Dynamic Translational Landscape Revealed by Genome-Wide Ribosome Profiling under Drought and Heat Stress in Potato
Abstract
1. Introduction
2. Results
2.1. Characteristics of Ribosome-Profiling Data
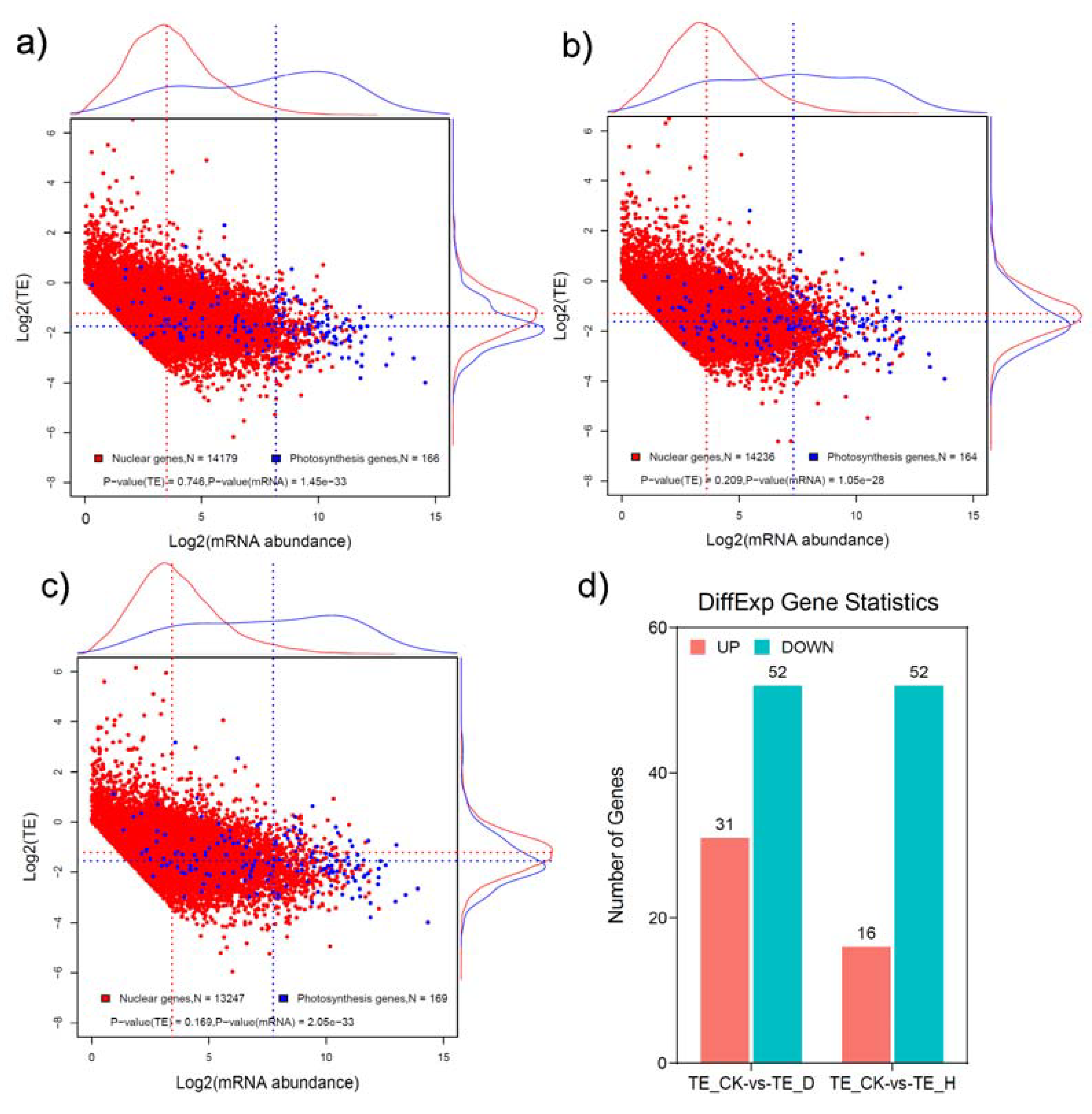
2.2. Expression Changes at the Transcriptional and Translational Levels in Response to Drought and Heat Stress
2.3. The Translational Efficiencies of a Large Number of Genes Were Significantly Changed in Response to Drought and Heat Stress
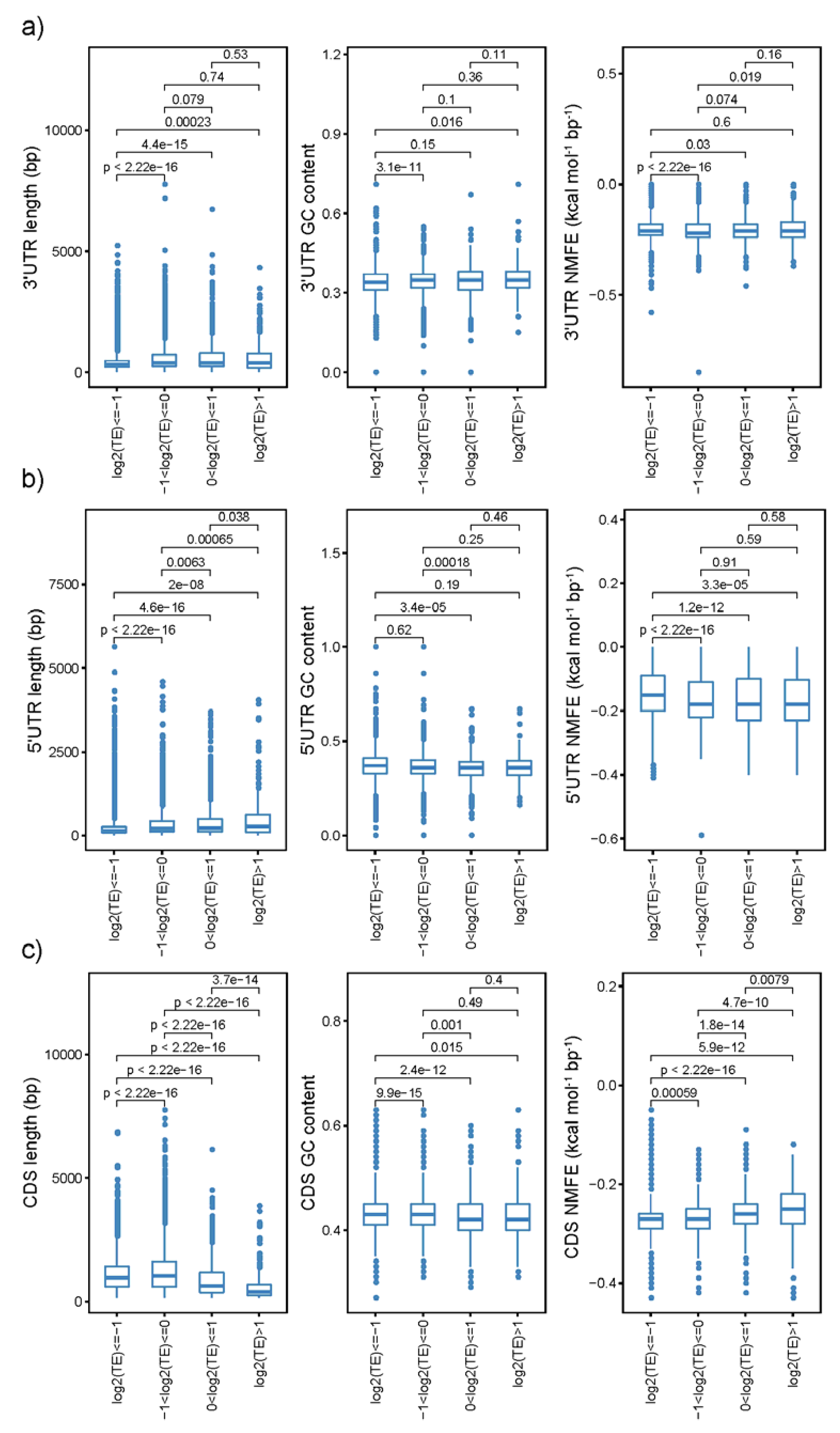
2.4. Gene-Sequence Features Significantly Affect the Translational Efficiencies
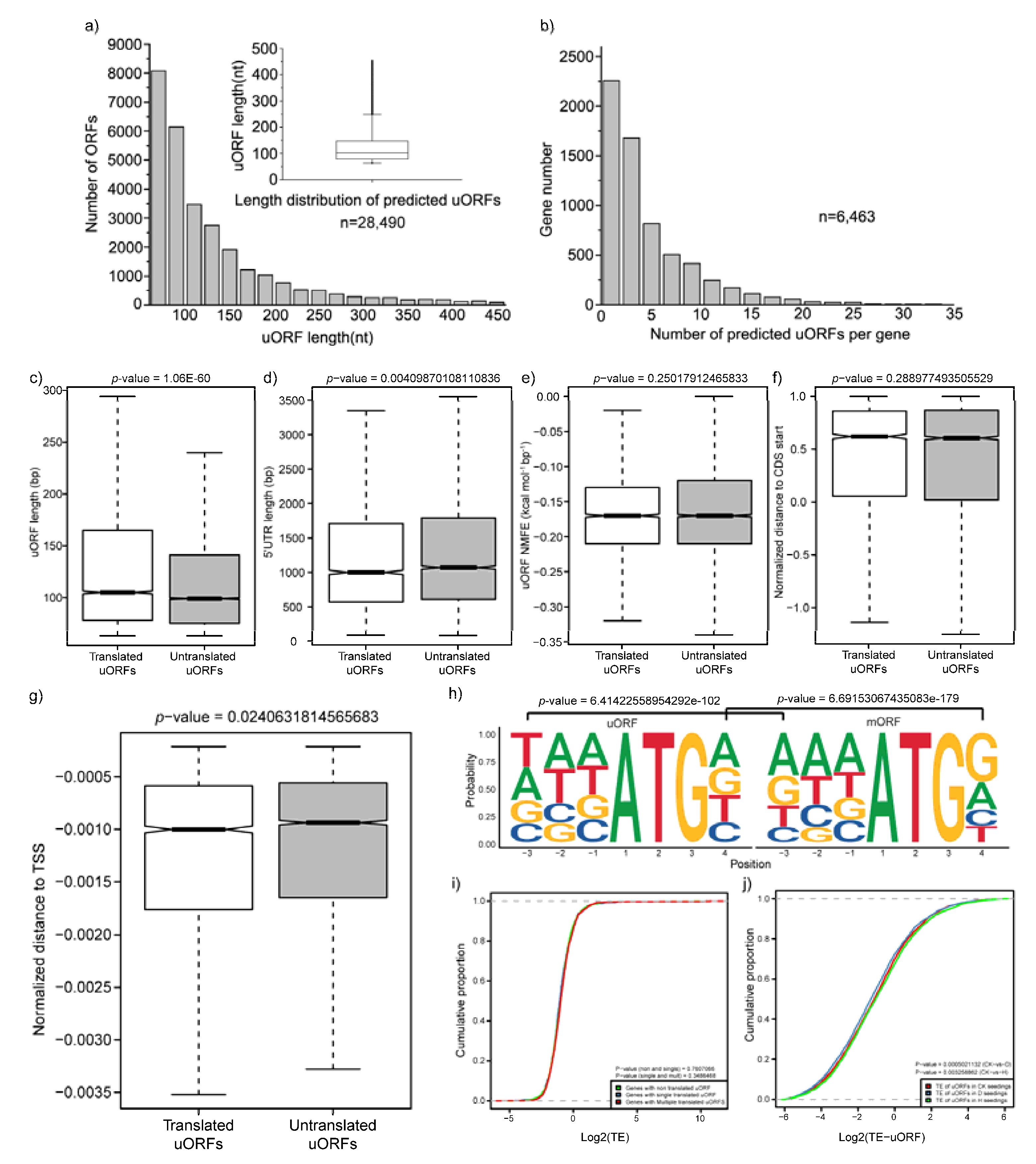
2.5. Characterization of uORFs and Their Impact on the Translation of mORFs
3. Discussion
3.1. Transcriptional and Translational Regulation Coordinate to Respond to Drought and Heat Stress
3.2. TE Was Significantly Affected by Sequence Features
4. Materials and Methods
4.1. Plant Materials
4.2. Ribosome Profiling
4.3. mRNA-Sequencing-Library Construction and Sequencing
4.4. Ribo-seq Data Analysis
4.5. Calculation of Translational Abundance and Calculation of Translational Efficiency (TE)
4.6. Correlation between Transcriptional and Translational Abundance
4.7. The Analysis of the Sequence Features among Four Groups of Different Translational Efficiency
4.8. Analysis of uORFs
4.9. qRT-PCR Validation
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Ciais, P.; Reichstein, M.; Viovy, N.; Granier, A.; Ogée, J.; Allard, V.; Aubinet, M.; Buchmann, N.; Bernhofer, C.; Carrara, A.; et al. Europe-wide reduction in primary productivity caused by the heat and drought in 2003. Nature 2005, 437, 529–533. [Google Scholar] [CrossRef] [PubMed]
- Moreno, A.A.; Orellana, A. The physiological role of the unfolded protein response in plants. Biol. Res. 2011, 44, 75–80. [Google Scholar] [CrossRef]
