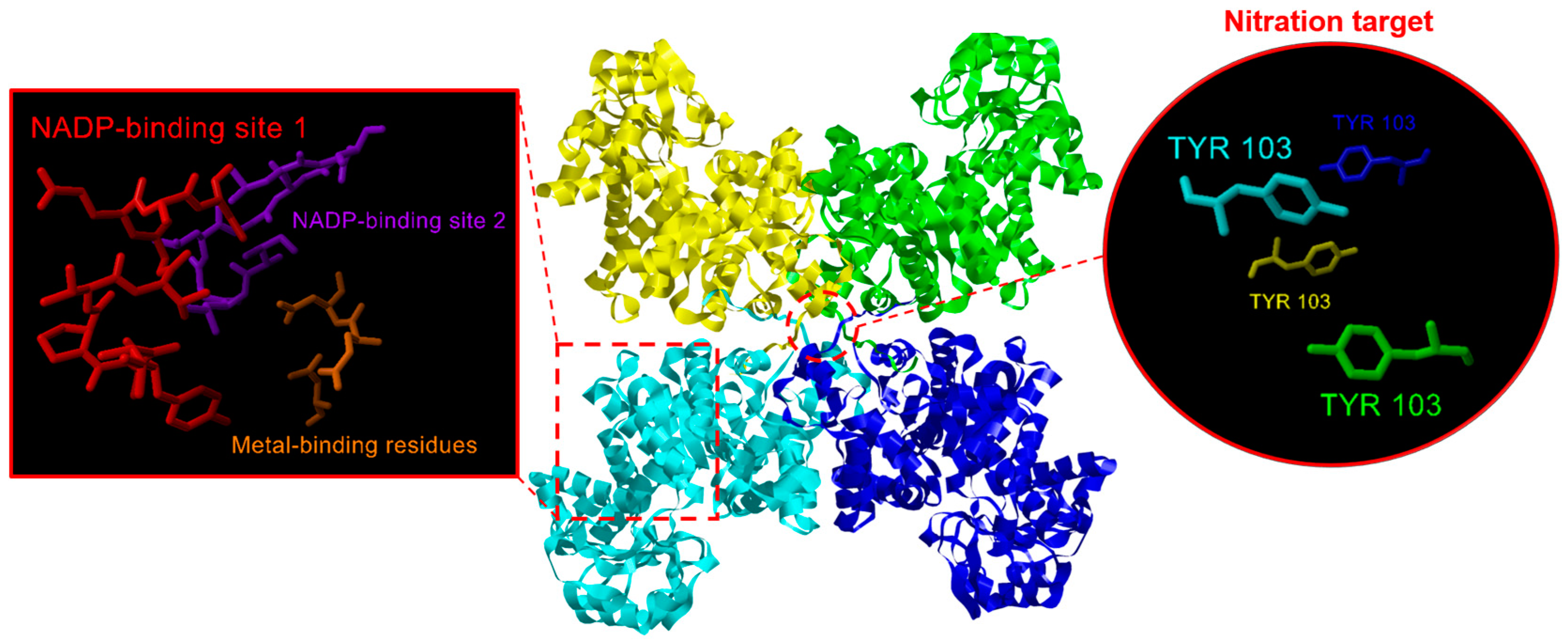
2.2. NADP-ME Proteins from Pepper: Sequence, Phylogenetic Analysis and Modeling
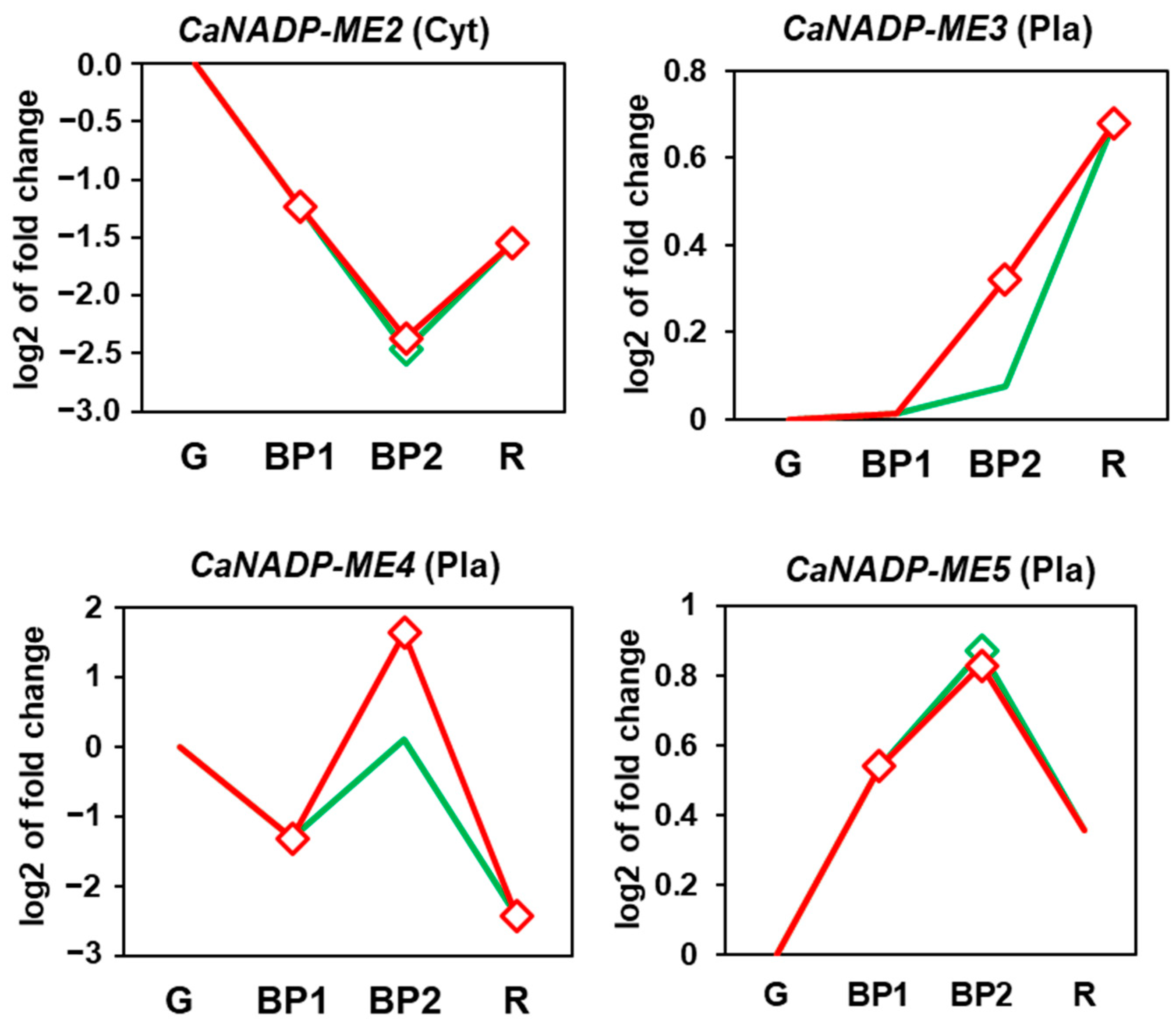
2.3. Fruit CaNADP-ME Genes: Expression during Ripening and Exogenous NO Treatment
2.4. Identification of the CaNADP-ME Isozymes in Pepper Fruits
3. Discussion
3.1. The Pepper Genome Contains Five CaNADP-ME Genes but Only Four Genes (CaNADP-ME2 to ME5) Are Expressed in Fruits
3.2. During Fruit Ripening, the Expression of CaNADP-ME2 and ME4 Is Downregulated, Whereas That of CaNADP-ME3 Is Upregulated, and Exogenous NO Gas Exerts a Negative Effect on CaNADP-ME3 and ME4
4. Materials and Methods
4.1. Identification of the NADP-ME Genes in Pepper (Capsicum annuum L.), Chromosomal Location and Synteny Analysis
4.2. Phylogenetic and Conserved Motif Analyses of NADP-ME Protein Sequences
4.3. Introns/Exons and Cis-Regulatory Elements Analysis of the CaNADP-ME Genes
4.4. Plant Material and Exogenous Nitric Oxide (NO) Gas Treatment
4.5. Library Preparation and RNA Sequencing
4.6. Fruit Extracts, Protein Assay, Protein Enrichment Using Ammonium–Sulfate [(NH4)2SO4] Fractionation and In-Gel Isozyme Profile of NADP-ME Activity
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Corpas, F.J.; Barroso, J.B. NADPH-Generating Dehydrogenases: Their Role in the Mechanism of Protection against Nitro-Oxidative Stress Induced by Adverse Environmental Conditions. Front. Environ. Sci. 2014, 2, 55. [Google Scholar] [CrossRef]
- Habashi, R.; Hacham, Y.; Dhakarey, R.; Matityahu, I.; Holland, D.; Tian, L.; Amir, R. Elucidating the Role of Shikimate Dehydrogenase in Controlling the Production of Anthocyanins and Hydrolysable Tannins in the Outer Peels of Pomegranate. BMC Plant Biol. 2019, 19, 476. [Google Scholar] [CrossRef] [PubMed]
- Meeks, K.R.; Tanner, J.J. Expression and Kinetic Characterization of PYCR3. Arch. Biochem. Biophys. 2023, 733, 109468. [Google Scholar] [CrossRef] [PubMed]
- Sagi, M.; Fluhr, R. Production of Reactive Oxygen Species by Plant NADPH Oxidases. Plant Physiol. 2006, 141, 336–340. [Google Scholar] [CrossRef] [PubMed]
- Kaya, H.; Takeda, S.; Kobayashi, M.J.; Kimura, S.; Iizuka, A.; Imai, A.; Hishinuma, H.; Kawarazaki, T.; Mori, K.; Yamamoto, Y.; et al. Comparative Analysis of the Reactive Oxygen Species-Producing Enzymatic Activity of Arabidopsis NADPH Oxidases. Plant J. 2019, 98, 291–300. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.; Li, P.; Hong, X.; Xu, C.; Wang, R.; Liang, Y. The Receptor-like Cytosolic Kinase RIPK Activates NADP-Malic Enzyme 2 to Generate NADPH for Fueling ROS Production. Mol. Plant 2022, 15, 887–903. [Google Scholar] [CrossRef]
- Corpas, F.J.; González-Gordo, S.; Palma, J.M. NO Source in Higher Plants: Present and Future of an Unresolved Question. Trends Plant Sci. 2022, 27, 116–119. [Google Scholar] [CrossRef] [PubMed]
- Edwards, G.E.; Andreo, C.S. NADP-Malic Enzyme from Plants. Phytochemistry 1992, 31, 1845–1857. [Google Scholar] [CrossRef]
- Drincovich, M.F.; Casati, P.; Andreo, C.S. NADP-Malic Enzyme from Plants: A Ubiquitous Enzyme Involved in Different Metabolic Pathways. FEBS Lett. 2001, 490, 1–6. [Google Scholar] [CrossRef]
- Yazdanpanah, F.; Maurino, V.G.; Mettler-Altmann, T.; Buijs, G.; Bailly, M.N.; Karimi Jashni, M.; Willems, L.; Sergeeva, L.I.; Rajjou, L.C.; Hilhorst, H.W.M.; et al. NADP-MALIC ENZYME 1 Affects Germination after Seed Storage in Arabidopsis thaliana. Plant Cell Physiol. 2019, 60, 318–328. [Google Scholar] [CrossRef]
- Airaki, M.; Leterrier, M.; Valderrama, R.; Chaki, M.; Begara-Morales, J.C.; Barroso, J.B.; del Río, L.A.; Palma, J.M.; Corpas, F.J. Spatial and Temporal Regulation of the Metabolism of Reactive Oxygen and Nitrogen Species during the Early Development of Pepper (Capsicum annuum) Seedlings. Ann. Bot. 2015, 116, 679–693. [Google Scholar] [CrossRef] [PubMed]
- Goodenough, P.W.; Prosser, I.M.; Young, K. NADP-Linked Malic Enzyme and Malate Metabolism in Ageing Tomato Fruit. Phytochemistry 1985, 24, 1157–1162. [Google Scholar] [CrossRef]
- Franke, K.E.; Adams, D.O. Cloning of a Full-Length CDNA for Malic Enzyme (EC 1.1.1.40) from Grape Berries. Plant Physiol. 1995, 107, 1009–1010. [Google Scholar] [CrossRef] [PubMed]
- Famiani, F.; Walker, R.P.; Técsi, L.; Chen, Z.H.; Proietti, P.; Leegood, R.C. An Immunohistochemical Study of the Compartmentation of Metabolism during the Development of Grape (Vitis vinifera L.) Berries. J. Exp. Bot. 2000, 51, 675–683. [Google Scholar] [PubMed]
- Famiani, F.; Casulli, V.; Baldicchi, A.; Battistelli, A.; Moscatello, S.; Walker, R.P. Development and Metabolism of the Fruit and Seed of the Japanese Plum Ozark Premier (Rosaceae). J. Plant Physiol. 2012, 169, 551–560. [Google Scholar] [CrossRef] [PubMed]
- Osorio, S.; Vallarino, J.G.; Szecowka, M.; Ufaz, S.; Tzin, V.; Angelovici, R.; Galili, G.; Fernie, A.R. Alteration of the Interconversion of Pyruvate and Malate in the Plastid or Cytosol of Ripening Tomato Fruit Invokes Diverse Consequences on Sugar but Similar Effects on Cellular Organic Acid, Metabolism, and Transitory Starch Accumulation. Plant Physiol. 2013, 161, 628–643. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Wang, B.; Ding, H.; Zhang, J.; Li, S. Review: The Role of NADP-Malic Enzyme in Plants under Stress. Plant Sci. 2019, 281, 206–212. [Google Scholar] [CrossRef]
- Valderrama, R.; Corpas, F.J.; Carreras, A.; Gómez-Rodríguez, M.V.; Chaki, M.; Pedrajas, J.R.; Fernández-Ocaña, A.; Del Río, L.A.; Barroso, J.B. The Dehydrogenase-Mediated Recycling of NADPH Is a Key Antioxidant System against Salt-Induced Oxidative Stress in Olive Plants. Plant Cell Environ. 2006, 29, 1449–1459. [Google Scholar] [CrossRef]
- Manai, J.; Gouia, H.; Corpas, F.J. Redox and Nitric Oxide Homeostasis Are Affected in Tomato (Solanum lycopersicum) Roots under Salinity-Induced Oxidative Stress. J. Plant Physiol. 2014, 171, 1028–1035. [Google Scholar] [CrossRef]