- Pastore, A.; Martin, S.R.; Politou, A.; Kondapalli, K.C.; Stemmler, T.; Temussi, P.A. Unbiased cold denaturation: Low- and high-temperature unfolding of yeast frataxin under physiological conditions. J. Am. Chem. Soc. 2007, 129, 5374–5375. [Google Scholar] [CrossRef]
- Königshofer, H.; Tromballa, H.W.; Löppert, H.G. Early events in signalling high-temperature stress in tobacco BY2 cells involve alterations in membrane fluidity and enhanced hydrogen peroxide production. Plant Cell Environ. 2008, 31, 1771–1780. [Google Scholar] [CrossRef] [PubMed]
- Saidi, Y.; Domini, M.; Choy, F.; Zryd, J.P.; Schwitzguebel, J.P.; Goloubinoff, P. Activation of the heat shock response in plants by chlorophenols: Transgenic Physcomitrella patens as a sensitive biosensor for organic pollutants. Plant Cell Environ. 2007, 30, 753–763. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Shi, J.; Chen, J.; Zhang, M.; Sun, S.; Xie, S.; Li, X.; Zeng, B.; Peng, L.; Hauck, A.; et al. Ribosome profiling reveals dynamic translational landscape in maize seedlings under drought stress. Plant J. 2015, 84, 1206–1218. [Google Scholar] [CrossRef]
- Zhao, X.; Li, C.; Wan, S.; Zhang, T.; Yan, C.; Shan, S. Transcriptomic analysis and discovery of genes in the response of Arachis hypogaea to drought stress. Mol. Biol. Rep. 2018, 45, 119–131. [Google Scholar] [CrossRef]
- An, J.P.; Zhang, X.W.; Bi, S.Q.; You, C.X.; Wang, X.F.; Hao, Y.J. The ERF transcription factor MdERF38 promotes drought stress-induced anthocyanin biosynthesis in apple. Plant J. 2020, 101, 573–589. [Google Scholar] [CrossRef]
- Tian, Y.; Gu, H.; Fan, Z.; Shi, G.; Yuan, J.; Wei, F.; Yang, Y.; Tian, B.; Cao, G.; Huang, J. Role of a cotton endoreduplication-related gene, GaTOP6B, in response to drought stress. Planta 2019, 249, 1119–1132. [Google Scholar] [CrossRef]