- Bouthour, D.; Kalai, T.; Chaffei, H.C.; Gouia, H.; Corpas, F.J. Differential Response of NADP-Dehydrogenases and Carbon Metabolism in Leaves and Roots of Two Durum Wheat (Triticum durum Desf.) Cultivars (Karim and Azizi) with Different Sensitivities to Salt Stress. J. Plant Physiol. 2015, 179, 56–63. [Google Scholar] [CrossRef]
- Signorelli, S.; Corpas, F.J.; Borsani, O.; Barroso, J.B.; Monza, J. Water Stress Induces a Differential and Spatially Distributed Nitro-Oxidative Stress Response in Roots and Leaves of Lotus Japonicus. Plant Sci. 2013, 201–202, 137–146. [Google Scholar] [CrossRef] [PubMed]
- Doubnerová Hýsková, V.; Miedzińska, L.; Dobrá, J.; Vankova, R.; Ryšlavá, H. Phosphoenolpyruvate Carboxylase, NADP-Malic Enzyme, and Pyruvate, Phosphate Dikinase Are Involved in the Acclimation of Nicotiana tabacum L. to Drought Stress. J. Plant Physiol. 2014, 171, 19–25. [Google Scholar] [CrossRef] [PubMed]
- Houmani, H.; Rodríguez-Ruiz, M.; Palma, J.M.; Corpas, F.J. Mechanical Wounding Promotes Local and Long Distance Response in the Halophyte Cakile Maritima through the Involvement of the ROS and RNS Metabolism. Nitric Oxide 2018, 74, 93–101. [Google Scholar] [CrossRef] [PubMed]
- Corpas, F.J.; Aguayo-Trinidad, S.; Ogawa, T.; Yoshimura, K.; Shigeoka, S. Activation of NADPH-Recycling Systems in Leaves and Roots of Arabidopsis Thaliana under Arsenic-Induced Stress Conditions Is Accelerated by Knock-out of Nudix Hydrolase 19 (AtNUDX19) Gene. J. Plant Physiol. 2016, 192, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Ruíz-Torres, C.; Feriche-Linares, R.; Rodríguez-Ruíz, M.; Palma, J.M.; Corpas, F.J. Arsenic-Induced Stress Activates Sulfur Metabolism in Different Organs of Garlic (Allium sativum L.) Plants Accompanied by a General Decline of the NADPH-Generating Systems in Roots. J. Plant Physiol. 2017, 211, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Ruiz, M.; Aparicio-Chacón, M.V.; Palma, J.M.; Corpas, F.J. Arsenate Disrupts Ion Balance, Sulfur and Nitric Oxide Metabolisms in Roots and Leaves of Pea (Pisum sativum L.) Plants. Environ. Exp. Bot. 2019, 161, 143–156. [Google Scholar] [CrossRef]
- Kharbech, O.; Houmani, H.; Chaoui, A.; Corpas, F.J. Alleviation of Cr(VI)-Induced Oxidative Stress in Maize (Zea mays L.) Seedlings by NO and H2S Donors through Differential Organ-Dependent Regulation of ROS and NADPH-Recycling Metabolisms. J. Plant Physiol. 2017, 219, 71–80. [Google Scholar] [CrossRef]
- de Freitas-Silva, L.; Rodríguez-Ruiz, M.; Houmani, H.; da Silva, L.C.; Palma, J.M.; Corpas, F.J. Glyphosate-Induced Oxidative Stress in Arabidopsis thaliana Affecting Peroxisomal Metabolism and Triggers Activity in the Oxidative Phase of the Pentose Phosphate Pathway (OxPPP) Involved in NADPH Generation. J. Plant Physiol. 2017, 218, 196–205. [Google Scholar] [CrossRef]
- Begara-Morales, J.C.; Sánchez-Calvo, B.; Gómez-Rodríguez, M.V.; Chaki, M.; Valderrama, R.; Mata-Pérez, C.; López-Jaramillo, J.; Corpas, F.J.; Barroso, J.B. Short-Term Low Temperature Induces Nitro-Oxidative Stress That Deregulates the NADP-Malic Enzyme Function by Tyrosine Nitration in Arabidopsis thaliana. Antioxidants 2019, 8, 448. [Google Scholar] [CrossRef]
- Sánchez-McSweeney, A.; González-Gordo, S.; Aranda-Sicilia, M.N.; Rodríguez-Rosales, M.P.; Venema, K.; Palma, J.M.; Corpas, F.J. Loss of Function of the Chloroplast Membrane K+/H+ Antiporters AtKEA1 and AtKEA2 Alters the ROS and NO Metabolism but Promotes Drought Stress Resilience. Plant Physiol. Biochem. 2021, 160, 106–119. [Google Scholar] [CrossRef]
- Houmani, H.; Debez, A.; de Freitas-Silva, L.; Abdelly, C.; Palma, J.M.; Corpas, F.J. Potassium (K+) Starvation-Induced Oxidative Stress Triggers a General Boost of Antioxidant and NADPH-Generating Systems in the Halophyte Cakile Maritima. Antioxidants 2022, 11, 401. [Google Scholar] [CrossRef] [PubMed]
- Guevara, L.; Domínguez-Anaya, M.Á.; Ortigosa, A.; González-Gordo, S.; Díaz, C.; Vicente, F.; Corpas, F.J.; Pérez Del Palacio, J.; Palma, J.M. Identification of Compounds with Potential Therapeutic Uses from Sweet Pepper (Capsicum annuum L.) Fruits and Their Modulation by Nitric Oxide (NO). Int. J. Mol. Sci. 2021, 22, 4476. [Google Scholar] [CrossRef] [PubMed]
- González-Gordo, S.; Bautista, R.; Claros, M.G.; Cañas, A.; Palma, J.M.; Corpas, F.J. Nitric Oxide-Dependent Regulation of Sweet Pepper Fruit Ripening. J. Exp. Bot. 2019, 70, 4557–4570. [Google Scholar] [CrossRef] [PubMed]
- González-Gordo, S.; Rodríguez-Ruiz, M.; Palma, J.M.; Corpas, F.J. Superoxide Radical Metabolism in Sweet Pepper (Capsicum annuum L.) Fruits Is Regulated by Ripening and by a NO-Enriched Environment. Front. Plant Sci. 2020, 11, 485. [Google Scholar] [CrossRef] [PubMed]
- González-Gordo, S.; Rodríguez-Ruiz, M.; López-Jaramillo, J.; Muñoz-Vargas, M.A.; Palma, J.M.; Corpas, F.J. Nitric Oxide (NO) Differentially Modulates the Ascorbate Peroxidase (APX) Isozymes of Sweet Pepper (Capsicum annuum L.) Fruits. Antioxidants 2022, 11, 765. [Google Scholar] [CrossRef]
- González-Gordo, S.; Cañas, A.; Muñoz-Vargas, M.A.; Palma, J.M.; Corpas, F.J. Lipoxygenase (LOX) in Sweet and Hot Pepper (Capsicum annuum L.) Fruits during Ripening and under an Enriched Nitric Oxide (NO) Gas Atmosphere. Int. J. Mol. Sci. 2022, 23, 15211. [Google Scholar] [CrossRef]
- González-Gordo, S.; Palma, J.M.; Corpas, F.J. Small Heat Shock Protein (SHSP) Gene Family from Sweet Pepper (Capsicum annuum L.) Fruits: Involvement in Ripening and Modulation by Nitric Oxide (NO). Plants 2023, 12, 389. [Google Scholar] [CrossRef]
- Palma, J.M.; Terán, F.; Contreras-Ruiz, A.; Rodríguez-Ruiz, M.; Corpas, F.J. Antioxidant Profile of Pepper (Capsicum annuum L.) Fruits Containing Diverse Levels of Capsaicinoids. Antioxidants 2020, 9, 878. [Google Scholar] [CrossRef]
- Rodríguez-Ruiz, M.; Mioto, P.; Palma, J.M.; Corpas, F.J. S-Nitrosoglutathione Reductase (GSNOR) Activity Is down-Regulated during Pepper (Capsicum annuum L.) Fruit Ripening. Nitric Oxide 2017, 68, 51–55. [Google Scholar] [CrossRef]
- Rodríguez-Ruiz, M.; González-Gordo, S.; Cañas, A.; Campos, M.J.; Paradela, A.; Corpas, F.J.; Palma, J.M. Sweet Pepper (Capsicum annuum L.) Fruits Contain an Atypical Peroxisomal Catalase That Is Modulated by Reactive Oxygen and Nitrogen Species. Antioxidants 2019, 8, 374. [Google Scholar] [CrossRef]