- Fang, C.; Dou, L.; Liu, Y.; Yu, J.; Tu, J. Heat stress-responsive transcriptome analysis in heat susceptible and tolerant rice by high-throughput sequencing. Ecol. Genet. Genom. 2018, 6, 33–40. [Google Scholar] [CrossRef]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Gayacharan; Joel, A.J. Epigenetic responses to drought stress in rice (Oryza sativa L.). Physiol. Mol. Biol. Plants 2013, 19, 379–387. [Google Scholar] [CrossRef] [PubMed]
- de Nadal, E.; Ammerer, G.; Posas, F. Controlling gene expression in response to stress. Nat. Rev. Genet. 2011, 12, 833–845. [Google Scholar] [CrossRef] [PubMed]
- Kawaguchi, R.; Williams, A.J.; Bray, E.A.; Bailey-Serres, J. Water-deficit-induced translational control in Nicotiana tabacum. Plant Cell Environ. 2003, 26, 221–229. [Google Scholar] [CrossRef]
- Hsiao, T.C. Rapid Changes in Levels of Polyribosomes in Zea mays in Response to Water Stress. Plant Physiol. 1970, 46, 281–285. [Google Scholar] [CrossRef]
- Kumar, S.V.; Wigge, P.A. H2A.Z-containing nucleosomes mediate the thermosensory response in Arabidopsis. Cell 2010, 140, 136–147. [Google Scholar] [CrossRef]
- Ingolia, N.T.; Brar, G.A.; Rouskin, S.; McGeachy, A.M.; Weissman, J.S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 2012, 7, 1534–1550. [Google Scholar] [CrossRef]
- Juntawong, P.; Girke, T.; Bazin, J.; Bailey-Serres, J. Translational dynamics revealed by genome-wide profiling of ribosome footprints in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, E203–E212. [Google Scholar] [CrossRef]
- Shalgi, R.; Hurt, J.A.; Krykbaeva, I.; Taipale, M.; Lindquist, S.; Burge, C.B. Widespread regulation of translation by elongation pausing in heat shock. Mol. Cell 2013, 49, 439–452. [Google Scholar] [CrossRef]
- Liu, M.J.; Wu, S.H.; Wu, J.F.; Lin, W.D.; Wu, Y.C.; Tsai, T.Y.; Tsai, H.L.; Wu, S.H. Translational landscape of photomorphogenic Arabidopsis. Plant Cell 2013, 25, 3699–3710. [Google Scholar] [CrossRef]
- Gerashchenko, M.V.; Lobanov, A.V.; Gladyshev, V.N. Genome-wide ribosome profiling reveals complex translational regulation in response to oxidative stress. Proc. Natl. Acad. Sci. USA 2012, 109, 17394–17399. [Google Scholar] [CrossRef]