- Mateos, R.M.; Bonilla-Valverde, D.; del Río, L.A.; Palma, J.M.; Corpas, F.J. NADP-Dehydrogenases from Pepper Fruits: Effect of Maturation. Physiol. Plant 2009, 135, 130–139. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Vargas, M.A.; González-Gordo, S.; Palma, J.M.; Corpas, F.J. Inhibition of NADP-Malic Enzyme Activity by H2S and NO in Sweet Pepper (Capsicum annuum L.) Fruits. Physiol. Plant 2020, 168, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Rothermel, B.A.; Nelson, T. Primary Structure of the Maize NADP-Dependent Malic Enzyme. J. Biol. Chem. 1989, 264, 19587–19592. [Google Scholar] [CrossRef]
- Börsch, D.; Westhoff, P. Primary Structure of NADP-Dependent Malic Enzyme in the Dicotyledonous C4 Plant Flaveria Trinervia. FEBS Lett. 1990, 273, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Tronconi, M.A.; Andreo, C.S.; Drincovich, M.F. Chimeric Structure of Plant Malic Enzyme Family: Different Evolutionary Scenarios for NAD- and NADP-Dependent Isoforms. Front. Plant Sci. 2018, 9, 565. [Google Scholar] [CrossRef] [PubMed]
- Detarsio, E.; Alvarez, C.E.; Saigo, M.; Andreo, C.S.; Drincovich, M.F. Identification of Domains Involved in Tetramerization and Malate Inhibition of Maize C4-NADP-Malic Enzyme. J. Biol. Chem. 2007, 282, 6053–6060. [Google Scholar] [CrossRef]
- Saigo, M.; Alvarez, C.E.; Andreo, C.S.; Drincovich, M.F. Plastidial NADP-Malic Enzymes from Grasses: Unraveling the Way to the C4 Specific Isoforms. Plant Physiol. Biochem. 2013, 63, 39–48. [Google Scholar] [CrossRef]
- Zhao, H.; Wang, Y.; Lyu, M.-J.A.; Zhu, X.-G. Two Major Metabolic Factors for an Efficient NADP-Malic Enzyme Type C4 Photosynthesis. Plant Physiol. 2022, 189, 84–98. [Google Scholar] [CrossRef]
- Honda, H.; Akagi, H.; Shimada, H. An Isozyme of the NADP-Malic Enzyme of a CAM Plant, Aloe Arborescens, with Variation on Conservative Amino Acid Residues. Gene 2000, 243, 85–92. [Google Scholar] [CrossRef]
- Laporte, M.M.; Shen, B.; Tarczynski, M.C. Engineering for Drought Avoidance: Expression of Maize NADP-Malic Enzyme in Tobacco Results in Altered Stomatal Function. J. Exp. Bot. 2002, 53, 699–705. [Google Scholar] [CrossRef]
- Arias, C.L.; Andreo, C.S.; Drincovich, M.F.; Gerrard Wheeler, M.C. Fumarate and Cytosolic PH as Modulators of the Synthesis or Consumption of C(4) Organic Acids through NADP-Malic Enzyme in Arabidopsis Thaliana. Plant Mol. Biol. 2013, 81, 297–307. [Google Scholar] [CrossRef]
- Badia, M.B.; Arias, C.L.; Tronconi, M.A.; Maurino, V.G.; Andreo, C.S.; Drincovich, M.F.; Wheeler, M.C.G. Enhanced Cytosolic NADP-ME2 Activity in A. Thaliana Affects Plant Development, Stress Tolerance and Specific Diurnal and Nocturnal Cellular Processes. Plant Sci. 2015, 240, 193–203. [Google Scholar] [CrossRef]
- Aghdam, M.S.; Palma, J.M.; Corpas, F.J. NADPH as a Quality Footprinting in Horticultural Crops Marketability. Trends Food Sci. Technol. 2020, 103, 152–161. [Google Scholar] [CrossRef]