- Zhang, Y.; Xiao, Z.; Zou, Q.; Fang, J.; Wang, Q.; Yang, X.; Gao, N. Ribosome Profiling Reveals Genome-wide Cellular Translational Regulation upon Heat Stress in Escherichia coli. Genom. Proteom. Bioinform. 2017, 15, 324–330. [Google Scholar] [CrossRef]
- Bánfalvi, Z.; Molnár, A.; Kostyál, Z.; Lakatos, L.; Molnár, G. Comparative studies on potato tuber development using an in vitro tuber induction system. Acta Biol. Hung. 1997, 48, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Wan, J.; Shu, X.E.; Mao, Y.; Liu, X.-M.; Yuan, X.; Zhang, X.; Hess, M.E.; Brüning, J.C.; Qian, S.-B. N6-Methyladenosine Guides mRNA Alternative Translation during Integrated Stress Response. Mol. Cell 2018, 69, 636–647.e7. [Google Scholar] [CrossRef]
- Wu, L.Y.; Lv, Y.Q.; Ye, Y.; Liang, Y.R.; Ye, J.H. Transcriptomic and Translatomic Analyses Reveal Insights into the Developmental Regulation of Secondary Metabolism in the Young Shoots of Tea Plants (Camellia sinensis L.). J. Agric. Food Chem. 2020, 68, 10750–10762. [Google Scholar] [CrossRef] [PubMed]
- Ingolia, N.T.; Ghaemmaghami, S.; Newman, J.R.; Weissman, J.S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 2009, 324, 218–223. [Google Scholar] [CrossRef] [PubMed]
- Young David, J.; Guydosh Nicholas, R.; Zhang, F.; Hinnebusch Alan, G.; Green, R. Rli1/ABCE1 Recycles Terminating Ribosomes and Controls Translation Reinitiation in 3′UTRs In Vivo. Cell 2015, 162, 872–884. [Google Scholar] [CrossRef]
- Hayden, C.A.; Jorgensen, R.A. Identification of novel conserved peptide uORF homology groups in Arabidopsis and rice reveals ancient eukaryotic origin of select groups and preferential association with transcription factor-encoding genes. BMC Biol. 2007, 5, 32. [Google Scholar] [CrossRef]
- Barbosa, C.; Peixeiro, I.; Romão, L. Gene expression regulation by upstream open reading frames and human disease. PLoS Genet. 2013, 9, e1003529. [Google Scholar] [CrossRef]
- Dunn, J.G.; Foo, C.K.; Belletier, N.G.; Gavis, E.R.; Weissman, J.S. Ribosome profiling reveals pervasive and regulated stop codon readthrough in Drosophila melanogaster. eLife 2013, 2, e01179. [Google Scholar] [CrossRef]
- Vogel, C.; Abreu Rde, S.; Ko, D.; Le, S.Y.; Shapiro, B.A.; Burns, S.C.; Sandhu, D.; Boutz, D.R.; Marcotte, E.M.; Penalva, L.O. Sequence signatures and mRNA concentration can explain two-thirds of protein abundance variation in a human cell line. Mol. Syst. Biol. 2010, 6, 400. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.J.; Wu, S.H.; Chen, H.M.; Wu, S.H. Widespread translational control contributes to the regulation of Arabidopsis photomorphogenesis. Mol. Syst. Biol. 2012, 8, 566. [Google Scholar] [CrossRef] [PubMed]
- Ringner, M.; Krogh, M. Folding free energies of 5’-UTRs impact post-transcriptional regulation on a genomic scale in yeast. PLoS Comp. Biol. 2005, 1, 585–592. [Google Scholar] [CrossRef] [PubMed]
- Johnstone, T.G.; Bazzini, A.A.; Giraldez, A.J. Upstream ORFs are prevalent translational repressors in vertebrates. Embo J. 2016, 35, 706–723. [Google Scholar] [CrossRef]
- Ivanov, I.P.; Loughran, G.; Atkins, J.F. uORFs with unusual translational start codons autoregulate expression of eukaryotic ornithine decarboxylase homologs. Proc. Natl. Acad. Sci. USA 2008, 105, 10079–10084. [Google Scholar] [CrossRef]
- Kochetov, A.V. Alternative translation start sites and hidden coding potential of eukaryotic mRNAs. Bioessays 2008, 30, 683–691. [Google Scholar] [CrossRef]
- Sachs, M.S.; Geballe, A.P. Downstream control of upstream open reading frames. Genes Dev. 2006, 20, 915–921. [Google Scholar] [CrossRef]
- Kozak, M. Point mutations define a sequence flanking the AUG initiator codon that modulates translation by eukaryotic ribosomes. Cell 1986, 44, 283–292. [Google Scholar] [CrossRef]