- Maurino, V.G.; Saigo, M.; Andreo, C.S.; Drincovich, M.F. Non-Photosynthetic “malic Enzyme” from Maize: A Constituvely Expressed Enzyme That Responds to Plant Defence Inducers. Plant Mol. Biol. 2001, 45, 409–420. [Google Scholar] [CrossRef]
- Walter, M.H.; Grima-Pettenati, J.; Feuillet, C. Characterization of a Bean (Phaseolus vulgaris L.) Malic-Enzyme Gene. Eur. J. Biochem. 1994, 224, 999–1009. [Google Scholar] [CrossRef]
- Wheeler, M.C.G.; Tronconi, M.A.; Drincovich, M.F.; Andreo, C.S.; Flügge, U.-I.; Maurino, V.G. A Comprehensive Analysis of the NADP-Malic Enzyme Gene Family of Arabidopsis. Plant Physiol. 2005, 139, 39–51. [Google Scholar] [CrossRef]
- Jo, B.-S.; Choi, S.S. Introns: The Functional Benefits of Introns in Genomes. Genom. Inf. 2015, 13, 112–118. [Google Scholar] [CrossRef]
- Schmitz-Linneweber, C.; Lampe, M.-K.; Sultan, L.D.; Ostersetzer-Biran, O. Organellar Maturases: A Window into the Evolution of the Spliceosome. Biochim. Biophys. Acta 2015, 1847, 798–808. [Google Scholar] [CrossRef]
- Fu, Z.-Y.; Zhang, Z.-B.; Hu, X.-J.; Shao, H.-B.; Ping, X. Cloning, Identification, Expression Analysis and Phylogenetic Relevance of Two NADP-Dependent Malic Enzyme Genes from Hexaploid Wheat. C. R. Biol. 2009, 332, 591–602. [Google Scholar] [CrossRef]
- Tao, P.; Guo, W.; Li, B.; Wang, W.; Yue, Z.; Lei, J.; Zhao, Y.; Zhong, X. Genome-Wide Identification, Classification, and Analysis of NADP-ME Family Members from 12 Crucifer Species. Mol. Genet. Genom. 2016, 291, 1167–1180. [Google Scholar] [CrossRef]
- Wheeler, M.C.G.; Arias, C.L.; Tronconi, M.A.; Maurino, V.G.; Andreo, C.S.; Drincovitch, M.F. Arabidopsis thaliana NADP-Malic Enzyme Isoforms: High Degree of Identity but Clearly Distinct Properties. Plant Mol. Biol. 2008, 67, 231–242. [Google Scholar] [CrossRef]
- Centeno, D.C.; Osorio, S.; Nunes-Nesi, A.; Bertolo, A.L.F.; Carneiro, R.T.; Araújo, W.L.; Steinhauser, M.-C.; Michalska, J.; Rohrmann, J.; Geigenberger, P.; et al. Malate Plays a Crucial Role in Starch Metabolism, Ripening, and Soluble Solid Content of Tomato Fruit and Affects Postharvest Softening. Plant Cell 2011, 23, 162–184. [Google Scholar] [CrossRef] [PubMed]
- Ruffner, H.P.; Possner, D.; Brem, S.; Rast, D.M. The Physiological Role of Malic Enzyme in Grape Ripening. Planta 1984, 160, 444–448. [Google Scholar] [CrossRef] [PubMed]
- Carvajal, F.; Palma, F.; Jamilena, M.; Garrido, D. Preconditioning Treatment Induces Chilling Tolerance in Zucchini Fruit Improving Different Physiological Mechanisms against Cold Injury. Ann. Appl. Biol. 2015, 166, 340–354. [Google Scholar] [CrossRef]
- Batista Silva, W.; Cosme Silva, G.M.; Santana, D.B.; Salvador, A.R.; Medeiros, D.B.; Belghith, I.; da Silva, N.M.; Cordeiro, M.H.M.; Misobutsi, G.P. Chitosan Delays Ripening and ROS Production in Guava (Psidium guajava L.) Fruit. Food Chem. 2018, 242, 232–238. [Google Scholar] [CrossRef] [PubMed]
- Cherian, S.; Figueroa, C.R.; Nair, H. ‘Movers and Shakers’ in the Regulation of Fruit Ripening: A Cross-Dissection ff Climacteric Versus Non-Climacteric Fruit. J. Exp. Botany 2014, 65, 4705–4722. [Google Scholar] [CrossRef]
- Jarret, R.L.; Berke, T.; Baldwin, E.A.; Antonious, G.F. Variability for Free Sugars and Organic Acids in Capsicum Chinense. Chem. Biodivers. 2009, 6, 138–145. [Google Scholar] [CrossRef]
- de Ávila Silva, L.; Condori-Apfata, J.A.; Costa, P.M.D.A.; Martino, P.B.; Tavares, A.C.A.; Marcelino, M.M.; Raimundi, S.C.J.; Picoli, E.A.D.T.; Araújo, W.L.; Zsögön, A.; et al. Source Strength Modulates Fruit Set by Starch Turnover and Export of Both Sucrose and Amino Acids in Pepper. Plant Cell Physiol. 2019, 60, 2319–2330. [Google Scholar] [CrossRef]
- Selinski, J.; Scheibe, R. Malate Valves: Old Shuttles with New Perspectives. Plant Biol. 2019, 21 (Suppl. S1), 21–30. [Google Scholar] [CrossRef]
- Zhao, Y.; Yu, H.; Zhou, J.-M.; Smith, S.M.; Li, J. Malate Circulation: Linking Chloroplast Metabolism to Mitochondrial ROS. Trends Plant Sci. 2020, 25, 446–454. [Google Scholar] [CrossRef]
- Rödiger, A.; Agne, B.; Dobritzsch, D.; Helm, S.; Müller, F.; Pötzsch, N.; Baginsky, S. Chromoplast Differentiation in Bell Pepper (Capsicum annuum) Fruits. Plant J. 2021, 105, 1431–1442. [Google Scholar] [CrossRef] [PubMed]
- González-Gordo, S.; Palma, J.M.; Corpas, F.J. Peroxisomal Proteome Mining of Sweet Pepper (Capsicum annuum L.) Fruit Ripening Through Whole Isobaric Tags for Relative and Absolute Quantitation Analysis. Front. Plant Sci. 2022, 13, 893376. [Google Scholar] [CrossRef] [PubMed]
- González-Gordo, S.; Rodríguez-Ruiz, M.; Paradela, A.; Ramos-Fernández, A.; Corpas, F.J.; Palma, J.M. Mitochondrial Protein Expression during Sweet Pepper (Capsicum annuum L.) Fruit Ripening: ITRAQ-Based Proteomic Analysis and Role of Cytochrome c Oxidase. J. Plant Physiol. 2022, 274, 153734. [Google Scholar] [CrossRef] [PubMed]
- Corpas, F.J.; González-Gordo, S.; Rodríguez-Ruiz, M.; Muñoz-Vargas, M.A.; Palma, J.M. Thiol-Based Oxidative Posttranslational Modifications (OxiPTMs) of Plant Proteins. Plant Cell Physiol. 2022, 63, 889–900. [Google Scholar] [CrossRef]
- Hu, J.; Huang, X.; Chen, L.; Sun, X.; Lu, C.; Zhang, L.; Wang, Y.; Zuo, J. Site-Specific Nitrosoproteomic Identification of Endogenously S-Nitrosylated Proteins in Arabidopsis. Plant Physiol. 2015, 167, 1731–1746. [Google Scholar] [CrossRef] [PubMed]
- Aroca, A.; Benito, J.M.; Gotor, C.; Romero, L.C. Persulfidation Proteome Reveals the Regulation of Protein Function by Hydrogen Sulfide in Diverse Biological Processes in Arabidopsis. J. Exp. Bot. 2017, 68, 4915–4927. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-Scale Protein Function Classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the Sensitivity of Progressive Multiple Sequence Alignment through Sequence Weighting, Position-Specific Gap Penalties and Weight Matrix Choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Subramanian, B.; Gao, S.; Lercher, M.J.; Hu, S.; Chen, W.-H. Evolview v3: A Webserver for Visualization, Annotation, and Management of Phylogenetic Trees. Nucleic Acids Res. 2019, 47, W270–W275. [Google Scholar] [CrossRef]