- Kozak, M. Pushing the limits of the scanning mechanism for initiation of translation. Gene 2002, 299, 1–34. [Google Scholar] [CrossRef]
- Wu, H.L.; Song, G.; Walley, J.W.; Hsu, P.Y. The Tomato Translational Landscape Revealed by Transcriptome Assembly and Ribosome Profiling. Plant Physiol. 2019, 181, 367–380. [Google Scholar] [CrossRef]
- Hou, N.; Li, C.; He, J.; Liu, Y.; Yu, S.; Malnoy, M.; Mobeen Tahir, M.; Xu, L.; Ma, F.; Guan, Q. MdMTA-mediated m(6) A modification enhances drought tolerance by promoting mRNA stability and translation efficiency of genes involved in lignin deposition and oxidative stress. New Phytol. 2022, 234, 1294–1314. [Google Scholar] [CrossRef] [PubMed]
- Sonenberg, N.; Hinnebusch, A.G. Regulation of translation initiation in eukaryotes: Mechanisms and biological targets. Cell 2009, 136, 731–745. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.V.; Topper, S.E.; Hubler, S.L.; Hose, J.; Wenger, C.D.; Coon, J.J.; Gasch, A.P. A dynamic model of proteome changes reveals new roles for transcript alteration in yeast. Mol. Syst. Biol. 2011, 7, 514. [Google Scholar] [CrossRef] [PubMed]
- Kage, U.; Powell, J.J.; Gardiner, D.M.; Kazan, K. Ribosome profiling in plants: What is not lost in translation? J. Exp. Bot. 2020, 71, 5323–5332. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Si, X.; Ji, X.; Fan, R.; Liu, J.; Chen, K.; Wang, D.; Gao, C. Genome editing of upstream open reading frames enables translational control in plants. Nat. Biotechnol. 2018, 36, 894–898. [Google Scholar] [CrossRef]
- Liu, S.; Wang, Z.; Zhu, R.; Wang, F.; Cheng, Y.; Liu, Y. Three Differential Expression Analysis Methods for RNA Sequencing: Limma, EdgeR, DESeq2. J. Vis. Exp. 2021, 175, e62528. [Google Scholar]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Calviello, L.; Mukherjee, N.; Wyler, E.; Zauber, H.; Hirsekorn, A.; Selbach, M.; Landthaler, M.; Obermayer, B.; Ohler, U. Detecting actively translated open reading frames in ribosome profiling data. Nat. Methods 2016, 13, 165–170. [Google Scholar] [CrossRef]
- Lorenz, R.; Bernhart, S.H.; Höner Zu Siederdissen, C.; Tafer, H.; Flamm, C.; Stadler, P.F.; Hofacker, I.L. ViennaRNA Package 2.0. Algorithms Mol. Biol. 2011, 6, 26. [Google Scholar] [CrossRef]
- Jian, H.; Xie, L.; Wang, Y.; Cao, Y.; Wan, M.; Lv, D.; Li, J.; Lu, K.; Xu, X.; Liu, L. Characterization of cold stress responses in different rapeseed ecotypes based on metabolomics and transcriptomics analyses. PeerJ 2020, 8, e8704. [Google Scholar] [CrossRef] [PubMed]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jian, H.; Wen, S.; Liu, R.; Zhang, W.; Li, Z.; Chen, W.; Zhou, Y.; Khassanov, V.; Mahmoud, A.M.A.; Wang, J.; et al. Dynamic Translational Landscape Revealed by Genome-Wide Ribosome Profiling under Drought and Heat Stress in Potato. Plants 2023, 12, 2232. https://doi.org/10.3390/plants12122232
Jian H, Wen S, Liu R, Zhang W, Li Z, Chen W, Zhou Y, Khassanov V, Mahmoud AMA, Wang J, et al. Dynamic Translational Landscape Revealed by Genome-Wide Ribosome Profiling under Drought and Heat Stress in Potato. Plants. 2023; 12(12):2232. https://doi.org/10.3390/plants12122232
Chicago/Turabian StyleJian, Hongju, Shiqi Wen, Rongrong Liu, Wenzhe Zhang, Ziyan Li, Weixi Chen, Yonghong Zhou, Vadim Khassanov, Ahmed M. A. Mahmoud, Jichun Wang, and et al. 2023. "Dynamic Translational Landscape Revealed by Genome-Wide Ribosome Profiling under Drought and Heat Stress in Potato" Plants 12, no. 12: 2232. https://doi.org/10.3390/plants12122232
APA StyleJian, H., Wen, S., Liu, R., Zhang, W., Li, Z., Chen, W., Zhou, Y., Khassanov, V., Mahmoud, A. M. A., Wang, J., & Lyu, D. (2023). Dynamic Translational Landscape Revealed by Genome-Wide Ribosome Profiling under Drought and Heat Stress in Potato. Plants, 12(12), 2232. https://doi.org/10.3390/plants12122232