- BaileyPl, T.L.; Elkan, C. Fitting a Mixture Model by Expectation Maximization to Discover Motifs in Biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.-J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein Localization Predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef] [PubMed]
- Sahu, S.S.; Loaiza, C.D.; Kaundal, R. Plant-MSubP: A Computational Framework for the Prediction of Single- and Multi-Target Protein Subcellular Localization Using Integrated Machine-Learning Approaches. AoB Plants 2020, 12, plz068. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a Database of Plant Cis-Acting Regulatory Elements and a Portal to Tools for In Silico Analysis of Promoter Sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast Gapped-Read Alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup the Sequence Alignment/Map Format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Gayte, I.G.; Moreno, R.B.; Zonjic, P.S.; Claros, M.G. DEgenes Hunter—A Flexible R Pipeline for Automated RNA-Seq Studies in Organisms without Reference Genome. Genom. Comput. Biol. 2017, 3, 31. [Google Scholar] [CrossRef]


| Gene Name | Gene ID | Chr. | Genomic Location | Protein ID | Length (aa) | Da | Theoretical pI | Subcellular Localization |
|---|---|---|---|---|---|---|---|---|
| CaNADP-ME1 | 107861795 | 3 | 267357814–267362949 | XP_047266087.1 | 643 | 70,840 | 5.94 | Plastid |
| CaNADP-ME2 | 107843507 | 5 | 158139423–158145651 | XP_016543302.1 | 578 | 64,245 | 5.80 | Cytosol |
| CaNADP-ME3 | 107847755 | 8 | 26364857–26372640 | XP_016547705.1 | 618 | 68,468 | 6.57 | Plastid |
| CaNADP-ME4 | 107842298 | 9 | 199489875–199498766 | XP_016541547.1 | 590 | 65,360 | 6.13 | Plastid |
| CaNADP-ME5 | 107850438 | 12 | 97276100–97283614 | XP_016550447.1 | 644 | 70,709 | 6.28 | Plastid |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Taboada, J.; González-Gordo, S.; Muñoz-Vargas, M.A.; Palma, J.M.; Corpas, F.J. NADP-Dependent Malic Enzyme Genes in Sweet Pepper Fruits: Involvement in Ripening and Modulation by Nitric Oxide (NO). Plants 2023, 12, 2353. https://doi.org/10.3390/plants12122353
Taboada J, González-Gordo S, Muñoz-Vargas MA, Palma JM, Corpas FJ. NADP-Dependent Malic Enzyme Genes in Sweet Pepper Fruits: Involvement in Ripening and Modulation by Nitric Oxide (NO). Plants. 2023; 12(12):2353. https://doi.org/10.3390/plants12122353
Chicago/Turabian StyleTaboada, Jorge, Salvador González-Gordo, María A. Muñoz-Vargas, José M. Palma, and Francisco J. Corpas. 2023. "NADP-Dependent Malic Enzyme Genes in Sweet Pepper Fruits: Involvement in Ripening and Modulation by Nitric Oxide (NO)" Plants 12, no. 12: 2353. https://doi.org/10.3390/plants12122353
APA StyleTaboada, J., González-Gordo, S., Muñoz-Vargas, M. A., Palma, J. M., & Corpas, F. J. (2023). NADP-Dependent Malic Enzyme Genes in Sweet Pepper Fruits: Involvement in Ripening and Modulation by Nitric Oxide (NO). Plants, 12(12), 2353. https://doi.org/10.3390/plants12122353








